Structural Equation Modeling (SEM) Analysis of Sequence Variation and Green Plant Regeneration via Anther Culture in Barley
Abstract
1. Introduction
2. Materials and Methods
3. Results
3.1. Characterization of Input Data
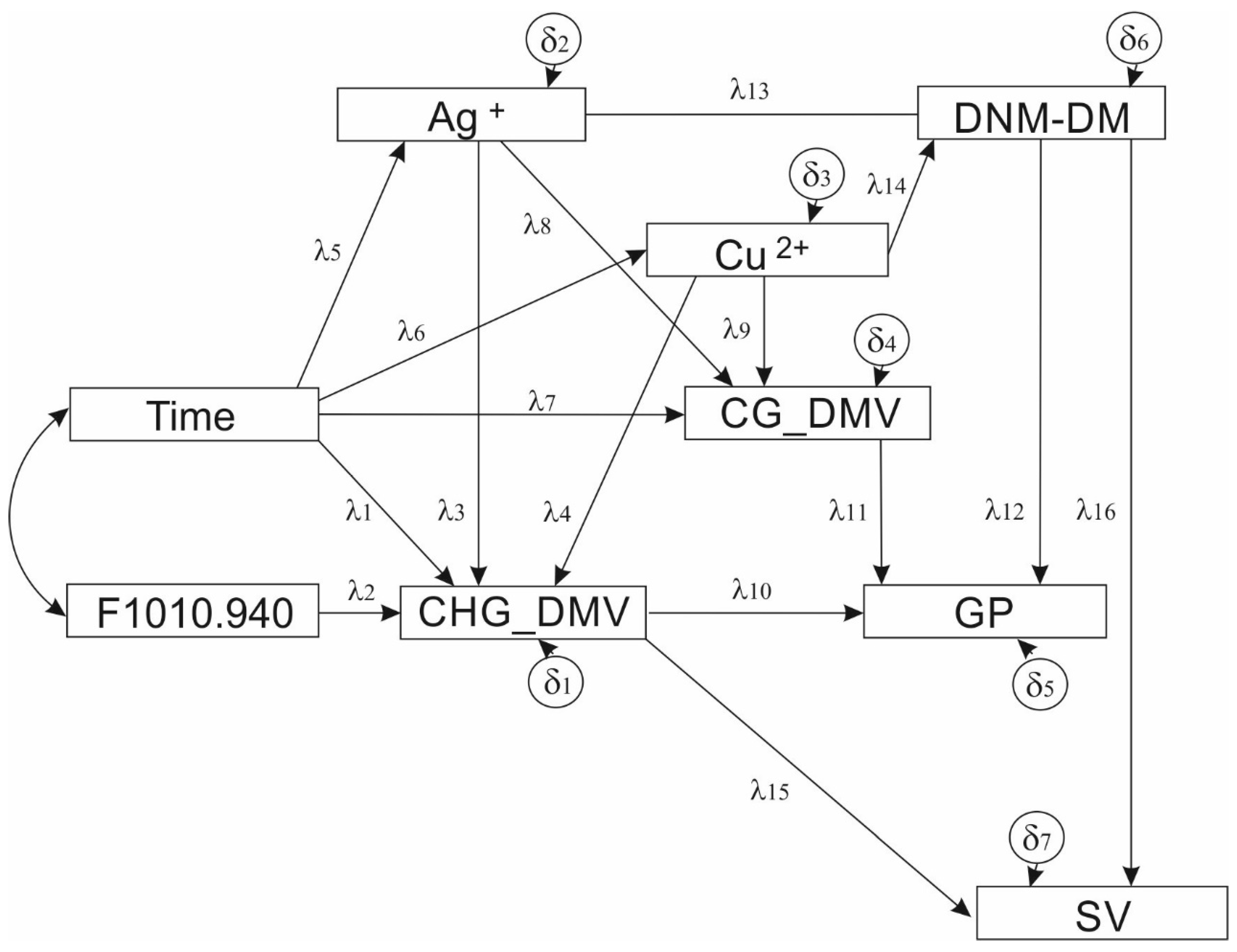
3.1.1. Model Specification and Estimation
3.1.2. Model Description
3.1.3. Model Matching
3.1.4. Estimation of Model Parameters
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| AGFI | Adjusted Goodness-of-Fit Index |
| AMPK | AMP-Activated Protein Kinase |
| ATP | Adenosine Triphosphate |
| ATR-FTIR | Attenuated Total Reflection Fourier Transform Infrared spectroscopy |
| CFI | Comparative Fit Index |
| DArTseqMet | Diversity Arrays Technology Sequencing Methylation Analysis |
| DM | Demethylation |
| DNM | De Novo methylation |
| ET | Endogenous ethylene |
| ETR1 | Ethylene receptor1 |
| GFI | Goodness-of-Fit Index |
| GP | Green Plant |
| IFI | Incremental Fit Index |
| metAFLP | Methylation-Sensitive Amplified Fragment Length Polymorphism |
| ML | Maximum Likelihood |
| MSAP | Methylation Sensitive Amplification Polymorphism |
| NFI | Normed Fit Index |
| NNFI | Non-Normed Fit Index |
| PCFI | Parsimonious Comparative Fit Index |
| PNFI | Parsimonious Normed Fit Index |
| RFI | Relative Fit Index |
| RMR | Root Mean Squares Residuals |
| RMSEA | Root Mean Square Error of Approximation |
| ROS | Reactive Oxygen Species |
| SAM | S-Adenosyl-L-Methionine |
| SEM | Structure Equatation Modeling |
| SRMR | Standardized Root Mean Squares Residuals |
| SV | Sequence Variation |
| TCA | Tricarboxylic Acid Cycle |
| TCIV | Tissue Culture-Induced Variation |
References
- Cistue, L.; Ramos, A.; Castillo, A.M. Influence of anther pretreatment and culture medium composition on the production of barley doubled haploids from model and low responding cultivars. Plant Cell Tissue Organ Cult. 1999, 55, 159–166. [Google Scholar] [CrossRef]
- Sood, S.; Dwivedi, S. Doubled Haploid Platform: An Accelerated Breeding Approach for Crop Improvement. In Plant Biology and Biotechnology: Volume II: Plant Genomics and Biotechnology; Bahadur, B., Venkat Rajam, M., Sahijram, L., Krishnamurthy, K.V., Eds.; Springer: New Delhi, India, 2015; pp. 89–111. [Google Scholar] [CrossRef]
- Huh, Y.S.; Lee, J.K.; Nam, S.Y. Effect of plant growth regulators and antioxidants on in vitro plant regeneration and callus induction from leaf explants of purple passion fruit (Passiflora edulis Sims). J. Plant Biotechnol. 2017, 44, 335–342. [Google Scholar] [CrossRef]
- Yildiz, M. The Prerequisite of the Success in Plant Tissue Culture: High Frequency Shoot Regeneration. In Recent Advances in Plant In Vitro Culture; Leva, A., Ed.; IntechOpen: London, UK, 2012. [Google Scholar] [CrossRef]
- Maraschin, S.F.; de Priester, W.; Spaink, H.P.; Wang, M. Androgenic switch: An example of plant embryogenesis from the male gametophyte perspective. J. Exp. Bot. 2005, 56, 1711–1726. [Google Scholar] [CrossRef] [PubMed]
- Ito, H.; Gaubert, H.; Bucher, E.; Mirouze, M.; Vaillant, I.; Paszkowski, J. An siRNA pathway prevents transgenerational retrotransposition in plants subjected to stress. Nature 2011, 472, 115–119. [Google Scholar] [CrossRef]
- Coronel, C.J.; González, A.I.; Ruiz, M.L.; Polanco, C. Analysis of somaclonal variation in transgenic and regenerated plants of Arabidopsis thaliana using methylation related metAFLP and TMD markers. Plant Cell Rep. 2018, 37, 137–152. [Google Scholar] [CrossRef]
