Impact of Water Stress on Microbial Community and Activity in Sandy and Loamy Soils
Abstract
1. Introduction
2. Materials and Methods
2.1. Experimental Setup
2.2. Chemical and Microbiological Analysis
2.3. Statistical Analysis
3. Results and Discussion
3.1. Impact of Water Stress on Nitrogen Forms and Nitrification Potential
3.2. Impact of Water Stress on Enzyme Activities
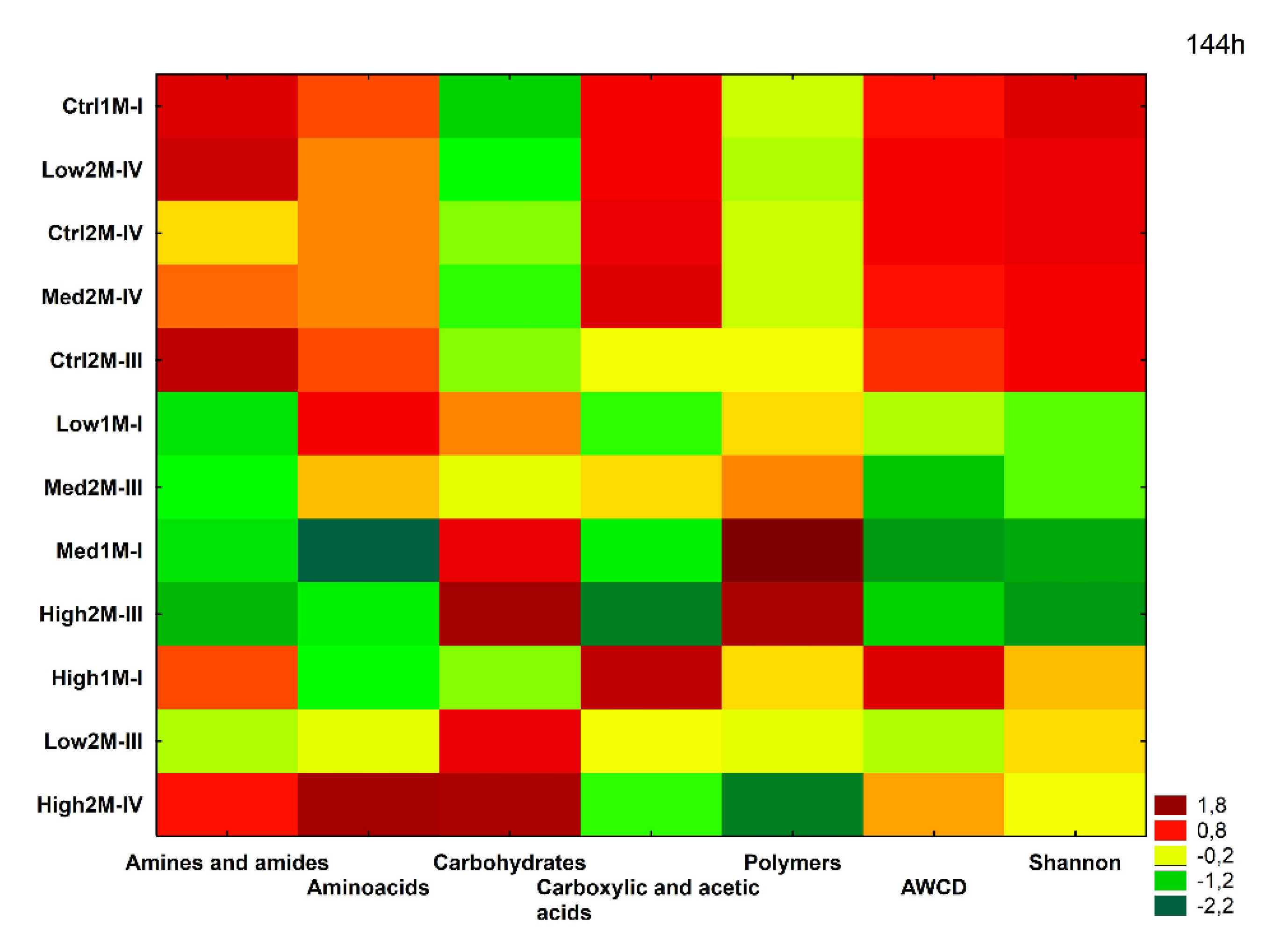
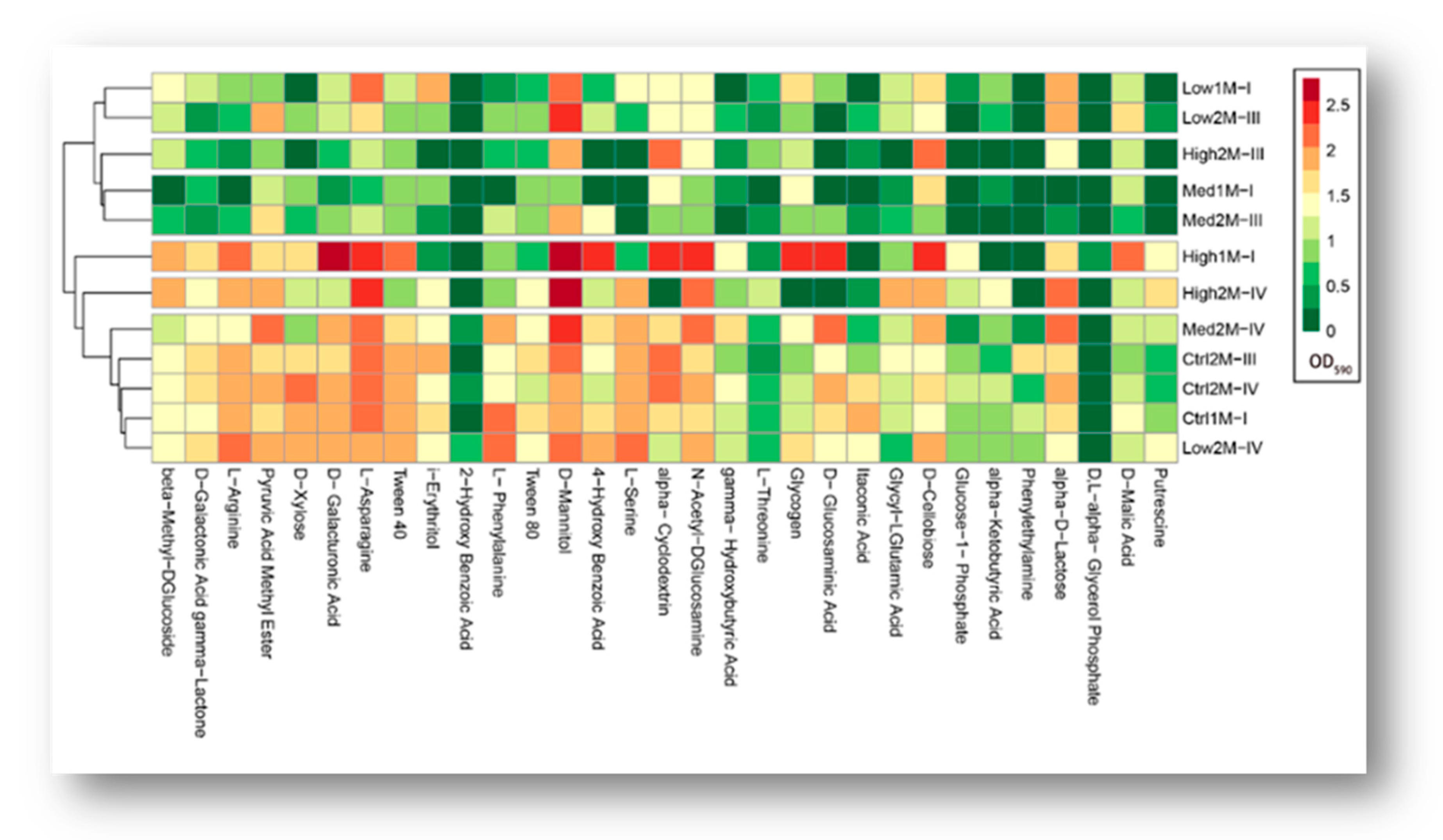
3.3. Impact of Water Stress on Functional Diversity of Bacteria
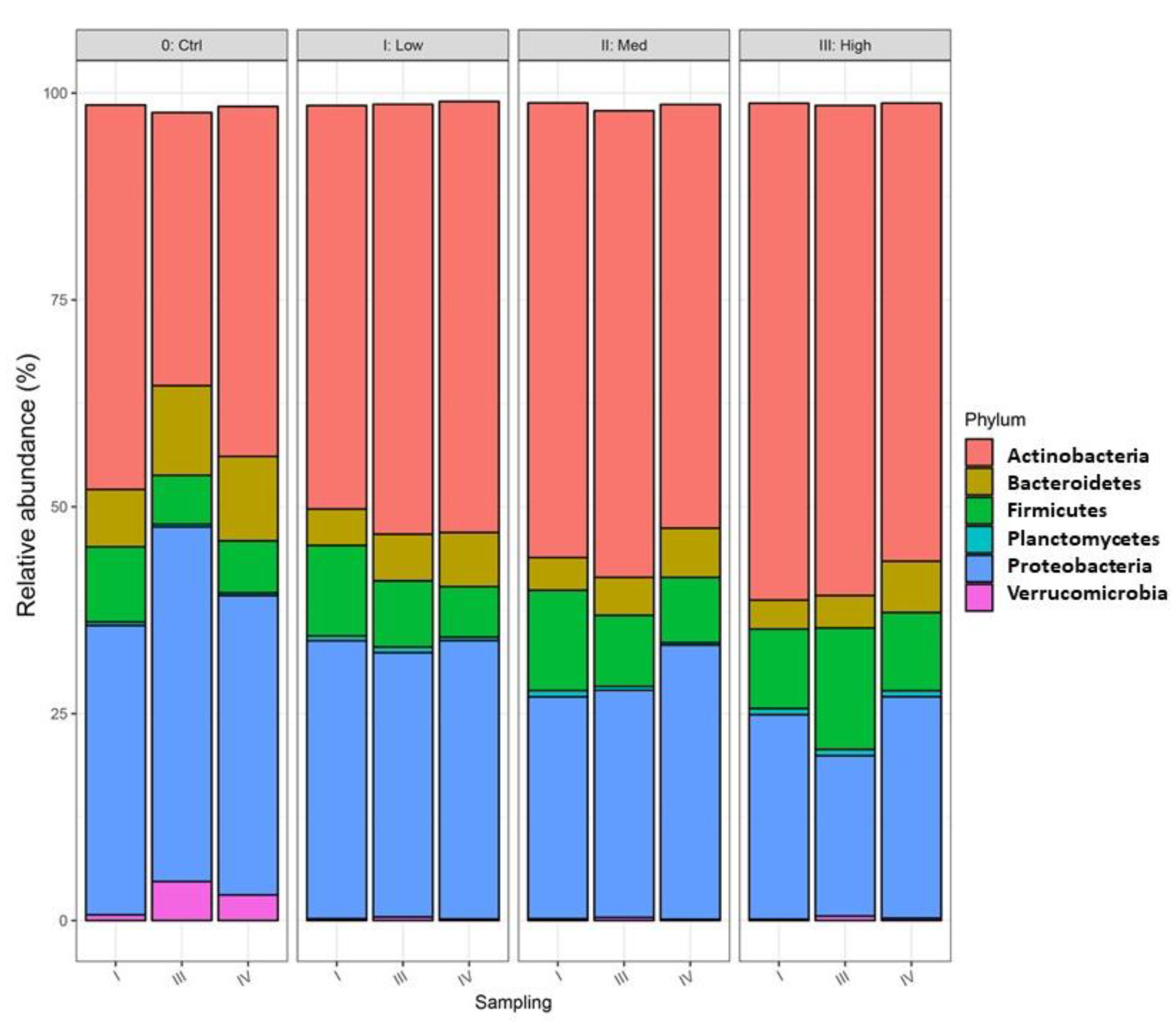
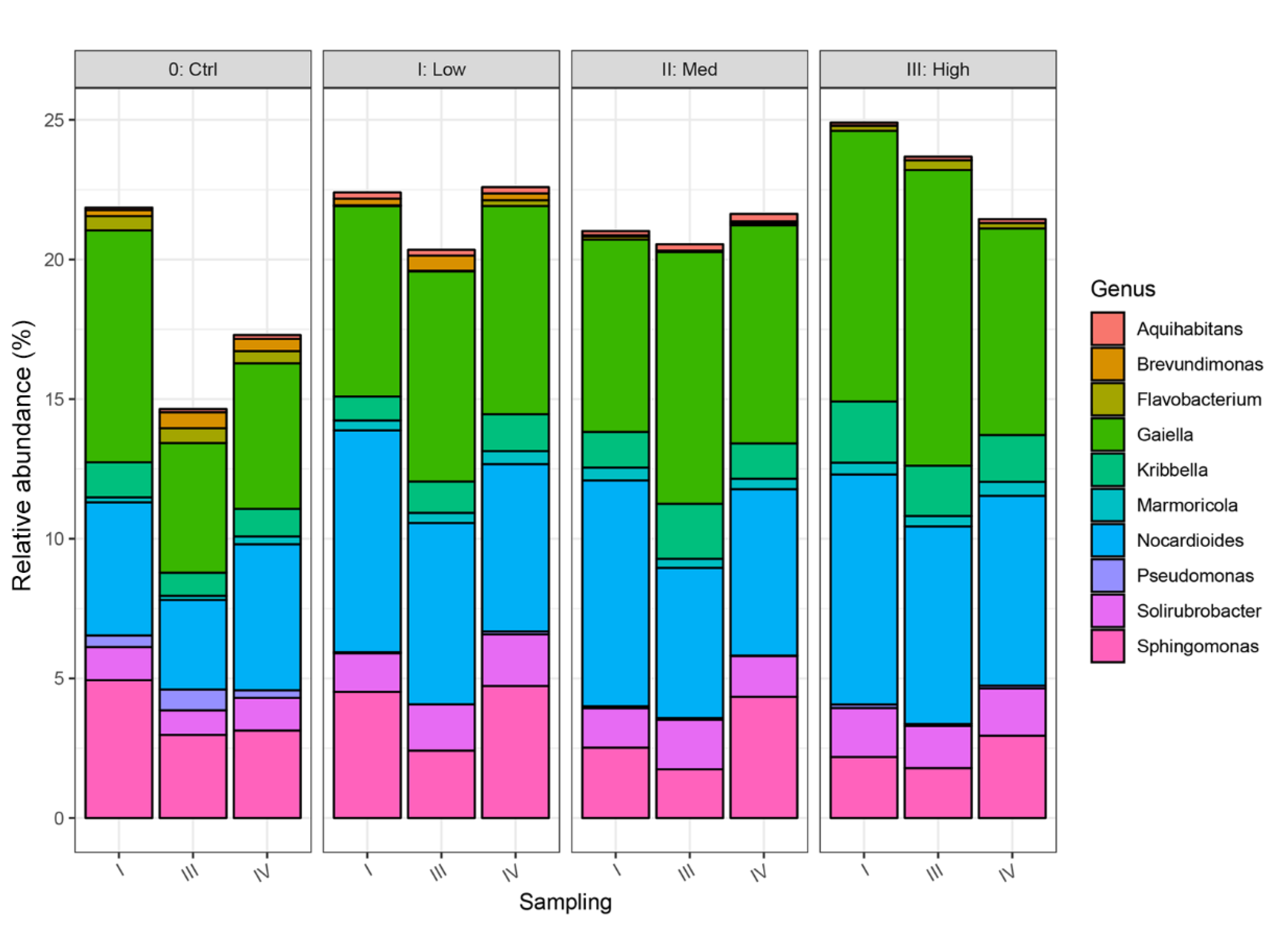
3.4. Impact of Water Stress on Genetic Diversity of Bacteria
3.5. Soil pH Change Under Water Stress
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Trenberth, K. Changes in precipitation with climate change. Clim. Res. 2011, 47, 123–138. [Google Scholar] [CrossRef]
- Hynčica, M.; Huth, R. Long-term changes in precipitation phase in Europe in cold half year. Atmos. Res. 2019, 227, 79–88. [Google Scholar] [CrossRef]
- Christensen, O.B.; Christensen, J.H. Intensification of extreme European summer precipitation in a warmer climate. Glob. Planet. Chang. 2004, 44, 107–117. [Google Scholar] [CrossRef]
- Grillakis, M.G. Increase in severe and extreme soil moisture droughts for Europe under climate change. Sci. Total Environ. 2019, 660, 1245–1255. [Google Scholar] [CrossRef] [PubMed]
- System of Monitoring of Agricultural Drought. 2020. Available online: http://www.susza.iung.pulawy.pl (accessed on 30 July 2020).
- Meisner, A.; Rousk, J.; Baath, E. Prolonged drought changes the bacterial growth response to rewetting. Soil Biol. Biochem. 2015, 88, 314–322. [Google Scholar] [CrossRef]
- Siebert, J.; Sünnemann, M.; Auge, H.; Berger, S.; Cesarz, S.; Ciobanu, M.; Guerrero-Ramírez, N.R.; Eisenhauer, N. The effects of drought and nutrient addition on soil organisms vary across taxonomic groups, but are constant across seasons. Sci. Rep. 2019, 9, 639. [Google Scholar] [CrossRef]
- Wang, S.; Cai, M.; Li, D.; Wu, B.; Li, J.; Huang, G.; Duan, H.; Wu, J. Heat waves intensify the effects of drought on bacterial diversity but not community composition in Solanum lycopersicum soil. J. Soils Sediments 2020. [Google Scholar] [CrossRef]
