Post-Transcriptional Regulation of Viral RNA through Epitranscriptional Modification
Abstract
1. Introduction
| Virus | Genome | m6A | m5C | ψ | m1A | ac4C | Nm | m7G | m1G | m6,6A |
|---|---|---|---|---|---|---|---|---|---|---|
| Adenovirus serotype 5 | DNA | [18] | ||||||||
| Dengue virus | RNA | [7,16] | [7] | [7] | [7] | [7] | [7] | [7] | [7] | [7] |
| Enterovirus 71 | RNA | [32] | ||||||||
| Epstein–Barr virus | DNA | [33] | ||||||||
| Hepatitis B virus | DNA | [8,34] | ||||||||
| Hepatitis C virus | RNA | [7,16,34] | [7] | [7] | [7] | [7] | [7] | [7] | [7] | [7] |
| HIV-1 | RNA | [6,7,9,10,14,35] | [6,7,36] | [7] | [6,7] | [7,11] | [6,7,15] | [6,7] | [6,7] | [6,7] |
| Human metapneumovirus | RNA | [37] | ||||||||
| Influenza A virus | RNA | [12,38,39] | ||||||||
| Kaposi’s sarcoma-associated herpesvirus | DNA | [40,41] | ||||||||
| Measles virus | RNA | [42] | ||||||||
| Murine leukaemia virus | RNA | [5] | [5,43] | [5] | [5] | [5] | [5] | [5] | [5] | |
| Poliovirus | RNA | [7] | [7] | [7] | [7] | [7] | [7] | [7] | [7] | [7] |
| Respiratory syncytial virus | RNA | [44] | ||||||||
| SARS-CoV-2 | RNA | [45] | [46] | |||||||
| Sendai virus | RNA | [42] | ||||||||
| Simian virus 40 | DNA | [13] | ||||||||
| Vesicular stomatitis virus | RNA | [42] | ||||||||
| West Nile virus | RNA | [16] | ||||||||
| Yellow fever virus | RNA | [16] | ||||||||
| Zika virus | RNA | [7,16,17] | [7] | [7] | [7] | [7] | [7] | [7] | [7] | [7] |
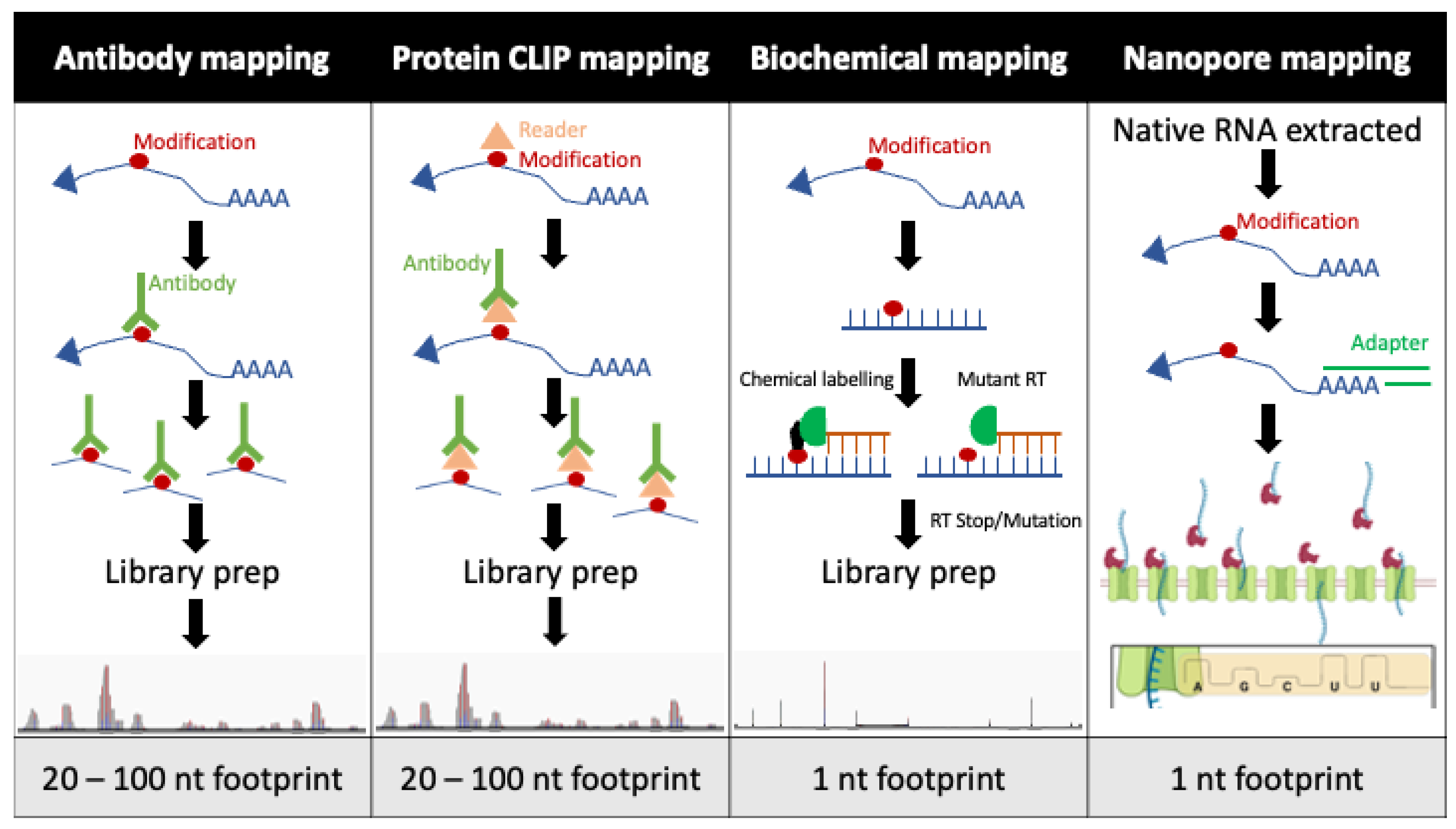
2. Viral Modification Mapping
2.1. Antibody Mapping
2.2. Protein CLIP Mapping
2.3. Biochemical Mapping
2.4. Nanopore Mapping
3. Viral RNA Trafficking
4. Degradation of Viral RNA
5. Splicing of Viral RNA
6. Immune Evasion by Viral RNA
7. Future Avenues of Research
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Li, X.; Xiong, X.; Yi, C. Epitranscriptome Sequencing Technologies: Decoding RNA Modifications. Nat. Methods 2016, 14, 23–31. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.-S.; Liu, C.; Ma, H.; Dai, Q.; Sun, H.-L.; Luo, G.; Zhang, Z.; Zhang, L.; Hu, L.; Dong, X.; et al. Transcriptome-Wide Mapping of Internal N7-Methylguanosine Methylome in Mammalian MRNA. Mol. Cell 2019, 1–13. [Google Scholar] [CrossRef]
- Dominissini, D.; Eyal, E.; Hershkovitz, V.; Salmon-Divon, M.; Clark, W.C.; Dai, Q.; Ben-Haim, M.S.; Solomon, O.; Amariglio, N.; Rechavi, G.; et al. The Dynamic N1-Methyladenosine Methylome in Eukaryotic Messenger RNA. Nature 2016, 530, 441–446. [Google Scholar] [CrossRef]
