Abstract
Pasture dieback is a syndrome of unknown cause affecting grasses in Australia, creating significant economic losses to farmers by reducing available livestock feed and paddock carrying capacity. RC3 is a commercial plant growth stimulant tri-sodium salt of trimercapto-S-triazine (TMT) and potassium humate as active ingredients. TMT is commonly used for soil and wastewater remediation by capturing and binding heavy metals, while potassium humate is an organic compound used as a plant growth promoter. We investigated the ability of RC3 to restore soil health and productivity under pasture dieback conditions. RC3 was applied on pasture dieback affected paddock replicate plots once, at a rate of 4 mL/m2, and soil core samples were taken weekly to analyse microbial communities. Plants were collected regularly to measure dry matter and plant morphometrics. Twenty weeks after a single application, dry matter increased in RC3 plots by 900 kg/ha compared to control plots, and at week 48, eleven months after the single application, RC3 plots showed a trend of more grass and dicot species than the control. Morphometric measures suggest minor improvements in dicotyledon plants. Alpha diversity did not change with the application of RC3. Temporal correlation analysis shows that RC3 steadily reduced the presence of genera predominant in poor soils and with extreme environmental conditions over time and prevented the decline of beneficial genera, such as Marmoricola, Actinomadura, Dactylosporangium, and mle1-7.
1. Introduction
Worldwide demand for beef as a source of nutritional protein is increasing [1]. The global cattle herd consists of almost one billion animals [2], providing 45% of the global protein supply for human consumption [3]. Australia had around 25 million cattle in 2019, with 45% raised by Queensland farmers [4].
Livestock production is highly dependent on feed availability, as this constitutes 70% of the total expenses of production [5]. Feed quality for ruminants can fluctuate according to feed intake [6], plant life stage [7], pasture composition [8], availability, and physiological stage of the animal. Queensland pastures are the source of feed for grass-fed cattle. The pasture growth and availability change according to the weather patterns [9]. Pasture quality depends on diversity and composition to deliver high nutritional value plants [10], management practices, and soil health.
Pasture dieback is a syndrome that causes plant decomposition and death, impacting grass production in Australia, affecting different grass species across distinct soil types, reducing the animal carrying capacity of paddocks, and considerably altering pasture availability and quality [11]. This disease appears as the yellowing or reddening on the tip of older leaves, soon affecting the rest of the plant. Necrotic and stunted roots are characteristics of this disorder, followed by plant death [12]. In many cases, the affected zone starts in patches of about 5 m2, spreading to encompass entire paddocks [13]. Although there are many working theories about the source of this disease, including insect infestation, climate change, reduced soil organic matter, damaged soil microbiota, poor management practices, loss of diversity, soil nutrient availability, and drought, the primary causes of this disease are yet unknown [13,14].
Climate change is gaining momentum in Queensland [15], the average temperature has increased by 1.5 °C compared to 1910, and the weather is becoming noticeably more extreme and unpredictable. The Central Queensland region where this study was completed has a sub-tropical climate with hot summers and warm winters, and the rainfall is highly seasonal, occurring primarily in the summer. Agriculture experts worldwide are developing policies to mitigate the effects of global warming and to prepare strategies to maintain agriculture sustainability under projected future environmental extremes [16,17,18]. These strategies include the development of heat and water stress tolerant plant varieties, improved management practices, more efficient water use, improved pest management, and communication of these strategies to farmers. Climate change has influenced Australian pastures [19], and climate simulation modelling predicts farms running at operation losses in the near future [19].
Facing major losses in pasture and reduced cattle carrying capacity, the farmers are looking for means to restore or improve the pasture quality and quantity. While it is clear that the permanent reversal of pasture dieback is unlikely to be a short-term investigation, plant growth stimulants present themselves as a possible solution capable of providing immediate results and quantifiable short-term pasture improvements. Our previous studies show significant pasture improvements due to some growth stimulants, such as sea minerals [20]. We investigated the capability of RC3, a commercial product whose active ingredients, potassium humate and TMT, are known to aid in soil remediation and health. The product claims that when combined in RC3, these two main ingredients amplify each other’s beneficial effects and perform better in synergy than in individual use and that it is a potent plant growth promotant [21].
The primary use of tri-sodium salt of TMT is the removal of heavy metals from wastewater and contaminated soils. However, it is widely used in several industries to precipitate and bind copper, cadmium, mercury, lead, silver, and nickel [22,23,24,25,26,27,28,29]. The novel use of RC3 is as a plant growth stimulant [16] in fruits. Products containing sulphur are used as plant growth stimulants to help plants cope with heat stress and to defend the plant from pathogenic microorganisms [30,31,32].
Potassium humate is a natural component of soil humus [33]. It affects soil’s physical and chemical properties by chelating heavy metals and enhancing soil biological and chemical properties, making nutrients available for plant use and helping the plant cope with environmental stress [34]. Humate has a positive effect on plant roots, especially in lateral roots and root hair initiation, as well as density, which enhances nutrient absorption [35]. Humate also interacts with the microbial community by recruiting and stimulating plant-associated beneficial bacteria to protect the plant from pathogens, thus affecting the soil’s microbial profile and bacteria reproduction [36,37].
We hypothesise that in pasture dieback, the weather extremes have disturbed microbial communities in the soil, especially the bacterial community, which is much more sensitive to heat than the fungal community [38]. The disturbance in microbial balance in the soil can reflect on the plant’s health. RC3 and both of its main ingredients have the ability to restore soil chemistry and microbiota and improve plant growth. In this experiment, we applied RC3 to pasture dieback-affected soil to assess its effects on the soil microbial community, plants, and possible use in the remediation of damaged pasture and soil. We used a temporal design to evaluate the capacity for long-term effects and 16S amplicon soil microbiota analysis, soil nutrient, pasture dry matter, and plant morphometric analysis to evaluate microbiota–mineral interactions, plant benefits, and long-term pasture capacity improvements.