- Machczyńska, J.; Zimny, J.; Bednarek, P. Tissue culture-induced genetic and epigenetic variation in triticale (× Triticosecale spp. Wittmack ex A. Camus 1927) regenerants. Plant Mol. Biol. 2015, 89, 279–292. [Google Scholar] [CrossRef]
- Orłowska, R.; Bednarek, P.T. Precise evaluation of tissue culture-induced variation during optimisation of in vitro regeneration regime in barley. Plant Mol. Biol. 2020, 103, 33–50. [Google Scholar] [CrossRef]
- Polanco, C.; Ruiz, M.L. AFLP analysis of somaclonal variation in Arabidopsis thaliana regenerated plants. Plant Sci. 2002, 162, 817–824. [Google Scholar] [CrossRef]
- Machczyńska, J.; Orłowska, R.; Mańkowski, D.R.; Zimny, J.; Bednarek, P.T. DNA methylation changes in triticale due to in vitro culture plant regeneration and consecutive reproduction. Plant Cell Tissue Organ Cult. 2014, 119, 289–299. [Google Scholar] [CrossRef]
- Orłowska, R.; Machczyńska, J.; Oleszczuk, S.; Zimny, J.; Bednarek, P.T. DNA methylation changes and TE activity induced in tissue cultures of barley (Hordeum vulgare L.). J. Biol. Res. 2016, 23, 19. [Google Scholar] [CrossRef]
- Orłowska, R.; Pachota, K.A.; Dynkowska, W.M.; Niedziela, A.; Bednarek, P.T. Androgenic-Induced Transposable Elements Dependent Sequence Variation in Barley. Int. J. Mol. Sci. 2021, 22, 6783. [Google Scholar] [CrossRef]
- Rodríguez-Serrano, M.; Bárány, I.; Prem, D.; Coronado, M.-J.; Risueño, M.C.; Testillano, P.S. NO, ROS, and cell death associated with caspase-like activity increase in stress-induced microspore embryogenesis of barley. J. Exp. Bot. 2012, 63, 2007–2024. [Google Scholar] [CrossRef]
- Foyer, C.H.; Noctor, G. Redox Homeostasis and Antioxidant Signaling: A Metabolic Interface between Stress Perception and Physiological Responses. Plant Cell 2005, 17, 1866–1875. [Google Scholar] [CrossRef] [PubMed]
- Das, K.; Roychoudhury, A. Reactive oxygen species (ROS) and response of antioxidants as ROS-scavengers during environmental stress in plants. Front. Environ. Sci. 2014, 2, 53. [Google Scholar] [CrossRef]
- Wani, S.H.; Kumar, V.; Shriram, V.; Sah, S.K. Phytohormones and their metabolic engineering for abiotic stress tolerance in crop plants. Crop J. 2016, 4, 162–176. [Google Scholar] [CrossRef]
- Durgud, M.; Gupta, S.; Ivanov, I.; Omidbakhshfard, M.A.; Benina, M.; Alseekh, S.; Staykov, N.; Hauenstein, M.; Dijkwel, P.P.; Hörtensteiner, S.; et al. Molecular Mechanisms Preventing Senescence in Response to Prolonged Darkness in a Desiccation-Tolerant Plant. Plant Physiol. 2018, 177, 1319–1338. [Google Scholar] [CrossRef] [PubMed]
- Kunz, H.H.; Scharnewski, M.; Feussner, K.; Feussner, I.; Flügge, U.I.; Fulda, M.; Gierth, M. The ABC transporter PXA1 and peroxisomal beta-oxidation are vital for metabolism in mature leaves of Arabidopsis during extended darkness. Plant Cell 2009, 21, 2733–2749. [Google Scholar] [CrossRef] [PubMed]
- Avin-Wittenberg, T.; Bajdzienko, K.; Wittenberg, G.; Alseekh, S.; Tohge, T.; Bock, R.; Giavalisco, P.; Fernie, A.R. Global analysis of the role of autophagy in cellular metabolism and energy homeostasis in Arabidopsis seedlings under carbon starvation. Plant Cell 2015, 27, 306–322. [Google Scholar] [CrossRef]
- Nemeth, C.; Freeman, J.; Jones, H.D.; Sparks, C.; Pellny, T.K.; Wilkinson, M.D.; Dunwell, J.; Andersson, A.A.M.; Åman, P.; Guillon, F.; et al. Down-regulation of the CSLF6 gene results in decreased (1,3;1,4)-beta-D-glucan in endosperm of wheat. Plant Physiol. 2010, 152, 1209–1218. [Google Scholar] [CrossRef] [PubMed]