- Suzuki, T.; Ito, S.; Horikawa, S.; Suzuki, T.; Kawauchi, H.; Tanaka, Y.; Suzuki, T. Human NAT10 Is an ATP-Dependent Rna Acetyltransferase Responsible for N4-Acetylcytidine Formation in 18 S Ribosomal RNA (RRNA). J. Biol. Chem. 2014, 289, 35724–35730. [Google Scholar] [CrossRef]
- Courtney, D.G.; Chalem, A.; Bogerd, H.P.; Law, B.A.; Kennedy, E.M.; Holley, C.L.; Cullen, B.R. Extensive Epitranscriptomic Methylation of A and C Residues on Murine Leukemia Virus Transcripts Enhances Viral Gene Expression. mBio 2019, 10, e01209-19. [Google Scholar] [CrossRef]
- Courtney, D.G.; Tsai, K.; Bogerd, H.P.; Kennedy, E.M.; Law, B.A.; Emery, A.; Swanstrom, R.; Holley, C.L.; Cullen, B.R. Epitranscriptomic Addition of M5C to HIV-1 Transcripts Regulates Viral Gene Expression. Cell Host Microbe 2019, 26, 217–227. [Google Scholar] [CrossRef]
- McIntyre, W.; Netzband, R.; Bonenfant, G.; Biegel, J.M.; Miller, C.; Fuchs, G.; Henderson, E.; Arra, M.; Canki, M.; Fabris, D.; et al. Positive-Sense RNA Viruses Reveal the Complexity and Dynamics of the Cellular and Viral Epitranscriptomes during Infection. Nucleic Acids Res. 2018, 46, 5776–5791. [Google Scholar] [CrossRef]
- Imam, H.; Khan, M.; Gokhale, N.S.; McIntyre, A.B.R.; Kim, G.-W.; Jang, J.Y.; Kim, S.-J.; Mason, C.E.; Horner, S.M.; Siddiqui, A. N6-Methyladenosine Modification of Hepatitis B Virus RNA Differentially Regulates the Viral Life Cycle. Proc. Natl. Acad. Sci. USA 2018, 115, 201808319. [Google Scholar] [CrossRef] [PubMed]
- Lichinchi, G.; Gao, S.; Saletore, Y.; Gonzalez, G.M.; Bansal, V.; Wang, Y.; Mason, C.E.; Rana, T.M. Dynamics of the Human and Viral M6A RNA Methylomes during HIV-1 Infection of T Cells. Nat. Microbiol. 2016, 1, 16011. [Google Scholar] [CrossRef] [PubMed]
- Kennedy, E.M.; Bogerd, H.P.; Kornepati, A.V.R.; Kang, D.; Ghoshal, D.; Marshall, J.B.; Poling, B.C.; Tsai, K.; Gokhale, N.S.; Horner, S.M.; et al. Posttranscriptional M6A Editing of HIV-1 MRNAs Enhances Viral Gene Expression. Cell Host Microbe 2016, 19, 675–685. [Google Scholar] [CrossRef]
- Tsai, K.; Jaguva Vasudevan, A.A.; Martinez Campos, C.; Emery, A.; Swanstrom, R.; Cullen, B.R. Acetylation of Cytidine Residues Boosts HIV-1 Gene Expression by Increasing Viral RNA Stability. Cell Host Microbe 2020. [Google Scholar] [CrossRef] [PubMed]
- Courtney, D.G.; Kennedy, E.M.; Dumm, R.E.; Bogerd, H.P.; Tsai, K.; Heaton, N.S.; Cullen, B.R. Epitranscriptomic Enhancement of Influenza A Virus Gene Expression and Replication. Cell Host Microbe 2017, 22, 377–386. [Google Scholar] [CrossRef]