- Milosević, N.A.; Marinković, J.B.; Tintor, B.B. Mitigating abiotic stress in crop plants by microorganisms. Proc. Nat. Sci. Matica Serpska Novi. Sad 2012, 123, 17–26. [Google Scholar] [CrossRef]
- Nguyen, L.T.T.; Osanai, Y.; Anderson, I.C.; Bange, M.P.; Tissue, D.T.; Singh, B.K. Flooding and prolonged drought have differential legacy impacts on soil nitrogen cycling, microbial communities and plant productivity. Plant. Soil 2018, 431, 371–387. [Google Scholar] [CrossRef]
- Young, I.M.; Ritz, K. Tillage, habitat space and function of soil microbes. Soil Tillage Res. 2000, 53, 201–213. [Google Scholar] [CrossRef]
- Gleeson, D.B.; Herrmann, A.M.; Livesley, S.J.; Murphy, D.V. Influence of water potential on nitrification and structure of nitrifying bacterial communities in semiarid soils. Appl. Soil Ecol. 2008, 40, 189–194. [Google Scholar] [CrossRef]
- Gianfreda, L.; Bollag, J.M. Enzyme activity in soil. Soil Biol. Biochem. 1996, 8, 123–192. [Google Scholar]
- Preece, C.; Verbruggen, E.; Liu, L.; Weedon, J.T.; Peñuelas, J. Effects of past and current drought on the composition and diversity of soil microbial communities. Soil Biol. Biochem. 2019, 131, 28–39. [Google Scholar] [CrossRef]
- Casida, L.; Klein, D.; Santoro, T. Soil dehydrogenase activity. Soil Sci. 1964, 98, 371–376. [Google Scholar] [CrossRef]
- Tabatabai, M.A.; Bremner, J.M. Use of p-nitrophenylphosphate for assay of soil phosphatase activity. Soil Biol. Biochem. 1969, 1, 301–307. [Google Scholar] [CrossRef]
- ISO 15685. Soil Quality—Determination of Potential Nitrification and Inhibition of Nitrification—Rapid Test by Ammonium Oxidation; International Organisation for Standardisation: Geneva, Switzerland, 2004. [Google Scholar]
- Garland, J.L.; Mills, A.L. Classification and characterization of heterotrophic microbial communities on the basis of patterns of community-level sole-carbon-source utilization. Appl. Environ. Microbiol. 1991, 57, 2351–2359. [Google Scholar] [CrossRef]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glöckner, F.O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013, 41, e1. [Google Scholar] [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef]
- R Core Team. A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018. [Google Scholar]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naive Bayesian Classifier for Rapid Assignment of rRNA Sequences into the New Bacterial Taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef]
