Biotransformation of Hydroxychalcones as a Method of Obtaining Novel and Unpredictable Products Using Whole Cells of Bacteria
Abstract
1. Introduction
2. Results and Discussion
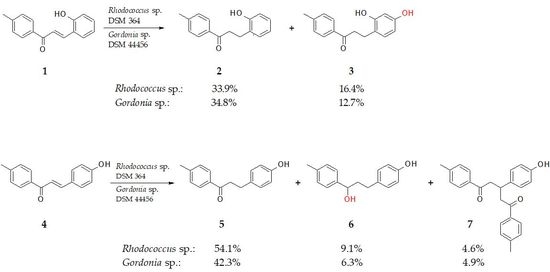
2.1. Biotransformation of 2-Hydroxy-4′-Methylchalcone (1)
2.2. Biotransformation of 4-Hydroxy-4′-Methylchalcone (4)
3. Materials and Methods
3.1. Chemicals
3.2. Synthesis of 2-Hydroxy-4′-Methylchalcone (1) and 4-Hydroxy-4′-Methylchalcone (4)
3.3. Biotransformations
3.3.1. Biotransformation in Small Scale
3.3.2. Biotransformation in Preparative Scale
3.4. Analytical Methods
3.5. Isolated Biotransformations Products
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Mai, C.W.; Yaeghoobi, M.; Abd-Rahman, N.; Kang, Y.B.; Pichika, M.R. Chalcones with electron-withdrawing and electron-donating substituents: Anticancer activity against TRAIL resistant cancer cells, structure-activity relationship analysis and regulation of apoptotic proteins. Eur. J. Med. Chem. 2014, 77, 378–387. [Google Scholar] [CrossRef] [PubMed]
- Moreira Osório, T.; Delle Monache, F.; Domeneghini Chiaradia, L.; Mascarello, A.; Regina Stumpf, T.; Roberto Zanetti, C.; Bardini Silveira, D.; Regina Monte Barardi, C.; De Fatima Albino Smânia, E.; Viancelli, A.; et al. Antibacterial activity of chalcones, hydrazones and oxadiazoles against methicillin-resistant Staphylococcus aureus. Bioorg. Med. Chem. Lett. 2012, 22, 225–230. [Google Scholar] [CrossRef] [PubMed]
- Nowakowska, Z. A review of anti-infective and anti-inflammatory chalcones. Eur. J. Med. Chem. 2007, 42, 125–137. [Google Scholar] [CrossRef] [PubMed]
- Singh, P.; Anand, A.; Kumar, V. Recent developments in biological activities of chalcones: A mini review. Eur. J. Med. Chem. 2014, 85, 758–777. [Google Scholar] [CrossRef] [PubMed]
- Panche, A.N.; Diwan, A.D.; Chandra, S.R. Flavonoids: An overview. J. Nutr. Sci. 2016, 5, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Sukumaran, S.; Chee, C.; Viswanathan, G.; Buckle, M.; Othman, R.; Rahman, N.A.; Chung, L. Synthesis, Biological Evaluation and Molecular Modelling of 2′-Hydroxychalcones as Acetylcholinesterase Inhibitors. Molecules 2016, 21, 955. [Google Scholar] [CrossRef]
- Barreca, D.; Bellocco, E.; Laganà, G.; Ginestra, G.; Bisignano, C. Biochemical and antimicrobial activity of phloretin and its glycosilated derivatives present in apple and kumquat. Food Chem. 2014, 160, 292–297. [Google Scholar] [CrossRef]
- Tomas-Barberan, F.A.; Clifford, M.N. Flavanones, chalcones and dihydrochalcones—Nature, occurrence and dietary burden. J. Sci. Food Agric. 2000, 80, 1073–1080. [Google Scholar] [CrossRef]
- Mortensen, A. Sweeteners permitted in the European Union: Safety aspects. Scand. J. Food Nutr. 2006, 50, 104–116. [Google Scholar] [CrossRef]
- Wang, R.; Huang, L.; Du, Z.; Feng, H. RhCl(CO)(PPh3)2 catalyzed α-alkylation of ketones with alcohols. J. Organomet. Chem. 2017, 846, 40–43. [Google Scholar] [CrossRef]
- Martinez, R.; Ramon, D.J.; Yus, M. Easy α-alkylation of ketones with alcohols through a hydrogen autotransfer process catalyzed by RuCl2(DMSO)4. Tetrahedron 2006, 62, 8988–9001. [Google Scholar] [CrossRef]
- Sheldon, R.A.; Woodley, J.M. Role of Biocatalysis in Sustainable Chemistry. Chem. Rev. 2018, 118. [Google Scholar] [CrossRef] [PubMed]
- Cao, H.; Chen, X.; Reza Jassbi, A.; Xiao, J. Microbial biotransformation of bioactive flavonoids. Biotechnol. Adv. 2015, 33, 214–223. [Google Scholar] [CrossRef]
