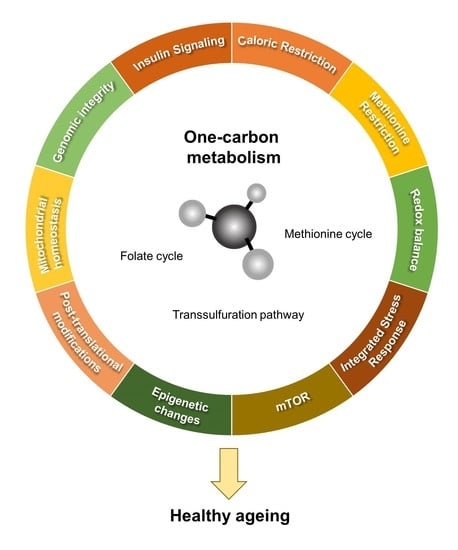
One-Carbon Metabolism: Pulling the Strings behind Aging and Neurodegeneration
Abstract
1. Introduction
2. One-Carbon Metabolic Pathways
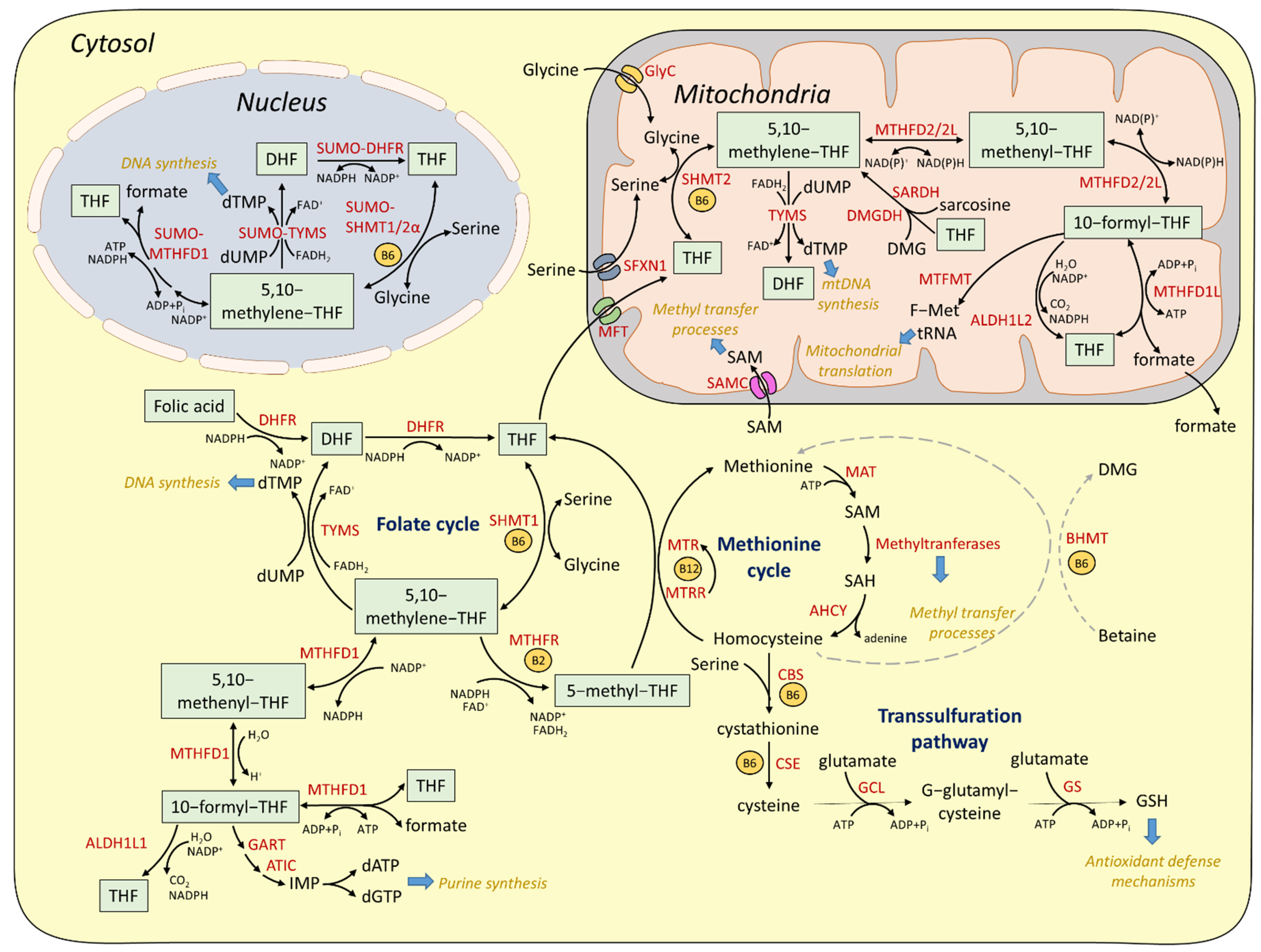
2.1. Cytosolic and Nuclear Pathways of One-Carbon Metabolism
2.2. Mitochondrial One-Carbon Metabolism
3. One-Carbon Metabolism in Aging
3.1. Folate Cycle in Aging
3.2. Methionine Cycle in Aging
3.3. Homocysteine in Aging
3.4. Transsulfuration Pathway in Aging
3.5. Glutathione in Aging
4. One-Carbon Metabolism and Neurodegeneration
4.1. One-Carbon Metabolism in the Pathogenesis of Alzheimer’s Disease
4.2. One-Carbon Metabolism in the Pathogenesis of Parkinson’s Disease
5. Conclusions and Future Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Ducker, G.S.; Rabinowitz, J.D. One-carbon metabolism in health and disease. Cell Metab. 2017, 25, 27–42. [Google Scholar] [CrossRef] [PubMed]
- Cossins, E.A.; Chen, L. Folates and one-carbon metabolism in plants and fungi. Phytochemistry 1997, 45, 437–452. [Google Scholar] [CrossRef]
- Bermingham, A.; Derrick, J.P. The folic acid biosynthesis pathway in bacteria: Evaluation of potential for antibacterial drug discovery. Bioessays 2002, 24, 637–648. [Google Scholar] [CrossRef]
- Hanson, A.D.; Gregory, J.F., 3rd. Folate biosynthesis, turnover, and transport in plants. Annu. Rev. Plant. Biol. 2011, 62, 105–125. [Google Scholar] [CrossRef]
- Garratt, L.C.; Ortori, C.A.; Tucker, G.A.; Sablitzky, F.; Bennett, M.J.; Barrett, D.A. Comprehensive metabolic profiling of mono- and polyglutamated folates and their precursors in plant and animal tissue using liquid chromatography/negative ion electrospray ionisation tandem mass spectrometry. Rapid Commun. Mass Spectrom. 2005, 19, 2390–2398. [Google Scholar] [CrossRef]
- Hou, Z.; Matherly, L.H. Biology of the major facilitative folate transporters slc19a1 and slc46a1. Curr. Top. Membr. 2014, 73, 175–204. [Google Scholar] [PubMed]
- Zhao, R.; Diop-Bove, N.; Visentin, M.; Goldman, I.D. Mechanisms of membrane transport of folates into cells and across epithelia. Annu. Rev. Nutr. 2011, 31, 177–201. [Google Scholar] [CrossRef] [PubMed]
- Qiu, A.; Jansen, M.; Sakaris, A.; Min, S.H.; Chattopadhyay, S.; Tsai, E.; Sandoval, C.; Zhao, R.; Akabas, M.H.; Goldman, I.D. Identification of an intestinal folate transporter and the molecular basis for hereditary folate malabsorption. Cell 2006, 127, 917–928. [Google Scholar] [CrossRef]
- Matherly, L.H.; Hou, Z.; Gangjee, A. The promise and challenges of exploiting the proton-coupled folate transporter for selective therapeutic targeting of cancer. Cancer Chemother. Pharm. 2018, 81, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Matherly, L.H.; Hou, Z. Structure and function of the reduced folate carrier a paradigm of a major facilitator superfamily mammalian nutrient transporter. Vitam. Horm. 2008, 79, 145–184. [Google Scholar] [PubMed]
- Alam, C.; Kondo, M.; O’Connor, D.L.; Bendayan, R. Clinical implications of folate transport in the central nervous system. Trends Pharm. Sci. 2020, 41, 349–361. [Google Scholar] [CrossRef]
- Ormazabal, A.; Casado, M.; Molero-Luis, M.; Montoya, J.; Rahman, S.; Aylett, S.B.; Hargreaves, I.; Heales, S.; Artuch, R. Can folic acid have a role in mitochondrial disorders? Drug Discov. Today 2015, 20, 1349–1354. [Google Scholar] [CrossRef]
- Serrano, M.; Pérez-Dueñas, B.; Montoya, J.; Ormazabal, A.; Artuch, R. Genetic causes of cerebral folate deficiency: Clinical, biochemical and therapeutic aspects. Drug Discov. Today 2012, 17, 1299–1306. [Google Scholar] [CrossRef]
- McCord, E.; Pawar, S.; Koneru, T.; Tatiparti, K.; Sau, S.; Iyer, A.K. Folate receptors’ expression in gliomas may possess potential nanoparticle-based drug delivery opportunities. ACS Omega 2021, 6, 4111–4118. [Google Scholar] [CrossRef]
- Tian, Y.; Wu, G.; Xing, J.C.; Tang, J.; Zhang, Y.; Huang, Z.M.; Jia, Z.C.; Zhao, R.; Tian, Z.Q.; Wang, S.F.; et al. A novel splice variant of folate receptor 4 predominantly expressed in regulatory t cells. BMC Immunol. 2012, 13, 1471–2172. [Google Scholar] [CrossRef] [PubMed]
- Tibbetts, A.S.; Appling, D.R. Compartmentalization of mammalian folate-mediated one-carbon metabolism. Annu. Rev. Nutr. 2010, 30, 57–81. [Google Scholar] [CrossRef] [PubMed]
- Anderson, D.D.; Quintero, C.M.; Stover, P.J. Identification of a de novo thymidylate biosynthesis pathway in mammalian mitochondria. Proc. Natl. Acad. Sci. USA 2011, 108, 15163–15168. [Google Scholar] [CrossRef] [PubMed]
- Chon, J.; Stover, P.J.; Field, M.S. Targeting nuclear thymidylate biosynthesis. Mol. Asp. Med. 2017, 53, 48–56. [Google Scholar] [CrossRef]
- Bottiglieri, T. S-adenosyl-l-methionine (same): From the bench to the bedside--molecular basis of a pleiotrophic molecule. Am. J. Clin. Nutr. 2002, 76. [Google Scholar] [CrossRef]
- Sanderson, S.M.; Gao, X.; Dai, Z.; Locasale, J.W. Methionine metabolism in health and cancer: A nexus of diet and precision medicine. Nat. Rev. Cancer 2019, 19, 625–637. [Google Scholar] [CrossRef]
- Pajares, M.A.; Pérez-Sala, D. Betaine homocysteine s-methyltransferase: Just a regulator of homocysteine metabolism? Cell Mol. Life Sci. 2006, 63, 2792–2803. [Google Scholar] [CrossRef] [PubMed]
- Sbodio, J.I.; Snyder, S.H.; Paul, B.D. Regulators of the transsulfuration pathway. Br. J. Pharm. 2019, 176, 583–593. [Google Scholar] [CrossRef]
- Kutzbach, C.; Stokstad, E.L. Mammalian methylenetetrahydrofolate reductase. Partial purification, properties, and inhibition by s-adenosylmethionine. Biochim. Biophys. Acta 1971, 250, 459–477. [Google Scholar] [CrossRef]
- Finkelstein, J.D. Metabolic regulatory properties of s-adenosylmethionine and s-adenosylhomocysteine. Clin. Chem. Lab. Med. 2007, 45, 1694–1699. [Google Scholar] [CrossRef] [PubMed]
- MacFarlane, A.J.; Perry, C.A.; McEntee, M.F.; Lin, D.M.; Stover, P.J. Mthfd1 is a modifier of chemically induced intestinal carcinogenesis. Carcinogenesis 2010, 32, 427–433. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Pietzke, M.; Meiser, J.; Vazquez, A. Formate metabolism in health and disease. Mol. Metab. 2020, 33, 23–37. [Google Scholar] [CrossRef]
- Ghannad-Zadeh, K.; Das, S. One-carbon metabolism associated vulnerabilities in glioblastoma: A review. Cancers 2021, 13, 3067. [Google Scholar] [CrossRef]
- Krupenko, N.I.; Holmes, R.S.; Tsybovsky, Y.; Krupenko, S.A. Aldehyde dehydrogenase homologous folate enzymes: Evolutionary switch between cytoplasmic and mitochondrial localization. Chem. Biol. Interact. 2015, 234, 12–17. [Google Scholar] [CrossRef]
- Anderson, D.D.; Stover, P.J. Shmt1 and shmt2 are functionally redundant in nuclear de novo thymidylate biosynthesis. PLoS ONE 2009, 4, e5839. [Google Scholar] [CrossRef]
- Anderson, D.D.; Woeller, C.F.; Chiang, E.-P.; Shane, B.; Stover, P.J. Serine hydroxymethyltransferase anchors de novo thymidylate synthesis pathway to nuclear lamina for DNA synthesis. J. Biol. Chem. 2012, 287, 7051–7062. [Google Scholar] [CrossRef]
- Field, M.S.; Kamynina, E.; Agunloye, O.C.; Liebenthal, R.P.; Lamarre, S.G.; Brosnan, M.E.; Brosnan, J.T.; Stover, P.J. Nuclear enrichment of folate cofactors and methylenetetrahydrofolate dehydrogenase 1 (mthfd1) protect de novo thymidylate biosynthesis during folate deficiency. J. Biol. Chem. 2014, 289, 29642–29650. [Google Scholar] [CrossRef] [PubMed]
- Woeller, C.F.; Anderson, D.D.; Szebenyi, D.M.; Stover, P.J. Evidence for small ubiquitin-like modifier-dependent nuclear import of the thymidylate biosynthesis pathway. J. Biol. Chem. 2007, 282, 17623–17631. [Google Scholar] [CrossRef]
- Kamynina, E.; Lachenauer, E.R.; DiRisio, A.C.; Liebenthal, R.P.; Field, M.S.; Stover, P.J. Arsenic trioxide targets mthfd1 and sumo-dependent nuclear de novo thymidylate biosynthesis. Proc. Natl. Acad. Sci. USA 2017, 114, E2319–E2326. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.K.; Koch, J.; Stokstad, E.L. Folate coenzyme pattern, folate linked enzymes and methionine biosynthesis in rat liver mitochondria. Biochem. Z. 1967, 346, 458–466. [Google Scholar]
- Titus, S.A.; Moran, R.G. Retrovirally mediated complementation of the glyb phenotype. Cloning of a human gene encoding the carrier for entry of folates into mitochondria. J. Biol. Chem. 2000, 275, 36811–36817. [Google Scholar] [CrossRef] [PubMed]
