Characterization of FOLH1 Expression in Renal Cell Carcinoma
Abstract
Simple Summary
Abstract
1. Introduction
2. Methods
2.1. Study Cohort
2.2. DNA/RNA Next-Generation Sequencing
2.3. RNA Expression Analyses
2.4. Immunohistochemistry
2.5. Tumor Mutational Burden
2.6. Survival Analysis
2.7. Statistical Analysis and Data Visualization
3. Results
3.1. Study Cohort and Patient Characteristics
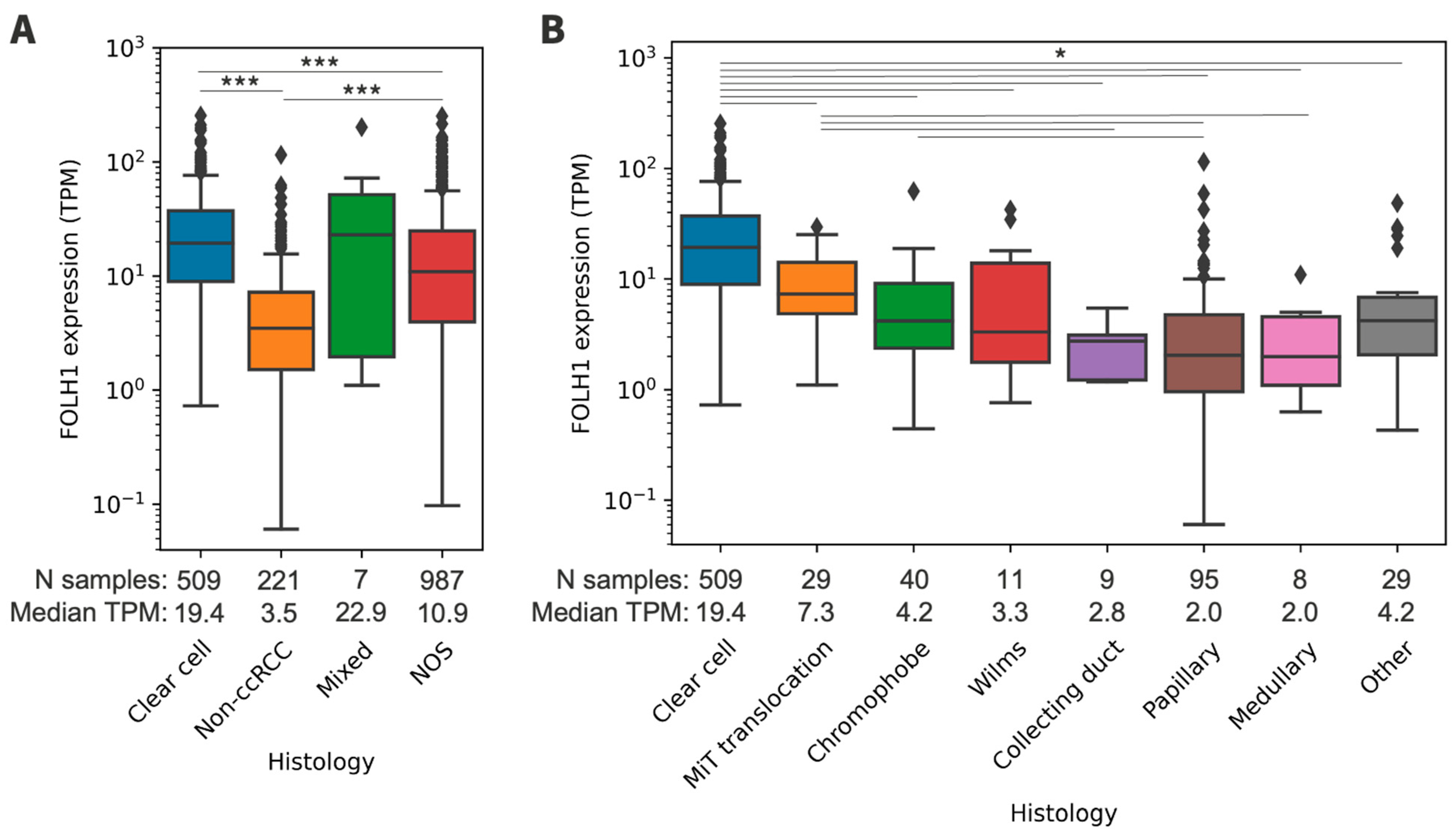
3.2. FOLH1 Expression across Histologic Subtypes
