Characterization and Drug Sensitivity of a New High-Grade Myxofibrosarcoma Cell Line
Abstract
1. Introduction
2. Materials and Methods
2.1. Patient History
2.2. Cell Culture Procedure
2.3. Cell Proliferation Assay
2.4. Immunohistochemical and Immunofluorescence Staining
2.5. Genome Array Comparative Genomic Hybridization (aCGH)
2.6. Gene Expression Analysis
2.7. Drugs Sensitivity Test
2.8. DNA Fragmentation Detection
2.9. Statistical Analysis
3. Results
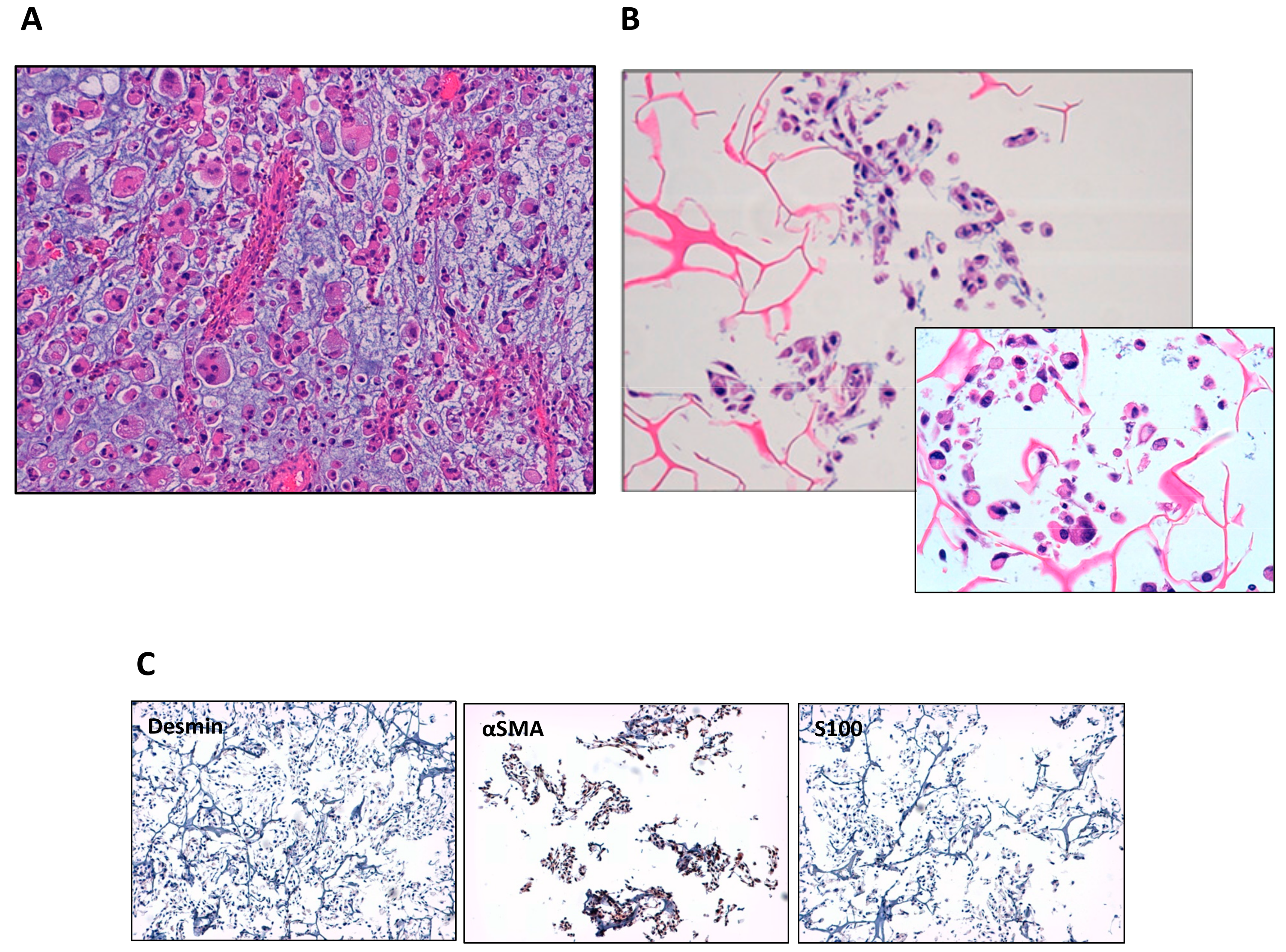
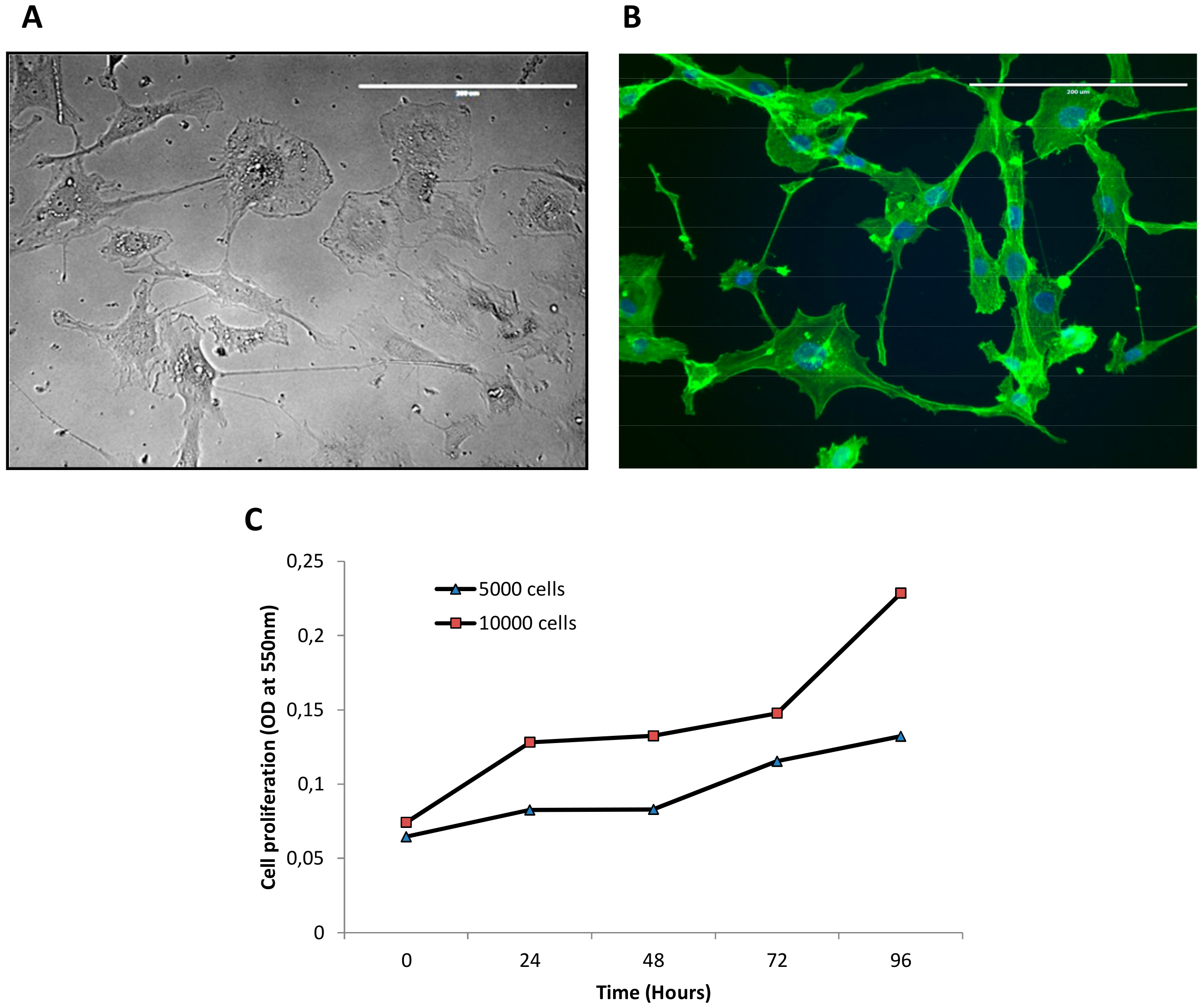
3.1. Establishment of IM-MFS-1 Myxofibrosarcoma Cell Line
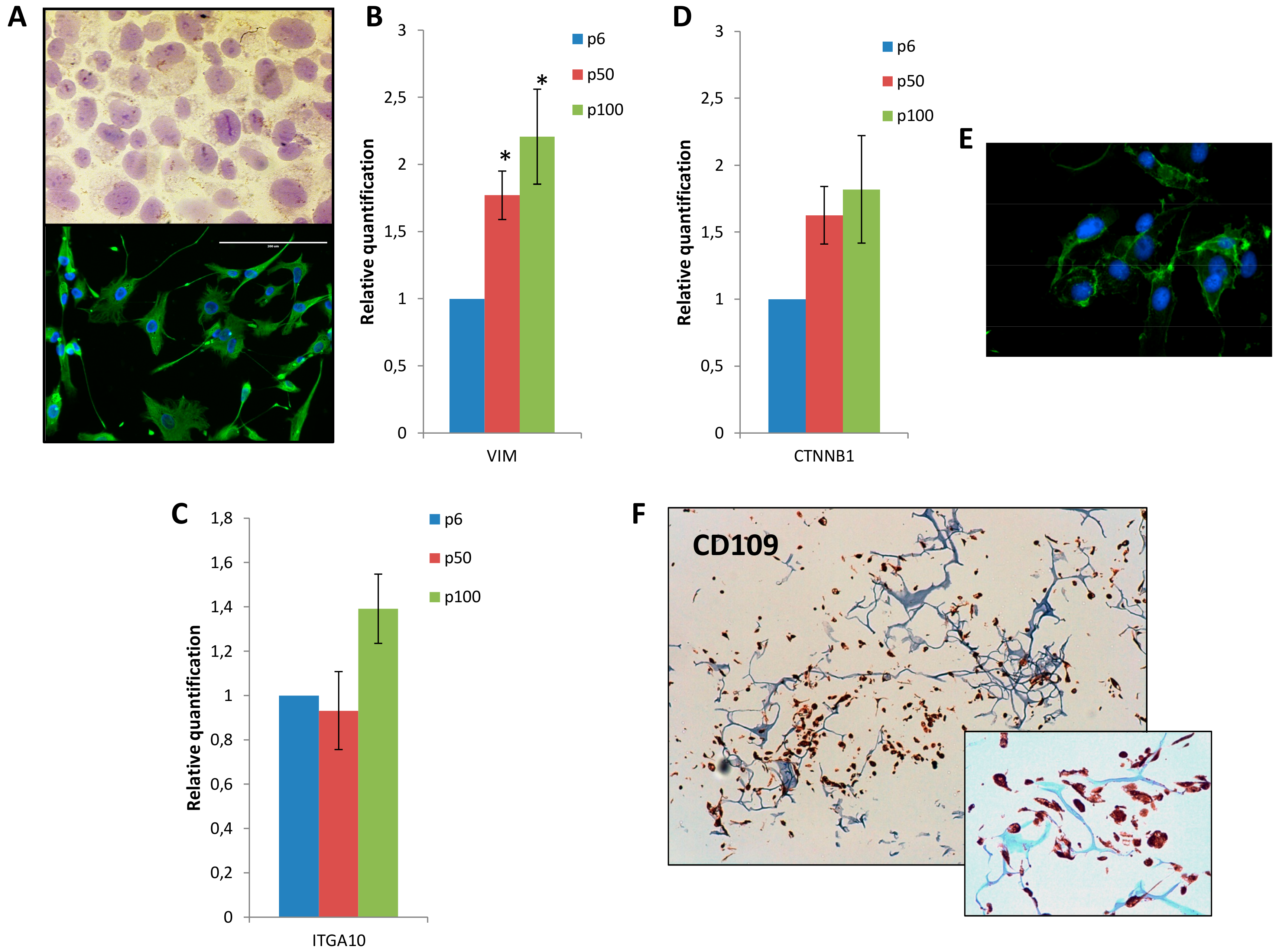
3.2. Mesenchymal Phenotype, β Catenin Distribution and CD109 Positivity of IM-MFS-1
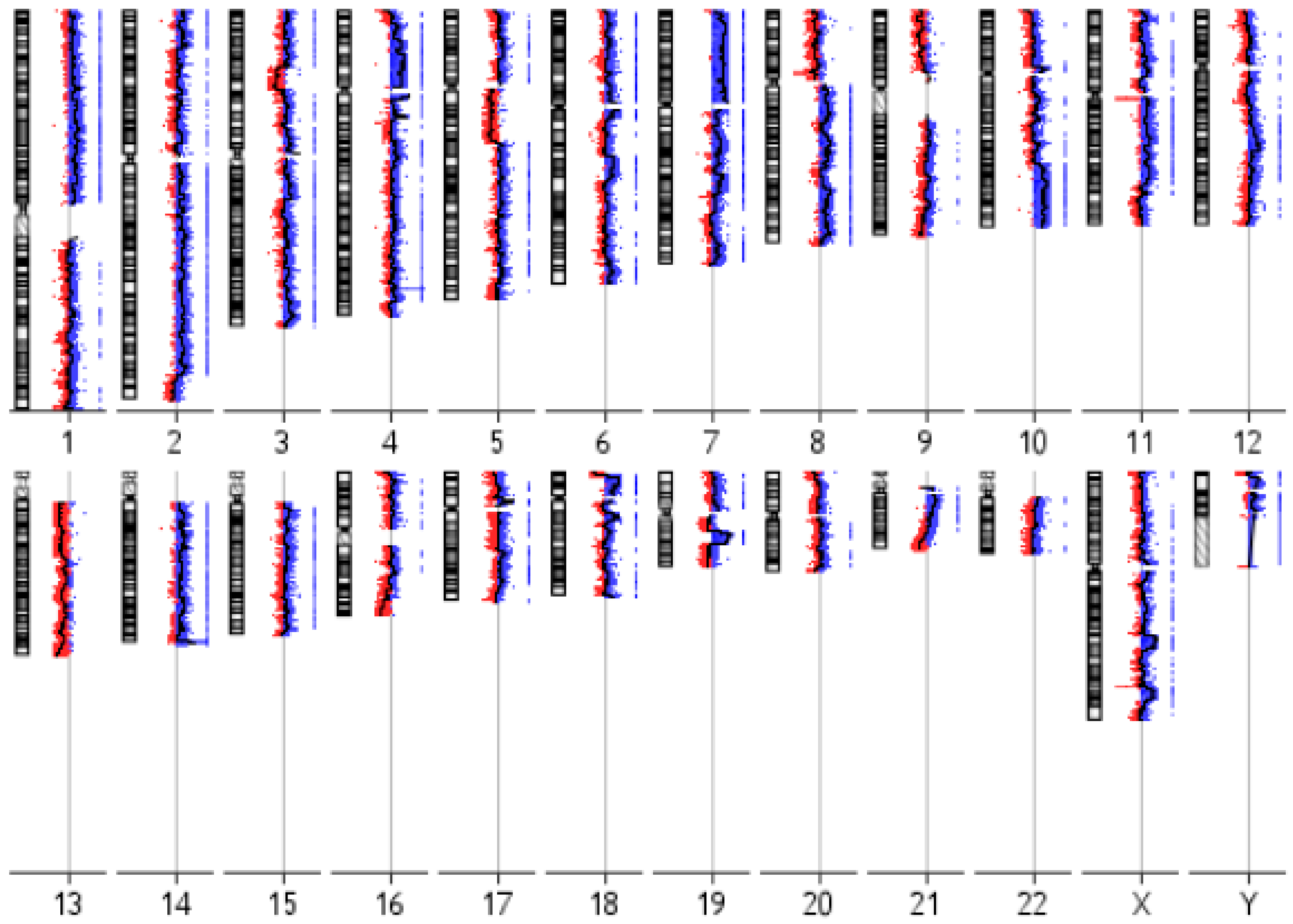
3.3. IM-MFS-1 Cytogenetic Evaluation
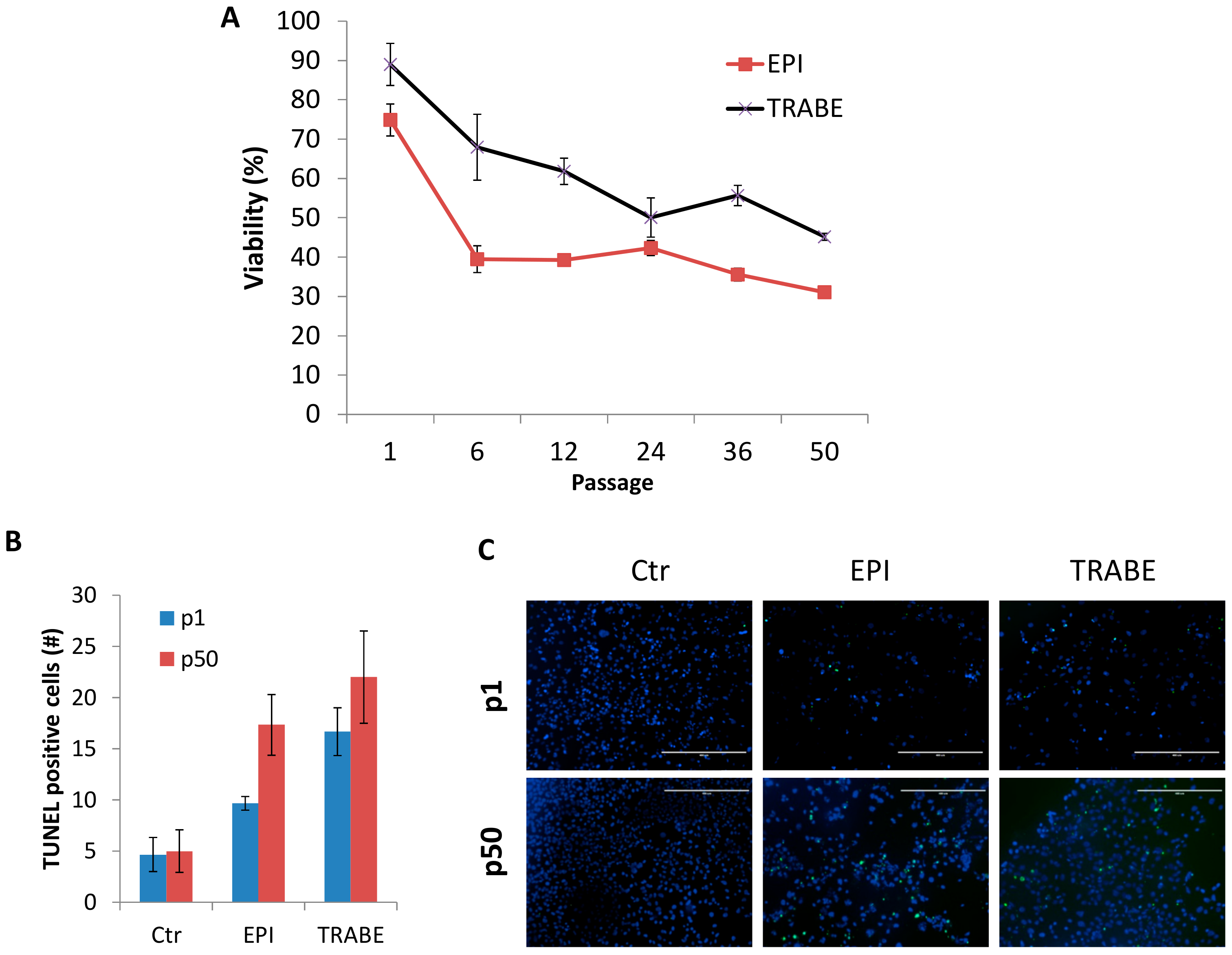
3.4. Increased Drug Sensitivity of IM-MFS-1
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Huang, H.Y.; Lal, P.; Qin, J.; Brennan, M.F.; Antonescu, C.R. Low-grade myxofibrosarcoma: A clinicopathologic analysis of 49 cases treated at a single institution with simultaneous assessment of the efficacy of 3-tier and 4-tier grading systems. Hum. Pathol. 2004, 35, 612–621. [Google Scholar] [CrossRef] [PubMed]
- Fletcher, C.D.M.; Hogendoorn, P.; Mertens, F. World Health Organization Classification of Tumours of Soft Tissue and Bone, 4th ed.; IARC Press: Lyon, France, 2013; pp. 264–274. [Google Scholar]
- Sanfilippo, R.; Miceli, R.; Grosso, F.; Fiore, M.; Puma, E.; Pennacchioli, E.; Barisella, M.; Sangalli, C.; Mariani, L.; Casali, P.G.; et al. Myxofibrosarcoma: Prognostic factors and survival in a series of patients treated at a single institution. Ann. Surg. Oncol. 2011, 18, 720–725. [Google Scholar] [CrossRef] [PubMed]
- Merck, C.; Angervall, L.; Kindblom, L.G.; Odén, A. A malignant soft tissue tumor of fibroblastic-histiocytic origin. A clinicopathologic and prognostic study of 110 cases using a multivariate analysis. Acta Pathol. Microbiol. Immunol. Scand. Suppl. 1983, 282, 1–40. [Google Scholar] [PubMed]
