Genome-Wide Association Study for Powdery Mildew and Rusts Adult Plant Resistance in European Spring Barley from Polish Gene Bank
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Material
2.2. Field Experiments and Phenotypic Evaluation
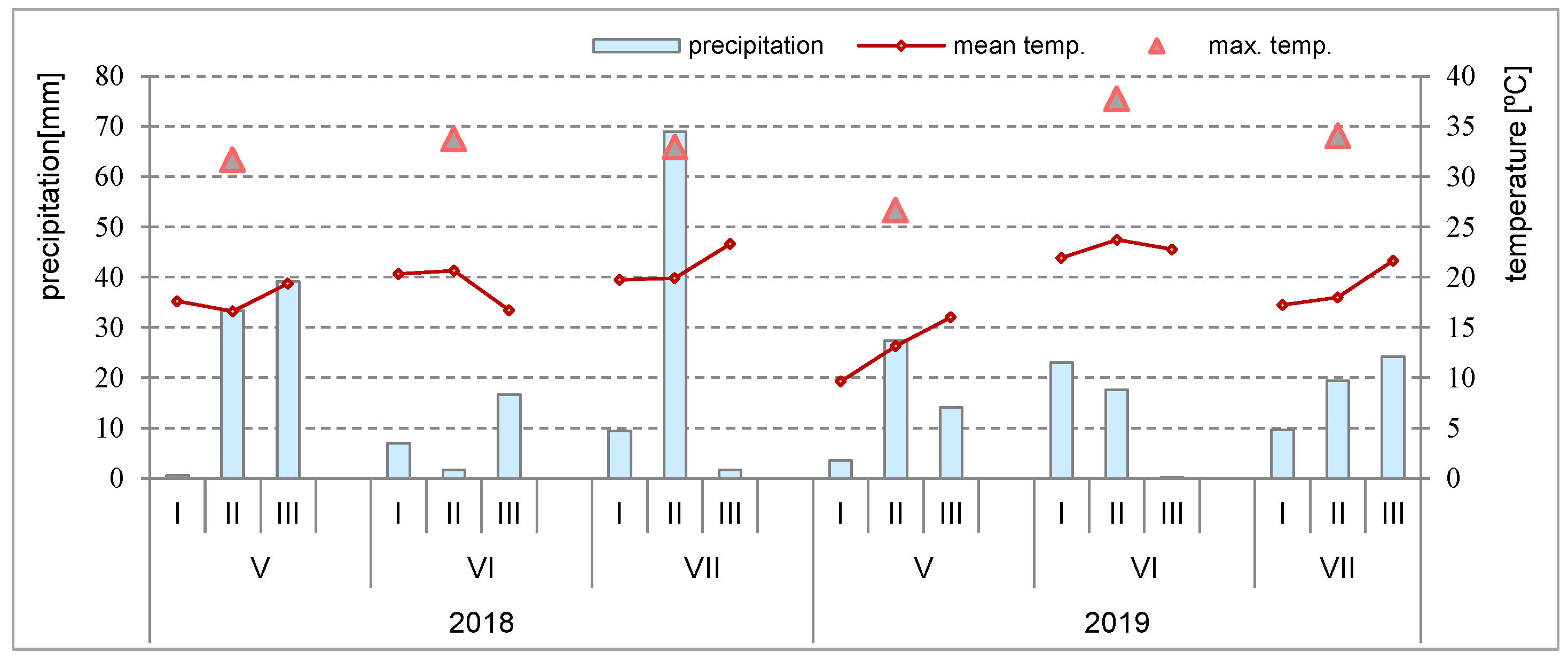
2.3. Weather Conditions
2.4. Statistical Analysis
2.5. Genotyping
2.6. Data Filtering Process
2.7. Genome-Wide Association Study (GWAS)
3. Results
3.1. Phenotypic Assessment of PM, SR, and BBR
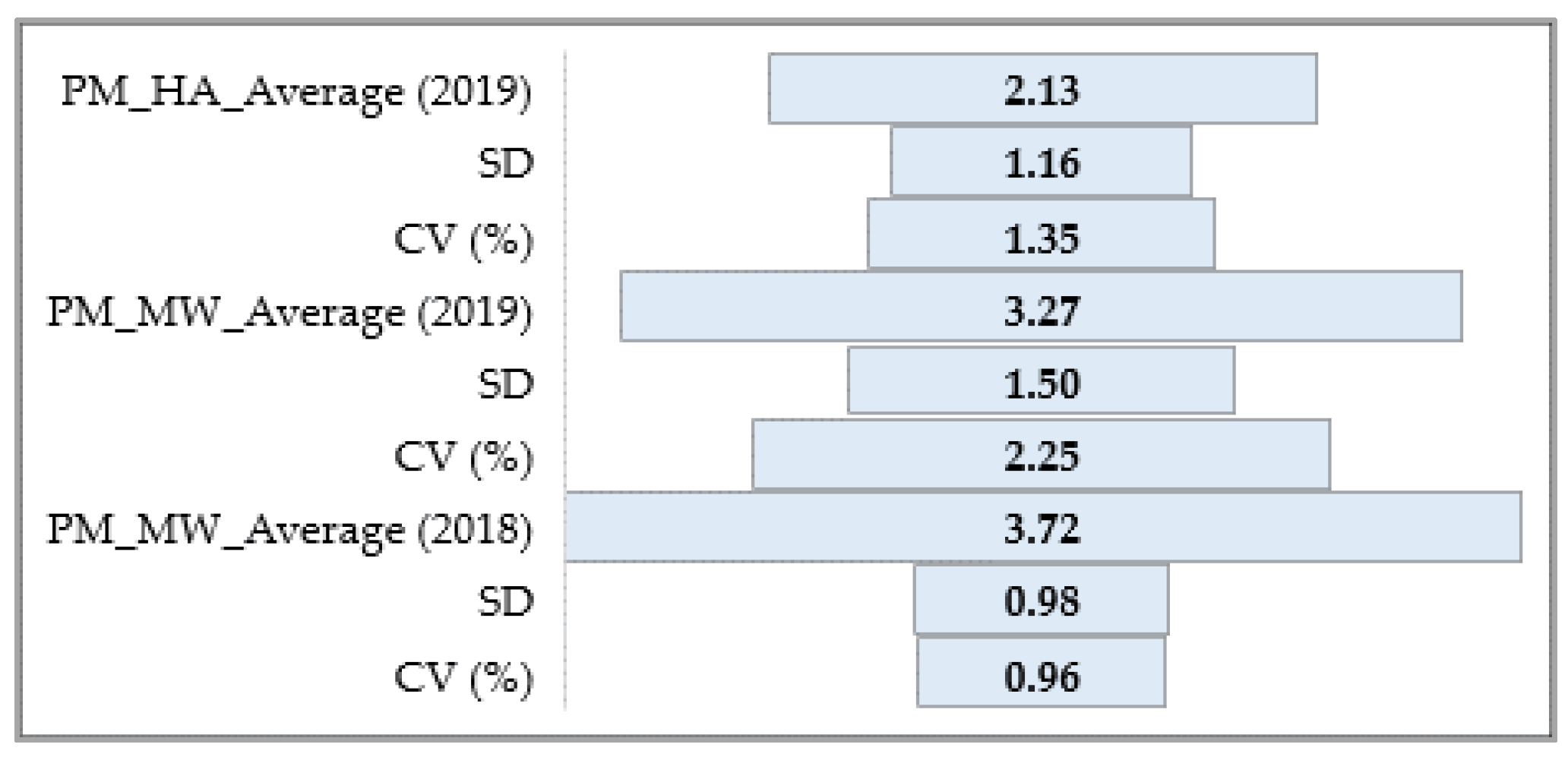
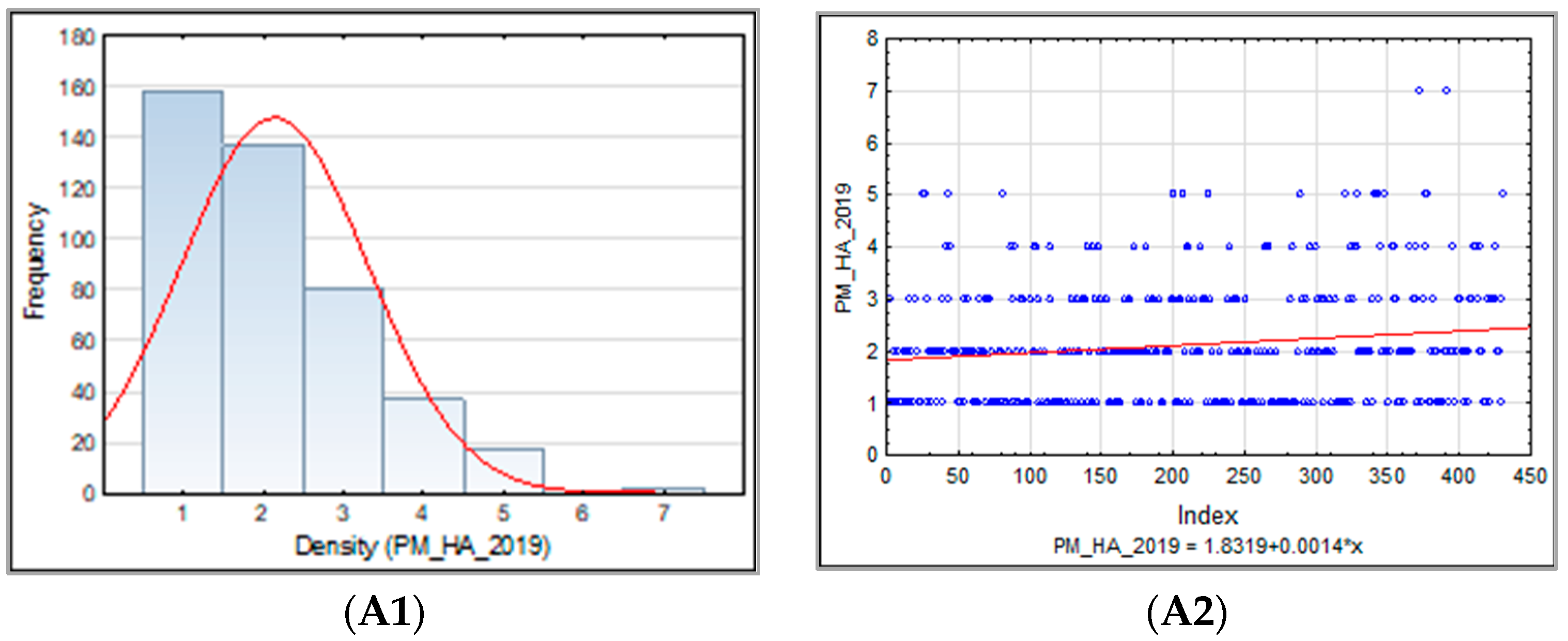
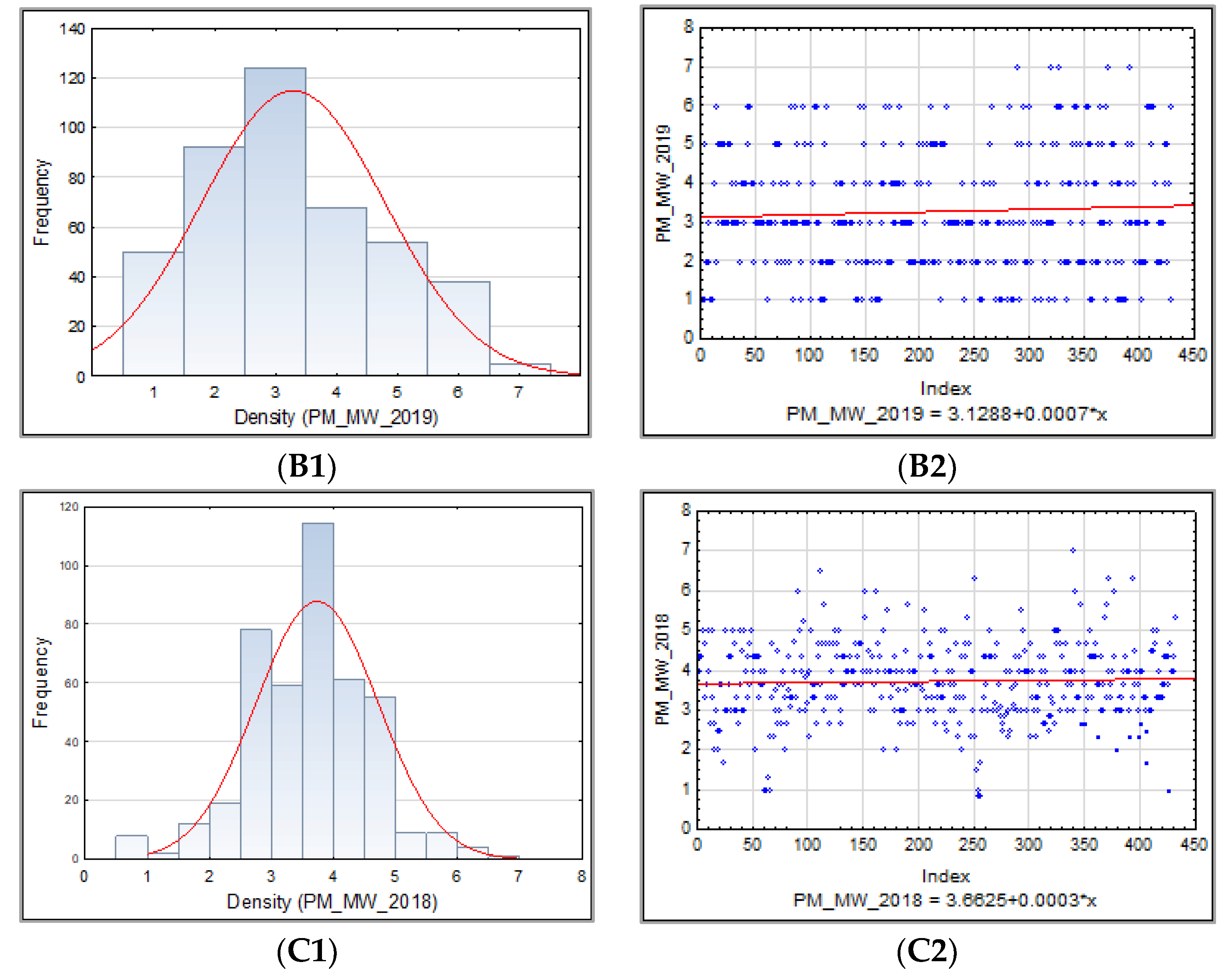
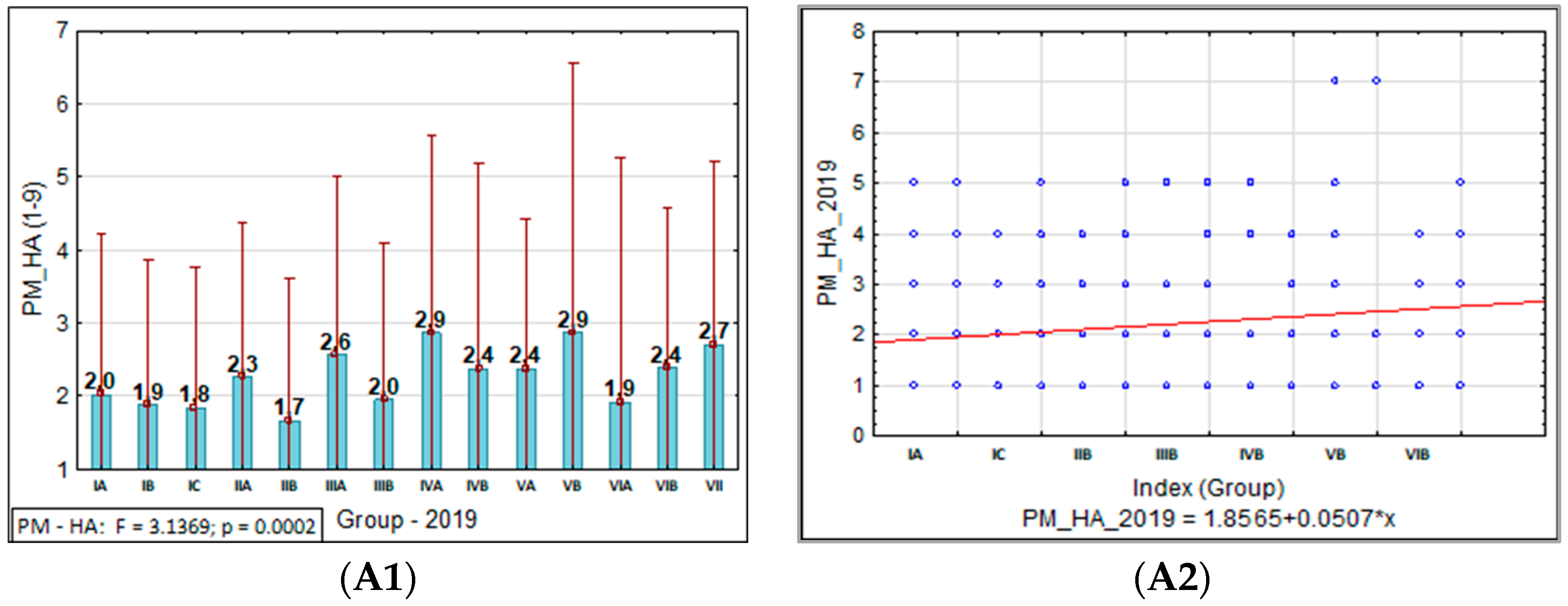
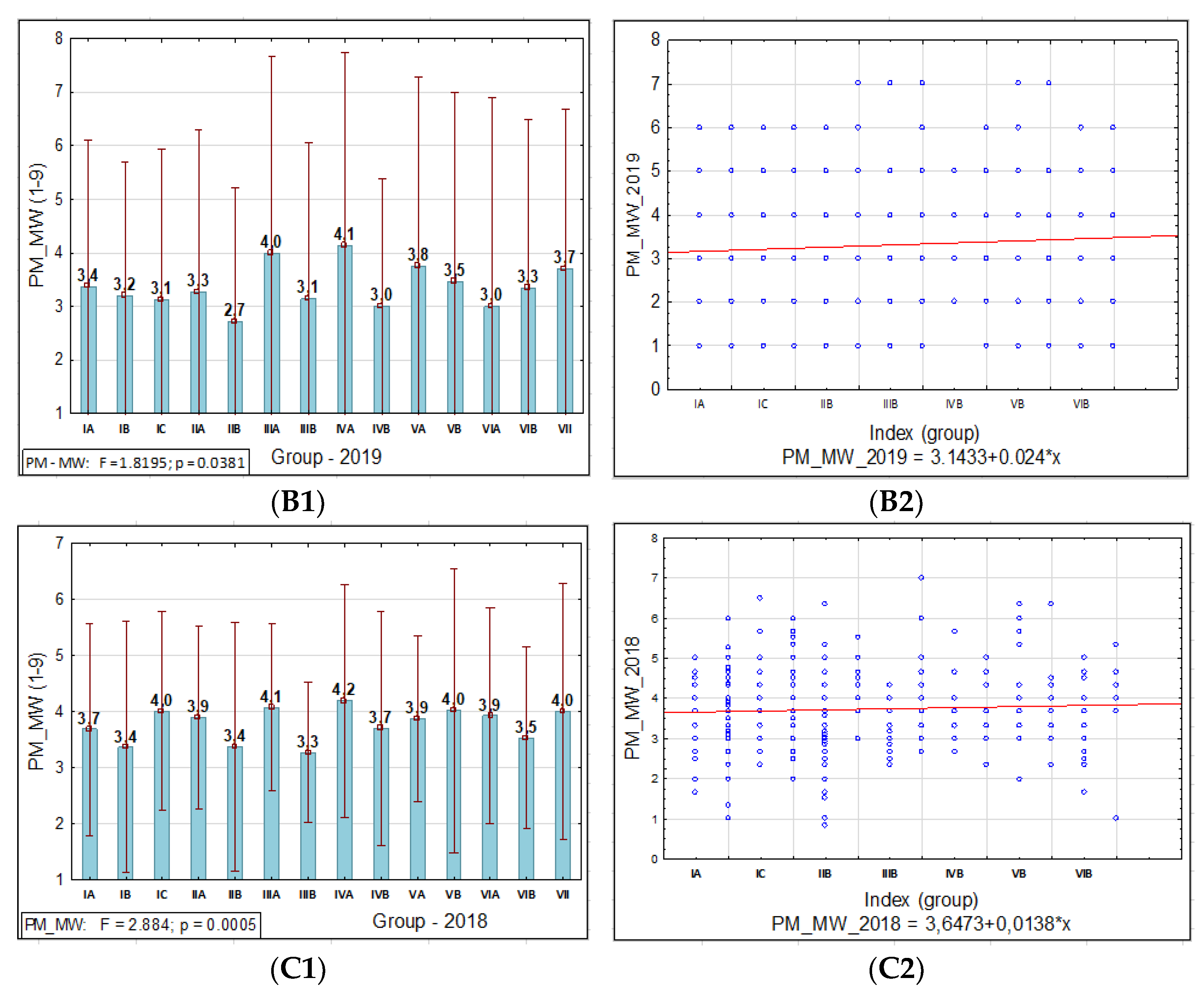
3.1.1. Powdery Mildew
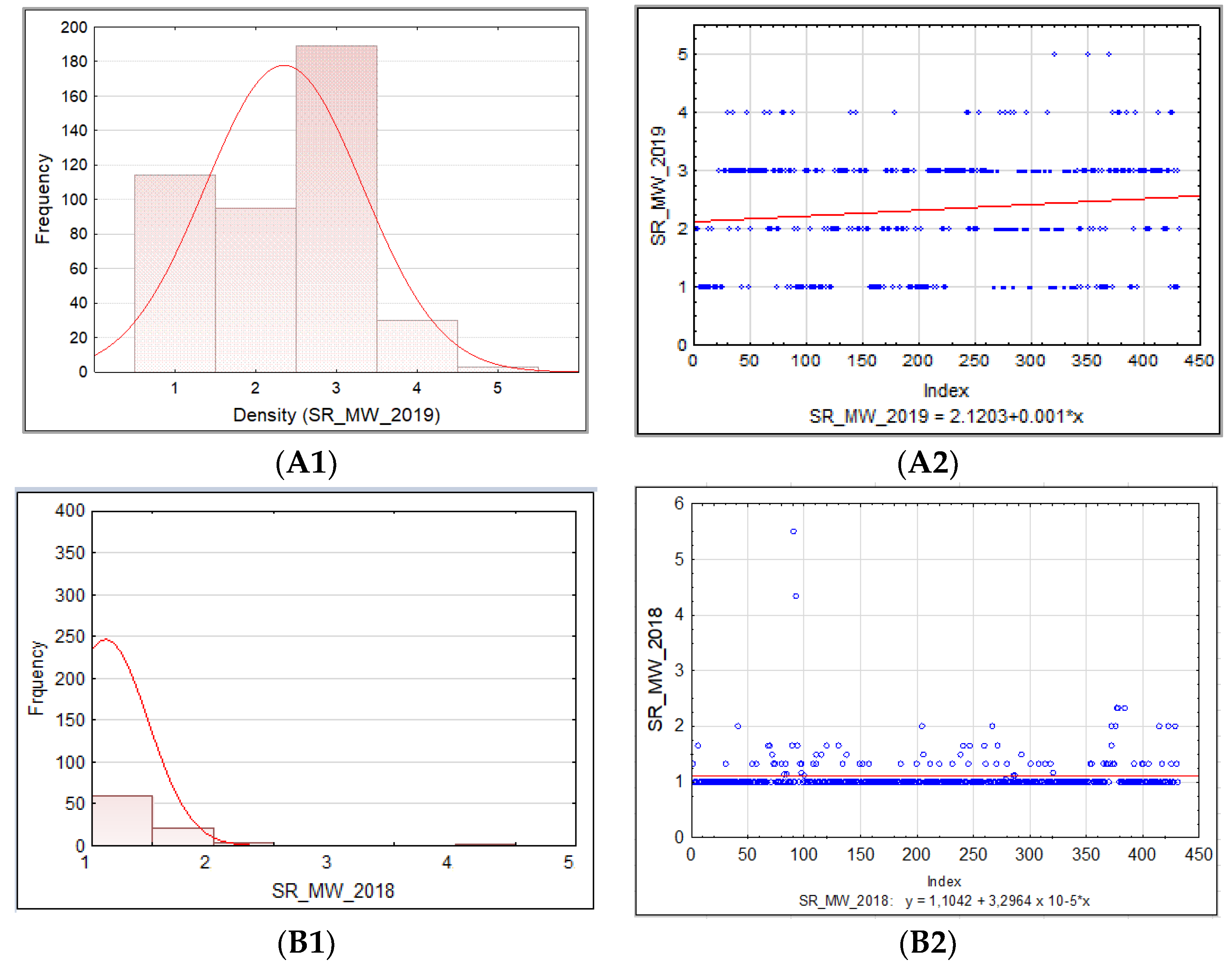
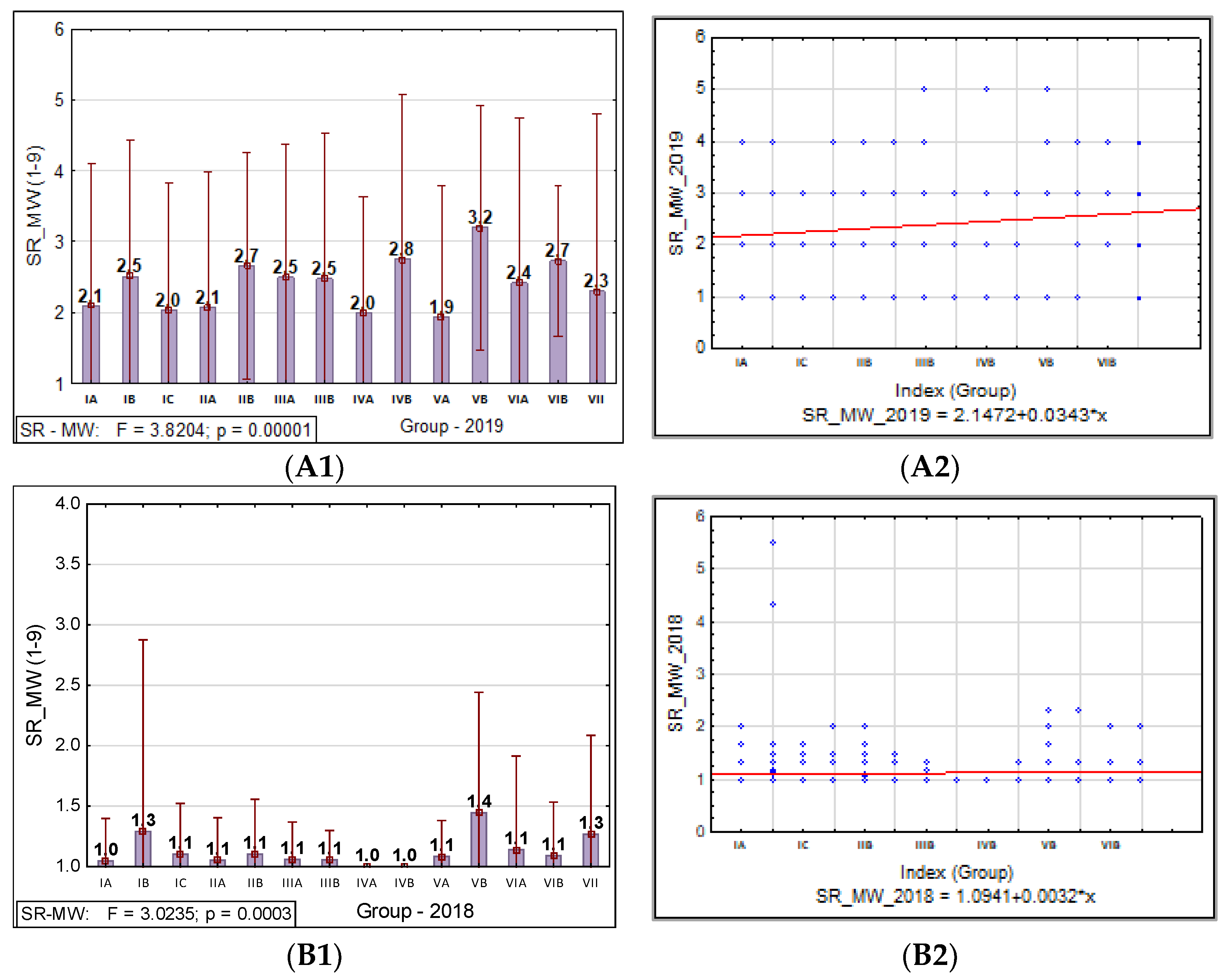
3.1.2. Barley Stem Rust
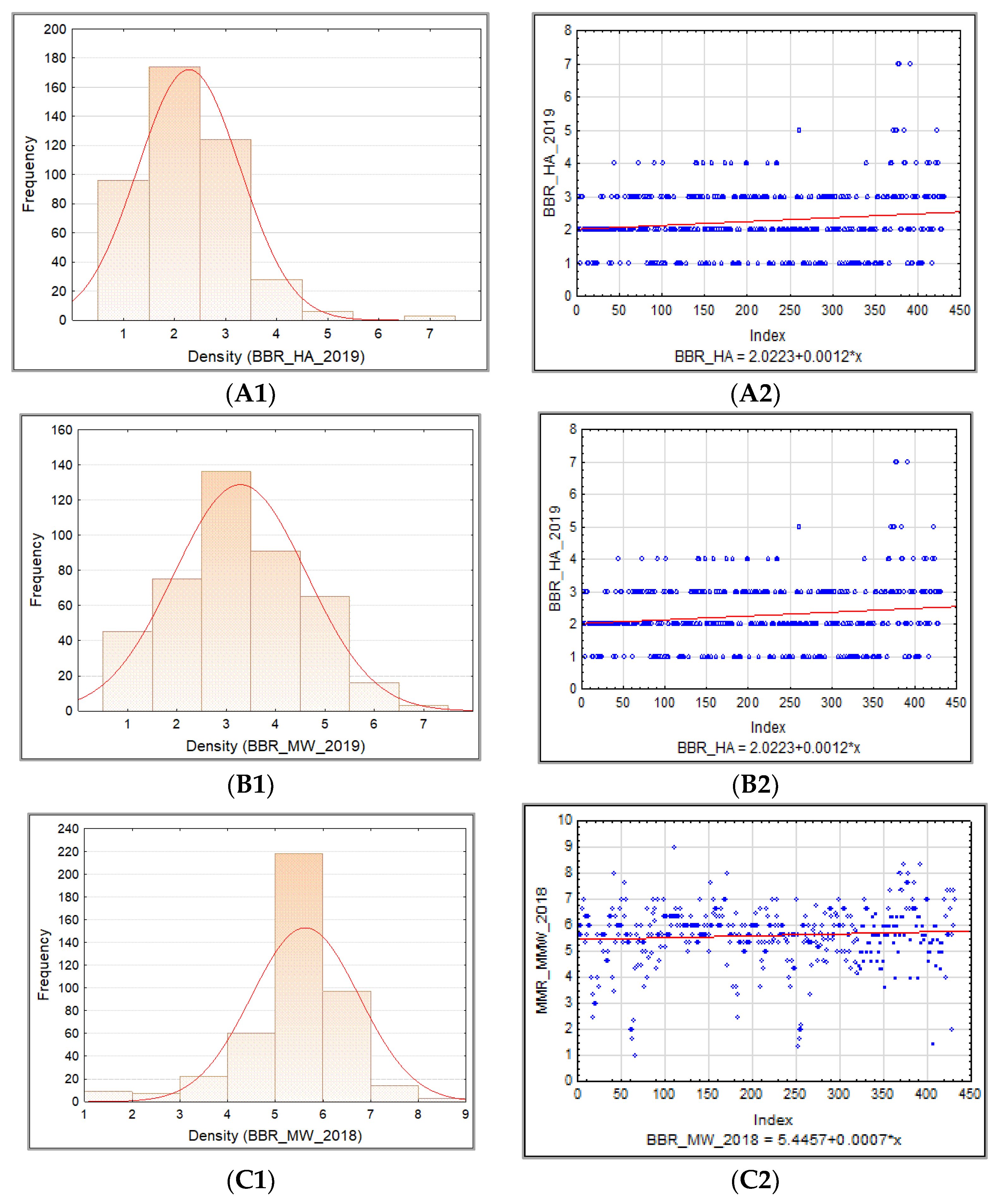
3.1.3. Barley Brown Rust
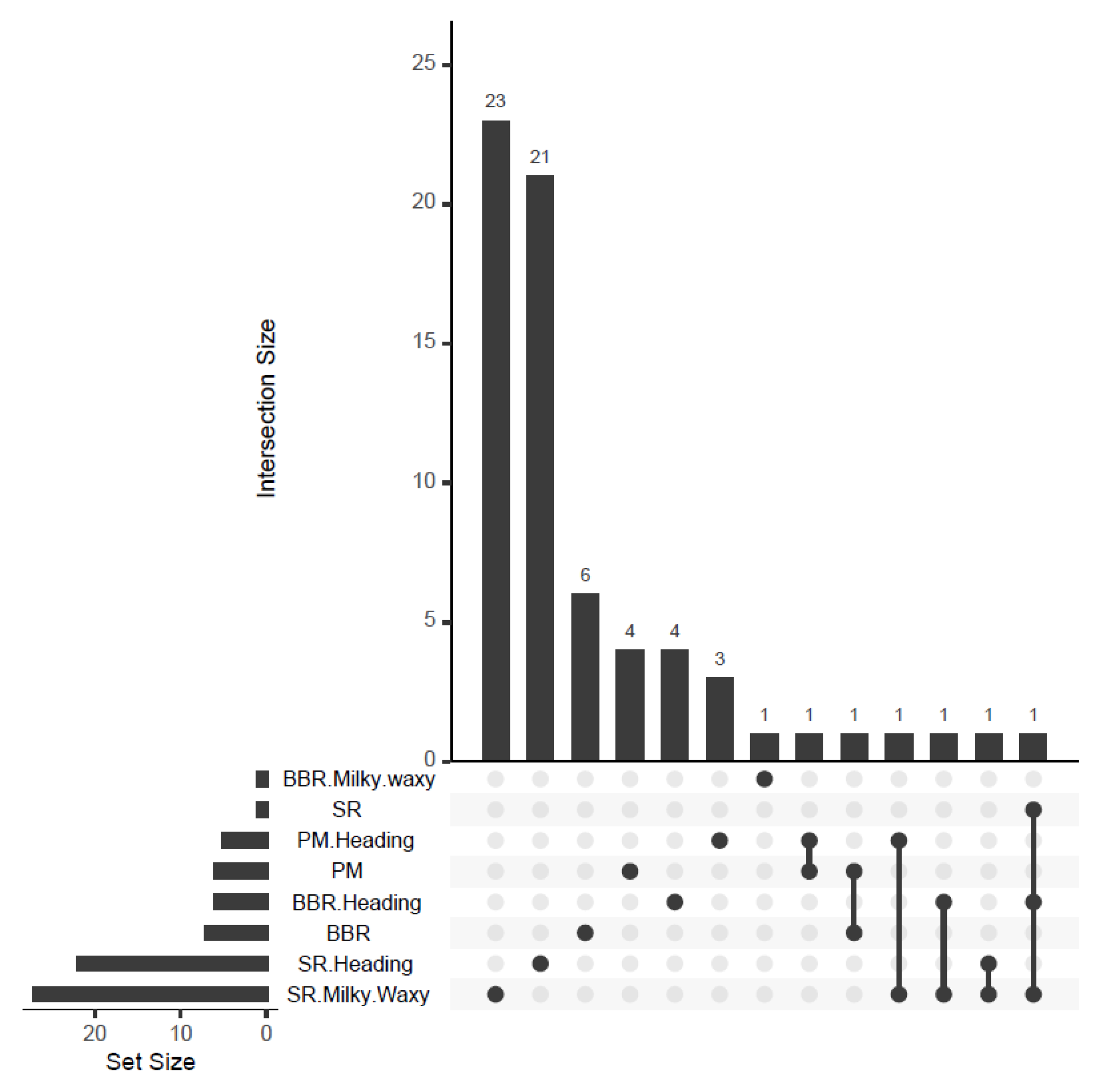
3.1.4. Relationship between PM, BBR, and SR
3.2. GWAS Analysis for Marker Trait Associations
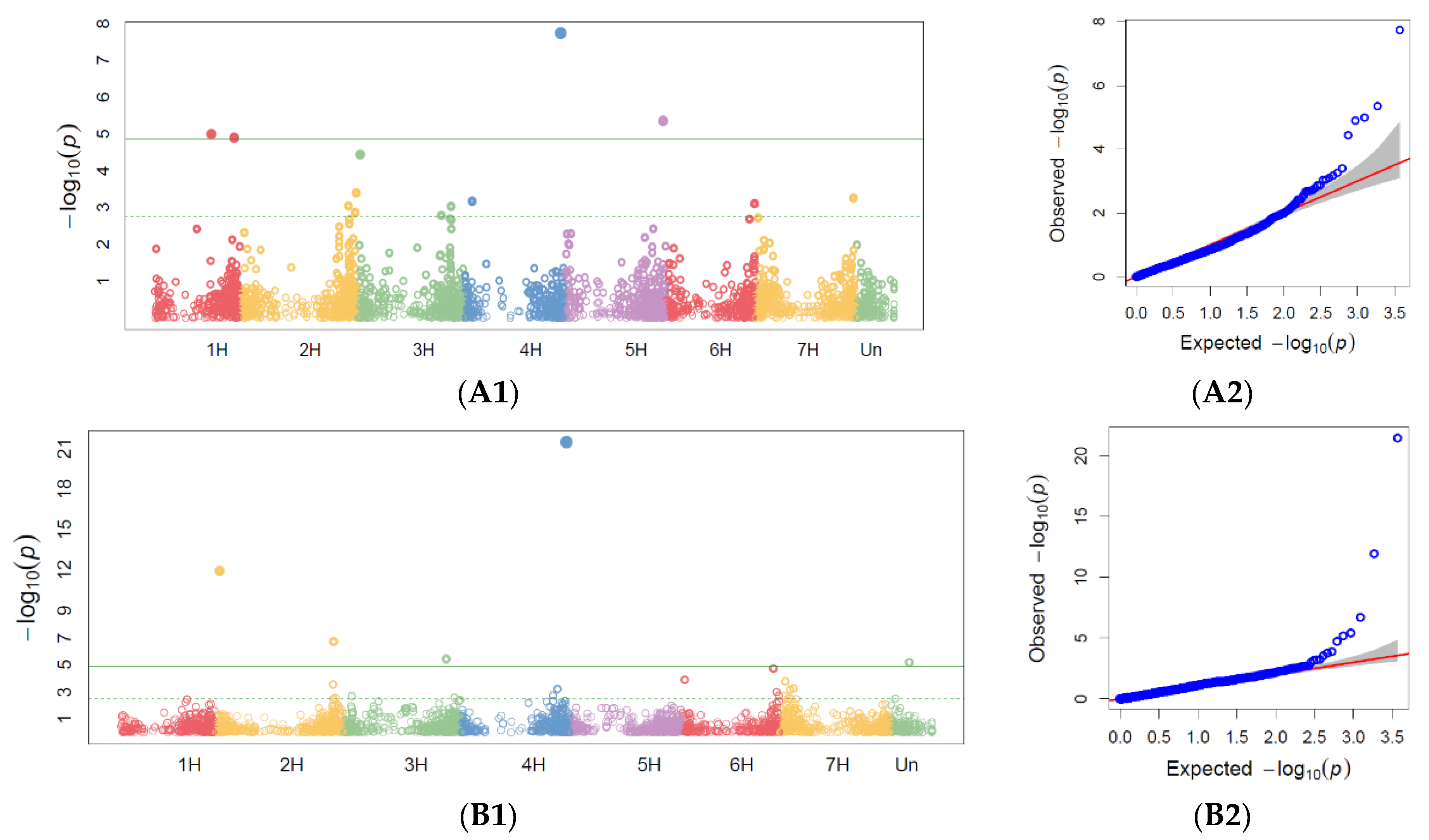
3.2.1. Powdery Mildew GWAS
3.2.2. Barley Brown Rust GWAS
3.2.3. Stem Rust GWAS
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Karagöz, A.; Özbek, K.; Akar, T.; Ergün, N.; Aydoğan, S.; Sayim, İ. Agro-morphological variation among an ancient world barley collection. Tarim Bilim. Derg. 2017, 23, 444–452. [Google Scholar] [CrossRef]
- Kumar, A.; Verma, R.P.S.; Singh, A.; Kumar Sharma, H.; Devi, G. Barley landraces: Ecological heritage for edaphic stress adaptations and sustainable production. Environ. Sustain. Indic. 2020, 6, 100035. [Google Scholar] [CrossRef]
- Dawson, I.K.; Russell, J.; Powell, W.; Steffenson, B.; Thomas, W.T.B.; Waugh, R. Barley: A translational model for adaptation to climate change. New Phytol. 2015, 206, 913–931. [Google Scholar] [CrossRef]
- Food, W. Statistics Statistical Yearbook; Food & Agriculture Organization: Rome, Italy, 2020; p. 366. ISBN 9789251333945. [Google Scholar]
- Wolfe, M.S.; Brändle, U.; Koller, B.; Limpert, E.; McDermott, J.M.; Müller, K.; Schaffner, D. Barley mildew in Europe: Population biology and host resistance. Euphytica 1992, 63, 125–139. [Google Scholar] [CrossRef]
- Agostinetto, L.; Casa, R.T.; Bogo, A.; Sachs, C.; Reis, E.M.; Kuhnem, P.R. Critical yield-point model to estimate damage caused by brown spot and powdery mildew in barley. Ciência Rural 2014, 44, 957–963. [Google Scholar] [CrossRef]
- Griffey, C.A.; Das, M.K.; Baldwin, R.E.; Waldenmaier, C.M. Yield losses in winter barley resulting from a new race of Puccinia hordei in North America. Plant Dis. 1994, 78, 256–260. [Google Scholar] [CrossRef]
- Park, R.F.; Golegaonkar, P.G.; Derevnina, L.; Sandhu, K.S.; Karaoglu, H.; Elmansour, H.M.; Dracatos, P.M.; Singh, D. Leaf Rust of Cultivated Barley: Pathology and Control. Annu. Rev. Phytopathol. 2015, 53, 565–589. [Google Scholar] [CrossRef]
- Murray, G.M.; Brennan, J.P. Estimating disease losses to the Australian barley industry. Australas. Plant Pathol. 2010, 39, 85–96. [Google Scholar] [CrossRef]
- Al-Abdallat, A.M.; Karadsheh, A.; Hadadd, N.I.; Akash, M.W.; Ceccarelli, S.; Baum, M.; Hasan, M.; Jighly, A.; Abu Elenein, J.M. Assessment of genetic diversity and yield performance in Jordanian barley (Hordeum vulgare L.) landraces grown under Rainfed conditions. BMC Plant Biol. 2017, 17, 191. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Çelik Oğuz, A.; Karakaya, A. Genetic Diversity of Barley Foliar Fungal Pathogens. Agronomy 2021, 11, 434. [Google Scholar] [CrossRef]
