Amaranthus palmeri a New Invasive Weed in Spain with Herbicide Resistant Biotypes
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material
2.2. Dose–Response Trials
2.3. Alternative Herbicides
2.4. ALS Gene Sequencing
2.5. ISSR DNA Fingerprinting
2.6. Data Analysis
3. Results
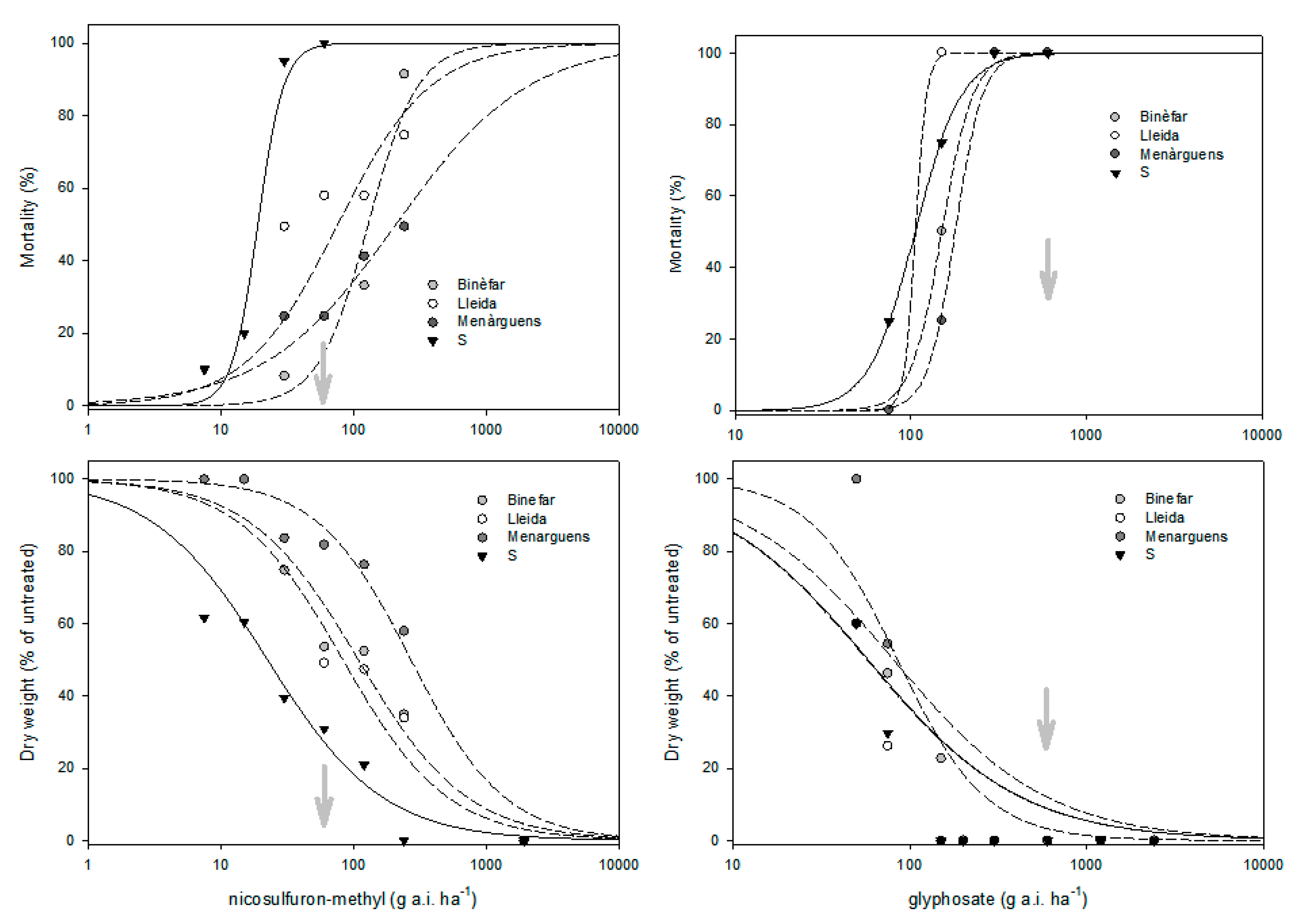
3.1. Dose–Response Trials
3.2. Alternative Herbicides
3.3. ALS Gene Sequencing
3.4. ISSR DNA Fingerprint
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Paini, D.R.; Sheppard, A.W.; Cook, D.C.; De Barro, P.J.; Worner, S.P.; Thomas, M.B. Global threat to agriculture from invasive species. Proc. Natl. Acad. Sci. USA 2016, 113, 7575–7579. [Google Scholar] [CrossRef] [PubMed]
- Pimentel, D.; Zuniga, R.; Morrison, D. Update on the environmental and economic costs associated with alien-invasive species in the United States. Ecol. Econ. 2005, 52, 273–288. [Google Scholar] [CrossRef]
- Boscutti, F.; Sigura, M.; De Simone, S.; Marini, L. Exotic plant invasion in agricultural landscapes: A matter of dispersal mode and disturbance intensity. Appl. Veg. Sci. 2018, 21, 250–257. [Google Scholar] [CrossRef]
- Lyons, K.; Prather, T. Invasive Plant Ecology in Natural and Agricultural Systems. Invasive Plant Sci. Manag. 2013, 6, 457. [Google Scholar] [CrossRef]
- Ehleringer, J. Ecophysiology of Amaranthus palmeri, a Sonoran Desert summer annual. Oecologia 1983, 57, 107–112. [Google Scholar] [CrossRef] [PubMed]
- Chaudhari, S.; Jordan, D.; York, A.; Jennings, K.M.; Cahoon, C.W.; Chandi, A.; Inman, M.D. Biology and management of glyphosate-resistant and glyphosate-susceptible Palmer amaranth (Amaranthus palmeri) phenotypes from a segregating population. Weed Sci. 2017, 65, 755–768. [Google Scholar] [CrossRef]
- Keeley, P.E.; Carter, C.H.; Thullen, R.J. Influence of planting date on growth of Palmer amaranth (Amaranthus palmeri). Weed Sci. 1987, 35, 199–204. [Google Scholar] [CrossRef]
- Smith, K.L.; Doherty, R.C.; Bullington, J.A.; Meier, J.R.; Bagavathiannan, M.V. Seed production potential of Palmer amaranth in Arkansas. In Summaries of Arkansas Cotton Research 2011; Oosterhuis, D.E., Ed.; University of Arkansas Division of Agriculture AAES Research Series 602: Little Rock, AR, USA, 2011; pp. 40–43. [Google Scholar]
- Sosnoskie, L.; Webster, T.; Culpepper, A. Glyphosate resistance does not affect Palmer amaranth (Amaranthus palmeri) seedbank longevity. Weed Sci. 2013, 61, 283–288. [Google Scholar] [CrossRef]