- Ariizumi, T.; Ogose, A.; Kawashima, H.; Hotta, T.; Umezu, H.; Endo, N. Multinucleation followed by an acytokinetic cell division in myxofibrosarcoma with giant cell proliferation. J. Exp. Clin. Cancer Res. 2009, 31. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, S.A.; Tepper, J.; Glatstein, E.; Costa, J.; Baker, A.; Brennan, M.; DeMoss, E.V.; Seipp, C.; Sindelar, W.F.; Sugarbaker, P.; et al. The treatment of soft-tissue sarcomas of the extremities: Prospective randomized evaluations of (1) limb-sparing surgery plus radiation therapy compared with amputation and (2) the role of adjuvant chemotherapy. Ann. Surg. 1982, 196, 305–315. [Google Scholar] [CrossRef] [PubMed]
- Gronchi, A.; Lo Vullo, S.; Colombo, C.; Collini, P.; Stacchiotti, S.; Mariani, L.; Fiore, M.; Casali, P.G. Extremity soft tissue sarcoma in a series of patients treated at a single institution: Local control directly impacts survival. Ann. Surg. 2010, 251, 506–511. [Google Scholar] [CrossRef] [PubMed]
- Manoso, M.W.; Pratt, J.; Healey, J.H.; Boland, P.J.; Athanasian, E.A. Infiltrative MRI pattern and incomplete initial surgery compromise local control of myxofibrosarcoma. Clin. Orthop. Relat. Res. 2006, 450, 89–94. [Google Scholar] [CrossRef] [PubMed]
- Sambri, A.; Bianchi, G.; Righi, A.; Ferrari, C.; Donati, D. Surgical margins do not affect prognosis in high grade myxofibrosarcoma. Eur. J. Surg. Oncol. 2016, 42, 1042–1048. [Google Scholar] [CrossRef] [PubMed]
- Haglund, K.E.; Raut, C.P.; Nascimento, A.F.; Wang, Q.; George, S.; Baldini, E.H. Recurrence patterns and survival for patients with intermediate- and high-grade myxofibrosarcoma. Int. J. Radiat. Oncol. Biol. Phys. 2012, 82, 361–367. [Google Scholar] [CrossRef] [PubMed]