2. Materials and Methods
2.1. Farm Description and Location
The property used for this experimental trial is located in Garnant, Queensland. The property is an organic cattle enterprise with up to 800 Bos indicus × Bos taurus breed beef cattle. The paddock used for the trial (38 ha) was productive until it was affected by pasture dieback in 2017. The paddock comprised multiple grass species, including Buffelgrass (Cenchrus ciliaris), forest blue grass (Bothriochloa bladhii ssp. glabra), black spear grass (Heteropogon contortus) Queensland bluegrass (Dichnthium sericeum), kangaroo grass (Themeda triandra), Indian couch (Bothriochloa pertusa), green panic (Megathyrsus maximus), and legume species like seca stylo (Stylosanthes scabra). The first signs of pasture dieback appeared in 2017 when the grass started to die off. By 2019, there was no new grass growth, and the paddock was overwhelmed with weeds. In 2020, the paddock seemed to be recovering with a minimal amount of grass and some native legume species such as phasey bean (Macroptilium lathyroides), and it seemed to be transitioning through several dominant species. By 2021, when this study was conducted, the predominant species were weeds such as wild sage (Salvia verbenaca L.), some woody plants such as currant bush (Carissa ovata and Carissa lanceolata), and the only grass species present was Indian couch (Bothriochloa pertusa). The cattle avoided dieback-affected pasture grazing only on the remaining Indian couch, significantly reducing the cattle carrying capacity.
2.2. Experimental Plots
The experimental paddock used in this trial presented similar pasture dieback characteristics. The entire paddock was affected by pasture dieback, and no healthy control was available. The experimental plots used for this trial were 5 m by 5 m, separated by a buffer zone to avoid drifting the product on application. The treatments were randomised [39] between RC3 treated and untreated pasture dieback -affected control (CTR). We used three replicate plots (5 m × 5 m) per treatment. RC3 (Green Earth Technology, Gin Gin, 4671, Australia) was diluted at the rate of 2 mL per litre of water and was applied 50 litres per plot resulting in a final dose rate of 4 mL/m2. The cost of RC3 in 2022 was AUD $66 per litre. The CTR plots were sprayed with the same amount of untreated water simultaneously with treating RC3 plots. The basic climate conditions, including the temperature, rainfall, and solar exposure, for the duration of the trial are provided in Supplementary Material Figure S1.
2.3. Sampling and Measurement
After a single product application, soil sampling was done by the core soil collection method [40], using a T-bar core sampler at 15 cm depth. The samples were taken before application, then weekly for six weeks, and then fortnightly until week 20. The samples were kept on ice and placed in a −80 °C freezer. Plant samples for dry matter measurements were collected before the application (week 0) and at week 20, using a quadrat thrown twice per plot and cutting and collecting all plant material as close to the ground as possible using scissors. Due to the nature of this syndrome (affected plant matter disintegrated on contact), separation into pasture components was not possible at this stage. However, 48 weeks after the single application, separating into monocotyledon, dicotyledon and litter was possible. Plant samples were dried at 160 °C for 24 h, and the weights were recorded to calculate and compare dry matter per hectare.
Plant samples for morphological studies were taken at week 20 and again eleven months after (week 48). A representative grass and a dominant weed (wild sage) plant were taken entirely from each plot and kept on ice until measurements were taken. The root widths were measured using a caliper, while root length, leaf size and plant height were measured with a ruler. The number of tillers, branches, roots, and seed heads were counted.
2.4. Soil Mineral Analysis
Soil samples for mineral analysis were outsourced to the University of Western Australia’s Earth and Environment Analysis Laboratory (EEAL) located within the School of Agriculture and Environment. The soil samples were analysed by inductively coupled plasma optical emission spectroscopy (ICP–OES). The minerals measurements taken were total Carbon/Nitrogen, Al, Ca, Cd, Co, Cr, Cu, Fe, K, Mg, Mn, Mo, Na, Ni, P, Pb, S, Zn, pH, and EC (electric conductivity).
2.5. DNA Sequencing and Bioinformatics
Soil DNA was extracted using DNeasy PowerSoil Pro Kit (QIAGEN, Hilden, Germany) following the manufacturer’s recommended protocols. After DNA extraction, the 16S rRNA gene amplicon sequencing library was prepared by amplifying the V3-V4 hypervariable region using forward primer 338F (5′-ACTCCTACGGGAGGCAGCAG-3′) and reverse primer 806R (5′-GGACTACHVGGGTWTCTAAT-3′) with spacers, barcodes, and Illumina sequencing linkers. The sequencing was outsourced to Azenta Life Sciences (Suzhou, China) and performed using a 2 × 300 bp paired-end sequencing with the Illumina MiSeq system.
Data analysis was performed using Qiime2 software, with denoising and chimaera checking performed by Dada2. We used better quality read with a minimum Phred score of 20 over the length of 200 nt. The ASV level data were clustered to an OTU level using 98% similarity. The samples with fewer than 4000 sequences were removed from the analysis, leaving 57 samples. OTU level data were rarefied to 4000 sequences per sample and further explored using a range of R packages, including Phyloseq, Vegan, and Microecco. Plant data, including dry matter and plant morphometrics, were analysed and presented using GraphPad Prism v9. We used non-parametric Mann–Whitney tests for all morphometric measurements and dry matter calculations.
3. Results
3.1. Plant Dry Matter, Morphometric Measurements and Soil Minerals
Dry matter increased between week 0 and week 20 (Figure 1) and based on the difference of means, showed an insignificant increase of 1600 kg/ha in CTR. There was a significant increase (p = 0.029) of 2100 kg/ha in RC3-treated plots between week 0 and week 20 (Figure 1). At 48 weeks after the application, there was no improvement in total dry matter (Figure 2A). There was no statistically significant improvement (p < 0.05) in the amount of grass (Figure 2C). However, based on the mean yields, there was 182 kg/ha more grass on RC3 compared to the CTR plots (Figure 2D).
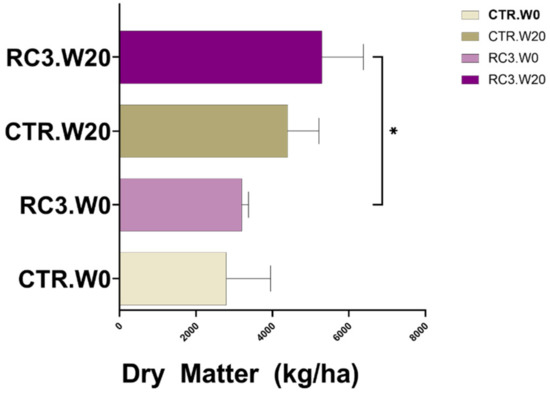
Figure 1.