- Ward, S.; Webster, T.; Steckel, L. Palmer amaranth (Amaranthus palmeri): A review. Weed Technol. 2013, 27, 12–27. [Google Scholar] [CrossRef]
- EPPO [European and Mediterranean Plant Protection Organization] (2020) EPPO Alert List. Available online: http://www.eppo.int/INVASIVE_PLANTS/ias_lists.htm#AlertList (accessed on 8 January 2020).
- Carretero, J.L. Amaranthus. In Flora Iberica Vol II: Platanaceae—Plumbaginaceae (Partim); Castroviejo, S., Ed.; Real Jardín Botánico CSIC: Madrid, Spain, 1990; pp. 559–569. [Google Scholar]
- Sánchez-Guillón, E.; Verloove, F. New records of interesting xenophytes in Spain. II. Lagascalia 2009, 29, 281–291. [Google Scholar]
- Verloove, F.; Sánchez-Guillón, E. New records of interesting xenophytes in the Iberian Peninsula. Acta Bot. Malac. 2008, 33, 147–167. [Google Scholar] [CrossRef][Green Version]
- Recasens, J.; Conesa, J.A. Presencia de la mala hierba Amaranthus palmeri en el NE de la península Ibérica. Una amenaza como potencial invasora de cultivos extensivos de regadío. Boletín De Sanid. Veg. Plagas 2011, 37, 129–132. [Google Scholar]
- Recasens, J.; Osuna, M.D.; Royo-Esnal, A.; Torra, J. Amaranthus palmeri S. Watson en Cataluña y Aragón. Tres poblaciones con un mismo origen? In Proceedings of the XVI Congreso de la Sociedad Española de Malherbología, Pamplona, Spain, 25–27 October 2017; pp. 15–20. [Google Scholar]
- Palma-Bautista, C.; Torra, J.; García, M.J.; Bracamonte, E.R.; Rojano-Delgado, A.M.; Alcántara-de la Cruz, R.; De Prado, R. Reduced absorption and impaired translocation endows glyphosate resistance in Amaranthus palmeri harvested in glyphosate-resistant soybean from Argentina. J. Agric. Food Chem. 2019, 67, 1052–1060. [Google Scholar] [CrossRef] [PubMed]
- Gaines, T.A.; Zhang, W.; Wang, D.; Bukun, B.; Chisholm, S.T.; Shaner, D.; Nissen, S.J.; Patzoldt, W.L.; Tranel, P.; Culpepper, A.S.; et al. Gene amplification confers glyphosate resistance in Amaranthus palmeri. Proc. Natl. Acad. Sci. USA 2009, 107, 1029–1034. [Google Scholar] [CrossRef] [PubMed]
- Domínguez-Valenzuela, J.A.; Gherekhloo, J.; Fernández-Moreno, P.T.; Cruz-Hipolito, H.E.; La Cruz, R.A.-D.; Sánchez-González, E.; De Prado, R. First confirmation and characterization of target and non-target site resistance to glyphosate in Palmer amaranth (Amaranthus palmeri) from Mexico. Plant Physiol. Biochem. 2017, 115, 212–218. [Google Scholar] [CrossRef] [PubMed]
- Heap, I. International Survey of Herbicide Resistant Weeds. Available online: http://www.weedscience.org (accessed on 15 January 2020).