- Lin, B.F.; Huang, R.F.; Shane, B. Regulation of folate and one-carbon metabolism in mammalian cells. Iii. Role of mitochondrial folylpoly-gamma-glutamate synthetase. J. Biol. Chem. 1993, 268, 21674–21679. [Google Scholar] [CrossRef]
- Kory, N.; Wyant, G.A.; Prakash, G.; Uit de Bos, J.; Bottanelli, F.; Pacold, M.E.; Chan, S.H.; Lewis, C.A.; Wang, T.; Keys, H.R.; et al. Sfxn1 is a mitochondrial serine transporter required for one-carbon metabolism. Science 2018, 362. [Google Scholar] [CrossRef]
- Lunetti, P.; Damiano, F.; De Benedetto, G.; Siculella, L.; Pennetta, A.; Muto, L.; Paradies, E.; Marobbio, C.M.T.; Dolce, V.; Capobianco, L. Characterization of human and yeast mitochondrial glycine carriers with implications for heme biosynthesis and anemia*. J. Biol. Chem. 2016, 291, 19746–19759. [Google Scholar] [CrossRef]
- Tan, Y.-L.; Sou, N.-L.; Tang, F.-Y.; Ko, H.-A.; Yeh, W.-T.; Peng, J.-H.; Chiang, E.-P.I. Tracing metabolic fate of mitochondrial glycine cleavage system derived formate in vitro and in vivo. Int. J. Mol. Sci. 2020, 21, 8808. [Google Scholar] [CrossRef]
- Porter, D.H.; Cook, R.J.; Wagner, C. Enzymatic properties of dimethylglycine dehydrogenase and sarcosine dehydrogenase from rat liver. Arch. BioChem. Biophys. 1985, 243, 396–407. [Google Scholar] [CrossRef]
- Zhu, Z.; Leung, G.K.K. More than a metabolic enzyme: Mthfd2 as a novel target for anticancer therapy? Front. Oncol. 2020, 10. [Google Scholar] [CrossRef] [PubMed]
- Ben-Sahra, I.; Hoxhaj, G.; Ricoult, S.J.H.; Asara, J.M.; Manning, B.D. Mtorc1 induces purine synthesis through control of the mitochondrial tetrahydrofolate cycle. Science 2016, 351, 728–733. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Garcia Canaveras, J.C.; Chen, Z.; Wang, L.; Liang, L.; Jang, C.; Mayr, J.A.; Zhang, Z.; Ghergurovich, J.M.; Zhan, L.; et al. Serine catabolism feeds nadh when respiration is impaired. Cell Metab. 2020, 31, 809–821.e806. [Google Scholar] [CrossRef] [PubMed]
- Maynard, A.G.; Kanarek, N. Nadh ties one-carbon metabolism to cellular respiration. Cell Metab. 2020, 31, 660–662. [Google Scholar] [CrossRef] [PubMed]
- Nicholls, P. Formate as an inhibitor of cytochrome c oxidase. BioChem. Biophys. Res. Commun. 1975, 67, 610–616. [Google Scholar] [CrossRef]
- Minton, D.R.; Nam, M.; McLaughlin, D.J.; Shin, J.; Bayraktar, E.C.; Alvarez, S.W.; Sviderskiy, V.O.; Papagiannakopoulos, T.; Sabatini, D.M.; Birsoy, K.; et al. Serine catabolism by shmt2 is required for proper mitochondrial translation initiation and maintenance of formylmethionyl-trnas. Mol. Cell 2018, 69, 610–621. [Google Scholar] [CrossRef]
- Esch, B.M.; Limar, S.; Bogdanowski, A.; Gournas, C.; More, T.; Sundag, C.; Walter, S.; Heinisch, J.J.; Ejsing, C.S.; André, B.; et al. Uptake of exogenous serine is important to maintain sphingolipid homeostasis in saccharomyces cerevisiae. PLOS Genet. 2020, 16, e1008745. [Google Scholar] [CrossRef]
- Horne, D.W.; Holloway, R.S.; Wagner, C. Transport of s-adenosylmethionine in isolated rat liver mitochondria. Arch. BioChem. Biophys. 1997, 343, 201–206. [Google Scholar] [CrossRef]
- Agrimi, G.; Di Noia, M.A.; Marobbio, C.M.; Fiermonte, G.; Lasorsa, F.M.; Palmieri, F. Identification of the human mitochondrial s-adenosylmethionine transporter: Bacterial expression, reconstitution, functional characterization and tissue distribution. BioChem. J. 2004, 379, 183–190. [Google Scholar] [CrossRef]
- Schober, F.A.; Moore, D.; Atanassov, I.; Moedas, M.F.; Clemente, P.; Végvári, Á.; Fissi, N.E.; Filograna, R.; Bucher, A.L.; Hinze, Y.; et al. The one-carbon pool controls mitochondrial energy metabolism via complex i and iron-sulfur clusters. Sci. Adv. 2021, 7. [Google Scholar] [CrossRef]
- Zheng, Y.; Lin, T.Y.; Lee, G.; Paddock, M.N.; Momb, J.; Cheng, Z.; Li, Q.; Fei, D.L.; Stein, B.D.; Ramsamooj, S.; et al. Mitochondrial one-carbon pathway supports cytosolic folate integrity in cancer cells. Cell 2018, 175, 1546–1560. [Google Scholar] [CrossRef]
- De Vitto, H.; Arachchige, D.B.; Richardson, B.C.; French, J.B. The intersection of purine and mitochondrial metabolism in cancer. Cells 2021, 10, 2603. [Google Scholar] [CrossRef] [PubMed]
- French, J.B.; Jones, S.A.; Deng, H.; Pedley, A.M.; Kim, D.; Chan, C.Y.; Hu, H.; Pugh, R.J.; Zhao, H.; Zhang, Y.; et al. Spatial colocalization and functional link of purinosomes with mitochondria. Science 2016, 351, 733–737. [Google Scholar] [CrossRef] [PubMed]
- Plummer, J.D.; Johnson, J.E. Extension of cellular lifespan by methionine restriction involves alterations in central carbon metabolism and is mitophagy-dependent. Front. Cell Dev. Biol. 2019, 7. [Google Scholar] [CrossRef] [PubMed]
- Fabrizio, P.; Longo, V.D. The chronological life span of saccharomyces cerevisiae. Methods Mol. Biol. 2007, 371, 89–95. [Google Scholar] [PubMed]
- Ruckenstuhl, C.; Netzberger, C.; Entfellner, I.; Carmona-Gutierrez, D.; Kickenweiz, T.; Stekovic, S.; Gleixner, C.; Schmid, C.; Klug, L.; Sorgo, A.G.; et al. Lifespan extension by methionine restriction requires autophagy-dependent vacuolar acidification. PLoS Genet. 2014, 10. [Google Scholar] [CrossRef]
- Johnson, J.E.; Johnson, F.B. Methionine restriction activates the retrograde response and confers both stress tolerance and lifespan extension to yeast, mouse and human cells. PLoS ONE 2014, 9. [Google Scholar] [CrossRef]
- McCormick, M.A.; Delaney, J.R.; Tsuchiya, M.; Tsuchiyama, S.; Shemorry, A.; Sim, S.; Chou, A.C.; Ahmed, U.; Carr, D.; Murakami, C.J.; et al. A comprehensive analysis of replicative lifespan in 4,698 single-gene deletion strains uncovers conserved mechanisms of aging. Cell Metab. 2015, 22, 895–906. [Google Scholar] [CrossRef]
- Laschober, G.T.; Ruli, D.; Hofer, E.; Muck, C.; Carmona-Gutierrez, D.; Ring, J.; Hutter, E.; Ruckenstuhl, C.; Micutkova, L.; Brunauer, R.; et al. Identification of evolutionarily conserved genetic regulators of cellular aging. Aging Cell 2010, 9, 1084–1097. [Google Scholar] [CrossRef] [PubMed]
- Hine, C.; Harputlugil, E.; Zhang, Y.; Ruckenstuhl, C.; Lee, B.C.; Brace, L.; Longchamp, A.; Treviño-Villarreal, J.H.; Mejia, P.; Ozaki, C.K.; et al. Endogenous hydrogen sulfide production is essential for dietary restriction benefits. Cell 2015, 160, 132–144. [Google Scholar] [CrossRef]
- Tello-Padilla, M.F.; Perez-Gonzalez, A.Y.; Canizal-García, M.; González-Hernández, J.C.; Cortes-Rojo, C.; Olivares-Marin, I.K.; Madrigal-Perez, L.A. Glutathione levels influence chronological life span of saccharomyces cerevisiae in a glucose-dependent manner. Yeast 2018, 35, 387–396. [Google Scholar] [CrossRef]
- Virk, B.; Correia, G.; Dixon, D.P.; Feyst, I.; Jia, J.; Oberleitner, N.; Briggs, Z.; Hodge, E.; Edwards, R.; Ward, J.; et al. Excessive folate synthesis limits lifespan in the c. Elegans: E. Coli aging model. BMC Biol. 2012, 10, 1741–7007. [Google Scholar] [CrossRef] [PubMed]
- Annibal, A.; Tharyan, R.G.; Schonewolff, M.F.; Tam, H.; Latza, C.; Auler, M.M.K.; Grönke, S.; Partridge, L.; Antebi, A. Regulation of the one carbon folate cycle as a shared metabolic signature of longevity. Nat. Commun. 2021, 12, 021–23856. [Google Scholar] [CrossRef] [PubMed]
- Anderson, E.N.; Corkins, M.E.; Li, J.C.; Singh, K.; Parsons, S.; Tucey, T.M.; Sorkaç, A.; Huang, H.; Dimitriadi, M.; Sinclair, D.A.; et al. C. Elegans lifespan extension by osmotic stress requires fudr, base excision repair, foxo, and sirtuins. Mech. Ageing Dev. 2016, 154, 30–42. [Google Scholar] [CrossRef]
- Liu, Y.J.; Janssens, G.E.; McIntyre, R.L.; Molenaars, M.; Kamble, R.; Gao, A.W.; Jongejan, A.; Weeghel, M.V.; MacInnes, A.W.; Houtkooper, R.H. Glycine promotes longevity in caenorhabditis elegans in a methionine cycle-dependent fashion. PLoS Genet. 2019, 15. [Google Scholar] [CrossRef]
- Hansen, M.; Hsu, A.L.; Dillin, A.; Kenyon, C. New genes tied to endocrine, metabolic, and dietary regulation of lifespan from a caenorhabditis elegans genomic rnai screen. PLoS Genet. 2005, 1, 119–128. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.C.; Lim, C.Y.; Lee, P.J.; Hsu, A.L.; Ching, T.T. S-adenosyl methionine synthetase sams-5 mediates dietary restriction-induced longevity in caenorhabditis elegans. PLoS ONE 2020, 15. [Google Scholar] [CrossRef]