- Tsai, K.; Courtney, D.G.; Cullen, B.R. Addition of M6A to SV40 Late MRNAs Enhances Viral Structural Gene Expression and Replication. PLoS Pathog. 2018, 14, e1006919. [Google Scholar] [CrossRef] [PubMed]
- Tirumuru, N.; Simen Zhao, B.; Lu, W.; Lu, Z.; He, C.; Wu, L.; Zhao, B.S.; Lu, W.; Lu, Z.; He, C.; et al. N6-Methyladenosine of HIV-1 RNA Regulates Viral Infection and HIV-1 Gag Protein Expression. eLife 2016, 5, e15528. [Google Scholar] [CrossRef]
- Ringeard, M.; Marchand, V.; Decroly, E.; Motorin, Y.; Bennasser, Y. FTSJ3 Is an RNA 2′-O-Methyltransferase Recruited by HIV to Avoid Innate Immune Sensing. Nature 2019, 565, 500–504. [Google Scholar] [CrossRef] [PubMed]
- Gokhale, N.S.; McIntyre, A.B.R.; McFadden, M.J.; Roder, A.E.; Kennedy, E.M.; Gandara, J.A.; Hopcraft, S.E.; Quicke, K.M.; Vazquez, C.; Willer, J.; et al. N6-Methyladenosine in Flaviviridae Viral RNA Genomes Regulates Infection. Cell Host Microbe 2016, 20, 654–665. [Google Scholar] [CrossRef]
- Lichinchi, G.; Zhao, B.S.; Wu, Y.; Lu, Z.; Qin, Y.; He, C.; Rana, T.M. Dynamics of Human and Viral RNA Methylation during Zika Virus Infection. Cell Host Microbe 2016, 20, 666–673. [Google Scholar] [CrossRef] [PubMed]
- Price, A.M.; Hayer, K.E.; McIntyre, A.B.R.; Gokhale, N.S.; Abebe, J.S.; della Fera, A.N.; Mason, C.E.; Horner, S.M.; Wilson, A.C.; Depledge, D.P.; et al. Direct RNA Sequencing Reveals M6A Modifications on Adenovirus RNA Are Necessary for Efficient Splicing. Nat. Commun. 2020, 11, 1–17. [Google Scholar] [CrossRef]
- Roundtree, I.A.; Luo, G.Z.; Zhang, Z.; Wang, X.; Zhou, T.; Cui, Y.; Sha, J.; Huang, X.; Guerrero, L.; Xie, P.; et al. YTHDC1 Mediates Nuclear Export of N6-Methyladenosine Methylated MRNAs. eLife 2017. [Google Scholar] [CrossRef] [PubMed]
- Wojtas, M.N.; Pandey, R.R.; Mendel, M.; Homolka, D.; Sachidanandam, R.; Pillai, R.S. Regulation of M6A Transcripts by the 3ʹ→5ʹ RNA Helicase YTHDC2 Is Essential for a Successful Meiotic Program in the Mammalian Germline. Mol. Cell 2017. [Google Scholar] [CrossRef]
- Amort, T.; Rieder, D.; Wille, A.; Khokhlova-Cubberley, D.; Riml, C.; Trixl, L.; Jia, X.Y.; Micura, R.; Lusser, A. Distinct 5-Methylcytosine Profiles in Poly(A) RNA from Mouse Embryonic Stem Cells and Brain. Genome Biol. 2017. [Google Scholar] [CrossRef] [PubMed]
- Choe, J.; Lin, S.; Zhang, W.; Liu, Q.; Wang, L.; Ramirez-Moya, J.; Du, P.; Kim, W.; Tang, S.; Sliz, P.; et al. MRNA Circularization by METTL3–EIF3h Enhances Translation and Promotes Oncogenesis. Nature 2018, 561, 556–560. [Google Scholar] [CrossRef] [PubMed]