- Horak, M.J.; Peterson, D.E. Biotypes of Palmer amaranth (Amaranthus palmeri) and common waterhemp (Amaranthus rudis) are resistant to imazethapyr and thifensulfuron. Weed Technol. 1995, 9, 192–195. [Google Scholar] [CrossRef]
- Larran, A.S.; Palmieri, V.E.; Perotti, V.E.; Lieber, L.; Tuesca, D.; Permingeat, H.R. Target-site resistance to acetolactate synthase (ALS)-inhibiting herbicides in Amaranthus palmeri from Argentina. Pest Manag. Sci. 2017, 73, 2578–2584. [Google Scholar] [CrossRef]
- Tranel, P.J.; Wright, T.R.; Heap, I.M. Mutations in Herbicide-Resistant Weeds to ALS Inhibitors. Available online: http://www.weedscience.com (accessed on 15 January 2020).
- Varanasi, V.; Brabham, C.; Korres, N.; Norsworthy, J. Nontarget site resistance in Palmer amaranth [Amaranthus palmeri (S.) Wats.] confers cross-resistance to protoporphyrinogen oxidase-inhibiting herbicides. Weed Technol. 2019, 33, 349–354. [Google Scholar] [CrossRef]
- Bravo, W.; Leon, R.G.; Ferrell, J.A.; Mulvaney, M.J.; Wood, C.W. Differentiation of life-history traits among Palmer amaranth populations (Amaranthus palmeri) and its relation to cropping systems and glyphosate sensitivity. Weed Sci. 2017, 65, 339–349. [Google Scholar] [CrossRef]
- Prado, M.D.; De Prado, R.; Franco, A.R. Design and optimization of degenerated universal primers for the cloning of the plant acetolactate synthase conserved domains. Weed Sci. 2004, 52, 487–491. [Google Scholar] [CrossRef]
- Nei, M. Genetic distance between populations. Am. Nat. 1972, 106, 283–292. [Google Scholar] [CrossRef]
- Rohlf, F.J. NTSYS-Numerical Taxonomy and Multivariate Analysis System; Exeter Publishing Setauket: New York, NY, USA, 1998. [Google Scholar]
- Kumar, V.; Liu, R.; Boyer, G.; Stahlman, P.W. Confirmation of 2,4-D resistance and identification of multiple resistance in a Kansas Palmer amaranth (Amaranthus palmeri) population. Pest Manag. Sci. 2019, 75, 2925–2933. [Google Scholar] [CrossRef] [PubMed]
- Mohseni-Moghadam, M.; Schroeder, J.; Heerema, R.; Ashigh, J. Resistance to glyphosate in Palmer amaranth (Amaranthus palmeri) populations from New Mexico pecan orchards. Weed Technol. 2013, 27, 85–91. [Google Scholar] [CrossRef]
- Kaundun, S.S.; Jackson, L.V.; Hutchings, S.-J.; Galloway, J.; Marchegiani, E.; Howell, A.; Carlin, R.; Mcindoe, E.; Tuesca, D.; Moreno, R. Evolution of Target-Site Resistance to Glyphosate in an Amaranthus palmeri Population from Argentina and Its Expression at Different Plant Growth Temperatures. Plants 2019, 8, 512. [Google Scholar] [CrossRef] [PubMed]
- Berger, S.; Madeira, P.; Ferrell, J.; Gettys, L.; Morichetti, S.; Cantero, J.; Nuñez, C. Palmer amaranth (Amaranthus palmeri) identification and documentation of ALS-resistance in Argentina. Weed Sci. 2016, 64, 312–320. [Google Scholar] [CrossRef]
- Yu, Q.; Powles, S.B. Resistance to AHAS inhibitor herbicides: Current understanding. Pest Manag. Sci. 2014, 70, 1340–1350. [Google Scholar] [CrossRef]