- McMurdie, P.J.; Holmes, S. Phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 2013, 8, e61217. [Google Scholar] [CrossRef]
- Abdi, H.; Williams, L. Principal components analysis. Wiley Interdiscip. Rev. Comput. Stat. 2010, 2, 439–459. [Google Scholar] [CrossRef]
- Kovach, W.L. MVSP—A Multivariate Statistical Package for Windows ver. 3.1; Kovach Computing Service: Pentraeth, UK, 1999. [Google Scholar]
- Fuchslueger, L.; Wild, B.; Mooshammer, M.; Takriti, M.; Kienzl, S.; Knoltsch, A.; Hofhansl, F.; Bahn, M.; Richter, A. Microbial carbon and nitrogen cycling responses to drought and temperature in differently managed mountain grasslands. Soil Biol. Biochem. 2019, 135, 144–153. [Google Scholar] [CrossRef]
- Maliszewska-Kordybach, B.; Klimkowicz-Pawlas, A.; Smreczak, B.; Janusauskaite, D. Ecotoxic effect of phenanthrene on nitrifying bacteria in soils of different properties. J. Environ. Qual. 2007, 36, 1635–1645. [Google Scholar] [CrossRef]
- Suszek-Łopatka, B.; Maliszewska-Kordybach, B.; Klimkowicz-Pawlas, A.; Smreczak, B. Influence of temperature on phenanthrene toxicity towards nitrifying bacteria in three soils with different properties. Environ. Poll. 2016, 216, 911–918. [Google Scholar] [CrossRef]
- Fuchslueger, L.; Kastl, E.M.; Bauer, F.; Kienzl, S.; Hasibeder, R.; Ladreiter-Knauss, T.; Schmidt, M.; Bahn, M.; Schloter, M.; Richter, A.; et al. Effect of drought on nitrogen turnover and abundances of ammonia-oxidizers in mountain grassland. Biogeosciences 2014, 11, 6003–6015. [Google Scholar] [CrossRef]
- Thion, C.; Prosser, J.I. Differential response of nonadapted ammonia-oxidising archea and bacteria to drying-rewetting stress. FEMS Microbiol. Ecol. 2014, 90, 380–389. [Google Scholar] [CrossRef] [PubMed]
- Bravo-Garza, M.R.; Bryan, R.B.; Voroney, P. Influence of wetting and drying cycles and maize residue addition on the formation of water stable aggregates in Vertisols. Geoderma 2009, 151, 150–156. [Google Scholar] [CrossRef]
- Xiang, S.-R.; Doyle, A.; Holden, P.A.; Schimel, J.P. Drying and rewetting effects on C and N mineralization and microbial activity in surface and subsurface California grassland soils. Soil Biol. Biochem. 2008, 40, 2281–2289. [Google Scholar] [CrossRef]
- Henry, H.A.L. Soil extracellular enzyme dynamics in a changing climate. Soil Biol. Biochem. 2012, 47, 53–59. [Google Scholar] [CrossRef]