- Arango, D.; Sturgill, D.; Alhusaini, N.; Dillman, A.A.; Sweet, T.J.; Hanson, G.; Hosogane, M.; Sinclair, W.R.; Nanan, K.K.; Mandler, M.D.; et al. Acetylation of Cytidine in MRNA Promotes Translation Efficiency. Cell 2018, 1–15. [Google Scholar] [CrossRef]
- Coots, R.A.; Liu, X.M.; Mao, Y.; Dong, L.; Zhou, J.; Wan, J.; Zhang, X.; Qian, S.B. M6A Facilitates EIF4F-Independent MRNA Translation. Mol. Cell 2017. [Google Scholar] [CrossRef]
- Meyer, K.D.; Patil, D.P.; Zhou, J.; Zinoviev, A.; Skabkin, M.A.; Elemento, O.; Pestova, T.V.; Qian, S.B.; Jaffrey, S.R. 5′ UTR M6A Promotes Cap-Independent Translation. Cell 2015, 163, 999–1010. [Google Scholar] [CrossRef]
- Wang, X.; Zhao, B.S.; Roundtree, I.A.; Lu, Z.; Han, D.; Ma, H.; Weng, X.; Chen, K.; Shi, H.; He, C. N6-Methyladenosine Modulates Messenger RNA Translation Efficiency. Cell 2015. [Google Scholar] [CrossRef]
- Lin, S.; Choe, J.; Du, P.; Triboulet, R.; Gregory, R.I. The M6A Methyltransferase METTL3 Promotes Translation in Human Cancer Cells. Mol. Cell 2016, 62, 335–345. [Google Scholar] [CrossRef]
- Yang, X.; Yang, Y.; Sun, B.F.; Chen, Y.S.; Xu, J.W.; Lai, W.Y.; Li, A.; Wang, X.; Bhattarai, D.P.; Xiao, W.; et al. 5-Methylcytosine Promotes MRNA Export-NSUN2 as the Methyltransferase and ALYREF as an m 5 C Reader. Cell Res. 2017, 27, 606–625. [Google Scholar] [CrossRef]
- Schwartz, S.; Mumbach, M.R.; Jovanovic, M.; Wang, T.; Maciag, K.; Bushkin, G.G.; Mertins, P.; Ter-Ovanesyan, D.; Habib, N.; Cacchiarelli, D.; et al. Perturbation of M6A Writers Reveals Two Distinct Classes of MRNA Methylation at Internal and 5′ Sites. Cell Rep. 2014. [Google Scholar] [CrossRef]
- Wang, X.; Lu, Z.; Gomez, A.; Hon, G.C.; Yue, Y.; Han, D.; Fu, Y.; Parisien, M.; Dai, Q.; Jia, G.; et al. N 6-Methyladenosine-Dependent Regulation of Messenger RNA Stability. Nature 2014. [Google Scholar] [CrossRef] [PubMed]
- Woo, H.-H.; Chambers, S.K. Human ALKBH3-Induced M1A Demethylation Increases the CSF-1 MRNA Stability in Breast and Ovarian Cancer Cells. Biochim. Et Biophys. Acta (Bba) - Gene Regul. Mech. 2018, 1862, 35–46. [Google Scholar] [CrossRef]
- Yao, M.; Dong, Y.; Wang, Y.; Liu, H.; Ma, H.; Zhang, H.; Zhang, L.; Cheng, L.; Lv, X.; Xu, Z.; et al. N6-Methyladenosine Modifications Enhance Enterovirus 71 ORF Translation through METTL3 Cytoplasmic Distribution. Biochem. Biophys. Res. Commun. 2020, 527, 297–304. [Google Scholar] [CrossRef]