- Edwards, C.; Canfield, J.; Copes, N.; Brito, A.; Rehan, M.; Lipps, D.; Brunquell, J.; Westerheide, S.D.; Bradshaw, P.C. Mechanisms of amino acid-mediated lifespan extension in caenorhabditis elegans. BMC Genet. 2015, 16, 015–0167. [Google Scholar] [CrossRef]
- Cabreiro, F.; Au, C.; Leung, K.Y.; Vergara-Irigaray, N.; Cochemé, H.M.; Noori, T.; Weinkove, D.; Schuster, E.; Greene, N.D.; Gems, D. Metformin retards aging in c. Elegans by altering microbial folate and methionine metabolism. Cell 2013, 153, 228–239. [Google Scholar] [CrossRef]
- Miller, D.L.; Roth, M.B. Hydrogen sulfide increases thermotolerance and lifespan in caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 2007, 104, 20618–20622. [Google Scholar] [CrossRef]
- Ng, L.T.; Ng, L.F.; Tang, R.M.Y.; Barardo, D.; Halliwell, B.; Moore, P.K.; Gruber, J. Lifespan and healthspan benefits of exogenous h2s in c. Elegans are independent from effects downstream of eat-2 mutation. NPJ Aging Mech. Dis. 2020, 6, 6. [Google Scholar] [CrossRef] [PubMed]
- Oh, S.I.; Park, J.K.; Park, S.K. Lifespan extension and increased resistance to environmental stressors by n-acetyl-l-cysteine in caenorhabditis elegans. Clinics 2015, 70, 380–386. [Google Scholar] [CrossRef]
- Gusarov, I.; Shamovsky, I.; Pani, B.; Gautier, L.; Eremina, S.; Katkova-Zhukotskaya, O.; Mironov, A.; Makarov, A.; Nudler, E. Dietary thiols accelerate aging of c. Elegans. Nat. Commun. 2021, 12, 021–24634. [Google Scholar] [CrossRef] [PubMed]
- Obata, F.; Miura, M. Enhancing s-adenosyl-methionine catabolism extends drosophila lifespan. Nat. Commun. 2015, 6, 8332. [Google Scholar] [CrossRef] [PubMed]
- Troen, A.M.; French, E.E.; Roberts, J.F.; Selhub, J.; Ordovas, J.M.; Parnell, L.D.; Lai, C.Q. Lifespan modification by glucose and methionine in drosophila melanogaster fed a chemically defined diet. Age 2007, 29, 29–39. [Google Scholar] [CrossRef]
- Lee, B.C.; Kaya, A.; Ma, S.; Kim, G.; Gerashchenko, M.V.; Yim, S.H.; Hu, Z.; Harshman, L.G.; Gladyshev, V.N. Methionine restriction extends lifespan of drosophila melanogaster under conditions of low amino-acid status. Nat. Commun. 2014, 5, 3592. [Google Scholar] [CrossRef]
- Kabil, H.; Kabil, O.; Banerjee, R.; Harshman, L.G.; Pletcher, S.D. Increased transsulfuration mediates longevity and dietary restriction in drosophila. Proc. Natl. Acad. Sci. USA 2011, 108, 16831–16836. [Google Scholar] [CrossRef]
- Shaposhnikov, M.; Proshkina, E.; Koval, L.; Zemskaya, N.; Zhavoronkov, A.; Moskalev, A. Overexpression of cbs and cse genes affects lifespan, stress resistance and locomotor activity in drosophila melanogaster. Aging 2018, 10, 3260–3272. [Google Scholar] [CrossRef]
- Orr, W.C.; Radyuk, S.N.; Prabhudesai, L.; Toroser, D.; Benes, J.J.; Luchak, J.M.; Mockett, R.J.; Rebrin, I.; Hubbard, J.G.; Sohal, R.S. Overexpression of glutamate-cysteine ligase extends life span in drosophila melanogaster. J. Biol. Chem. 2005, 280, 37331–37338. [Google Scholar] [CrossRef]
- Parkhitko, A.A.; Binari, R.; Zhang, N.; Asara, J.M.; Demontis, F.; Perrimon, N. Tissue-specific down-regulation of s-adenosyl-homocysteine via suppression of dahcyl1/dahcyl2 extends health span and life span in drosophila. Genes Dev. 2016, 30, 1409–1422. [Google Scholar] [CrossRef]
- Lemon, J.A.; Boreham, D.R.; Rollo, C.D. A complex dietary supplement extends longevity of mice. J. Gerontol. A Biol. Sci. Med. Sci. 2005, 60, 275–279. [Google Scholar] [CrossRef]
- Anisimov, V.N.; Berstein, L.M.; Popovich, I.G.; Zabezhinski, M.A.; Egormin, P.A.; Piskunova, T.S.; Semenchenko, A.V.; Tyndyk, M.L.; Yurova, M.N.; Kovalenko, I.G.; et al. If started early in life, metformin treatment increases life span and postpones tumors in female shr mice. Aging 2011, 3, 148–157. [Google Scholar] [CrossRef] [PubMed]
- Miller, R.A.; Buehner, G.; Chang, Y.; Harper, J.M.; Sigler, R.; Smith-Wheelock, M. Methionine-deficient diet extends mouse lifespan, slows immune and lens aging, alters glucose, t4, igf-i and insulin levels, and increases hepatocyte mif levels and stress resistance. Aging Cell 2005, 4, 119–125. [Google Scholar] [CrossRef] [PubMed]
- Orentreich, N.; Matias, J.R.; DeFelice, A.; Zimmerman, J.A. Low methionine ingestion by rats extends life span. J. Nutr. 1993, 123, 269–274. [Google Scholar] [PubMed]
- Richie, J.P., Jr.; Leutzinger, Y.; Parthasarathy, S.; Malloy, V.; Orentreich, N.; Zimmerman, J.A. Methionine restriction increases blood glutathione and longevity in f344 rats. FASEB J. 1994, 8, 1302–1307. [Google Scholar] [CrossRef]
- Ito, T.; Yoshikawa, N.; Inui, T.; Miyazaki, N.; Schaffer, S.W.; Azuma, J. Tissue depletion of taurine accelerates skeletal muscle senescence and leads to early death in mice. PLoS ONE 2014, 9, e107409. [Google Scholar] [CrossRef]
- Husemoen, L.L.; Skaaby, T.; Jørgensen, T.; Thuesen, B.H.; Fenger, M.; Grarup, N.; Sandholt, C.H.; Hansen, T.; Pedersen, O.; Linneberg, A. Mthfr c677t genotype and cardiovascular risk in a general population without mandatory folic acid fortification. Eur. J. Nutr. 2014, 53, 1549–1559. [Google Scholar] [CrossRef]
- Yang, Q.; Bailey, L.; Clarke, R.; Flanders, W.D.; Liu, T.; Yesupriya, A.; Khoury, M.J.; Friedman, J.M. Prospective study of methylenetetrahydrofolate reductase (mthfr) variant c677t and risk of all-cause and cardiovascular disease mortality among 6000 us adults. Am. J. Clin. Nutr. 2012, 95, 1245–1253. [Google Scholar] [CrossRef]
- Varela-Moreiras, G.; Pérez-Olleros, L.; García-Cuevas, M.; Ruiz-Roso, B. Effects of ageing on folate metabolism in rats fed a long-term folate deficient diet. Int. J. Vitam. Nutr. Res. 1994, 64, 294–299. [Google Scholar]
- Challet, E.; Dumont, S.; Mehdi, M.K.; Allemann, C.; Bousser, T.; Gourmelen, S.; Sage-Ciocca, D.; Hicks, D.; Pévet, P.; Claustrat, B. Aging-like circadian disturbances in folate-deficient mice. NeuroBiol. Aging 2013, 34, 1589–1598. [Google Scholar] [CrossRef]
- Bahous, R.H.; Cosín-Tomás, M.; Deng, L.; Leclerc, D.; Malysheva, O.; Ho, M.K.; Pallàs, M.; Kaliman, P.; Bedell, B.J.; Caudill, M.A.; et al. Early manifestations of brain aging in mice due to low dietary folate and mild mthfr deficiency. Mol. NeuroBiol. 2019, 56, 4175–4191. [Google Scholar] [CrossRef]
- Ukraintseva, S.; Yashin, A.; Arbeev, K.; Kulminski, A.; Akushevich, I.; Wu, D.; Joshi, G.; Land, K.C.; Stallard, E. Puzzling role of genetic risk factors in human longevity: "Risk alleles" as pro-longevity variants. Biogerontology 2016, 17, 109–127. [Google Scholar] [CrossRef]
- Hall, H.; Cooper, B.R.; Qi, G.; Wijeratne, A.B.; Mosley, A.L.; Weake, V.M. Quantitative proteomic and metabolomic profiling reveals altered mitochondrial metabolism and folate biosynthesis pathways in the aging drosophila eye. Mol. Cell Proteom. 2021, 20, 100127. [Google Scholar] [CrossRef] [PubMed]
- Virk, B.; Jia, J.; Maynard, C.A.; Raimundo, A.; Lefebvre, J.; Richards, S.A.; Chetina, N.; Liang, Y.; Helliwell, N.; Cipinska, M.; et al. Folate acts in e. coli to accelerate c. elegans aging independently of bacterial biosynthesis. Cell Rep. 2016, 14, 1611–1620. [Google Scholar] [CrossRef] [PubMed]
- Onken, B.; Driscoll, M. Metformin induces a dietary restriction-like state and the oxidative stress response to extend c. Elegans healthspan via ampk, lkb1, and skn-1. PLoS ONE 2010, 5, 0008758. [Google Scholar] [CrossRef] [PubMed]
- Blount, B.C.; Mack, M.M.; Wehr, C.M.; MacGregor, J.T.; Hiatt, R.A.; Wang, G.; Wickramasinghe, S.N.; Everson, R.B.; Ames, B.N. Folate deficiency causes uracil misincorporation into human DNA and chromosome breakage: Implications for cancer and neuronal damage. Proc. Natl. Acad. Sci. USA 1997, 94, 3290–3295. [Google Scholar] [CrossRef]
- Chow, H.M.; Herrup, K. Genomic integrity and the ageing brain. Nat. Rev. NeuroSci. 2015, 16, 672–684. [Google Scholar] [CrossRef]
- Steinberg, S.E.; Fonda, S.; Campbell, C.L.; Hillman, R.S. Cellular abnormalities of folate deficiency. Br. J. Haematol. 1983, 54, 605–612. [Google Scholar] [CrossRef]
- Kesavan, V.; Pote, M.S.; Batra, V.; Viswanathan, G. Increased folate catabolism following total body gamma-irradiation in mice. J. Radiat. Res. 2003, 44, 141–144. [Google Scholar] [CrossRef][Green Version]
- Batra, V.; Kesavan, V.; Mishra, K.P. Modulation of enzymes involved in folate dependent one-carbon metabolism by gamma-radiation stress in mice. J. Radiat. Res. 2004, 45, 527–533. [Google Scholar] [CrossRef]
