Molecular Marker-Assisted Breeding of High-Quality and Salt-Tolerant Hybrid Japonica Rice Combination Shenyanyou 1
Abstract
1. Introduction
2. Materials and Methods
2.1. Test Materials and Field Cultivation Management
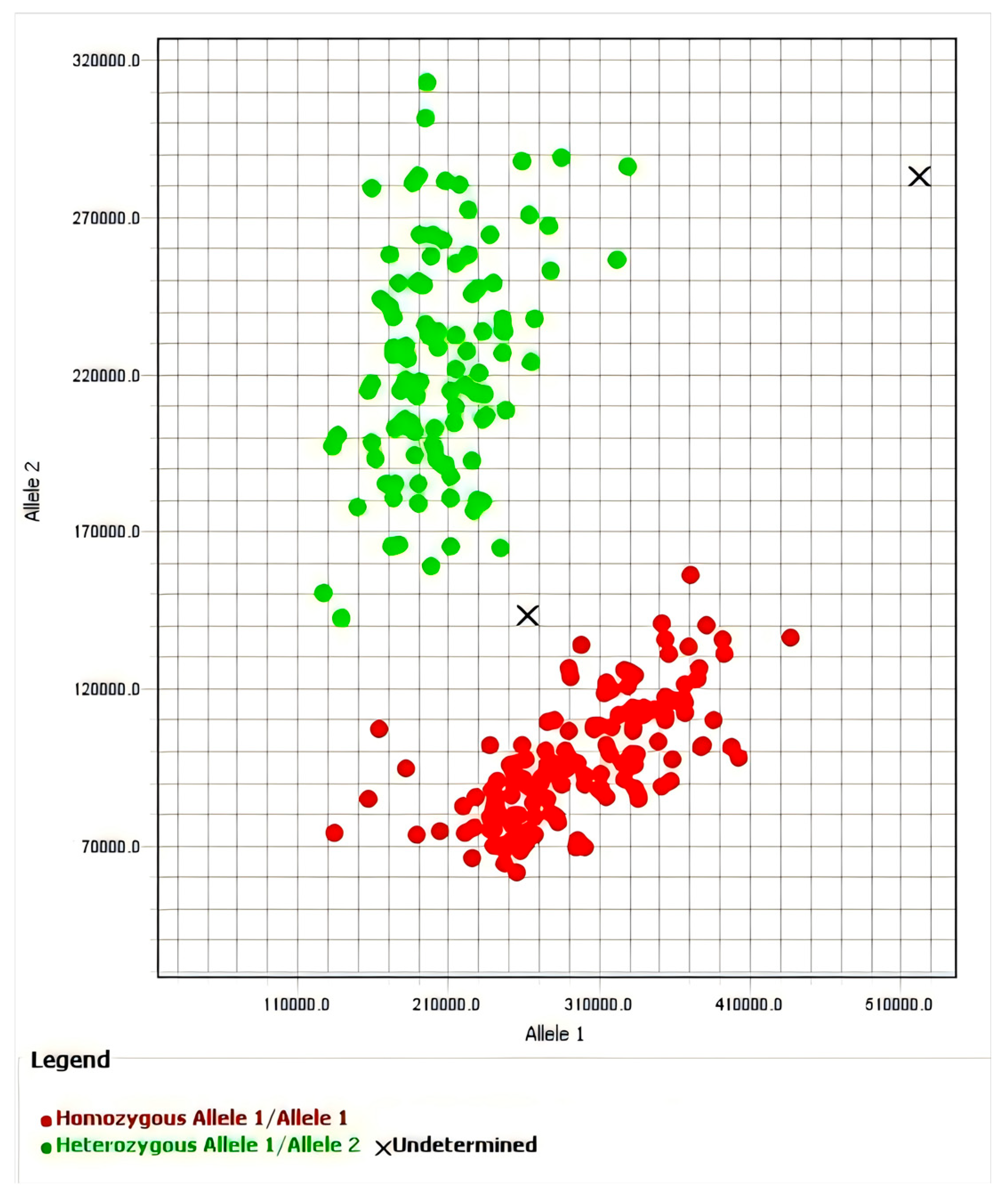
2.2. KASP Molecular Marker Detection
2.3. Rice Anther Culture
2.4. Gene Chip Detection
- A.