- Zheng, X.; Wang, J.; Zhang, X.; Fu, Y.; Peng, Q.; Lu, J.; Wei, L.; Li, Z.; Liu, C.; Wu, Y.; et al. RNA M6 A Methylation Regulates Virus-Host Interaction and EBNA2 Expression during Epstein-Barr Virus Infection. Immun. Inflamm. Dis. 2021. [Google Scholar] [CrossRef]
- Kim, G.W.; Imam, H.; Khan, M.; Siddiqui, A. N6-Methyladenosine Modification of Hepatitis B and C Viral RNAs Attenuates Host Innate Immunity via RIG-I Signaling. J. Biol. Chem. 2020, 295, 13123–13133. [Google Scholar] [CrossRef]
- Chen, S.; Kumar, S.; Espada, C.E.; Tirumuru, N.; Cahill, M.P.; Hu, L.; He, C.; Wu, L. N6-Methyladenosine Modification of HIV-1 RNA Suppresses Type-I Interferon Induction in Differentiated Monocytic Cells and Primary Macrophages. PLoS Pathog. 2021, 17, e1009421. [Google Scholar] [CrossRef] [PubMed]
- Kong, W.; Biswas, A.; Zhou, D.; Fiches, G.; Fujinaga, K.; Santoso, N.; Zhu, J. Nucleolar Protein NOP2/NSUN1 Suppresses HIV-1 Transcription and Promotes Viral Latency by Competing with Tat for TAR Binding and Methylation. PLoS Pathog. 2020, 16, e1008430. [Google Scholar] [CrossRef] [PubMed]
- Lu, M.; Zhang, Z.; Xue, M.; Zhao, B.S.; Harder, O.; Li, A.; Liang, X.; Gao, T.Z.; Xu, Y.; Zhou, J.; et al. N6-Methyladenosine Modification Enables Viral RNA to Escape Recognition by RNA Sensor RIG-I. Nat. Microbiol. 2020, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Narayan, P.; Ayers, D.F.; Rottman, F.M.; Maroney, P.A.; Nilsen, T.W. Unequal Distribution of N6-Methyladenosine in Influenza Virus MRNAs. Mol. Cell. Biol. 1987, 7, 1572–1575. [Google Scholar] [CrossRef]
- Krug, R.M.; Morgan, M.A.; Shatkin, A.J. Influenza Viral MRNA Contains Internal N6-Methyladenosine and 5′-Terminal 7-Methylguanosine in Cap Structures. J. Virol. 1976, 20, 45–53. [Google Scholar] [CrossRef]
- Hesser, C.R.; Karijolich, J.; Dominissini, D.; He, C.; Glaunsinger, B.A. N6-Methyladenosine Modification and the YTHDF2 Reader Protein Play Cell Type Specific Roles in Lytic Viral Gene Expression during Kaposi’s Sarcoma-Associated Herpesvirus Infection. PLoS Pathog. 2018. [Google Scholar] [CrossRef]
- Tan, B.; Liu, H.; Zhang, S.; da Silva, S.R.; Zhang, L.; Meng, J.; Cui, X.; Yuan, H.; Sorel, O.; Zhang, S.W.; et al. Viral and Cellular N6-Methyladenosine and N6,2′-O-Dimethyladenosine Epitranscriptomes in the KSHV Life Cycle. Nat. Microbiol. 2017, 3, 108–120. [Google Scholar] [CrossRef] [PubMed]