- Norsworthy, J.K.; Ward, S.M.; Shaw, D.R.; Llewellyn, R.S.; Nichols, R.L.; Webster, T.M.; Bradley, K.W.; Frisvold, G.; Powles, S.B.; Burgos, N.R.; et al. Reducing the risks of herbicide resistance: Best management practices and recommendations. Weed Sci. 2012, 60, 31–62. [Google Scholar] [CrossRef]
- Slotta, T.A.B. What we know about weeds: Insights from genetic markers. Weed Sci. 2008, 56, 322–326. [Google Scholar] [CrossRef]
- Dinelli, G.; Bonetti, A.; Marotti, I.; Minelli, M.; Catizone, P. Characterization of Italian populations of Lolium spp. resistant and susceptible to diclofop by inter simple sequence repeat. Weed Sci. 2004, 52, 554–563. [Google Scholar] [CrossRef]
- Mengistu, L.A.W.; Messersmith, C.G.; Christoffers, M.J. Genetic diversity of herbicide-resistant and -susceptible Avena fatua populations in North Dakota and Minnesota. Weed Res. 2005, 45, 413–423. [Google Scholar] [CrossRef]
- Küpper, A.; Borgato, E.; Patterson, E.; Gonçalves Netto, A.; Nicolai, M.; de Carvalho, S.J.P.; Nissen, S.J.; Gaines, T.A.; Christoffoleti, P.J. Multiple resistance to glyphosate and acetolactate synthase inhibitors in Palmer amaranth (Amaranthus palmeri) identified in Brazil. Weed Sci. 2017, 65, 317–326. [Google Scholar] [CrossRef]
- Patzoldt, W.; Tranel, P. Multiple ALS mutations confer herbicide resistance in waterhemp (Amaranthus tuberculatus). Weed Sci. 2007, 55, 421–428. [Google Scholar] [CrossRef]
- Culpepper, A.S.; Grey, T.L.; Vencill, W.K.; Kichler, J.M.; Webster, T.M.; Brown, S.M.; York, A.C.; Davis, J.W.; Hanna, W.W. Glyphosate-resistant Palmer amaranth (Amaranthus palmeri) confirmed in Georgia. Weed Sci. 2006, 54, 620–626. [Google Scholar] [CrossRef]
- DOGC. Ordre ARP/172/2019, de 10 de setembre, per la qual es declara l’existència de la mala herba Amaranthus palmeri i es qualifica d’utilitat pública la lluita contra aquesta; Diari Oficial de la Generalitat de Catalunya núm. 7959; Entitat Autònoma del Diari Oficial i de Publicacions (EADOP): Barcelona, Spain, 13 September 2019. [Google Scholar]
- Recasens, J.; Conesa, J.A.; Juárez-Escario, A. Las invasiones vegetales en sistemas agrícolas. Retrospectiva de los últimos 40 años en Cataluña. Inf. Tec. Econ. Agrar. 2019. [Google Scholar] [CrossRef]

| Name | Sequence |
|---|---|
| Ama BE 2 Forward | 5′-TGCTATTGGAGCTGCTGTTG-3′ |
| Ama BE 1 Reverse | 5′-CCTTCTTCCATCACCCTCTG-3′ |
| Ama CAD 1 Forward | 5′-CGCCCTCTTCAAATCTCATC-3′ |
| Ama CAD Reverse | 5′-CAATCAAAACAGGTCCAGGTC-3′ |
| Name | Sequence |
|---|---|
| ISSR 808 | 5′-AGAGAGAGAGAGAGAGC-3′ |
| ISSR 889 | 5′-DBDACACACACACACAC-3′ |
| ISSR 890 | 5′-VHVGTGTGTGTGTGTGT-3′ |
| ISSR 857 | 5′-ACACACACACACACACYG-3′ |
| ISSR 836 | 5′-AGAGAGAGAGAGAGAGYA-3′ |
| ISSR 827 | 5′-ACACACACACACACACG-3′ |
| ISSR 880 | 5′-GGAGAGGAGAGGAGA-3′ |
| ISSR 810 | 5′-GAGAGAGAGAGAGAGAT-3′ |
| Mortality (%) | Dry Weight Reduction (%) | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Herbicide | Field Dose | Population | b | LD50 (g a.i.·ha−1) | RI | p-Value | b | GR50 (g a.i.·ha−1) | RI | p-Value |
