Screening and Comparative Characterization of Microorganisms from Iranian Soil Samples Showing ω-Transaminase Activity toward a Plethora of Substrates
Abstract
1. Introduction
2. Results and Discussion
2.1. Microorganisms Screening and Identification
2.2. Enzymatic Properties and Substrate Specificity
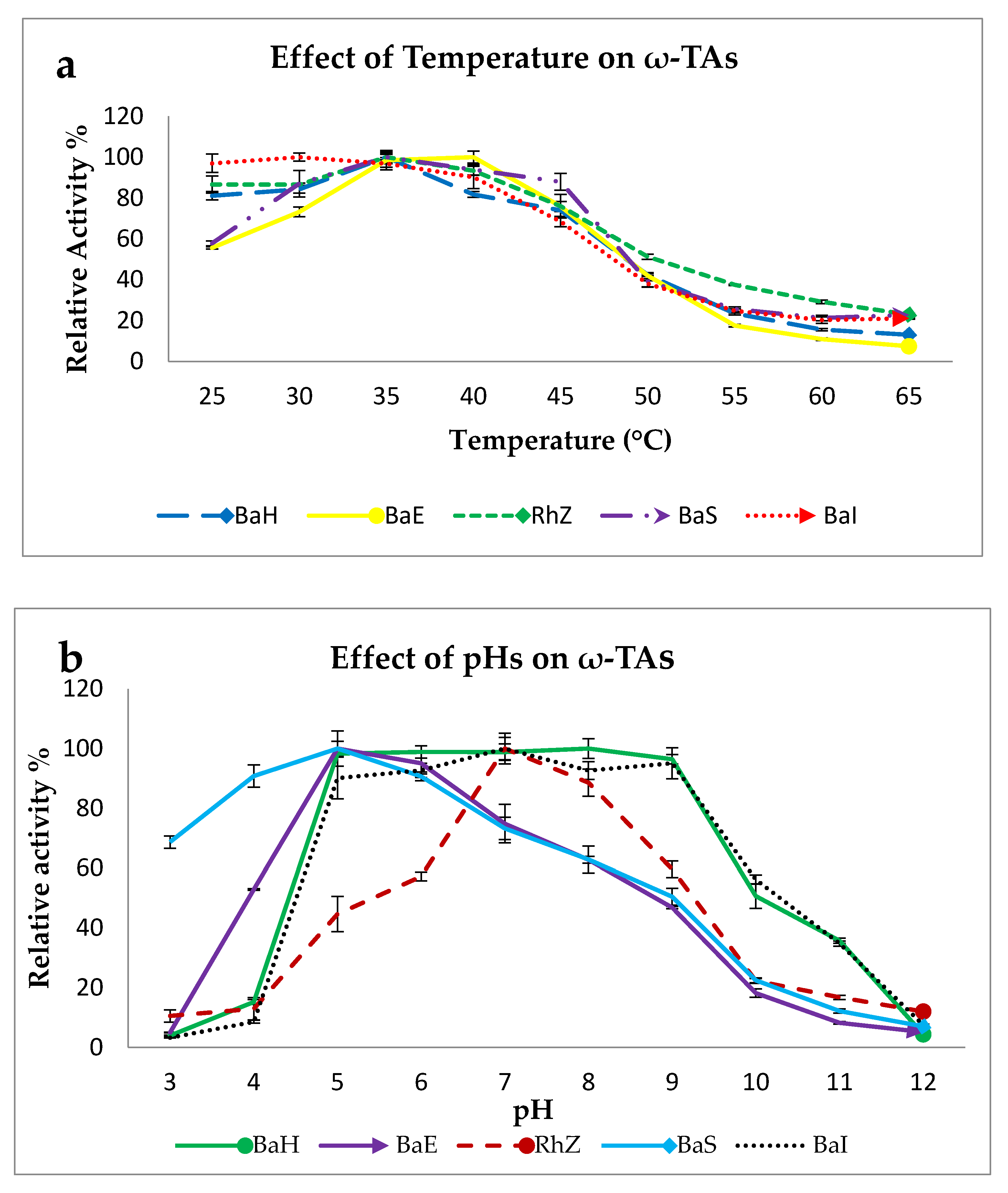
2.2.1. Effect of Temperature on ω-TA Activity
2.2.2. Effect of pH on ω-TA Activity
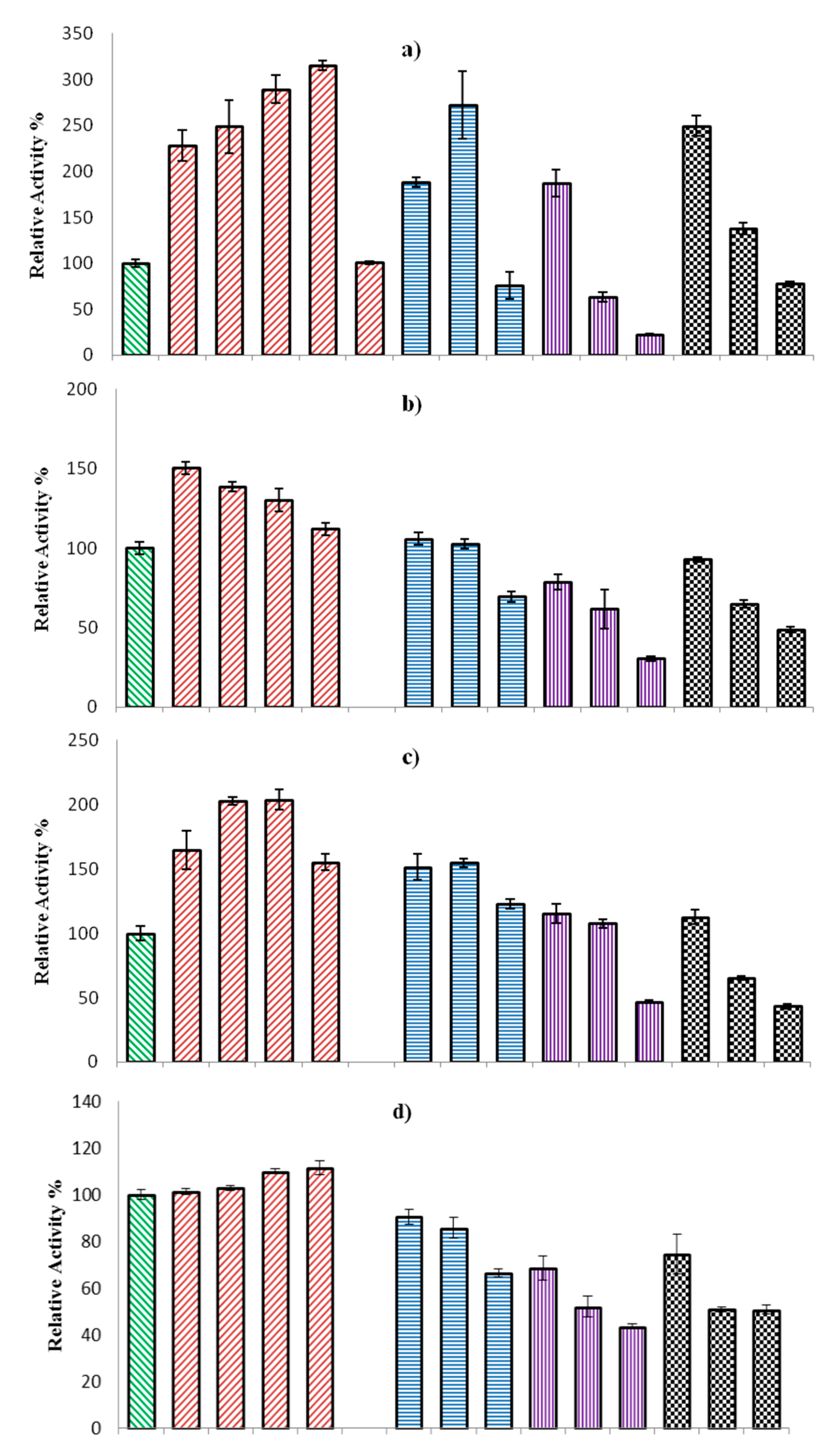
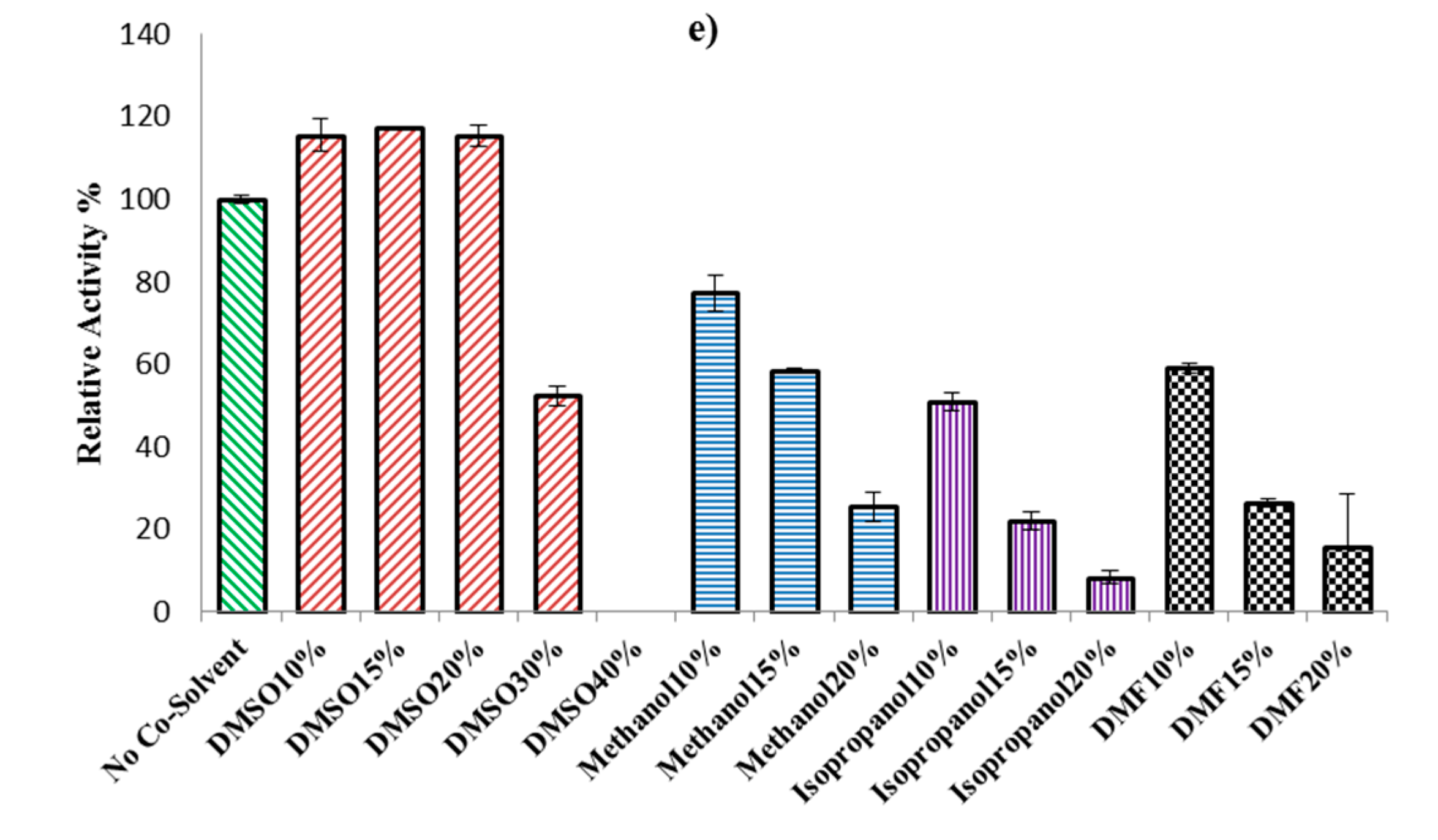
2.2.3. Effect of Organic Solvents on ω-TA Activity
2.2.4. Substrate Specificity and Enantioselectivity
3. Materials and Methods
3.1. Chemicals
3.2. Screening and Identification
3.2.1. Enrichment of Microorganisms on (rac)-α-MBA as Sole Nitrogen Source
3.2.2. Selection of the Most Promising Strains
3.2.3. Identification of Bacteria
3.3. Enzymatic Properties and Substrate Specificity
3.3.1. Preparation of Crude Extract
3.3.2. Effect of Temperature, pH, and Co-Solvent
3.3.3. Substrate Specificity and Enantioselectivity
3.4. HPLC Analytics
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Gomm, A.; Lewis, W.; Green, A.P.; O’Reilly, E. A New Generation of Smart Amine Donors for Transaminase-Mediated Biotransformations. Chem. A Eur. J. 2016, 22, 12692–12695. [Google Scholar] [CrossRef]
- Pawar, S.V.; Hallam, S.J.; Yadav, V.G. Metagenomic Discovery of a Novel Transaminase for Valorization of Monoaromatic Compounds. RSC Adv. 2018, 8, 22490–22497. [Google Scholar] [CrossRef]
- Aleku, G.A.; France, S.P.; Man, H.; Mangas-Sanchez, J.; Montgomery, S.L.; Sharma, M.; Leipold, F.; Hussain, S.; Grogan, G.; Turner, N.J. A Reductive Aminase from Aspergillus Oryzae. Nat. Chem. 2017, 9, 961. [Google Scholar] [CrossRef]
- Kim, G.H.; Jeon, H.; Khobragade, T.P.; Patil, M.D.; Sung, S.; Yoon, S.; Won, Y.; Choi, I.S.; Yun, H. Enzymatic Synthesis of Sitagliptin Intermediate Using a Novel ω-Transaminase. Enzym. Microb. Technol. 2019, 120, 52–60. [Google Scholar] [CrossRef]