- Lu, M.; Xue, M.; Wang, H.-T.; Kairis, E.L.; Ahmad, S.; Wei, J.; Zhang, Z.; Liu, Q.; Zhang, Y.; Gao, Y.; et al. Nonsegmented Negative-Sense RNA Viruses Utilize N6-Methyladenosine (m 6 A) as a Common Strategy To Evade Host Innate Immunity. J. Virol. 2021, 95. [Google Scholar] [CrossRef]
- Eckwahl, M.; Xu, R.; Michalkiewicz, J.; Zhang, W.; Patel, P.; Cai, Z.; Pan, T. 5-Methylcytosine RNA Modifications Promote Retrovirus Replication in an ALYREF Reader Protein-Dependent Manner. J. Virol. 2020. [Google Scholar] [CrossRef]
- Xue, M.; Zhao, B.S.; Zhang, Z.; Lu, M.; Harder, O.; Chen, P.; Lu, Z.; Li, A.; Ma, Y.; Xu, Y.; et al. Viral N 6-Methyladenosine Upregulates Replication and Pathogenesis of Human Respiratory Syncytial Virus. Nat. Commun. 2019, 10, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Xu, Y.-P.; Li, K.; Ye, Q.; Zhou, H.-Y.; Sun, H.; Li, X.; Yu, L.; Deng, Y.-Q.; Li, R.-T.; et al. The M6A Methylome of SARS-CoV-2 in Host Cells. Cell Res. 2021. [Google Scholar] [CrossRef]
- Kim, D.; Lee, J.Y.; Yang, J.S.; Kim, J.W.; Kim, V.N.; Chang, H. The Architecture of SARS-CoV-2 Transcriptome. Cell 2020, 181, 914–921.e10. [Google Scholar] [CrossRef] [PubMed]
- Dominissini, D.; Moshitch-Moshkovitz, S.; Schwartz, S.; Salmon-Divon, M.; Ungar, L.; Osenberg, S.; Cesarkas, K.; Jacob-Hirsch, J.; Amariglio, N.; Kupiec, M.; et al. Topology of the Human and Mouse M6A RNA Methylomes Revealed by M6A-Seq. Nature 2012, 485, 201–206. [Google Scholar] [CrossRef]
- Edelheit, S.; Schwartz, S.; Mumbach, M.R.; Wurtzel, O.; Sorek, R. Transcriptome-Wide Mapping of 5-Methylcytidine RNA Modifications in Bacteria, Archaea, and Yeast Reveals M5C within Archaeal MRNAs. PLoS Genet. 2013. [Google Scholar] [CrossRef]
- Chen, K.; Lu, Z.; Wang, X.; Fu, Y.; Luo, G.Z.; Liu, N.; Han, D.; Dominissini, D.; Dai, Q.; Pan, T.; et al. High-Resolution N6-Methyladenosine (M6A) Map Using Photo-Crosslinking-Assisted M6A Sequencing. Angew. Chem. Int. Ed. 2015, 54, 1587–1590. [Google Scholar] [CrossRef] [PubMed]
- Reid, R.; Greene, P.J.; Santi, D.V. Exposition of a Family of RNA M5C Methyltransferases from Searching Genomic and Proteomic Sequences. Nucleic Acids Res. 1999, 27, 3138–3145. [Google Scholar] [CrossRef] [PubMed]
- Bohnsack, K.E.; Höbartner, C.; Bohnsack, M.T. Eukaryotic 5-Methylcytosine (M 5 C) RNA Methyltransferases: Mechanisms, Cellular Functions, and Links to Disease. Genes 2019, 10, 102. [Google Scholar] [CrossRef]
- King, M.Y.; Redman, K.L. RNA Methyltransferases Utilize Two Cysteine Residues in the Formation of 5-Methylcytosine. Biochemistry 2002, 41, 11218–11225. [Google Scholar] [CrossRef]