- Parra-Vega, V.; Corral-Martínez, P.; Rivas-Sendra, A.; Seguí-Simarro, J.M. Induction of Embryogenesis in Brassica Napus Microspores Produces a Callosic Subintinal Layer and Abnormal Cell Walls with Altered Levels of Callose and Cellulose. Front. Plant Sci. 2015, 6, 1018. [Google Scholar] [CrossRef]
- Li, D.; Calderone, R. Exploiting mitochondria as targets for the development of new antifungals. Virulence 2017, 8, 159–168. [Google Scholar] [CrossRef] [PubMed]
- Roulin, S.; Buchala, A.J.; Fincher, G.B. Induction of (1→3,1→4)-β-d-glucan hydrolases in leaves of dark-incubated barley seedlings. Planta 2002, 215, 51–59. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sima, P.; Richter, J.; Vetvicka, V. Glucans as New Anticancer Agents. Anticancer. Res. 2019, 39, 3373–3378. [Google Scholar] [CrossRef] [PubMed]
- Able, A.J.; Sutherland, M.W.; Guest, D.I. Production of reactive oxygen species during non-specific elicitation, non-host resistance and field resistance expression in cultured tobacco cells. Funct. Plant Biol. 2003, 30, 91–99. [Google Scholar] [CrossRef] [PubMed]
- Toro, G.; Pinto, M. Plant respiration under low oxygen. Chil. J. Agric. Res. 2015, 75, 57–70. [Google Scholar] [CrossRef]
- Printz, B.; Lutts, S.; Hausman, J.-F.; Sergeant, K. Copper Trafficking in Plants and Its Implication on Cell Wall Dynamics. Front. Plant Sci. 2016, 7, 601. [Google Scholar] [CrossRef] [PubMed]
- Ravet, K.; Pilon, M. Copper and Iron Homeostasis in Plants: The Challenges of Oxidative Stress. Antioxid. Redox Signal 2013, 19, 919–932. [Google Scholar] [CrossRef] [PubMed]
- Pádua, M.; Cavaco, A.M.; Aubert, S.; Bligny, R.; Casimiro, A. Effects of copper on the photosynthesis of intact chloroplasts: Interaction with manganese. Physiol. Plant. 2010, 138, 301–311. [Google Scholar] [CrossRef]
- Tan, Y.-F.; O’Toole, N.; Taylor, N.L.; Millar, A.H. Divalent metal ions in plant mitochondria and their role in interactions with proteins and oxidative stress-induced damage to respiratory function. Plant Physiol. 2010, 152, 747–761. [Google Scholar] [CrossRef] [PubMed]
- Hirayama, T.; Alonso, J.M. Ethylene Captures a Metal! Metal Ions Are Involved in Ethylene Perception and Signal Transduction. Plant Cell Physiol. 2000, 41, 548–555. [Google Scholar] [CrossRef]
- Bowler, C.; Montagu, M.V.; Inze, D. Superoxide dismutase and stress tolerance. Annu. Rev. Plant Biol. 1992, 43, 83–116. [Google Scholar] [CrossRef]
- Lauer, M.M.; de Oliveira, C.B.; Yano, N.L.I.; Bianchini, A. Copper effects on key metabolic enzymes and mitochondrial membrane potential in gills of the estuarine crab Neohelice granulata at different salinities. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2012, 156, 140–147. [Google Scholar] [CrossRef]
- Cong, W.; Miao, Y.; Xu, L.; Zhang, Y.; Yuan, C.; Wang, J.; Zhuang, T.; Lin, X.; Jiang, L.; Wang, N.; et al. Transgenerational memory of gene expression changes induced by heavy metal stress in rice (Oryza sativa L.). BMC Plant Biol. 2019, 19, 282. [Google Scholar] [CrossRef]
- Svirshchevskaya, A.M.; Bormotov, V.E. Characteristic property of regeneration under gynogenesis of sugar-beet. Doklady Akademii Nauk Belarusi 1994, 38, 57–59. [Google Scholar]
- Trifonova, A.; Madsen, S.; Olesen, A. Agrobacterium-mediated transgene delivery and integration into barley under a range of in vitro culture conditions. Plant Sci. 2001, 161, 871–880. [Google Scholar] [CrossRef]
- Caredda, S.; Clément, C. Androgenesis and albinism in Poaceae: Influence of genotype and carbohydrates. In Anther and Pollen; Springer: Berlin/Heidelberg, Germany, 1999; pp. 211–228. [Google Scholar]
