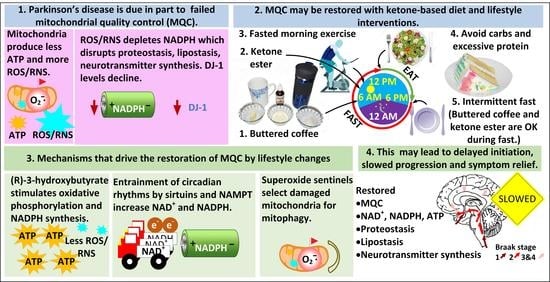
NADPH and Mitochondrial Quality Control as Targets for a Circadian-Based Fasting and Exercise Therapy for the Treatment of Parkinson’s Disease
Abstract
1. A Strong Rationale for Targeting MQC in PD
2. Interventional Strategies to Correct Dysfunctional MQC and Downstream Processes
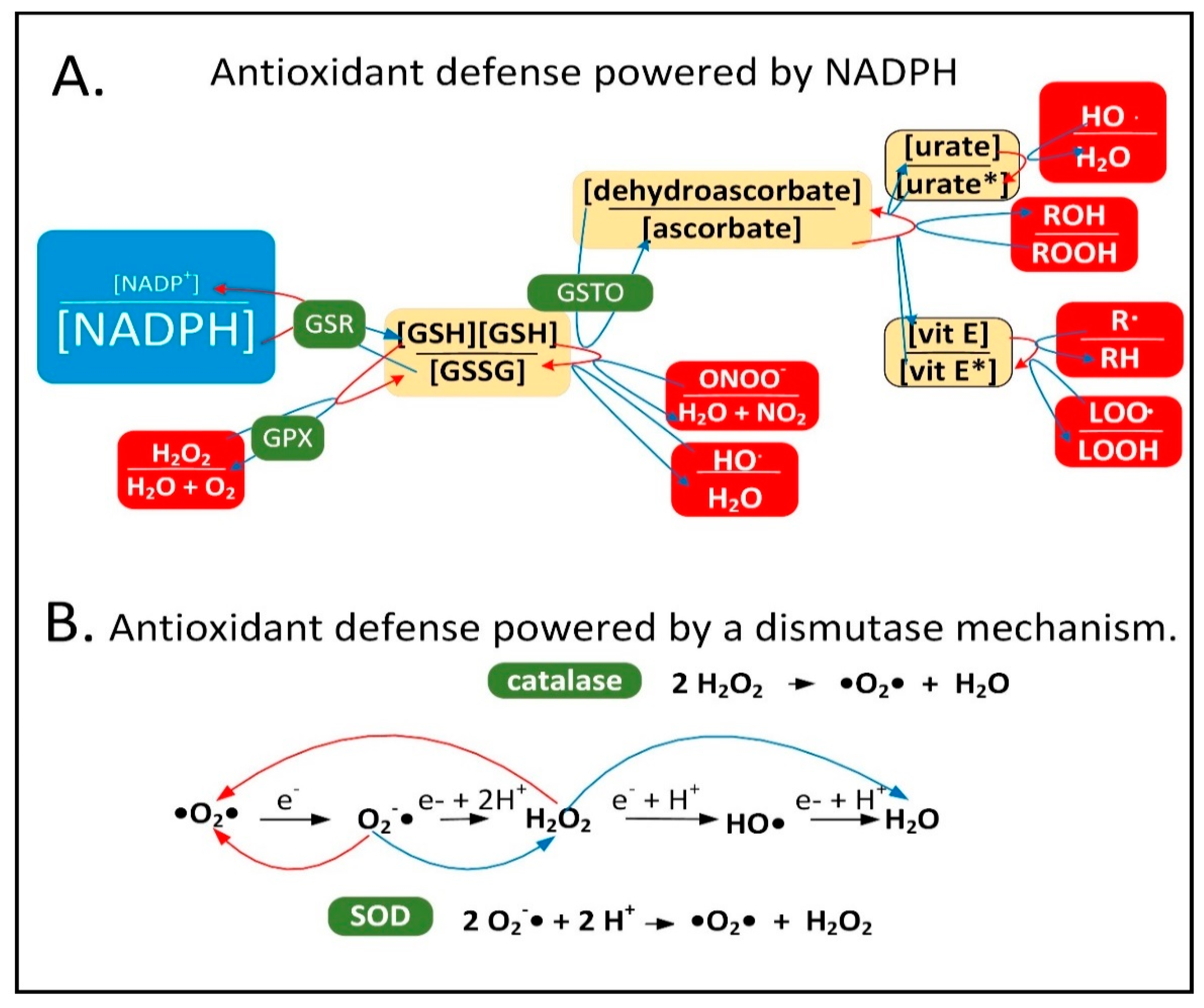
2.1. NADPH Is Required for Proper MQC
2.2. Mechanisms of Mitochondrial Protein Turnover
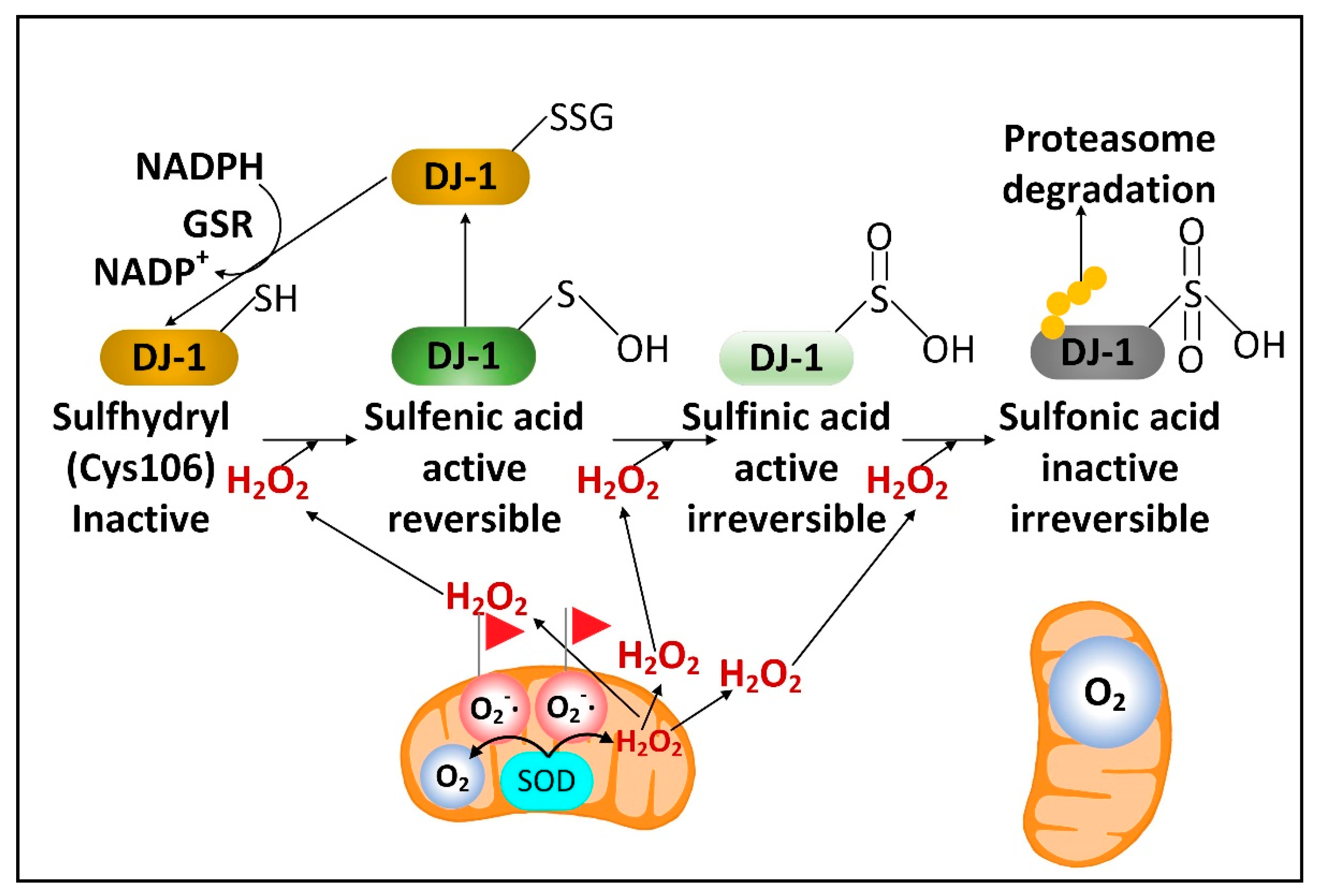
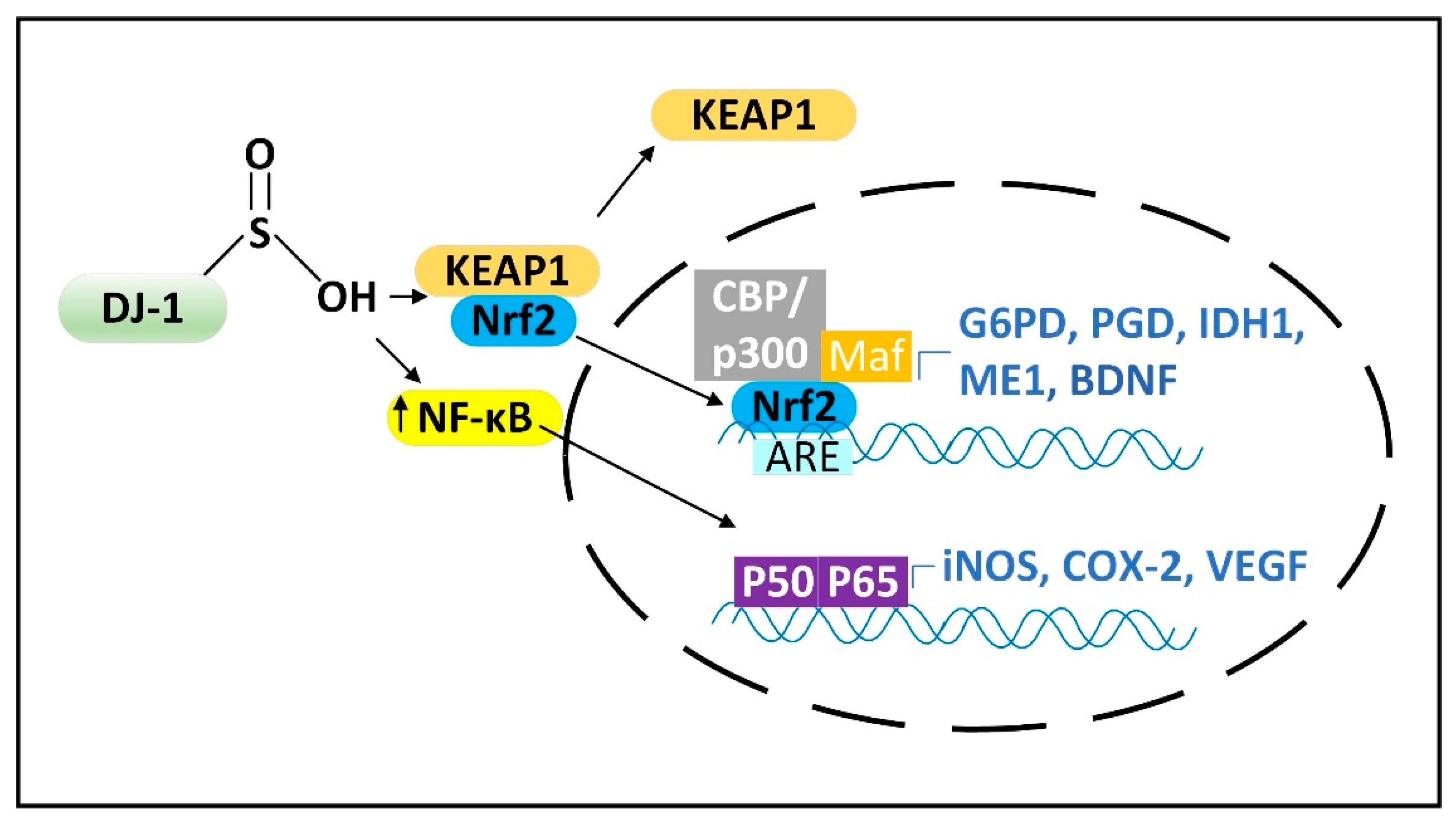
2.3. DJ-1 Is Regulated by Cytoplasmic NADPH and Glutathione and Controls Nrf2 Activation in Astrocytes
2.4. Fasting Increases the Hepatic Levels of ATF4 to Increase NADPH, FGF21, and Parkin
2.5. Fasting Activates Glutamate Dehydrogenase to Increase the Levels of the Anti-Aging Metabolite αkg
2.6. Fasting Decreases the Cytoplasmic [NADP+]/[NADPH] Ratio in Liver, but Its Effects on Neural Cell [NADP+]/[NADPH] Ratios Are Not Yet Known
2.7. Fasting May Improve PD Symptoms by Decreasing the Number of Senescent Astrocytes and Microglia without Affecting Substantia Nigra Mitochondrial DNA Deletion Levels
2.8. A Requirement for Fasting in Dietary Restriction-Induced Longevity
2.9. Evidence That Combining Fasting and Exercise Is Neuroprotective
2.10. Exercise and Its Effects on PD Brain and Muscles
2.11. Greater Metabolic Changes Induced by Exercise When the Exercise Occurs Early in the Active Period of the 24 h Circadian Cycle
2.12. Entrainment of the Circadian-Regulated Production of NAD+ in the Morning
2.13. Circadian Regulation of Mitophagy and Mitochondrial Dynamics May Play a Role in PD
3. Dopaminergic Neurons Are Vulnerable to Dysfunctional MQC
3.1. The Superoxide Sentinel Hypothesis of MQC
3.2. Neurons with Low-Quality Damaged Mitochondria Likely Show Decreased Proteolysis of α-Synuclein
3.3. Limited Clearance of ROS/RNS Leads to the Oxidation and Depletion of Cardiolipin and Plasmalogens
3.4. NADPH May Decrease in PD Neurons and Glia Limiting the Synthesis of Serotonin, Melatonin, Epinephrine, Norepinephrine, and •NO
3.5. Insulin Released after Feeding Increases Cytoplasmic NADPH Oxidation by Stimulating Fatty Acid Synthesis
3.6. Experiments Are Needed to Determine the (Free) Cytoplasmic [NADP+]/[NADPH] Ratio in Neural Cells from Aged and PD Model Mice and How These Are Altered by Fasting
3.7. Dysfunctional MQC Leads to Loss of Circadian Entrainment of NAD+ Synthesis
4. A Five-Pronged Strategy to Improve MQC for the Treatment of PD
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Panicker, N.; Ge, P.; Dawson, V.L.; Dawson, T.M. The Cell Biology of Parkinson’s Disease. J. Cell Biol. 2021, 220, e202012095. [Google Scholar] [CrossRef] [PubMed]
- Singh, F.; Ganley, I.G. Parkinson’s Disease and Mitophagy: An Emerging Role for LRRK2. Biochem. Soc. Trans. 2021, 49, 551–562. [Google Scholar] [CrossRef] [PubMed]
- Georgakopoulos, N.D.; Wells, G.; Campanella, M. The Pharmacological Regulation of Cellular Mitophagy. Nat. Chem. Biol. 2017, 13, 136–146. [Google Scholar] [CrossRef] [PubMed]
- Braak, H.; Del Tredici, K.; Rüb, U.; de Vos, R.A.I.; Jansen Steur, E.N.H.; Braak, E. Staging of Brain Pathology Related to Sporadic Parkinson’s Disease. Neurobiol. Aging 2003, 24, 197–211. [Google Scholar] [CrossRef]
- Butkovich, L.M.; Houser, M.C.; Chalermpalanupap, T.; Porter-Stransky, K.A.; Iannitelli, A.F.; Boles, J.S.; Lloyd, G.M.; Coomes, A.S.; Eidson, L.N.; De Sousa Rodrigues, M.E.; et al. Transgenic Mice Expressing Human α-Synuclein in Noradrenergic Neurons Develop Locus Ceruleus Pathology and Nonmotor Features of Parkinson’s Disease. J. Neurosci. 2020, 40, 7559–7576. [Google Scholar] [CrossRef]
- Eldeeb, M.A.; Thomas, R.A.; Ragheb, M.A.; Fallahi, A.; Fon, E.A. Mitochondrial Quality Control in Health and in Parkinson’s Disease. Physiol. Rev. 2022, 102, 1721–1755. [Google Scholar] [CrossRef]
- Rafaeloff-Phail, R.; Ding, L.; Conner, L.; Yeh, W.-K.; McClure, D.; Guo, H.; Emerson, K.; Brooks, H. Biochemical Regulation of Mammalian AMP-Activated Protein Kinase Activity by NAD and NADH. J. Biol. Chem. 2004, 279, 52934–52939. [Google Scholar] [CrossRef] [PubMed]
- Egan, D.F.; Shackelford, D.B.; Mihaylova, M.M.; Gelino, S.; Kohnz, R.A.; Mair, W.; Vasquez, D.S.; Joshi, A.; Gwinn, D.M.; Taylor, R.; et al. Phosphorylation of ULK1 (HATG1) by AMP-Activated Protein Kinase Connects Energy Sensing to Mitophagy. Science 2011, 331, 456–461. [Google Scholar] [CrossRef]
- Li, W.; Kou, J.; Qin, J.; Li, L.; Zhang, Z.; Pan, Y.; Xue, Y.; Du, W. NADPH Levels Affect Cellular Epigenetic State by Inhibiting HDAC3–Ncor Complex. Nat. Metab. 2021, 3, 75–89. [Google Scholar] [CrossRef]
- Sun, X.-Y.; Qu, Y.; Ni, A.-R.; Wang, G.-X.; Huang, W.-B.; Chen, Z.-P.; Lv, Z.-F.; Zhang, S.; Lindsay, H.; Zhao, S.; et al. Novel Histone Deacetylase Inhibitor N25 Exerts Anti-Tumor Effects and Induces Autophagy in Human Glioma Cells by Inhibiting HDAC3. Oncotarget 2017, 8, 75232–75242. [Google Scholar] [CrossRef][Green Version]
- Huang, S.; Chen, G.; Sun, J.; Chen, Y.; Wang, N.; Dong, Y.; Shen, E.; Hu, Z.; Gong, W.; Jin, L.; et al. Histone Deacetylase 3 Inhibition Alleviates Type 2 Diabetes Mellitus-Induced Endothelial Dysfunction via Nrf2. Cell Commun. Signal. 2021, 19, 35. [Google Scholar] [CrossRef] [PubMed]
- Vogelauer, M.; Krall, A.S.; McBrian, M.A.; Li, J.-Y.; Kurdistani, S.K. Stimulation of Histone Deacetylase Activity by Metabolites of Intermediary Metabolism. J. Biol. Chem. 2012, 287, 32006–32016. [Google Scholar] [CrossRef] [PubMed]
- Murari, A.; Goparaju, N.S.V.; Rhooms, S.-K.; Hossain, K.F.B.; Liang, F.G.; Garcia, C.J.; Osei, C.; Liu, T.; Li, H.; Kitsis, R.N.; et al. IDH2-Mediated Regulation of the Biogenesis of the Oxidative Phosphorylation System. Sci. Adv. 2022, 8, eabl8716. [Google Scholar] [CrossRef] [PubMed]
- Abdrakhmanova, A.; Zwicker, K.; Kerscher, S.; Zickermann, V.; Brandt, U. Tight Binding of NADPH to the 39-KDa Subunit of Complex I Is Not Required for Catalytic Activity but Stabilizes the Multiprotein Complex. Biochim. Biophys. Acta Bioenerg. 2006, 1757, 1676–1682. [Google Scholar] [CrossRef]
- McCarty, M.F.; Lerner, A. Nutraceuticals Targeting Generation and Oxidant Activity of Peroxynitrite May Aid Prevention and Control of Parkinson’s Disease. Int. J. Mol. Sci. 2020, 21, 3624. [Google Scholar] [CrossRef]
- Rodriguez-Enriquez, S.; He, L.; Lemasters, J.J. Role of Mitochondrial Permeability Transition Pores in Mitochondrial Autophagy. Int. J. Biochem. Cell Biol. 2004, 36, 2463–2472. [Google Scholar] [CrossRef]
- Lemasters, J.J.; Nieminen, A.-L.; Qian, T.; Trost, L.C.; Elmore, S.P.; Nishimura, Y.; Crowe, R.A.; Cascio, W.E.; Bradham, C.A.; Brenner, D.A.; et al. The Mitochondrial Permeability Transition in Cell Death: A Common Mechanism in Necrosis, Apoptosis and Autophagy. Biochim. Biophys. Acta Bioenerg. 1998, 1366, 177–196. [Google Scholar] [CrossRef]
- Salvi, M.; Battaglia, V.; Brunati, A.M.; Rocca, N.L.; Tibaldi, E.; Pietrangeli, P.; Marcocci, L.; Mondovi, B.; Rossi, C.A.; Toninello, A. Catalase Takes Part in Rat Liver Mitochondria Oxidative Stress Defense. J. Biol. Chem. 2007, 282, 24407–24415. [Google Scholar] [CrossRef] [PubMed]
- Radi, R.; Turrens, J.F.; Chang, L.Y.; Bush, K.M.; Crapo, J.D.; Freeman, B.A. Detection of Catalase in Rat Heart Mitochondria. J. Biol. Chem. 1991, 266, 22028–22034. [Google Scholar] [CrossRef]
- Menzies, R.A.; Gold, P.H. The Turnover of Mitochondria in a Variety of Tissues of Young Adult and Aged Rats. J. Biol. Chem. 1971, 246, 2425–2429. [Google Scholar] [CrossRef]
- Menzies, R.A.; Gold, P.H. The Apparent Turnover of Mitochondria, Ribosomes and SRNA of the Brain in Young Adult and Aged Rats. J. Neurochem. 1972, 19, 1671–1683. [Google Scholar] [CrossRef] [PubMed]
- Beattie, D.S.; Basford, R.E.; Koritz, S.B. The Turnover of the Protein Components of Mitochondria from Rat Liver, Kidney, and Brain. J. Biol. Chem. 1967, 242, 4584–4586. [Google Scholar] [CrossRef]
- Stotland, A.; Gottlieb, R.A. Mitochondrial Quality Control: Easy Come, Easy Go. Biochim. Biophys. Acta Mol. Cell Res. 2015, 1853, 2802–2811. [Google Scholar] [CrossRef] [PubMed]
- Bomba-Warczak, E.; Edassery, S.L.; Hark, T.J.; Savas, J.N. Long-Lived Mitochondrial Cristae Proteins in Mouse Heart and Brain. J. Cell Biol. 2021, 220, e202005193. [Google Scholar] [CrossRef]
- Krishna, S.; Arrojo E Drigo, R.; Capitanio, J.S.; Ramachandra, R.; Ellisman, M.; Hetzer, M.W. Identification of Long-Lived Proteins in the Mitochondria Reveals Increased Stability of the Electron Transport Chain. Dev. Cell 2021, 56, 2952–2965.e9. [Google Scholar] [CrossRef]
- Twig, G.; Elorza, A.; Molina, A.J.A.; Mohamed, H.; Wikstrom, J.D.; Walzer, G.; Stiles, L.; Haigh, S.E.; Katz, S.; Las, G.; et al. Fission and Selective Fusion Govern Mitochondrial Segregation and Elimination by Autophagy. EMBO J. 2008, 27, 433–446. [Google Scholar] [CrossRef]
- Kluever, V.; Russo, B.; Mandad, S.; Kumar, N.H.; Alevra, M.; Ori, A.; Rizzoli, S.O.; Urlaub, H.; Schneider, A.; Fornasiero, E.F. Protein Lifetimes in Aged Brains Reveal a Proteostatic Adaptation Linking Physiological Aging to Neurodegeneration. Sci. Adv. 2022, 8, eabn4437. [Google Scholar] [CrossRef]
- Misgeld, T.; Schwarz, T.L. Mitostasis in Neurons: Maintaining Mitochondria in an Extended Cellular Architecture. Neuron 2017, 96, 651–666. [Google Scholar] [CrossRef]
- Lamberts, J.T.; Hildebrandt, E.N.; Brundin, P. Spreading of α-Synuclein in the Face of Axonal Transport Deficits in Parkinson’s Disease: A Speculative Synthesis. Neurobiol. Dis. 2015, 77, 276–283. [Google Scholar] [CrossRef]
- Gomez-Fabra Gala, M.; Vögtle, F. Mitochondrial Proteases in Human Diseases. FEBS Lett. 2021, 595, 1205–1222. [Google Scholar] [CrossRef]