- Czembor, H.J.; Czembor, J.H. Leaf rust resistance in spring barley cultivars and breeding lines. Plant Breed. Seed Sci. 2007, 55, 5–20. [Google Scholar]
- Yang, L.; Zhang, X.; Zhang, X.; Wang, J.; Luo, M.; Yang, M.; Wang, H.; Xiang, L.; Zeng, F.; Yu, D.; et al. Identification and evaluation of resistance to powdery mildew and yellow rust in a wheat mapping population. PLoS ONE 2017, 12, e0177905. [Google Scholar] [CrossRef] [Green Version]
- Lamichhane, J.R.; Arseniuk, E.; Boonekamp, P.; Czembor, J.; Decroocq, V.; Enjalbert, J.; Finckh, M.R.; Korbin, M.; Koppel, M.; Kudsk, P.; et al. Advocating a need for suitable breeding approaches to boost integrated pest management: A European perspective. Pest Manag. Sci. 2018, 74, 1219–1227. [Google Scholar] [CrossRef]
- Hoseinzadeh, P.; Zhou, R.; Mascher, M.; Himmelbach, A.; Niks, R.E.; Schweizer, P.; Stein, N. High resolution genetic and physical mapping of a major powdery mildew resistance locus in barley. Front. Plant Sci. 2019, 10, 146. [Google Scholar] [CrossRef] [Green Version]
- McDonald, B.A.; Linde, C. Pathogen population genetics, evolutionary potential, and durable resistance. Annu. Rev. Phytopathol. 2002, 40, 349–379. [Google Scholar] [CrossRef] [Green Version]
- Forsberg, N.E.G.; Russell, J.; Macaulay, M.; Leino, M.W.; Hagenblad, J. Farmers without borders - genetic structuring in century old barley (Hordeum vulgare). Heredity 2015, 114, 195–206. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marcel, T.C.; Varshney, R.K.; Barbieri, M.; Jafary, H.; De Kock, M.J.D.; Graner, A.; Niks, R.E. A high-density consensus map of barley to compare the distribution of QTLs for partial resistance to Puccinia hordei and of defence gene homologues. Theor. Appl. Genet. 2007, 114, 487–500. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arabi, M.I.E.; Al-Shehadah, E.; Jawhar, M. Diallel analysis of barley resistance to Blumeria graminis. Cereal Res. Commun. 2019, 47, 455–462. [Google Scholar] [CrossRef]
- Dreiseitl, A. Specific resistance of barley to powdery mildew, its use and beyond. A concise critical review. Genes 2020, 11, 971. [Google Scholar] [CrossRef] [PubMed]
- Dreiseitl, A. Differences in powdery mildew epidemics in spring and winter barley based on 30-year variety trials. Ann. Appl. Biol. 2011, 159, 49–57. [Google Scholar] [CrossRef]
- Ziems, L.A.; Hickey, L.T.; Platz, G.J.; Franckowiak, J.D.; Dracatos, P.M.; Singh, D.; Park, R.F. Characterization of Rph24: A gene conferring adult plant resistance to Puccinia hordei in barley. Phytopathology 2017, 107, 834–841. [Google Scholar] [CrossRef] [Green Version]
- Dreiseitl, A.; Zavřelová, M. Identification of barley powdery mildew resistances in gene bank accessions and the use of gene diversity for verifying seed purity and authenticity. PLoS ONE 2018, 13, e0208719. [Google Scholar] [CrossRef] [PubMed]
- Piechota, U.; Czembor, P.C.; Słowacki, P.; Czembor, J.H. Identifying a novel powdery mildew resistance gene in a barley landrace from Morocco. J. Appl. Genet. 2019, 60, 243–254. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Piechota, U.; Słowacki, P.; Czembor, P.C. Identification of a novel recessive gene for resistance to powdery mildew (Blumeria graminis f. sp. hordei) in barley (Hordeum vulgare). Plant Breed. 2020, 139, 730–742. [Google Scholar] [CrossRef]