- Billingsley, K.G.; Burt, M.E.; Jara, E.; Ginsberg, R.J.; Woodruff, J.M.; Leung, D.H.; Brennan, M.F. Pulmonary metastases from soft tissue sarcoma: Analysis of patterns of diseases and postmetastasis survival. Ann. Surg. 1999, 229, 602–610. [Google Scholar] [CrossRef] [PubMed]
- Waters, B.; Panicek, D.M.; Lefkowitz, R.A.; Antonescu, C.R.; Healey, J.H.; Athanasian, E.A.; Brennan, M.F. Low-grade myxo-fibrosarcoma: CT and MRI patterns in recurrent disease. AJR Am. J. Roentgenol. 2007, 188, W193–W198. [Google Scholar] [CrossRef] [PubMed]
- Colia, A.; Fiore, M.; Provenzano, S.; Fumagalli, E.; Bertulli, R.; Morosi, C.; Dei Tos, A.P.; Barisella, M.; Gronchi, A.; Casali, P.G.; et al. Activity of anthracycline- and ifosfamide-based chemotherapy in a series of patients affected by advanced myxofibrosarcoma. Clin. Sarcoma Res. 2017, 22. [Google Scholar] [CrossRef] [PubMed]
- Look Hong, N.J.; Hornicek, F.J.; Raskin, K.A.; Yoon, S.S.; Szymonifka, J.; Yeap, B.; Chen, Y.L.; DeLaney, T.F.; Nielsen, G.P.; Mullen, J.T. Prognostic factors and outcomes of patients with myxofibrosarcoma. Ann. Surg. Oncol. 2013, 20, 80–86. [Google Scholar] [CrossRef] [PubMed]
- Salawu, A.; Fernando, M.; Hughes, D.; Reed, M.W.; Woll, P.; Greaves, C.; Day, C.; Alhajimohammed, M.; Sisley, K. Establishment and molecular characterization of seven novel soft-tissue sarcoma cell lines. Br. J. Cancer 2016, 115, 1058–1068. [Google Scholar] [CrossRef] [PubMed]
- Cree, I.A.; Glaysher, S.; Harvey, A.L. Efficacy of anti-cancer agents in cell lines versus human primary tumour tissue. Curr. Opin. Pharmacol. 2010, 10, 375–379. [Google Scholar] [CrossRef] [PubMed]
- De Vita, A.; Mercatali, L.; Recine, F.; Pieri, F.; Riva, N.; Bongiovanni, A.; Liverani, C.; Spadazzi, C.; Miserocchi, G.; Amadori, D.; et al. Current classification, treatment options, and new perspectives in the management of adipocytic sarcomas. Onco-Targets Ther. 2016, 9, 6233–6246. [Google Scholar] [CrossRef] [PubMed]
- De Vita, A.; Mercatali, L.; Miserocchi, G.; Liverani, C.; Spadazzi, C.; Recine, F.; Bongiovanni, A.; Pieri, F.; Cavaliere, D.; Fausti, V.; et al. Establishment of a primary culture of patient-derived soft tissue sarcoma. J. Vis. Exp. 2018, 134. [Google Scholar] [CrossRef] [PubMed]