Total dry matter before the application (W0) and at week 20. * p < 0.05.
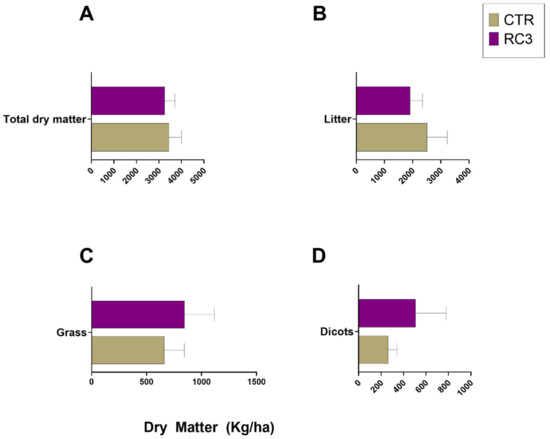
Figure 2.
Dry matter measured in kg/ha, panel (A) = total dry matter, (B) = litter, (C) = grass and (D) = dicotyledon.
Plant morphometric measurements were taken at week 20 and then again at week 48 to observe the long-term effect of the product. Although there were no statistically significant differences between RC3 and CTR plant measurements, in dicotyledons (Figure 3), based on a comparison of the group means, at week 20, there was a trend of higher root thickness, length of youngest leaf, the total number of roots, seed heads, number of branches, and longest root length. Eleven months later (W48), the leaf size, the number of branches and seed heads based on the group means remained higher in RC3 than in CTR plants, but this did not reach statistical significance. Grass morphology is provided in Supplementary Material Figure S2, and it shows a trend of a higher mean of the total number of roots and tillers in RC3 treated plots compared to control at 20 weeks post-application, but this did not persist to week 48 (Supplementary Material Figure S2).
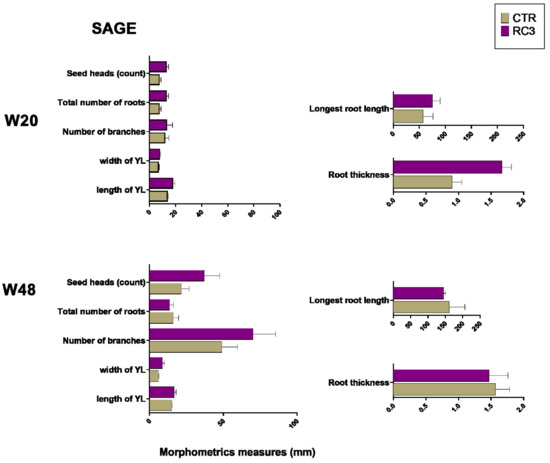
Figure 3.
Plant morphometric measurements at week 20 and week 48 for sage (dicotyledons representative species). YL = youngest leaf.
There were no significant differences in mineral concentrations before and 12 weeks after the application of RC3 (Supplementary Material Figure S3), although concentrations of Ca, Ni, Pb, C, and P were less in RC3 plots, while concentrations of K, Fe, Mg, and Mo were increased (Supplementary Material Figure S3).
3.2. Soil Microbial Community Structure
The most abundant genera in the soil were Rubrobacter, Solirubrobacterales 67-14, Unknown Gaielalles, Bacillus, Conexibacter, and in lower abundance Gaiella, Un. Xanthobacteraceae, MB-A2-108, Solirubrobacter, Un. Micromonosporaceae, Un. Bacillales, Un. Solirubrobacterales, and TK-10 (Figure 4).
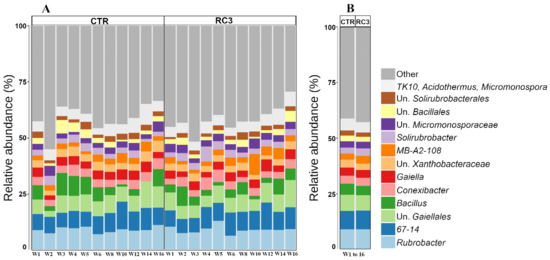
Figure 4.
Microbial community composition at the genus level. Plot (A) displays the stacked average of 3 plots per week from week 0 to week 16. Panel (B) shows the stacked bar chart showing the mean abundance of top genera in control (CTR) and RC3-treated soil.
The richness and alpha diversity indices measured with ACE, Chao1, Observed, Fisher, and Inverse Simpson (Supplementary Material Figure S4) show that the application of RC3 did not alter microbial richness or alpha diversity in the soil.
LEfSe (linear discriminant analysis effect size) plot, showing only genera with p < 0.05 and an absolute LDA score higher than 4, presented in Supplementary Material Figure S4 shows an increase of Actinobacteria and Solirubrobacter in RC3-treated soil, while genera Chloroflexi, MB-A2-108, Bacillus, Gaiellales, and Solirubrobacteracea have higher representation in the CTR plots.
3.3. Correlation Analysis
Correlation analysis of taxa with time was performed at genus-level to examine temporal fluctuations in the microbiota community (Table 1). The genera significantly negatively correlated with time in CTR plots were Marmoricola, Actinoplanes, BIrii41, Ellin6067, Reyranella, mle1-7, Actinomadura, Asanoa, Blastococcus, Cryptosporangium, Dactylosporangium, Dongia, MND1, Pyxidicoccus, Saccharopolyspora, Skermanella, Subgroup 10, and Vicinamibacteraceae. The genera significantly negatively correlated with time in RC3-treated soils were Blastococcus, Conexibacter, Craurococcus-Caldovatus, Planosporangium, and RB41.

Table 1.
Genus level temporal correlation in RC3 and control plots. Genera correlating with RC3 are in purple, and genera correlating with CTR are in dark yellow.
3.4. Beta Diversity
The analysis of weighted and unweighted Unifrac distances indicated no major effect of RC3 on the beta diversity. However, time significantly affected beta diversity (Table 2).

Table 2.
PREMNAOVA on Weighted and Unweighted distances.
The table shows a higher significance of the Time variable using unweighted than weighted Unifrac, indicating that membership is altered more than abundance.