- Chen, Y.; Azad, M.B.; Gibson, S.B. Superoxide Is the Major Reactive Oxygen Species Regulating Autophagy. Cell Death Differ. 2009, 16, 1040–1052. [Google Scholar] [CrossRef] [PubMed]
- Bandopadhyay, R.; Kingsbury, A.E.; Cookson, M.R.; Reid, A.R.; Evans, I.M.; Hope, A.D.; Pittman, A.M.; Lashley, T.; Canet-Aviles, R.; Miller, D.W.; et al. The Expression of DJ-1 (PARK7) in Normal Human CNS and Idiopathic Parkinson’s Disease. Brain 2004, 127, 420–430. [Google Scholar] [CrossRef] [PubMed]
- Blackinton, J.; Lakshminarasimhan, M.; Thomas, K.J.; Ahmad, R.; Greggio, E.; Raza, A.S.; Cookson, M.R.; Wilson, M.A. Formation of a Stabilized Cysteine Sulfinic Acid Is Critical for the Mitochondrial Function of the Parkinsonism Protein DJ-1. J. Biol. Chem. 2009, 284, 6476–6485. [Google Scholar] [CrossRef] [PubMed]
- Holmström, K.M.; Kostov, R.V.; Dinkova-Kostova, A.T. The Multifaceted Role of Nrf2 in Mitochondrial Function. Curr. Opin. Toxicol. 2016, 1, 80–91. [Google Scholar] [CrossRef]
- Yao, W.; Lin, S.; Su, J.; Cao, Q.; Chen, Y.; Chen, J.; Zhang, Z.; Hashimoto, K.; Qi, Q.; Zhang, J. Activation of BDNF by Transcription Factor Nrf2 Contributes to Antidepressant-like Actions in Rodents. Transl. Psychiatry 2021, 11, 140. [Google Scholar] [CrossRef]
- Dolgacheva, L.P.; Berezhnov, A.V.; Fedotova, E.I.; Zinchenko, V.P.; Abramov, A.Y. Role of DJ-1 in the Mechanism of Pathogenesis of Parkinson’s Disease. J. Bioenerg. Biomembr. 2019, 51, 175–188. [Google Scholar] [CrossRef]
- Guzman, J.N.; Sanchez-Padilla, J.; Wokosin, D.; Kondapalli, J.; Ilijic, E.; Schumacker, P.T.; Surmeier, D.J. Oxidant Stress Evoked by Pacemaking in Dopaminergic Neurons Is Attenuated by DJ-1. Nature 2010, 468, 696–700. [Google Scholar] [CrossRef]
- Xu, S.; Yang, X.; Qian, Y.; Xiao, Q. Parkinson’s Disease-Related DJ-1 Modulates the Expression of Uncoupling Protein 4 against Oxidative Stress. J. Neurochem. 2018, 145, 312–322. [Google Scholar] [CrossRef]
- He, C.H.; Gong, P.; Hu, B.; Stewart, D.; Choi, M.E.; Choi, A.M.K.; Alam, J. Identification of Activating Transcription Factor 4 (ATF4) as an Nrf2-Interacting Protein: Implication for Heme Oxygenase-1 Gene Regulation. J. Biol. Chem. 2001, 276, 20858–20865. [Google Scholar] [CrossRef]
- Zong, Z.-H.; Du, Z.-X.; Li, N.; Li, C.; Zhang, Q.; Liu, B.-Q.; Guan, Y.; Wang, H.-Q. Implication of Nrf2 and ATF4 in Differential Induction of CHOP by Proteasome Inhibition in Thyroid Cancer Cells. Biochim. Biophys. Acta Mol. Cell Res. 2012, 1823, 1395–1404. [Google Scholar] [CrossRef]
- Almeida, L.M.; Pinho, B.R.; Duchen, M.R.; Oliveira, J.M.A. The PERKs of Mitochondria Protection during Stress: Insights for PERK Modulation in Neurodegenerative and Metabolic Diseases. Biol. Rev. Camb. Philos. Soc. 2022; early view. [Google Scholar] [CrossRef]
- Lange, P.S.; Chavez, J.C.; Pinto, J.T.; Coppola, G.; Sun, C.-W.; Townes, T.M.; Geschwind, D.H.; Ratan, R.R. ATF4 Is an Oxidative Stress–Inducible, Prodeath Transcription Factor in Neurons in Vitro and in Vivo. J. Exp. Med. 2008, 205, 1227–1242. [Google Scholar] [CrossRef] [PubMed]
- Quirós, P.M.; Prado, M.A.; Zamboni, N.; D’Amico, D.; Williams, R.W.; Finley, D.; Gygi, S.P.; Auwerx, J. Multi-Omics Analysis Identifies ATF4 as a Key Regulator of the Mitochondrial Stress Response in Mammals. J. Cell Biol. 2017, 216, 2027–2045. [Google Scholar] [CrossRef] [PubMed]
- Kasai, S.; Yamazaki, H.; Tanji, K.; Engler, M.J.; Matsumiya, T.; Itoh, K. Role of the ISR-ATF4 Pathway and Its Cross Talk with Nrf2 in Mitochondrial Quality Control. J. Clin. Biochem. Nutr. 2019, 64, 18–37. [Google Scholar] [CrossRef] [PubMed]
- Torrence, M.E.; MacArthur, M.R.; Hosios, A.M.; Valvezan, A.J.; Asara, J.M.; Mitchell, J.R.; Manning, B.D. The MTORC1-Mediated Activation of ATF4 Promotes Protein and Glutathione Synthesis Downstream of Growth Signals. eLife 2021, 10, e63326. [Google Scholar] [CrossRef]
- Renz, P.F.; Valdivia-Francia, F.; Sendoel, A. Some like It Translated: Small ORFs in the 5′UTR. Exp. Cell Res. 2020, 396, 112229. [Google Scholar] [CrossRef]
- Endo, J.; Sano, M.; Katayama, T.; Hishiki, T.; Shinmura, K.; Morizane, S.; Matsuhashi, T.; Katsumata, Y.; Zhang, Y.; Ito, H.; et al. Metabolic Remodeling Induced by Mitochondrial Aldehyde Stress Stimulates Tolerance to Oxidative Stress in the Heart. Circ. Res. 2009, 105, 1118–1127. [Google Scholar] [CrossRef]
- Wang, X.; Zhang, G.; Dasgupta, S.; Niewold, E.L.; Li, C.; Li, Q.; Luo, X.; Tan, L.; Ferdous, A.; Lorenzi, P.L.; et al. ATF4 Protects the Heart From Failure by Antagonizing Oxidative Stress. Circ. Res. 2022, 131, 91–105. [Google Scholar] [CrossRef]
- Ben-Sahra, I.; Hoxhaj, G.; Ricoult, S.J.H.; Asara, J.M.; Manning, B.D. MTORC1 Induces Purine Synthesis Through Control of the Mitochondrial Tetrahydrofolate Cycle. Science 2016, 351, 728–733. [Google Scholar] [CrossRef]
- Hill, C.M.; Albarado, D.C.; Coco, L.G.; Spann, R.A.; Khan, M.S.; Qualls-Creekmore, E.; Burk, D.H.; Burke, S.J.; Collier, J.J.; Yu, S.; et al. FGF21 Is Required for Protein Restriction to Extend Lifespan and Improve Metabolic Health in Male Mice. Nat. Commun. 2022, 13, 1897. [Google Scholar] [CrossRef]
- Zhang, Y.; Xie, Y.; Berglund, E.D.; Coate, K.C.; He, T.T.; Katafuchi, T.; Xiao, G.; Potthoff, M.J.; Wei, W.; Wan, Y.; et al. The Starvation Hormone, Fibroblast Growth Factor-21, Extends Lifespan in Mice. eLife 2012, 1, e00065. [Google Scholar] [CrossRef]
- Klaus, S.; Igual Gil, C.; Ost, M. Regulation of Diurnal Energy Balance by Mitokines. Cell Mol. Life Sci. 2021, 78, 3369–3384. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.H.; Kim, S.H.; Min, Y.-K.; Yang, H.-M.; Lee, J.-B.; Lee, M.-S. Acute Exercise Induces FGF21 Expression in Mice and in Healthy Humans. PLoS ONE 2013, 8, e63517. [Google Scholar] [CrossRef] [PubMed]
- Wan, X.; Lu, X.; Xiao, Y.; Lin, Y.; Zhu, H.; Ding, T.; Yang, Y.; Huang, Y.; Zhang, Y.; Liu, Y.-L.; et al. ATF4- and CHOP-Dependent Induction of FGF21 through Endoplasmic Reticulum Stress. BioMed Res. Int. 2014, 2014, e807874. [Google Scholar] [CrossRef] [PubMed]
- Bouman, L.; Schlierf, A.; Lutz, A.K.; Shan, J.; Deinlein, A.; Kast, J.; Galehdar, Z.; Palmisano, V.; Patenge, N.; Berg, D.; et al. Parkin Is Transcriptionally Regulated by ATF4: Evidence for an Interconnection between Mitochondrial Stress and ER Stress. Cell Death. Differ. 2011, 18, 769–782. [Google Scholar] [CrossRef]
- Li, W.; Li, X.; Miller, R.A. ATF4 Activity: A Common Feature Shared by Many Kinds of Slow-Aging Mice. Aging Cell 2014, 13, 1012–1018. [Google Scholar] [CrossRef]
- Koyanagi, S.; Hamdan, A.M.; Horiguchi, M.; Kusunose, N.; Okamoto, A.; Matsunaga, N.; Ohdo, S. CAMP-Response Element (CRE)-Mediated Transcription by Activating Transcription Factor-4 (ATF4) Is Essential for Circadian Expression of the Period2 Gene. J. Biol. Chem. 2011, 286, 32416–32423. [Google Scholar] [CrossRef]
- Pathak, S.S.; Liu, D.; Li, T.; de Zavalia, N.; Zhu, L.; Li, J.; Karthikeyan, R.; Alain, T.; Liu, A.C.; Storch, K.-F.; et al. The EIF2α Kinase GCN2 Modulates Period and Rhythmicity of the Circadian Clock by Translational Control of Atf4. Neuron 2019, 104, 724–735.e6. [Google Scholar] [CrossRef]
- Haigis, M.C.; Mostoslavsky, R.; Haigis, K.M.; Fahie, K.; Christodoulou, D.C.; Murphy, A.J.; Valenzuela, D.M.; Yancopoulos, G.D.; Karow, M.; Blander, G.; et al. SIRT4 Inhibits Glutamate Dehydrogenase and Opposes the Effects of Calorie Restriction in Pancreatic β Cells. Cell 2006, 126, 941–954. [Google Scholar] [CrossRef]
- Shaw, E.; Talwadekar, M.; Rashida, Z.; Mohan, N.; Acharya, A.; Khatri, S.; Laxman, S.; Kolthur-Seetharam, U. Anabolic SIRT4 Exerts Retrograde Control over TORC1 Signaling by Glutamine Sparing in the Mitochondria. Mol. Cell Biol. 2020, 40, e00212-19. [Google Scholar] [CrossRef]
- de Goede, P.; Wüst, R.C.I.; Schomakers, B.V.; Denis, S.; Vaz, F.M.; Pras-Raves, M.L.; van Weeghel, M.; Yi, C.-X.; Kalsbeek, A.; Houtkooper, R.H. Time-Restricted Feeding during the Inactive Phase Abolishes the Daily Rhythm in Mitochondrial Respiration in Rat Skeletal Muscle. FASEB J. 2022, 36, e22133. [Google Scholar] [CrossRef]
- Chin, R.M.; Fu, X.; Pai, M.Y.; Vergnes, L.; Hwang, H.; Deng, G.; Diep, S.; Lomenick, B.; Meli, V.S.; Monsalve, G.C.; et al. The Metabolite Alpha-Ketoglutarate Extends Lifespan by Inhibiting the ATP Synthase and TOR. Nature 2014, 510, 397–401. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Wang, T.; Wu, N.; Li, D.; Fan, X.; Xu, Z.; Mishra, S.K.; Yang, M. Alpha-Ketoglutarate Extends Drosophila Lifespan by Inhibiting MTOR and Activating AMPK. Aging 2019, 11, 4183–4197. [Google Scholar] [CrossRef] [PubMed]
- Shahmirzadi, A.A.; Edgar, D.; Liao, C.-Y.; Hsu, Y.-M.; Lucanic, M.; Shahmirzadi, A.A.; Wiley, C.D.; Gan, G.; Kim, D.E.; Kasler, H.G.; et al. Alpha-Ketoglutarate, an Endogenous Metabolite, Extends Lifespan and Compresses Morbidity in Aging Mice. Cell Metab. 2020, 32, 447–456.e6. [Google Scholar] [CrossRef] [PubMed]
- Harrison, A.P.; Pierzynowski, S.G. Biological Effects of 2-Oxoglutarate with Particular Emphasis on the Regulation of Protein, Mineral and Lipid Absorption/Metabolism, Muscle Performance, Kidney Function, Bone Formation and Cancerogenesis, All Viewed from a Healthy Ageing Perspective State of the Art—Review Article. J. Physiol. Pharmacol. 2008, 59, 91–106. [Google Scholar]
- Lin, A.-L.; Zhang, W.; Gao, X.; Watts, L. Caloric Restriction Increases Ketone Bodies Metabolism and Preserves Blood Flow in Aging Brain. Neurobiol. Aging 2015, 36, 2296–2303. [Google Scholar] [CrossRef]
- Shrimali, N.M.; Agarwal, S.; Kaur, S.; Bhattacharya, S.; Bhattacharyya, S.; Prchal, J.T.; Guchhait, P. α-Ketoglutarate Inhibits Thrombosis and Inflammation by Prolyl Hydroxylase-2 Mediated Inactivation of Phospho-Akt. eBioMedicine 2021, 73, 103672. [Google Scholar] [CrossRef]