- Choi, S.W.; Kim, Y.I.; Weitzel, J.N.; Mason, J.B. Folate depletion impairs DNA excision repair in the colon of the rat. Gut 1998, 43, 93–99. [Google Scholar] [CrossRef] [PubMed]
- Simon, K.W.; Ma, H.; Dombkowski, A.A.; Cabelof, D.C. Aging alters folate homeostasis and DNA damage response in colon. Mech. Ageing Dev. 2012, 133, 75–82. [Google Scholar] [CrossRef]
- Dorling, J.L.; Martin, C.K.; Redman, L.M. Calorie restriction for enhanced longevity: The role of novel dietary strategies in the present obesogenic environment. Ageing Res. Rev. 2020, 64, 25. [Google Scholar] [CrossRef] [PubMed]
- Lee, B.C.; Kaya, A.; Gladyshev, V.N. Methionine restriction and life-span control. Ann. NY Acad. Sci. 2016, 10. [Google Scholar] [CrossRef] [PubMed]
- Parkhitko, A.A.; Jouandin, P.; Mohr, S.E.; Perrimon, N. Methionine metabolism and methyltransferases in the regulation of aging and lifespan extension across species. Aging Cell 2019, 18, 28. [Google Scholar] [CrossRef]
- Mota-Martorell, N.; Jové, M.; Berdún, R.; Pamplona, R. Plasma methionine metabolic profile is associated with longevity in mammals. Commun. Biol. 2021, 4, 725. [Google Scholar] [CrossRef]
- Sanz, A.; Caro, P.; Ayala, V.; Portero-Otin, M.; Pamplona, R.; Barja, G. Methionine restriction decreases mitochondrial oxygen radical generation and leak as well as oxidative damage to mitochondrial DNA and proteins. FASEB J. 2006, 20, 1064–1073. [Google Scholar] [CrossRef] [PubMed]
- Lees, E.K.; Król, E.; Grant, L.; Shearer, K.; Wyse, C.; Moncur, E.; Bykowska, A.S.; Mody, N.; Gettys, T.W.; Delibegovic, M. Methionine restriction restores a younger metabolic phenotype in adult mice with alterations in fibroblast growth factor 21. Aging Cell 2014, 13, 817–827. [Google Scholar] [CrossRef]
- Stone, K.P.; Wanders, D.; Orgeron, M.; Cortez, C.C.; Gettys, T.W. Mechanisms of increased in vivo insulin sensitivity by dietary methionine restriction in mice. Diabetes 2014, 63, 3721–3733. [Google Scholar] [CrossRef]
- Wanders, D.; Forney, L.A.; Stone, K.P.; Burk, D.H.; Pierse, A.; Gettys, T.W. Fgf21 mediates the thermogenic and insulin-sensitizing effects of dietary methionine restriction but not its effects on hepatic lipid metabolism. Diabetes 2017, 66, 858–867. [Google Scholar] [CrossRef]
- Wanders, D.; Stone, K.P.; Forney, L.A.; Cortez, C.C.; Dille, K.N.; Simon, J.; Xu, M.; Hotard, E.C.; Nikonorova, I.A.; Pettit, A.P.; et al. Role of gcn2-independent signaling through a noncanonical perk/nrf2 pathway in the physiological responses to dietary methionine restriction. Diabetes 2016, 65, 1499–1510. [Google Scholar] [CrossRef] [PubMed]
- Gu, X.; Orozco, J.M.; Saxton, R.A.; Condon, K.J.; Liu, G.Y.; Krawczyk, P.A.; Scaria, S.M.; Harper, J.W.; Gygi, S.P.; Sabatini, D.M. Samtor is an s-adenosylmethionine sensor for the mtorc1 pathway. Science 2017, 358, 813–818. [Google Scholar] [CrossRef]
- Stone, K.P.; Ghosh, S.; Kovalik, J.P.; Orgeron, M.; Wanders, D.; Sims, L.C.; Gettys, T.W. The acute transcriptional responses to dietary methionine restriction are triggered by inhibition of ternary complex formation and linked to erk1/2, mtor, and atf4. Sci. Rep. 2021, 11, 021–83380. [Google Scholar] [CrossRef]
- Aissa, A.F.; Gomes, T.D.; Almeida, M.R.; Hernandes, L.C.; Darin, J.D.; Bianchi, M.L.; Antunes, L.M. Methionine concentration in the diet has a tissue-specific effect on chromosomal stability in female mice. Food Chem. Toxicol. 2013, 62, 456–462. [Google Scholar] [CrossRef] [PubMed]
- Bárcena, C.; López-Otín, C.; Kroemer, G. Methionine restriction for improving progeria: Another autophagy-inducing anti-aging strategy? Autophagy 2019, 15, 558–559. [Google Scholar] [CrossRef] [PubMed]
- Zou, K.; Rouskin, S.; Dervishi, K.; McCormick, M.A.; Sasikumar, A.; Deng, C.; Chen, Z.; Kaeberlein, M.; Brem, R.B.; Polymenis, M.; et al. Life span extension by glucose restriction is abrogated by methionine supplementation: Cross-talk between glucose and methionine and implication of methionine as a key regulator of life span. Sci. Adv. 2020, 6. [Google Scholar] [CrossRef] [PubMed]
- Tain, L.S.; Jain, C.; Nespital, T.; Froehlich, J.; Hinze, Y.; Grönke, S.; Partridge, L. Longevity in response to lowered insulin signaling requires glycine n-methyltransferase-dependent spermidine production. Aging Cell 2020, 19, e13043. [Google Scholar] [CrossRef] [PubMed]
- Brown-Borg, H.M.; Rakoczy, S.; Wonderlich, J.A.; Borg, K.E.; Rojanathammanee, L. Metabolic adaptation of short-living growth hormone transgenic mice to methionine restriction and supplementation. Ann. NY Acad. Sci. 2018, 1418, 118–136. [Google Scholar] [CrossRef] [PubMed]
- Jang, H.S.; Shin, W.J.; Lee, J.E.; Do, J.T. Cpg and non-cpg methylation in epigenetic gene regulation and brain function. Genes 2017, 8, 148. [Google Scholar] [CrossRef] [PubMed]
- Greenberg, M.V.C.; Bourc’his, D. The diverse roles of DNA methylation in mammalian development and disease. Nat. Rev. Mol. Cell Biol. 2019, 20, 590–607. [Google Scholar] [CrossRef] [PubMed]
- Bollati, V.; Schwartz, J.; Wright, R.; Litonjua, A.; Tarantini, L.; Suh, H.; Sparrow, D.; Vokonas, P.; Baccarelli, A. Decline in genomic DNA methylation through aging in a cohort of elderly subjects. Mech. Ageing Dev. 2009, 130, 234–239. [Google Scholar] [CrossRef] [PubMed]
- Horvath, S.; Zhang, Y.; Langfelder, P.; Kahn, R.S.; Boks, M.P.M.; van Eijk, K.; van den Berg, L.H.; Ophoff, R.A. Aging effects on DNA methylation modules in human brain and blood tissue. Genome Biol. 2012, 13, R97. [Google Scholar] [CrossRef] [PubMed]
- Horvath, S.; Gurven, M.; Levine, M.E.; Trumble, B.C.; Kaplan, H.; Allayee, H.; Ritz, B.R.; Chen, B.; Lu, A.T.; Rickabaugh, T.M.; et al. An epigenetic clock analysis of race/ethnicity, sex, and coronary heart disease. Genome Biol. 2016, 17, 171. [Google Scholar] [CrossRef] [PubMed]
- Singhal, R.P.; Mays-Hoopes, L.L.; Eichhorn, G.L. DNA methylation in aging of mice. Mech. Ageing Dev. 1987, 41, 199–210. [Google Scholar] [CrossRef]
- Vanyushin, B.F.; Nemirovsky, L.E.; Klimenko, V.V.; Vasiliev, V.K.; Belozersky, A.N. The 5-methylcytosine in DNA of rats. Tissue and age specificity and the changes induced by hydrocortisone and other agents. Gerontologia 1973, 19, 138–152. [Google Scholar] [CrossRef]
- Cole, J.J.; Robertson, N.A.; Rather, M.I.; Thomson, J.P.; McBryan, T.; Sproul, D.; Wang, T.; Brock, C.; Clark, W.; Ideker, T.; et al. Diverse interventions that extend mouse lifespan suppress shared age-associated epigenetic changes at critical gene regulatory regions. Genome Biol. 2017, 18, 58. [Google Scholar] [CrossRef]
- Wilkinson, G.S.; Adams, D.M.; Haghani, A.; Lu, A.T.; Zoller, J.; Breeze, C.E.; Arnold, B.D.; Ball, H.C.; Carter, G.G.; Cooper, L.N.; et al. DNA methylation predicts age and provides insight into exceptional longevity of bats. Nat. Commun. 2021, 12, 1615. [Google Scholar] [CrossRef]
- Lozoya, O.A.; Martinez-Reyes, I.; Wang, T.; Grenet, D.; Bushel, P.; Li, J.; Chandel, N.; Woychik, R.P.; Santos, J.H. Mitochondrial nicotinamide adenine dinucleotide reduced (nadh) oxidation links the tricarboxylic acid (tca) cycle with methionine metabolism and nuclear DNA methylation. PLoS Biol. 2018, 16. [Google Scholar] [CrossRef]
- Han, S.; Brunet, A. Histone methylation makes its mark on longevity. Trends Cell Biol. 2012, 22, 42–49. [Google Scholar] [CrossRef]
- Kumar, A.; Palfrey, H.A.; Pathak, R.; Kadowitz, P.J.; Gettys, T.W.; Murthy, S.N. The metabolism and significance of homocysteine in nutrition and health. Nutr. Metab. 2017, 14, 017–0233. [Google Scholar] [CrossRef]
- Ostrakhovitch, E.A.; Tabibzadeh, S. Homocysteine and age-associated disorders. Ageing Res. Rev. 2019, 49, 144–164. [Google Scholar] [CrossRef] [PubMed]
- Perna, A.F.; Ingrosso, D. Homocysteine and Chronic Kidney Disease: An Ongoing Narrative. J Nephrol. 2019, 32, 673–675. [Google Scholar] [CrossRef] [PubMed]
- van Ede, A.E.; Laan, R.F.; Blom, H.J.; Boers, G.H.; Haagsma, C.J.; Thomas, C.M.; De Boo, T.M.; van de Putte, L.B. Homocysteine and folate status in methotrexate-treated patients with rheumatoid arthritis. Rheumatology 2002, 41, 658–665. [Google Scholar] [CrossRef]
- Schwaninger, M.; Ringleb, P.; Winter, R.; Kohl, B.; Fiehn, W.; Rieser, P.A.; Walter-Sack, I. Elevated plasma concentrations of homocysteine in antiepileptic drug treatment. Epilepsia 1999, 40, 345–350. [Google Scholar] [CrossRef] [PubMed]
