1. Introduction
Huge volumes of data are being created regularly, such as content generated on online social media networks and news portals. To facilitate the processing, selection, and comprehension of the huge data volumes that are available, we propose a methodology that uses intelligent knowledge extraction (KE) using natural language processing (NLP) technologies. In this work, we focus on named entity extraction from the pharmaceutical domain, specifically entities that represent
Pharmaceutical Organizations and
Drugs. According to [
1,
2], this NLP job is known as named entity recognition (NER). Its goal is to find entities of a specific kind inside text corpora. NER occupies a prominent position in many NLP systems as a foundational task in information extraction, question answering, and other processes.
Our interest in this subject stems from a challenge we are currently working on with our LinkedDrugs dataset [
3], where the manufacturers (
Pharmaceutical Organization) and active ingredients (
Drug entities) of the collected drug products can be expressed in varying forms, depending on the data source, country of registration, language, etc. Encouraged by the results of our preliminary research [
4], we wish to build on it. Given the ambiguity in entity naming in our drug products dataset, we aimed to greatly enhance the dataset’s quality, as well as the outcomes of any downstream analytical work by utilizing NER to normalize these name values for the active components and manufacturers.
Recently, NER accuracy has been improving due to advances in neural network architectures, particularly due to bidirectional long short-term memory (LSTM) networks [
5,
6], convolutional networks [
7], and recently, transformer architectures [
8]. Over the years, several language-processing libraries from academia and business have been made accessible to the public [
9]. These libraries are fit with incredibly precise pre-trained models for the extraction of common entity classes, such as
Person,
Date,
Location,
Organization, etc. These models should either be improved or re-trained using relevant datasets for the desired entity types, since a particular business may need to recognize more specific entities in text.
In order to train a model with a high level of accuracy, a significant amount of labeled training data must be obtained. Although several carefully labeled, extremely precise, and general datasets are available online [
10], their use may not be practical for the purpose at hand. It may not be possible to manually classify relevant data, or relevant data may not be available on the Internet.
To solve this issue, we provide a way to automatically generate labeled datasets for unique entity types, as seen in texts from the pharmaceutical industry. In our instance, this tactic is used on texts from the pharmaceutical domain, i.e., in news articles from the domain. The research described in [
11], where the basic findings about named entity recognition and knowledge extraction from pharmaceutical texts were reported, is expanded in this paper.
By labeling Drug entities and analyzing the results, we demonstrate that it may be expanded to tagging additional custom entities in other texts in the pharmaceutical domain. The primary focus is on the automatic application of common language-processing tasks, including tokenization, handling stop words and punctuation, lemmatization, and the potential application of custom, business-specific text-processing functions, such as performing text similarity calculations or tagging multi-token entities by joining consecutive tokens.
Two well-known language-processing libraries, spaCy [
12] and AllenNLP [
13], come with a pre-trained model based on convolutional layers with residual connections and a pre-trained model based on ELMo embeddings [
14], respectively. We used them as the baseline to assess the overall applicability and accuracy of the proposed methodology. The custom-trained models exhibit high tagging accuracy in tagging the custom entity
Pharmaceutical Organization when compared to the initial pre-trained models’ accuracy while tagging the more generic
Organization entity over the same testing dataset. Likewise, a model trained on fine-tuned BERT is used for obtaining a more in-depth insight into the results. Finally, a fine-tuned BioBERT [
15], a model based on the BERT architecture and pre-trained on biomedical text corpora, was used to further benchmark the proposed methodology.
The contributions of the work presented in this paper are:
The extension of our prior work [
3,
4] based on the existing BioBERT model to be able to extract two new entity types, namely
Drug and
Pharmaceutical Organization, and proposing a technique for automatically building the training set, which is beneficial to multiple downstream tasks.
We show how to create the labeled dataset, if we know many class representatives that we want to learn, which can be considered as a semi-supervised method.