- Mathew, S.; Nadarajan, S.P.; Chung, T.; Park, H.H.; Yun, H. Biochemical Characterization of Thermostable ω-Transaminase from Sphaerobacter Thermophilus and its Application for Producing Aromatic β-and γ-Amino Acids. Enzym. Microb. Technol. 2016, 87, 52–60. [Google Scholar] [CrossRef]
- Haycock-Lewandowski, S.J.; Wilder, A.; Åhman, J. Development of a Bulk Enabling Route to Maraviroc (UK-427,857), a CCR-5 Receptor Antagonist. Org. Process. Res. Dev. 2008, 12, 1094–1103. [Google Scholar] [CrossRef]
- Slomka, C.; Zhong, S.; Fellinger, A.; Engel, U.; Syldatk, C.; Bräse, S.; Rudat, J. Chemical Synthesis and Enzymatic, Stereoselective Hydrolysis of a Functionalized Dihydropyrimidine for the Synthesis of β-Amino Acids. AMB Express 2015, 5, 85. [Google Scholar] [CrossRef]
- Weiner, B.; Szymański, W.; Janssen, D.B.; Minnaard, A.J.; Feringa, B.L. Recent Advances in the Catalytic Asymmetric Synthesis of β-Amino Acids. Chem. Soc. Rev. 2010, 39, 1656–1691. [Google Scholar] [CrossRef]
- Turner, N.J.; Truppo, M.D. Biocatalytic Routes to Nonracemic Chiral Amines. In Chiral Amine Synthesis: Methods, Developments and Applications; Nugent, T.C., Ed.; Wiley-VCH: Weinheim, Germany, 2010; pp. 431–459. [Google Scholar]
- Mathew, S.; Jeong, S.S.; Chung, T.; Lee, S.H.; Yun, H. Asymmetric Synthesis of Aaromatic β-Amino Acids Using ω-Transaminase: Optimizing the Lipase Concentration to Obtain Thermodynamically Unstable β-Keto Acids. Biotechnol. J. 2016, 11, 185–190. [Google Scholar] [CrossRef]
- Malik, M.S.; Park, E.S.; Shin, J.S. Features and Technical Applications of ω-Transaminases. Appl. Microbiol. Biotechnol. 2012, 94, 1163–1171. [Google Scholar] [CrossRef]
- Jiang, J.; Chen, X.; Feng, J.; Wu, Q.; Zhu, D. Substrate Profile of an ω-Transaminase from Burkholderia Vietnamiensis and its Potential for the Production of Optically Pure Amines and Unnatural Amino Acids. J. Mol. Catal. B Enzym. 2014, 100, 32–39. [Google Scholar] [CrossRef]
- Cerioli, L.; Planchestainer, M.; Cassidy, J.; Tessaro, D.; Paradisi, F. Characterization of a Novel Amine Transaminase from Halomonas Elongata. J. Mol. Catal. B Enzym. 2015, 120, 141–150. [Google Scholar] [CrossRef]