- Steinwag, J.M.; Dukes, J.S.; Wallenstein, M.D. Modeling the effects of temperature and moisture on soil enzyme activity: Linking laboratory assays to continuous field data. Soil Biol. Biochem. 2012, 55, 85–92. [Google Scholar] [CrossRef]
- Sardans, J.; Peñuelas, J. Drought decreases soil enzyme activity in a Mediterranean Quercus ilex L. forest. Soil Biol. Biochem. 2005, 37, 455–461. [Google Scholar] [CrossRef]
- Siebielec, G.; Siebielec, S.; Lipski, D. Long-term impact of sewage sludge, digestate and mineral fertilizers on plant yield and soil biological activity. J. Clean. Prod. 2018, 187, 372–379. [Google Scholar] [CrossRef]
- Fernandes, S.A.; Bettiol, W.; Cerri, C.C. Effect of sewage sludge on microbial biomass, basal respiration, metabolic quotient and soil enzymatic activity. Appl. Soil Ecol. 2005, 30, 65–77. [Google Scholar] [CrossRef]
- Pascual, I.; Antolin, M.C.; Garcia, C.; Polo, A.; Sanchez-Diaz, M. Effect of water deficit on microbial characteristics in soil amended with sewage sludge or inorganic fertilizer under laboratory conditions. Bioresour. Technol. 2007, 98, 140–144. [Google Scholar] [CrossRef] [PubMed]
- Arif, M.S.; Riaz, M.; Shahzad, S.M.; Yasmeen, T.; Ashraf, M.; Siddique, M.; Mubarik, M.S.; Bragazza, L.; Buttler, A. Fresh and composted industrial sludge restore soil functions in surface soil of degraded agricultural land. Sci. Total Environ. 2018, 619, 517–527. [Google Scholar] [CrossRef]
- García-Gil, J.C.; Plaza, C.; Soler-Rovira, P.; Polo, A. Long-term effects of municipal solid waste compost application on soil enzyme activities and microbial biomass. Soil Biol. Biochem. 2000, 32, 1907–1913. [Google Scholar] [CrossRef]
- Crecchio, C.; Curci, M.; Pizzigallo, M.D.R.; Ricciuti, P.; Ruggiero, P. Effects of municipal solid waste compost amendments on soil enzyme activities and bacterial genetic diversity. Soil Biol. Biochem. 2004, 36, 1595–1605. [Google Scholar] [CrossRef]
- Garland, J.L. Analytical approaches to the characterization of samples of microbial communities using patterns of potential C source utilization. Soil Biol. Biochem. 1996, 28, 213–221. [Google Scholar] [CrossRef]
- Preece, C.; Farré-Armengol, G.; Peñuelas, J. Drought is a stronger driver of soil respiration and microbial communities than nitrogen or phosphorus addition in two Mediterranean tree species. Sci. Total Environ. 2020, 735, 139554. [Google Scholar] [CrossRef]