- Konstantinova, S.V.; Tell, G.S.; Vollset, S.E.; Nygård, O.; Bleie, Ø.; Ueland, P.M. Divergent associations of plasma choline and betaine with components of metabolic syndrome in middle age and elderly men and women. J. Nutr. 2008, 138, 914–920. [Google Scholar] [CrossRef] [PubMed]
- Huang, B.X.; Zhu, Y.Y.; Tan, X.Y.; Lan, Q.Y.; Li, C.L.; Chen, Y.M.; Zhu, H.L. Serum betaine is inversely associated with low lean mass mainly in men in a chinese middle-aged and elderly community-dwelling population. Br. J. Nutr. 2016, 115, 2181–2188. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.J.; Noormohammadi, A.; Koyuncu, S.; Calculli, G.; Simic, M.S.; Herholz, M.; Trifunovic, A.; Vilchez, D. Prostaglandin signals from adult germ stem cells delay somatic aging of caenorhabditis elegans. Nat. Metab. 2019, 1, 790–810. [Google Scholar] [CrossRef] [PubMed]
- Mota-Martorell, N.; Jové, M.; Borrás, C.; Berdún, R.; Obis, È.; Sol, J.; Cabré, R.; Pradas, I.; Galo-Licona, J.D.; Puig, J.; et al. Methionine transsulfuration pathway is upregulated in long-lived humans. Free Radic. Biol. Med. 2021, 162, 38–52. [Google Scholar] [CrossRef]
- Sokolov, A.S.; Nekrasov, P.V.; Shaposhnikov, M.V.; Moskalev, A.A. Hydrogen sulfide in longevity and pathologies: Inconsistency is malodorous. Ageing Res. Rev. 2021, 67, 28. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Tang, Z.H.; Ren, Z.; Qu, S.L.; Liu, M.H.; Liu, L.S.; Jiang, Z.S. Hydrogen sulfide, the next potent preventive and therapeutic agent in aging and age-associated diseases. Mol. Cell Biol. 2013, 33, 1104–1113. [Google Scholar] [CrossRef]
- Dziegelewska, M.; Holtze, S.; Vole, C.; Wachter, U.; Menzel, U.; Morhart, M.; Groth, M.; Szafranski, K.; Sahm, A.; Sponholz, C.; et al. Low sulfide levels and a high degree of cystathionine β-synthase (cbs) activation by s-adenosylmethionine (sam) in the long-lived naked mole-rat. Redox Biol. 2016, 8, 192–198. [Google Scholar] [CrossRef] [PubMed]
- Wei, Y.; Kenyon, C. Roles for ros and hydrogen sulfide in the longevity response to germline loss in caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 2016, 113, E2832–E2841. [Google Scholar] [CrossRef] [PubMed]
- Bithi, N.; Link, C.; Henderson, Y.O.; Kim, S.; Yang, J.; Li, L.; Wang, R.; Willard, B.; Hine, C. Dietary restriction transforms the mammalian protein persulfidome in a tissue-specific and cystathionine γ-lyase-dependent manner. Nat. Commun. 2021, 12, 1–20. [Google Scholar] [CrossRef] [PubMed]
- Pow, D.V.; Sullivan, R.; Reye, P.; Hermanussen, S. Localization of taurine transporters, taurine, and (3)h taurine accumulation in the rat retina, pituitary, and brain. Glia 2002, 37, 153–168. [Google Scholar] [CrossRef] [PubMed]
- Ames, B.N. Prolonging healthy aging: Longevity vitamins and proteins. Proc. Natl. Acad. Sci. USA 2018, 115, 10836–10844. [Google Scholar] [CrossRef] [PubMed]
- Du, G.; Liu, Z.; Yu, Z.; Zhuo, Z.; Zhu, Y.; Zhou, J.; Li, Y.; Chen, H. Taurine represses age-associated gut hyperplasia in drosophila via counteracting endoplasmic reticulum stress. Aging Cell 2021, 20, e13319. [Google Scholar] [CrossRef]
- Liu, Y.; Hyde, A.S.; Simpson, M.A.; Barycki, J.J. Emerging regulatory paradigms in glutathione metabolism. Adv. Cancer Res. 2014, 122, 69–101. [Google Scholar] [PubMed]
- Go, Y.M.; Jones, D.P. Redox theory of aging: Implications for health and disease. Clin. Sci. 2017, 131, 1669–1688. [Google Scholar] [CrossRef]
- Schulz, T.J.; Zarse, K.; Voigt, A.; Urban, N.; Birringer, M.; Ristow, M. Glucose restriction extends caenorhabditis elegans life span by inducing mitochondrial respiration and increasing oxidative stress. Cell Metab. 2007, 6, 280–293. [Google Scholar] [CrossRef]
- Elshorbagy, A.K.; Jernerén, F.; Scudamore, C.L.; McMurray, F.; Cater, H.; Hough, T.; Cox, R.; Refsum, H. Exploring the lean phenotype of glutathione-depleted mice: Thiol, amino acid and fatty acid profiles. PLoS ONE 2016, 11. [Google Scholar] [CrossRef]
- Kumar, P.; Liu, C.; Hsu, J.W.; Chacko, S.; Minard, C.; Jahoor, F.; Sekhar, R.V. Glycine and n-acetylcysteine (glynac) supplementation in older adults improves glutathione deficiency, oxidative stress, mitochondrial dysfunction, inflammation, insulin resistance, endothelial dysfunction, genotoxicity, muscle strength, and cognition: Results of a pilot clinical trial. Clin. Transl. Med. 2021, 11, 372. [Google Scholar]
- El-Hafidi, M.; Franco, M.; Ramírez, A.R.; Sosa, J.S.; Flores, J.A.P.; Acosta, O.L.; Salgado, M.C.; Cardoso-Saldaña, G. Glycine increases insulin sensitivity and glutathione biosynthesis and protects against oxidative stress in a model of sucrose-induced insulin resistance. Oxid. Med. Cell Longev. 2018, 21. [Google Scholar] [CrossRef] [PubMed]
- Sekhar, R.V.; McKay, S.V.; Patel, S.G.; Guthikonda, A.P.; Reddy, V.T.; Balasubramanyam, A.; Jahoor, F. Glutathione synthesis is diminished in patients with uncontrolled diabetes and restored by dietary supplementation with cysteine and glycine. Diabetes Care 2011, 34, 162–167. [Google Scholar] [CrossRef] [PubMed]
- Dekhne, A.S.; Hou, Z.; Gangjee, A.; Matherly, L.H. Therapeutic targeting of mitochondrial one-carbon metabolism in cancer. Mol. Cancer Ther. 2020, 19, 2245–2255. [Google Scholar] [CrossRef] [PubMed]
- Korsmo, H.W.; Jiang, X. One carbon metabolism and early development: A diet-dependent destiny. Trends Endocrinol. Metab. 2021, 32, 579–593. [Google Scholar] [CrossRef]
- Reina-Campos, M.; Diaz-Meco, M.T.; Moscat, J. The complexity of the serine glycine one-carbon pathway in cancer. J. Cell Biol. 2019, 219. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Vousden, K.H. Serine and one-carbon metabolism in cancer. Nat. Rev. Cancer 2016, 16, 650–662. [Google Scholar] [CrossRef] [PubMed]
- Imbard, A.; Benoist, J.-F.; Blom, H.J. Neural tube defects, folic acid and methylation. Int. J. Environ. Res. Public Health 2013, 10, 4352–4389. [Google Scholar] [CrossRef] [PubMed]
- Naninck, E.F.G.; Stijger, P.C.; Brouwer-Brolsma, E.M. The importance of maternal folate status for brain development and function of offspring. Adv. Nutr. 2019, 10, 502–519. [Google Scholar] [CrossRef]
- Troen, A.M. Folate and vitamin b12: Function and importance in cognitive development. Nestle Nutr. Inst. Workshop Ser. 2012, 70, 161–171. [Google Scholar]
- Kronenberg, G.; Colla, M.; Endres, M. Folic acid, neurodegenerative and neuropsychiatric disease. Curr. Mol. Med. 2009, 9, 315–323. [Google Scholar] [CrossRef] [PubMed]
- Coppedè, F. One-carbon epigenetics and redox biology of neurodegeneration. Free Radic. Biol. Med. 2021, 170, 19–33. [Google Scholar] [CrossRef] [PubMed]
- Knopman, D.S.; Amieva, H.; Petersen, R.C.; Chételat, G.; Holtzman, D.M.; Hyman, B.T.; Nixon, R.A.; Jones, D.T. Alzheimer disease. Nat. Rev. Dis. Primers 2021, 7, 33. [Google Scholar] [CrossRef] [PubMed]
- Otaegui-Arrazola, A.; Amiano, P.; Elbusto, A.; Urdaneta, E.; Martínez-Lage, P. Diet, cognition, and alzheimer’s disease: Food for thought. Eur. J. Nutr. 2014, 53, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Corrada, M.M.; Kawas, C.H.; Hallfrisch, J.; Muller, D.; Brookmeyer, R. Reduced risk of alzheimer’s disease with high folate intake: The baltimore longitudinal study of aging. Alzheimers Dement. 2005, 1, 11–18. [Google Scholar] [CrossRef]
- Luchsinger, J.A.; Tang, M.-X.; Miller, J.; Green, R.; Mayeux, R. Relation of higher folate intake to lower risk of alzheimer disease in the elderly. Arch. Neurol. 2007, 64, 86–92. [Google Scholar] [CrossRef] [PubMed]
- Ma, F.; Wu, T.; Zhao, J.; Song, A.; Liu, H.; Xu, W.; Huang, G. Folic acid supplementation improves cognitive function by reducing the levels of peripheral inflammatory cytokines in elderly chinese subjects with mci. Sci. Rep. 2016, 6, 37486. [Google Scholar] [CrossRef]
- Sofi, F.; Cesari, F.; Abbate, R.; Gensini, G.F.; Casini, A. Adherence to mediterranean diet and health status: Meta-analysis. BMJ 2008, 337, a1344. [Google Scholar] [CrossRef]
- Ravaglia, G.; Forti, P.; Maioli, F.; Martelli, M.; Servadei, L.; Brunetti, N.; Porcellini, E.; Licastro, F. Homocysteine and folate as risk factors for dementia and alzheimer disease. Am. J. Clin. Nutr. 2005, 82, 636–643. [Google Scholar] [CrossRef] [PubMed]
