A LEA Gene from a Vietnamese Maize Landrace Can Enhance the Drought Tolerance of Transgenic Maize and Tobacco
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials and Drought-Stimulating Growth Conditions
2.2. Isolation of the Putative LEA Coding Region in Tevang 1 Landrace Maize
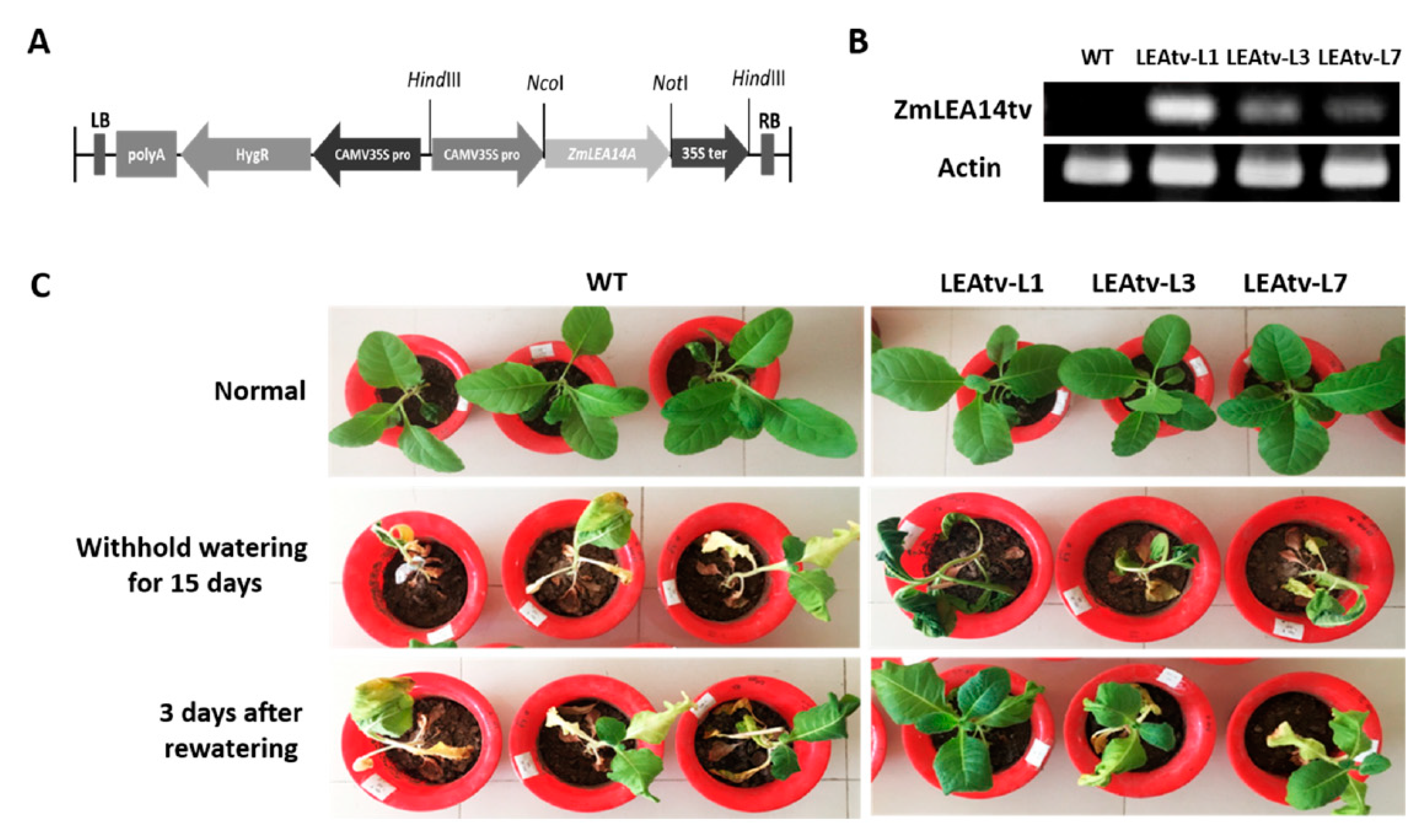
2.3. ZmLEA14tv Expression Vector Construction and Transgenic Plant Generation
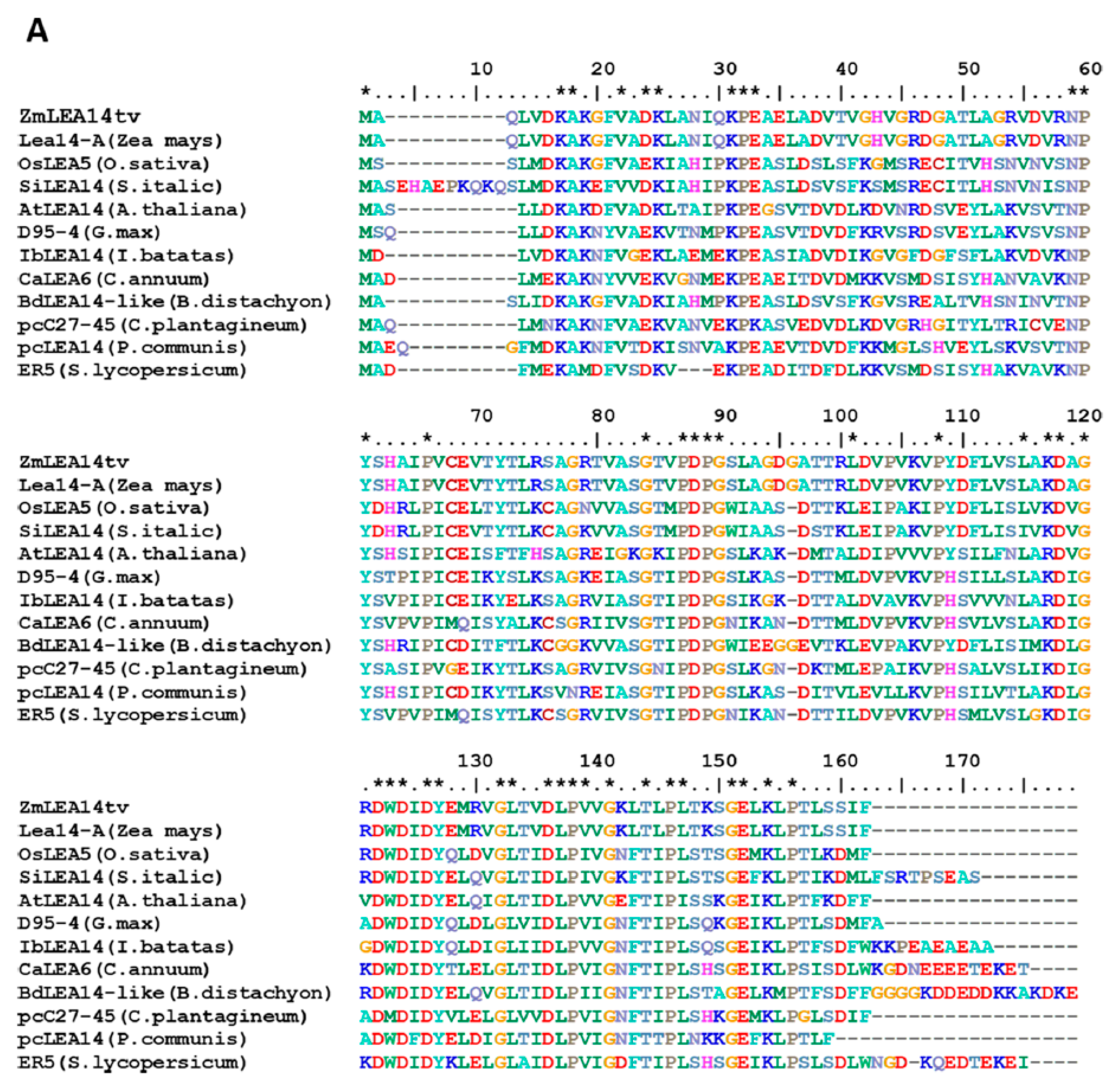
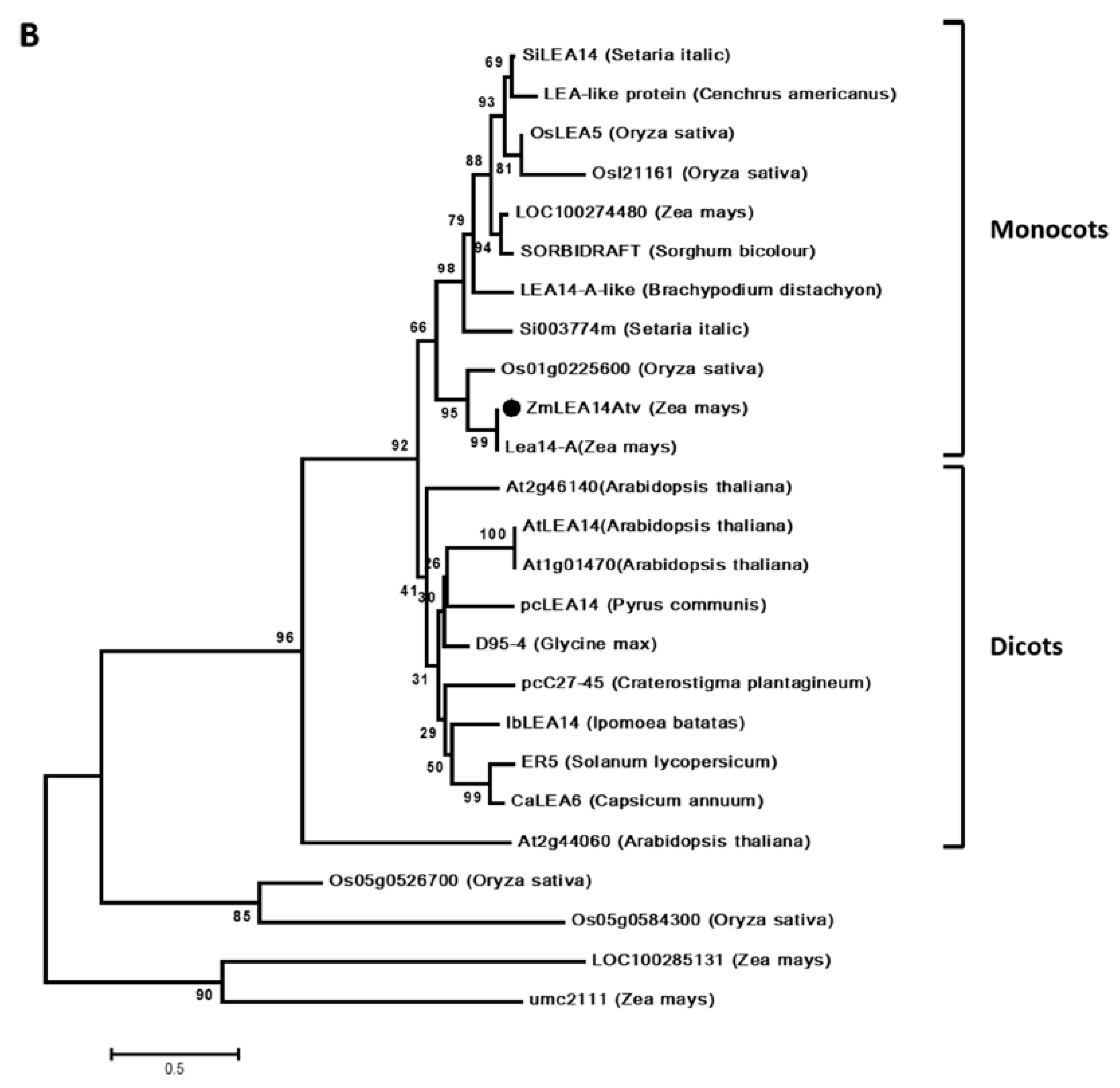
2.4. Sequence Alignment and Gene Evolution Analysis
3. Results
3.1. Isolation and Vector Construction of ZmLEA14tv Gene
3.2. ZmLEA14tv Gene Expression in Drought Resistance for Transgenic Tobacco
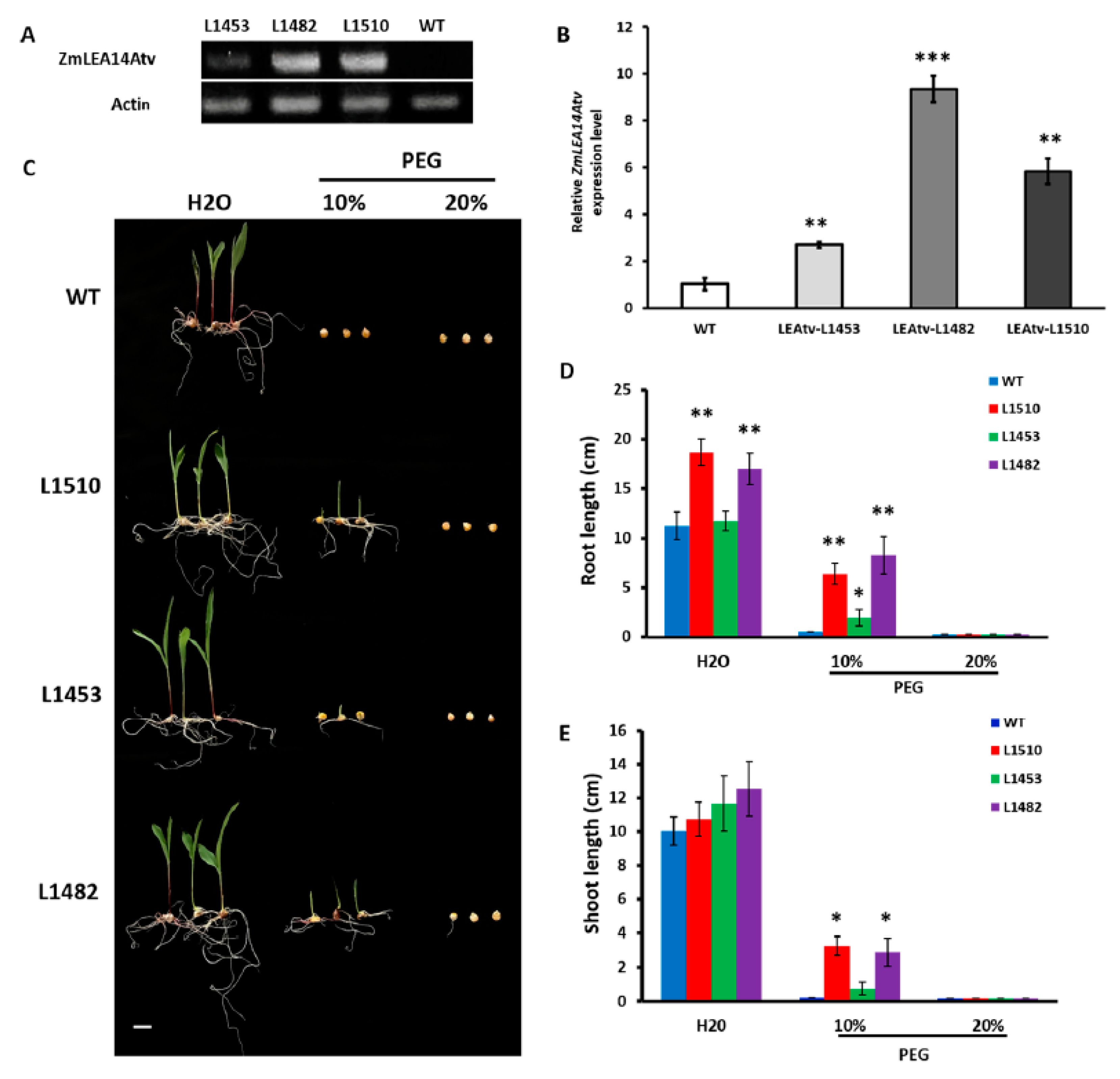
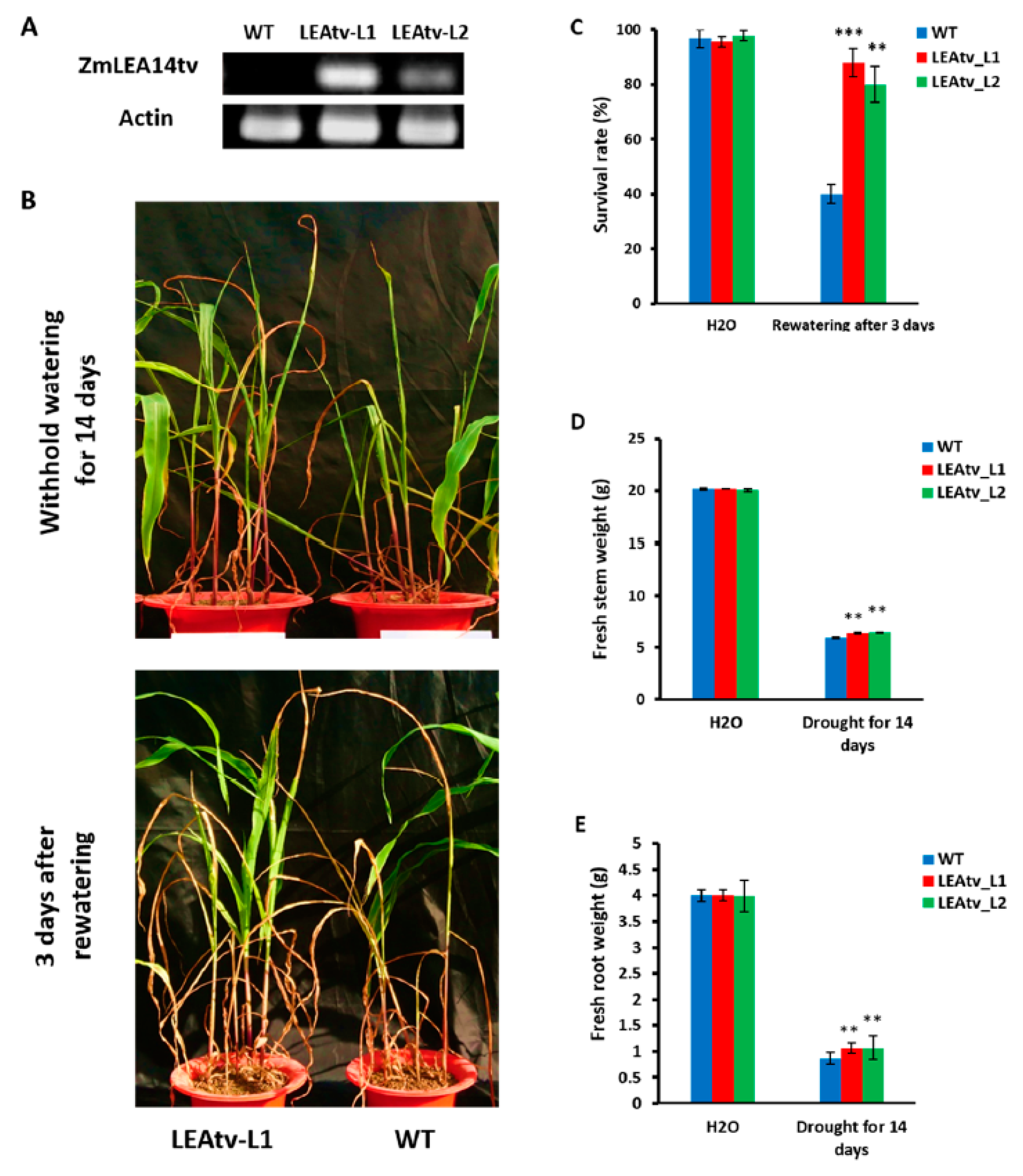
3.3. Transgenic Maize with ZmLEA14tv Gene in Drought Tolerance
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Chandler, P.M.; Robertson, M. Gene expression regulated by abscisic acid and its relation to stress tolerance. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1994, 45, 113–141. [Google Scholar] [CrossRef]