- Hussain, S.; Sajini, A.A.; Blanco, S.; Dietmann, S.; Lombard, P.; Sugimoto, Y.; Paramor, M.; Gleeson, J.G.; Odom, D.T.; Ule, J.; et al. NSun2-Mediated Cytosine-5 Methylation of Vault Noncoding RNA Determines Its Processing into Regulatory Small RNAs. Cell Rep. 2013, 4, 255–261. [Google Scholar] [CrossRef]
- Linder, B.; Grozhik, A.V.; Olarerin-George, A.O.; Meydan, C.; Mason, C.E.; Jaffrey, S.R. Single-Nucleotide-Resolution Mapping of M6A and M6Am throughout the Transcriptome. Nat. Methods 2015, 12, 767–772. [Google Scholar] [CrossRef] [PubMed]
- Schaefer, M.; Pollex, T.; Hanna, K.; Lyko, F. RNA Cytosine Methylation Analysis by Bisulfite Sequencing. Nucleic Acids Res. 2009, 37. [Google Scholar] [CrossRef] [PubMed]
- Huang, T.; Chen, W.; Liu, J.; Gu, N.; Zhang, R. Genome-Wide Identification of MRNA 5-Methylcytosine in Mammals. Nat. Struct. Mol. Biol. 2019, 26. [Google Scholar] [CrossRef] [PubMed]
- Carlile, T.M.; Rojas-Duran, M.F.; Zinshteyn, B.; Shin, H.; Bartoli, K.M.; Gilbert, W.V. Pseudouridine Profiling Reveals Regulated MRNA Pseudouridylation in Yeast and Human Cells. Nature 2014, 515, 143–146. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, S.; Bernstein, D.A.; Mumbach, M.R.; Jovanovic, M.; Herbst, R.H.; León-Ricardo, B.X.; Engreitz, J.M.; Guttman, M.; Satija, R.; Lander, E.S.; et al. Transcriptome-Wide Mapping Reveals Widespread Dynamic-Regulated Pseudouridylation of NcRNA and MRNA. Cell 2014, 159, 148–162. [Google Scholar] [CrossRef]
- Marchand, V.; Blanloeil-Oillo, F.; Helm, M.; Motorin, Y. Illumina-Based RiboMethSeq Approach for Mapping of 2′-O-Me Residues in RNA. Nucleic Acids Res. 2016, 44, e135. [Google Scholar] [CrossRef]
- Hsu, P.J.; Fei, Q.; Dai, Q.; Shi, H.; Dominissini, D.; Ma, L.; He, C. Single Base Resolution Mapping of 2′-O-Methylation Sites in Human MRNA and in 3′ Terminal Ends of Small RNAs. Methods 2018. [Google Scholar] [CrossRef]
- Thalalla Gamage, S.; Sas-Chen, A.; Schwartz, S.; Meier, J.L. Quantitative Nucleotide Resolution Profiling of RNA Cytidine Acetylation by Ac4C-Seq. Nat. Protoc. 2021, 16, 2286–2307. [Google Scholar] [CrossRef]
- Sas-Chen, A.; Thomas, J.M.; Matzov, D.; Taoka, M.; Nance, K.D.; Nir, R.; Bryson, K.M.; Shachar, R.; Liman, G.L.S.; Burkhart, B.W.; et al. Dynamic RNA Acetylation Revealed by Quantitative Cross-Evolutionary Mapping. Nature 2020, 583, 638–643. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Rauch, S.; Dai, Q.; Cui, X.; Zhang, Z.; Nachtergaele, S.; Sepich, C.; He, C.; Dickinson, B.C. Evolution of a Reverse Transcriptase to Map N 1-Methyladenosine in Human Messenger RNA. Nat. Methods 2019, 16, 1281–1288. [Google Scholar] [CrossRef]