- Mercatali, L.; La Manna, F.; Groenewoud, A.; Casadei, R.; Recine, F.; Miserocchi, G.; Pieri, F.; Liverani, C.; Bongiovanni, A.; Spadazzi, C.; et al. Development of a patient-derived xenograft (PDX) of breast cancer bone metastasis in a zebrafish model. Int. J. Mol. Sci. 2016, 17, 1375. [Google Scholar] [CrossRef] [PubMed]
- Liverani, C.; La Manna, F.; Groenewoud, A.; Mercatali, L.; Van Der Pluijm, G.; Pieri, F.; Cavaliere, D.; De Vita, A.; Spadazzi, C.; Miserocchi, G.; et al. Innovative approaches to establish and characterize primary cultures: An ex vivo 3D system and the zebrafish model. Biol. Open 2017, 6, 133–140. [Google Scholar] [CrossRef] [PubMed]
- Fogli, S.; Danesi, R.; Gennari, A.; Donati, S.; Conte, P.F.; Del Tacca, M. Gemcitabine, epirubicin and paclitaxel: Pharmacokinetic and pharmacodynamic interactions in advanced breast cancer. Ann. Oncol. 2002, 13, 919–927. [Google Scholar] [CrossRef] [PubMed]
- Robert, J.; Vrignaud, P.; Nguyen-Ngoc, T.; Iliadis, A.; Mauriac, L.; Hurteloup, P. Comparative pharmacokinetics and metabolism of doxorubicin and epirubicin in patients with metastatic breast cancer. Cancer Treat. Rep. 1985, 69, 633–640. [Google Scholar] [PubMed]
- Danesi, R.; Innocenti, F.; Fogli, S.; Gennari, A.; Baldini, E.; Di Paolo, A. Pharmacokinetics and pharmacodynamics of combination chemotherapy with paclitaxel and epirubicinin breast cancer patients. Br. J. Clin. Pharmacol. 2002, 53, 508–518. [Google Scholar] [CrossRef] [PubMed]
- van Kesteren, C.; de Vooght, M.M.; López-Lázaro, L.; Mathôt, R.A.; Schellens, J.H.; Jimeno, J.M.; Beijnen, J.H. Yondelis (trabectedin, ET-743): The development of an anticancer agent of marine origin. Anticancer Drugs 2003, 14, 487–502. [Google Scholar] [CrossRef] [PubMed]