- Winkler, C.K.; Clay, D.; Van Heerden, E.; Faber, K. Overcoming Co-Product Inhibition in the Nicotinamide Independent Asymmetric Bioreduction of Activated C=C-Bonds Using Flavin-Dependent Ene-Reductases. Biotechnol. Bioeng. 2013, 110, 3085–3092. [Google Scholar] [CrossRef] [PubMed]
- Williams, R.E.; Bruce, N.C. “New uses for an Old Enzyme” The Old Yellow Enzyme family of flavoenzymes. Microbiology 2002, 148, 1607–1614. [Google Scholar] [CrossRef]
- Filippucci, S.; Tasselli, G.; Labbani, F.K.; Turchetti, B.; Cramarossa, M.R.; Buzzini, P.; Forti, L. Non-Conventional Yeasts as Sources of Ene-Reductases for the Bioreduction of Chalcones. Fermentation 2020, 6, 29. [Google Scholar] [CrossRef]
- Scholtissek, A.; Tischler, D.; Westphal, A.; van Berkel, W.; Paul, C.; van Berkel, W. Old Yellow Enzyme-Catalysed Asymmetric Hydrogenation: Linking Family Roots with Improved Catalysis. Catalysts 2017, 7, 130. [Google Scholar] [CrossRef]
- Janeczko, T.; Gładkowski, W.; Kostrzewa-Susłow, E. Microbial transformations of chalcones to produce food sweetener derivatives. J. Mol. Catal. B Enzym. 2013, 98, 55–61. [Google Scholar] [CrossRef]
- Żyszka-Haberecht, B.; Poliwoda, A.; Lipok, J. Biocatalytic hydrogenation of the C=C bond in the enone unit of hydroxylated chalcones—Process arising from cyanobacterial adaptations. Appl. Microbiol. Biotechnol. 2018, 102, 7097–7111. [Google Scholar] [CrossRef]
- Kostrzewa-Susłow, E.; Dymarska, M.; Guzik, U.; Wojcieszyńska, D.; Janeczko, T. Stenotrophomonas maltophilia: A gram-negative bacterium useful for transformations of flavanone and chalcone. Molecules 2017, 22, 1830. [Google Scholar] [CrossRef]
- Li, Z.; Van Beilen, J.B.; Duetz, W.A.; Schmid, A.; De Raadt, A.; Griengl, H.; Witholt, B. Oxidative biotransformations using oxygenases. Curr. Opin. Chem. Biol. 2002, 6, 136–144. [Google Scholar] [CrossRef]
- Kostrzewa-Susłow, E.; Dmochowska-Gładysz, J.; Białońska, A.; Ciunik, Z. Microbial transformations of flavanone by Aspergillus niger and Penicillium chermesinum cultures. J. Mol. Catal. B Enzym. 2008, 52, 34–39. [Google Scholar] [CrossRef]
- Roh, C.; Seo, S.H.; Choi, K.Y.; Cha, M.; Pandey, B.P.; Kim, J.H.; Park, J.S.; Kim, D.H.; Chang, I.S.; Kim, B.G. Regioselective hydroxylation of isoflavones by Streptomyces avermitilis MA-4680. J. Biosci. Bioeng. 2009, 108, 41–46. [Google Scholar] [CrossRef] [PubMed]
- Schlangen, K.; Miosic, S.; Topuz, F.; Muster, G.; Marosits, T.; Seitz, C.; Halbwirth, H. Chalcone 3-hydroxylation is not a general property of flavonoid 3’-hydroxylase. Plant Sci. 2009, 177, 97–102. [Google Scholar] [CrossRef]
- Kozłowska, J.; Potaniec, B.; Żarowska, B.; Anioł, M. Microbial transformations of 4′-methylchalcones as an efficient method of obtaining novel alcohol and dihydrochalcone derivatives with antimicrobial activity. RSC Adv. 2018, 8, 30379–30386. [Google Scholar] [CrossRef]
- Amir, M.; Kumar, H.; Khan, S.A. Synthesis and pharmacological evaluation of pyrazoline derivatives as new anti-inflammatory and analgesic agents. Bioorg. Med. Chem. Lett. 2008, 18, 918–922. [Google Scholar] [CrossRef]
- Ohta, H.; Konishi, J.; Tsuchihashi, G. Selective Hydrogenation of Carbon-Carbon Double bonds of Chalcones by Corynebacterium equi IFO 3730. Agric. Biol. Chem. 1985, 49, 665–669. [Google Scholar] [CrossRef]
- Shang, X.; Zhou, X.; Zhang, W.; Wan, C.; Chen, J. Tosylhydrazine mediated conjugate reduction and sequential reductive coupling cyclization: Synthesis of 2-arylchromans. Tetrahedron 2015, 71, 8187–8193. [Google Scholar] [CrossRef]