- Zhang, C.; Tayyab, M.; Abubakar, A.Y.; Yang, Z.; Pang, Z.; Islam, W.; Lin, Z.; Li, S.; Luo, J.; Fan, X.; et al. Bacteria with different assemblages in the soil profile drive the diverse nutrient cycles in the sugarcane straw retention ecosystem. Diversity 2019, 11, 194. [Google Scholar] [CrossRef]
- Fierer, N.; Bradford, M.A.; Jackson, R.B. Toward an ecological classification of soil bacteria. Ecology 2007, 88, 1354–1364. [Google Scholar] [CrossRef] [PubMed]
- Van Horn, D.J.; Okie, J.G.; Buelow, H.N.; Gooseff, M.N.; Barrett, J.E.; Takacs-Vesbach, C.D. Soil microbial responses to increased moisture and organic resources along a salinity gradient in a polar desert. Appl. Environ. Microbiol. 2014, 80, 3034–3043. [Google Scholar] [CrossRef]
- Veach, A.M.; Zeglin, L.H. Historical drought affects microbial population dynamics and activity during soil drying and re-wet. Microb. Ecol. 2020, 79, 662–674. [Google Scholar] [CrossRef] [PubMed]
- Zeng, Q.; Dong, Y.; An, S. Bacterial community responses to soils along a latitudinal and vegetation gradient on the Loess Plateau, China. PLoS ONE 2016, 11, e0152894. [Google Scholar] [CrossRef] [PubMed]
- Bergmann, G.T.; Bates, S.T.; Eilers, K.G.; Lauber, C.L.; Caporaso, J.G.; Walters, W.A.; Knight, R.; Fierer, N. The under-recognized dominance of Verrucomicrobia in soil bacterial communities. Soil Biol. Biochem. 2011, 43, 1450–1455. [Google Scholar] [CrossRef]
- Dastager, S.G.; Lee, J.C.; Ju, Y.J.; Park, D.J.; Kim, C.J. Nocardioides halotolerans sp. nov., isolated from soil on Bigeum Island, Korea. Syst. Appl. Microbiol. 2008, 31, 24–29. [Google Scholar] [CrossRef]
- Manandhar, P.; Zhang, G.; Lama, A.; Liu, F.; Hu, Y. Sphingomonas montana sp. nov., isolated from a soil sample from the Tanggula Mountain in the Qinghai Tibetan Plateau. Antonie van Leeuwenhoek 2017, 110, 1659–1668. [Google Scholar] [CrossRef]
- Coppotelli, B.M.; Ibarrolaza, A.; Del Panno, M.T.; Morelli, I.S. Effects of the inoculant strain Sphingomonas paucimobilis 20006FA on soil bacterial community and biodegradation in phenanthrene-contaminated soil. Microb. Ecol. 2008, 55, 173–183. [Google Scholar] [CrossRef]