- Voss, M.; Das, D.; Genz, M.; Kumar, A.; Kulkarni, N.; Kustosz, J.; Kumar, P.; Bornscheuer, U.T.; Höhne, M. In Silico Based Engineering Approach to Improve Transaminases for the Conversion of Bulky Substrates. ACS Catal. 2018, 8, 11524–11533. [Google Scholar] [CrossRef]
- Rudat, J.; Brucher, B.R.; Syldatk, C. Transaminases for the Synthesis of Enantiopure Beta-Amino Acids. AMB Express 2012, 2, 11. [Google Scholar] [CrossRef]
- Pavlidis, I.V.; Weiß, M.S.; Genz, M.; Spurr, P.; Hanlon, S.P.; Wirz, B.; Iding, H.; Bornscheuer, U.T. Identification of (S)-Selective Transaminases for the Asymmetric Synthesis of Bulky Chiral Amines. Nat. Chem. 2016, 8, 1076–1082. [Google Scholar] [CrossRef]
- Kelly, S.A.; Magill, D.J.; Megaw, J.; Skvortsov, T.; Allers, T.; McGrath, J.W.; Allen, C.C.; Moody, T.S.; Gilmore, B.F. Characterisation of a Solvent-Tolerant Haloarchaeal (R)-Selective Transaminase Isolated from a Triassic Period Salt Mine. Appl. Microbiol. Biotechnol. 2019, 103, 5727–5737. [Google Scholar] [CrossRef]
- Leipold, L.; Dobrijevic, D.; Jeffries, J.W.; Bawn, M.; Moody, T.S.; Ward, J.M.; Hailes, H.C. The Identification and Use of Robust Transaminases from a Domestic Drain Metagenome. Green Chem. 2019, 21, 75–86. [Google Scholar] [CrossRef]
- Ferrandi, E.E.; Previdi, A.; Bassanini, I.; Riva, S.; Peng, X.; Monti, D. Novel Thermostable Amine Transferases from Hot Spring Metagenomes. Appl. Microbiol. Biotechnol. 2017, 101, 4963–4979. [Google Scholar] [CrossRef]
- Genz, M.; Vickers, C.; van den Bergh, T.; Joosten, H.-J.; Dörr, M.; Höhne, M.; Bornscheuer, U. Alteration of the Donor/Acceptor Spectrum of the (S)-Amine Transaminase from Vibrio Fluvialis. Int. J. Mol. Sci. 2015, 16, 26953–26963. [Google Scholar] [CrossRef]
- Kelly, S.A.; Megaw, J.; Caswell, J.; Scott, C.J.; Allen, C.C.; Moody, T.S.; Gilmore, B.F. Isolation and Characterisation of a Halotolerant ω-Transaminase from a Triassic Period Salt Mine and Its Application to Biocatalysis. ChemistrySelect 2017, 2, 9783–9791. [Google Scholar] [CrossRef]
