EGFR Status Assessment for Better Care of Early Stage Non-Small Cell Lung Carcinoma: What Is Changing in the Daily Practice of Pathologists?
Abstract
:1. Introduction
2. Short Overview of Studies concerning the Use of EGFR TKIs in Early Stage NSCLC in Neoadjuvant and Adjuvant Setting
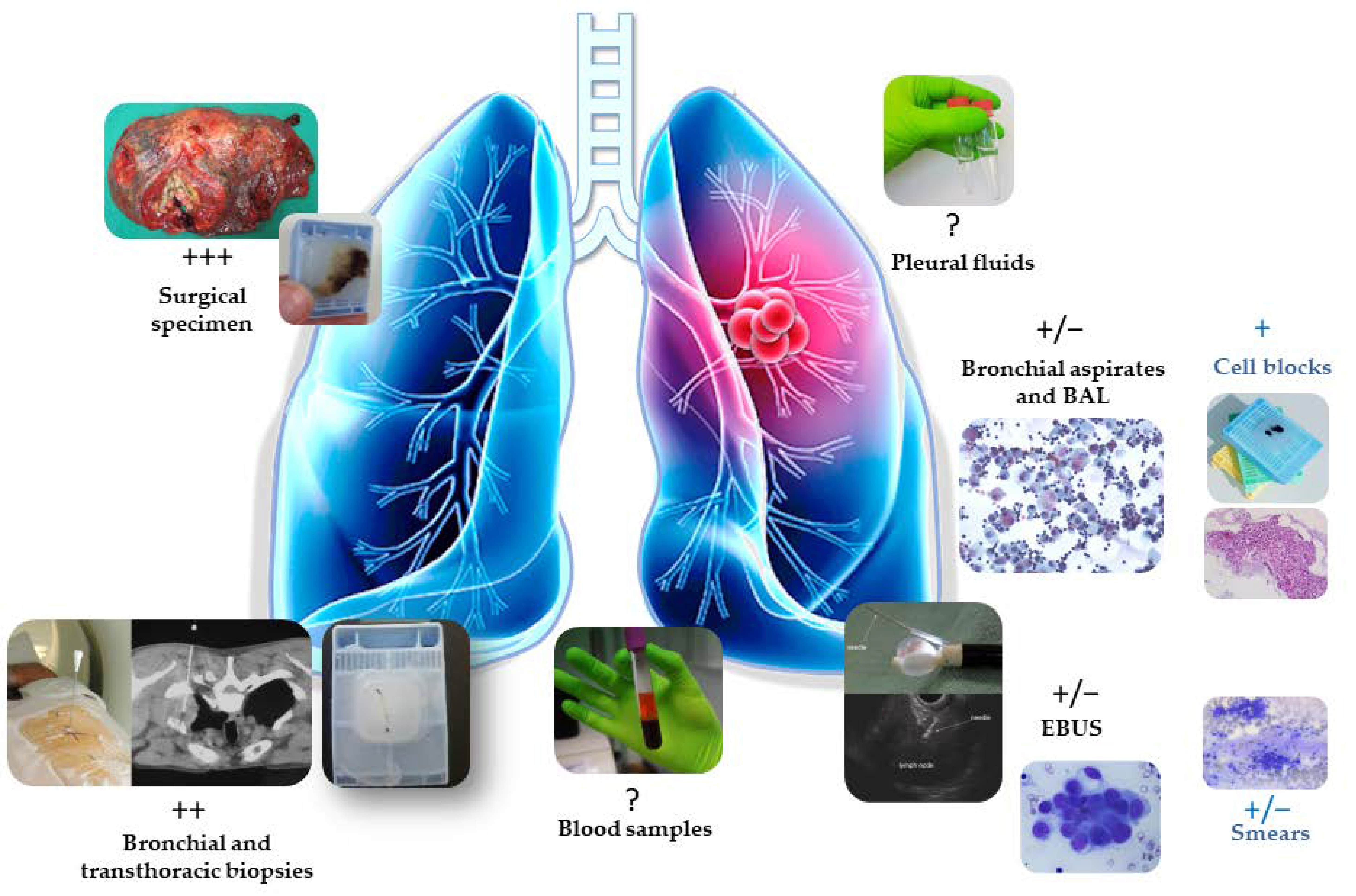
3. Which Biological Sample for Evaluation of the EGFR Status of Early Stage NSCLC?
4. Which Approach(es) for Sequencing Technology?
5. Issues concerning the Cost and Reimbursement
6. Neoadjuvant Immunotherapy or Neoadjuvant Immuno/Chemotherapy versus Adjuvant Targeted Therapy against EGFR: How to Make the Right Choice?