- Hong-Bo, S.; Zong-Suo, L.; Ming-An, S. LEA proteins in higher plants: Structure, function, gene expression and regulation. Colloids Surf. B Biointerfaces 2005, 45, 131–135. [Google Scholar] [CrossRef] [PubMed]
- Kimura, M.; Yamamoto Yoshiharu, Y.; Seki, M.; Sakurai, T.; Sato, M.; Abe, T.; Yoshida, S.; Manabe, K.; Shinozaki, K.; Matsui, M. Identification of arabidopsis genes regulated by high light–stress using cDNA microarray. Photochem. Photobiol. 2007, 77, 226–233. [Google Scholar]
- Tunnacliffe, A.; Wise, M.J. The continuing conundrum of the LEA proteins. Naturwissenschaften 2007, 94, 791–812. [Google Scholar] [CrossRef]
- Baker, J.; Van Dennsteele, C.; Dure, L. Sequence and characterization of 6 LEA proteins and their genes from cotton. Plant Mol. Biol. 1988, 11, 277–291. [Google Scholar] [CrossRef] [PubMed]
- Dure, L.; Galau, G.A. Developmental Biochemistry of Cottonseed Embryogenesis and Germination. Plant Physiol. 1981, 68, 187–194. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, E.; Amara, I.; Bellido, D.; Odena, M.A.; Dominguez, E.; Pages, M.; Goday, A. LC-MSMS identification of Arabidopsis thaliana heat-stable seed proteins: Enriching for LEA-type proteins by acid treatment. J. Mass Spectrom. 2007, 42, 1485–1495. [Google Scholar] [CrossRef] [PubMed]
- Hundertmark, M.; Hincha, D.K. LEA (Late Embryogenesis Abundant) proteins and their encoding genes in Arabidopsis thaliana. BMC Genom. 2008, 9, 118–139. [Google Scholar] [CrossRef] [PubMed]
- Garay-Arroyo, A.; Colmenero-Flores, J.M.; Garciarrubio, A.; Covarrubias, A.A. Highly hydrophilic proteins in prokaryotes and eukaryotes are common during conditions of water deficit. J. Biol. Chem. 2000, 275, 5668–5674. [Google Scholar] [CrossRef] [PubMed]
- Yale, J.; Bohnert, H.J. Transcript expression in Saccharomyces cerevisiae at high salinity. J. Biol. Chem. 2001, 276, 15996–16007. [Google Scholar] [CrossRef]
- Battaglia, M.; Olvera-Carrillo, Y.; Garciarrubio, A.; Campos, F.; Covarrubias, A.A. The enigmatic LEA proteins and other hydrophilins. Plant Physiol. 2008, 148, 6–24. [Google Scholar] [CrossRef] [PubMed]