- Shin, J.S.; Kim, B.G. Kinetic Resolution of α-Methylbenzylamine with o-Transaminase Screened from Soil Microorganisms: Application of a Biphasic System to Overcome Product Inhibition. Biotechnol. Bioeng. 1997, 55, 348–358. [Google Scholar] [CrossRef]
- Pavkov-Keller, T.; Strohmeier, G.A.; Diepold, M.; Peeters, W.; Smeets, N.; Schürmann, M.; Gruber, K.; Schwab, H.; Steiner, K. Discovery and Structural Characterisation of New Fold Type IV-Transaminases Exemplify the Diversity of This Eenzyme Fold. Sci. Rep. 2016, 6, 38183. [Google Scholar] [CrossRef] [PubMed]
- Yazdi, A.; Emami, M.H.; Shafiee, S.M. Dasht-E Lut in Iran, the Most Complete Collection of Beautiful Geomorphological Phenomena of Desert. Open J. Geol. 2014, 4, 249–261. [Google Scholar] [CrossRef][Green Version]
- Ataee, N.; Fooladi, J.; Namaei, M.H.; Rezadoost, H.; Mirzajani, F. Biocatalysts Screening of Papaver Bracteatum Flora for Thebaine Transformation to Codeine and Morphine. Biocatal. Agric. Biotechnol. 2017, 9, 127–133. [Google Scholar] [CrossRef]
- Ghasemi, Y.; Rasoul-Amini, S.; Ebrahiminezhad, A.; Kazemi, A.; Shahbazi, M.; Talebnia, N. Screening and Isolation of Extracellular Protease Producing Bacteria from the Maharloo Salt Lake. Iran. J. Pharm. Sci. 2011, 7, 175–180. [Google Scholar]
- Alghabpoor, S.S.; Panosyan, H.; Trchounian, A.; Popov, Y. Purification and Characterization of a Novel Thermostable and Acid Stable α-Amylase from Bacillus Sp. Iranian S1. Int. J. Eng. Trans. B Appl. 2013, 26, 815–820. [Google Scholar] [CrossRef]
- Shirsalimian, M.; Amoozegar, M.; Sepahy, A.A.; Kalantar, S.; Dabbagh, R. Isolation of Extremely Halophilic Archaea from a Saline River in the Lut Desert of Iran, Moderately Resistant to Desiccation and Gamma Radiation. Microbiology 2017, 86, 403–411. [Google Scholar] [CrossRef]
- Shin, J.S.; Kim, B.G. Comparison of the ω-Transaminases from Different Microorganisms and Application to Production of Chiral Amines. Biosci. Biotechnol. Biochem. 2001, 65, 1782–1788. [Google Scholar] [CrossRef]
- Mathew, S.; Bea, H.; Nadarajan, S.P.; Chung, T.; Yun, H. Production of Chiral β-Amino Acids Using ω-Transaminase from Burkholderia Graminis. J. Biotechnol. 2015, 196, 1–8. [Google Scholar] [CrossRef]