- Van Der Geize, R.; Hessels, G.I.; Nienhuis-Kuiper, M.; Dijkhuizen, L. Characterization of a Second Rhodococcus erythropolis SQ1 3-Ketosteroid 9α-Hydroxylase Activity Comprising a Terminal Oxygenase Homologue, KshA2, Active with Oxygenase-Reductase Component KshB. Appl. Environ. Microbiol. 2008, 74, 7197–7203. [Google Scholar] [CrossRef]
- Petrusma, M.; Dijkhuizen, L.; Van Der Geize, R. Rhodococcus rhodochrous DSM 43269 3-Ketosteroid 9α-Hydroxylase, a Two-Component Iron-Sulfur-Containing Monooxygenase with Subtle Steroid Substrate Specificity. Appl. Environ. Microbiol. 2009, 75, 5300–5307. [Google Scholar] [CrossRef]
- Koppolu, S.R.; Balamurugan, R. In situ formed acetals facilitated direct Michael addition of unactivated ketones. New J. Chem. 2017, 41, 1186–1192. [Google Scholar] [CrossRef]
- Yanagisawa, A.; Takahashi, H.; Arai, T. One-pot synthesis of 1,5-diketones catalyzed by barium isopropoxide. Tetrahedron 2007, 63, 8581–8585. [Google Scholar] [CrossRef]
- Das, G.C.; Hursthouse, M.B.; Malik, K.M.A.; Rahman, M.M.; Rahman, M.T.; Olsson, T. Preparation, spectral studies and X-ray crystal structure of 1,3,5-triphenyl-1,5-pentanedione, C23H2002. J. Chem. Crystallogr. 1994, 24, 511–515. [Google Scholar] [CrossRef]
- Paul, N.; Shanmugam, M.J.; Muthusubramanian, S. Facile Microwave-Assisted Michael Addition of Diphenacyl Sulfides to Chalcones Under Solvent-Free Conditions: Generation of Symmetrical and Unsymmetrical 1,5-Diketones. Synth. Commun. 2013, 43, 129–138. [Google Scholar] [CrossRef]
- Busch, H.; Hagedoorn, P.; Hanefeld, U. Rhodococcus as a Versatile Biocatalyst in Organic Synthesis. Int. J. Mol. Sci. 2019, 20, 4787. [Google Scholar] [CrossRef] [PubMed]
- O’Reilly, E.; Corbett, M.; Hussain, S.; Kelly, P.P.; Richardson, D.; Flitsch, S.L.; Turner, N.J. Substrate promiscuity of cytochrome P450 RhF. Catal. Sci. Technol. 2013, 3, 1490–1492. [Google Scholar] [CrossRef]
- Tiwari, U.; Ameta, C.; Rawal, M.K.; Ameta, R.; Punjabi, P.B. MW assisted synthesis of some pyrazoles containing benzotriazole moiety: An environmentally benign approach. Indian J. Chem. Sect. B Org. Med. Chem. 2013, 52B, 432–439. [Google Scholar] [CrossRef]
- Bai, X.; Shi, W.Q.; Chen, H.F.; Zhang, P.; Li, Y.; Yin, S.F. Synthesis and antitumor activity of 1-acetyl-3-(4-phenyl)-4,5-dihydro-2-pyrazoline-5-phenylursolate and 4-chalcone ursolate derivatives. Chem. Nat. Compd. 2012, 48, 60–65. [Google Scholar] [CrossRef]




© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kozłowska, J.; Potaniec, B.; Anioł, M. Biotransformation of Hydroxychalcones as a Method of Obtaining Novel and Unpredictable Products Using Whole Cells of Bacteria. Catalysts 2020, 10, 1167. https://doi.org/10.3390/catal10101167
Kozłowska J, Potaniec B, Anioł M. Biotransformation of Hydroxychalcones as a Method of Obtaining Novel and Unpredictable Products Using Whole Cells of Bacteria. Catalysts. 2020; 10(10):1167. https://doi.org/10.3390/catal10101167
Chicago/Turabian StyleKozłowska, Joanna, Bartłomiej Potaniec, and Mirosław Anioł. 2020. "Biotransformation of Hydroxychalcones as a Method of Obtaining Novel and Unpredictable Products Using Whole Cells of Bacteria" Catalysts 10, no. 10: 1167. https://doi.org/10.3390/catal10101167
APA StyleKozłowska, J., Potaniec, B., & Anioł, M. (2020). Biotransformation of Hydroxychalcones as a Method of Obtaining Novel and Unpredictable Products Using Whole Cells of Bacteria. Catalysts, 10(10), 1167. https://doi.org/10.3390/catal10101167