- Xu, J.; Kloepper, J.W.; Huang, P.; McInroy, J.A.; Hu, C.H. Isolation and characterization of N2-fixing bacteria from giant reed and switchgrass for plant growth promotion and nutrient uptake. J. Basic Microbiol. 2018, 58, 459–471. [Google Scholar] [CrossRef]
- Park, Y.H.; Yoon, J.H.; Shin, Y.K.; Suzuki, K.; Kudo, T.; Seino, A.; Kim, H.J.; Lee, J.S.; Lee, S.T. Classification of ’Nocardioides fulvus’ IFO 14399 and Nocardioides sp. ATCC 39419 in Kribbella gen. nov., as Kribbella flavida sp. nov. and Kribbella sandramycini sp. nov. Int. J. Syst. Bacteriol. 1999, 49, 743–752. [Google Scholar] [CrossRef]
- Mohammadipanah, F.; Hamedi, J.; Goker, M.; Fiebig, A.; Pukall, R.; Sproer, C.; Klenk, H.-P. Kribbella shirazensis sp. nov., isolated from Iranian soil. Int. J. Syst. Evol. Microbiol. 2013, 63, 3369–3374. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Shi, L.; Lu, H.; Shao, Y.; Liu, S.; Fu, S. Drought promotes soil phosphorus transformation and reduces phosphorus bioavailability in a temperate forest. Sci. Total Environ. 2020, 732, 139295. [Google Scholar] [CrossRef] [PubMed]


| Stress Level | Ammonia-N (mg kg−1) | Nitrite-N (mg kg−1) | Nitrate-N (mg kg−1) | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| I 1 | II | III | IV | I | II | III | IV | I | II | III | IV | |
| Loamy soil | ||||||||||||
| Ctrl | 0.27 c2 | 0.00 b | 0.06 c | 0.43 a | 0.07 c | 0.07 b | 0.11 c | 0.07 a | 87.4 a | 84.9 a | 14.2 b | 9.6 c |
| Low | 5.94 b | 0.00 b | 2.64 bc | 0.72 a | 3.45 b | 0.09 b | 2.12 b | 0.22 a | 165.7 a | 95.2 a | 137.1 a | 36.6 a |
| Med | 14.1 a | 0.00 b | 4.24 b | 0.00 a | 8.81 a | 0.04 b | 3.08 b | 0.07 a | 190.4 a | 43.1 a | 102.6 ab | 13.8 bc |
| High | 12.2 a | 8.74 a | 12.31 a | 0.00 a | 6.81 a | 6.72 a | 8.31 a | 0.10 a | 170.6 a | 50.4 a | 57.7 ab | 26.4 ab |
| Sandy soil | ||||||||||||
| Ctrl | 1.95 b | 0.45 b | 0.60 c | 1.05 a | 0.25 a | 0.22 b | 0.67 a | 0.44 a | 154.4 a | 48.3 b | 60.8 b | 50.3 ab |
| Low | 4.72 a | 1.11 b | 2.19 b | 0.95 a | 0.59 a | 0.18 b | 0.34 a | 0.12 b | 132.0 a | 165.8 a | 212.7 a | 76.8 a |
| Med | 4.74 a | 0.77 b | 3.71 a | 1.20 a | 1.25 a | 0.15 b | 0.46 a | 0.08 b | 161.9 a | 35.3 b | 214.4 a | 24.6 bc |
| High | 4.16 a | 3.32 a | 4.96 a | 1.47 a | 1.19 a | 0.96 a | 0.79 a | 0.16 b | 171.0 a | 126.3 a | 86.4 b | 10.9 c |
| Sandy soil amended with compost | ||||||||||||