- Robinson, N.; Grabowski, P.; Rehman, I. Alzheimer’s disease pathogenesis: Is there a role for folate? Mech. Ageing Dev. 2018, 174, 86–94. [Google Scholar] [CrossRef]
- Smith, A.D.; Refsum, H.; Bottiglieri, T.; Fenech, M.; Hooshmand, B.; McCaddon, A.; Miller, J.W.; Rosenberg, I.H.; Obeid, R. Homocysteine and dementia: An international consensus statement. J. Alzheimers Dis. 2018, 62, 561–570. [Google Scholar] [CrossRef]
- Kennedy, B.P.; Bottiglieri, T.; Arning, E.; Ziegler, M.G.; Hansen, L.A.; Masliah, E. Elevated s-adenosylhomocysteine in alzheimer brain: Influence on methyltransferases and cognitive function. J. Neural. Transm. 2004, 111, 547–567. [Google Scholar] [CrossRef] [PubMed]
- Fuso, A.; Seminara, L.; Cavallaro, R.A.; D’Anselmi, F.; Scarpa, S. S-adenosylmethionine/homocysteine cycle alterations modify DNA methylation status with consequent deregulation of ps1 and bace and beta-amyloid production. Mol. Cell. Neurosci. 2005, 28, 195–204. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, R.J.; Wong, P.C. Amyloid precursor protein processing and alzheimer’s disease. Annu. Rev. NeuroSci. 2011, 34, 185–204. [Google Scholar] [CrossRef]
- Fuso, A.; Nicolia, V.; Cavallaro, R.A.; Ricceri, L.; D’Anselmi, F.; Coluccia, P.; Calamandrei, G.; Scarpa, S. B-vitamin deprivation induces hyperhomocysteinemia and brain s-adenosylhomocysteine, depletes brain s-adenosylmethionine, and enhances ps1 and bace expression and amyloid-β deposition in mice. Mol. Cell. Neurosci. 2008, 37, 731–746. [Google Scholar] [CrossRef]
- Do Carmo, S.; Hanzel, C.E.; Jacobs, M.L.; Machnes, Z.; Iulita, M.F.; Yang, J.; Yu, L.; Ducatenzeiler, A.; Danik, M.; Breuillaud, L.S.; et al. Rescue of early bace-1 and global DNA demethylation by s-adenosylmethionine reduces amyloid pathology and improves cognition in an alzheimer’s model. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.C.; Oelze, B.; Schumacher, A. Age-specific epigenetic drift in late-onset alzheimer’s disease. PLoS ONE 2008, 3, 0002698. [Google Scholar] [CrossRef] [PubMed]
- Iwata, A.; Nagata, K.; Hatsuta, H.; Takuma, H.; Bundo, M.; Iwamoto, K.; Tamaoka, A.; Murayama, S.; Saido, T.; Tsuji, S. Altered cpg methylation in spoRadic. alzheimer’s disease is associated with app and mapt dysregulation. Hum. Mol. Genet. 2013, 23, 648–656. [Google Scholar] [CrossRef]
- Fuso, A.; Nicolia, V.; Pasqualato, A.; Fiorenza, M.T.; Cavallaro, R.A.; Scarpa, S. Changes in presenilin 1 gene methylation pattern in diet-induced b vitamin deficiency. NeuroBiol. Aging 2011, 32, 187–199. [Google Scholar] [CrossRef] [PubMed]
- Fuso, A.; Nicolia, V.; Ricceri, L.; Cavallaro, R.A.; Isopi, E.; Mangia, F.; Fiorenza, M.T.; Scarpa, S. S-adenosylmethionine reduces the progress of the alzheimer-like features induced by b-vitamin deficiency in mice. NeuroBiol. Aging 2012, 33, 4. [Google Scholar] [CrossRef]
- Monti, N.; Cavallaro, R.A.; Stoccoro, A.; Nicolia, V.; Scarpa, S.; Kovacs, G.G.; Fiorenza, M.T.; Lucarelli, M.; Aronica, E.; Ferrer, I.; et al. Cpg and non-cpg presenilin1 methylation pattern in course of neurodevelopment and neurodegeneration is associated with gene expression in human and murine brain. Epigenetics 2020, 15, 781–799. [Google Scholar] [CrossRef]
- Tian, T.; Bai, D.; Li, W.; Huang, G.-W.; Liu, H. Effects of folic acid on secretases involved in aβ deposition in app/ps1 mice. Nutrients 2016, 8, 556. [Google Scholar] [CrossRef] [PubMed]
- Fuso, A.; Nicolia, V.; Cavallaro, R.A.; Scarpa, S. DNA methylase and demethylase activities are modulated by one-carbon metabolism in alzheimer’s disease models. J. Nutr. Biochem. 2011, 22, 242–251. [Google Scholar] [CrossRef]
- Li, W.; Liu, H.; Yu, M.; Zhang, X.; Zhang, M.; Wilson, J.X.; Huang, G. Folic acid administration inhibits amyloid β-peptide accumulation in app/ps1 transgenic mice. J. Nutr. Biochem. 2015, 26, 883–891. [Google Scholar] [CrossRef] [PubMed]
- Barrachina, M.; Ferrer, I. DNA methylation of alzheimer disease and tauopathy-related genes in postmortem brain. J. NeuroPathol. Exp. Neurol. 2009, 68, 880–891. [Google Scholar] [CrossRef]
- Stoccoro, A.; Tannorella, P.; Migliore, L.; Coppedè, F. Polymorphisms of genes required for methionine synthesis and DNA methylation influence mitochondrial DNA methylation. Epigenomics 2020, 12, 1003–1012. [Google Scholar] [CrossRef]
- Blanch, M.; Mosquera, J.L.; Ansoleaga, B.; Ferrer, I.; Barrachina, M. Altered mitochondrial DNA methylation pattern in alzheimer disease-related pathology and in parkinson disease. Am. J. Pathol. 2016, 186, 385–397. [Google Scholar] [CrossRef]
- Janssens, V.; Longin, S.; Goris, J. Pp2a holoenzyme assembly: In cauda venenum (the sting is in the tail). Trends BioChem. Sci. 2008, 33, 113–121. [Google Scholar] [CrossRef] [PubMed]
- Sontag, E.; Hladik, C.; Montgomery, L.; Luangpirom, A.; Mudrak, I.; Ogris, E.; White, C.L., 3rd. Downregulation of protein phosphatase 2a carboxyl methylation and methyltransferase may contribute to alzheimer disease pathogenesis. J. NeuroPathol. Exp. Neurol. 2004, 63, 1080–1091. [Google Scholar] [CrossRef]
- Zhou, X.W.; Gustafsson, J.A.; Tanila, H.; Bjorkdahl, C.; Liu, R.; Winblad, B.; Pei, J.J. Tau hyperphosphorylation correlates with reduced methylation of protein phosphatase 2a. NeuroBiol. Dis. 2008, 31, 386–394. [Google Scholar] [CrossRef]
- Sontag, J.M.; Nunbhakdi-Craig, V.; Montgomery, L.; Arning, E.; Bottiglieri, T.; Sontag, E. Folate deficiency induces in vitro and mouse brain region-specific downregulation of leucine carboxyl methyltransferase-1 and protein phosphatase 2a b(alpha) subunit expression that correlate with enhanced tau phosphorylation. J. NeuroSci. 2008, 28, 11477–11487. [Google Scholar] [CrossRef]
- Zheng, M.; Zou, C.; Li, M.; Huang, G.; Gao, Y.; Liu, H. Folic acid reduces tau phosphorylation by regulating pp2a methylation in streptozotocin-induced diabetic mice. Int. J. Mol. Sci. 2017, 18, 861. [Google Scholar] [CrossRef] [PubMed]
- Park, H.J.; Lee, K.W.; Oh, S.; Yan, R.; Zhang, J.; Beach, T.G.; Adler, C.H.; Voronkov, M.; Braithwaite, S.P.; Stock, J.B.; et al. Protein phosphatase 2a and its methylation modulating enzymes lcmt-1 and pme-1 are dysregulated in tauopathies of progressive supranuclear palsy and alzheimer disease. J. NeuroPathol. Exp. Neurol. 2018, 77, 139–148. [Google Scholar] [CrossRef] [PubMed]
- Nasa, I.; Kettenbach, A.N. Effects of carboxyl-terminal methylation on holoenzyme function of the pp2a subfamily. BioChem. Soc. Trans. 2020, 48, 2015–2027. [Google Scholar] [CrossRef]
- Li, W.; Jiang, M.; Xiao, Y.; Zhang, X.; Cui, S.; Huang, G. Folic acid inhibits tau phosphorylation through regulation of pp2a methylation in sh-sy5y cells. J. Nutr. Health Aging 2015, 19, 123–129. [Google Scholar] [CrossRef] [PubMed]
- Sontag, E.; Nunbhakdi-Craig, V.; Sontag, J.M.; Diaz-Arrastia, R.; Ogris, E.; Dayal, S.; Lentz, S.R.; Arning, E.; Bottiglieri, T. Protein phosphatase 2a methyltransferase links homocysteine metabolism with tau and amyloid precursor protein regulation. J. NeuroSci. 2007, 27, 2751–2759. [Google Scholar] [CrossRef] [PubMed]
- Taleski, G.; Schuhmacher, D.; Su, H.; Sontag, J.-M.; Sontag, E. Disturbances in pp2a methylation and one-carbon metabolism compromise fyn distribution, neuritogenesis, and app regulation. J. Biol. Chem. 2021, 296. [Google Scholar] [CrossRef] [PubMed]
- Sontag, J.-M.; Wasek, B.; Taleski, G.; Smith, J.; Arning, E.; Sontag, E.; Bottiglieri, T. Altered protein phosphatase 2a methylation and tau phosphorylation in the young and aged brain of methylenetetrahydrofolate reductase (mthfr) deficient mice. Front. Aging NeuroSci. 2014, 6, 214. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.E.; Tian, Q.; Wei, W.; Peng, J.H.; Liu, G.P.; Zhou, X.W.; Wang, Q.; Wang, D.W.; Wang, J.Z. Homocysteine induces tau phosphorylation by inactivating protein phosphatase 2a in rat hippocampus. NeuroBiol. Aging 2008, 29, 1654–1665. [Google Scholar] [CrossRef]
- Ho, P.I.; Collins, S.C.; Dhitavat, S.; Ortiz, D.; Ashline, D.; Rogers, E.; Shea, T.B. Homocysteine potentiates beta-amyloid neurotoxicity: Role of oxidative stress. J. NeuroChem. 2001, 78, 249–253. [Google Scholar] [CrossRef]
- McCaddon, A.; Hudson, P.; Hill, D.; Barber, J.; Lloyd, A.; Davies, G.; Regland, B.j. Alzheimer’s disease and total plasma aminothiols. Biol. Psychiatry 2003, 53, 254–260. [Google Scholar] [CrossRef]