- DNA Fragmentation: Amplified DNA was enzymatically fragmented into small segments at 37 °C for 1 h.
- B.
- DNA Purification: Fragmented DNA was mixed with isopropanol and incubated at 4 °C for 30 min. Samples were centrifuged at 3000× g for 20 min. Supernatants were discarded, and pellets were air-dried at room temperature for 1 h.
- C.
- DNA Resuspension: Pellets were resuspended in RA1 buffer (containing 10–30% formamide) and incubated at 48 °C for 1 h.
- D.
- DNA Denaturation: Resuspended DNA was heat-denatured at 95 °C for 20 min using a dry bath.
- E.
- Chip Hybridization: Denatured DNA was loaded onto designated positions of the SNP array and hybridized at 48 °C for 16–24 h in a hybridization oven.
- F.
- Single-Base Extension (SBE) and Staining: Hybridized chips underwent single-base extension with fluorescence-labeled dNTPs. Post-extension staining amplified fluorescence signals for enhanced detection.
- G.
- Array Scanning: Processed chips were scanned using an Illumina iScan® system, with a scanning time of approximately 0.5 min for each sample.
- H.
- Data Genotyping: Raw fluorescence data were analyzed and genotyped using Illumina GenomeStudio® software (v2.0 or higher).
2.5. Salt Tolerance Evaluation of Shenyanyou 1
3. Results
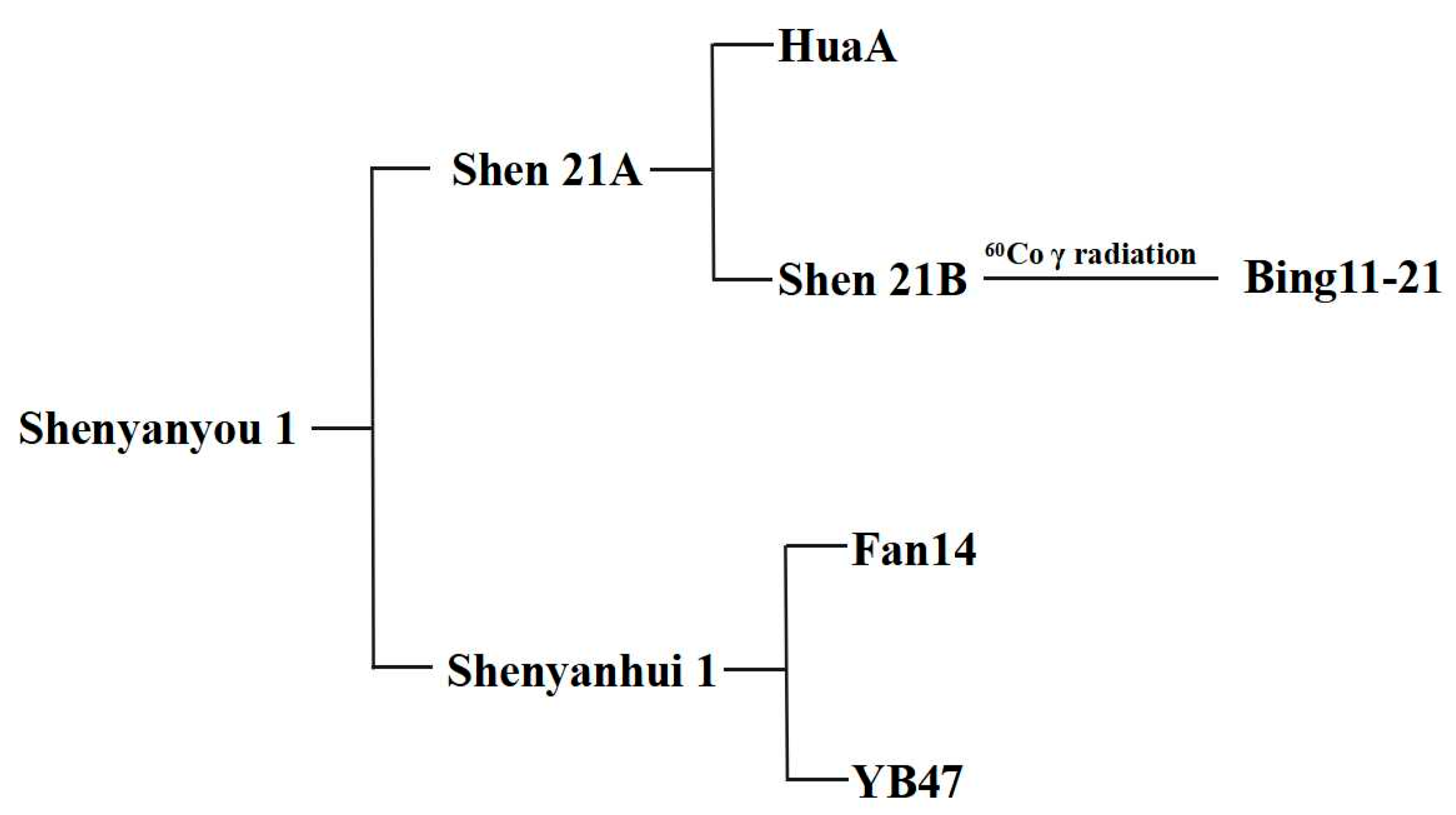
3.1. Breeding Process of Shenyanhui 1
3.2. Breeding Process of Shenyanyou 1
3.3. Characteristics of Shenyanyou 1
3.3.1. Main Agronomic Traits of Shenyanyou 1
3.3.2. The Salt Tolerance Evaluation of Shenyanyou 1
3.3.3. Quality of Shenyanyou 1
3.4. Production of Shenyanyou 1
3.5. Genotypic Analysis of Shenyanyou 1
4. Discussion
4.1. Shenyanyou 1 Provides an Excellent Option for Salt Tolerance Improvement in Saline–Alkali Land
4.2. Modern Biotechnological Breeding Techniques Have Accelerated the Development of New Salt-Tolerant Rice Varieties
4.3. Enhanced Comprehensive Evaluation of Salinity and Alkalinity Tolerance in Shenyanyou 1
4.4. Pyramiding Major and Minor Effect Salt Tolerance Genes/QTLs to Enhance Saline–Alkali Tolerance of Bred Varieties
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Wang, C.L.; Zhang, Y.D.; Zhao, L.; Lu, K.; Zhu, Z.; Chen, T.; Zhao, Q.Y.; Yao, S.; Zhou, L.H.; Zhao, C.F.; et al. Research status, problems and suggestions on saline-alkali tolerant rice. China Rice 2019, 25, 1–6. [Google Scholar]
- Leng, C.X.; Zheng, F.Y.; Zhao, B.P.; Liu, H.Y.; Wang, Y.J. Advances on Alkaline Tolerance of Rice. Biotechnol. Bull. 2020, 36, 103–111. [Google Scholar]
- Ma, G.H.; Zheng, D.F.; Mu, D.W.; Wang, F.B.; Dai, Q.G.; Wei, Z.W.; Feng, N.J.; Wang, C.L. Research Progress and Prospect of Saline-alkali Tolerant Rice. Hybrid Rice 2024, 39, 1–10. [Google Scholar]