We show that we can optimize the performance of the whole learning process, especially in low-resource domains. This includes reducing the effort of time-consuming tasks such as manual labeling owing to the visualization tool. This time is multiple orders of magnitude lower than the performance of any of the current models.
The remainder of the paper is structured as follows. In
Section 2, we review relevant works in the NER domain. Then, in
Section 3, we describe the proposed methodology, and afterwards, in
Section 4, we illustrate how it is applied and fine-tuned in the pharmaceutical domain.
Section 5 explains how the knowledge graph can be generated and enriched. Finally,
Section 6 and
Section 7 discuss the main contributions of the paper and the limitations of our study and conclude the work, while providing ideas for future work.
2. Related Work
By adding hierarchical identification, named entity recognition (NER), a crucial part of NLP systems for tagging entities with their appropriate classes, enhances the semantic context of the words. There is much new research being performed in this area right now, particularly in the area of neural network label sequencing optimization, which outperforms earlier NER systems based on domain dictionaries, lexicons, orthographic feature extraction, and semantic rules. Neural network NER systems with minimum feature engineering have gained popularity since [
16], because of the results they provide. They do this by proposing unified n-dimensional word representations and convolutional-neural-network (CNN)-based neural-sequence-labeling models.
Character-level models treat text as distributions over characters, and they can generate embeddings for any string of characters within any textual context. With this, they improve the model’s generalization on both frequent and unseen words, making them popular in the biomedical domain. A model based on stacked bidirectional long short-term memory (LSTM) was introduced in [
17]. This model inputs characters and outputs tag probabilities for each character, achieving state-of-the-art NER performance in seven languages without using additional lexicons and hand-engineered features. In [
18], the authors presented a language model composed of a CNN and LSTM, where they used characters as the input to form a word representation for each token in the sentence; thus, it outperformed word/morpheme-level LSTM baselines.
The authors of [
19] proposed a biomedical named entity recognition (Bio-NER) method that is based on a deep neural network architecture, which utilizes word representations pre-trained on unlabeled data collected from the PubMed database with a skip-gram language model. In [
20], the authors developed a general model based on the long short-term memory network-conditional random field (LSTM-CRF), which outperforms cutting-edge entity-specific NER technologies. Word embedding techniques were used to capture the semantics of the terms in the phrase.
T5 [
21] and XLNet [
22] are state-of-the-art natural language processing (NLP) models that have been developed by Google. Text-to-text transfer transformer (T5) is based on the transformer architecture and is a general-purpose model that can be fine-tuned for various NLP tasks. XLNet, on the other hand, is a generalized autoregressive pretraining method that uses permutation language modeling to learn bidirectional representations from unlabeled text data. One key difference between the two models is that T5 is trained on a single task or direction of text, while XLNet can be trained on multiple tasks and can handle text in either direction. In terms of performance, both models have achieved state-of-the-art results on a range of benchmarks, but XLNet has shown particularly strong performance on natural language understanding tasks.
Since 2018, sequence-to-sequence (Seq2Seq) architectures that work with text have become a popular topic in NLP, due to their powerful ability to transform a given sequence of elements into another sequence. This concept fits well in machine translation. Transformers are models that implement the Seq2Seq architecture by using an encoder–decoder structure.
The launch of Google’s BERT [
8], which is built on a transformer architecture and incorporates an attention mechanism, is one of the most-recent achievements in this development. Due to its capacity to recognize contextual relationships between words (or sub-words) in a text, it excels in various NLP tasks, including NER, and is thus useful in the biomedical and pharmaceutical industries. For the recognition of biomedical named entities for content in Spanish, Hakala and Pyysalo [
23] proposed a method based on conditional random fields (CRFs) and multilingual BERT. The authors investigate feature-based and fine-tuning training methods for the BERT model for NER in Portuguese in [
24]. A method for question answering in the biomedical sector was presented by Lamurias and Couto in their paper [
25]. It was based on a transformer architecture.