- Wolkers, W.F.; McCready, S.; Brandt, W.F.; Lindsey, G.G.; Hoekstra, F.A. Isolation and characterization of a D-7 LEA protein from pollen that stabilizes glasses in vitro. Biochim. Biophys. Acta Protein Struct. Mol. Enzymol. 2001, 1544, 196–206. [Google Scholar] [CrossRef]
- Wang, M.; Li, P.; Li, C.; Pan, Y.; Jiang, X.; Zhu, D.; Zhao, Q.; Yu, J. SiLEA14, a novel atypical LEA protein, confers abiotic stress resistance in foxtail millet. BMC Plant Biol. 2014, 14, 290–305. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Wu, L.; Wang, J.; Sun, J.; Xia, X.; Geng, X.; Wang, X.; Xu, Z.; Xu, Q. Genome sequencing of rice subspecies and genetic analysis of recombinant lines reveals regional yield- and quality-associated loci. BMC Biol. 2018, 16, 102. [Google Scholar] [CrossRef] [PubMed]
- Hirsch, C.N.; Hirsch, C.D.; Brohammer, A.B.; Bowman, M.J.; Soifer, I.; Barad, O.; Shem-Tov, D.; Baruch, K.; Lu, F.; Hernandez, A.G.; et al. Draft assembly of elite inbred line ph207 provides insights into genomic and transcriptome diversity in maize. Plant Cell 2016, 28, 2700–2714. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Fan, G.; Lu, C.; Xiao, G.; Zou, C.; Kohel, R.J.; Ma, Z.; Shang, H.; Ma, X.; Wu, J.; et al. Genome sequence of cultivated Upland cotton (Gossypium hirsutum TM-1) provides insights into genome evolution. Nat. Biotechnol. 2015, 33, 524–530. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.S.; Zhu, H.B.; Jin, G.L.; Liu, H.L.; Wu, W.R.; Zhu, J. Genome-scale identification and analysis of LEA genes in rice (Oryza sativa L.). Plant Sci. 2007, 172, 414–420. [Google Scholar] [CrossRef]
- Magwanga, R.O.; Lu, P.; Kirungu, J.N.; Lu, H.; Wang, X.; Cai, X.; Zhou, Z.; Zhang, Z.; Salih, H.; Wang, K.; et al. Characterization of the late embryogenesis abundant (LEA) proteins family and their role in drought stress tolerance in upland cotton. BMC Genet. 2018, 19, 6–37. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Cao, J. Late Embryogenesis Abundant (LEA) gene family in maize: Identification, evolution, and expression profiles. Plant Mol. Biol. Rep. 2016, 34, 15–28. [Google Scholar] [CrossRef]
- Galau, G.A.; Wang, H.Y.; Hughes, D.W. Cotton Lea5 and Lea14 encode atypical late embryogenesis-abundant proteins. Plant Physiol. 1993, 101, 695–696. [Google Scholar] [CrossRef]
- Maitra, N.; Cushman, J.C. Isolation and characterization of a drought-induced soybean cDNA encoding a D95 family late-embryogenesis-abundant protein. Plant Physiol. 1994, 106, 805–806. [Google Scholar] [CrossRef] [PubMed]
- Zegzouti, H.; Jones, B.; Marty, C.; Lelievre, J.M.; Latche, A.; Pech, J.C.; Bouzayen, M. ER5, a tomato cDNA encoding an ethylene-responsive LEA-like protein: Characterization and expression in response to drought, ABA and wounding. Plant Mol. Biol. 1997, 35, 847–854. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.S.; Lee, J.H.; Kim, J.J.; Kim, C.H.; Jun, S.S.; Hong, Y.N. Molecular and functional characterization of CaLEA6, the gene for a hydrophobic LEA protein from Capsicum annuum. Gene 2005, 344, 115–123. [Google Scholar] [CrossRef] [PubMed]