- Wang, S.Y.; Mao, T.; Zhang, L.L.; Li, X.; Liu, Y.; Zhao, Y.Z.; Ni, S.J.; Zhong, S.C.; Wang, B.Q.; Zhang, Z. Advances in Salt Tolerance Breeding of Rice. North Rice 2021, 51, 48–51. [Google Scholar] [CrossRef]
- Zhao, J.W.; Lei, C.S.; Liu, Y.Q.; Zhang, L.; Cheng, Y.F.; Wang, X.L. Primary Exploration on Saline-alkali Tolerance of Haidao 86 in Germination Period. China Rice 2018, 24, 87–92. [Google Scholar]
- Zhang, Q.; Jiang, Y.J.; Lin, Q.S.; Yu, N.; Wei, J.; Gao, Y.; Liu, Z.X. Breeding and Application of Salt-tolerant Rice Variety Guanghong 3. China Rice 2024, 30, 133–135. [Google Scholar]
- Niu, F.A.; Zhou, J.H.; Cao, L.M.; Cheng, C.; Tu, R.J.; Hu, X.J.; Yang, J.; Sun, B.; Luo, Z.Y.; Zhao, Z.P.; et al. Development and Utilization of KASP Marker for Low Amylose Content Gene, Wxmq, in Rice. Mol. Plant Breed. 2019, 17, 8125–8131. [Google Scholar]
- Shao, Y.; Peng, Y.; Mao, B.G.; Lv, Q.M.; Yuan, D.Y.; Liu, X.L.; Zhao, B.R. Allelic variations of the Wx locus in cultivated rice and their use in the development of hybrid rice in China. PLoS ONE 2020, 15, e0232279. [Google Scholar] [CrossRef]
- Addison, C.K.; Angira, B.; Kongchum, M.; Harrell, D.L.; Baisakh, N.; Linscombe, S.D.; Famoso, A.N. Characterization of Haplotype Diversity in the BADH2 Aroma Gene and Development of a KASP SNP Assay for Predicting Aroma in U.S. Rice. Rice 2020, 13, 47–55. [Google Scholar] [CrossRef]
- Qiu, S.Q.; Lu, Q.; Yu, H.H.; Ni, X.M.; Zhang, G.Y.; He, H.; Xie, W.B.; Zhou, F.S. The development and application of rice whole genome selection breeding platform. Chin. Bull. Life Sci. 2018, 30, 1120–1128. [Google Scholar]
- Ding, Z.Q.; Pan, Y.Y.; Shi, Y.; Huang, H.X. Comprehensive Evaluation and Comparative Analysis of Jiahe Series Long-Grain japonica Rice with High Eating Quality Based on Gene Chip Technology. Chin. J. Rice Sci. 2024, 38, 397–408. [Google Scholar]
- Chen, H.D.; Xie, W.B.; He, H.; Yu, H.H.; Chen, W.; Li, J.; Yu, R.B.; Yao, Y.; Zhang, W.H.; He, Y.Q.; et al. A high-density SNP genotyping array for rice biology and molecular breeding. Mol. Plant 2014, 7, 541–553. [Google Scholar] [CrossRef]
- Cao, L.M.; Cheng, C.; Zhou, J.H.; Chu, H.W.; Niu, F.A.; Yuan, Q. Development and prospect of hybrid japonica rice industry in Shanghai. China Seed Ind. 2018, 9, 19–22. [Google Scholar]
- Niu, F.A.; Cheng, C.; Sun, B.; Chu, H.W.; Zhou, J.H.; Deng, H.Z.; Zhong, Q.Q.; Cao, L.M. Breeding and utilization of japonica male sterile line Shen 21A with high quality, disease resistance and high combining ability. Acta Agric. Shanghai 2022, 38, 32–36. [Google Scholar]