- Wang, Y.; Deng, P.; Liu, Y.; Wu, Y.; Chen, Y.; Guo, Y.; Zhang, S.; Zheng, X.; Zhou, L.; Liu, W.; et al. Alpha-Ketoglutarate Ameliorates Age-Related Osteoporosis via Regulating Histone Methylations. Nat. Commun. 2020, 11, 5596. [Google Scholar] [CrossRef]
- Satpute, R.; Lomash, V.; Kaushal, M.; Bhattacharya, R. Neuroprotective Effects of Alpha-Ketoglutarate and Ethyl Pyruvate against Motor Dysfunction and Oxidative Changes Caused by Repeated 1-Methyl-4-Phenyl-1,2,3,6 Tetrahydropyridine Exposure in Mice. Hum. Exp. Toxicol. 2013, 32, 747–758. [Google Scholar] [CrossRef]
- Yang, Y.R.; Kwon, K.-S. Potential Roles of Exercise-Induced Plasma Metabolites Linking Exercise to Health Benefits. Front. Physiol. 2020, 11, 602748. [Google Scholar] [CrossRef]
- Daneshmandi, S.; Cassel, T.; Higashi, R.M.; Fan, T.W.-M.; Seth, P. 6-Phosphogluconate Dehydrogenase (6PGD), a Key Checkpoint in Reprogramming of Regulatory T Cells Metabolism and Function. eLife 2021, 10, e67476. [Google Scholar] [CrossRef]
- Liu, Q.; Zhu, F.; Liu, X.; Lu, Y.; Yao, K.; Tian, N.; Tong, L.; Figge, D.A.; Wang, X.; Han, Y.; et al. Non-Oxidative Pentose Phosphate Pathway Controls Regulatory T Cell Function by Integrating Metabolism and Epigenetics. Nat. Metab. 2022, 4, 559–574. [Google Scholar] [CrossRef] [PubMed]
- Moebus, S.; Göres, L.; Lösch, C.; Jöckel, K.-H. Impact of Time since Last Caloric Intake on Blood Glucose Levels. Eur. J. Epidemiol. 2011, 26, 719–728. [Google Scholar] [CrossRef] [PubMed]
- Gannon, M.C.; Nuttall, F.Q.; Lane, J.T.; Fang, S.; Gupta, V.; Sandhofer, C.R. Effect of 24 Hours of Starvation on Plasma Glucose and Insulin Concentrations in Subjects with Untreated Non—Insulin-Dependent Diabetes Mellitus. Metabolism 1996, 45, 492–497. [Google Scholar] [CrossRef]
- Owen, O.E.; Felig, P.; Morgan, A.P.; Wahren, J.; Cahill, G.F. Liver and Kidney Metabolism during Prolonged Starvation. J. Clin. Investig. 1969, 48, 574–583. [Google Scholar] [CrossRef] [PubMed]
- Cahill, G.F., Jr. Fuel Metabolism in Starvation. Annu. Rev. Nutr. 2006, 26, 1–22. [Google Scholar] [CrossRef]
- Minich, T.; Yokota, S.; Dringen, R. Cytosolic and Mitochondrial Isoforms of NADP+-Dependent Isocitrate Dehydrogenases Are Expressed in Cultured Rat Neurons, Astrocytes, Oligodendrocytes and Microglial Cells. J. Neurochem. 2003, 86, 605–614. [Google Scholar] [CrossRef]
- Kurz, G.M.; Wiesinger, H.; Hamprecht, B. Purification of Cytosolic Malic Enzyme from Bovine Brain, Generation of Monoclonal Antibodies, and Immunocytochemical Localization of the Enzyme in Glial Cells of Neural Primary Cultures. J. Neurochem. 1993, 60, 1467–1474. [Google Scholar] [CrossRef]
- Jiménez-Jiménez, F.J.; Alonso-Navarro, H.; García-Martín, E.; Agúndez, J.A.G. Cerebrospinal and Blood Levels of Amino Acids as Potential Biomarkers for Parkinson’s Disease: Review and Meta-analysis. Eur. J. Neurol. 2020, 27, 2336–2347. [Google Scholar] [CrossRef]
- Stanton, R.C. Glucose-6-Phosphate Dehydrogenase, NADPH, and Cell Survival. IUBMB Life 2012, 64, 362–369. [Google Scholar] [CrossRef]
- Zhang, Z.; TeSlaa, T.; Xu, X.; Zeng, X.; Yang, L.; Xing, G.; Tesz, G.J.; Clasquin, M.F.; Rabinowitz, J.D. Serine Catabolism Generates Liver NADPH and Supports Hepatic Lipogenesis. Nat. Metab. 2021, 3, 1608–1620. [Google Scholar] [CrossRef]
- Veech, R.L.; Eggleston, L.V.; Krebs, H.A. The Redox State of Free Nicotinamide–Adenine Dinucleotide Phosphate in the Cytoplasm of Rat Liver. Biochem. J. 1969, 115, 609–619. [Google Scholar] [CrossRef] [PubMed]
- Wasselin, T.; Zahn, S.; Maho, Y.L.; Dorsselaer, A.V.; Raclot, T.; Bertile, F. Exacerbated Oxidative Stress in the Fasting Liver According to Fuel Partitioning. Proteomics 2014, 14, 1905–1921. [Google Scholar] [CrossRef]
- Pawlosky, R.J.; Kemper, M.F.; Kashiwaya, Y.; King, M.T.; Mattson, M.P.; Veech, R.L. Effects of a Dietary Ketone Ester on Hippocampal Glycolytic and TCA Cycle Intermediates and Amino Acids in a 3xTgAD Mouse Model of Alzheimer’s Disease. J. Neurochem. 2017, 141, 195–207. [Google Scholar] [CrossRef] [PubMed]
- Someya, S.; Yu, W.; Hallows, W.C.; Xu, J.; Vann, J.M.; Leeuwenburgh, C.; Tanokura, M.; Denu, J.M.; Prolla, T.A. Sirt3 Mediates Reduction of Oxidative Damage and Prevention of Age-Related Hearing Loss under Caloric Restriction. Cell 2010, 143, 802–812. [Google Scholar] [CrossRef] [PubMed]
- Mary, C.; Soflaee, M.H.; Kesavan, R.; Gelin, M.; Brown, H.; Zacharias, G.; Mathews, T.P.; Lemoff, A.; Lionne, C.; Labesse, G.; et al. Crystal Structure of Human NADK2 Reveals a Dimeric Organization and Active Site Occlusion by Lysine Acetylation. Mol. Cell, 2022, in press. [CrossRef] [PubMed]
- Verma, D.K.; Seo, B.A.; Ghosh, A.; Ma, S.-X.; Hernandez-Quijada, K.; Andersen, J.K.; Ko, H.S.; Kim, Y.-H. Alpha-Synuclein Preformed Fibrils Induce Cellular Senescence in Parkinson’s Disease Models. Cells 2021, 10, 1694. [Google Scholar] [CrossRef] [PubMed]
- Chinta, S.J.; Woods, G.; Demaria, M.; Rane, A.; Zou, Y.; McQuade, A.; Rajagopalan, S.; Limbad, C.; Madden, D.T.; Campisi, J.; et al. Cellular Senescence Is Induced by the Environmental Neurotoxin Paraquat and Contributes to Neuropathology Linked to Parkinson’s Disease. Cell Rep. 2018, 22, 930–940. [Google Scholar] [CrossRef]
- Martini, H.; Passos, J.F. Cellular Senescence: All Roads Lead to Mitochondria. FEBS J. 2022; early view. [Google Scholar] [CrossRef]
- Zhao, Y.; Liu, B.; Xu, L.; Yu, S.; Fu, J.; Wang, J.; Yan, X.; Su, J. ROS-Induced MtDNA Release: The Emerging Messenger for Communication between Neurons and Innate Immune Cells during Neurodegenerative Disorder Progression. Antioxidants 2021, 10, 1917. [Google Scholar] [CrossRef]
- Xian, H.; Watari, K.; Sanchez-Lopez, E.; Offenberger, J.; Onyuru, J.; Sampath, H.; Ying, W.; Hoffman, H.M.; Shadel, G.S.; Karin, M. Oxidized DNA Fragments Exit Mitochondria via MPTP- and VDAC-Dependent Channels to Activate NLRP3 Inflammasome and Interferon Signaling. Immunity, 2022, in press. [CrossRef]
- Salnikov, V.; Lukyánenko, Y.O.; Frederick, C.A.; Lederer, W.J.; Lukyánenko, V. Probing the Outer Mitochondrial Membrane in Cardiac Mitochondria with Nanoparticles. Biophys. J. 2007, 92, 1058–1071. [Google Scholar] [CrossRef]
- Rauckhorst, A.J.; Pfeiffer, D.R.; Broekemeier, K.M. The IPLA2γ Is Identified as the Membrane Potential Sensitive Phospholipase in Liver Mitochondria. FEBS Lett. 2015, 589, 2367–2371. [Google Scholar] [CrossRef] [PubMed]
- Patrushev, M.; Kasymov, V.; Patrusheva, V.; Ushakova, T.; Gogvadze, V.; Gaziev, A. Mitochondrial Permeability Transition Triggers the Release of MtDNA Fragments. CMLS Cell. Mol. Life Sci. 2004, 61, 3100–3103. [Google Scholar] [CrossRef] [PubMed]
- García, N.; García, J.J.; Correa, F.; Chávez, E. The Permeability Transition Pore as a Pathway for the Release of Mitochondrial DNA. Life Sci. 2005, 76, 2873–2880. [Google Scholar] [CrossRef] [PubMed]
- Hou, Y.; Wei, Y.; Lautrup, S.; Yang, B.; Wang, Y.; Cordonnier, S.; Mattson, M.P.; Croteau, D.L.; Bohr, V.A. NAD+ Supplementation Reduces Neuroinflammation and Cell Senescence in a Transgenic Mouse Model of Alzheimer’s Disease via CGAS–STING. Proc. Natl. Acad. Sci. USA 2021, 118, e2011226118. [Google Scholar] [CrossRef]
- Bender, A.; Krishnan, K.J.; Morris, C.M.; Taylor, G.A.; Reeve, A.K.; Perry, R.H.; Jaros, E.; Hersheson, J.S.; Betts, J.; Klopstock, T.; et al. High Levels of Mitochondrial DNA Deletions in Substantia Nigra Neurons in Aging and Parkinson Disease. Nat. Genet. 2006, 38, 515–517. [Google Scholar] [CrossRef] [PubMed]
- Kraytsberg, Y.; Kudryavtseva, E.; McKee, A.C.; Geula, C.; Kowall, N.W.; Khrapko, K. Mitochondrial DNA Deletions Are Abundant and Cause Functional Impairment in Aged Human Substantia Nigra Neurons. Nat. Genet. 2006, 38, 518–520. [Google Scholar] [CrossRef] [PubMed]
- Dölle, C.; Flønes, I.; Nido, G.S.; Miletic, H.; Osuagwu, N.; Kristoffersen, S.; Lilleng, P.K.; Larsen, J.P.; Tysnes, O.-B.; Haugarvoll, K.; et al. Defective Mitochondrial DNA Homeostasis in the Substantia Nigra in Parkinson Disease. Nat. Commun. 2016, 7, 13548. [Google Scholar] [CrossRef]
- Manini, A.; Abati, E.; Comi, G.P.; Corti, S.; Ronchi, D. Mitochondrial DNA Homeostasis Impairment and Dopaminergic Dysfunction: A Trembling Balance. Ageing Res. Rev. 2022, 76, 101578. [Google Scholar] [CrossRef]
- Wiley, C.D.; Velarde, M.C.; Lecot, P.; Liu, S.; Sarnoski, E.A.; Freund, A.; Shirakawa, K.; Lim, H.W.; Davis, S.S.; Ramanathan, A.; et al. Mitochondrial Dysfunction Induces Senescence with a Distinct Secretory Phenotype. Cell Metab. 2016, 23, 303–314. [Google Scholar] [CrossRef]
- Kujoth, G.C.; Hiona, A.; Pugh, T.D.; Someya, S.; Panzer, K.; Wohlgemuth, S.E.; Hofer, T.; Seo, A.Y.; Sullivan, R.; Jobling, W.A.; et al. Mitochondrial DNA Mutations, Oxidative Stress, and Apoptosis in Mammalian Aging. Science 2005, 309, 481–484. [Google Scholar] [CrossRef]
- Tyynismaa, H.; Mjosund, K.P.; Wanrooij, S.; Lappalainen, I.; Ylikallio, E.; Jalanko, A.; Spelbrink, J.N.; Paetau, A.; Suomalainen, A. Mutant Mitochondrial Helicase Twinkle Causes Multiple MtDNA Deletions and a Late-Onset Mitochondrial Disease in Mice. Proc. Natl. Acad. Sci. USA 2005, 102, 17687–17692. [Google Scholar] [CrossRef] [PubMed]
- Vermulst, M.; Wanagat, J.; Kujoth, G.C.; Bielas, J.H.; Rabinovitch, P.S.; Prolla, T.A.; Loeb, L.A. DNA Deletions and Clonal Mutations Drive Premature Aging in Mitochondrial Mutator Mice. Nat. Genet. 2008, 40, 392–394. [Google Scholar] [CrossRef] [PubMed]
- Someya, S.; Kujoth, G.C.; Kim, M.-J.; Hacker, T.A.; Vermulst, M.; Weindruch, R.; Prolla, T.A. Effects of Calorie Restriction on the Lifespan and Healthspan of POLG Mitochondrial Mutator Mice. PLoS ONE 2017, 12, e0171159. [Google Scholar] [CrossRef]