- Kavanagh, P.J.; Singh, D.; Bansal, U.K.; Park, R.F. Inheritance and characterization of the new and rare gene Rph25 conferring seedling resistance in Hordeum vulgare against Puccinia hordei. Plant Breed. 2017, 136, 908–912. [Google Scholar] [CrossRef]
- Fazlikhani, L.; Keilwagen, J.; Kopahnke, D.; Deising, H.; Ordon, F.; Perovic, D. High resolution mapping of RphMBR1012 conferring resistance to Puccinia hordei in barley (Hordeum vulgare L.). Front. Plant Sci. 2019, 10, 640. [Google Scholar] [CrossRef] [PubMed]
- Niks, R.E.; Walther, U.; Jaiser, H.; Martinez, F.; Rubiales, D.; Andersen, O.; Flath, K.; Gymer, P.; Heinrichs, F.; Jonsson, R.; et al. Resistance against barley leaf rust (Puccinia hordei) in West-European spring barley germplasm. Agronomie 2000, 20, 769–782. [Google Scholar] [CrossRef] [Green Version]
- Dracatos, P.M.; Haghdoust, R.; Singh, R.P.; Huerta Espino, J.; Barnes, C.W.; Forrest, K.; Hayden, M.; Niks, R.E.; Park, R.F.; Singh, D. High-density mapping of triple rust resistance in barley using DArT-seq markers. Front. Plant Sci. 2019, 10, 467. [Google Scholar] [CrossRef]
- Sharma Poudel, R.; Al-Hashel, A.F.; Gross, T.; Gross, P.; Brueggeman, R. Pyramiding rpg4- And Rpg1-mediated stem rust resistance in barley requires the Rrr1 gene for both to function. Front. Plant Sci. 2018, 871, 1789. [Google Scholar] [CrossRef] [Green Version]
- Czembor, J.H.; Czembor, E. Mlo resistance to powdery mildew (Blumeria graminis f. sp. hordei) in barley landraces collected in Yemen. Agronomy 2021, 11, 1582. [Google Scholar] [CrossRef]
- Jørgensen, I.H. Discovery, characterization and exploitation of Mlo powdery mildew resistance in barley. Euphytica 1992, 63, 141–152. [Google Scholar] [CrossRef]
- Czembor, J.H.; Czembor, E. Mlo resistance of barley to powdery mildew (Blumeria graminis f.sp. hordei). Part I. Genetics, phenotype, mechanism and molecular investigations. Biul. IHAR 2003, 230, 335–354. [Google Scholar]
- Ge, C.; Wentzel, E.; D’Souza, N.; Chen, K.; Oliver, R.P.; Ellwood, S.R. Adult resistance genes to barley powdery mildew confer basal penetration resistance associated with broad-spectrum resistance. Plant Genome 2021, 1–13. [Google Scholar] [CrossRef]
- Hickey, L.T.; Lawson, W.; Platz, G.J.; Dieters, M.; Arief, V.N.; Germán, S.; Fletcher, S.; Park, R.F.; Singh, D.; Pereyra, S.; et al. Mapping Rph20: A gene conferring adult plant resistance to Puccinia hordei in barley. Theor. Appl. Genet. 2011, 123, 55–68. [Google Scholar] [CrossRef]
- Liu, F.; Gupta, S.; Zhang, X.Q.; Jones, M.; Loughman, R.; Lance, R.; Li, C. PCR markers for selection of adult plant leaf rust resistance in barley (Hordeum vulgare L.). Mol. Breed. 2011, 28, 657–666. [Google Scholar] [CrossRef]
- Singh, D.; Ziems, L.A.; Dracatos, P.M.; Pourkheirandish, M.; Tshewang, S.; Czembor, P.; German, S.; Fowler, R.A.; Snyman, L.; Platz, G.J.; et al. Genome-wide association studies provide insights on genetic architecture of resistance to leaf rust in a worldwide barley collection. Mol. Breed. 2018, 38, 43. [Google Scholar] [CrossRef]
- Singh, D.; Dracatos, P.; Derevnina, L.; Zhou, M.; Park, R.F. Rph23: A new designated additive adult plant resistance gene to leaf rust in barley on chromosome 7H. Plant Breed. 2015, 134, 62–69. [Google Scholar] [CrossRef]
- Mehnaz, M.; Dracatos, P.; Pham, A.; March, T.; Maurer, A.; Pillen, K.; Forrest, K.; Kulkarni, T.; Pourkheirandish, M.; Park, R.F.; et al. Discovery and fine mapping of Rph28: A new gene confereing resistance to Puccinia hordei from wild barley. Theor. Plant Genet. 2021, 134, 2167–2179. [Google Scholar] [CrossRef] [PubMed]