- De Vita, A.; Recine, F.; Mercatali, L.; Miserocchi, G.; Spadazzi, C.; Liverani, C.; Bongiovanni, A.; Pieri, F.; Casadei, R.; Riva, N.; et al. Primary culture of undifferentiated pleomorphic sarcoma: Molecular characterization and response to anticancer agents. Int. J. Mol. Sci. 2017, 18, 2662. [Google Scholar] [CrossRef] [PubMed]
- Mercatali, L.; Spadazzi, C.; Miserocchi, G.; Liverani, C.; De Vita, A.; Bongiovanni, A.; Recine, F.; Amadori, D.; Ibrahim, T. The effect of everolimus in an in vitro model of triple negative breast cancer and osteoclasts. Int. J. Mol. Sci. 2016, 17, 1827. [Google Scholar] [CrossRef] [PubMed]
- Lv, D.; Yu, S.C.; Ping, Y.F.; Wu, H.; Zhao, X.; Zhang, H.; Cui, Y.; Chen, B.; Zhang, X.; Dai, J.; et al. A three-dimensional collagen scaffold cell culture system for screening anti-glioma therapeutics. Oncotarget 2016, 7, 56904–56914. [Google Scholar] [CrossRef] [PubMed]
- Okada, T.; Lee, A.Y.; Qin, L.X.; Agaram, N.; Mimae, T.; Shen, Y.; O’Connor, R.; López-Lago, M.A.; Craig, A.; Miller, M.L.; et al. Integrin-α10 Dependency Identifies RAC and RICTOR as Therapeutic Targets in High-Grade Myxofibrosarcoma. Cancer Discov. 2016, 6, 1148–1165. [Google Scholar] [CrossRef] [PubMed]
- Shang, S.; Hua, F.; Hu, Z.W. The regulation of β-catenin activity and function in cancer: Therapeutic opportunities. Oncotarget 2017, 8, 33972–33989. [Google Scholar] [CrossRef] [PubMed]
- Emori, M.; Tsukahara, T.; Murata, K.; Sugita, S.; Sonoda, T.; Kaya, M.; Soma, T.; Sasaki, M.; Nagoya, S.; Hasegawa, T.; et al. Prognostic impact of CD109 expression in myxofibrosarcoma. J. Surg. Oncol. 2015, 111, 975–979. [Google Scholar] [CrossRef] [PubMed]
- Bizet, A.A.; Liu, K.; Tran-Khanh, N.; Saksena, A.; Vorstenbosch, J.; Finnson, K.W.; Buschmann, M.D.; Philip, A. The TGF-beta co-receptor, CD109, promotes internalization and degradation of TGF-betareceptors. Biochim. Biophys. Acta 2011, 1813, 742–753. [Google Scholar] [CrossRef] [PubMed]