A domain-specific language representation called BioBERT [
15] was pre-trained on sizable biomedical corpora. Using the BERT architecture, it was pre-trained on large general domain datasets (English books, Wikipedia, etc.) and biomedical domain corpora (PubMed abstracts, PMC full-text articles). This language model offers better outcomes for NER and other biological text-mining applications.
The problem of co-reference resolution [
14] was also discussed in [
26], further stressing the application of it in downstream tasks, as well as the challenges associated with it, particularly with rarer and under-resourced languages. Therefore, the authors proposed a method to overcome this and apply it to process e-health records from the reception at a Lithuanian hospital.
In neural networks, transfer learning as a machine learning technique introduces the idea of re-usability, where a model created for one task may be utilized as the starting point for training a different problem with a much smaller training set. Transfer learning has been one of the most widely used methods for computer vision and NLP applications in recent years, because it consistently outperforms state-of-the-art models while using much less computational power and training data.
Over the past few years, transfer learning has helped the F1-score for co-reference resolution tasks rise, enabling it to attain a gratifying average of 73%. This assignment aims to group textual mentions of the same underlying real-world objects into groups. Different methods employ biLSTM and attention processes to calculate span representations, and then, a softmax mention ranking model [
27] is used to locate co-reference chains. The F1-score significantly improved with the addition of ELMo and coarse-to-fine and second-order inference, obtaining the aforementioned average of 73%. This task was evaluated with the OntoNotes co-reference annotations from the CONLL2012 shared task [
28], which involved predicting co-references in English, Chinese, and Arabic, using the final version (5.0) of the OntoNotes corpus. It provides an accurate and integrated annotation of multiple levels of the shallow semantic structure in text in multiple languages.
On the other hand, using transfer learning for semantic role labeling demonstrates that using a straightforward BERT-based model can produce state-of-the-art results compared to earlier neural models that included lexical and syntactic features such as parts-of-speech tags and dependency trees [
29]. The reason is that, out of the four tasks that make up semantic role labeling, predicate detection, predicate sense disambiguation, argument identification, and argument classification, the predicate disambiguation task, which can be formulated as a sequence labeling task and is where BERT really excels, is focused on determining the correct meaning of a predicate in a given context.
There are multiple ways to construct an RDF-based knowledge graph (KG), which generally depend on the source data. In our case, we worked with extracted and labeled data to utilize existing solutions that recognize and match the entities in our data with their corresponding version in other publicly available KGs. One such tool is DBpedia Spotlight, an open-source solution for automatic annotation of DBpedia entities in natural language text [
30]. It provides phrase spotting and disambiguation, i.e., entity linking, for the provided input. Its disambiguation algorithm is based on cosine similarities and a modification of the TF-IDF weights. The main phrase spotting algorithm is exact string matching, which uses LingPipe’s (
http://alias-i.com/lingpipe, accessed on 1 November 2022) Aho–Corasick implementation.
Many systems, such as AllenNLP [
13] and Spacy [
12], attempt to provide demo sites for NLP model testing, as well as code snippets for machine learning experts to use more conveniently. On the other hand, libraries such as Hugging Face Transformers [
31] and Deep Pavlov AI [
32], considerably speed up prototyping and make it easier to develop new solutions based on the NLP models that already exist.
In the past year, we have seen several models developed that try to solve a similar problem to the one we target. Ruijie et al. [
33] developed an entity-recognition model, which they used on abstracts of biomedical scientific papers. However, even though their model showed very good F1-score and accuracy values on the two datasets they used, their approach is focused on general pharmaceutical entities: genes, diseases, metabolic processes, etc., in contrast to our focus on pharmaceutical organizations and drugs. Colombo and Oliveira [
34] developed a system that extracts information from pharmaceutical package inserts, to help health professionals guide their patients. Their approach targets the extraction of drugs, diseases, and people, and even though the overall F1 value was comparable to ours, their F1 value for detecting drugs was only 59.14%.