7. Perspectives and Issues
8. Conclusions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Herbst, R.S.; Morgensztern, D.; Boshoff, C. The biology and management of non-small cell lung cancer. Nature 2018, 553, 446–454. [Google Scholar] [CrossRef]
- Howlader, N.; Forjaz, G.; Mooradian, M.J.; Meza, R.; Kong, C.Y.; Cronin, K.A.; Mariotto, A.B.; Lowy, D.R.; Feuer, E.J. The Effect of Advances in Lung-Cancer Treatment on Population Mortality. N. Engl. J. Med. 2020, 383, 640–649. [Google Scholar] [CrossRef]
- Chaft, J.E.; Rimner, A.; Weder, W.; Azzoli, C.G.; Kris, M.G.; Cascone, T. Evolution of systemic therapy for stages I-III non-metastatic non-small-cell lung cancer. Nat. Rev. Clin. Oncol. 2021, 18, 547–557. [Google Scholar] [CrossRef]
- Howington, J.A.; Blum, M.G.; Chang, A.C.; Balekian, A.A.; Murthy, S.C. Treatment of stage I and II non-small cell lung cancer: Diagnosis and management of lung cancer, 3rd ed: American College of Chest Physicians evidence-based clinical practice guidelines. Chest 2013, 143 (Suppl. S5), e278S–e313S. [Google Scholar] [CrossRef]
- Spaans, J.N.; Goss, G.D. Epidermal growth factor receptor tyrosine kinase inhibitors in early-stage nonsmall cell lung cancer. Curr. Opin. Oncol. 2015, 27, 102–107. [Google Scholar] [CrossRef] [PubMed]
- Ahern, E.; Solomon, B.J.; Hui, R.; Pavlakis, N.; O’Byrne, K.; Hughes, B.G.M. Neoadjuvant immunotherapy for non-small cell lung cancer: Right drugs, right patient, right time? J. Immunother. Cancer 2021, 9, e002248. [Google Scholar] [CrossRef] [PubMed]
- Attili, I.; Passaro, A.; Spaggiari, L.; Spitaleri, G.; de Marinis, F. Adjuvant EGFR TKIs in NSCLC harboring EGFR mutations: Looking for a consensus way. Ann. Transl. Med. 2020, 8, 1111. [Google Scholar] [CrossRef]
- Broderick, S.R. Adjuvant and Neoadjuvant Immunotherapy in Non-Small Cell Lung Cancer. Thorac. Surg. Clin. 2020, 30, 215–220. [Google Scholar] [CrossRef] [PubMed]
- Kang, J.; Zhang, C.; Zhong, W.Z. Neoadjuvant immunotherapy for non-small cell lung cancer: State of the art. Cancer Commun. 2021, 41, 287–302. [Google Scholar] [CrossRef]
- Krishnamoorthy, M.; Lenehan, J.G.; Maleki Vareki, S. Neoadjuvant Immunotherapy for High-Risk, Resectable Malignancies: Scientific Rationale and Clinical Challenges. J. Natl. Cancer Inst. 2021, 113, 823–832. [Google Scholar] [CrossRef] [PubMed]
- Pall, G. Neoadjuvant immunotherapy in nonsmall cell lung cancer. Curr. Opin. Oncol. 2021, 33, 59–63. [Google Scholar] [CrossRef] [PubMed]
- Shukla, N.; Hanna, N. Neoadjuvant and Adjuvant Immunotherapy in Early-Stage Non-Small Cell Lung Cancer. Lung Cancer 2021, 12, 51–60. [Google Scholar] [CrossRef] [PubMed]
- Uprety, D.; Mandrekar, S.J.; Wigle, D.; Roden, A.C.; Adjei, A.A. Neoadjuvant Immunotherapy for NSCLC: Current Concepts and Future Approaches. J. Thorac. Oncol. 2020, 15, 1281–1297. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.L.; Tsuboi, M.; He, J.; John, T.; Grohe, C.; Majem, M.; Goldman, J.W.; Laktionov, K.; Kim, S.W.; Kato, T.; et al. Osimertinib in Resected EGFR-Mutated Non-Small-Cell Lung Cancer. N. Engl. J. Med. 2020, 383, 1711–1723. [Google Scholar] [CrossRef]
- Hofman, P. PD-L1 immunohistochemistry for non-small cell lung carcinoma: Which strategy should be adopted? Expert. Rev. Mol. Diagn. 2017, 17, 1097–1108. [Google Scholar] [CrossRef]
- Hofman, P.; Barlesi, F. Companion diagnostic tests for treatment of lung cancer patients: What are the current and future challenges? Expert Rev. Mol. Diagn. 2019, 19, 429–438. [Google Scholar] [CrossRef]
- Xu, Y.; Wan, B.; Chen, X.; Zhan, P.; Zhao, Y.; Zhang, T.; Liu, H.; Afzal, M.Z.; Dermime, S.; Hochwald, S.N.; et al. Transl. Lung Cancer Res. The association of PD-L1 expression with the efficacy of anti-PD-1/PD-L1 immunotherapy and survival of non-small cell lung cancer patients: A meta-analysis of randomized controlled trials. Transl. Lung Cancer Res. 2019, 8, 413–428. [Google Scholar] [CrossRef]
- Lin, C.; Hu, F.; Chu, H.; Ren, P.; Ma, S.; Wang, J.; Bai, J.; Han, X.; Ma, S. The role of EGFR-TKIs as adjuvant therapy in EGFR mutation-positive early-stage NSCLC: A meta-analysis. Thorac. Cancer 2021, 12, 1084–1095. [Google Scholar] [CrossRef]
- Mitsudomi, T.; Soh, J. Adjuvant therapy of operable nonsmall cell lung cancer: An update. Curr. Opin. Oncol. 2021, 33, 47–54. [Google Scholar] [CrossRef]
- Ortega-Franco, A.; Calvo, V.; Franco, F.; Provencio, M.; Califano, R. Integrating immune checkpoint inhibitors and targeted therapies in the treatment of early stage non-small cell lung cancer: A narrative review. Transl. Lung Cancer Res. 2020, 9, 2656–2673. [Google Scholar] [CrossRef]
- Zhong, W.Z.; Wang, Q.; Mao, W.M.; Xu, S.T.; Wu, L.; Wei, Y.C.; Liu, Y.Y.; Chen, C.; Cheng, Y.; Yin, R.; et al. Gefitinib Versus Vinorelbine Plus Cisplatin as Adjuvant Treatment for Stage II-IIIA (N1-N2) EGFR-Mutant NSCLC: Final Overall Survival Analysis of CTONG1104 Phase III Trial. J. Clin. Oncol. 2021, 39, 713–722. [Google Scholar] [CrossRef] [PubMed]
- Marquez-Medina, D.; Popat, S. Eventual role of EGFR-tyrosine kinase inhibitors in early-stage non-small-cell lung cancer. Future Oncol. 2016, 12, 815–825. [Google Scholar] [CrossRef]
- Tsuboi, M.; Weder, W.; Escriu, C.; Blakely, C.; He, J.; Dacic, S.; Yatabe, Y.; Zeng, L.; Walding, A.; Chaft, J.E. Neoadjuvant osimertinib with/without chemotherapy versus chemotherapy alone for EGFR-mutated resectable non-small-cell lung cancer: NeoADAURA. Future Oncol. 2021. [CrossRef]
- FDA Approves Osimertinib for Adjuvant Treatment of Early-Stage EGFR-Mutated NSCLC. 16 February 2021. Available online: https://www.ajmc.com/view/fda-approves-tagrisso-for-adjuvant-treatment-of-early-stage-egfr-mutated-nsclc (accessed on 2 August 2021).
- AstraZeneca’s Tagrisso Recommended by EMA Committee in Adjuvant NSCLC. 26 April 2021. Available online: https://www.precisiononcologynews.com/cancer/astrazenecas-tagrisso-recommended-ema-committee-adjuvant-nsclc# (accessed on 2 August 2021).