- Ohguri, T.; Hisaoka, M.; Kawauchi, S.; Sasaki, K.; Aoki, T.; Kanemitsu, S.; Matsuyama, A.; Korogi, Y.; Hashimoto, H. Cytogenetic analysis of myxoid liposarcoma and myxofibrosarcoma by array-based comparative genomic hybridisation. J. Clin. Pathol. 2006, 59, 978–983. [Google Scholar] [CrossRef] [PubMed]
- Kawashima, H.; Ogose, A.; Gu, W.; Nishio, J.; Kudo, N.; Kondo, N.; Hotta, T.; Umezu, H.; Tohyama, T.; Nishijima, H.; et al. Establishment and characterization of a novel myxofibrosarcoma cell line. Cancer Genet. Cytogenet. 2005, 161, 28–35. [Google Scholar] [CrossRef] [PubMed]
- Choong, P.F.; Mandahl, N.; Mertens, F.; Willén, H.; Alvegård, T.; Kreicbergs, A.; Mitelman, F.; Rydholm, A. 19p+ marker chromosome correlates with relapse in malignant fibrous histiocytoma. Gene Chromosomes Cancer 1996, 16, 88–93. [Google Scholar] [CrossRef]
- Weiss, S.W.; Enzinger, F.M. Myxoid variant of malignant fibrous histiocytoma. Cancer 1977, 39, 1672–1685. [Google Scholar] [CrossRef]
- Hatano, H.; Tokunaga, K.; Ogose, A.; Imaizumi, S.; Hayami, T.; Yamagiwa, H.; Hotta, T.; Endo, N.; Takahashi, H.E.; Naito, M. Origin of histiocyte-like cells and multinucleated giant cells in malignant fibrous histiocytoma: Neoplastic or reactive? Pathol. Int. 1999, 49, 14–22. [Google Scholar] [CrossRef] [PubMed]
- Lohberger, B.; Stuendl, N.; Leithner, A.; Rinner, B.; Sauer, S.; Kashofer, K.; Liegl-Atzwanger, B. Establishment of a novel cellular model for myxofibrosarcoma heterogeneity. Sci. Rep. 2017, 7, 44700. [Google Scholar] [CrossRef] [PubMed]
- Lohberger, B.; Stuendl, N.; Wolf, E.; Liegl-Atzwanger, B.; Leithner, A.; Rinner, B. The novel myxofibrosarcoma cell line MUG-Myx1 expresses a tumourigenic stem-like cell population with high aldehyde dehydrogenase 1 activity. BMC Cancer 2013, 13, 563. [Google Scholar] [CrossRef] [PubMed]
- De Vita, A.; Recine, F.; Mercatali, L.; Miserocchi, G.; Liverani, C.; Spadazzi, C.; Casadei, R.; Bongiovanni, A.; Pieri, F.; Riva, N.; et al. Myxofibrosarcoma primary cultures: Molecular and pharmacological profile. Ther. Adv. Med. Oncol. 2017, 9, 755–767. [Google Scholar] [CrossRef] [PubMed]