- Guo, F.; Berglund, P. Transaminase Biocatalysis: Optimization and Application. Green Chem. 2017, 19, 333–360. [Google Scholar] [CrossRef]
- Dreßen, A.; Hilberath, T.; Mackfeld, U.; Billmeier, A.; Rudat, J.; Pohl, M. Phenylalanine Ammonia Lyase from Arabidopsis Thaliana (AtPAL2): A Potent MIO-Enzyme for the Synthesis of Non-Canonical Aromatic Alpha-Amino Acids: Part I: Comparative Characterization to the Enzymes from Petroselinum Crispum (PcPAL1) and Rhodosporidium Toruloides (RtPAL). J. Biotechnol. 2017, 258, 148–157. [Google Scholar]
- Schätzle, S.; Höhne, M.; Robins, K.; Bornscheuer, U.T. Conductometric Method for the Rapid Characterization of the Substrate Specificity of Amine-Transaminases. Anal. Chem. 2010, 82, 2082–2086. [Google Scholar] [CrossRef]
- Schätzle, S.; Höhne, M.; Redestad, E.; Robins, K.; Bornscheuer, U.T. Rapid and Sensitive Kinetic Assay for Characterization of ω-Transaminases. Anal. Chem. 2009, 81, 8244–8248. [Google Scholar] [CrossRef] [PubMed]
- Koszelewski, D.; Tauber, K.; Faber, K.; Kroutil, W. ω-Transaminases for the Synthesis of Non-Racemic α-Chiral Primary Amines. Trends Biotechnol. 2010, 28, 324–332. [Google Scholar] [CrossRef] [PubMed]
- Buß, O.; Dold, S.M.; Obermeier, P.; Litty, D.; Muller, D.; Grüninger, J.; Rudat, J. Enantiomer Discrimination in β-Phenylalanine Degradation by a Newly Isolated Paraburkholderia Strain BS115 and Type Strain PsJN. AMB Express 2018, 8, 149. [Google Scholar] [CrossRef] [PubMed]
- Conti, E.; Stachelhaus, T.; Marahiel, M.A.; Brick, P. Structural Basis for the Activation of Phenylalanine in the Non-Ribosomal Biosynthesis of Gramicidin S. EMBO J. 1997, 16, 4174–4183. [Google Scholar] [CrossRef] [PubMed]
- Yamada, M.; Kurahashi, K. Further Purification and Properties of Adenosine Triphosphate-Dependent Phenylalanine Racemase of Bacillus Brevis Nagano. J. Biochem. 1969, 66, 529–540. [Google Scholar] [CrossRef]
- Yun, H.; Lim, S.; Cho, B.K.; Kim, B.G. ω-Amino Acid: Pyruvate Transaminase from Alcaligenes Denitrificans Y2k-2: A New Catalyst for Kinetic Resolution of β-Amino Acids and Amines. Appl. Environ. Microbiol. 2004, 70, 2529–2534. [Google Scholar] [CrossRef]