- Chaft, J.E.; Dagogo-Jack, I.; Santini, F.C.; Eng, J.; Yeap, B.Y.; Izar, B.; Chin, E.; Jones, D.R.; Kris, M.G.; Shaw, A.T.; et al. Clinical outcomes of patients with resected, early-stage ALK-positive lung cancer. Lung Cancer 2018, 122, 67–71. [Google Scholar] [CrossRef]
- Cheema, P.K.; Gomes, M.; Banerji, S.; Joubert, P.; Leighl, N.B.; Melosky, B.; Sheffield, B.S.; Stockley, T.; Ionescu, D.N. Consensus recommendations for optimizing biomarker testing to identify and treat advanced EGFR-mutated non-small-cell lung cancer. Curr. Oncol. 2020, 27, 321–329. [Google Scholar] [CrossRef] [PubMed]
- Cao, C.; Guo, A.; Chen, C.; Chakos, A.; Bott, M.; Yang, C.J.; Zielinski, R.; Melfi, F. Systematic Review of Neoadjuvant Immunotherapy for Patients With Non-Small Cell Lung Cancer. Semin. Thorac. Cardiovasc. Surg. 2021. [CrossRef]
- Compton, C.C.; Robb, J.A.; Anderson, M.W.; Berry, A.B.; Birdsong, G.G.; Bloom, K.J.; Branton, P.A.; Crothers, J.W.; Cushman-Vokoun, A.M.; Hicks, D.G.; et al. Preanalytics and Precision Pathology: Pathology Practices to Ensure Molecular Integrity of Cancer Patient Biospecimens for Precision Medicine. Arch. Pathol. Lab. Med. 2019, 143, 1346–1363. [Google Scholar] [CrossRef] [Green Version]
- Ilie, M.; Hofman, P. Pitfalls in lung cancer molecular pathology: How to limit them in routine practice? Curr. Med. Chem. 2012, 19, 2638–2651. [Google Scholar] [CrossRef]
- Kim, L.; Tsao, M.S. Tumour tissue sampling for lung cancer management in the era of personalised therapy: What is good enough for molecular testing? Eur. Respir. J. 2014, 44, 1011–1022. [Google Scholar] [CrossRef]
- Hofman, P. The challenges of evaluating predictive biomarkers using small biopsy tissue samples and liquid biopsies from non-small cell lung cancer patients. J. Thorac. Dis. 2019, 11 (Suppl. S1), S57–S64. [Google Scholar] [CrossRef]
- He, D.; Wang, D.; Lu, P.; Yang, N.; Xue, Z.; Zhu, X.; Zhang, P.; Fan, G. Single-cell RNA sequencing reveals heterogeneous tumor and immune cell populations in early-stage lung adenocarcinomas harboring EGFR mutations. Oncogene 2021, 40, 355–368. [Google Scholar] [CrossRef]
- Passaro, A.; Malapelle, U.; Del Re, M.; Attili, I.; Russo, A.; Guerini-Rocco, E.; Fumagalli, C.; Pisapia, P.; Pepe, F.; De Luca, C.; et al. Understanding EGFR heterogeneity in lung cancer. ESMO Open 2020, 5, e000919. [Google Scholar] [CrossRef]
- Sakurada, A.; Lara-Guerra, H.; Liu, N.; Shepherd, F.A.; Tsao, M.S. Tissue heterogeneity of EGFR mutation in lung adenocarcinoma. J. Thorac. Oncol. 2008, 3, 527–529. [Google Scholar] [CrossRef] [Green Version]
- Yatabe, Y.; Matsuo, K.; Mitsudomi, T. Heterogeneous distribution of EGFR mutations is extremely rare in lung adenocarcinoma. J. Clin. Oncol. 2011, 29, 2972–2977. [Google Scholar] [CrossRef] [PubMed]
- Mohar, B.; Smojver Ježek, S.; Rajković Molek, K.; Štemberger, C.; Kurpis, M.; Kupanovac, Ž.; Samaržija, M.; Jonjić, N.; Grahovac, B. Detection of an EGFR mutation in cytological specimens of lung adenocarcinoma. Cytopathology 2016, 27, 444–451. [Google Scholar] [CrossRef] [PubMed]
- Vigliar, E.; Malapelle, U.; Bellevicine, C.; de Luca, C.; Troncone, G. Outsourcing cytological samples to a referral laboratory for EGFR testing in non-small cell lung cancer: Does theory meet practice? Cytopathology 2015, 26, 312–317. [Google Scholar] [CrossRef]
- Aggarwal, C.; Rolfo, C.D.; Oxnard, G.R.; Gray, J.E.; Sholl, L.M.; Gandara, D.R. Strategies for the successful implementation of plasma-based NSCLC genotyping in clinical practice. Nat. Rev. Clin. Oncol. 2021, 8, 56–62. [Google Scholar] [CrossRef] [PubMed]
- Hofman, P. Liquid Biopsy and Therapeutic Targets: Present and Future Issues in Thoracic Oncology. Cancers 2017, 9, 154. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hofman, V.; Hofman, P. Resistances to EGFR tyrosine kinase inhibitors in lung cancer-how to routinely track them in a molecular pathology laboratory? J. Thorac. Dis. 2019, 11 (Suppl. S1), S65–S70. [Google Scholar] [CrossRef] [PubMed]
- Hofman, P. Next-Generation Sequencing with Liquid Biopsies from Treatment-Naïve Non-Small Cell Lung Carcinoma Patients. Cancers 2021, 13, 2049. [Google Scholar] [CrossRef] [PubMed]
- Lam, V.K.; Zhang, J.; Wu, C.C.; Tran, H.T.; Li, L.; Diao, L.; Wang, J.; Rinsurongkawong, W.; Raymond, V.M.; Lanman, R.B.; et al. Genotype-Specific Differences in Circulating Tumor DNA Levels in Advanced NSCLC. J. Thorac. Oncol. 2021, 16, 601–609. [Google Scholar] [CrossRef]
- Pérez-Barrios, C.; Sánchez-Herrero, E.; Garcia-Simón, N.; Barquín, M.; Clemente, M.B.; Provencio, M.; Romero, A. ctDNA from body fluids is an adequate source for EGFR biomarker testing in advanced lung adenocarcinoma. Clin. Chem. Lab. Med. 2021, 59, 1221–1229. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.; Tang, J.; Tao, H.; Guo, L.; Wu, W.; Wu, H.; Liu, Z.; Tong, L.; Wu, W.; Li, H.; et al. Detection of EGFR Mutations in Cerebrospinal Fluid of EGFR-Mutant Lung Adenocarcinoma with Brain Metastases. Front. Oncol. 2021, 11, 622142. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Wu, X.; Tang, M.; Nie, X.; Li, L. Detection of EGFR gene mutation status from pleural effusions and other body fluid specimens in patients with lung adenocarcinoma. Thorac. Cancer 2019, 10, 2218–2224. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bettegowda, C.; Sausen, M.; Leary, R.J.; Kinde, I.; Wang, Y.; Agrawal, N.; Bartlett, B.R.; Wang, H.; Luber, B.; Alani, R.M.; et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci. Transl. Med. 2014, 6, 224ra24. [Google Scholar] [CrossRef] [Green Version]