- Colman, R.J.; Beasley, T.M.; Kemnitz, J.W.; Johnson, S.C.; Weindruch, R.; Anderson, R.M. Caloric Restriction Reduces Age-Related and All-Cause Mortality in Rhesus Monkeys. Nat. Commun. 2014, 5, 3557. [Google Scholar] [CrossRef] [PubMed]
- McKiernan, S.H.; Colman, R.J.; Aiken, E.; Evans, T.D.; Beasley, T.M.; Aiken, J.M.; Weindruch, R.; Anderson, R.M. Cellular Adaptation Contributes to Calorie Restriction-Induced Preservation of Skeletal Muscle in Aged Rhesus Monkeys. Exp. Gerontol. 2012, 47, 229–236. [Google Scholar] [CrossRef]
- Finley, L.W.S.; Lee, J.; Souza, A.; Desquiret-Dumas, V.; Bullock, K.; Rowe, G.C.; Procaccio, V.; Clish, C.B.; Arany, Z.; Haigis, M.C. Skeletal Muscle Transcriptional Coactivator PGC-1α Mediates Mitochondrial, but Not Metabolic, Changes during Calorie Restriction. Proc. Natl. Acad. Sci. USA 2012, 109, 2931–2936. [Google Scholar] [CrossRef]
- Guo, X.; Kudryavtseva, E.; Bodyak, N.; Nicholas, A.; Dombrovsky, I.; Yang, D.; Kraytsberg, Y.; Simon, D.K.; Khrapko, K. Mitochondrial DNA Deletions in Mice in Men: Substantia Nigra Is Much Less Affected in the Mouse. Biochim. Biophys. Acta 2010, 1797, 1159–1162. [Google Scholar] [CrossRef]
- Dai, Y.; Kiselak, T.; Clark, J.; Clore, E.; Zheng, K.; Cheng, A.; Kujoth, G.C.; Prolla, T.A.; Maratos-Flier, E.; Simon, D.K. Behavioral and Metabolic Characterization of Heterozygous and Homozygous POLG Mutator Mice. Mitochondrion 2013, 13, 282–291. [Google Scholar] [CrossRef]
- Green, C.L.; Lamming, D.W.; Fontana, L. Molecular Mechanisms of Dietary Restriction Promoting Health and Longevity. Nat. Rev. Mol. Cell Biol. 2022, 23, 56–73. [Google Scholar] [CrossRef]
- Mitchell, S.J.; Bernier, M.; Mattison, J.A.; Aon, M.A.; Kaiser, T.A.; Anson, R.M.; Ikeno, Y.; Anderson, R.M.; Ingram, D.K.; de Cabo, R. Daily Fasting Improves Health and Survival in Male Mice Independent of Diet Composition and Calories. Cell Metab. 2019, 29, 221–228.e3. [Google Scholar] [CrossRef]
- Shimokawa, I.; Komatsu, T.; Hayashi, N.; Kim, S.-E.; Kawata, T.; Park, S.; Hayashi, H.; Yamaza, H.; Chiba, T.; Mori, R. The Life-Extending Effect of Dietary Restriction Requires Foxo3 in Mice. Aging Cell 2015, 14, 707–709. [Google Scholar] [CrossRef] [PubMed]
- Edwards, C.; Canfield, J.; Copes, N.; Rehan, M.; Lipps, D.; Bradshaw, P.C. D-Beta-Hydroxybutyrate Extends Lifespan in C. elegans. Aging 2014, 6, 621–644. [Google Scholar] [CrossRef] [PubMed]
- Mattson, M.P.; Moehl, K.; Ghena, N.; Schmaedick, M.; Cheng, A. Intermittent Metabolic Switching, Neuroplasticity and Brain Health. Nat. Rev. Neurosci. 2018, 19, 81–94. [Google Scholar] [CrossRef] [PubMed]
- Hvingelby, V.S.; Glud, A.N.; Sørensen, J.C.H.; Tai, Y.; Andersen, A.S.M.; Johnsen, E.; Moro, E.; Pavese, N. Interventions to Improve Gait in Parkinson’s Disease: A Systematic Review of Randomized Controlled Trials and Network Meta-Analysis. J. Neurol. 2022, 269, 4068–4079. [Google Scholar] [CrossRef] [PubMed]
- Uhrbrand, A.; Stenager, E.; Pedersen, M.S.; Dalgas, U. Parkinson’s Disease and Intensive Exercise Therapy—A Systematic Review and Meta-Analysis of Randomized Controlled Trials. J. Neurol. Sci. 2015, 353, 9–19. [Google Scholar] [CrossRef]
- Stillman, C.M.; Cohen, J.; Lehman, M.E.; Erickson, K.I. Mediators of Physical Activity on Neurocognitive Function: A Review at Multiple Levels of Analysis. Front. Hum. Neurosci. 2016, 10, 626. [Google Scholar] [CrossRef]
- Petzinger, G.M.; Walsh, J.P.; Akopian, G.; Hogg, E.; Abernathy, A.; Arevalo, P.; Turnquist, P.; Vučković, M.; Fisher, B.E.; Togasaki, D.M.; et al. Effects of Treadmill Exercise on Dopaminergic Transmission in the 1-Methyl-4-Phenyl-1,2,3,6-Tetrahydropyridine-Lesioned Mouse Model of Basal Ganglia Injury. J. Neurosci. 2007, 27, 5291–5300. [Google Scholar] [CrossRef]
- Stranahan, A.M.; Lee, K.; Martin, B.; Maudsley, S.; Golden, E.; Cutler, R.G.; Mattson, M.P. Voluntary Exercise and Caloric Restriction Enhance Hippocampal Dendritic Spine Density and BDNF Levels in Diabetic Mice. Hippocampus 2009, 19, 951–961. [Google Scholar] [CrossRef]
- Lee, J.; Duan, W.; Mattson, M.P. Evidence That Brain-Derived Neurotrophic Factor Is Required for Basal Neurogenesis and Mediates, in Part, the Enhancement of Neurogenesis by Dietary Restriction in the Hippocampus of Adult Mice. J. Neurochem. 2002, 82, 1367–1375. [Google Scholar] [CrossRef]
- Vivar, C.; Potter, M.C.; Choi, J.; Lee, J.; Stringer, T.P.; Callaway, E.M.; Gage, F.H.; Suh, H.; van Praag, H. Monosynaptic Inputs to New Neurons in the Dentate Gyrus. Nat. Commun. 2012, 3, 1107. [Google Scholar] [CrossRef]
- Estrada, N.M.; Isokawa, M. Metabolic Demand Stimulates CREB Signaling in the Limbic Cortex: Implication for the Induction of Hippocampal Synaptic Plasticity by Intrinsic Stimulus for Survival. Front. Syst. Neurosci. 2009, 3, 5. [Google Scholar] [CrossRef]
- Yang, J.-L.; Lin, Y.-T.; Chuang, P.-C.; Bohr, V.A.; Mattson, M.P. BDNF and Exercise Enhance Neuronal DNA Repair by Stimulating CREB-Mediated Production of Apurinic/Apyrimidinic Endonuclease 1. Neuromol. Med. 2014, 16, 161–174. [Google Scholar] [CrossRef] [PubMed]
- Marosi, K.; Mattson, M.P. BDNF Mediates Adaptive Brain and Body Responses to Energetic Challenges. Trends Endocrinol. Metab. TEM 2014, 25, 89–98. [Google Scholar] [CrossRef] [PubMed]
- Marosi, K.; Kim, S.W.; Moehl, K.; Scheibye-Knudsen, M.; Cheng, A.; Cutler, R.; Camandola, S.; Mattson, M.P. 3-Hydroxybutyrate Regulates Energy Metabolism and Induces BDNF Expression in Cerebral Cortical Neurons. J. Neurochem. 2016, 139, 769–781. [Google Scholar] [CrossRef] [PubMed]
- Sleiman, S.F.; Henry, J.; Al-Haddad, R.; El Hayek, L.; Abou Haidar, E.; Stringer, T.; Ulja, D.; Karuppagounder, S.S.; Holson, E.B.; Ratan, R.R.; et al. Exercise Promotes the Expression of Brain Derived Neurotrophic Factor (BDNF) through the Action of the Ketone Body β-Hydroxybutyrate. eLife 2016, 5, e15092. [Google Scholar] [CrossRef]
- Hood, D.A.; Tryon, L.D.; Carter, H.N.; Kim, Y.; Chen, C.C.W. Unravelling the Mechanisms Regulating Muscle Mitochondrial Biogenesis. Biochem. J. 2016, 473, 2295–2314. [Google Scholar] [CrossRef]
- Di Benedetto, S.; Müller, L.; Wenger, E.; Düzel, S.; Pawelec, G. Contribution of Neuroinflammation and Immunity to Brain Aging and the Mitigating Effects of Physical and Cognitive Interventions. Neurosci. Biobehav. Rev. 2017, 75, 114–128. [Google Scholar] [CrossRef]
- Simeone, T.A.; Simeone, K.A.; Rho, J.M. Ketone Bodies as Anti-Seizure Agents. Neurochem. Res. 2017, 42, 2011–2018. [Google Scholar] [CrossRef]
- Ngandu, T.; Lehtisalo, J.; Solomon, A.; Levälahti, E.; Ahtiluoto, S.; Antikainen, R.; Bäckman, L.; Hänninen, T.; Jula, A.; Laatikainen, T.; et al. A 2 Year Multidomain Intervention of Diet, Exercise, Cognitive Training, and Vascular Risk Monitoring versus Control to Prevent Cognitive Decline in at-Risk Elderly People (FINGER): A Randomised Controlled Trial. Lancet 2015, 385, 2255–2263. [Google Scholar] [CrossRef]
- Grammatikopoulou, M.G.; Tousinas, G.; Balodimou, C.; Anastasilakis, D.A.; Gkiouras, K.; Dardiotis, E.; Evangeliou, A.E.; Bogdanos, D.P.; Goulis, D.G. Ketogenic Therapy for Parkinson’s Disease: A Systematic Review and Synthesis without Meta-Analysis of Animal and Human Trials. Maturitas 2022, 163, 46–61. [Google Scholar] [CrossRef]
- Choi, A.; Hallett, M.; Ehrlich, D. Nutritional Ketosis in Parkinson’s Disease—A Review of Remaining Questions and Insights. Neurotherapeutics 2021, 18, 1637–1649. [Google Scholar] [CrossRef] [PubMed]
- Kwon, J.H.; Moon, K.M.; Min, K.-W. Exercise-Induced Myokines Can Explain the Importance of Physical Activity in the Elderly: An Overview. Healthcare 2020, 8, 378. [Google Scholar] [CrossRef] [PubMed]
- Hao, Z.; Zhang, X.; Chen, P. Effects of Ten Different Exercise Interventions on Motor Function in Parkinson’s Disease Patients—A Network Meta-Analysis of Randomized Controlled Trials. Brain Sci. 2022, 12, 698. [Google Scholar] [CrossRef]
- Bindoff, L.A.; Birch-Machin, M.A.; Cartlidge, N.E.F.; Parker, W.D.; Turnbull, D.M. Respiratory Chain Abnormalities in Skeletal Muscle from Patients with Parkinson’s Disease. J. Neurol. Sci. 1991, 104, 203–208. [Google Scholar] [CrossRef]
- Blin, O.; Desnuelle, C.; Rascol, O.; Borg, M.; Peyro Saint Paul, H.; Azulay, J.P.; Billé, F.; Figarella, D.; Coulom, F.; Pellissier, J.F. Mitochondrial Respiratory Failure in Skeletal Muscle from Patients with Parkinson’s Disease and Multiple System Atrophy. J. Neurol. Sci. 1994, 125, 95–101. [Google Scholar] [CrossRef]
- Cardellach, F.; Martí, M.J.; Fernández-Solá, J.; Marín, C.; Hoek, J.B.; Tolosa, E.; Urbano-Márquez, A. Mitochondrial Respiratory Chain Activity in Skeletal Muscle from Patients with Parkinson’s Disease. Neurology 1993, 43, 2258–2262. [Google Scholar] [CrossRef]
- Mann, V.M.; Cooper, J.M.; Kridge, D.; Daniel, S.E.; Schapira, A.H.V.; Marsden, C.D. Brain, Skeletal Muscle and Platelet Homogenate Mitochondrial Function in Parkinson’s Disease. Brain 1992, 115, 333–342. [Google Scholar] [CrossRef]
- Harper, C.; Gopalan, V.; Goh, J. Exercise Rescues Mitochondrial Coupling in Aged Skeletal Muscle: A Comparison of Different Modalities in Preventing Sarcopenia. J. Transl. Med. 2021, 19, 71. [Google Scholar] [CrossRef]
- Erlich, A.T.; Brownlee, D.M.; Beyfuss, K.; Hood, D.A. Exercise Induces TFEB Expression and Activity in Skeletal Muscle in a PGC-1α-Dependent Manner. Am. J. Physiol. Cell Physiol. 2018, 314, C62–C72. [Google Scholar] [CrossRef]
- Gendi, F.; Pei, F.; Wang, Y.; Li, H.; Fu, J.; Chang, C. Mitochondrial Proteins Unveil the Mechanism by Which Physical Exercise Ameliorates Memory, Learning and Motor Activity in Hypoxic Ischemic Encephalopathy Rat Model. Int. J. Mol. Sci. 2022, 23, 4235. [Google Scholar] [CrossRef]