- Zafrilla, P.; Mulero, J.; Xandri, M.J.; Santo, E.; Caravaca, G.; Morillas, M.J. Oxidative stress in alzheimer patients in different stages of the disease. Curr. Med. Chem. 2006, 13, 1075–1083. [Google Scholar] [CrossRef] [PubMed]
- Resende, R.; Moreira, P.I.; Proença, T.; Deshpande, A.; Busciglio, J.; Pereira, C.; Oliveira, C.R. Brain oxidative stress in a triple-transgenic mouse model of alzheimer disease. Free Radic. Biol. Med. 2008, 44, 2051–2057. [Google Scholar] [CrossRef] [PubMed]
- Cankurtaran, M.; Yesil, Y.; Kuyumcu, M.E.; Oztürk, Z.A.; Yavuz, B.B.; Halil, M.; Ulger, Z.; Cankurtaran, E.S.; Arıoğul, S. Altered levels of homocysteine and serum natural antioxidants links oxidative damage to alzheimer’s disease. J. Alzheimers Dis. 2013, 33, 1051–1058. [Google Scholar] [CrossRef]
- Ghodake, S.R.; Suryakar, A.N.; Kulhalli, P.M.; Shaikh, A.K. The study of homocysteine and its relationship with oxidative stress biomarkers in alzheimer’s disease. Int. J. Med. Sci. Clin. Invent. 2016. [Google Scholar] [CrossRef]
- Hama, Y.; Hamano, T.; Shirafuji, N.; Hayashi, K.; Ueno, A.; Enomoto, S.; Nagata, M.; Kimura, H.; Matsunaga, A.; Ikawa, M.; et al. Influences of folate supplementation on homocysteine and cognition in patients with folate deficiency and cognitive impairment. Nutrients 2020, 12, 3138. [Google Scholar] [CrossRef]
- Román, G.C. Mthfr gene mutations: A potential marker of late-onset alzheimer’s disease? J. Alzheimers Dis. 2015, 47, 323–327. [Google Scholar] [CrossRef]
- Beyer, K.; Lao, J.I.; Carrato, C.; Rodriguez-Vila, A.; Latorre, P.; Mataró, M.; Llopis, M.A.; Mate, J.L.; Ariza, A. Cystathionine beta synthase as a risk factor for alzheimer disease. Curr. Alzheimer Res. 2004, 1, 127–133. [Google Scholar] [CrossRef]
- The role of cystathionine β-synthase in homocysteine metabolism. Antioxid. Redox Signal. 2005, 7, 813–822. [CrossRef]
- Paul, B.D. Neuroprotective roles of the reverse transsulfuration pathway in alzheimer’s disease. Front. Aging NeuroSci. 2021, 13. [Google Scholar] [CrossRef]
- Tjiattas, L.; Ortiz, D.O.; Dhivant, S.; Mitton, K.; Rogers, E.; Shea, T.B. Folate deficiency and homocysteine induce toxicity in cultured dorsal root ganglion neurons via cytosolic calcium accumulation. Aging Cell 2004, 3, 71–76. [Google Scholar] [CrossRef] [PubMed]
- Poddar, R.; Paul, S. Homocysteine-nmda receptor-mediated activation of extracellular signal-regulated kinase leads to neuronal cell death. J. NeuroChem. 2009, 110, 1095–1106. [Google Scholar] [CrossRef]
- Danysz, W.; Parsons, C.G. Alzheimer’s disease, β-amyloid, glutamate, nmda receptors and memantine–Searching for the connections. Br. J. Pharm. 2012, 167, 324–352. [Google Scholar] [CrossRef]
- Övey, İ.S.; Naziroğlu, M. Homocysteine and cytosolic gsh depletion induce apoptosis and oxidative toxicity through cytosolic calcium overload in the hippocampus of aged mice: Involvement of trpm2 and trpv1 channels. Neuroscience 2015, 284, 225–233. [Google Scholar] [CrossRef]
- Kruman, I.I.; Culmsee, C.; Chan, S.L.; Kruman, Y.; Guo, Z.; Penix, L.; Mattson, M.P. Homocysteine elicits a DNA damage response in neurons that promotes apoptosis and hypersensitivity to excitotoxicity. J. Neurosci. 2000, 20, 6920–6926. [Google Scholar] [CrossRef] [PubMed]
- Vanzin, C.S.; Manfredini, V.; Marinho, A.E.; Biancini, G.B.; Ribas, G.S.; Deon, M.; Wyse, A.T.; Wajner, M.; Vargas, C.R. Homocysteine contribution to DNA damage in cystathionine β-synthase-deficient patients. Gene 2014, 539, 270–274. [Google Scholar] [CrossRef]
- Andra, A.; Tanigawa, S.; Bito, T.; Ishihara, A.; Watanabe, F.; Yabuta, Y. Effects of vitamin b(12) deficiency on amyloid-β toxicity in caenorhabditis elegans. Antioxidants 2021, 10, 962. [Google Scholar] [CrossRef]
- Lam, A.B.; Kervin, K.; Tanis, J.E. Vitamin b12 impacts amyloid beta-induced proteotoxicity by regulating the methionine/s-adenosylmethionine cycle. Cell Rep. 2021, 36, 109753. [Google Scholar] [CrossRef]
- Leiteritz, A.; Dilberger, B.; Wenzel, U.; Fitzenberger, E. Betaine reduces β-amyloid-induced paralysis through activation of cystathionine-β-synthase in an alzheimer model of caenorhabditis elegans. Genes Nutr. 2018, 13, 018–0611. [Google Scholar] [CrossRef]
- Bermejo, P.; Martín-Aragón, S.; Benedí, J.; Susín, C.; Felici, E.; Gil, P.; Ribera, J.M.; Villar, A.M. Peripheral levels of glutathione and protein oxidation as markers in the development of alzheimer’s disease from mild cognitive impairment. Free Radic. Res. 2008, 42, 162–170. [Google Scholar] [CrossRef] [PubMed]
- Mandal, P.K.; Saharan, S.; Tripathi, M.; Murari, G. Brain glutathione levels--a novel biomarker for mild cognitive impairment and alzheimer’s disease. Biol. Psychiatry 2015, 78, 702–710. [Google Scholar] [CrossRef]
- Ansari, M.A.; Scheff, S.W. Oxidative stress in the progression of alzheimer disease in the frontal cortex. J. NeuroPathol. Exp. Neurol. 2010, 69, 155–167. [Google Scholar] [CrossRef] [PubMed]
- Saharan, S.; Mandal, P.K. The emerging role of glutathione in alzheimer’s disease. J. Alzheimers Dis. 2014, 40, 519–529. [Google Scholar] [CrossRef] [PubMed]
- Poewe, W.; Seppi, K.; Tanner, C.M.; Halliday, G.M.; Brundin, P.; Volkmann, J.; Schrag, A.-E.; Lang, A.E. Parkinson disease. Nat. Rev. Dis. Primers 2017, 3, 17013. [Google Scholar] [CrossRef]
- Day, J.O.; Mullin, S. The genetics of parkinson’s disease and implications for clinical practice. Genes 2021, 12, 1006. [Google Scholar] [CrossRef] [PubMed]
- Murray, L.K.; Jadavji, N.M. The role of one-carbon metabolism and homocysteine in parkinson’s disease onset, pathology and mechanisms. Nutr. Res. Rev. 2019, 32, 218–230. [Google Scholar] [CrossRef] [PubMed]
- Fan, X.; Zhang, L.; Li, H.; Chen, G.; Qi, G.; Ma, X.; Jin, Y. Role of homocysteine in the development and progression of parkinson’s disease. Ann. Clin. Transl. Neurol. 2020, 7, 2332–2338. [Google Scholar] [CrossRef] [PubMed]
- Paul, R.; Borah, A. L-dopa-induced hyperhomocysteinemia in parkinson’s disease: Elephant in the room. Biochim. Biophys. Acta (BBA)-Gen. Subj. 2016, 1860, 1989–1997. [Google Scholar] [CrossRef] [PubMed]
- van der Steen, W.; den Heijer, T.; Groen, J. Vitamin b6 deficiency caused by the use of levodopa. Ned. Tijdschr. Geneeskd. 2018, 23, 162. [Google Scholar]
- Rojo-Sebastián, A.; González-Robles, C.; García de Yébenes, J. Vitamin b6 deficiency in patients with parkinson disease treated with levodopa/carbidopa. Clin. Neuropharmacol 2020, 43, 151–157. [Google Scholar] [CrossRef]
- Miller, J.W.; Selhub, J.; Nadeau, M.R.; Thomas, C.A.; Feldman, R.G.; Wolf, P.A. Effect of l-dopa on plasma homocysteine in pd patients: Relationship to b-vitamin status. Neurology 2003, 60, 1125–1129. [Google Scholar] [CrossRef]
- dos Santos, E.F.; Busanello, E.N.; Miglioranza, A.; Zanatta, A.; Barchak, A.G.; Vargas, C.R.; Saute, J.; Rosa, C.; Carrion, M.J.; Camargo, D.; et al. Evidence that folic acid deficiency is a major determinant of hyperhomocysteinemia in parkinson’s disease. Metab. Brain Dis. 2009, 24, 257–269. [Google Scholar] [CrossRef]
- Murakami, K.; Miyake, Y.; Sasaki, S.; Tanaka, K.; Fukushima, W.; Kiyohara, C.; Tsuboi, Y.; Yamada, T.; Oeda, T.; Miki, T.; et al. Dietary intake of folate, vitamin b6, vitamin b12 and riboflavin and risk of parkinson’s disease: A case-control study in japan. Br. J. Nutr. 2010, 104, 757–764. [Google Scholar] [CrossRef] [PubMed]
- Shen, L. Associations between b vitamins and parkinson’s disease. Nutrients 2015, 7, 7197–7208. [Google Scholar] [CrossRef] [PubMed]
- Christine, C.W.; Auinger, P.; Joslin, A.; Yelpaala, Y.; Green, R. Vitamin b12 and homocysteine levels predict different outcomes in early parkinson’s disease. Mov. Disord. 2018, 33, 762–770. [Google Scholar] [CrossRef]
- Luthra, N.S.; Marcus, A.H.; Hills, N.K.; Christine, C.W. Vitamin b12 measurements across neurodegenerative disorders. J. Clin. Mov. Disord. 2020, 7, 3. [Google Scholar] [CrossRef] [PubMed]
- Ibrahimagic, O.C.; Smajlovic, D.; Dostovic, Z.; Pasic, Z.; Kunic, S.; Iljazovic, A.; Hajdarevic, D.S. Hyperhomocysteinemia and its treatment in patients with parkinson’s disease. Mater. Sociomed 2016, 28, 303–306. [Google Scholar] [CrossRef]