- Wan, L.; Liu, Q.; Liang, D.; Guo, Y.; Liu, G.; Ren, J.; He, Y.; Shan, B. Circulating Tumor Cell and Metabolites as Novel Biomarkers for Early-Stage Lung Cancer Diagnosis. Front. Oncol. 2021, 31, 630672. [Google Scholar] [CrossRef]
- Gerber, T.; Taschner-Mandl, S.; Saloberger-Sindhöringer, L.; Popitsch, N.; Heitzer, E.; Witt, V.; Geyeregger, R.; Hutter, C.; Schwentner, R.; Ambros, I.M.; et al. Assessment of Pre-Analytical Sample Handling Conditions for Comprehensive Liquid Biopsy Analysis. J. Mol. Diagn. 2020, 22, 1070–1086. [Google Scholar] [CrossRef]
- Krug, A.K.; Enderle, D.; Karlovich, C.; Priewasser, T.; Bentink, S.; Spiel, A.; Brinkmann, K.; Emenegger, J.; Grimm, D.G.; Castellanos-Rizaldos, E.; et al. Improved EGFR mutation detection using combined exosomal RNA and circulating tumor DNA in NSCLC patient plasma. Ann. Oncol. 2018, 29, 700–706. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Weber, S.; Spiegl, B.; Perakis, S.O.; Ulz, C.M.; Abuja, P.M.; Kashofer, K.; Leest, P.V.; Azpurua, M.A.; Tamminga, M.; Brudzewsky, D.; et al. Technical Evaluation of Commercial Mutation Analysis Platforms and Reference Materials for Liquid Biopsy Profiling. Cancers 2020, 12, 1588. [Google Scholar] [CrossRef] [PubMed]
- Chaudhuri, A.A.; Chabon, J.J.; Lovejoy, A.F.; Newman, A.M.; Stehr, H.; Azad, T.D.; Khodadoust, M.S.; Esfahani, M.S.; Liu, C.L.; Zhou, L.; et al. Early Detection of Molecular Residual Disease in Localized Lung Cancer by Circulating Tumor DNA Profiling. Cancer Discov. 2017, 7, 1394–1403. [Google Scholar] [CrossRef] [Green Version]
- Amiri, A.; Pourhanifeh, M.H.; Mirzaei, H.R.; Nahand, J.S.; Moghoofei, M.; Sahebnasagh, R.; Mirzaei, H.; Hamblin, M.R. Exosomes and Lung Cancer: Roles in Pathophysiology, Diagnosis and Therapeutic Applications. Curr. Med. Chem. 2021, 28, 308–328. [Google Scholar] [CrossRef] [PubMed]
- Cui, S.; Cheng, Z.; Qin, W.; Jiang, L. Exosomes as a liquid biopsy for lung cancer. Lung Cancer 2018, 116, 46–54. [Google Scholar] [CrossRef] [PubMed]
- Jouida, A.; McCarthy, C.; Fabre, A.; Keane, M.P. Exosomes: A new perspective in EGFR-mutated lung cancer. Cancer Metastasis Rev. 2021, 40, 589–601. [Google Scholar] [CrossRef] [PubMed]
- Yin, L.; Liu, X.; Shao, X.; Feng, T.; Xu, J.; Wang, Q.; Hua, S. The role of exosomes in lung cancer metastasis and clinical applications: An updated review. J. Transl. Med. 2021, 19, 312. [Google Scholar] [CrossRef] [PubMed]
- Radonic, T.; Dickhoff, C.; Mino-Kenudson, M.; Lely, R.; Paul, R.; Thunnissen, E. Gross handling of pulmonary resection specimen: Maintaining the 3-dimensional orientation. J. Thorac. Dis. 2019, 11 (Suppl. S1), S37–S44. [Google Scholar] [CrossRef]
- Do, H.; Molania, R.; Mitchell, P.L.; Vaiskunaite, R.; Murdoch, J.D.; Dobrovic, A. Reducing Artifactual EGFR T790M Mutations in DNA from Formalin-Fixed Paraffin-Embedded Tissue by Use of Thymine-DNA Glycosylase. Clin. Chem. 2017, 63, 1506–1514. [Google Scholar] [CrossRef]
- Cheung, A.H.; Tong, J.H.; Chung, L.Y.; Chau, S.L.; Ng, C.S.; Wan, I.Y.P.; To, K.F. EGFR mutation exists in squamous cell lung carcinoma. Pathology 2020, 52, 323–328. [Google Scholar] [CrossRef]
- Anand, K.; Phung, T.L.; Bernicker, E.H.; Cagle, P.T.; Olsen, R.J.; Thomas, J.S. Clinical Utility of Reflex Ordered Testing for Molecular Biomarkers in Lung Adenocarcinoma. Clin. Lung Cancer 2020, 21, 437–442. [Google Scholar] [CrossRef]
- Cheema, P.K.; Menjak, I.B.; Winterton-Perks, Z.; Raphael, S.; Cheng, S.Y.; Verma, S.; Muinuddin, A.; Freedman, R.; Toor, N.; Perera, J.; et al. Impact of Reflex EGFR/ALK Testing on Time to Treatment of Patients with Advanced Nonsquamous Non-Small-Cell Lung Cancer. J. Oncol. Pract. 2017, 13, e130–e138. [Google Scholar] [CrossRef] [PubMed]
- Cheema, P.K.; Raphael, S.; El-Maraghi, R.; Li, J.; McClure, R.; Zibdawi, L.; Chan, A.; Victor, J.C.; Dolley, A.; Dziarmaga, A. Rate of EGFR mutation testing for patients with nonsquamous non-small-cell lung cancer with implementation of reflex testing by pathologists. Curr. Oncol. 2017, 24, 16–22. [Google Scholar] [CrossRef] [Green Version]
- Ellis, P.M.; Verma, S.; Sehdev, S.; Younus, J.; Leighl, N.B. Challenges to implementation of an epidermal growth factor receptor testing strategy for non-small-cell lung cancer in a publicly funded health care system. J. Thorac. Oncol. 2013, 8, 1136–1141. [Google Scholar] [CrossRef] [Green Version]