| Ctrl | 1.17 b | 0.28 b | 0.63 c | 0.93 ab | 0.16 b | 0.14 b | 0.56 a | 0.51 a | 112.0 b | 65.2 b | 37.2 b | 54.9 ab |
| Low | 4.95 a | 0.60 b | 2.06 bc | 0.47 b | 0.68 ab | 0.21 b | 0.70 a | 0.11 a | 322.5 ab | 211.1 a | 361.3 a | 89.7 a |
| Med | 6.61 a | 0.32 b | 3.54 ab | 0.99 ab | 1.55 a | 0.09 b | 0.88 a | 0.16 a | 326.6 ab | 52.9 b | 327.7 a | 31.2 bc |
| High | 5.42 a | 3.78 a | 4.83 a | 1.27 a | 1.09 a | 1.22 a | 1.00 a | 0.14 a | 410.5 a | 167.0 a | 105.2 b | 5.5 c |
| Stress Level | Nitrification Potential (µg NO2−·g DM−1) | |||
|---|---|---|---|---|
| I 1 | II | III | IV | |
| Loamy soil | ||||
| Ctrl | 7.47 a2 | 8.87 a | 6.89 a | 6.85 a |
| Low | 2.12 b | 8.36 a | 3.42 b | 8.77 a |
| Med | 2.12 b | 7.25 a | 2.89 b | 8.54 a |
| High | 2.50 b | 2.00 b | 2.03 b | 5.85 a |
| Sandy soil | ||||
| Ctrl | 4.71 a | 6.08 a | 10.92 a | 8.66 a |
| Low | 1.17 b | 3.89 b | 1.41 b | 3.25 b |
| Med | 0.97 b | 4.19 b | 1.89 b | 3.36 b |
| High | 1.06 b | 1.01 c | 1.46 b | 3.19 b |
| Sandy soil amended with compost | ||||
| Ctrl | 4.73 a | 5.78 a | 10.99 a | 9.28 a |
| Low | 1.22 b | 4.39 ab | 1.43 b | 3.70 b |
| Med | 0.98 b | 4.80 a | 1.77 b | 6.15 ab |
| High | 1.02 b | 1.57 b | 1.56 b | 4.13 b |
| Stress Level | Dehydrogenases (μg g DM−1 24 h−1) | Acidic Phosphatase (μg g DM−1 h−1) | Alkaline Phosphatase (μg g DM−1 h−1) | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| I 1 | II | III | IV | I | II | III | IV | I | II | III | IV | |
| Loamy soil | ||||||||||||
| Ctrl | 163.1 a2 | 160.7 a | 159.3 a | 181.7 a | 19.2 a | 15.1 a | 151.8 a | 138.5 a | 8.1 a | 7.8 a | 61.4 a | 84.0 a |
| Low | 73.7 b | 101.6 b | 35.7 b | 95.3 b | 12.4 b | 14.9 a | 98.2 b | 146.6 a | 5.2 ab | 7.4 a | 44.8 b | 72.9 a |
| Med | 51.6 b | 97.8 c | 33.9 b | 81.1 b | 11.5 b | 13.9 a | 104.5 b | 146.6 a | 3.9 b | 5.1 a | 43.4 b | 68.3 a |
| High | 46.0 b | 60.2 c | 33.1 b | 78.9 b | 11.4 b | 12.4 a | 96.9 b | 121.6 a | 3.4 b | 4.1 a | 37.8 b | 59.9 a |
| Sandy soil | ||||||||||||
| Ctrl | 45.9 a | 35.9 a | 42.0 a | 40.6 a | 3.4 a | 3.2 a | 30.5 a | 41.7 a | 2.4 a | 2.0 a | 16.6 a | 14.8 a |
| Low | 4.2 b | 7.8 b | 1.8 b | 2.7 b | 2.3 b | 2.8 a | 16.9 b | 23.8 b | 1.0 b | 1.3 b | 5.0 b | 5.7 b |
| Med | 3.1 b | 5.3 b | 3.7 b | 5.0 b | 2.2 b | 3.0 a | 18.5 b | 23.4 b | 1.0 b | 1.3 b | 5.1 b | 7.0 b |
| High | 3.4 b | 2.4 b | 3.0 b | 2.6 b | 2.1 b | 2.5 a | 17.8 b | 26.8 b | 1.1 b | 1.1 b | 3.8 b | 8.0 b |