- Mesquita, P.H.C.; Lamb, D.A.; Parry, H.A.; Moore, J.H.; Smith, M.A.; Vann, C.G.; Osburn, S.C.; Fox, C.D.; Ruple, B.A.; Huggins, K.W.; et al. Acute and Chronic Effects of Resistance Training on Skeletal Muscle Markers of Mitochondrial Remodeling in Older Adults. Physiol. Rep. 2020, 8, e14526. [Google Scholar] [CrossRef] [PubMed]
- Lamb, D.A.; Moore, J.H.; Mesquita, P.H.C.; Smith, M.A.; Vann, C.G.; Osburn, S.C.; Fox, C.D.; Lopez, H.L.; Ziegenfuss, T.N.; Huggins, K.W.; et al. Resistance Training Increases Muscle NAD+ and NADH Concentrations as Well as NAMPT Protein Levels and Global Sirtuin Activity in Middle-Aged, Overweight, Untrained Individuals. Aging 2020, 12, 9447–9460. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Mascher, H.; Psilander, N.; Blomstrand, E.; Sahlin, K. Resistance Exercise Enhances the Molecular Signaling of Mitochondrial Biogenesis Induced by Endurance Exercise in Human Skeletal Muscle. J. Appl. Physiol. 2011, 111, 1335–1344. [Google Scholar] [CrossRef] [PubMed]
- Gibala, M.J.; Little, J.P.; MacDonald, M.J.; Hawley, J.A. Physiological Adaptations to Low-Volume, High-Intensity Interval Training in Health and Disease. J. Physiol. 2012, 590, 1077–1084. [Google Scholar] [CrossRef]
- Wahl, P.; Bloch, W.; Proschinger, S. The Molecular Signature of High-Intensity Training in the Human Body. Int. J. Sports Med. 2022, 43, 195–205. [Google Scholar] [CrossRef]
- Ma, J.K.; Scribbans, T.D.; Edgett, B.A.; Boyd, J.C.; Simpson, C.A.; Little, J.P.; Gurd, B.J. Extremely Low-Volume, High-Intensity Interval Training Improves Exercise Capacity and Increases Mitochondrial Protein Content in Human Skeletal Muscle. Open J. Mol. Integr. Physiol. 2013, 3, 202–210. [Google Scholar] [CrossRef]
- Santiago, J.A.; Quinn, J.P.; Potashkin, J.A. Physical Activity Rewires the Human Brain against Neurodegeneration. Int. J. Mol. Sci. 2022, 23, 6223. [Google Scholar] [CrossRef]
- Palma, J.-A.; Kaufmann, H. Treatment of Autonomic Dysfunction in Parkinson Disease and Other Synucleinopathies. Mov. Disord. 2018, 33, 372–390. [Google Scholar] [CrossRef]
- Pfeiffer, R.F. Autonomic Dysfunction in Parkinson’s Disease. Neurotherapeutics 2020, 17, 1464–1479. [Google Scholar] [CrossRef]
- Nalls, M.A.; McLean, C.Y.; Rick, J.; Eberly, S.; Hutten, S.J.; Gwinn, K.; Sutherland, M.; Martinez, M.; Heutink, P.; Williams, N.; et al. Diagnosis of Parkinson’s Disease on the Basis of Clinical–Genetic Classification: A Population-Based Modelling Study. Lancet Neurol. 2015, 14, 1002–1009. [Google Scholar] [CrossRef]
- De Abreu, R.M.; Rehder-Santos, P.; Simões, R.P.; Catai, A.M. Can High-Intensity Interval Training Change Cardiac Autonomic Control? A Systematic Review. Braz. J. Phys. Ther. 2019, 23, 279–289. [Google Scholar] [CrossRef] [PubMed]
- Harvey, M.; Weston, K.L.; Gray, W.K.; O’Callaghan, A.; Oates, L.L.; Davidson, R.; Walker, R.W. High-Intensity Interval Training in People with Parkinson’s Disease: A Randomized, Controlled Feasibility Trial. Clin. Rehabil. 2019, 33, 428–438. [Google Scholar] [CrossRef] [PubMed]
- Sato, S.; Dyar, K.A.; Treebak, J.T.; Jepsen, S.L.; Ehrlich, A.M.; Ashcroft, S.P.; Trost, K.; Kunzke, T.; Prade, V.M.; Small, L.; et al. Atlas of Exercise Metabolism Reveals Time-Dependent Signatures of Metabolic Homeostasis. Cell Metab. 2022, 34, 329–345.e8. [Google Scholar] [CrossRef] [PubMed]
- Williamson, D.H.; Lund, P.; Krebs, H.A. The Redox State of Free Nicotinamide-Adenine Dinucleotide in the Cytoplasm and Mitochondria of Rat Liver. Biochem. J. 1967, 103, 514–527. [Google Scholar] [CrossRef]
- Goodman, R.P.; Markhard, A.L.; Shah, H.; Sharma, R.; Skinner, O.S.; Clish, C.B.; Deik, A.; Patgiri, A.; Hsu, Y.-H.; Masia, R.; et al. Hepatic NADH Reductive Stress Underlies Common Variation in Metabolic Traits. Nature 2020, 583, 122–126. [Google Scholar] [CrossRef]
- Naghizadeh, F. Oxidation of Alpha-Hydroxybutyrate by Human Serum. Clin. Chim. Acta 1977, 81, 277–282. [Google Scholar] [CrossRef]
- Lu, S.C. Glutathione Synthesis. Biochim. Biophys. Acta Gen. Subj. 2013, 1830, 3143–3153. [Google Scholar] [CrossRef]
- Hine, C.; Harputlugil, E.; Zhang, Y.; Ruckenstuhl, C.; Lee, B.C.; Brace, L.; Longchamp, A.; Trevino-Villarreal, J.H.; Mejia, P.; Ozaki, C.K.; et al. Endogenous Hydrogen Sulfide Production Is Essential for Dietary Restriction Benefits. Cell 2015, 160, 132–144. [Google Scholar] [CrossRef]
- Acosta-Rodríguez, V.; Rijo-Ferreira, F.; Izumo, M.; Xu, P.; Wight-Carter, M.; Green, C.B.; Takahashi, J.S. Circadian Alignment of Early Onset Caloric Restriction Promotes Longevity in Male C57BL/6J Mice. Science 2022, 376, 1192–1202. [Google Scholar] [CrossRef]
- Peluso, A.; Damgaard, M.V.; Mori, M.A.S.; Treebak, J.T. Age-Dependent Decline of NAD+—Universal Truth or Confounded Consensus? Nutrients 2021, 14, 101. [Google Scholar] [CrossRef]
- Conlon, N.; Ford, D. A Systems-Approach to NAD+ Restoration. Biochem. Pharmacol. 2022, 198, 114946. [Google Scholar] [CrossRef] [PubMed]
- Zeidler, J.D.; Hogan, K.A.; Agorrody, G.; Peclat, T.R.; Kashyap, S.; Kanamori, K.S.; Gomez, L.S.; Mazdeh, D.Z.; Warner, G.M.; Thompson, K.L.; et al. The CD38 Glycohydrolase and the NAD Sink: Implications for Pathological Conditions. Am. J. Physiol. Cell Physiol. 2022, 322, C521–C545. [Google Scholar] [CrossRef] [PubMed]
- Strømland, Ø.; Diab, J.; Ferrario, E.; Sverkeli, L.J.; Ziegler, M. The Balance between NAD+ Biosynthesis and Consumption in Ageing. Mech. Ageing Dev. 2021, 199, 111569. [Google Scholar] [CrossRef] [PubMed]
- Reiten, O.K.; Wilvang, M.A.; Mitchell, S.J.; Hu, Z.; Fang, E.F. Preclinical and Clinical Evidence of NAD+ Precursors in Health, Disease, and Ageing. Mech. Ageing Dev. 2021, 199, 111567. [Google Scholar] [CrossRef] [PubMed]
- Magni, G.; Orsomando, G.; Raffaelli, N. Structural and Functional Properties of NAD Kinase, a Key Enzyme in NADP Biosynthesis. MRMC 2006, 6, 739–746. [Google Scholar] [CrossRef]
- Du, J.; Estrella, M.; Solorio-Kirpichyan, K.; Jeffrey, P.D.; Korennykh, A. Structure of Human NADK2 Reveals Atypical Assembly and Regulation of NAD Kinases from Animal Mitochondria. Proc. Natl. Acad. Sci. USA 2022, 119, e2200923119. [Google Scholar] [CrossRef]
- Murata, K. Polyphosphate-Dependent Nicotinamide Adenine Dinucleotide (NAD) Kinase: A Novel Missing Link in Human Mitochondria. Proc. Jpn Acad. Ser. B Phys. Biol. Sci. 2021, 97, 479–498. [Google Scholar] [CrossRef]
- Veech, R.L.; Todd King, M.; Pawlosky, R.; Kashiwaya, Y.; Bradshaw, P.C.; Curtis, W. The “Great” Controlling Nucleotide Coenzymes. IUBMB Life 2019, 71, 565–579. [Google Scholar] [CrossRef]
- Pollak, N.; Niere, M.; Ziegler, M. NAD Kinase Levels Control the NADPH Concentration in Human Cells. J. Biol. Chem. 2007, 282, 33562–33571. [Google Scholar] [CrossRef]
- Shats, I.; Williams, J.G.; Liu, J.; Makarov, M.V.; Wu, X.; Lih, F.B.; Deterding, L.J.; Lim, C.; Xu, X.; Randall, T.A.; et al. Bacteria Boost Mammalian Host NAD Metabolism by Engaging the Deamidated Biosynthesis Pathway. Cell Metab. 2020, 31, 564–579.e7. [Google Scholar] [CrossRef]
- Covarrubias, A.J.; Perrone, R.; Grozio, A.; Verdin, E. NAD+ Metabolism and Its Roles in Cellular Processes during Ageing. Nat. Rev. Mol. Cell. Biol. 2021, 22, 119–141. [Google Scholar] [CrossRef]
- Liu, L.; Su, X.; Quinn, W.J.; Hui, S.; Krukenberg, K.; Frederick, D.W.; Redpath, P.; Zhan, L.; Chellappa, K.; White, E.; et al. Quantitative Analysis of NAD Synthesis-Breakdown Fluxes. Cell Metab. 2018, 27, 1067–1080.e5. [Google Scholar] [CrossRef]
- Brakedal, B.; Dölle, C.; Riemer, F.; Ma, Y.; Nido, G.S.; Skeie, G.O.; Craven, A.R.; Schwarzlmüller, T.; Brekke, N.; Diab, J.; et al. The NADPARK Study: A Randomized Phase I Trial of Nicotinamide Riboside Supplementation in Parkinson’s Disease. Cell Metab. 2022, 34, 396–407.e6. [Google Scholar] [CrossRef] [PubMed]
- Perrin, L.; Loizides-Mangold, U.; Chanon, S.; Gobet, C.; Hulo, N.; Isenegger, L.; Weger, B.D.; Migliavacca, E.; Charpagne, A.; Betts, J.A.; et al. Transcriptomic Analyses Reveal Rhythmic and CLOCK-Driven Pathways in Human Skeletal Muscle. eLife 2018, 7, e34114. [Google Scholar] [CrossRef] [PubMed]
- Schug, T.T.; Li, X. Sirtuin 1 in Lipid Metabolism and Obesity. Ann. Med. 2011, 43, 198–211. [Google Scholar] [CrossRef] [PubMed]
- Luna, A.; McFadden, G.B.; Aladjem, M.I.; Kohn, K.W. Predicted Role of NAD Utilization in the Control of Circadian Rhythms during DNA Damage Response. PLoS Comput. Biol. 2015, 11, e1004144. [Google Scholar] [CrossRef] [PubMed]
- Ramsey, K.M.; Yoshino, J.; Brace, C.S.; Abrassart, D.; Kobayashi, Y.; Marcheva, B.; Hong, H.-K.; Chong, J.L.; Buhr, E.D.; Lee, C.; et al. Circadian Clock Feedback Cycle Through NAMPT-Mediated NAD+ Biosynthesis. Science 2009, 324, 651–654. [Google Scholar] [CrossRef]
- van Moorsel, D.; Hansen, J.; Havekes, B.; Scheer, F.A.J.L.; Jörgensen, J.A.; Hoeks, J.; Schrauwen-Hinderling, V.B.; Duez, H.; Lefebvre, P.; Schaper, N.C.; et al. Demonstration of a Day-Night Rhythm in Human Skeletal Muscle Oxidative Capacity. Mol. Metab. 2016, 5, 635–645. [Google Scholar] [CrossRef]
- Qiu, Z.; Ming, H.; Lei, S.; Zhou, B.; Zhao, B.; Yu, Y.; Xue, R.; Xia, Z. Roles of HDAC3-Orchestrated Circadian Clock Gene Oscillations in Diabetic Rats Following Myocardial Ischaemia/Reperfusion Injury. Cell Death Dis. 2021, 12, 43. [Google Scholar] [CrossRef]
- Goodman, R.P.; Calvo, S.E.; Mootha, V.K. Spatiotemporal Compartmentalization of Hepatic NADH and NADPH Metabolism. J. Biol. Chem. 2018, 293, 7508–7516. [Google Scholar] [CrossRef]
- Vallée, A.; Lecarpentier, Y.; Guillevin, R.; Vallée, J.-N. Circadian Rhythms, Neuroinflammation and Oxidative Stress in the Story of Parkinson’s Disease. Cells 2020, 9, 314. [Google Scholar] [CrossRef] [PubMed]
- Hunt, J.; Coulson, E.J.; Rajnarayanan, R.; Oster, H.; Videnovic, A.; Rawashdeh, O. Sleep and Circadian Rhythms in Parkinson’s Disease and Preclinical Models. Mol. Neurodegener. 2022, 17, 2. [Google Scholar] [CrossRef] [PubMed]