The ecological relationship between the samples is presented as a cladogram based on weighted Unifrac distance in Figure 5. The figure shows a conspicuous level of samples clustering by treatment.
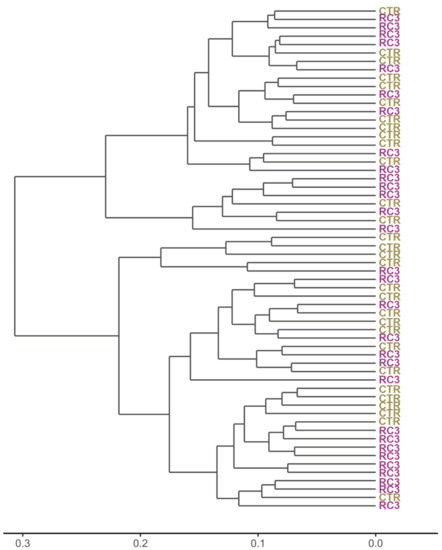
Figure 5.
Weighted Unifrac cladogram. Soil sample IDs were replaced with the treatment.
Microeco R package was used to explore the interactions of soil chemistry parameters with the microbial community. Environmental and biota interactions were visualised using a redundancy analysis (RDA) plot at the genus level (Figure 6). The mineral–genus correlation heatmap based on unweighted Unifrac distance, provided in Figure 7, shows the different responses of genera to mineral concentrations in soil in the presence and the absence of RC3. For example, Vicinamibacteraceae increased significantly in the presence of Cr in CTR plots but decreased in RC3 plots. On the contrary, in RC3 plots, the presence of K and Al significantly positively correlated with this genus (Vicinamibacteraceae), but it was reduced in CTR plots. Another example is that the Nocardioides genus increased significantly in the presence of N, C, Cr, Ni, and S, only in RC3 plots.
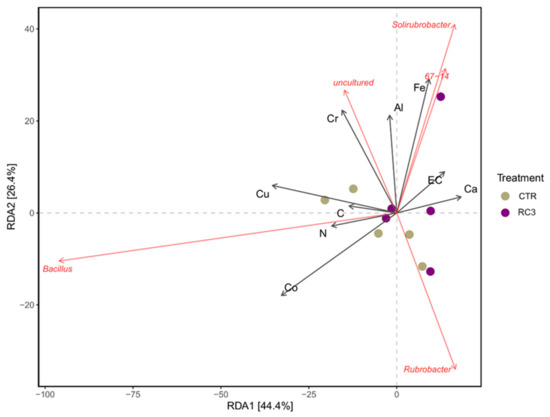
Figure 6.
RDA plot at the genus level.
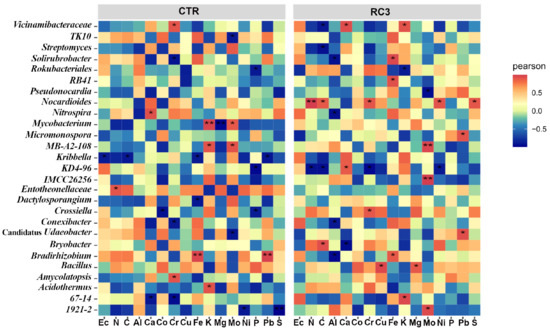
Figure 7.
Heatmap of significant mineral correlations with soil genera. Only significant correlations are shown on the map with an asterisk (* = p < 0.05; ** = p < 0.01).
4. Discussion
With millions of hectares of pasture affected and severe economic loss, pasture dieback has been a serious problem for Queensland farmers. Despite continuous efforts from scientists and significant investment from the government and non-government sectors, a clear cause and remedy for this problem have not been identified. The results of this study have shown that RC3 could be a candidate for short-term improvement in the productivity of dieback-affected pastures. Although there was an improvement in total dry matter production at week 20 after the application of RC3, the improvement did not last until week 48.
A single application of RC3 produced a significant net increment of 900 kg/ha in the dry matter at week 20 compared to the dry matter in control plots. All other changes were insignificant but showed a trend toward RC3-induced improvements in dry matter and plant morphometrics, which should be further investigated. The mineral concentration in soil did not change significantly, but there was a slight reduction in Pb, Ni, and Cr concentrations at week 12. This was expected as the active compound of RC3 is known for capturing and binding heavy metals from wastewater and contaminated soils, as mentioned previously.
The community structure shows genera commonly found in soil, some benign, e.g., plant-promoting Bacillus [41,42,43] and Conexibacter, known as nitrate-reducing [44]. However, there were genera characteristics of stressed soils, e.g., Rubrobacter genus, known as radiation and high temperature tolerant [45], and Solirubrobacter, which is usually found in poor and disturbed soils [46]. Gaiellales are found in extreme environments [47], and Blastococcus are typical for arid and desert soils [48,49].
Based on the data presented in Table 1, temporal correlation shows three genera positively correlated with time only in RC3 plots, indicating that their promotion is RC3 dependent. All of them are associated with extreme environments, e.g., RB41, typically found in stressed and low nutrient soils [50,51], Planosporangium, usually found in hot springs [52], and Conexibacter involved in nitrate reduction [44]. Craurococcus caldovatus, negatively correlated with time only in RC3 plots, is associated with amoeba presence [53]. This indicates that RC3 may have antiparasitic effects, which could affect microbial community structure and plant health. Amoeba and bacteria have a strong antagonistic relationship, with amoeba being predator of bacteria. It was estimated that 1 g of pasture soil contains 109 bacteria and 105 protozoa [54]. Rosenberg and coauthors [55] reported that amoeba in the soil prey on specific taxa, which results in a prompt change in the bacterial community composition [55]. Amoeba decreases the abundance of Betaproteobacteria and Firmicutes and increases Actinobacteria, Nitrospira, Verrucomicrobia, and Planctomycetes, and these alterations have pronounced effects on the structure of the rhizosphere bacterial community.