| Sandy soil amended with compost | ||||||||||||
| Ctrl | 49.4 a | 45.7 a | 35.1 a | 38.0 a | 3.4 a | 3.8 a | 28.4 a | 29.2 a | 3.0 a | 2.3 a | 32.6 a | 68.4 a |
| Low | 20.4 b | 27.2 ab | 6.7 c | 17.8 c | 2.1 b | 3.0 b | 16.1 b | 23.3 b | 1.4 b | 1.7 a | 7.9 a | 19.3 b |
| Med | 23.4 b | 22.0 b | 8.5 bc | 23.8 b | 2.2 b | 3.1 b | 16.6 b | 23.6 b | 1.6 b | 2.1 a | 16.8 a | 15.7 b |
| High | 22.2 b | 21.4 b | 11.5 b | 28.8 b | 2.3 b | 2.2 c | 17.6 b | 23.1 b | 1.7 b | 2.0 a | 12.8 a | 19.4 b |
| Stress Level | Soil pH | |||
|---|---|---|---|---|
| I 1 | II | III | IV | |
| Loamy soil | ||||
| Ctrl | 7.20 a2 | 6.57 ab | 6.63 a | 6.97 a |
| Low | 6.60 b | 6.53 ab | 6.40 ab | 6.63 b |
| Med | 6.43 bc | 6.63 a | 6.27 b | 6.60 b |
| High | 6.37 c | 6.47 c | 6.40 ab | 6.53 b |
| Sandy soil | ||||
| Ctrl | 6.87 a | 7.03 a | 6.77 a | 6.70 c |
| Low | 6.70 ab | 6.87 b | 6.60 b | 6.73 bc |
| Med | 6.60 b | 7.07 a | 6.47 c | 6.90 ab |
| High | 6.57 b | 6.83 b | 6.43 c | 6.93 a |
| Sandy soil amended with compost | ||||
| Ctrl | 6.87a | 7.03a | 6.97a | 6.93b |
| Low | 6.60b | 6.80b | 6.60b | 6.90b |
| Med | 6.53 b | 7.03 a | 6.63 b | 7.00 a |
| High | 6.57 b | 6.73 b | 6.67 b | 7.10 a |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Siebielec, S.; Siebielec, G.; Klimkowicz-Pawlas, A.; Gałązka, A.; Grządziel, J.; Stuczyński, T. Impact of Water Stress on Microbial Community and Activity in Sandy and Loamy Soils. Agronomy 2020, 10, 1429. https://doi.org/10.3390/agronomy10091429
Siebielec S, Siebielec G, Klimkowicz-Pawlas A, Gałązka A, Grządziel J, Stuczyński T. Impact of Water Stress on Microbial Community and Activity in Sandy and Loamy Soils. Agronomy. 2020; 10(9):1429. https://doi.org/10.3390/agronomy10091429
Chicago/Turabian StyleSiebielec, Sylwia, Grzegorz Siebielec, Agnieszka Klimkowicz-Pawlas, Anna Gałązka, Jarosław Grządziel, and Tomasz Stuczyński. 2020. "Impact of Water Stress on Microbial Community and Activity in Sandy and Loamy Soils" Agronomy 10, no. 9: 1429. https://doi.org/10.3390/agronomy10091429
APA StyleSiebielec, S., Siebielec, G., Klimkowicz-Pawlas, A., Gałązka, A., Grządziel, J., & Stuczyński, T. (2020). Impact of Water Stress on Microbial Community and Activity in Sandy and Loamy Soils. Agronomy, 10(9), 1429. https://doi.org/10.3390/agronomy10091429