- Lisberg, A.; Cummings, A.; Goldman, J.W.; Bornazyan, K.; Reese, N.; Wang, T.; Coluzzi, P.; Ledezma, B.; Mendenhall, M.; Hunt, J.; et al. A Phase II Study of Pembrolizumab in EGFR-Mutant, PD-L1+, Tyrosine Kinase Inhibitor Naïve Patients with Advanced NSCLC. J. Thorac. Oncol. 2018, 13, 1138–1145. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Proto, C.; Ferrara, R.; Signorelli, D.; Lo Russo, G.; Galli, G.; Imbimbo, M.; Prelaj, A.; Zilembo, N.; Ganzinelli, M.; Pallavicini, L.M.; et al. Choosing wisely first line immunotherapy in non-small cell lung cancer (NSCLC): What to add and what to leave out. Cancer Treat. Rev. 2019, 75, 39–51. [Google Scholar] [CrossRef] [PubMed]
- Heeke, S.; Hofman, P. EGFR Mutation Analysis in Non-small Cell Lung Carcinoma from Tissue Samples Using the Fully Automated Idylla™ qPCR System. Methods Mol. Biol. 2019, 2054, 147–155. [Google Scholar] [CrossRef]
- Lassalle, S.; Hofman, V.; Heeke, S.; Benzaquen, J.; Long, E.; Poudenx, M.; Lantéri, E.; Boutros, J.; Tanga, V.; Zahaf, K.; et al. Targeted Assessment of the EGFR Status as Reflex Testing in Treatment-Naive Non-Squamous Cell Lung Carcinoma Patients: A Single Laboratory Experience (LPCE, Nice, France). Cancers 2020, 12, 955. [Google Scholar] [CrossRef] [Green Version]
- Heeke, S.; Benzaquen, J.; Hofman, V.; Ilié, M.; Allegra, M.; Long-Mira, E.; Lassalle, S.; Tanga, V.; Salacroup, C.; Bonnetaud, C.; et al. Critical Assessment in Routine Clinical Practice of Liquid Biopsy for EGFR Status Testing in Non-Small-Cell Lung Cancer: A Single-Laboratory Experience (LPCE, Nice, France). Clin. Lung Cancer 2020, 21, 56–65.e8. [Google Scholar] [CrossRef] [Green Version]
- Passaro, A.; Mok, T.; Peters, S.; Popat, S.; Ahn, M.J.; de Marinis, F. Recent Advances on the Role of EGFR Tyrosine Kinase Inhibitors in the Management of NSCLC With Uncommon, Non Exon 20 Insertions, EGFR Mutations. J. Thorac. Oncol. 2021, 16, 764–773. [Google Scholar] [CrossRef] [PubMed]
- Hastings, K.; Yu, H.A.; Wei, W.; Sanchez-Vega, F.; DeVeaux, M.; Choi, J.; Rizvi, H.; Lisberg, A.; Truini, A.; Lydon, C.A.; et al. EGFR mutation subtypes and response to immune checkpoint blockade treatment in non-small-cell lung cancer. Ann. Oncol. 2019, 30, 1311–1320. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Foggetti, G.; Li, C.; Cai, H.; Hellyer, J.A.; Lin, W.Y.; Ayeni, D.; Hastings, K.; Choi, J.; Wurtz, A.; Andrejka, L.; et al. Genetic Determinants of EGFR-Driven Lung Cancer Growth and Therapeutic Response In Vivo. Cancer Discov. 2021. [CrossRef]
- Kim, I.A.; Hur, J.Y.; Kim, H.J.; Park, J.H.; Hwang, J.J.; Lee, S.A.; Lee, S.E.; Kim, W.S.; Lee, K.Y. Targeted Next-Generation Sequencing Analysis for Recurrence in Early-Stage Lung Adenocarcinoma. Ann. Surg. Oncol. 2021, 28, 3983–3993. [Google Scholar] [CrossRef]
- Leonetti, A.; Sharma, S.; Minari, R.; Perego, P.; Giovannetti, E.; Tiseo, M. Resistance mechanisms to osimertinib in EGFR-mutated non-small cell lung cancer. Br. J. Cancer 2019, 121, 725–737. [Google Scholar] [CrossRef] [PubMed]
- Marcoux, N.; Gettinger, S.N.; O’Kane, G.; Arbour, K.C.; Neal, J.W.; Husain, H.; Evans, T.L.; Brahmer, J.R.; Muzikansky, A.; Bonomi, P.D.; et al. EGFR-Mutant Adenocarcinomas That Transform to Small-Cell Lung Cancer and Other Neuroendocrine Carcinomas: Clinical Outcomes. J. Clin. Oncol. 2019, 37, 278–285. [Google Scholar] [CrossRef] [PubMed]
- Skoulidis, F.; Heymach, J.V. Co-occurring genomic alterations in non-small-cell lung cancer biology and therapy. Nat. Rev. Cancer 2019, 19, 495–509. [Google Scholar] [CrossRef] [PubMed]
- Blakely, C.M.; Watkins, T.B.K.; Wu, W.; Gini, B.; Chabon, J.J.; McCoach, C.E.; McGranahan, N.; Wilson, G.A.; Birkbak, N.J.; Olivas, V.R.; et al. Evolution and clinical impact of co-occurring genetic alterations in advanced-stage EGFR-mutant lung cancers. Nat. Genet. 2017, 49, 1693–1704. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Xu, Y.; Zhao, J.; Zhong, W.; Zhang, L.; Bi, Y.; Wang, M. Concurrent Driver Gene Mutations as Negative Predictive Factors in Epidermal Growth Factor Receptor-Positive Non-Small Cell Lung Cancer. EBioMedicine 2019, 42, 304–310. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jung, J.H.; Lee, H.; Zeng, S.X.; Lu, H. RBM10, a New Regulator of p53. Cells 2020, 9, 2107. [Google Scholar] [CrossRef]
- Karachaliou, N.; Paulina Bracht, J.W.; Rosell, R. ARID1A Gene Driver Mutations in Lung Adenocarcinomas. J. Thorac. Oncol. 2018, 13, e255–e257. [Google Scholar] [CrossRef] [Green Version]
- Yu, H.A.; Suzawa, K.; Jordan, E.; Zehir, A.; Ni, A.; Kim, R.; Kris, M.G.; Hellmann, M.D.; Li, B.T.; Somwar, R.; et al. Concurrent Alterations in EGFR-Mutant Lung Cancers Associated with Resistance to EGFR Kinase Inhibitors and Characterization of MTOR as a Mediator of Resistance. Clin. Cancer Res. 2018, 24, 3108–3118. [Google Scholar] [CrossRef] [Green Version]
- Friedlaender, A.; Addeo, A.; Russo, A.; Gregorc, V.; Cortinovis, D.; Rolfo, C.D. Targeted Therapies in Early Stage NSCLC: Hype or Hope? Int. J. Mol. Sci. 2020, 21, 6329. [Google Scholar] [CrossRef]
- Bruno, R.; Fontanini, G. Next Generation Sequencing for Gene Fusion Analysis in Lung Cancer: A Literature Review. Diagnostics 2020, 10, 521. [Google Scholar] [CrossRef]
- Hofman, P.; Rouleau, E.; Sabourin, J.C.; Denis, M.; Deleuze, J.F.; Barlesi, F.; Laurent-Puig, P. Predictive molecular pathology in non-small cell lung cancer in France: The past, the present and the perspectives. Cancer Cytopathol. 2020, 128, 601–610. [Google Scholar] [CrossRef] [PubMed]