- Temple, K.L.; Colmer, A.R. The Autotrophic Oxidation of Iron by a New Bacterium: Thiobacillus Ferrooxidans. J. Bacteriol. 1951, 62, 605. [Google Scholar]
- Buß, O.; Voss, M.; Delavault, A.; Gorenflo, P.; Syldatk, C.; Bornscheuer, U.; Rudat, J. β-Phenylalanine Ester Synthesis from Stable β-Keto Ester Substrate Using Engineered ω-Transaminases. Molecules 2018, 23, 1211. [Google Scholar] [CrossRef]
- Yoon, S.H.; Ha, S.M.; Kwon, S.; Lim, J.; Kim, Y.; Seo, H.; Chun, J. Introducing EzBioCloud: A taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int. J. Syst. Evol. Microbiol. 2017, 67, 1613. [Google Scholar] [PubMed]
- Brucher, B.; Rudat, J.; Syldatk, C.; Vielhauer, O. Enantioseparation of Aromatic β³-Amino Acid by Precolumn Derivatization with o-Phthaldialdehyde and N-Isobutyryl-l-Cysteine. Chromatographia 2010, 71, 1063–1067. [Google Scholar] [CrossRef]
- Buß, O.; Muller, D.; Jager, S.; Rudat, J.; Rabe, K.S. Improvement in the Thermostability of a β-Amino Acid Converting ω-Transaminase by Using FoldX. ChemBioChem 2018, 19, 379–387. [Google Scholar] [CrossRef] [PubMed]
- Gao, S.; Su, Y.; Zhao, L.; Li, G.; Zheng, G. Characterization of a (R)-Selective Amine Transaminase from Fusarium Oxysporum. Process Biochem. 2017, 63, 130–136. [Google Scholar] [CrossRef]
| Isolate | Closest Relative in GenBank | % Similarity/Sequence Length (bp) |
|---|---|---|
| BaH | Bacillus halotolerans (LPVF01000003) | 99.9/1388 |
| BaE | Bacillus endophyticus (AF295302) | 99.7/1273 |
| BaS | Bacillus subtilis subsp. stercoris (JHCA01000027) | 99/1385 |
| BaI | Bacillus subtilis subsp. inaquosorum (AMXN01000021) | 99.9/1406 |
| RhZ | Rhizobium radiobacter (AJ389904) | 99.4/1338 |
| Amino Donor | Relative Activity% | |||||||
|---|---|---|---|---|---|---|---|---|
| BaH | BaE | BaS | BaI | RhZ | ||||
| 1 | (rac) | α-methylbenzylamine | 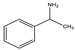 | 53 | 48 | 45 | 60 | 36 |
| 1s | (S) | 100 | 100 | 100 | 100 | 100 | ||