- Prudon, B.; Duncan, G.W.; Khoo, T.K.; Yarnall, A.J.; Burn, D.J.; Anderson, K.N. Primary Sleep Disorder Prevalence in Patients with Newly Diagnosed Parkinson’s Disease. Mov. Disord. 2014, 29, 259–262. [Google Scholar] [CrossRef] [PubMed]
- Korshunov, K.S.; Blakemore, L.J.; Trombley, P.Q. Dopamine: A Modulator of Circadian Rhythms in the Central Nervous System. Front. Cell. Neurosci. 2017, 11, 91. [Google Scholar] [CrossRef] [PubMed]
- Mendoza, J.; Challet, E. Circadian Insights into Dopamine Mechanisms. Neuroscience 2014, 282, 230–242. [Google Scholar] [CrossRef]
- Hood, S.; Cassidy, P.; Cossette, M.-P.; Weigl, Y.; Verwey, M.; Robinson, B.; Stewart, J.; Amir, S. Endogenous Dopamine Regulates the Rhythm of Expression of the Clock Protein PER2 in the Rat Dorsal Striatum via Daily Activation of D2 Dopamine Receptors. J. Neurosci. 2010, 30, 14046–14058. [Google Scholar] [CrossRef]
- Yujnovsky, I.; Hirayama, J.; Doi, M.; Borrelli, E.; Sassone-Corsi, P. Signaling Mediated by the Dopamine D2 Receptor Potentiates Circadian Regulation by CLOCK:BMAL1. Proc. Natl. Acad. Sci. USA 2006, 103, 6386–6391. [Google Scholar] [CrossRef]
- Imbesi, M.; Yildiz, S.; Arslan, A.D.; Sharma, R.; Manev, H.; Uz, T. Dopamine Receptor-Mediated Regulation of Neuronal “Clock” Gene Expression. Neuroscience 2009, 158, 537–544. [Google Scholar] [CrossRef]
- Parekh, P.K.; Ozburn, A.R.; McClung, C.A. Circadian Clock Genes: Effects on Dopamine, Reward and Addiction. Alcohol 2015, 49, 341–349. [Google Scholar] [CrossRef]
- Breen, D.P.; Vuono, R.; Nawarathna, U.; Fisher, K.; Shneerson, J.M.; Reddy, A.B.; Barker, R.A. Sleep and Circadian Rhythm Regulation in Early Parkinson Disease. JAMA Neurol. 2014, 71, 589–595. [Google Scholar] [CrossRef]
- Cai, Y.; Liu, S.; Sothern, R.B.; Xu, S.; Chan, P. Expression of Clock Genes Per1 and Bmal1 in Total Leukocytes in Health and Parkinson’s Disease: Clock Genes in PD. Eur. J. Neurol. 2010, 17, 550–554. [Google Scholar] [CrossRef]
- Li, S.-Y.; Wang, Y.-L.; Liu, W.-W.; Lyu, D.-J.; Wang, F.; Mao, C.-J.; Yang, Y.-P.; Hu, L.-F.; Liu, C.-F. Long-Term Levodopa Treatment Accelerates the Circadian Rhythm Dysfunction in a 6-Hydroxydopamine Rat Model of Parkinson’s Disease. Chin. Med. J. 2017, 130, 1085–1092. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Song, S.; Wang, Y.; Huang, C.; Zhang, F.; Liu, J.; Hong, J.-S. Low-Grade Inflammation Aggravates Rotenone Neurotoxicity and Disrupts Circadian Clock Gene Expression in Rats. Neurotox. Res. 2019, 35, 421–431. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Wei, S.; Huang, G.; Liu, L.; Gu, C.; Shen, Y.; Wang, X.; Xia, S.; Xie, A.; Hu, L.; et al. BMAL1 Regulation of Microglia-mediated Neuroinflammation in MPTP-induced Parkinson’s Disease Mouse Model. FASEB J. 2020, 34, 6570–6581. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Jang, S.; Choi, M.; Chung, S.; Choe, Y.; Choe, H.K.; Son, G.H.; Rhee, K.; Kim, K. Abrogation of the Circadian Nuclear Receptor REV-ERBα Exacerbates 6-Hydroxydopamine-Induced Dopaminergic Neurodegeneration. Mol. Cells 2018, 41, 742–752. [Google Scholar] [CrossRef] [PubMed]
- Shkodina, A.D.; Tan, S.C.; Hasan, M.M.; Abdelgawad, M.; Chopra, H.; Bilal, M.; Boiko, D.I.; Tarianyk, K.A.; Alexiou, A. Roles of Clock Genes in the Pathogenesis of Parkinson’s Disease. Ageing Res. Rev. 2022, 74, 101554. [Google Scholar] [CrossRef]
- de Goede, P.; Wefers, J.; Brombacher, E.C.; Schrauwen, P.; Kalsbeek, A. Circadian Rhythms in Mitochondrial Respiration. J. Mol. Endocrinol. 2018, 60, R115–R130. [Google Scholar] [CrossRef]
- Jacobi, D.; Liu, S.; Burkewitz, K.; Kory, N.; Knudsen, N.H.; Alexander, R.K.; Unluturk, U.; Li, X.; Kong, X.; Hyde, A.; et al. Hepatic Bmal1 Regulates Rhythmic Mitochondrial Dynamics and Promotes Metabolic Fitness. Cell Metab. 2015, 22, 709–720. [Google Scholar] [CrossRef]
- Schmitt, K.; Grimm, A.; Dallmann, R.; Oettinghaus, B.; Restelli, L.M.; Witzig, M.; Ishihara, N.; Mihara, K.; Ripperger, J.A.; Albrecht, U.; et al. Circadian Control of DRP1 Activity Regulates Mitochondrial Dynamics and Bioenergetics. Cell Metab. 2018, 27, 657–666.e5. [Google Scholar] [CrossRef]
- Rabinovich-Nikitin, I.; Rasouli, M.; Reitz, C.J.; Posen, I.; Margulets, V.; Dhingra, R.; Khatua, T.N.; Thliveris, J.A.; Martino, T.A.; Kirshenbaum, L.A. Mitochondrial Autophagy and Cell Survival Is Regulated by the Circadian Clock Gene in Cardiac Myocytes during Ischemic Stress. Autophagy 2021, 17, 3794–3812. [Google Scholar] [CrossRef]
- Galmozzi, A.; Mitro, N.; Ferrari, A.; Gers, E.; Gilardi, F.; Godio, C.; Cermenati, G.; Gualerzi, A.; Donetti, E.; Rotili, D.; et al. Inhibition of Class I Histone Deacetylases Unveils a Mitochondrial Signature and Enhances Oxidative Metabolism in Skeletal Muscle and Adipose Tissue. Diabetes 2013, 62, 732–742. [Google Scholar] [CrossRef] [PubMed]
- Sengupta, S.; Yang, G.; O’Donnell, J.C.; Hinson, M.D.; McCormack, S.E.; Falk, M.J.; La, P.; Robinson, M.B.; Williams, M.L.; Yohannes, M.T.; et al. The Circadian Gene Rev-Erbα Improves Cellular Bioenergetics and Provides Preconditioning for Protection against Oxidative Stress. Free Radic. Biol. Med. 2016, 93, 177–189. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Newman, J.C.; Verdin, E. β-Hydroxybutyrate. Annu. Rev. Nutr. 2017, 37, 51–76. [Google Scholar] [CrossRef] [PubMed]
- Shen, R.-S.; Abell, C.W.; Gessner, W.; Brossi, A. Serotonergic Conversion of MPTP and Dopaminergic Accumulation of MPP+. FEBS Lett. 1985, 189, 225–230. [Google Scholar] [CrossRef]
- Ramsay, R.R.; Dadgar, J.; Trevor, A.; Singer, T.P. Energy-Driven Uptake of N-Methyl-4-Phenylpyridine by Brain Mitochondria Mediates the Neurotoxicity of MPTP. Life Sci. 1986, 39, 581–588. [Google Scholar] [CrossRef]
- Devi, L.; Raghavendran, V.; Prabhu, B.M.; Avadhani, N.G.; Anandatheerthavarada, H.K. Mitochondrial Import and Accumulation of α-Synuclein Impair Complex I in Human Dopaminergic Neuronal Cultures and Parkinson Disease Brain. J. Biol. Chem. 2008, 283, 9089–9100. [Google Scholar] [CrossRef]
- Cassarino, D.S.; Parks, J.K.; Parker, W.D.; Bennett, J.P. The Parkinsonian Neurotoxin MPP+ Opens the Mitochondrial Permeability Transition Pore and Releases Cytochrome c in Isolated Mitochondria via an Oxidative Mechanism. Biochim. Biophys. Acta Mol. Basis Dis. 1999, 1453, 49–62. [Google Scholar] [CrossRef]
- Klaidman, L.K.; Adams, J.D.; Leung, A.C.; Sam Kim, S.; Cadenas, E. Redox Cycling of MPP+: Evidence for a New Mechanism Involving Hydride Transfer with Xanthine Oxidase, Aldehyde Dehydrogenase, and Lipoamide Dehydrogenase. Free. Radic. Biol. Med. 1993, 15, 169–179. [Google Scholar] [CrossRef]
- Adams, J.D., Jr.; Klaidman, L.K.; Leung, A.C. MPP+ and MPDP+ Induced Oxygen Radical Formation with Mitochondrial Enzymes. Free Radic. Biol. Med. 1993, 15, 181–186. [Google Scholar] [CrossRef]
- Makino, K.; Hagiwara, T.; Murakami, A. A Mini Review: Fundamental Aspects of Spin Trapping with DMPO. Int. J. Radiat. Appl. Instrum. Part C Radiat. Phys. Chem. 1991, 37, 657–665. [Google Scholar] [CrossRef]
- Hoehne, M.N.; Jacobs, L.J.H.C.; Lapacz, K.J.; Calabrese, G.; Murschall, L.M.; Marker, T.; Kaul, H.; Trifunovic, A.; Morgan, B.; Fricker, M.; et al. Spatial and Temporal Control of Mitochondrial H2O2 Release in Intact Human Cells. EMBO J. 2022, 41, e109169. [Google Scholar] [CrossRef] [PubMed]
- Xiao, B.; Deng, X.; Lim, G.G.Y.; Xie, S.; Zhou, Z.D.; Lim, K.-L.; Tan, E.-K. Superoxide Drives Progression of Parkin/PINK1-Dependent Mitophagy Following Translocation of Parkin to Mitochondria. Cell Death Dis. 2017, 8, e3097. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Wang, B.; Wang, H.; He, H.; Wu, Q.; Qin, X.; Yang, X.; Chen, L.; Xu, G.; Yuan, Z.; et al. Disruption of the Superoxide Anions-Mitophagy Regulation Axis Mediates Copper Oxide Nanoparticles-Induced Vascular Endothelial Cell Death. Free Radic. Biol. Med. 2018, 129, 268–278. [Google Scholar] [CrossRef] [PubMed]
- Aon, M.A.; Cortassa, S.; Marbán, E.; O’Rourke, B. Synchronized Whole Cell Oscillations in Mitochondrial Metabolism Triggered by a Local Release of Reactive Oxygen Species in Cardiac Myocytes. J. Biol. Chem. 2003, 278, 44735–44744. [Google Scholar] [CrossRef]
- Marchissio, M.J.; Francés, D.E.A.; Carnovale, C.E.; Marinelli, R.A. Mitochondrial Aquaporin-8 Knockdown in Human Hepatoma HepG2 Cells Causes ROS-Induced Mitochondrial Depolarization and Loss of Viability. Toxicol. Appl. Pharmacol. 2012, 264, 246–254. [Google Scholar] [CrossRef]
- Almasalmeh, A.; Krenc, D.; Wu, B.; Beitz, E. Structural Determinants of the Hydrogen Peroxide Permeability of Aquaporins. FEBS J. 2014, 281, 647–656. [Google Scholar] [CrossRef]
- Wang, Y.; Yen, F.S.; Zhu, X.G.; Timson, R.C.; Weber, R.; Xing, C.; Liu, Y.; Allwein, B.; Luo, H.; Yeh, H.-W.; et al. SLC25A39 Is Necessary for Mitochondrial Glutathione Import in Mammalian Cells. Nature 2021, 599, 136–140. [Google Scholar] [CrossRef]
- Gialluisi, A.; Reccia, M.G.; Modugno, N.; Nutile, T.; Lombardi, A.; Di Giovannantonio, L.G.; Pietracupa, S.; Ruggiero, D.; Scala, S.; Gambardella, S.; et al. Identification of Sixteen Novel Candidate Genes for Late Onset Parkinson’s Disease. Mol. Neurodegener. 2021, 16, 35. [Google Scholar] [CrossRef]
- Francisco, A.; Engel, D.F.; Figueira, T.R.; Rogério, F.; de Bem, A.F.; Castilho, R.F. Mitochondrial NAD(P)+ Transhydrogenase Is Unevenly Distributed in Different Brain Regions, and Its Loss Causes Depressive-like Behavior and Motor Dysfunction in Mice. Neuroscience 2020, 440, 210–229. [Google Scholar] [CrossRef]
- Palacios-Callender, M.; Quintero, M.; Hollis, V.S.; Springett, R.J.; Moncada, S. Endogenous NO Regulates Superoxide Production at Low Oxygen Concentrations by Modifying the Redox State of Cytochrome c Oxidase. Proc. Natl. Acad. Sci. USA 2004, 101, 7630–7635. [Google Scholar] [CrossRef]