- Weise, S.; Lohwasser, U.; Oppermann, M. Document or lose it—on the importance of information management for genetic resources conservation in genebanks. Plants 2020, 9, 1050. [Google Scholar] [CrossRef] [PubMed]
- FAO. The Second Report on the State of the World’s Animal Genetic Resources for Food and Agriculture; FAO: Rome, Italy, 2015; p. 562. ISBN 9789251065341. [Google Scholar]
- Milner, S.G.; Jost, M.; Taketa, S.; Mazón, E.R.; Himmelbach, A.; Oppermann, M.; Weise, S.; Knüpffer, H.; Basterrechea, M.; König, P.; et al. Genebank genomics highlights the diversity of a global barley collection. Nat. Genet. 2019, 51, 319–326. [Google Scholar] [CrossRef] [Green Version]
- Wang, X.; Xu, Y.; Hu, Z.; Xu, C. Genomic selection methods for crop improvement: Current status and prospects. Crop J. 2018, 6, 330–340. [Google Scholar] [CrossRef]
- Rasheed, A.; Hao, Y.; Xia, X.; Khan, A.; Xu, Y.; Varshney, R.K.; He, Z. Crop Breeding Chips and Genotyping Platforms: Progress, Challenges, and Perspectives. Mol. Plant 2017, 10, 1047–1064. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Crossa, J.; Pérez-Rodríguez, P.; Cuevas, J.; Montesinos-López, O.; Jarquín, D.; de los Campos, G.; Burgueño, J.; González-Camacho, J.M.; Pérez-Elizalde, S.; Beyene, Y.; et al. Genomic Selection in Plant Breeding: Methods, Models, and Perspectives. Trends Plant Sci. 2017, 22, 961–975. [Google Scholar] [CrossRef] [PubMed]
- König, P.; Beier, S.; Basterrechea, M.; Schüler, D.; Arend, D.; Mascher, M.; Stein, N.; Scholz, U.; Lange, M. BRIDGE—A Visual Analytics Web Tool for Barley Genebank Genomics. Front. Plant Sci. 2020, 11, 701. [Google Scholar] [CrossRef] [PubMed]
- Jing, R.; Ambrose, M.A.; Knox, M.R.; Smykal, P.; Hybl, M.; Ramos, Á.; Caminero, C.; Burstin, J.; Duc, G.; van Soest, L.J.M.; et al. Genetic diversity in European Pisum germplasm collections. Theor. Appl. Genet. 2012, 125, 367–380. [Google Scholar] [CrossRef] [Green Version]
- Newton, A.C.; Akar, T.; Baresel, J.P.; Bebeli, P.J.; Bettencourt, E.; Bladenopoulos, K.V.; Czembor, J.H.; Fasoula, D.A.; Katsiotis, A.; Koutis, K.; et al. Cereal landraces for sustainable agriculture. A review. Agron. Sustain. Dev. 2010, 30, 237–269. [Google Scholar] [CrossRef] [Green Version]
- Dreiseitl, A. Heterogeneity of powdery mildew resistance revealed in accessions of the icarda wild barley collection. Front. Plant Sci. 2017, 8, 202. [Google Scholar] [CrossRef]
- Parzies, H.K.; Spoor, W.; Ennos, R.A. Genetic diversity of barley landrace accessions (Hordeum vulgare ssp. vulgare) conserved for different lengths of time in ex situ gene banks. Heredity 2000, 84, 476–486. [Google Scholar] [CrossRef]
- Amezrou, R.; Gyawali, S.; Belqadi, L.; Chao, S.; Arbaoui, M.; Mamidi, S.; Rehman, S.; Sreedasyam, A.; Verma, R.P.S. Molecular and phenotypic diversity of ICARDA spring barley (Hordeum vulgare L.) collection. Genet. Resour. Crop Evol. 2018, 65, 255–269. [Google Scholar] [CrossRef]
- Brbaklić, L.; Trkulja, D.; Mikić, S.; Mirosavljević, M.; Momčilović, V.; Dudić, B.; Procházková, L.; Aćin, V. Genetic diversity and population structure of serbian barley (Hordeum vulgare L.) collection during a 40-year long breeding period. Agronomy 2021, 11, 118. [Google Scholar] [CrossRef]