- Li, Y.; Cheng, C.; Zhou, J.H.; Niu, F.A.; Yao, Y.; Huang, Y.W.; Zhang, A.P.; Sun, B.; Cao, L.M.; Chu, H.W. Development and Utilization of KASP Marker for Salt Tolerance Gene SKC1 in Rice. Mol. Plant Breed. 2024, 22, 2923–2929. [Google Scholar]
- NY/T 3692-2020; Technical Code of Practice for Identification of Salt Tolerance in Rice. China Agriculture Press: Beijing, China, 2020.
- Komatsu, K.; Maekawa, M.; Ujiie, S.; Satake, Y.; Furutani, I.; Okamoto, H.; Shimamoto, K.; Kyozuka, J. LAX and SPA: Major regulators of shoot branching in rice. Proc. Natl. Acad. Sci. USA 2003, 100, 11765–11770. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.K.; Li, S.; Liu, Q.; Wu, K.; Zhang, J.Q.; Wang, S.S.; Wang, Y.; Chen, X.B.; Zhang, Y.; Gao, C.X.; et al. The OsSPL16-GW7 regulatory module determines grain shape and simultaneously improves rice yield and grain quality. Nat. Genet. 2015, 47, 949–954. [Google Scholar] [CrossRef]
- Wang, Y.X.; Xiong, G.S.; Hu, J.; Jiang, L.; Yu, H.; Xu, J.; Fang, Y.X.; Zeng, L.J.; Xu, E.B.; Xu, J.; et al. Copy number variation at the GL7 locus contributes to grain size diversity in rice. Nat. Genet. 2015, 47, 944–948. [Google Scholar] [CrossRef] [PubMed]
- Tong, J.P.; Han, Z.S.; Han, A.N.; Liu, X.J.; Zhang, S.Y.; Fu, B.Y.; Hu, J.; Su, J.P.; Li, S.Q.; Wang, S.J.; et al. Sdt97: A point mutation in the 5′ untranslated region confers semidwarfism in rice. G3 Genes Genomes Genet. 2016, 6, 1491–1502. [Google Scholar] [CrossRef]
- Wang, Q.; Liu, Y.Q.; He, J.; Zheng, X.M.; Hu, J.L.; Liu, Y.L.; Dai, H.M.; Zhang, Y.X.; Wang, B.X.; Wu, W.X.; et al. STV11 encodes a sulphotransferase and confers durable resistance to rice stripe virus. Nat. Commun. 2014, 5, 4768–4775. [Google Scholar] [CrossRef]
- Shabbir, G.; Nazir, H.; Bhatti, M.K.; Afzal, A.; Shakir, M.A. Salt tolerance potential of some selected fine rice cultivars. J. Biol. Sci. 2001, 1, 1175–1177. [Google Scholar] [CrossRef]
- Singh, R.K.; Kota, S.; Flowers, T.J. Salt tolerance in rice: Seedling and reproductive stage QTL mapping come of age. Theor. Appl. Genet. 2021, 134, 3495–3533. [Google Scholar] [CrossRef]
- Lu, H.Y.; Zhou, L.; Lin, F.; Wang, R.; Wang, F.G.; Zhao, H. Development of efficient KASP molecular markers based on high throughput sequencing in maize. Acta Agron. Sin. 2019, 45, 872–878. [Google Scholar]
- Semagn, K.; Babu, R.; Hearne, S.; Olsen, M. Single nucleotide polymorphism genotyping using Kompetitive Allele Specific PCR (KASP): Overview of the technology and its application in crop improvement. Mol. Breed. 2014, 33, 1–14. [Google Scholar] [CrossRef]