- Brown, G.C. Nitric Oxide and Mitochondrial Respiration. Biochim. Biophys. Acta Bioenerg. 1999, 1411, 351–369. [Google Scholar] [CrossRef]
- Heck, D.; Kagan, V.; Shvedova, A.; Laskin, J. An Epigrammatic (Abridged) Recounting of the Myriad Tales of Astonishing Deeds and Dire Consequences Pertaining to Nitric Oxide and Reactive Oxygen Species in Mitochondria with an Ancillary Missive Concerning the Origins of Apoptosis. Toxicology 2005, 208, 259–271. [Google Scholar] [CrossRef] [PubMed]
- MacMillan-Crow, L.A.; Thompson, J.A. Tyrosine Modifications and Inactivation of Active Site Manganese Superoxide Dismutase Mutant (Y34F) by Peroxynitrite. Arch. Biochem. Biophys. 1999, 366, 82–88. [Google Scholar] [CrossRef]
- Radi, R.; Cassina, A.; Hodara, R.; Quijano, C.; Castro, L. Peroxynitrite Reactions and Formation in Mitochondria. Free. Radic. Biol. Med. 2002, 33, 1451–1464. [Google Scholar] [CrossRef]
- Forsmark-Andrée, P.; Persson, B.; Radi, R.; Dallner, G.; Ernster, L. Oxidative Modification of Nicotinamide Nucleotide Transhydrogenase in Submitochondrial Particles: Effect of Endogenous Ubiquinol. Arch. Biochem. Biophys. 1996, 336, 113–120. [Google Scholar] [CrossRef]
- Zago, E.B.; Castilho, R.F.; Vercesi, A.E. The Redox State of Endogenous Pyridine Nucleotides Can Determine Both the Degree of Mitochondrial Oxidative Stress and the Solute Selectivity of the Permeability Transition Pore. FEBS Lett. 2000, 478, 29–33. [Google Scholar] [CrossRef]
- Ronchi, J.A.; Figueira, T.R.; Ravagnani, F.G.; Oliveira, H.C.F.; Vercesi, A.E.; Castilho, R.F. A Spontaneous Mutation in the Nicotinamide Nucleotide Transhydrogenase Gene of C57BL/6J Mice Results in Mitochondrial Redox Abnormalities. Free. Radic. Biol. Med. 2013, 63, 446–456. [Google Scholar] [CrossRef]
- Thomas, M.H.; Karout, M.; Rodriguez, B.P.; Gui, Y.; Jaeger, C.; Michelucci, A.; Kollmus, H.; Schughart, K.; Coowar, D.; Balling, R.; et al. Strain- and Age-Dependent Features of the Nigro-Striatal Circuit in Three Common Laboratory Mouse Strains, C57BL/6J, A/J, and DBA/2J—Implications for Parkinson’s Disease Modeling. bioRxiv 2020. [Google Scholar] [CrossRef]
- Chen, L.; Zhang, Z.; Hoshino, A.; Zheng, H.D.; Morley, M.; Arany, Z.; Rabinowitz, J.D. NADPH Production by the Oxidative Pentose-Phosphate Pathway Supports Folate Metabolism. Nat. Metab. 2019, 1, 404–415. [Google Scholar] [CrossRef]
- Willems, P.H.G.M.; Rossignol, R.; Dieteren, C.E.J.; Murphy, M.P.; Koopman, W.J.H. Redox Homeostasis and Mitochondrial Dynamics. Cell Metab. 2015, 22, 207–218. [Google Scholar] [CrossRef]
- Terešak, P.; Lapao, A.; Subic, N.; Boya, P.; Elazar, Z.; Simonsen, A. Regulation of PRKN-Independent Mitophagy. Autophagy 2022, 18, 24–39. [Google Scholar] [CrossRef]
- Lopes da Fonseca, T.; Villar-Piqué, A.; Outeiro, T.F. The Interplay between Alpha-Synuclein Clearance and Spreading. Biomolecules 2015, 5, 435–471. [Google Scholar] [CrossRef]
- Peth, A.; Nathan, J.A.; Goldberg, A.L. The ATP Costs and Time Required to Degrade Ubiquitinated Proteins by the 26 S Proteasome. J. Biol. Chem. 2013, 288, 29215–29222. [Google Scholar] [CrossRef]
- Peth, A.; Uchiki, T.; Goldberg, A.L. ATP-Dependent Steps in the Binding of Ubiquitin Conjugates to the 26S Proteasome That Commit to Degradation. Mol. Cell 2010, 40, 671–681. [Google Scholar] [CrossRef]
- Singh, R.; Cuervo, A.M. Autophagy in the Cellular Energetic Balance. Cell Metab. 2011, 13, 495–504. [Google Scholar] [CrossRef]
- Bell, S.M.; Burgess, T.; Lee, J.; Blackburn, D.J.; Allen, S.P.; Mortiboys, H. Peripheral Glycolysis in Neurodegenerative Diseases. Int. J. Mol. Sci. 2020, 21, 8924. [Google Scholar] [CrossRef]
- Beg, M.; Abdullah, N.; Thowfeik, F.S.; Altorki, N.K.; McGraw, T.E. Distinct Akt Phosphorylation States Are Required for Insulin Regulated Glut4 and Glut1-Mediated Glucose Uptake. eLife 2017, 6, e26896. [Google Scholar] [CrossRef]
- Cai, R.; Zhang, Y.; Simmering, J.E.; Schultz, J.L.; Li, Y.; Fernandez-Carasa, I.; Consiglio, A.; Raya, A.; Polgreen, P.M.; Narayanan, N.S.; et al. Enhancing Glycolysis Attenuates Parkinson’s Disease Progression in Models and Clinical Databases. J. Clin. Investig. 2019, 129, 4539–4549. [Google Scholar] [CrossRef]
- Rhea, E.M.; Banks, W.A. Interactions of Lipids, Lipoproteins, and Apolipoproteins with the Blood-Brain Barrier. Pharm. Res. 2021, 38, 1469–1475. [Google Scholar] [CrossRef]
- Panov, A.; Orynbayeva, Z.; Vavilin, V.; Lyakhovich, V. Fatty Acids in Energy Metabolism of the Central Nervous System. Biomed Res. Int. 2014, 2014, 472459. [Google Scholar] [CrossRef]
- Barber, C.N.; Raben, D.M. Lipid Metabolism Crosstalk in the Brain: Glia and Neurons. Front. Cell Neurosci. 2019, 13, 212. [Google Scholar] [CrossRef]
- Ioannou, M.S.; Jackson, J.; Sheu, S.-H.; Chang, C.-L.; Weigel, A.V.; Liu, H.; Pasolli, H.A.; Xu, C.S.; Pang, S.; Matthies, D.; et al. Neuron-Astrocyte Metabolic Coupling Protects against Activity-Induced Fatty Acid Toxicity. Cell 2019, 177, 1522–1535.e14. [Google Scholar] [CrossRef]
- Dedkova, E.N.; Blatter, L.A. Role of β-Hydroxybutyrate, Its Polymer Poly-β-Hydroxybutyrate and Inorganic Polyphosphate in Mammalian Health and Disease. Front. Physiol. 2014, 5, 260. [Google Scholar] [CrossRef]
- Koronowski, K.B.; Greco, C.M.; Huang, H.; Kim, J.-K.; Fribourgh, J.L.; Crosby, P.; Mathur, L.; Ren, X.; Partch, C.L.; Jang, C.; et al. Ketogenesis Impact on Liver Metabolism Revealed by Proteomics of Lysine β-Hydroxybutyrylation. Cell Rep. 2021, 36, 109487. [Google Scholar] [CrossRef]
- Terranova, C.J.; Stemler, K.M.; Barrodia, P.; Jeter-Jones, S.L.; Ge, Z.; de la Cruz Bonilla, M.; Raman, A.; Cheng, C.-W.; Allton, K.L.; Arslan, E.; et al. Reprogramming of H3K9bhb at Regulatory Elements Is a Key Feature of Fasting in the Small Intestine. Cell Rep. 2021, 37, 110044. [Google Scholar] [CrossRef]
- Benderdour, M.; Charron, G.; deBlois, D.; Comte, B.; Rosiers, C.D. Cardiac Mitochondrial NADP+-Isocitrate Dehydrogenase Is Inactivated through 4-Hydroxynonenal Adduct Formation: An Event that Precedes Hypertrophy Development. J. Biol. Chem. 2003, 278, 45154–45159. [Google Scholar] [CrossRef]
- Perez, M.A.; Magtanong, L.; Dixon, S.J.; Watts, J.L. Dietary Lipids Induce Ferroptosis in Caenorhabditis Elegans and Human Cancer Cells. Dev. Cell 2020, 54, 447–454.e4. [Google Scholar] [CrossRef]
- Yamada, N.; Karasawa, T.; Kimura, H.; Watanabe, S.; Komada, T.; Kamata, R.; Sampilvanjil, A.; Ito, J.; Nakagawa, K.; Kuwata, H.; et al. Ferroptosis Driven by Radical Oxidation of N-6 Polyunsaturated Fatty Acids Mediates Acetaminophen-Induced Acute Liver Failure. Cell Death Dis. 2020, 11, 144. [Google Scholar] [CrossRef]
- Tripathi, A.; Fanning, S.; Dettmer, U. Lipotoxicity Downstream of α-Synuclein Imbalance: A Relevant Pathomechanism in Synucleinopathies? Biomolecules 2021, 12, 40. [Google Scholar] [CrossRef]
- Kagan, V.E.; Tyurin, V.A.; Jiang, J.; Tyurina, Y.Y.; Ritov, V.B.; Amoscato, A.A.; Osipov, A.N.; Belikova, N.A.; Kapralov, A.A.; Kini, V.; et al. Cytochrome c Acts as a Cardiolipin Oxygenase Required for Release of Proapoptotic Factors. Nat. Chem. Biol. 2005, 1, 223–232. [Google Scholar] [CrossRef]
- Jenkins, C.M.; Yang, K.; Liu, G.; Moon, S.H.; Dilthey, B.G.; Gross, R.W. Cytochrome c Is an Oxidative Stress–Activated Plasmalogenase That Cleaves Plasmenylcholine and Plasmenylethanolamine at the Sn-1 Vinyl Ether Linkage. J. Biol. Chem. 2018, 293, 8693–8709. [Google Scholar] [CrossRef] [PubMed]
- Hsu, Y.-H.; Dumlao, D.S.; Cao, J.; Dennis, E.A. Assessing Phospholipase A2 Activity toward Cardiolipin by Mass Spectrometry. PLoS ONE 2013, 8, e59267. [Google Scholar] [CrossRef]
- Mancuso, D.J.; Kotzbauer, P.; Wozniak, D.F.; Sims, H.F.; Jenkins, C.M.; Guan, S.; Han, X.; Yang, K.; Sun, G.; Malik, I.; et al. Genetic Ablation of Calcium-Independent Phospholipase A2γ Leads to Alterations in Hippocampal Cardiolipin Content and Molecular Species Distribution, Mitochondrial Degeneration, Autophagy, and Cognitive Dysfunction. J. Biol. Chem. 2009, 284, 35632–35644. [Google Scholar] [CrossRef] [PubMed]
- Mahajan, M.; Bharambe, N.; Shang, Y.; Lu, B.; Mandal, A.; Madan Mohan, P.; Wang, R.; Boatz, J.C.; Manuel Martinez Galvez, J.; Shnyrova, A.V.; et al. NMR Identification of a Conserved Drp1 Cardiolipin-Binding Motif Essential for Stress-Induced Mitochondrial Fission. Proc. Natl. Acad. Sci. USA 2021, 118, e2023079118. [Google Scholar] [CrossRef] [PubMed]
- Chu, C.T.; Ji, J.; Dagda, R.K.; Jiang, J.F.; Tyurina, Y.Y.; Kapralov, A.A.; Tyurin, V.A.; Yanamala, N.; Shrivastava, I.H.; Mohammadyani, D.; et al. Cardiolipin Externalization to the Outer Mitochondrial Membrane Acts as an Elimination Signal for Mitophagy in Neuronal Cells. Nat. Cell Biol. 2013, 15, 1197–1205. [Google Scholar] [CrossRef] [PubMed]
- Iyer, S.S.; He, Q.; Janczy, J.R.; Elliott, E.I.; Zhong, Z.; Olivier, A.K.; Sadler, J.J.; Knepper-Adrian, V.; Han, R.; Qiao, L.; et al. Mitochondrial Cardiolipin Is Required for Nlrp3 Inflammasome Activation. Immunity 2013, 39, 311–323. [Google Scholar] [CrossRef]
- Monteiro-Cardoso, V.F.; Oliveira, M.M.; Melo, T.; Domingues, M.R.M.; Moreira, P.I.; Ferreiro, E.; Peixoto, F.; Videira, R.A. Cardiolipin Profile Changes Are Associated to the Early Synaptic Mitochondrial Dysfunction in Alzheimer’s Disease. J. Alzheimer’s Dis. 2015, 43, 1375–1392. [Google Scholar] [CrossRef]
- Kimura, T.; Kimura, A.K.; Ren, M.; Monteiro, V.; Xu, Y.; Berno, B.; Schlame, M.; Epand, R.M. Plasmalogen Loss Caused by Remodeling Deficiency in Mitochondria. Life Sci. Alliance 2019, 2, e201900348. [Google Scholar] [CrossRef]
- Senanayake, V.; Goodenowe, D.B. Plasmalogen Deficiency and Neuropathology in Alzheimer’s Disease: Causation or Coincidence? Alzheimers Dement. 2019, 5, 524–532. [Google Scholar] [CrossRef]