- Marone, D.; Russo, M.A.; Mores, A.; Ficco, D.B.M.; Laidò, G.; Mastrangelo, A.M.; Borrelli, G.M. Importance of landraces in cereal breeding for stress tolerance. Plants 2021, 10, 1267. [Google Scholar] [CrossRef]
- Piechota, U.; Czembor, P.C.; Czembor, J.H. Evaluating barley landraces collected in North Africa and the Middle East for powdery mildew infection at seedling and adult plant stages. Cereal Res. Commun. 2020, 48, 179–185. [Google Scholar] [CrossRef] [Green Version]
- Jones, H.; Civá, P.; Cockram, J.; Leigh, F.J.; Smith, L.M.J.; Jones, M.K.; Charles, M.P.; Molina-Cano, J.L.; Powell, W.; Jones, G.; et al. Evolutionary history of barley cultivation in Europe revealed by genetic analysis of extant landraces. BMC Evol. Biol. 2011, 11, 320. [Google Scholar] [CrossRef] [Green Version]
- Aoki, K. A three-population wave-of-advance model for the European early Neolithic. PLoS ONE 2020, 15, e0233184. [Google Scholar] [CrossRef] [PubMed]
- Malysheva-Otto, L.; Ganal, M.W.; Law, J.R.; Reeves, J.C.; Röder, M.S. Temporal trends of genetic diversity in European barley cultivars (Hordeum vulgare L.). Mol. Breed. 2007, 20, 309–322. [Google Scholar] [CrossRef]
- 58 Zadoks, J.C.; Chang, T.T.; Konzak, C.F. A decimal code for the growth stages of cereals. Weed Res. 1974, 14, 415–421. [Google Scholar] [CrossRef]
- Elshire, R.J.; Glaubitz, J.C.; Sun, Q.; Poland, J.A.; Kawamoto, K.; Buckler, E.S.; Mitchell, S.E. A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS ONE 2011, 6, e19379. [Google Scholar] [CrossRef] [Green Version]
- Mascher, M.; Gundlach, H.; Himmelbach, A.; Beier, S.; Twardziok, S.O.; Wicker, T.; Radchuk, V.; Dockter, C.; Hedley, P.E.; Russell, J.; et al. A chromosome conformation capture ordered sequence of the barley genome. Nature 2017, 544, 427–433. [Google Scholar] [CrossRef] [Green Version]
- Czembor, E.; Czembor, J.H.; Suchecki, R.; Watson-Haigh, N.S. DArT-based evaluation of soybean germplasm from Polish Gene Bank. BMC Res. Notes 2021, 14, 343. [Google Scholar] [CrossRef]
- Gruber, B.; Unmack, P.J.; Berry, O.F.; Georges, A. dartr: An r package to facilitate analysis of SNP data generated from reduced representation genome sequencing. Mol. Ecol. Resour. 2018, 18, 691–699. [Google Scholar] [CrossRef] [PubMed]
- Paradis, E. Pegas: An R package for population genetics with an integrated-modular approach. Bioinformatics 2010, 26, 419–420. [Google Scholar] [CrossRef] [Green Version]
- Huang, M.; Liu, X.; Zhou, Y.; Summers, R.M.; Zhang, Z. BLINK: A package for the next level of genome-wide association studies with both individuals and markers in the millions. Gigascience 2018, 8, giy154. [Google Scholar] [CrossRef]
- Alqudah, A.M.; Sallam, A.; Stephen Baenziger, P.; Börner, A. GWAS: Fast-forwarding gene identification and characterization in temperate cereals: Lessons from Barley—A review. J. Adv. Res. 2020, 22, 119–135. [Google Scholar] [CrossRef]
- Alqudah, A.M.; Schnurbusch, T. Heading date is not flowering time in spring barley. Front. Plant Sci. 2017, 8, 896. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Monteagudo, A.; Casas, A.M.; Cantalapiedra, C.P.; Contreras-Moreira, B.; Gracia, M.P.; Igartua, E. Harnessing novel diversity from landraces to improve an elite barley variety. Front. Plant Sci. 2019, 10, 434. [Google Scholar] [CrossRef] [PubMed]
- Czembor, J.H.; Pietrusinska, A.; Piechota, U. Hordeum bulbosum—Jako źródło efektywnej odporności na rdzę karłową jęczmienia. Biul. IHAR 2017, 5, 57–58. [Google Scholar]