- Chen, H.; Zhang, S.M.; Schwarzschild, M.A.; Hernán, M.A.; Logroscino, G.; Willett, W.C.; Ascherio, A. Folate intake and risk of parkinson’s disease. Am. J. Epidemiol. 2004, 160, 368–375. [Google Scholar] [CrossRef] [PubMed]
- de Lau, L.M.; Koudstaal, P.J.; Witteman, J.C.; Hofman, A.; Breteler, M.M. Dietary folate, vitamin b12, and vitamin b6 and the risk of parkinson disease. Neurology 2006, 67, 315–318. [Google Scholar] [CrossRef]
- Diao, H.-M.; Song, Z.-F.; Xu, H.-D. Association between mthfr genetic polymorphism and parkinson’s disease susceptibility: A meta-analysis. Open Med. 2019, 14, 613–624. [Google Scholar] [CrossRef]
- Liu, L.; Zhang, L.; Guo, L.; Yu, Q.; Li, H.; Teng, J.; Xie, A. Mthfr c677t and a1298c polymorphisms may contribute to the risk of parkinson’s disease: A meta-analysis of 19 studies. NeuroSci. Lett. 2018, 662, 339–345. [Google Scholar] [CrossRef]
- Fong, C.S.; Shyu, H.Y.; Shieh, J.C.; Fu, Y.P.; Chin, T.Y.; Wang, H.W.; Cheng, C.W. Association of mthfr, mtr, and mtrr polymorphisms with parkinson’s disease among ethnic chinese in taiwan. Clin. Chim Acta 2011, 412, 332–338. [Google Scholar] [CrossRef]
- Miranda-Morales, E.; Meier, K.; Sandoval-Carrillo, A.; Salas-Pacheco, J.; Vázquez-Cárdenas, P.; Arias-Carrión, O. Implications of DNA methylation in parkinson’s disease. Front. Mol. NeuroSci. 2017, 10. [Google Scholar] [CrossRef] [PubMed]
- Henderson, A.R.; Wang, Q.; Meechoovet, B.; Siniard, A.L.; Naymik, M.; De Both, M.; Huentelman, M.J.; Caselli, R.J.; Driver-Dunckley, E.; Dunckley, T. DNA methylation and expression profiles of whole blood in parkinson’s disease. Front. Genet. 2021, 12. [Google Scholar] [CrossRef]
- Jowaed, A.; Schmitt, I.; Kaut, O.; Wüllner, U. Methylation regulates alpha-synuclein expression and is decreased in parkinson’s disease patients’ brains. J. NeuroSci. 2010, 30, 6355–6359. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, L.; Takuma, H.; Tamaoka, A.; Kurisaki, H.; Date, H.; Tsuji, S.; Iwata, A. Cpg demethylation enhances alpha-synuclein expression and affects the pathogenesis of parkinson’s disease. PLoS ONE 2010, 5, 0015522. [Google Scholar] [CrossRef] [PubMed]
- Ai, S.X.; Xu, Q.; Hu, Y.C.; Song, C.Y.; Guo, J.F.; Shen, L.; Wang, C.R.; Yu, R.L.; Yan, X.X.; Tang, B.S. Hypomethylation of snca in blood of patients with spoRadic. parkinson’s disease. J. Neurol. Sci. 2014, 337, 123–128. [Google Scholar] [CrossRef]
- Guhathakurta, S.; Evangelista, B.A.; Ghosh, S.; Basu, S.; Kim, Y.-S. Hypomethylation of intron1 of α-synuclein gene does not correlate with parkinson’s disease. Mol. Brain 2017, 10, 6. [Google Scholar] [CrossRef] [PubMed]
- Gu, J.; Barrera, J.; Yun, Y.; Murphy, S.K.; Beach, T.G.; Woltjer, R.L.; Serrano, G.E.; Kantor, B.; Chiba-Falek, O. Cell-type specific changes in DNA methylation of snca intron 1 in synucleinopathy brains. Front. Neurosci. 2021, 15. [Google Scholar] [CrossRef]
- Obeid, R.; Schadt, A.; Dillmann, U.; Kostopoulos, P.; Fassbender, K.; Herrmann, W. Methylation status and neurodegenerative markers in parkinson disease. Clin. Chem. 2009, 55, 1852–1860. [Google Scholar] [CrossRef] [PubMed]
- Desplats, P.; Spencer, B.; Coffee, E.; Patel, P.; Michael, S.; Patrick, C.; Adame, A.; Rockenstein, E.; Masliah, E. Alpha-synuclein sequesters dnmt1 from the nucleus: A novel mechanism for epigenetic alterations in lewy body diseases. J. Biol. Chem. 2011, 286, 9031–9037. [Google Scholar] [CrossRef]
- Lüth, T.; Wasner, K.; Klein, C.; Schaake, S.; Tse, R.; Pereira, S.L.; Laß, J.; Sinkkonen, L.; Grünewald, A.; Trinh, J. Nanopore single-molecule sequencing for mitochondrial DNA methylation analysis: Investigating parkin-associated parkinsonism as a proof of concept. Front. Aging NeuroSci. 2021, 13. [Google Scholar] [CrossRef]
- Malpartida, A.B.; Williamson, M.; Narendra, D.P.; Wade-Martins, R.; Ryan, B.J. Mitochondrial dysfunction and mitophagy in parkinson’s disease: From mechanism to therapy. Trends Biochem. Sci. 2021, 46, 329–343. [Google Scholar] [CrossRef]
- Guo, J.D.; Zhao, X.; Li, Y.; Li, G.R.; Liu, X.L. Damage to dopaminergic neurons by oxidative stress in parkinson’s disease (review). Int. J. Mol. Med. 2018, 41, 1817–1825. [Google Scholar] [CrossRef]
- Bjørklund, G.; Peana, M.; Maes, M.; Dadar, M.; Severin, B. The glutathione system in parkinson’s disease and its progression. Neurosci. Biobehav. Rev. 2021, 120, 470–478. [Google Scholar] [CrossRef] [PubMed]
- Chinta, S.J.; Kumar, M.J.; Hsu, M.; Rajagopalan, S.; Kaur, D.; Rane, A.; Nicholls, D.G.; Choi, J.; Andersen, J.K. Inducible alterations of glutathione levels in adult dopaminergic midbrain neurons result in nigrostriatal degeneration. J. NeuroSci. 2007, 27, 13997–14006. [Google Scholar] [CrossRef] [PubMed]
- Srivastav, S.; Singh, S.K.; Yadav, A.K.; Srikrishna, S. Folic acid supplementation ameliorates oxidative stress, metabolic functions and developmental anomalies in a novel fly model of parkinson’s disease. Neurochem. Res. 2015, 40, 1350–1359. [Google Scholar] [CrossRef]
- Meiser, J.; Delcambre, S.; Wegner, A.; Jäger, C.; Ghelfi, J.; d’Herouel, A.F.; Dong, X.; Weindl, D.; Stautner, C.; Nonnenmacher, Y.; et al. Loss of dj-1 impairs antioxidant response by altered glutamine and serine metabolism. Neurobiol. Dis. 2016, 89, 112–125. [Google Scholar] [CrossRef] [PubMed]
| Genetic Manipulation | Environmental Manipulation | Effect on Lifespan | References |
|---|---|---|---|
| Yeast | |||
| Methionine restriction | CLS extension | [54,55] | |
| met2 deletion | CLS extension | [56,57] | |
| met15 deletion | CLS extension | [56,57] | |
| met3 deletion | RLS extension | [58] | |
| sam1 deletion | RLS extension | [58] | |
| cys4 deletion | CLS extension | [59] | |
| H2S | CLS extension | [60] | |
| gsh1 deletion | 10% glucose | CLS shortening | [61] |
| Nematodes | |||
| sulfamethoxazole | Extension | [62] | |
| dhfr-1 RNAi | extension | [63] | |
| tyms-1 RNAi | extension | [63,64] | |
| daf-2(e1370) | MTHF5 supplementation | Reverses longevity of daf-2 mutants | [63] |
| mel-32 RNAi | extension | [65] | |
| sams-1 RNAi | extension | [66,67] | |
| sams-5 RNAi | extension | [67] | |
| Glycine supplementation | extension | [65,68] | |
| Serine supplementation | extension | [65,68] | |
| Metformin | extension | [69] | |
| sams-1 mutant | Metformin | Reverse of metformin’s benefits | [69] |
| metr-1 mutant | Metformin | Reverse of metformin’s benefits | [69] |
| H2S | extension | [70,71] | |
| cbs-1 overexpression | extension | [60] | |
| NAC from day 3 | extension | [72] | |
| NAC from L4 | shortening | [73] | |
| taurine | extension | [68] | |
| Acivicin (GSH restriction) | extension | [73] | |
| Flies | |||
| GNMT overexpression | extension | [74] | |
| Methionine restriction | extension | [75,76] | |
| Sams depletion | extension | [74] | |
| Cbs overexpression | extension | [77] | |
| Cbs depletion | Caloric restriction | Reversed CR-driven longevity | [78] |
| Gclc overexpression | extension | [79] | |
| Gclm overexpression | extension | [79] | |
| dAhcyL1/dAhcyL2 suppression | Extension | [80] | |
| Mammals | |||
| Transgenic growth hormone mice | Supplementation of several forms of folate | Extension | [81] |
| wt | Supplementation of several forms of folate | Extension | [81] |
| Female SHR mice | metformin | Extension | [82] |
| Rats, mice | Dietary Methionine restriction | Extension | [83,84,85] |
| Tissue-specific taurine transporter depleted mice | Shortening | [86] | |
| Humans | |||
| C667T MTHFR | Associates with decrease in all-cause mortality | [87,88] | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lionaki, E.; Ploumi, C.; Tavernarakis, N. One-Carbon Metabolism: Pulling the Strings behind Aging and Neurodegeneration. Cells 2022, 11, 214. https://doi.org/10.3390/cells11020214
Lionaki E, Ploumi C, Tavernarakis N. One-Carbon Metabolism: Pulling the Strings behind Aging and Neurodegeneration. Cells. 2022; 11(2):214. https://doi.org/10.3390/cells11020214
Chicago/Turabian StyleLionaki, Eirini, Christina Ploumi, and Nektarios Tavernarakis. 2022. "One-Carbon Metabolism: Pulling the Strings behind Aging and Neurodegeneration" Cells 11, no. 2: 214. https://doi.org/10.3390/cells11020214
APA StyleLionaki, E., Ploumi, C., & Tavernarakis, N. (2022). One-Carbon Metabolism: Pulling the Strings behind Aging and Neurodegeneration. Cells, 11(2), 214. https://doi.org/10.3390/cells11020214