Furthermore, genera significantly reduced in CTR plots but not temporarily altered in RC3, Marmoricola, Actinomadura, and Dactilosporangium, are known to produce antibiotics and antimicrobial metabolites [56,57,58]. This also points to a possible mechanism of RC3 induced alterations in the microbiota that indirectly resulted in plant morphometric improvements. Ellin 6067, Reyranella, Mle1-7, and Brii41, also temporarily reduced in CTR only, are involved in the nitrogen cycle, converting ammonia to nitrite, producing nitrate, fixing N, breaking down proteins, oxidating ammonia, and denitrifying, thus making N available for plant absorption [59,60,61,62,63,64]. Actinoplanes is known for phosphate solubilisation [65], Asanoa genus hydrolyse cellulose [66], and all genera are valuable for maintaining soil health. This suggests that RC3 could have prevented the temporal decline of these beneficial genera.
The bacterial community interacted with minerals differently in RC3 and CTR (Figure 7). Unknown Vicinamibacteraceae, a degrader of complex organic matter and chitin [67,68], correlated with Cr in control and with Al and K in RC3. Nocardioides, a genus that has the ability to reduce carbon, was positively correlated with carbon in RC3 only but not in the CTR [69]. KD4-96, a genus associated with heavy metal pollution [59,70], showed a negative temporal correlation with heavy metals Cr and Mo in RC3 only, supporting the known effect of RC3 in heavy metal remediation.
5. Conclusions
Pasture dieback is affecting pasture production and generating losses in the cattle industry. In this trial, we presented the use of a known product to remediate the effects of pasture dieback in the soil microbial community and plants. Our microbiota data demonstrate the beneficial effects of RC3 in poor, damaged, and stressed soils via a significant reduction of genera used as biomarkers of poor soil. More research is needed to find a more permanent solution, with a particular focus on the regeneration of pioneer legume species. Our research fits with the global effort to formulate new strategies in agriculture to prepare for the potentially devastating effects of climate change and to maintain agriculture’s sustainability under altered environmental conditions. Future work could include investigating synergies between known plant growth promotants, developing novel plant and microbiota beneficial treatments, and focusing on identifying mechanisms, functions, and pathways in the soil microbial community and in plants that play a role in both damage and remediation of the syndromes emerging from the altered climate conditions.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/agronomy12123153/s1, SupplementaryMaterial.pdf containing Figures S1 to S4. Figure S1: Plant morphometrics for grass. Figure S2: Mineral composition did not change related to treatment. Figure S3. Alpha diversity measures show microbial community variation does not involve changes in diversity. Figure S4: LEfSe plot showing differential taxa at all taxonomic levels.
Author Contributions
M.M.W.: investigation, methodology, formal analysis, project administration, writing—original draft. X.R.: investigation, methodology, writing—review & editing. S.J.Y.: investigation, methodology, writing—review & editing T.T.: methodology review & editing Y.S.B.: investigation, methodology, formal analysis, project administration, writing—review & editing. D.S.: conceptualisation, methodology, formal analysis, funding acquisition, writing—review & editing. All authors have read and agreed to the published version of the manuscript.
Funding
We want to thank David Tomlinson, who donated funding towards the scholarship for MMW. There was no other funding involved.
Data Availability Statement
Raw sequences are publically available on NCBI Sequence Read Archive (SRA) database under accession number PRJNA887675.
Acknowledgments
The data were analysed using the Marie Curie High-Performance Computing System at Central Queensland University. We acknowledge and appreciate Jason Bell’s help in all aspects of High-Performance Computing. We also wish to acknowledge continual support in our pasture dieback investigations by the Fitzroy Basin Association and numerous local farmers. We also thank David Tomlinson, Robert Alder (Geoleak Solutions), Mick and Noela Alexander for their continual advisory and in-kind help with the project.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Smith, S.B.; Gotoh, T.; Greenwood, P.L. Current situation and future prospects for global beef production: Overview of special issue. Asian-Australas. J. Anim. Sci. 2018, 31, 927. [Google Scholar] [CrossRef] [PubMed]
- Terry, S.A.; Basarab, J.A.; Guan, L.L.; McAllister, T.A. Strategies to improve the efficiency of beef cattle production. Can. J. Anim. Sci. 2021, 101, 1–19. [Google Scholar] [CrossRef]
- Mottet, A.; Teillard, F.; Boettcher, P.; De’Besi, G.; Besbes, B. Domestic herbivores and food security: Current contribution, trends and challenges for a sustainable development. Animal 2018, 12, s188–s198. [Google Scholar] [CrossRef] [PubMed]
- Meat & Livestock Australia. State of the Industry Report-2020; MLA: North Sydney, Autralia, 2020. [Google Scholar]
- Hassan, F.-u.; Arshad, M.A.; Li, M.; Rehman, M.S.-u.; Loor, J.J.; Huang, J. Potential of Mulberry Leaf Biomass and Its Flavonoids to Improve Production and Health in Ruminants: Mechanistic Insights and Prospects. Animals 2020, 10, 2076. [Google Scholar] [CrossRef]
- Llonch, P.; Somarriba, M.; Duthie, C.A.; Troy, S.; Roehe, R.; Rooke, J.; Haskell, M.J.; Turner, S.P. Temperament and dominance relate to feeding behaviour and activity in beef cattle: Implications for performance and methane emissions. Animal 2018, 12, 2639–2648. [Google Scholar] [CrossRef]
- Insua, J.R.; Agnusdei, M.G.; Utsumi, S.A.; Berone, G.D. Morphological, environmental and management factors affecting nutritive value of tall fescue (Lolium arundinaceum). Crop Pasture Sci. 2018, 69, 1165–1172. [Google Scholar] [CrossRef]
- Tharmaraj, J.; Chapman, D.F.; Nie, Z.N.; Lane, A.P. Herbage accumulation, botanical composition, and nutritive value of five pasture types for dairy production in southern Australia. Aust. J. Agric. Res. 2008, 59, 127–138. [Google Scholar] [CrossRef]
- Cobon, D.H.; Stone, G.; Carter, J.; McKeon, G.; Zhang, B.S.; Heidemann, H. Native pastures and beef cattle show a spatially variable response to a changing climate in Queensland, Australia. Eur. J. Agron. 2020, 114. [Google Scholar] [CrossRef]
- Dineen, M.; McCarthy, B.; Ross, D.; Ortega, A.; Dillon, P.; Van Amburgh, M.E. Characterization of the nutritive value of perennial ryegrass (Lolium perenne L.) dominated pastures using updated chemical methods with application for the Cornell Net Carbohydrate and Protein System. Anim. Feed Sci. Technol. 2021, 272, 114752. [Google Scholar] [CrossRef]
- Buck, S. What Is Pasture Dieback? Queenland Government: Brisbane, Australia, 2021.