| 1r | (R) | 23 | 0 | 0 | 6 | 0 | ||
| 2 | (rac) | 3-amino-3-(4-chlorophenyl) propionic acid | 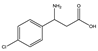 | 19 | 0 | 0 | 0 | 0 |
| 2s | (S) | 20 | 0 | 0 | 0 | 0 | ||
| 2r | (R) | 16 | 0 | 0 | 0 | 0 | ||
| 3 | (rac) | 3-amino-3-(3,4-dichloro-phenyl)propionic acid | 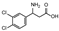 | 20 | 0 | 0 | 0 | 0 |
| 4 | (rac) | 3-amino-3-(4-nitrophenyl)propionic acid | 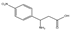 | 51 | 0 | 0 | 0 | 0 |
| 4r | (R) | 19 | 0 | 0 | 0 | 0 | ||
| 5r | (R) | (R)3-amino-3-(2-nitro-phenyl)-propionic acid | 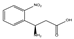 | 16 | 0 | 0 | 0 | 0 |
| 6 | (rac) | 3-amino-3-(4-methoxyphenyl) propionic acid |  | 21 | 0 | 0 | 0 | 0 |
| 6s | (S) | 17 | 0 | 0 | 0 | 0 | ||
| 7s | (S) | 3-amino-3-(4-hydroxy-phenyl)propionic acid | 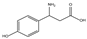 | 18 | 0 | 0 | 0 | 0 |
| 7r | (R) | 15 | 0 | 0 | 0 | 0 | ||
| 8 | (rac) | 3-amino-3-(4-bromophenyl)-propionic acid |  | 21 | 0 | 0 | 0 | 0 |
| 8r | (R) | 20 | 0 | 0 | 0 | 0 | ||
| 9 | (rac) | 3-amino-3-(4-fluorophenyl)propionic acid |  | 43 | 0 | 0 | 0 | 0 |
| 9s | (S) | 16 | 0 | 0 | 0 | 0 | ||
| 10 | (rac) | Methyl(-3-amino-3-phenyl propanoate hydrochloride) | 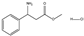 | 11 | 0 | 0 | 0 | 0 |
| 11 | (rac) | 3-amino-3-(4-isopropylphenyl) propionic acid |  | 20 | 0 | 0 | 0 | 0 |
| 12 | (rac) | 1-aminocyclopropane-1-carboxylic acid | 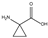 | 17 | 17 | 0 | 30 | 0 |
| 13 | (rac) | β-phenylalanine | 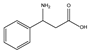 | 17 | 0 | 0 | 0 | 0 |
| 13s | (S) | 16 | 0 | 0 | 0 | 0 | ||
| 13r | (R) | 22 | 0 | 0 | 0 | 0 | ||
| 14 | (rac) | β-Homophenylalanine |  | 16 | 0 | 0 | 0 | 0 |
| 15 | Isopropylamine |  | 20 | 0 | 0 | 21 | 0 | |
| 16 | Sec-butylamine |  | 25 | 7 | 0 | 27 | 0 | |
| 17s | (S) | 3-amino butyric acid | 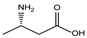 | 91 | 59 | 54 | 20 | 135 |
| Amino Acceptors | Relative Activity% | ||||||
|---|---|---|---|---|---|---|---|
| BaH | BaE | BaS | BaI | RhZ | |||