- Sauter, J.L.; Butnor, K.J. Clinical and Cost Implications of Universal Versus Locally Advanced-Stage and Advanced-Stage-Only Molecular Testing for Epidermal Growth Factor Receptor Mutations and Anaplastic Lymphoma Kinase Rearrangements in Non- Small Cell Lung Carcinoma: A Tertiary Academic Institution Experience. Arch. Pathol. Lab. Med. 2016, 140, 358–361. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thunnissen, E.; Weynand, B.; Udovicic-Gagula, D.; Brcic, L.; Szolkowska, M.; Hofman, P.; Smojver-Ježek, S.; Anttila, S.; Calabrese, F.; Kern, I.; et al. Lung cancer biomarker testing: Perspective from Europe. Transl. Lung Cancer Res. 2020, 9, 887–897. [Google Scholar] [CrossRef] [PubMed]
- Mehrad, M.; Roy, S.; Bittar, H.T.; Dacic, S. Next-Generation Sequencing Approach to Non-Small Cell Lung Carcinoma Yields More Actionable Alterations. Arch. Pathol. Lab. Med. 2018, 142, 353–357. [Google Scholar] [CrossRef] [Green Version]
- Pennell, N.A.; Mutebi, A.; Zhou, Z.-H. Economic impact of next-generation sequencing versus single-gene testing to detect geneomic alterations in metastatic non-small cell lung cancer using a decision analytic model. JCO Precis. Oncol. 2019, 3, 1–9. [Google Scholar] [CrossRef]
- Pruneri, G.; De Braud, F.; Sapino, A.; Aglietta, M.; Vecchione, A.; Giusti, R.; Marchiò, C.; Scarpino, S.; Baggi, A.; Bonetti, G.; et al. Next-Generation Sequencing in Clinical Practice: Is It a Cost-Saving Alternative to a Single-Gene Testing Approach? Pharmacoecon. Open 2021, 5, 285–298. [Google Scholar] [CrossRef]
- Graham, R.P.; Treece, A.L.; Lindeman, N.I.; Vasalos, P.; Shan, M.; Jennings, L.J.; Rimm, D.L. Worldwide Frequency of Commonly Detected EGFR Mutations. Arch. Pathol. Lab. Med. 2018, 142, 163–167. [Google Scholar] [CrossRef] [Green Version]
- Li, W.; Guo, H.; Li, L.; Cui, J. Comprehensive Comparison Between Adjuvant Targeted Therapy and Chemotherapy for EGFR-Mutant NSCLC Patients: A Cost-Effectiveness Analysis. Front. Oncol. 2021, 11, 619376. [Google Scholar] [CrossRef] [PubMed]
- Yotsukura, M.; Nakagawa, K.; Suzuki, K.; Takamochi, K.; Ito, H.; Okami, J.; Aokage, K.; Shiono, S.; Yoshioka, H.; Aoki, T.; et al. Recent advances and future perspectives in adjuvant and neoadjuvant immunotherapies for lung cancer. Jpn. J. Clin. Oncol. 2021, 51, 28–36. [Google Scholar] [CrossRef]
- Ilié, M.; Ngo-Mai, M.; Long-Mira, E.; Lassalle, S.; Butori, C.; Bence, C.; Hamila, M.; Hofman, V.; Hofman, P. Using 22C3 Anti-PD-L1 Antibody Concentrate on Biopsy and Cytology Samples from Non-small Cell Lung Cancer Patients. J. Vis. Exp. 2018, 139, 58082. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ilie, M.; Juco, J.; Huang, L.; Hofman, V.; Khambata-Ford, S.; Hofman, P. Use of the 22C3 anti-programmed death-ligand 1 antibody to determine programmed death-ligand 1 expression in cytology samples obtained from non-small cell lung cancer patients. Cancer Cytopathol. 2018, 126, 264–274. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tejerina, E.; Garca Tobar, L.; Echeveste, J.I.; de Andrea, C.E.; Vigliar, E.; Lozano, M.D. PD-L1 in Cytological Samples: A Review and a Practical Approach. Front. Med. 2021, 8, 668612. [Google Scholar] [CrossRef]
- Qiao, M.; Jiang, T.; Liu, X.; Mao, S.; Zhou, F.; Li, X.; Zhao, C.; Chen, X.; Su, C.; Ren, S.; et al. Immune Checkpoint Inhibitors in EGFR-Mutated NSCLC: Dusk or Dawn? J. Thorac. Oncol. 2021, 16, 1267–1288. [Google Scholar] [CrossRef] [PubMed]
- Kneuertz, P.J.; Carbone, D.P.; D’Souza, D.M.; Shilo, K.; Abdel-Rasoul, M.; Zhao, W.; Williams, T.M.; Jones, D.; Merritt, R.E. Prognostic value and therapeutic implications of expanded molecular testing for resected early stage lung adenocarcinoma. Lung Cancer 2020, 143, 60–66. [Google Scholar] [CrossRef]
- Ilie, M.; Long-Mira, E.; Bence, C.; Butori, C.; Lassalle, S.; Bouhlel, L.; Fazzalari, L.; Zahaf, K.; Lalvée, S.; Washetine, K.; et al. Comparative study of the PD-L1 status between surgically resected specimens and matched biopsies of NSCLC patients reveal major discordances: A potential issue for anti-PD-L1 therapeutic strategies. Ann. Oncol. 2016, 27, 147–153. [Google Scholar] [CrossRef] [PubMed]
- Mansfield, A.S.; Dong, H. Implications of Programmed Cell Death 1 Ligand 1 Heterogeneity in the Selection of Patients With Non-Small Cell Lung Cancer to Receive Immunotherapy. Clin. Pharmacol. Ther. 2016, 100, 220–222. [Google Scholar] [CrossRef] [PubMed]
- Dong, Z.Y.; Zhang, J.T.; Liu, S.Y.; Su, J.; Zhang, C.; Xie, Z.; Zhou, Q.; Tu, H.Y.; Xu, C.R.; Yan, L.X.; et al. EGFR mutation correlates with uninflamed phenotype and weak immunogenicity, causing impaired response to PD-1 blockade in non-small cell lung cancer. Oncoimmunology 2017, 6, e1356145. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- To, K.K.W.; Fong, W.; Cho, W.C.S. Immunotherapy in Treating EGFR-Mutant Lung Cancer: Current Challenges and New Strategies. Front. Oncol. 2021, 11, 635007. [Google Scholar] [CrossRef]