Currently, the main challenge is not only building an architecture of a model, but also obtaining a labeled training dataset. Therefore, the novelty of our work is that we provide the methodology and source code to crawl a large dataset of drugs and diseases, which can be later used and fine-tuned to obtain even larger labeled datasets. To the best of our knowledge, there is no full solution for knowledge extraction in the pharmaceutical domain that is focused on the needs of professionals and allows for the visualization of the outcomes in a manner that they can comprehend. We provide a solution in the form of a platform that aims to close this gap.
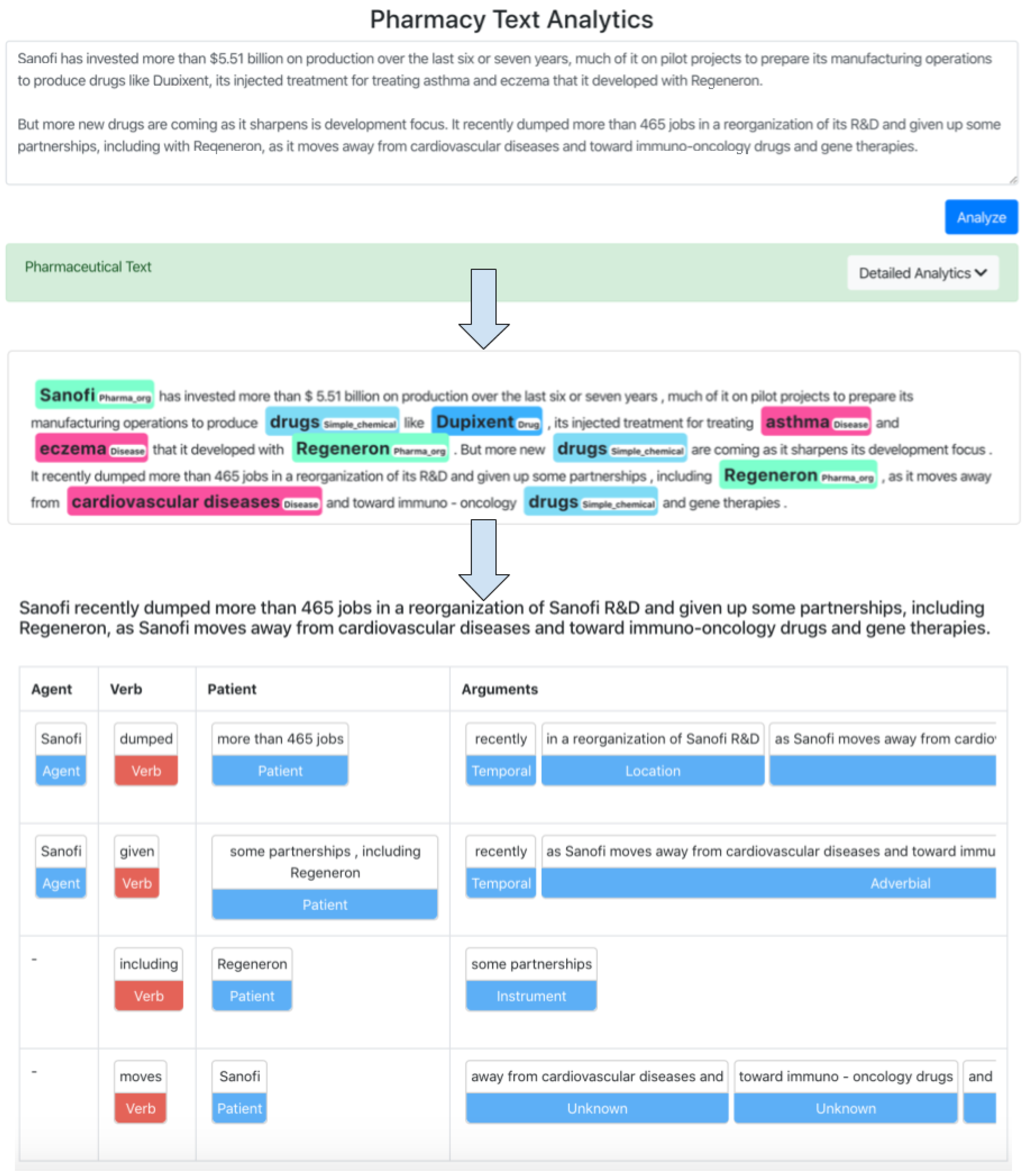
The next sections provide a full overview of the labeled-dataset-generation process, followed by an assessment of the custom model training. The extracted entities can aid in the documents and news filtering, but this is insufficient in the age of “data overload”. Consequently, we take things a step further and include these findings in a platform that later extracts and presents the knowledge associated with these entities. Currently, this platform integrates state-of-the-art NLP models for co-reference resolution [
14] and semantic role labeling (SRL) [
29] to extract the context in which the entities of interest appear. This platform additionally offers convenient visualization of the obtained findings, which brings the relevant concepts closer to the people who use the platform.