- Borrell, B. How accurate are cancer cell lines? Nature 2010, 463, 858. [Google Scholar] [CrossRef] [PubMed]
- Gao, C.; Wang, Y.; Broaddus, R.; Sun, L.; Xue, F.; Zhang, W. Exon 3 mutations of CTNNB1 drive tumorigenesis: A review. Oncotarget 2017, 9, 5492–5508. [Google Scholar] [CrossRef] [PubMed]
- Ogura, K.; Hosoda, F.; Arai, Y.; Nakamura, H.; Hama, N.; Totoki, Y.; Yoshida, A.; Nagai, M.; Kato, M.; Arakawa, E.; et al. Integrated genetic and epigenetic analysis of myxofibrosarcoma. Nat Commun. 2018, 9, 2765. [Google Scholar] [CrossRef] [PubMed]
- Jamieson, C.; Sharma, M.; Henderson, B.R. Targeting the β-catenin nuclear transport pathway in cancer. Semin. Cancer Biol. 2014, 27, 20–9. [Google Scholar] [CrossRef] [PubMed]
- Bezdieniezhnykh, N.; Lykhova, A.; Semesiuk, N.; Okhrimenko, R.; Kudryavets, Y. Establishment and characterization of new breast and ovarian cancer cell lines as a model for studying cellular plasticity in vitro. Exp. Oncol. 2016, 38, 94–100. [Google Scholar] [PubMed]
- Recine, F.; Bongiovanni, A.; Riva, N.; Fausti, V.; De Vita, A.; Mercatali, L.; Liverani, C.; Miserocchi, G.; Amadori, D.; Ibrahim, T. Update on the role of trabectedin in the treatment of intractable soft tissue sarcomas. Onco-Targets Ther. 2017, 10, 1155–1164. [Google Scholar] [CrossRef] [PubMed]
- Miserocchi, G.; Mercatali, L.; Liverani, C.; De Vita, A.; Spadazzi, C.; Pieri, F.; Bongiovanni, A.; Recine, F.; Amadori, D.; Ibrahim, T. Management and potentialities of primary cancer cultures in preclinical and translational studies. J. Transl. Med. 2017, 15, 229. [Google Scholar] [CrossRef] [PubMed]
- De Vita, A.; Miserocchi, G.; Recine, F.; Mercatali, L.; Pieri, F.; Medri, L.; Bongiovanni, A.; Cavaliere, D.; Liverani, C.; Spadazzi, C.; et al. Activity of eribulin in a primary culture of well-differentiated/dedifferentiated adipocytic sarcoma. Molecules 2016, 21, 1662. [Google Scholar] [CrossRef] [PubMed]