- Wood, P.L.; Smith, T.; Lane, N.; Khan, M.A.; Ehrmantraut, G.; Goodenowe, D.B. Oral Bioavailability of the Ether Lipid Plasmalogen Precursor, PPI-1011, in the Rabbit: A New Therapeutic Strategy for Alzheimer’s Disease. Lipids Health Dis. 2011, 10, 227. [Google Scholar] [CrossRef]
- Mawatari, S.; Ohara, S.; Taniwaki, Y.; Tsuboi, Y.; Maruyama, T.; Fujino, T. Improvement of Blood Plasmalogens and Clinical Symptoms in Parkinson’s Disease by Oral Administration of Ether Phospholipids: A Preliminary Report. Parkinson’s Dis. 2020, 2020, 2671070. [Google Scholar] [CrossRef] [PubMed]
- Dorninger, F.; Forss-Petter, S.; Wimmer, I.; Berger, J. Plasmalogens, Platelet-Activating Factor and beyond—Ether Lipids in Signaling and Neurodegeneration. Neurobiol. Dis. 2020, 145, 105061. [Google Scholar] [CrossRef] [PubMed]
- Jo, D.S.; Cho, D.-H. Peroxisomal Dysfunction in Neurodegenerative Diseases. Arch. Pharm. Res. 2019, 42, 393–406. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Bai, H. Peroxisomal Stress Response and Inter-Organelle Communication in Cellular Homeostasis and Aging. Antioxidants 2022, 11, 192. [Google Scholar] [CrossRef] [PubMed]
- Salem, N.; Litman, B.; Kim, H.-Y.; Gawrisch, K. Mechanisms of Action of Docosahexaenoic Acid in the Nervous System. Lipids 2001, 36, 945–959. [Google Scholar] [CrossRef]
- Hoffman-Kuczynski, B.; Reo, N.V. Administration of Myo-Inositol Plus Ethanolamine Elevates Phosphatidylethanolamine Plasmalogen in the Rat Cerebellum. Neurochem. Res. 2005, 30, 47–60. [Google Scholar] [CrossRef]
- Torres, S.; García-Ruiz, C.M.; Fernandez-Checa, J.C. Mitochondrial Cholesterol in Alzheimer’s Disease and Niemann–Pick Type C Disease. Front. Neurol. 2019, 10, 1168. [Google Scholar] [CrossRef]
- Jayashankar, V.; Selwan, E.; Hancock, S.E.; Verlande, A.; Goodson, M.O.; Eckenstein, K.H.; Milinkeviciute, G.; Hoover, B.M.; Chen, B.; Fleischman, A.G.; et al. Drug-like Sphingolipid SH-BC-893 Opposes Ceramide-induced Mitochondrial Fission and Corrects Diet-induced Obesity. EMBO Mol. Med. 2021, 13, e13086. [Google Scholar] [CrossRef]
- Brekk, O.R.; Honey, J.R.; Lee, S.; Hallett, P.J.; Isacson, O. Cell Type-Specific Lipid Storage Changes in Parkinson’s Disease Patient Brains Are Recapitulated by Experimental Glycolipid Disturbance. Proc. Natl. Acad. Sci. USA 2020, 117, 27646–27654. [Google Scholar] [CrossRef]
- Zhu, J.; Schwörer, S.; Berisa, M.; Kyung, Y.J.; Ryu, K.W.; Yi, J.; Jiang, X.; Cross, J.R.; Thompson, C.B. Mitochondrial NADP(H) Generation Is Essential for Proline Biosynthesis. Science 2021, 372, 968–972. [Google Scholar] [CrossRef]
- Abraham, M.A.; Lam, T.K.T. Glucagon Action in the Brain. Diabetologia 2016, 59, 1367–1371. [Google Scholar] [CrossRef] [PubMed]
- Berwick, D.C.; Hers, I.; Heesom, K.J.; Moule, S.K.; Tavareá, J.M. The Identification of ATP-Citrate Lyase as a Protein Kinase B (Akt) Substrate in Primary Adipocytes. J. Biol. Chem. 2002, 277, 33895–33900. [Google Scholar] [CrossRef] [PubMed]
- Brownsey, R.W.; Boone, A.N.; Elliott, J.E.; Kulpa, J.E.; Lee, W.M. Regulation of Acetyl-CoA Carboxylase. Biochem. Soc. Trans. 2006, 34, 5. [Google Scholar] [CrossRef]
- Hoxhaj, G.; Ben-Sahra, I.; Lockwood, S.E.; Timson, R.C.; Byles, V.; Henning, G.T.; Gao, P.; Selfors, L.M.; Asara, J.M.; Manning, B.D. Direct Stimulation of NADP+ Synthesis through Akt-Mediated Phosphorylation of NAD Kinase. Science 2019, 363, 1088–1092. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Xu, Y.; Lu, W.; Li, J.; Yu, S.; Brown, E.J.; Stanger, B.Z.; Rabinowitz, J.D.; Yang, X. G6PD-Mediated Increase in De Novo NADP+ Biosynthesis Promotes Antioxidant Defense and Tumor Metastasis. Sci. Adv. 2022, 8, eabo0404. [Google Scholar] [CrossRef]
- Peng, I.-C.; Chen, Z.; Sun, W.; Li, Y.-S.; Marin, T.L.; Hsu, P.-H.; Su, M.-I.; Cui, X.; Pan, S.; Lytle, C.Y.; et al. Glucagon Regulates ACC Activity in Adipocytes through the CAMKKβ/AMPK Pathway. Am. J. Physiol. Endocrinol. Metab. 2012, 302, E1560–E1568. [Google Scholar] [CrossRef][Green Version]
- Wang, H.; Dou, S.; Zhu, J.; Shao, Z.; Wang, C.; Xu, X.; Cheng, B. Ghrelin Protects against Rotenone-Induced Cytotoxicity: Involvement of Mitophagy and the AMPK/SIRT1/PGC1α Pathway. Neuropeptides 2021, 87, 102134. [Google Scholar] [CrossRef]
- Kobilo, T.; Guerrieri, D.; Zhang, Y.; Collica, S.C.; Becker, K.G.; van Praag, H. AMPK Agonist AICAR Improves Cognition and Motor Coordination in Young and Aged Mice. Learn. Mem. 2014, 21, 119–126. [Google Scholar] [CrossRef]
- Ramamurthy, S.; Chang, E.; Cao, Y.; Zhu, J.; Ronnett, G.V. AMPK Activation Regulates Neuronal Structure in Developing Hippocampal Neurons. Neuroscience 2014, 259, 13–24. [Google Scholar] [CrossRef]
- McCarthy, C.G.; Waigi, E.W.; Yeoh, B.S.; Mell, B.; Vijay-Kumar, M.; Wenceslau, C.F.; Joe, B. Low-Dose 1,3-Butanediol Reverses Age-Associated Vascular Dysfunction Independent of Ketone Body β-Hydroxybutyrate. Am. J. Physiol. Heart Circ. Physiol. 2022, 322, H466–H473. [Google Scholar] [CrossRef]
- McReynolds, M.R.; Chellappa, K.; Chiles, E.; Jankowski, C.; Shen, Y.; Chen, L.; Descamps, H.C.; Mukherjee, S.; Bhat, Y.R.; Lingala, S.R.; et al. NAD+ Flux Is Maintained in Aged Mice despite Lower Tissue Concentrations. Cell Syst. 2021, 12, 1160–1172.e4. [Google Scholar] [CrossRef] [PubMed]
- Merrill, D.K.; Guynn, R.W. The Calculation of the Cytoplasmic Free [NADP+]/[NADPH] Ratio in Brain: Effect of Electroconvulsive Seizure. Brain Res. 1981, 221, 307–318. [Google Scholar] [CrossRef]
- Parihar, M.S.; Kunz, E.A.; Brewer, G.J. Age-Related Decreases in NAD(P)H and Glutathione Cause Redox Declines before ATP Loss during Glutamate Treatment of Hippocampal Neurons. J. Neurosci. Res. 2008, 86, 2339–2352. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, D.; Levault, K.R.; Brewer, G.J. Relative Importance of Redox Buffers GSH and NAD(P)H in Age-Related Neurodegeneration and Alzheimer Disease-like Mouse Neurons. Aging Cell 2014, 13, 631–640. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, D.; LeVault, K.R.; Barnett, A.J.; Brewer, G.J. A Reversible Early Oxidized Redox State That Precedes Macromolecular ROS Damage in Aging Nontransgenic and 3xTg-AD Mouse Neurons. J. Neurosci. 2012, 32, 5821–5832. [Google Scholar] [CrossRef]
- Blacker, T.S.; Duchen, M.R. Investigating Mitochondrial Redox State Using NADH and NADPH Autofluorescence. Free Radic. Biol. Med. 2016, 100, 53–65. [Google Scholar] [CrossRef]
- Subrahmanian, N.; LaVoie, M.J. Is There a Special Relationship between Complex I Activity and Nigral Neuronal Loss in Parkinson’s Disease? A Critical Reappraisal. Brain Res. 2021, 1767, 147434. [Google Scholar] [CrossRef]
- Borst, P. The Malate–Aspartate Shuttle (Borst Cycle): How It Started and Developed into a Major Metabolic Pathway. IUBMB Life 2020, 72, 2241–2259. [Google Scholar] [CrossRef]
- Ren, X.; Chen, J.-F. Caffeine and Parkinson’s Disease: Multiple Benefits and Emerging Mechanisms. Front. Neurosci. 2020, 14, 602697. [Google Scholar] [CrossRef]
- Zhang, Q.; Wu, Y.; Zhang, P.; Sha, H.; Jia, J.; Hu, Y.; Zhu, J. Exercise Induces Mitochondrial Biogenesis after Brain Ischemia in Rats. Neuroscience 2012, 205, 10–17. [Google Scholar] [CrossRef]
- Park, J.; Kim, J.; Mikami, T. Exercise-Induced Lactate Release Mediates Mitochondrial Biogenesis in the Hippocampus of Mice via Monocarboxylate Transporters. Front. Physiol. 2021, 12, 736905. [Google Scholar] [CrossRef] [PubMed]
- Yanagi, S.; Sato, T.; Kangawa, K.; Nakazato, M. The Homeostatic Force of Ghrelin. Cell Metab. 2018, 27, 786–804. [Google Scholar] [CrossRef]
- Liu, C.; Xue, Y.; Liu, M.-F.; Wang, Y.; Chen, L. Orexin and Parkinson’s Disease: A Protective Neuropeptide with Therapeutic Potential. Neurochem. Int. 2020, 138, 104754. [Google Scholar] [CrossRef]
- Elamin, M.; Ruskin, D.N.; Masino, S.A.; Sacchetti, P. Ketone-Based Metabolic Therapy: Is Increased NAD+ a Primary Mechanism? Front. Mol. Neurosci. 2017, 10, 377. [Google Scholar] [CrossRef] [PubMed]
- Izuta, Y.; Imada, T.; Hisamura, R.; Oonishi, E.; Nakamura, S.; Inagaki, E.; Ito, M.; Soga, T.; Tsubota, K. Ketone Body 3-hydroxybutyrate Mimics Calorie Restriction via the Nrf2 Activator, Fumarate, in the Retina. Aging Cell 2018, 17, e12699. [Google Scholar] [CrossRef] [PubMed]
- Youm, Y.-H.; Nguyen, K.Y.; Grant, R.W.; Goldberg, E.L.; Bodogai, M.; Kim, D.; D’Agostino, D.; Planavsky, N.; Lupfer, C.; Kanneganti, T.D.; et al. Ketone Body β-Hydroxybutyrate Blocks the NLRP3 Inflammasome-Mediated Inflammatory Disease. Nat. Med. 2015, 21, 263–269. [Google Scholar] [CrossRef] [PubMed]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Curtis, W.M.; Seeds, W.A.; Mattson, M.P.; Bradshaw, P.C. NADPH and Mitochondrial Quality Control as Targets for a Circadian-Based Fasting and Exercise Therapy for the Treatment of Parkinson’s Disease. Cells 2022, 11, 2416. https://doi.org/10.3390/cells11152416
Curtis WM, Seeds WA, Mattson MP, Bradshaw PC. NADPH and Mitochondrial Quality Control as Targets for a Circadian-Based Fasting and Exercise Therapy for the Treatment of Parkinson’s Disease. Cells. 2022; 11(15):2416. https://doi.org/10.3390/cells11152416
Chicago/Turabian StyleCurtis, William M., William A. Seeds, Mark P. Mattson, and Patrick C. Bradshaw. 2022. "NADPH and Mitochondrial Quality Control as Targets for a Circadian-Based Fasting and Exercise Therapy for the Treatment of Parkinson’s Disease" Cells 11, no. 15: 2416. https://doi.org/10.3390/cells11152416
APA StyleCurtis, W. M., Seeds, W. A., Mattson, M. P., & Bradshaw, P. C. (2022). NADPH and Mitochondrial Quality Control as Targets for a Circadian-Based Fasting and Exercise Therapy for the Treatment of Parkinson’s Disease. Cells, 11(15), 2416. https://doi.org/10.3390/cells11152416