- Makiela, S.; Harrower, K.M. Overview of the current status of buffel grass dieback. Australas. Plant Dis. Notes 2008, 3, 12–16. [Google Scholar] [CrossRef]
- Hauxwell, C.; McNicholl, D. Mealybugs and Pasture Dieback; Meat and Livestock Australia: North Sydney, Australia, 2018. [Google Scholar]
- Thomson, M. Hidden Gems: An Epidemiological Investigation into the Association of Ground Pearls with Pasture Dieback; University of Queensland: Brisbane, Australia, 2019. [Google Scholar]
- Queensland Department of Environment and Sciences. Climate Change in Australia. Available online: https://www.climatechangeinaustralia.gov.au/ (accessed on 15 November 2022).
- Albiac, J.; Kahil, T.; Notivol, E.; Calvo, E. Agriculture and climate change: Potential for mitigation in Spain. Sci. Total Environ. 2017, 592, 495–502. [Google Scholar] [CrossRef]
- Pathak, H. Impact, adaptation, and mitigation of climate change in Indian agriculture. Environ. Monit. Assess. 2022, 195, 52. [Google Scholar] [CrossRef]
- Zilli, M.; Scarabello, M.; Soterroni, A.C.; Valin, H.; Mosnier, A.; Leclere, D.; Havlik, P.; Kraxner, F.; Lopes, M.A.; Ramos, F.M. The impact of climate change on Brazil’s agriculture. Sci. Total Environ. 2020, 740, 139384. [Google Scholar] [CrossRef]
- Moore, A.D.; Ghahramani, A. Climate change and broadacre livestock production across southern Australia. 1. Impacts of climate change on pasture and livestock productivity, and on sustainable levels of profitability. Glob. Chang. Biol. 2013, 19, 1440–1455. [Google Scholar] [CrossRef]
- Whitton, M.M.; Ren, X.; Yu, S.J.; Irving, A.D.; Trotter, T.T.; Bajagai, Y.S.; Stanley, D. Sea minerals reduce dysbiosis, improve pasture productivity and plant morphometrics in pasture dieback affected soils. Sustainability 2022, 14, 14873. [Google Scholar] [CrossRef]
- Alder, R.; Tomlinson, D. Use of composition as growth promotant for plants. 2019361542, 16 October 2019. [Google Scholar]
- Yazici, E.; Ahlatci, F.; Yilmaz, E.; Celep, O.; Deveci, H. Precipitation of zinc from cyanide leach solutions using Trimercapto-s-triazine (TMT). Hydrometallurgy 2020, 191, 105206. [Google Scholar] [CrossRef]
- Ayoub, G.M.; Koopman, B.; Bitton, G.; Riedesel, K. Heavy metal detoxification by trimercapto-s-triazine (TMT) as evaluated by a bacterial enzyme assay. Environ. Toxicol. Chem. Int. J. 1995, 14, 193–196. [Google Scholar] [CrossRef]
- Wang, Q.; Zheng, C.; Zhang, J.; He, F.; Yao, Y.; Zhang, T.C.; He, C. Insights into the adsorption of Pb (II) over trimercapto-s-triazine trisodium salt-modified lignin in a wide pH range. Chem. Eng. J. Adv. 2020, 1, 100002. [Google Scholar] [CrossRef]
- Zhao, N.; Li, B.; Huang, H.; Lv, X.; Zhang, M.; Cao, L. Modification of kelp and sludge biochar by TMT-102 and NaOH for cadmium adsorption. J. Taiwan Inst. Chem. Eng. 2020, 116, 101–111. [Google Scholar] [CrossRef]
- Pohl, A. Removal of heavy metal ions from water and wastewaters by sulfur-containing precipitation agents. Water Air Soil Pollut. 2020, 231, 1–17. [Google Scholar] [CrossRef]
- Wang, D.-F.; Li, S.-H.; Wang, X.-Q.; Li, L.-X.; Zhang, X. Synergistic passivation of fly ash and TMT on heavy metals in sewage sludge. Sustainability 2018, 10, 4731. [Google Scholar] [CrossRef]
- Yang, Y.; Li, M.; Sun, Y.; Gao, H.; Mao, L.; Zhang, H.; Tao, H. Optimization of Solidification and Stabilization Efficiency of Heavy Metal Contaminated Sediment Based on Response Surface Methodology. Sustainability 2022, 14, 3306. [Google Scholar] [CrossRef]
- Ahlatci, F.; Yazici, E.; Yilmaz, E.; Celep, O.; Deveci, H. A novel method for the recovery of silver from thiosulphate leach solutions using trimercapto-s-triazine (TMT). Miner. Eng. 2022, 177, 107373. [Google Scholar] [CrossRef]
- Ihsan, M.Z.; Daur, I.; Alghabari, F.; Alzamanan, S.; Rizwan, S.; Ahmad, M.; Waqas, M.; Shafqat, W. Heat stress and plant development: Role of sulphur metabolites and management strategies. Acta Agric. Scand. Sect. B—Soil Plant Sci. 2019, 69, 332–342. [Google Scholar] [CrossRef]
- Rency, A.S.; Pandian, S.; Ramesh, M. Influence of adenine sulphate on multiple shoot induction in Clitoria ternatea L. and analysis of phyto-compounds in in vitro grown plants. Biocatal. Agric. Biotechnol. 2018, 16, 181–191. [Google Scholar] [CrossRef]
- Tiku, A.R. Antimicrobial compounds and their role in plant defense. In Molecular Aspects of Plant-Pathogen Interaction; Springer: Berlin/Heidelberg, Germany, 2018; pp. 283–307. [Google Scholar]
- Pope, J.; Eichenberg, R.; Birthisel, T. Use of humate dispersible granule technology as a soil amendment in turfgrass and horticultural soils. Appl. Turfgrass Sci. 2013, 10, 1. [Google Scholar] [CrossRef]