- Shi, H.; Wei, J.; He, C. Where, When, and How: Context-Dependent Functions of RNA Methylation Writers, Readers, and Erasers. Mol. Cell 2019, 74, 640–650. [Google Scholar] [CrossRef] [PubMed]
- Durbin, A.F.; Wang, C.; Marcotrigiano, J.; Gehrke, L. RNAs Containing Modified Nucleotides Fail to Trigger RIG-I Conformational Changes for Innate Immune Signaling. mBio 2016, 7, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Karikó, K.; Buckstein, M.; Ni, H.; Weissman, D. Suppression of RNA Recognition by Toll-like Receptors: The Impact of Nucleoside Modification and the Evolutionary Origin of RNA. Immunity 2005, 23, 165–175. [Google Scholar] [CrossRef] [PubMed]
| Modification | Writers | Readers | Roles on mRNAs |
|---|---|---|---|
| N6-methyladenosine (m6A) | METTL3 METTL4 METTL16 | YTHDF1-3 YTHDC1-2 | Splicing Stability Translation Localisation |
| 5-methylcytidine (m5C) | NSUN2 DNMT2 | YBX1 | Splicing Translation |
| 2ʹO-methylated nucleosides (Nm) | FTSJ3 | Unknown | Structure Stability |
| N4-acetylcytidine (ac4C) | NAT10 | Unknown | Stability Translation |
| Pseudouridine (ψ) | PUS7 TRUB1 DKC1 | Unknown | Codon misreading |
| 7-methylguanosine (m7G) | METTL1 | Unknown | Translation |
| N1-methyladenosine (m1A) | TRMT6/61A | YTHDF1-3 | Translation |
| 1-methylguanosine (m1G) | TRMT10A/B | Unknown | Unknown |
| N6,N6-dimethyladenosine (m6,6A) | Unknown | Unknown | Unknown |
| Mapping Method | Advantages | Disadvantages |
|---|---|---|
| Antibody mapping |
|
|
| Protein CLIP mapping |
|
|
| Biochemical mapping |
|
|
| Nanopore mapping |
|
|
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Courtney, D.G. Post-Transcriptional Regulation of Viral RNA through Epitranscriptional Modification. Cells 2021, 10, 1129. https://doi.org/10.3390/cells10051129
Courtney DG. Post-Transcriptional Regulation of Viral RNA through Epitranscriptional Modification. Cells. 2021; 10(5):1129. https://doi.org/10.3390/cells10051129
Chicago/Turabian StyleCourtney, David G. 2021. "Post-Transcriptional Regulation of Viral RNA through Epitranscriptional Modification" Cells 10, no. 5: 1129. https://doi.org/10.3390/cells10051129
APA StyleCourtney, D. G. (2021). Post-Transcriptional Regulation of Viral RNA through Epitranscriptional Modification. Cells, 10(5), 1129. https://doi.org/10.3390/cells10051129