- Yao, J.L.; Cohen, D. Multiple gene control of plastome-genome incompatibility and plastid DNA inheritance in interspecific hybrids of Zantedeschia. Theor. Appl. Genet. 2000, 101, 400–406. [Google Scholar] [CrossRef]
- Makowska, K.; Oleszczuk, S. Albinism in barley androgenesis. Plant Cell Rep. 2014, 33, 385–392. [Google Scholar] [CrossRef]
- Cohu, C.M.; Pilon, M. Cell Biology of Copper. In Cell Biology of Metals and Nutrients; Hell, R., Mendel, R.-R., Eds.; Springer: Berlin/Heidelberg, Germany, 2010; pp. 55–74. [Google Scholar] [CrossRef]
- Jacquard, C.; Nolin, F.; Hecart, C.; Grauda, D.; Rashal, I.; Dhondt-Cordelier, S.; Sangwan, R.S.; Devaux, P.; Mazeyrat-Gourbeyre, F.; Clement, C. Microspore embryogenesis and programmed cell death in barley: Effects of copper on albinism in recalcitrant cultivars. Plant Cell Rep. 2009, 28, 1329–1339. [Google Scholar] [CrossRef]
- Orłowska, R.; Pachota, K.A.; Machczyńska, J.; Niedziela, A.; Makowska, K.; Zimny, J.; Bednarek, P.T. Improvement of anther cultures conditions using the Taguchi method in three cereal crops. Electron. J. Biotechnol. 2020, 43, 8–15. [Google Scholar] [CrossRef]
- Puchkova, L.V.; Broggini, M.; Polishchuk, E.V.; Ilyechova, E.Y.; Polishchuk, R.S. Silver Ions as a Tool for Understanding Different Aspects of Copper Metabolism. Nutrients 2019, 11, 1364. [Google Scholar] [CrossRef]
- Costa, C.S.; Ronconi, J.V.V.; Daufenbach, J.F.; Gonçalves, C.L.; Rezin, G.T.; Streck, E.L.; da Silva Paula, M.M. In vitro effects of silver nanoparticles on the mitochondrial respiratory chain. Mol. Cell. Biochem. 2010, 342, 51–56. [Google Scholar] [CrossRef] [PubMed]
- McDaniel, B.K.; Binder, B.M. Ethylene Receptor 1 (ETR1) Is Sufficient and Has the Predominant Role in Mediating Inhibition of Ethylene Responses by Silver in Arabidopsis thaliana. J. Biol. Chem. 2012, 287, 26094–26103. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez, F.I.; Esch, J.J.; Hall, A.E.; Binder, B.M.; Schaller, G.E.; Bleecker, A.B. A copper cofactor for the ethylene receptor ETR1 from Arabidopsis. Science 1999, 283, 996–998. [Google Scholar] [CrossRef] [PubMed]
- Bednarek, P.T.; Orłowska, R. CG Demethylation Leads to Sequence Mutations in an Anther Culture of Barley Due to the Presence of Cu, Ag Ions in the Medium and Culture Time. Int. J. Mol. Sci. 2020, 21, 4401. [Google Scholar] [CrossRef] [PubMed]
- Baker, M.J.; Trevisan, J.; Bassan, P.; Bhargava, R.; Butler, H.J.; Dorling, K.M.; Fielden, P.R.; Fogarty, S.W.; Fullwood, N.J.; Heys, K.A.; et al. Using Fourier transform IR spectroscopy to analyze biological materials. Nat. Protoc. 2014, 9, 1771–1791. [Google Scholar] [CrossRef] [PubMed]
- Bednarek, P.T.; Zebrowski, J.; Orłowska, R. Exploring the Biochemical Origin of DNA Sequence Variation in Barley Plants Regenerated via in Vitro Anther Culture. Int. J. Mol. Sci. 2020, 21, 5770. [Google Scholar] [CrossRef]
- Legner, N.; Meinen, C.; Rauber, R. Root Differentiation of Agricultural Plant Cultivars and Proveniences Using FTIR Spectroscopy. Front. Plant Sci. 2018, 9, 748. [Google Scholar] [CrossRef]
- Bureau, S.; Cozzolino, D.; Clark, C.J. Contributions of Fourier-transform mid infrared (FT-MIR) spectroscopy to the study of fruit and vegetables: A review. Postharvest Biol. Technol. 2019, 148, 1–14. [Google Scholar] [CrossRef]