- Taha, S.S.; Osman, A.S. Influence of potassium humate on biochemical and agronomic attributes of bean plants grown on saline soil. J. Hortic. Sci. Biotechnol. 2018, 93, 545–554. [Google Scholar] [CrossRef]
- El-Naqma, K. The role of humate substances in controlling synergism and antagonism of nutrients uptake by potato plants. Environ. Biodivers. Soil Secur. 2020, 4, 149–165. [Google Scholar] [CrossRef]
- Jin, Q.; Zhang, Y.; Wang, Q.; Li, M.; Sun, H.; Liu, N.; Zhang, L.; Zhang, Y.; Liu, Z. Effects of potassium fulvic acid and potassium humate on microbial biodiversity in bulk soil and rhizosphere soil of Panax ginseng. Microbiol. Res. 2022, 254, 126914. [Google Scholar] [CrossRef]
- da Silva, M.S.R.D.; dos Santos, B.D.S.; da Silva, C.S.R.D.; da Silva, C.S.R.D.; Antunes, L.F.D.; dos Santos, R.M.; Santos, C.H.B.; Rigobelo, E.C. Humic Substances in Combination With Plant Growth-Promoting Bacteria as an Alternative for Sustainable Agriculture. Front. Microbiol. 2021, 12. [Google Scholar] [CrossRef]
- de Vries, F.T.; Griffiths, R.I.; Bailey, M.; Craig, H.; Girlanda, M.; Gweon, H.S.; Hallin, S.; Kaisermann, A.; Keith, A.M.; Kretzschmar, M.; et al. Soil bacterial networks are less stable under drought than fungal networks. Nat. Commun. 2018, 9, 3033. [Google Scholar] [CrossRef]
- Yang, Y.; Tilman, D.; Furey, G.; Lehman, C. Soil carbon sequestration accelerated by restoration of grassland biodiversity. Nat. Commun. 2019, 10. [Google Scholar] [CrossRef]
- Ferguson, R.B.; Hergert, G.W.; Shapiro, C.A.; Wortmann, C.S. Guidelines for Soil Sampling; University of Nebraska–Lincoln: Lincoln, NE, USA, 2007. [Google Scholar]
- Wagi, S.; Ahmed, A. Bacillus spp.: Potent microfactories of bacterial IAA. PeerJ 2019, 7. [Google Scholar] [CrossRef]
- Idris, E.E.; Iglesias, D.J.; Talon, M.; Borriss, R. Tryptophan-dependent production of indole-3-acetic acid (IAA) affects level of plant growth promotion by Bacillus amyloliquefaciens FZB42. Mol. Plant-Microbe Interact. 2007, 20, 619–626. [Google Scholar] [CrossRef]
- Zhu, H.Z.; Zhang, Z.F.; Zhou, N.; Jiang, C.Y.; Wang, B.J.; Cai, L.; Liu, S.J. Diversity, Distribution and Co-occurrence Patterns of Bacterial Communities in a Karst Cave System. Front. Microbiol. 2019, 10, 1726. [Google Scholar] [CrossRef] [PubMed]
- Seki, T.; Matsumoto, A.; Ōmura, S.; Takahashi, Y. Distribution and isolation of strains belonging to the order Solirubrobacterales. J. Antibiot. 2015, 68, 763–766. [Google Scholar] [CrossRef]
- Egas, C.; Barroso, C.; Froufe, H.J.C.; Pacheco, J.; Albuquerque, L.; da Costa, M.S. Complete genome sequence of the Radiation-Resistant bacterium Rubrobacter radiotolerans RSPS-4. Stand. Genomic. Sci. 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Goswami, A.; Adkins-Jablonsky, S.; Barreto Filho, M.M.; Schilling, M.; Dawson, A.; Heiser, S.; O’Connor, A.; Walker, M.; Roberts, Q.; Morris, J. Heavy metal pollution impacts soil bacterial community structure and antimicrobial resistance at the Birmingham 35th Avenue Superfund Site. bioRxiv 2022. [Google Scholar] [CrossRef]
- Peng, M.; Jia, H.; Wang, Q. The effect of land use on bacterial communities in saline–alkali soil. Curr. Microbiol. 2017, 74, 325–333. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.-W.; Asem, M.D.; Li, X.; Li, L.-Y.; Salam, N.; Alkhalifah, D.H.M.; Hozzein, W.N.; Nie, G.-X.; Li, W.-J. Blastococcus deserti sp. nov., isolated from a desert sample. Arch. Microbiol. 2019, 201, 193–198. [Google Scholar] [CrossRef]
- Castro, J.F.; Nouioui, I.; Sangal, V.; Choi, S.; Yang, S.-J.; Kim, B.-Y.; Trujillo, M.E.; Riesco, R.; del Carmen Montero-Calasanz, M.; Rahmani, T.P. Blastococcus atacamensis sp. nov., a novel strain adapted to life in the Yungay core region of the Atacama Desert. Int. J. Syst. Evol. Microbiol. 2018, 68, 2712–2721. [Google Scholar] [CrossRef] [PubMed]
- Foesel, B.U.; Rohde, M.; Overmann, J. Blastocatella fastidiosa gen. nov., sp. nov., isolated from semiarid savanna soil—The first described species of Acidobacteria subdivision 4. Syst. Appl. Microbiol. 2013, 36, 82–89. [Google Scholar] [CrossRef] [PubMed]
- Stone, B.W.; Li, J.H.; Koch, B.J.; Blazewicz, S.J.; Dijkstra, P.; Hayer, M.; Hofmockel, K.S.; Liu, X.J.A.; Mau, R.L.; Morrissey, E.M.; et al. Nutrients cause consolidation of soil carbon flux to small proportion of bacterial community. Nat. Commun. 2021, 12. [Google Scholar] [CrossRef]
- Thawai, C.; Thamsathit, W.; Kudo, T. Planosporangium thailandense sp. nov., isolated from soil from a Thai hot spring. Int. J. Syst. Evol. Microbiol. 2013, 63, 1051–1055. [Google Scholar] [CrossRef] [PubMed]
- Scaturro, M.; Chierico, F.D.; Motro, Y.; Chaldoupi, A.; Flountzi, A.; Moran-Gilad, J.; Girolamo, A.; Koutsiomani, T.; Krogulska, B.; Lindsay, D.; et al. Bacterial communities of premise plumbing systems in four European cities, and their association with culturable Legionella. bioRxiv 2008. Available online: https://www.biorxiv.org/content/10.1101/2022.08.12.503735v1 (accessed on 31 October 2022). [CrossRef]