- Park, S.C.; Kim, Y.H.; Jeong, J.C.; Kim, C.Y.; Lee, H.S.; Bang, J.W.; Kwak, S.S. Sweetpotato late embryogenesis abundant 14 (IbLEA14) gene influences lignification and increases osmotic- and salt stress-tolerance of transgenic calli. Planta 2011, 233, 621–634. [Google Scholar] [CrossRef] [PubMed]
- He, S.; Tan, L.; Hu, Z.; Chen, G.; Wang, G.; Hu, T. Molecular characterization and functional analysis by heterologous expression in E. coli under diverse abiotic stresses for OsLEA5, the atypical hydrophobic LEA protein from Oryza sativa L. Mol. Genet. Genom. 2012, 287, 39–54. [Google Scholar] [CrossRef]
- Sharma, A.; Kumar, D.; Kumar, S.; Rampuria, S.; Reddy, A.R.; Kirti, P.B. Ectopic expression of an atypical hydrophobic group 5 LEA protein from wild peanut, Arachis diogoi confers abiotic stress tolerance in tobacco. PLoS ONE 2016, 11, e0150609. [Google Scholar] [CrossRef]
- FAOSTAT. 2017. Available online: http://www.fao.org/faostat/en/#data/QC (accessed on 6 December 2018).