| Nicosulfuron-methyl | 60 g a.i.·ha−1 | Susceptible | 4.5 | 19.0 | – | 0.016 | −1.0 | 22.5 | – | 0.003 |
| Binéfar | 2.3 | 128.9 | 6.8 | 0.001 | −1.1 | 108.8 | 4.8 | 0.003 | ||
| Menàrguens | 1.2 | 76.0 | 4.0 | 0.011 | −1.1 | 83.1 | 3.7 | 0.019 | ||
| Lleida | 0.9 | 208.7 | 11.0 | 0.004 | −1.2 | 271.0 | 12.1 | 0.002 | ||
| 600 g a.i.·ha−1 | Susceptible | 3.3 | 105.6 | – | 0.006 | −1.0 | 57.4 | – | 0.029 | |
| Glyphosate | Binéfar | 4.9 | 150.0 | 1.4 | 0.015 | −1.0 | 58.3 | 1.0 | 0.026 | |
| Menàrguens | 5.6 | 178.8 | 1.7 | 0.017 | −1.7 | 88.6 | 1.5 | 0.005 | ||
| Lleida | 13.6 | 106.5 | 1.0 | 0.014 | −1.0 | 81.2 | 1.4 | 0.005 | ||
| HRAC | Field Rate | Application | Mortality (%) | ||||
|---|---|---|---|---|---|---|---|
| Herbicide | Group | SoA * | (g a.i. ha−1) | Time | Binéfar | Menàrguens | Lleida |
| S-metolachlor | K3 | VLCFA | 960 | Pre-emergence | 100 ± 0 | 100 ± 0 | 100 ± 0 |
| Isoxaflutole | F2 | HPPD | 96 | Pre-emergence | 100 ± 0 | 100 ± 0 | 100 ± 0 |
| Mesotrione | F2 | HPPD | 150 | Post-emergence | 100 ± 0 | 100 ± 0 | 100 ± 0 |
| Sulcotrione | F2 | HPPD | 300 | Post-emergence | 100 ± 0 | 100 ± 0 | 100 ± 0 |
| Dicamba | O | SAH | 288 | Post-emergence | 100 ± 0 | 100 ± 0 | 100 ± 0 |
| Thifensulfuron | B | ALS | 30 | Post-emergence | 32 ± 12 | 56 ± 7 | 8 ± 5 |
| Nicosulfuron | B | ALS | 60 | Post-emergence | 25 ± 10 | 25 ± 10 | 50 ± 11 |
| ALS | Population (Number of Plants) | |||||
|---|---|---|---|---|---|---|
| Domain | Position | Amino Acid | Genotype | Lleida | Binéfar | Menàrguens |
| BE | Trp574 | Leu | RS | 5 | 1 | 1 |
| RR | 3 | 0 | 0 | |||
| Ser653 | Asn | RS | 0 | 0 | 0 | |
| RR | 3 | 0 | 0 | |||
| Ile | RS | 0 | 0 | 0 | ||
| RR | 0 | 0 | 1 | |||
| CAD | Pro197 | Thr | RS | 0 | 2 | 4 |
| RR | 0 | 6 | 0 | |||
| Ser | RS | 0 | 0 | 0 | ||
| RR | 0 | 0 | 1 | |||
| Susceptible * | 9 | 11 | 13 | |||
| Resistant | 11 | 9 | 7 | |||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Torra, J.; Royo-Esnal, A.; Romano, Y.; Osuna, M.D.; León, R.G.; Recasens, J. Amaranthus palmeri a New Invasive Weed in Spain with Herbicide Resistant Biotypes. Agronomy 2020, 10, 993. https://doi.org/10.3390/agronomy10070993
Torra J, Royo-Esnal A, Romano Y, Osuna MD, León RG, Recasens J. Amaranthus palmeri a New Invasive Weed in Spain with Herbicide Resistant Biotypes. Agronomy. 2020; 10(7):993. https://doi.org/10.3390/agronomy10070993
Chicago/Turabian StyleTorra, Joel, Aritz Royo-Esnal, Yolanda Romano, María Dolores Osuna, Ramón G. León, and Jordi Recasens. 2020. "Amaranthus palmeri a New Invasive Weed in Spain with Herbicide Resistant Biotypes" Agronomy 10, no. 7: 993. https://doi.org/10.3390/agronomy10070993
APA StyleTorra, J., Royo-Esnal, A., Romano, Y., Osuna, M. D., León, R. G., & Recasens, J. (2020). Amaranthus palmeri a New Invasive Weed in Spain with Herbicide Resistant Biotypes. Agronomy, 10(7), 993. https://doi.org/10.3390/agronomy10070993