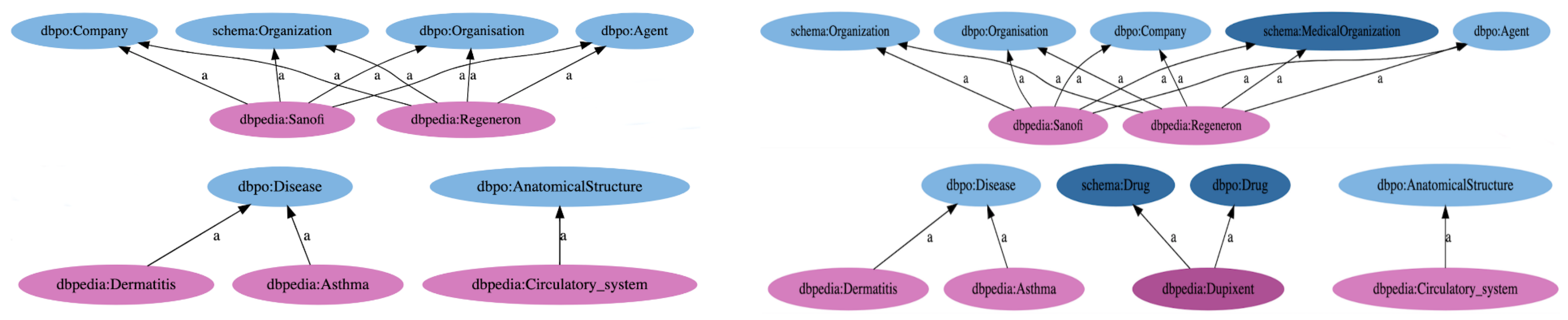
The Resource Description Framework (RDF) [
35] is then used to create a knowledge graph (KG), which is a graph-oriented knowledge representation of the entities and their relations. This provides two main advantages: the RDF graph data model enables the platform to seamlessly integrate the results of multiple knowledge extraction processes from a variety of news sources and, at the same time, links the extracted entities to their counterparts in DBpedia [
36] and the rest of the linked data on the web [
37]. This gives platform users uniform access to all knowledge collected from the platform and relevant connected knowledge already existing in knowledge graphs that are open to the general public.
4. Entity Recognition for Pharmaceutical Organizations and Drugs
Our methodology starts with a text corpora from the pharmaceutical domain and a closed collection of things that are members of a particular class. In our case, we utilized entities that stand for
Pharmaceutical Organizations and
Drugs. We demonstrate that we can train models that can extract even unseen entities from the class of interest using just these two preconditions.
Figure 2 visualizes the whole process.