- Rana, R.; Herz, K.; Bruelheide, H.; Dietz, S.; Haider, S.; Jandt, U.; Pena, R. Leaf Attenuated Total Reflection Fourier Transform Infrared (ATR-FTIR) biochemical profile of grassland plant species related to land-use intensity. Ecol. Indic. 2018, 84, 803–810. [Google Scholar] [CrossRef]
- Pereira, W.J.; Pappas, M.d.C.R.; Grattapaglia, D.; Pappas, G.J., Jr. A cost-effective approach to DNA methylation detection by Methyl Sensitive DArT sequencing. PLoS ONE 2020, 15, e0233800. [Google Scholar] [CrossRef]
- Hayes, A.F. Introduction to Mediation, Moderation, and Conditional Process Analysis. A Regression Bases Approach; A Division of Guilford Publications, Inc.: New York, NY, USA, 2018; p. 507. [Google Scholar]
- Bednarek, P.T.; Orłowska, R. Time of In Vitro Anther Culture May Moderate Action of Copper and Silver Ions that Affect the Relationship between DNA Methylation Change and the Yield of Barley Green Regenerants. Plants 2020, 9, 1064. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.F.; Hoffman, N.E. Ethylene Biosynthesis and its Regulation in Higher Plants. Annu. Rev. Plant Physiol. 1984, 35, 155–189. [Google Scholar] [CrossRef]
- Sauter, M.; Moffatt, B.; Saechao, M.C.; Hell, R.; Wirtz, M. Methionine salvage and S-adenosylmethionine: Essential links between sulfur, ethylene and polyamine biosynthesis. Biochem. J. 2013, 451, 145–154. [Google Scholar] [CrossRef]
- Kende, H. Ethylene Biosynthesis. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1993, 44, 283–307. [Google Scholar] [CrossRef]
- Lee, D.H.; O’Connor, T.R.; Pfeifer, G.P. Oxidative DNA damage induced by copper and hydrogen peroxide promotes CG-->TT tandem mutations at methylated CpG dinucleotides in nucleotide excision repair-deficient cells. Nucleic Acids Res. 2002, 30, 3566–3573. [Google Scholar] [CrossRef]
- McDonald, R.P. A simple comprehensive model for the analysis of covariance structures. Br. J. Math. Stat. Psychol. 1978, 31, 59–72. [Google Scholar] [CrossRef]
- Jöreskog, K.G. Structural equation models in the social sciences: Specification, estimation and testing. In Applications in Statistics; Krishnaiah, P.R., Ed.; Applications in Statistics: New York, NY, USA, 1977; pp. 265–287. [Google Scholar]
- Shipley, B. Cause and Correlation in Biology: A User’s Guide to Path Analysis, Structural Equations and Causal Inference; Cambridge University Press: Cambridge, UK, 2000. [Google Scholar] [CrossRef]
- Kozak, M.; Kang, M.S. Note on modern path analysis in application to crop science. Commun. Biometry Crop Sci. 2006, 1, 32–34. [Google Scholar]
- Lamb, E.G.; Shirtliffe, S.J.; May, W.E. Structural equation modeling in the plant sciences: An example using yield components in oat. Can. J. Plant Sci. 2011, 91, 603–619. [Google Scholar] [CrossRef]
- Arbuckle, J.L. Amos; Version 23.0; IBM SPSS: Chicago, IL, USA, 2014. [Google Scholar]
- Levenberg, K. A method for the solution of certain non linear problems in least squares. Q. Appl. Math. 1944, 2, 164–168. [Google Scholar] [CrossRef]
- Madsen, K.; Nielsen, H.B.; Tingleff, O. Methods for Non-Linear Least Squares Problems. Informatics and Mathematical Modeling; Technical University of Denmark: Lyngby, Denmark, 2004. [Google Scholar]
- Marquardt, D.W. An Algorithm for Least-Squares Estimation of Nonlinear Parameters. J. Soc. Ind. Appl. Math. 1963, 11, 431–441. [Google Scholar] [CrossRef]
- Taboga, M. Lectures on Probability Theory and Mathematical Statistics; CreateSpace Independent Publishing Platform: North Charleston, SC, USA, 2012. [Google Scholar]
- Kenny, D.A. Measuring Model Fit. Available online: http://davidakenny.net/cm/fit.htm (accessed on 12 October 2019).