- Czembor, J.H.; Czembor, H.J.; Attene, G.; Papa, R.; Agraria, V.; Nicola, V.D.; Dago, M. Leaf rust resistance in selections from barley landraces collected in Sardinia. Plant Breed. Seed Sci. 2007, 56, 73–84. [Google Scholar]
- Dreiseitl, A. A novel way to identify specific powdery mildew resistance genes in hybrid barley cultivars. Sci. Rep. 2020, 10, 18930. [Google Scholar] [CrossRef] [PubMed]
- Tsai, H.Y.; Janss, L.L.; Andersen, J.R.; Orabi, J.; Jensen, J.D.; Jahoor, A.; Jensen, J. Genomic prediction and GWAS of yield, quality and disease-related traits in spring barley and winter wheat. Sci. Rep. 2020, 10, 3347. [Google Scholar] [CrossRef]
- Hickey, L.T.; Lawson, W.; Platz, G.J.; Fowler, R.A.; Arief, V.; Dieters, M.; German, S.; Fletcher, S.; Park, R.F.; Singh, D.; et al. Mapping Quantitative Trait. Loci for Partial Resistance to Powdery Mildew in an Australian Barley Population. Crop Sci. 2012, 52, 1021–1032. [Google Scholar] [CrossRef]
- Turuspekov, Y.; Ormanbekova, D.; Rsaliev, A.; Abugalieva, S. Genome-wide association study on stem rust resistance in Kazakh spring barley lines. BMC Plant Biol. 2016, 16, 13–21. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ledesma-Ramírez, L.; Solís-Moya, E.; Iturriaga, G.; Sehgal, D.; Reyes-Valdes, M.H.; Montero-Tavera, V.; Sansaloni, C.P.; Burgueño, J.; Ortiz, C.; Aguirre-Mancilla, C.L.; et al. GWAS to Identify Genetic Loci for Resistance to Yellow Rust in Wheat Pre-Breeding Lines Derived from Diverse Exotic Crosses. Front. Plant. Sci. 2019, 10, 1390. [Google Scholar] [CrossRef]
- Almerekova, S.; Genievskaya, Y.; Abugalieva, S.; Sato, K.; Turuspekov, Y. Population structure and genetic diversity of two-rowed barley accessions from Kazakhstan based on snp genotyping data. Plants 2021, 10, 2025. [Google Scholar] [CrossRef] [PubMed]
- Brueggeman, R.; Rostoks, N.; Kudrna, D.; Kilian, A.; Han, F.; Chen, J.; Druka, A.; Steffenson, B.; Kleinhofs, A. The barley stem rust-resistance gene Rpg1 is a novel disease-resistance gene with homology to receptor kinases. Proc. Natl. Acad. Sci. USA 2002, 99, 9328–9333. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wenz, P.; Raman, R.; Wang, J.; Zhou, M.; Huttner, E.; Kilian, A. A DArT platform for quantitative bulked segregant analysis. BMC Genom. 2007, 8, 196. [Google Scholar] [CrossRef]
- Kilian, A.; Huttneri, E.; Wenzl, P.; Jaccoud, D.; Carling, J.; Gaig, V.; Eversi, M.; Heller-Uszynska, K.; Uszynski, G.; Cayla, C.; et al. The Fast and the Cheap: SNP and DArT-Based Whole Genome Profiling for Crop Improvement. In Proceedings of the International Congress “In the Wake of the Double Helix: From the Green Revolution to the Gene Revolution”, Bologna, Italy, 27–31 May 2003; Tuberosa, R., Phillips, R.L., Gale, M., Eds.; Avenue Media: Bologna, Italy, 2005; pp. 443–461. [Google Scholar]





| Sub-Collection | Country of Origin | Group | Germplasm Numbers (n) | |
|---|---|---|---|---|
| Polish | POL | I | A | 51 |
| B | 49 | |||
| C | 37 | |||
| West-Central European | CSK, DEU, DNK, AUT | II | A | 91 |
| B | 59 | |||
| French | FRA | III | A | 14 |
| B | 21 | |||
| Great British | GBR, IRL | IV | A | 22 |
| B | 8 | |||
| Other European countries | NLD, FIN, NOR | V | A | 16 |
| B | 15 | |||
| Swedish | SWE | VI | A | 12 |
| B | 26 | |||
| Other non-European countries | VII | 10 | ||
| Total | 431 | |||
| Chromosome | Powdery Mildew (PM) | Stem Rust (SR) | Barley Brown Rust (BBR) | ||||||
|---|---|---|---|---|---|---|---|---|---|