The text corpora from the pharmaceutical domain that may contain the items from the class of interest are where we begin. The news in this text corpus was gathered from the following websites that are relevant to pharmacies:
FiercePharma (
https://www.fiercepharma.com/, accessed on 1 November 2022),
Pharmacist (
https://www.pharmacist.com/, accessed on 1 November 2022), and
Pharmaceutical Journal (
https://www.pharmaceutical-journal.com/, accessed on 1 November 2022). Next, we tokenized the text such that we extracted the words and, then, attempted to annotate each word in relation to the collection of entities from the needed type. We used the cosine similarity and Levenshtein distance to determine whether the word is comparable to some of the entities [
43]. Each token in the text is given a start position and an end position throughout the annotation process. After finishing this stage, we will have created a labeled dataset, denoted as
MD.
4.1. Creating a Labeled Dataset
One of the main challenges is that the Pharmaceutical Organization entity type can be found in a given text as multi-word phrases, such as “Sanofi Pharmaceuticals Ltd. Spain”, or as a single word: “Sanofi”. Additionally, the name of the Pharmaceutical Organization can contain pharmacy-related keywords, such as “Pharmaceuticals”, “Pharma”, “Medical”, “Biotech”, etc., which are not part of the core name of the organization and can either be found along with it in the sentence or not at all. This means that we should not classify the countries, legal entities, and pharmacy-related words as parts of the Pharmaceutical Organization type. Therefore, the annotation process sequentially performs use-case-specific token filtering during the creation of the MD dataset.
A non-entity list, which comprises all tokens that need to be ignored, is used to do this. In our instance, the list includes all nations, as well as business legal forms (such as “Ltd.”, “Inc.”, “GmbH”, “Corp.”, etc.) and terms related to pharmacies. In our scenario, after removing the tokens from the non-entity list, only “Sanofi” will be left, and we can be sure that the core name has been fully extracted. The same lists are then used to identify any neighbor tokens for multi-token names that may be present as parts of the organization name using text similarity metrics once the core name has been matched in the text.
After the application of the custom, use-case-related filtering, the MD dataset consists of the core entities that have high text similarity. Only the entities that have similarity above the customized threshold are labeled as members of the target class. In our experiments, we used a similarity threshold of 0.9. Some Pharmaceutical Organization entities consist of multiple, consecutive tokens, such as “J & J”. We solved this by token concatenation of consecutive relevant tokens, using a custom function applied on the MD.
After applying all custom text-processing functions, the state of the
MD is as shown in
Table 2.
4.2. Model Fine-Tuning
Next, a model that can extract named entities from the specified class was trained using the MD dataset. The training dataset does not need to contain a large number of varied entities since NER models take into account the context in which the entities appear in a phrase. Here, we used small to moderate quantities of labeled data to enhance the general knowledge language model for the more particular job.
In our case, we fine-tuned the spaCy, AllenNLP, BERT, and BioBERT models. However, each of these models requires a different data format. SpaCy requires an array of sentences with respective tagged entities for each sentence and their start and end positions. AllenNLP requires a dataset in BIOUL or BIO notations (
https://natural-language-understanding.fandom.com/wiki/Named_entity_recognition, accessed on 1 November 2022), which differentiate the following token annotations:
Multi-word entity beginning token: (B);
Multi-word entity inside tokens: (I);
Multi-word entity ending token: (L);
Single-token entities: (U);
Non-entity tokens: (O).
Regardless of the number of tokens, the dataset customized for BERT and BioBERT labels the entities with I–PH_ORG, while all other tokens are tagged with O. As a result, we exported the training and test datasets for the fine-tuning procedure in the needed format using various dataset serializers. For the Drug entity type, labeled datasets are produced using the same process. In this instance, we made use of the same text corpora, but they were annotated with a somewhat bigger collection of Drug entities. After finishing the process of fine-tuning, we have named entity recognition models that can extract the entities from a given type.
4.3. Evaluation
Approximately 5000 news items from a collection of pharmacy-related news were used to gauge the accuracy of our suggested method. The
Drug entities set had 20,266 distinct drug brand names, whereas the
Pharmaceutical Organization entities set had 3633 unique values. As a part of our earlier effort [
3,
4], these sets were already extracted and released.