- Bentler, P.M.; Bonett, D.G. Significance tests and goodness-of-fit in the analysis of covariance structures. Psychol. Bull. 1980, 88, 588–600. [Google Scholar] [CrossRef]
- Kenny, D.A.; McCoach, D.B. Effect of the Number of Variables on Measures of Fit in Structural Equation Modeling. Struct. Equ. Model. A Multidiscip. J. 2003, 10, 333–351. [Google Scholar] [CrossRef]
- MacCallum, R.C. Working with imperfect models. Multivar. Behav. Res. 2003, 1, 13–149. [Google Scholar]
- Ho, R. Handbook of Univariate and Multivariate Data Analysis and Interpretation with SPSS; Central Queensland University: Rockhampton, Australia, 2006. [Google Scholar]
- Williams, L.J.; Holahan, P.J. Parsimony-based fit indices for multiple-indicator models: Do they work? Struct. Equ. Model. A Multidiscip. J. 1994, 1, 161–189. [Google Scholar] [CrossRef]
- Hu, L.-T.; Bentler, P.M. Fit indices in covariance structure modeling: Sensitivity to underparameterized model misspecification. Psychol. Methods 1998, 3, 424–453. [Google Scholar] [CrossRef]
- Hu, L.-T.; Bentler, P.M. Cutoff criteria for fit indices in covariance structure analysis: Conventional criteria versus new alternatives. Struct. Equ. Model. 1999, 6, 1–55. [Google Scholar] [CrossRef]
- Mulaik, S.A.; James, L.R.; Van Alstine, J.; Bennett, N.; Lind, S.; Stilwell, C.D. Evaluation of goodness-of-fit indices for structural equation models. Psychol. Bull. 1989, 105, 430–445. [Google Scholar] [CrossRef]
- MacCallum, R.C.; Browne, M.W.; Sugawara, H.M. Power analysis and determination of sample size for covariance structure modeling. Psychol. Methods 1996, 1, 130–149. [Google Scholar] [CrossRef]
- Mansilla, N.; Racca, S.; Gras, D.E.; Gonzalez, D.H.; Welchen, E. The Complexity of Mitochondrial Complex IV: An Update of Cytochrome c Oxidase Biogenesis in Plants. Int. J. Mol. Sci. 2018, 19, 662. [Google Scholar] [CrossRef]
- Horn, D.; Barrientos, A. Mitochondrial copper metabolism and delivery to cytochrome c oxidase. IUBMB Life 2008, 60, 421–429. [Google Scholar] [CrossRef]
- Zhang, Y.; Fernie, A.R. On the role of the tricarboxylic acid cycle in plant productivity. J. Integr. Plant Biol. 2018, 60, 1199–1216. [Google Scholar] [CrossRef] [PubMed]
- Touraev, A.; Pfosser, M.; Heberle-Bors, E. The microspore: A haploid multipurpose cell. Adv. Bot. Res. 2001, 35, 53–109. [Google Scholar]
- Moffatt, B.A.; Weretilnyk, E.A. Sustaining S-adenosyl-l-methionine-dependent methyltransferase activity in plant cells. Physiol. Plant. 2001, 113, 435–442. [Google Scholar] [CrossRef]
- Ogawa, T.; Tsubakiyama, R.; Kanai, M.; Koyama, T.; Fujii, T.; Iefuji, H.; Soga, T.; Kume, K.; Miyakawa, T.; Hirata, D.; et al. Stimulating S-adenosyl-l-methionine synthesis extends lifespan via activation of AMPK. Proc. Natl. Acad. Sci. USA 2016, 113, 11913–11918. [Google Scholar] [CrossRef]
- Powell, W.; Thomas, W.T.B.; Baird, E.; Lawrence, P.; Booth, A.; Harrower, B.; McNicol, J.W.; Waugh, R. Analysis of quantitative traits in barley by the use of Amplified Fragment Length Polymorphisms. Heredity 1997, 79, 48–59. [Google Scholar] [CrossRef]
| Variable | Descriptive Statistics | |||
|---|---|---|---|---|
| Mean | Variance | Skewness | Kurtosis | |
| [F1010.940] 1 | 0.035 | 0.000 | 0.531 | −0.689 |
| [Cu2+] | 4.751 | 17.205 | 0.123 | −1.573 |