- Bryan, G.T.; Wu, K.S.; Farrall, L.; Jia, Y.L.; Hershey, H.P.; McAdams, S.A.; Faulk, K.N.; Donaldson, G.K.; Tarchini, R.; Valent, B. A single amino acid difference distinguishes resistant and susceptible alleles of the rice blast resistance gene Pi-ta. Plant Cell 2000, 12, 2033–2046. [Google Scholar] [CrossRef]
- Zhou, B.; Qu, S.H.; Liu, G.F.; Dolan, M.; Sakai, H.; Lu, G.D.; Bellizzi, M.; Wang, G.L. The eight amino-acid differences within three leucine-rich repeats between Pi2 and Piz-t resistance proteins determine the resistance specificity to magnaporthe grisea. Mol. Plant-Microbe Interact. 2006, 19, 1216–1228. [Google Scholar] [CrossRef]
- Chen, H.L.; Chen, B.T.; Zhang, D.P.; Xie, Y.F.; Zhang, Q.F. Pathotypes of pyricularia grisea in rice fields of central and southern China. Plant Dis. 2001, 85, 843–850. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.X.; Zhang, S.B.; Hu, J.X.; Sun, W.X.; Padilla, J.; He, Y.L.; Li, Y.; Yin, Z.Y.; Liu, X.Y.; Wang, W.H.; et al. Phosphorylation-guarded light-harvesting complex II contributes to broad-spectrum blast resistance in rice. Proc. Natl. Acad. Sci. USA 2019, 116, 17572–17577. [Google Scholar] [CrossRef] [PubMed]
- Ren, Z.H.; Gao, J.P.; Li, L.G.; Cai, X.L.; Huang, W.; Chao, D.Y.; Zhu, M.Z.; Wang, Z.Y.; Luan, S.; Lin, H.X. A rice quantitative trait locus for salt tolerance encodes a sodium transporter. Nat. Genet. 2005, 37, 1141–1146. [Google Scholar] [CrossRef]
- Ganapati, R.K.; Naveed, S.A.; Zafar, S.; Wang, W.S.; Xu, J.L. Saline-alkali tolerance in rice: Physiological response, molecular mechanism, and QTL identification and application to breeding. Rice Sci. 2022, 29, 412–434. [Google Scholar] [CrossRef]
- Sun, P.Y.; Zhang, W.H.; Shu, F.; He, Q.; Zhang, L.; Yang, Z.H.; Peng, Z.R.; Xie, Y.; Deng, H.F. rehensive evaluation of salt-alkali tolerance of rice germplasms at germination and seedling stages and analysis of salt-tolerant genes. Chin. J. Biotech. 2022, 38, 252–263. [Google Scholar]
- Guan, B.; Zhou, D.; Zhang, H.; Tian, Y.; Japhet, W.; Wang, P. Germination responses of Medicago ruthenica seeds to salinity, alkalinity, and temperature. J. Arid. Environ. 2009, 73, 135–138. [Google Scholar] [CrossRef]
- Liang, W.J.; Ma, X.L.; Wan, P.; Liu, L.Y. Plant salt-tolerance mechanism: A review. Biochem. Biophys. Res. Commun. 2018, 495, 286–291. [Google Scholar] [CrossRef] [PubMed]


| Step | Description | Temperature | Time | No. of Cycle |
|---|---|---|---|---|
| 1 | Activation | 95 °C | 10 min | 1 |
| 2 | Denaturation | 95 °C | 20 s | 10 |
| Annealing/Elongation | 61–55 °C | 60 s | ||
| 3 | Denaturation | 95 °C | 20 s | 27 |
| Annealing/Elongation | 55 °C | 60 s | ||