| 1b | Ethyl benzoylacetate | 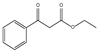 | 40.95 | 0 | 20.84 | 14.87 | 0 |
| 2b | Ethyl 3-(4-methoxyphenyl)-3-oxopropanoate | 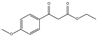 | 0 | 0 | 0 | 60.48 | 0 |
| 3b | Sodium 3-oxo-3-phenylpropanoate | 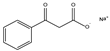 | 0 | 0 | 0 | 0 | 0 |
| 4b | Ethyl(4-flurobenzoyl)acetate | 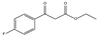 | 0 | 0 | 0 | 0 | 0 |
| 5b | Ethyl(4-chlorobenzoyl)acetate |  | 0 | 0 | 0 | 0 | 0 |
| 6b | Ethyl 4-nitrobenzoylacetate | 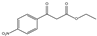 | 0 | 0 | 0 | 0 | 0 |
| 7b | Pyruvate |  | 100 | 100 | 43.73 | 100 | 36.39 |
| 8b | α-ketoglutarate | 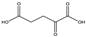 | 17.09 | 7.31 | 100 | 0 | 100 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gord Noshahri, N.; Fooladi, J.; Syldatk, C.; Engel, U.; Heravi, M.M.; Zare Mehrjerdi, M.; Rudat, J. Screening and Comparative Characterization of Microorganisms from Iranian Soil Samples Showing ω-Transaminase Activity toward a Plethora of Substrates. Catalysts 2019, 9, 874. https://doi.org/10.3390/catal9100874
Gord Noshahri N, Fooladi J, Syldatk C, Engel U, Heravi MM, Zare Mehrjerdi M, Rudat J. Screening and Comparative Characterization of Microorganisms from Iranian Soil Samples Showing ω-Transaminase Activity toward a Plethora of Substrates. Catalysts. 2019; 9(10):874. https://doi.org/10.3390/catal9100874
Chicago/Turabian StyleGord Noshahri, Najme, Jamshid Fooladi, Christoph Syldatk, Ulrike Engel, Majid M. Heravi, Mohammad Zare Mehrjerdi, and Jens Rudat. 2019. "Screening and Comparative Characterization of Microorganisms from Iranian Soil Samples Showing ω-Transaminase Activity toward a Plethora of Substrates" Catalysts 9, no. 10: 874. https://doi.org/10.3390/catal9100874
APA StyleGord Noshahri, N., Fooladi, J., Syldatk, C., Engel, U., Heravi, M. M., Zare Mehrjerdi, M., & Rudat, J. (2019). Screening and Comparative Characterization of Microorganisms from Iranian Soil Samples Showing ω-Transaminase Activity toward a Plethora of Substrates. Catalysts, 9(10), 874. https://doi.org/10.3390/catal9100874