- Finlay, B.J.; Black, H.I.; Brown, S.; Clarke, K.J.; Esteban, G.F.; Hindle, R.M.; Olmo, J.L.; Rollett, A.; Vickerman, K. Estimating the growth potential of the soil protozoan community. Protist 2000, 151, 69–80. [Google Scholar] [CrossRef]
- Rosenberg, K.; Bertaux, J.; Krome, K.; Hartmann, A.; Scheu, S.; Bonkowski, M. Soil amoebae rapidly change bacterial community composition in the rhizosphere of Arabidopsis thaliana. ISME J. 2009, 3, 675–684. [Google Scholar] [CrossRef] [PubMed]
- Pan, X.; Zhang, S.; Zhong, Q.; Gong, G.; Wang, G.; Guo, X.; Xu, X. Effects of soil chemical properties and fractions of Pb, Cd, and Zn on bacterial and fungal communities. Sci. Total Environ. 2020, 715, 136904. [Google Scholar] [CrossRef]
- Bill, M.; Chidamba, L.; Gokul, J.K.; Labuschagne, N.; Korsten, L. Bacterial community dynamics and functional profiling of soils from conventional and organic cropping systems. Appl. Soil. Ecol. 2021, 157. [Google Scholar] [CrossRef]
- Niemhom, N.; Thawai, C. Screening and Identification of Rare Actinomycetes Isolated from Soil in Pho Hin Dad Waterfall (Namtok Sam Lan National Park, Saraburi Province) for Antimicrobial Activities. In Proceedings of the RSU International Research Conference 2020, Pathum Thani, Thailand, 1 May 2020; pp. 654–664. [Google Scholar]
- Wang, Y.; Du, L.; Liu, H.; Long, D.; Huang, M.; Wang, Y.; Huang, S.; Jin, D. Halosulfuron methyl did not have a significant effect on diversity and community of sugarcane rhizosphere microflora. J. Hazard. Mater. 2020, 399, 123040. [Google Scholar] [CrossRef]
- Li, Q.; Wan, F.; Zhao, M. Distinct soil microbial communities under Ageratina adenophora invasions. Plant Biol. 2022, 24, 430–439. [Google Scholar] [CrossRef]
- Inaba, T.; Hori, T.; Navarro, R.R.; Ogata, A.; Hanajima, D.; Habe, H. Revealing sludge and biofilm microbiomes in membrane bioreactor treating piggery wastewater by non-destructive microscopy and 16S rRNA gene sequencing. Chem. Eng. J. 2018, 331, 75–83. [Google Scholar] [CrossRef]
- Canisares, L.P.; Poffenbarger, H.; Brodie, E.L.; Sorensen, P.O.; Karaoz, U.; Villegas, D.M.; Arango, J.; Momesso, L.; Crusciol, C.A.C.; Cantarella, H. Legacy effects of intercropping and nitrogen fertilization on soil N cycling, nitrous oxide emissions, and the soil microbial community in tropical maize production. Front. Soil Sci. 2021, 17. [Google Scholar] [CrossRef]
- Gschwend, F.; Hartmann, M.; Mayerhofer, J.; Hug, A.S.; Enkerli, J.; Gubler, A.; Meuli, R.G.; Frey, B.; Widmer, F. Site and land-use associations of soil bacteria and fungi define core and indicative taxa. FEMS Microbiol. Ecol. 2021, 97, fiab165. [Google Scholar] [CrossRef] [PubMed]
- Yajun, W.; Chongchong, G.; Tianjing, C.; Jinshou, L.; Yan, X.; Dafang, F. Adaptability of enhanced bioretention cell for nitrogen and phosphorus removal under two antibiotics stress. Ecotoxicol. Environ. Saf. 2022, 230, 113114. [Google Scholar] [CrossRef]
- Bhatti, A.A.; Haq, S.; Bhat, R.A. Actinomycetes benefaction role in soil and plant health. Microb. Pathog. 2017, 111, 458–467. [Google Scholar] [CrossRef]
- Putri, A.L.; Setiawan, R. Isolation and screening of actinomycetes producing cellulase and xylanase from Mamasa soil, West Sulawesi. In Proceedings of the IOP Conference Series: Earth and Environmental Science; IOP Publishing Ltd.: Bristol, UK, 2019; p. 012035. [Google Scholar]
- Zhang, Y.; Wu, X.-h.; LIi, X.-g.; Duan, T.-t.; Xu, J.; Dong, F.-s.; Liu, X.-g.; Zheng, Y.-q. The Response of Soil Microbial Community to Repeated Application Clomazone. Biotechnol. Bull. 2020, 36, 64. [Google Scholar]
- Zhang, K.; Li, K.; Tong, M.; Xia, Y.; Cui, Y.; Liu, Z.; Chen, Q.; Li, Q.; Hu, F.; Yang, F. Distribution Pattern and Influencing Factors of Heavy Metal Resistance Genes in the Yellow River Sediments of Henan Section. Int. J. Environ. Res. Public Health 2022, 19, 10724. [Google Scholar] [CrossRef] [PubMed]
- Meena, K.K.; Bitla, U.M.; Sorty, A.M.; Singh, D.P.; Gupta, V.K.; Wakchaure, G.; Kumar, S. Mitigation of salinity stress in wheat seedlings due to the application of phytohormone-rich culture filtrate extract of methylotrophic actinobacterium Nocardioides sp. NIMMe6. Front. Microbiol. 2020, 11, 2091. [Google Scholar] [CrossRef] [PubMed]
- Girardot, F.; Allégra, S.; Pfendler, S.; Conord, C.; Rey, C.; Gillet, B.; Hughes, S.; Bouchardon, A.E.; Hua, A.; Paran, F. Bacterial diversity on an abandoned, industrial wasteland contaminated by polychlorinated biphenyls, dioxins, furans and trace metals. Sci. Total Environ. 2020, 748, 141242. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).