- Daryanto, S.; Wang, L.; Jacinthe, P.A. Global synthesis of drought effects on maize and wheat production. PLoS ONE 2016, 11, e0156362. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wang, L.; Xing, X.; Sun, L.; Pan, J.; Kong, X.; Zhang, M.; Li, D. ZmLEA3, a multifunctional group 3 LEA protein from maize (Zea mays L.), is involved in biotic and abiotic stresses. Plant Cell Physiol. 2013, 54, 944–959. [Google Scholar] [CrossRef]
- Topping, J.F. Tobacco transformation. In Plant Virology Protocols: From Virus Isolation to Transgenic Resistance; Foster, G.D., Taylor, S.C., Eds.; Humana Press: Totowa, NJ, USA, 1998; pp. 365–372. [Google Scholar]
- Frame, B.; Main, M.; Schick, R.; Wang, K. Genetic transformation using maize immature zygotic embryos. In Plant Embryo Culture: Methods and Protocols; Thorpe, T.A., Yeung, E.C., Eds.; Humana Press: Totowa, NJ, USA, 2011; pp. 327–341. [Google Scholar]
- Tamura, K.; Filipski, A.; Peterson, D.; Stecher, G.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Goyal, K.; Walton, L.J.; Tunnacliffe, A. LEA proteins prevent protein aggregation due to water stress. Biochem. J. 2005, 388, 151–157. [Google Scholar] [CrossRef]
- Kumar, M.; Choi, J.; An, G.; Kim, S.R. Ectopic expression of OsSta2 enhances salt stress tolerance in rice. Front. Plant Sci. 2017, 8, 316. [Google Scholar] [CrossRef] [PubMed]
- Kumar, M.; Lee, S.C.; Kim, J.Y.; Kim, S.J.; Aye, S.S.; Kim, S.R. Over-expression of dehydrin gene, OsDhn1, improves drought and salt stress tolerance through scavenging of reactive oxygen species in rice (Oryza sativa L.). J. Plant Biol. 2014, 57, 383–393. [Google Scholar] [CrossRef]

© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Minh, B.M.; Linh, N.T.; Hanh, H.H.; Hien, L.T.T.; Thang, N.X.; Hai, N.V.; Hue, H.T.T. A LEA Gene from a Vietnamese Maize Landrace Can Enhance the Drought Tolerance of Transgenic Maize and Tobacco. Agronomy 2019, 9, 62. https://doi.org/10.3390/agronomy9020062
Minh BM, Linh NT, Hanh HH, Hien LTT, Thang NX, Hai NV, Hue HTT. A LEA Gene from a Vietnamese Maize Landrace Can Enhance the Drought Tolerance of Transgenic Maize and Tobacco. Agronomy. 2019; 9(2):62. https://doi.org/10.3390/agronomy9020062
Chicago/Turabian StyleMinh, Bui Manh, Nguyen Thuy Linh, Ha Hong Hanh, Le Thi Thu Hien, Nguyen Xuan Thang, Nong Van Hai, and Huynh Thi Thu Hue. 2019. "A LEA Gene from a Vietnamese Maize Landrace Can Enhance the Drought Tolerance of Transgenic Maize and Tobacco" Agronomy 9, no. 2: 62. https://doi.org/10.3390/agronomy9020062
APA StyleMinh, B. M., Linh, N. T., Hanh, H. H., Hien, L. T. T., Thang, N. X., Hai, N. V., & Hue, H. T. T. (2019). A LEA Gene from a Vietnamese Maize Landrace Can Enhance the Drought Tolerance of Transgenic Maize and Tobacco. Agronomy, 9(2), 62. https://doi.org/10.3390/agronomy9020062