| 4 | Read | 25 °C | 30 s | 1 |
| Treatments | Germination Rate/% | Average Germination Rate/% | Relative Salt Damage Rate/% | ||||
|---|---|---|---|---|---|---|---|
| I | II | III | IV | V | |||
| H2O | 89.0 | 86.0 | 92.0 | 94.0 | 88.0 | 89.8 | |
| 1.5% NaCl | 44.0 | 41.0 | 36.0 | 42.0 | 38.0 | 40.2 | 55.23 |
| Variety | Harvested Area (m2) | Fresh Grain Yield (kg) | Moisture Content (%) | Impurity Content (%) | Actual Yield (kg/hm2) | Yield Increase Rate (%) |
|---|---|---|---|---|---|---|
| W087 | 6.5 | 2.95 | 15.7 | 2.0 | 4387.5 | 45.4 |
| Shenyanyou 1 | 6.5 | 2.43 | 15.1 | 2.0 | 3640.5 | 20.7 |
| W103 | 6.5 | 2.2 | 13.6 | 2.0 | 3354.0 | 11.2 |
| Yuanzhong 62 | 6.5 | 2.07 | 15.5 | 2.0 | 3085.5 | 2.3 |
| Xingeng 2 | 6.5 | 2.02 | 13.7 | 2.0 | 3075.0 | 2.0 |
| Zhongbao 8 | 6.5 | 2 | 13.8 | 2.0 | 3042.0 | 0.8 |
| Yandao 21 (CK) | 6.5 | 1.99 | 14.1 | 2.0 | 3016.5 | / |
| Inspection Items | Unit | Standard Formulation | Test Result | Single Item Judgment | Testing Basis |
|---|---|---|---|---|---|
| Whole-head rice rate | % | ≥69.0 | 71.2 | Level 1 | NY/T 2334-2013 |
| Chalkiness | % | ≤5.0 | 4.2 | Level 3 | NY/T 2334-2013 |
| Transparency | Grade | ≤2 | 2 | Level 2 | NY/T 2334-2013 |
| Alkali spreading value | Grade | ≥6.0 | 6.3 | Level 3 | NY/T 83-2017 |
| Gel consistency | mm | ≥70 | 81 | Level 1 | GB/T 22294-2008 |
| Amylose content | % | 13.0–18.0 | 15.8 | Level 1 | NY/T 2639-2014 |
| Grain length | mm | / | 5.6 | / | NY/T 2334-2013 |
| Aspect ratio | / | / | 2.1 | / | NY/T 2334-2013 |
| Brown rice percentage | % | / | 82.3 | / | NY/T 83-2017 |
| Polished rice rate | % | / | 74.7 | / | NY/T 83-2017 |
| Chalky grain rate | % | / | 24 | / | NY/T 2334-2013 |
| Protein | Score | / | 6.65 | / | NY/T 596-2002 |
| Location | Fresh Grain Yield (kg/hm2) | Yield Increase Rate (%) | |
|---|---|---|---|
| Shenyanyou 1 | Huayou 14 | ||
| Kangfeng, Jinshan, Shanghai | 9214.5 | 9637.5 | −4.39 |
| Nanqiao, Fengxian, Shanghai | 10,150.2 | 9643.4 | 5.26 |
| Kunshan, Jiangsu | 9099.0 | 9009.0 | 1.00 |
| Hefei, Anhui | 7760.0 | 7847.5 | −1.12 |
| Average | 9055.9 | 9034.3 | 0.24 |
| Gene | Chr. | Traits Regulated | Function of Superior Alleles | Parental Lines Carrying Superior Alleles |
|---|---|---|---|---|
| LAX1 | 1 | Yield components | Increases grain number per panicle | Paternal Line |
| GW5 | 5 | Yield components | Increases grain width | Both Parental Lines |
| GW7 | 7 | Yield components | Increases grain length | Paternal Line |
| GW8/OsSPL16 | 8 | Yield components | Increases grain weight | Paternal Line |
| Wxb | 6 | Taste quality | Reduces amylose content | Both Parental Lines |