| [Ag+] | 20.286 | 667.546 | 0.900 | −1.129 |
| [DNM-DM] | 0.794 | 6.089 | −0.804 | −0.048 |
| [CG_DMV] | 0.391 | 0.056 | 0.170 | 0.219 |
| [CHG_DMV] | 0.940 | 0.098 | −2.393 | 5.433 |
| [SV] | 3.952 | 9.585 | 2.849 | 7.049 |
| [GP] | 1.123 | 0.715 | 0.835 | −0.424 |
| [Time] | 27.800 | 35.988 | 0.057 | −1.657 |
| Variable | [F1010.940] 1 | [Cu2+] | [Ag+] | [DNM_DM] | [CG_DMV] | [CHG_DMV] | [SV] | [GP] | [T] |
|---|---|---|---|---|---|---|---|---|---|
| [F1010.940] | 1.000 | ||||||||
| [Cu2+] | 0.338 a * | 1.000 | |||||||
| [Ag+] | −0.017 | −0.107 | 1.000 | ||||||
| [DNM_DM] | −0.128 | 0.476 ** | 0.166 | 1.000 | |||||
| [CG_DMV] | −0.157 | −0.055 | −0.208 | −0.023 | 1.000 | ||||
| [CHG_DMV] | −0.405 * | −0.438** | −0.508 ** | −0.231 | 0.590 ** | 1.000 | |||
| [SV] | 0.472 ** | 0.418 * | 0.478 ** | 0.080 | −0.391 * | −0.887 ** | 1.000 | ||
| [GP] | 0.050 | 0.347 * | 0.157 | 0.210 | −0.315 | −0.251 | 0.240 | 1.000 | |
| [Time] | −0.079 | −0.002 | 0.160 | 0.114 | 0.593 ** | 0.306 | −0.252 | −0.138 | 1.000 |
| Parameter | Postulated Model |
|---|---|
| Degrees of freedom (df) | 19 |
| Chi-square | 21.125 |
| p-value | 0.330 |
| Root Mean Squares Residuals (RMR) | 2.836 |
| Standardized Root Mean Squares Residuals (SRMR) | 0.134 |
| Goodness-of-Fit Index (GFI) | 0.862 |
| Adjusted Goodness-of-Fit Index (AGFI) | 0.673 |
| Normed Fit Index (NFI) | 0.647 |
| Relative Fit Index (RFI) | 0.331 |
| Incremental Fit Index (IFI) | 0.948 |
| Non-Normed Fit Index (NNFI) | 0.831 |
| Comparative Fit Index (CFI) | 0.911 |
| Parsimonious Normed Fit Index (PNFI) | 0.341 |
| Parsimonious Comparative Fit Index (PCFI) | 0.481 |
| Root Mean Square Error of Approximation (RMSEA) | 0.057 |
| Parameter | Effect | Estimate (b) | Standard Error | Test Statistic | Standardized Estimate (β) | ||
|---|---|---|---|---|---|---|---|
| Path coefficients | |||||||
| λ1 | [Time] | → | [CHG_DMV] | 0.014 | 0.005 | 2.641 ** | 0.284 |
| λ2 | [F1010.940] | → | [CHG_DMV] | −6.230 | 5.127 | −1.215 | −0.139 |
| λ3 | [Ag+] | → | [CHG_DMV] | −0.007 | 0.001 | −6.508 ** | −0.695 |
| λ4 | [Cu2+] | → | [CHG_DMV] | −0.037 | 0.009 | −4.180 ** | −0.481 |
| λ5 | [Time] | → | [Ag+] | 0.800 | 0.834 | 0.959 | 0.173 |
| λ6 | [Time] | → | [Cu2+] | −0.023 | 0.127 | −0.179 | −0.036 |
| λ7 | [Time] | → | [CG_DMV] | 0.018 | 0.005 | 3.555 ** | 0.592 |
| λ8 | [Ag+] | → | [CG_DMV] | −0.003 | 0.001 | −2.404 * | −0.402 |
| λ9 | [Cu2+] | → | [CG_DMV] | −0.009 | 0.009 | −1.011 | −0.179 |
| λ10 | [CHG_DMV] | → | [GP] | 0.055 | 0.676 | 0.082 | 0.019 |
| λ11 | [CG_DMV] | → | [GP] | −2.596 | 0.953 | −2.723 ** | −0.563 |
| λ12 | [DNM_DM] | → | [GP] | 0.046 | 0.068 | 0.684 | 0.141 |
| λ13 | [Ag+] | → | [DNM-DM] | 0.021 | 0.013 | 1.586 | 0.220 |
| λ14 | [Cu2+] | → | [DNM-DM] | 0.454 | 0.109 | 4.161 ** | 0.646 |
| λ15 | [CHG_DMV] | → | [SV] | −9.803 | 1.047 | −9.366 ** | −0.985 |
| λ16 | [DNM_DM] | → | [SV] | −0.200 | 0.106 | −1.880 | 0.141 |
| Covariances | |||||||
| φ1 | [F1010.940] | ↔ | [Time] | 0.002 | 0.007 | 0.347 | 0.074 |
| Variances | |||||||
| δ1 | 0.018 | 0.005 | 3.346 ** | ||||
| δ2 | 616.775 | 151.285 | 4.077 ** | ||||
| δ3 | 11.636 | 3.407 | 3.415 ** | ||||
| δ4 | 0.015 | 0.005 | 3.018 ** | ||||
| δ5 | 0.396 | 0.124 | 3.199 ** | ||||
| δ6 | 3.083 | 0.930 | 3.313 ** | ||||
| δ7 | 1.120 | 0.328 | 3.418 ** | ||||