Both entity types were subjected to two separate assessment situations. Without taking into account the distribution of the entities within them, we divided the news articles from the dataset into training and test parts with sizes of 70% and 30%, respectively. To mitigate the risk of bias and randomness, the whole process was repeated 10 times, and the results presented in this paper are the averages from the 10 repetitions. This approach helped us evaluate the refined model’s overall accuracy.
We assessed our approach’s generalizability in the second evaluation scenario. In this case, we divided the training and test parts according to the entities they each include, ensuring that there was no entity overlap between the two. For testing purposes, we extracted the news articles that included 30% of the entities, while the remaining news was used for training. However, with this, more than 30% of the entire news document set was in the testing part. Therefore, the test component was decreased to comprise exactly 30% of the news articles, while the entities in the remaining documents were changed to other entities that did not belong to the entity set used in the testing portion in order to create a 70% to 30% ratio between the training and test portions.
4.3.1. Entity Recognition for Pharmaceutical Organizations
The obtained fine-tuned models for detecting
Pharmaceutical Organization entities using spaCy, AllenNLP, BERT, and BioBERT were evaluated accordingly. The results were compared to the initial models prior to their fine-tuning, where the task was the extraction of
Organization entities. The results are given in
Table 3, indicating that the fine-tuned models are able to achieve significantly higher F1-scores compared to the original models. Furthermore, we can outline that AllenNLP outperforms spaCy in this NER task, a result that can be attributed to the different neural architectures used by both libraries, while the BERT model is able to outperform both. However, the pre-trained BioBERT on biomedical text is able to slightly outperform BERT in every evaluation.
The sentence context in which the entities occur is taken into account by the pre-trained models, but we can assess the enhanced model generalization capabilities by producing a test dataset that only comprises the entities that were not present during the training. In order to do this, we selected a sample of entities at random from the joint dataset of pharmacy-related news in order to establish a split of 70% to 30% between the training and test datasets, where the test dataset comprises entities that were absent from the training dataset.
The SpaCy, AllenNLP, BERT, and BioBERT models were also trained using these datasets, and the results are given in
Table 4. To better visualize the accuracy,
Figure 3 denotes a sentence extracted from pharmacy-related news where the
Pharmaceutical Organization entities are recognized as expected.
4.3.2. Entity Recognition for Drugs
The SpaCy, AllenNLP, BERT, and BioBERT models were also created for recognizing
Drug entities in texts. The evaluation results are given in
Table 5 for the scenario where the same
Drug entity can be present in both the training and the test dataset, while
Table 6 shows the results when the test dataset does not contain any of the entities used in the training phase. Again, the training–test dataset ratio is 70–30%. To better visualize the accuracy,
Figure 4 denotes a sentence extracted from pharmacy-related news, where the
Drug entity is recognized as expected.
6. Discussion
The platform outlined in this work places a strong emphasis on a strategy for integrating the top NLP models and applying them to a new domain. We employed a modular strategy, wherein each model is a distinct stage in the information extraction pipeline, enabling a simple upgrade with new and potentially better models, ultimately enhancing platform performance.
In contrast to [
12,
13,
31,
32], our platform’s objective is to offer a knowledge extraction solution for the pharmaceutical industry that makes cutting-edge NLP accomplishments more accessible to those who examine enormous volumes of text. Because the PharmKE platform is human-centric, it is primarily intended for users who need to extract knowledge. Users can better grasp the procedure for capturing and connecting this knowledge since each phase’s results are shown. We are also releasing an application programming interface (API) that exposes the outcomes from our platform to other applications because the web browser might not be the most-practical tool for domain experts to use in the process of knowledge extraction, particularly when they analyze texts from various sources. By doing this, we make it possible to create editor plugins that might one day extract and display the knowledge contained in the tools that professionals now work with regularly.