| SKC1 | 1 | Abiotic Stress | Enhances salt tolerance | Paternal Line |
| NRT1.1B | 10 | Abiotic Stress | Improves nitrogen use efficiency | Paternal Line |
| Pi2 | 6 | Biotic Stress | Enhances blast resistance | Both Parental Lines |
| Pita | 12 | Biotic Stress | Enhances blast resistance | Maternal Line |
| Pizt | 6 | Biotic Stress | Enhances blast resistance | Paternal Line |
| LHCB5 | 11 | Biotic Stress | Enhances blast resistance | Paternal Line |
| STV11 | 11 | Biotic Stress | Confers durable resistance to rice stripe virus | Maternal Line |
| Xa26/Xa3 | 11 | Biotic Stress | Enhances bacterial blight resistance | Paternal Line |
| qLTG3-1 | 3 | Abiotic Stress | Enhances cold tolerance and low-temperature germination vigor | Maternal Line |
| OsFRO1 | 4 | Abiotic Stress | Improves iron tolerance | Maternal Line |
| qUVR-10 | 10 | Abiotic Stress | Enhances light repair activity | Maternal Line |
| BET1 | 4 | Abiotic Stress | Increases boron toxicity tolerance | Paternal Line |
| SaF | 1 | Fertility | Confers wide-compatibility (japonica compatibility) | Both Parental Lines |
| Sdt97 | 6 | Plant type | Confers semi-dwarfism | Maternal Line |
| Rf2 | 2 | Yield components | Restores fertility | Paternal Line |
| Hd17/Hd3b | 6 | Heading date | Delaying heading date | Paternal Line |
| sh4 | 4 | Others | Prevents grain shattering | Both Parental Lines |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Niu, F.; Zhang, A.; Cheng, C.; Chu, H.; Fang, J.; Zhou, J.; Sun, B.; Dai, Y.; Zhang, J.; Feng, Z.; et al. Molecular Marker-Assisted Breeding of High-Quality and Salt-Tolerant Hybrid Japonica Rice Combination Shenyanyou 1. Agronomy 2025, 15, 2006. https://doi.org/10.3390/agronomy15082006
Niu F, Zhang A, Cheng C, Chu H, Fang J, Zhou J, Sun B, Dai Y, Zhang J, Feng Z, et al. Molecular Marker-Assisted Breeding of High-Quality and Salt-Tolerant Hybrid Japonica Rice Combination Shenyanyou 1. Agronomy. 2025; 15(8):2006. https://doi.org/10.3390/agronomy15082006
Chicago/Turabian StyleNiu, Fuan, Anpeng Zhang, Can Cheng, Huangwei Chu, Jun Fang, Jihua Zhou, Bin Sun, Yuting Dai, Jianming Zhang, Zhizun Feng, and et al. 2025. "Molecular Marker-Assisted Breeding of High-Quality and Salt-Tolerant Hybrid Japonica Rice Combination Shenyanyou 1" Agronomy 15, no. 8: 2006. https://doi.org/10.3390/agronomy15082006
APA StyleNiu, F., Zhang, A., Cheng, C., Chu, H., Fang, J., Zhou, J., Sun, B., Dai, Y., Zhang, J., Feng, Z., & Cao, L. (2025). Molecular Marker-Assisted Breeding of High-Quality and Salt-Tolerant Hybrid Japonica Rice Combination Shenyanyou 1. Agronomy, 15(8), 2006. https://doi.org/10.3390/agronomy15082006