| [Time] | 29.723 | 7.845 | 3.789 ** | ||||
| [F1010.940] | 0.000 | 0.000 | 3.015 ** | ||||
| Effect | Estimates (b) | Standardized Estimates (β) | ||||||
|---|---|---|---|---|---|---|---|---|
| Direct Effect | Indirect Effects | Total Effects | Direct Effect | Indirect Effects | Total Effects | |||
| [CHG_DMV] | ||||||||
| [Time] | → | [CHG_DMV] | 0.014 | −0.005 | 0.009 | 0.284 | −0.103 | 0.181 |
| [F1010.940] | → | [CHG_DMV] | −6.230 | — | −6.230 | −0.139 | — | −0.139 |
| [Ag+] | → | [CHG_DMV] | 0.007 | — | 0.007 | −0.695 | — | −0.695 |
| [Cu2+] | → | [CHG_DMV] | −0.037 | — | −0.037 | −0.481 | — | −0.481 |
| [Ag+] | ||||||||
| [Time] | → | [Ag+] | 0.800 | — | 0.800 | 0.173 | — | 0.173 |
| [Cu2+] | ||||||||
| [Time] | → | [Cu2+] | −0.023 | — | −0.023 | −0.036 | — | −0.036 |
| [SV] | ||||||||
| [CHG_DMV] | → | [SV] | −9.803 | — | −9.803 | −0.985 | — | −0.985 |
| [Time] | → | [SV] | — | −0.088 | −0.088 | — | −0.181 | −0.181 |
| [F1010.940] | → | [SV] | — | 61.073 | 61.073 | — | 0.137 | 0.137 |
| [Ag+] | → | [SV] | — | 0.067 | 0.067 | — | 0.644 | 0.644 |
| [Cu2+] | → | [SV] | — | 0.275 | 0.275 | — | 0.356 | 0.356 |
| [DNM_DM] | → | [SV] | −0.200 | — | −0.200 | −0.182 | — | −0.182 |
| [GP] | ||||||||
| [CHG_DMV] | → | [GP] | 0.055 | — | 0.055 | 0.019 | — | 0.019 |
| [Time] | → | [GP] | — | −0.042 | −0.042 | — | −0.292 | −0.292 |
| [F1010.940] | → | [GP] | — | −0.344 | −0.344 | — | −0.003 | −0.003 |
| [Ag+] | → | [GP] | — | 0.008 | 0.008 | — | 0.245 | 0.245 |
| [Cu2+] | → | [GP] | — | 0.042 | 0.042 | — | 0.183 | 0.183 |
| [CG_DMV] | → | [GP] | −2.596 | — | −2.596 | −0.563 | — | −0.563 |
| [DNM_DM] | → | [GP] | 0.046 | — | 0.046 | 0.141 | — | 0.141 |
| [CG_DMV] | ||||||||
| [Ag+] | → | [CG_DMV] | −0.003 | — | −0.003 | −0.402 | — | −0.402 |
| [Cu2+] | → | [CG_DMV] | −0.009 | — | −0.009 | −0.179 | — | −0.179 |
| [Time] | → | [CG_DMV] | 0.017 | — | 0.017 | 0.591 | — | 0.529 |
| [DNM_DM] | ||||||||
| [Ag+] | → | [DNM_DM] | 0.021 | — | 0.021 | 0.220 | — | 0.220 |
| [Cu2+] | → | [DNM_DM] | 0.454 | — | 0.454 | 0.646 | — | 0.646 |
| [Time] | → | [DNM_DM] | — | 0.006 | 0.006 | — | 0.015 | 0.015 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bednarek, P.T.; Orłowska, R.; Mańkowski, D.R.; Oleszczuk, S.; Zebrowski, J. Structural Equation Modeling (SEM) Analysis of Sequence Variation and Green Plant Regeneration via Anther Culture in Barley. Cells 2021, 10, 2774. https://doi.org/10.3390/cells10102774
Bednarek PT, Orłowska R, Mańkowski DR, Oleszczuk S, Zebrowski J. Structural Equation Modeling (SEM) Analysis of Sequence Variation and Green Plant Regeneration via Anther Culture in Barley. Cells. 2021; 10(10):2774. https://doi.org/10.3390/cells10102774
Chicago/Turabian StyleBednarek, Piotr Tomasz, Renata Orłowska, Dariusz Rafał Mańkowski, Sylwia Oleszczuk, and Jacek Zebrowski. 2021. "Structural Equation Modeling (SEM) Analysis of Sequence Variation and Green Plant Regeneration via Anther Culture in Barley" Cells 10, no. 10: 2774. https://doi.org/10.3390/cells10102774
APA StyleBednarek, P. T., Orłowska, R., Mańkowski, D. R., Oleszczuk, S., & Zebrowski, J. (2021). Structural Equation Modeling (SEM) Analysis of Sequence Variation and Green Plant Regeneration via Anther Culture in Barley. Cells, 10(10), 2774. https://doi.org/10.3390/cells10102774