- Offin, M.; Rizvi, H.; Tenet, M.; Ni, A.; Sanchez-Vega, F.; Li, B.T.; Drilon, A.; Kris, M.G.; Rudin, C.M.; Schultz, N. Tumor Mutation Burden and Efficacy of EGFR-Tyrosine Kinase Inhibitors in Patients with EGFR-Mutant Lung Cancers. Clin. Cancer Res. 2019, 25, 1063–1069. [Google Scholar] [CrossRef] [Green Version]
- Smeltzer, M.P.; Wynes, M.W.; Lantuejoul, S.; Soo, R.; Ramalingam, S.S.; Varella-Garcia, M.; Meadows Taylor, M.; Richeimer, K.; Wood, K.; Howell, K.E.; et al. The International Association for the Study of Lung Cancer Global Survey on Molecular Testing in Lung Cancer. J. Thorac. Oncol. 2020, 15, 1434–1448. [Google Scholar] [CrossRef]
- Mehta, A.; Vasudevan, S. Rare epidermal growth factor receptor gene alterations in non-small cell lung cancer patients, tyrosine kinase inhibitor response and outcome analysis. Cancer Treat. Res. Commun. 2021, 28, 100398. [Google Scholar] [CrossRef]
- Volckmar, A.L.; Christopoulos, P.; Kirchner, M.; Allgäuer, M.; Neumann, O.; Budczies, J.; Rempel, E.; Horak, P.; Glade, J.; Goldschmid, H.; et al. Targeting rare and non-canonical driver variants in NSCLC-An uncharted clinical field. Lung Cancer 2021, 154, 131–141. [Google Scholar] [CrossRef] [PubMed]
- Nagasaka, M.; Zhu, V.W.; Lim, S.M.; Greco, M.; Wu, F.; Ou, S.I. Beyond Osimertinib: The Development of Third-Generation EGFR Tyrosine Kinase Inhibitors For Advanced EGFR+ NSCLC. J. Thorac. Oncol. 2021, 16, 740–763. [Google Scholar] [CrossRef]
- Nagasaka, M.; Uddin, M.H.; Al-Hallak, M.N.; Rahman, S.; Balasubramanian, S.; Sukari, A.; Azmi, A.S. Liquid biopsy for therapy monitoring in early-stage non-small cell lung cancer. Mol. Cancer 2021, 20, 82. [Google Scholar] [CrossRef]
- Peng, M.; Huang, Q.; Yin, W.; Tan, S.; Chen, C.; Liu, W.; Tang, J.; Wang, X.; Zhang, B.; Zou, M.; et al. Circulating Tumor DNA as a Prognostic Biomarker in Localized Non-small Cell Lung Cancer. Front. Oncol. 2020, 10, 561598. [Google Scholar] [CrossRef] [PubMed]
- Suda, K.; Mitsudomi, T.; Shintani, Y.; Okami, J.; Ito, H.; Ohtsuka, T.; Toyooka, S.; Mori, T.; Watanabe, S.I.; Asamura, H.; et al. Clinical Impacts of EGFR Mutation Status: Analysis of 5780 Surgically Resected Lung Cancer Cases. Ann. Thorac. Surg. 2021, 111, 269–276. [Google Scholar] [CrossRef]
- Pei, G.; Li, M.; Min, X.; Liu, Q.; Li, D.; Yang, Y.; Wang, S.; Wang, X.; Wang, H.; Cheng, H.; et al. Molecular Identification and Genetic Characterization of Early-Stage Multiple Primary Lung Cancer by Large-Panel Next-Generation Sequencing Analysis. Front. Oncol. 2021, 11, 653988. [Google Scholar] [CrossRef]
- Fligor, S.C.; Tsikis, S.T.; Wang, S.; Ore, A.S.; Allar, B.G.; Whitlock, A.E.; Calvillo-Ortiz, R.; Arndt, K.; Callery, M.P.; Gangadharan, S.P. Time to surgery in thoracic cancers and prioritization during COVID-19: A systematic review. J. Thorac. Dis. 2020, 12, 6640–6654. [Google Scholar] [CrossRef] [PubMed]
- Hofman, P.; Ilié, M.; Chamorey, E.; Brest, P.; Schiappa, R.; Nakache, V.; Antoine, M.; Barberis, M.; Begueret, H.; Bibeau, F.; et al. Clinical and molecular practice of European thoracic pathology laboratories during the COVID-19 pandemic. The past and the near future. ESMO Open. 2021, 6, 100024. [Google Scholar] [CrossRef] [PubMed]
- Johnson, B.A.; Waddimba, A.C.; Ogola, G.O.; Fleshman, J.W., Jr.; Preskitt, J.T. A systematic review and meta-analysis of surgery delays and survival in breast, lung and colon cancers: Implication for surgical triage during the COVID-19 pandemic. Am. J. Surg. 2021, 222, 311–318. [Google Scholar] [CrossRef] [PubMed]
- Malapelle, U.; Pisapia, P.; Iaccarino, A.; Barberis, M.; Bellevicine, C.; Brunnström, H.; de Biase, D.; De Maglio, G.; Ericson Lindquist, K.; Fassan, M.; et al. Predictive molecular pathology in the time of coronavirus disease (COVID-19) in Europe. J. Clin. Pathol. 2020, 74, 391–395. [Google Scholar] [CrossRef] [PubMed]
- Patton, S.; Normanno, N.; Blackhall, F.; Murray, S.; Kerr, K.M.; Dietel, M.; Filipits, M.; Benlloch, S.; Popat, S.; Stahel, R.; et al. Assessing standardization of molecular testing for non-small-cell lung cancer: Results of a worldwide external quality assessment (EQA) scheme for EGFR mutation testing. Br. J. Cancer 2014, 111, 413–420. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bank, P.C.D.; Jacobs, L.H.J.; van den Berg, S.A.A.; van Deutekom, H.W.M.; Hamann, D.; Molenkamp, R.; Ruivenkamp, C.A.L.; Swen, J.J.; Tops, B.B.J.; Wamelink, M.M.C.; et al. The end of the laboratory developed test as we know it? Recommendations from a national multidisciplinary taskforce of laboratory specialists on the interpretation of the IVDR and its complications. Clin. Chem. Lab. Med. 2020, 59, 491–497. [Google Scholar] [CrossRef] [PubMed]
- Barberis, M. In vitro diagnostic medical device regulation (IVDR): The end of laboratory developed tests (LDT)? Pathologica 2021, 113, 68–69. [Google Scholar] [CrossRef] [PubMed]
- Vogeser, M.; Brüggemann, M. Complex analytical procedures in diagnostic laboratories and the IVDR. Clin. Chem. Lab. Med. 2020, 59, 457–458. [Google Scholar] [CrossRef]
- Han, J.; Yu, R.; Duan, J.; Li, J.; Zhao, W.; Feng, G.; Bai, H.; Wang, Y.; Zhang, X.; Wan, R.; et al. Weighting tumor-specific TCR repertoires as a classifier to stratify the immunotherapy delivery in non-small cell lung cancers. Sci. Adv. 2021, 7, eabd6971. [Google Scholar] [CrossRef]