| 2019 | Maximum Scores Across All Replicates 2018–2019 | 2019 | Maximum Scores Across All Replicates 2018–2019 | 2019 | Maximum Scores Across All Replicates 2018–2019 | ||||
| Heading Stage (HA) | Milky-Waxy Stage (MW) | Heading Stage (HA) | Milky-Waxy Stage (MW) | Heading Stage (HA) | Milky-Waxy Stage (MW) | ||||
| 1H | 2 | 0 | 0 | 2 | 6 | 0 | 2 | 0 | 0 |
| 2H | 0 | 0 | 2 | 5 | 4 | 0 | 1 | 0 | 1 |
| 3H | 1 | 0 | 1 | 2 | 0 | 0 | 0 | 0 | 3 |
| 4H | 1 | 0 | 1 | 4 | 1 | 0 | 0 | 0 | 0 |
| 5H | 1 | 0 | 0 | 3 | 3 | 0 | 2 | 0 | 1 |
| 6H | 0 | 0 | 1 | 3 | 3 | 0 | 0 | 1 | 0 |
| 7H | 0 | 0 | 0 | 3 | 9 | 1 | 1 | 0 | 2 |
| Total | 5 | 0 | 5 | 22 | 26 | 1 | 6 | 1 | 7 |
| Disease | Year | Growth Stage | Powdery Mildew (PM) | Barley Brown Rust (BBR) | Stem Rust (SR) | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| 2019 | 2018 | 2019 | 2018 | 2019 | ||||||
| HA | MW | MW | HA | MW | MW | HA | MW | |||
| Powdery mildew (PM) | 2019 | HA | 1.000 | |||||||
| MW | 0.771 *** | 1.000 | ||||||||
| 2018 | MW | 0.315 ** | 0.186 * | 1.000 | ||||||
| Barley brown rust (BBR) | 2019 | HA | 0.309 ** | 0.194 * | 0.149 * | 1.000 | ||||
| MW | 0.215 ** | 0.123 * | 0.089 | 0.695 *** | 1.000 | |||||
| 2018 | MW | 0.144 * | 0.050 | 0.488 *** | 0.329 ** | 0.122 * | 1.000 | |||
| Stem rust (SR) | 2019 | HA | 0.174* | 0.126 * | 0.022 | 0.166 * | 0.133 * | 0.054 | 1.000 | |
| MW | 0.174 * | 0.092 | −0.012 | 0.277 ** | 0.231 ** | −0.012 | 0.074 | 1.000 | ||
| 2018 | MW | 0.100 * | 0.036 | 0.347 *** | 0.227 ** | 0.174 * | 0.383 *** | 0.062 | −0.019 | |
| Trait | SNP ID | Chromosome | Alleles | SNP Physical Localization | p-Value | FDR_Adjusted_p-Values |
|---|---|---|---|---|---|---|
| PM-HA | 3911365-45 | 1H | G/A | 368985776 | <0.0001 | 0.0698 |
| PM-HA | 3262583-22 | 1H | A/C | 519071852 | 0.7863 | 0.9757 |
| PM-HA | 3259998-68 | 3H | G/C | 17670801 | 0.0114 | 0.6563 |
| PM-HA | 3432490-28 | 4H | T/C | 627441392 | 0.3438 | 0.8421 |
| PM-HA | 4169584-26 | 5H | G/T | 650318560 | 0.0362 | 0.6563 |
| PM | 6278248-38 | 2H | T/C | 706966122 | <0.0001 | <0.0001 |
| PM | 3255391-29 | 3H | G/A | 611444890 | <0.0001 | <0.0001 |
| PM | 3432490-28 | 4H | T/C | 627441392 | <0.0001 | <0.0001 |
| PM | 6270346-59 | 6H | C/G | 352566440 | 0.0826 | 0.7632 |
| PM | 3254946-37 | 2H | C/T | 69199631 | 0.2554 | 0.9025 |
| Trait | SNP ID | Chromosome | Alleles | SNP Physical Localization | p-Value | FDR_Adjusted_p-Value |
|---|---|---|---|---|---|---|
| BBR-MW | 3432368-45 | 6H | A/G | 541093516 | <0.0001 | <0.0001 |
| BBR | 3254780-21 | 2H | C/T | 745071336 | 0.2754 | 0.7942 |
| BBR | 3255391-29 | 3H | G/A | 611444890 | 0.5133 | 0.9340 |
| BBR | 3398205-7 | 3H | A/G | 611554417 | 0.5133 | 0.9340 |
| BBR | 3432906-23 | 3H | T/G | 173445890 | 0.1095 | 0.7760 |
| BBR | 3256738-34 | 5H | A/G | 371708131 | 0.4108 | 0.8968 |
| BBR | 3919209-42 | 7H | G/C | 632737838 | 0.3551 | 0.8500 |
| BBR | 5260969-17 | 7H | T/G | 638276193 | 0.6841 | 0.9699 |
| Trait | SNP ID | Chromosome | Alleles | SNP Physical Localization | p-Value | FDR_Adjusted_p-Value |
|---|---|---|---|---|---|---|
| SR-HA | 3916722-6 | 1H | G/A | 443692735 | <0.0001 | <0.0001 |
| SR-HA | 7242068-56 | 1H | C/G | 54281150 | <0.0001 | <0.0001 |
| SR-HA | 3257827-54 | 2H | A/G | 612548328 | <0.0001 | <0.0001 |
| SR-HA | 3254861-28 | 2H | T/G | 672603288 | <0.0001 | <0.0001 |
| SR-HA | 3920041-34 | 3H | G/A | 553941251 | <0.0001 | <0.0001 |