| Cytoband | Gain/Loss | Size (kb 1) | Candidate Gene |
|---|---|---|---|
| 1q25.2 | Loss | 327 | ABL2 |
| 1p32.3 | Gain | 4756 | CDKN2C |
| 2q33.3 | Loss | 20 | CREB1 |
| 3p13 | Loss | 639 | FOXP1 |
| 5q22.2 | Gain | 304 | APC |
| 5q32 | Gain | 33 | PDGFRB |
| 6p22.3 | Loss | 2198 | SOX4 |
| 7q31.2 | Loss | 520 | CAV1, MET |
| 7q34 | Gain | 150 | BRAF |
| 8q24.11-q24.13 | Gain | 5875 | EXT1 |
| 8q24.21 | Loss | 430 | MYC |
| 10q11.22 | Loss | 61 | MAPK8 |
| 10q23.2 | Loss | 224 | BMPR1A |
| 11q23.3 | Loss | 201 | SIK3 |
| 12p12.1 | Loss | 534 | CASC1 |
| 13q14.2 | Loss | 1238 | RB1 |
| 18q11.2 | Loss | 2230 | SS18 |
| 20q12-q13.11 | Gain | 4072 | MAFB |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Miserocchi, G.; De Vita, A.; Mercatali, L.; Recine, F.; Liverani, C.; Spadazzi, C.; Pieri, F.; Riva, N.; Bongiovanni, A.; Casadei, R.; et al. Characterization and Drug Sensitivity of a New High-Grade Myxofibrosarcoma Cell Line. Cells 2018, 7, 186. https://doi.org/10.3390/cells7110186
Miserocchi G, De Vita A, Mercatali L, Recine F, Liverani C, Spadazzi C, Pieri F, Riva N, Bongiovanni A, Casadei R, et al. Characterization and Drug Sensitivity of a New High-Grade Myxofibrosarcoma Cell Line. Cells. 2018; 7(11):186. https://doi.org/10.3390/cells7110186
Chicago/Turabian StyleMiserocchi, Giacomo, Alessandro De Vita, Laura Mercatali, Federica Recine, Chiara Liverani, Chiara Spadazzi, Federica Pieri, Nada Riva, Alberto Bongiovanni, Roberto Casadei, and et al. 2018. "Characterization and Drug Sensitivity of a New High-Grade Myxofibrosarcoma Cell Line" Cells 7, no. 11: 186. https://doi.org/10.3390/cells7110186
APA StyleMiserocchi, G., De Vita, A., Mercatali, L., Recine, F., Liverani, C., Spadazzi, C., Pieri, F., Riva, N., Bongiovanni, A., Casadei, R., Fausti, V., & Ibrahim, T. (2018). Characterization and Drug Sensitivity of a New High-Grade Myxofibrosarcoma Cell Line. Cells, 7(11), 186. https://doi.org/10.3390/cells7110186