In the most-recent release of the PharmKE platform described in this paper, we improved the named entity recognition module to extract two new entity types in addition to those previously recognized by the better BioBERT model, namely Drug and Pharmaceutical Organization. Using a text corpus from the pharmaceutical domain and a closed set of entity instances from the kinds of interest, we demonstrate a technique for automatically building the training set for the recognition of Pharmaceutical Organization and Drug during the fine-tuning phase. This technology allows for the recognition of entities that are not included in the training set, which is a promising outcome, according to the evaluation of the fine-tuned model.
The goal of our paper was not to optimize the model performances for a specific task, such as the text analysis, but to show that we can optimize the performance of the whole process, which includes time-consuming tasks such as manual labeling, especially in low-resource domains. This time is multiple orders of magnitude lower than the performance of any of the current models. Therefore, in this paper, we focused on how to create the labeled dataset, if we know many class representatives that we want to learn. The speedup that the proposed methodology introduces, in combination with the visualization tool, is quite significant, especially in the data-labeling phase.
The knowledge graph that we constructed and enhanced at the last step of the pipeline aims to demonstrate the potential for packaging and reusing the knowledge produced by the pipeline in other software solutions. Because an RDF knowledge graph is created as the last stage in the platform’s process, even if it is human-centric, the outcomes may be saved, shared, merged with other RDF knowledge graphs, and (re)used programmatically outside of the platform. Due to the nature of RDF and knowledge graphs, it is possible to practically seamlessly combine platform findings with additional RDF data that are available externally or internally in the user environment.
The PharmKE platform is receptive to ongoing developments in the NLP industry. The coupling of the relations acquired by the SRL model with the relevant attributes in the knowledge graph is one of the essential steps in the knowledge extraction process that is not addressed by the present models. In addition to adding any model that will produce better outcomes in some of the present jobs, our team will attempt to address this difficulty in its future study. The platform’s modular construction makes all of this feasible. Cleaning up the knowledge graph from incorrect inferences produced by the pipeline, which is a common and anticipated issue with NLP, would be another obstacle.
One limitation of the study is that we do not redistribute the dataset because we do not have such a license. However, we do provide the source code (see [
38,
39]) so that interested readers can execute it and recreate the dataset by themselves. With this code, all experiments performed after the creation of the dataset can also be reproduced.
7. Conclusions
Using cutting-edge models for text categorization, pharmaceutical domain named entity recognition (NER), co-reference resolution (CRR), semantic role labeling (SRL), and knowledge extraction, we built our modular PharmKE platform [
38,
39]. The platform is primarily intended for human users. Pharmaceutical domain specialists may easily identify the information extracted from the input texts thanks to PharmKE’s visualization of the findings from each of the integrated models.
The PharmKE platform’s modular architecture makes it simple to incorporate additional and potentially improved models, which is one of our strategic goals. One such move in this manner was our addition of the Pharmaceutical Organization and Drug entity type identification to the more modern BioBERT model for NER.
Additionally, the platform is open-source and publicly accessible [
38,
39], providing the reproducibility of our findings. This also implies that, as a result of the platform’s modular architecture, other researchers can alter their own copies of it, use them to run their own instances, and even re-purpose it.
The proposed methodology could be used in mobile and pervasive systems since it enables patients to scan the medication instructions for their prescriptions, which can give them more pertinent and understandable information. The proposed methodology may also be used to check whether a drug from a different vendor is compatible with the patient’s prescription medication. The potential of patient empowerment lies on such methods.
The absence of labeled datasets for testing and training custom models for language comprehension tasks in text is a prevalent problem. We offer a way to automate the process of producing labeled datasets for training models for custom entity tagging in order to address this problem. SpaCy, AllenNLP, BERT, and BioBERT were used to train custom models for named entity identification in order to evaluate the technique. The findings show that the newly trained models perform better at identifying custom entities than the pre-trained models.