| SR-HA | 3920041-34 | 3H | G/A | 553941251 | <0.0001 | <0.0001 |
| SR-HA | 100017008-29 | 3H | T/G | 236509272 | <0.0001 | <0.0001 |
| SR-HA | 3260326-22 | 5H | C/T | 584321761 | <0.0001 | <0.0001 |
| SR-HA | 3911523-28 | 5H | G/A | 582296115 | <0.0001 | <0.0001 |
| SR-HA | 3926286-9 | 6H | T/C | 582688589 | <0.0001 | <0.0001 |
| SR-HA | 8658544-48 | 6H | C/A | 19549390 | <0.0001 | <0.0001 |
| SR-HA | 3665999-27 | 6H | G/A | 89415341 | 0.0018 | 0.2114 |
| SR-HA | 6437148-27 | 7H | G/C | 625898188 | 0.9998 | 1.0000 |
| SR-HA | 7242117-49 | 7H | G/T | 576216690 | <0.0001 | 0.0086 |
| SR-MW | 3911365-45 | 1H | G/A | 368985776 | <0.0001 | 0.0059 |
| SR-MW | 3924215-36 | 1H | C/T | 368983947 | 0.0001 | 0.0157 |
| SR-MW | 3255467-46 | 1H | A/G | 361648993 | 0.0001 | 0.0200 |
| SR-MW | 6429360-14 | 1H | C/T | 361649059 | 0.0001 | 0.0200 |
| SR-MW | 4007106-27 | 1H | T/A | 87353500 | 0.0001 | 0.0269 |
| SR-MW | 7242068-56 | 1H | C/G | 54281150 | 0.0003 | 0.0462 |
| SR-MW | 3256098-57 | 2H | A/G | 724576571 | 0.0002 | 0.0300 |
| SR-MW | 4329845-19 | 2H | G/C | 724652573 | 0.0002 | 0.0344 |
| SR-MW | 3256445-31 | 2H | G/T | 651160368 | 0.0002 | 0.0398 |
| SR-MW | 3255089-30 | 2H | G/C | 760930749 | 0.0003 | 0.0462 |
| SR-MW | 3924288-41 | 4H | A/G | 36945544 | 0.0001 | 0.0160 |
| SR-MW | 14350408-63 | 5H | T/C | 453639849 | 0.0001 | 0.0183 |
| SR-MW | 3254700-12 | 5H | C/G | 622959955 | 0.0002 | 0.0360 |
| SR-MW | 3398368-6 | 5H | C/G | 623060574 | 0.0003 | 0.0398 |
| SR-MW | 7244989-23 | 6H | G/A | 56338324 | <0.0001 | 0.0115 |
| SR-MW | 3261100-7 | 6H | G/A | 549171429 | 0.0001 | 0.0200 |
| SR-MW | 4790439-14 | 6H | C/G | 548288936 | 0.0002 | 0.0360 |
| SR-MW | 3925588-20 | 7H | T/G | 472163680 | <0.0001 | 0.0049 |
| SR-MW | 4792770-43 | 7H | A/G | 384858359 | <0.0001 | 0.0049 |
| SR-MW | 3918497-64 | 7H | A/G | 552832603 | <0.0001 | 0.0049 |
| SR-MW | 3920255-11 | 7H | G/A | 553641369 | <0.0001 | 0.0049 |
| SR-MW | 6277097-23 | 7H | C/G | 555109356 | <0.0001 | 0.0049 |
| SR-MW | 3432458-59 | 7H | T/C | 357545360 | <0.0001 | 0.0049 |
| SR-MW | 5258884-14 | 7H | G/T | 372119583 | <0.0001 | 0.0049 |
| SR-MW | 13142583-10 | 7H | G/A | 556433100 | <0.0001 | 0.0049 |
| SR-MW | 3258004-51 | 7H | T/C | 606094789 | 0.0003 | 0.0445 |
| SR | 3918497-64 | 7H | A/G | 552832603 | <0.0001 | 0.0049 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Czembor, J.H.; Czembor, E.; Suchecki, R.; Watson-Haigh, N.S. Genome-Wide Association Study for Powdery Mildew and Rusts Adult Plant Resistance in European Spring Barley from Polish Gene Bank. Agronomy 2022, 12, 7. https://doi.org/10.3390/agronomy12010007
Czembor JH, Czembor E, Suchecki R, Watson-Haigh NS. Genome-Wide Association Study for Powdery Mildew and Rusts Adult Plant Resistance in European Spring Barley from Polish Gene Bank. Agronomy. 2022; 12(1):7. https://doi.org/10.3390/agronomy12010007
Chicago/Turabian StyleCzembor, Jerzy H., Elzbieta Czembor, Radoslaw Suchecki, and Nathan S. Watson-Haigh. 2022. "Genome-Wide Association Study for Powdery Mildew and Rusts Adult Plant Resistance in European Spring Barley from Polish Gene Bank" Agronomy 12, no. 1: 7. https://doi.org/10.3390/agronomy12010007
APA StyleCzembor, J. H., Czembor, E., Suchecki, R., & Watson-Haigh, N. S. (2022). Genome-Wide Association Study for Powdery Mildew and Rusts Adult Plant Resistance in European Spring Barley from Polish Gene Bank. Agronomy, 12(1), 7. https://doi.org/10.3390/agronomy12010007