- Hofman, P.; Heeke, S.; Alix-Panabières, C.; Pantel, K. Liquid biopsy in the era of immuno-oncology: Is it ready for prime-time use for cancer patients? Ann. Oncol. 2019, 30, 1448–1459. [Google Scholar] [CrossRef] [Green Version]
- Qiu, Y.; Liu, L.; Yang, H.; Chen, H.; Deng, Q.; Xiao, D.; Lin, Y.; Zhu, C.; Li, W.; Shao, D.; et al. Integrating Histologic and Genomic Characteristics to Predict Tumor Mutation Burden of Early-Stage Non-Small-Cell Lung Cancer. Front. Oncol. 2021, 10, 608989. [Google Scholar] [CrossRef]
- Yang, H.; Wang, Y.; Jia, Z.; Wang, Y.; Yang, X.; Wu, P.; Song, Y.; Xu, H.; Gu, D.; Chen, R.; et al. Characteristics of T-Cell Receptor Repertoire and Correlation with EGFR Mutations in All Stages of Lung Cancer. Front. Oncol. 2021, 11, 537735. [Google Scholar] [CrossRef]
- Yu, H.; Chen, Z.; Ballman, K.V.; Watson, M.A.; Govindan, R.; Lanc, I.; Beer, D.G.; Bueno, R.; Chirieac, L.R.; Chui, M.H.; et al. Correlation of PD-L1 Expression with Tumor Mutation Burden and Gene Signatures for Prognosis in Early-Stage Squamous Cell Lung Carcinoma. J. Thorac. Oncol. 2019, 14, 25–36. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stenzinger, A.; Alber, M.; Allgäuer, M.; Jurmeister, P.; Bockmayr, M.; Budczies, J.; Lennerz, J.; Eschrich, J.; Kazdal, D.; Schirmacher, P.; et al. Artificial intelligence and pathology: From principles to practice and future applications in histomorphology and molecular profiling. Semin. Cancer Biol. 2021. [CrossRef]

| Should we only look for the EGFR status on pre-operative biopsies (bronchial and transthoracic biopsies)? |
| Should we only look for the EGFR status on surgically resected specimens? |
| Should we look for the EGFR status systematically on both biopsies and surgically resected specimens? |
| Is it acceptable to look for the EGFR status on cytological samples only? |
| What is the added value of integrating a liquid biopsy before and/or after surgery for early stage NSCLC patients? |
| Does the assessment of EGFR have to be done only by targeted (RT-PCR) or next generation sequencing technologies? |
| What about the systematic evaluation of the other predictive biomarkers on biopsies (PD-L1, ALK, and other genes)? |
| How can we master the turnaround time in obtaining results, notably in the context of neoadjuvant immunotherapy versus adjuvant EGFR TKIs administration? |
| How should we consider the landscape of concomitant mutation (notably in TP53, RB, RBM10, CTNNB1, FAT1, ABCB1, ARID1A) in EGFR 19 del19/L858R mutated tumors and their association with response to EGFR-TKIs in early stage NSCLC patients? |
| How should we consider EGFR mutations out of EGFR 19 del19/L858R? |
| How should we take care of patients with early stage squamous cell lung carcinoma with an EGFR mutation? |
| How should we reimburse the molecular biology reflex tests and receive coverage for the full cost? |
| How should we integrate the induced workload and new infrastructural challenges in the pathology laboratories? |
| How should we anticipate the issues associated with the development of the In Vitro Diagnostic Regulation in Europe? |
| Should we assess the ct-DNA level before and/or after surgery? |
| What is the best timing for blood sampling after complete surgery? |
| How should we define a cutoff for the ct-DNA level after complete surgery? |
| How should we set up a liquid biopsy in new daily practice? |
| How should we determine false and true negative results for EGFR mutation in a liquid biopsy? |
| How should we determine false positive results for EGFR mutation in a liquid biopsy? |
| What are the best practices for mastering the different pre-analytical phases? |
| How should we distinguish cf-DNA and ct-DNA in routine clinical practice? |
| How should we set up blood methods for standardization and validation of molecular biology testing? |
| How should we obtain a mandatory accreditation (such as the ISO 15,189 accreditation in Europe)? |
| How should we reimburse the liquid biopsy in the health care system? |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hofman, P. EGFR Status Assessment for Better Care of Early Stage Non-Small Cell Lung Carcinoma: What Is Changing in the Daily Practice of Pathologists? Cells 2021, 10, 2157. https://doi.org/10.3390/cells10082157
Hofman P. EGFR Status Assessment for Better Care of Early Stage Non-Small Cell Lung Carcinoma: What Is Changing in the Daily Practice of Pathologists? Cells. 2021; 10(8):2157. https://doi.org/10.3390/cells10082157
Chicago/Turabian StyleHofman, Paul. 2021. "EGFR Status Assessment for Better Care of Early Stage Non-Small Cell Lung Carcinoma: What Is Changing in the Daily Practice of Pathologists?" Cells 10, no. 8: 2157. https://doi.org/10.3390/cells10082157
APA StyleHofman, P. (2021). EGFR Status Assessment for Better Care of Early Stage Non-Small Cell Lung Carcinoma: What Is Changing in the Daily Practice of Pathologists? Cells, 10(8), 2157. https://doi.org/10.3390/cells10082157