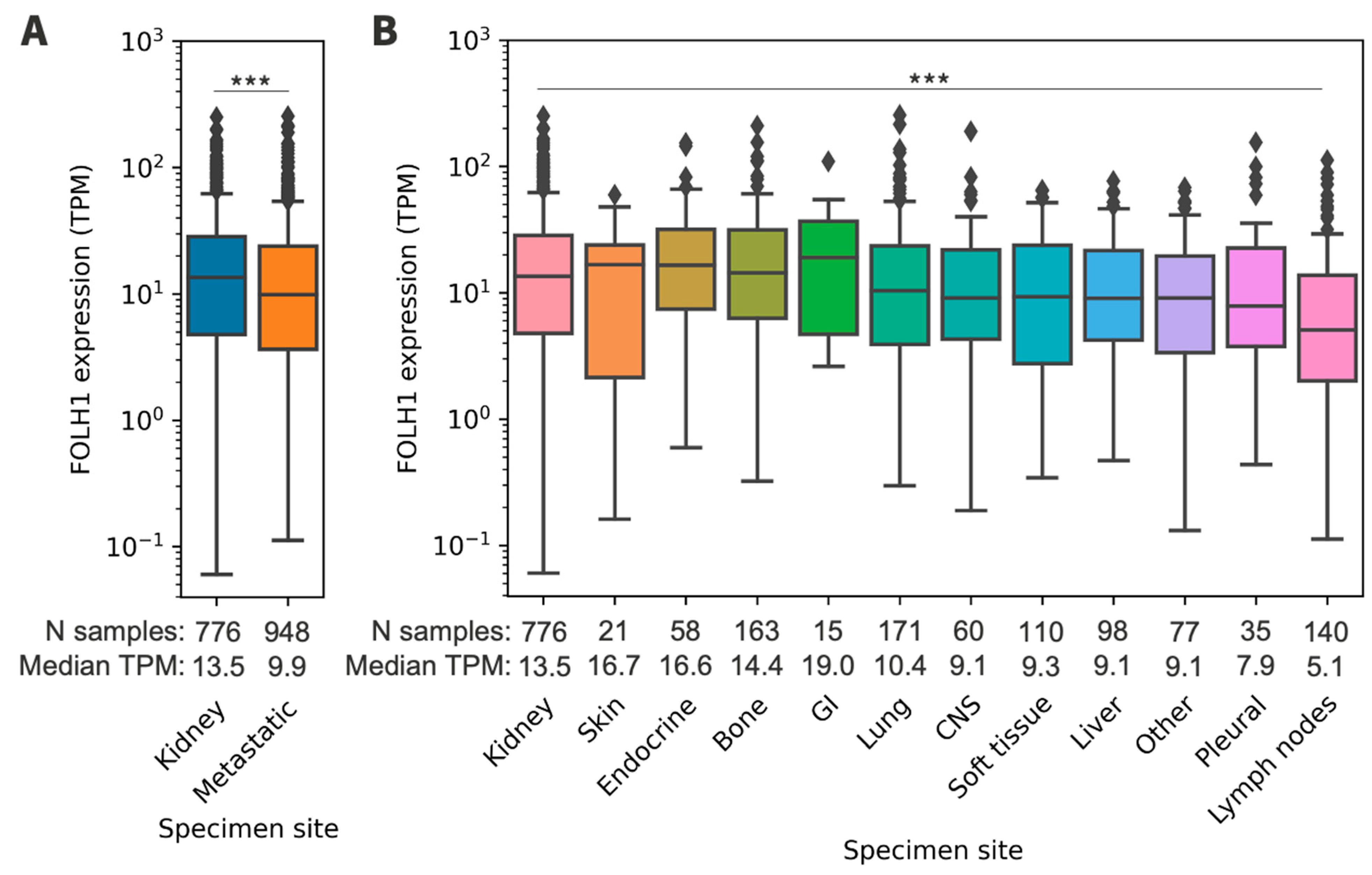
3.3. FOLH1 Expression across Sites of Metastasis
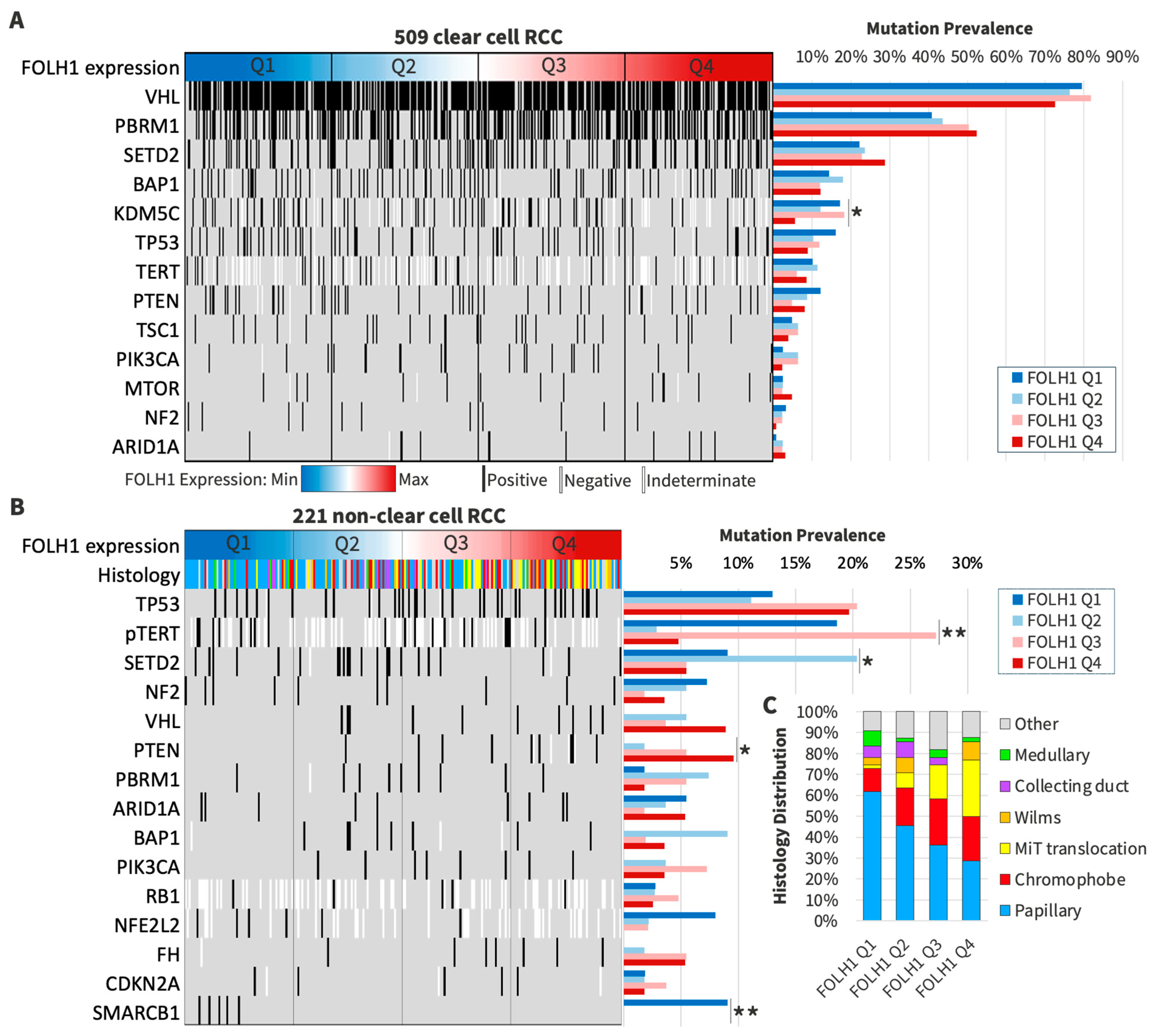
3.4. Patterns of DNA Profiling in FOLH1 Expression Groups
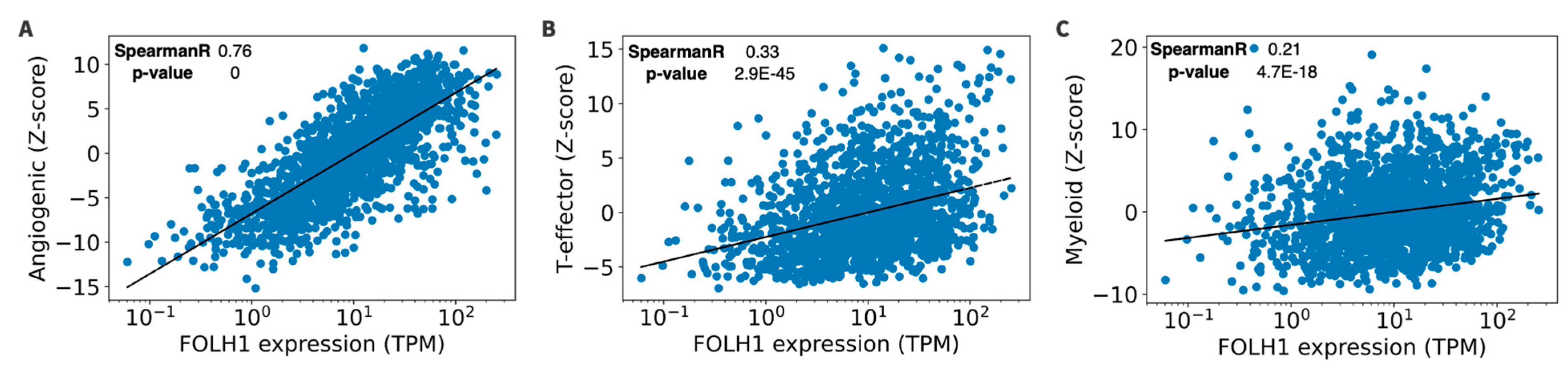
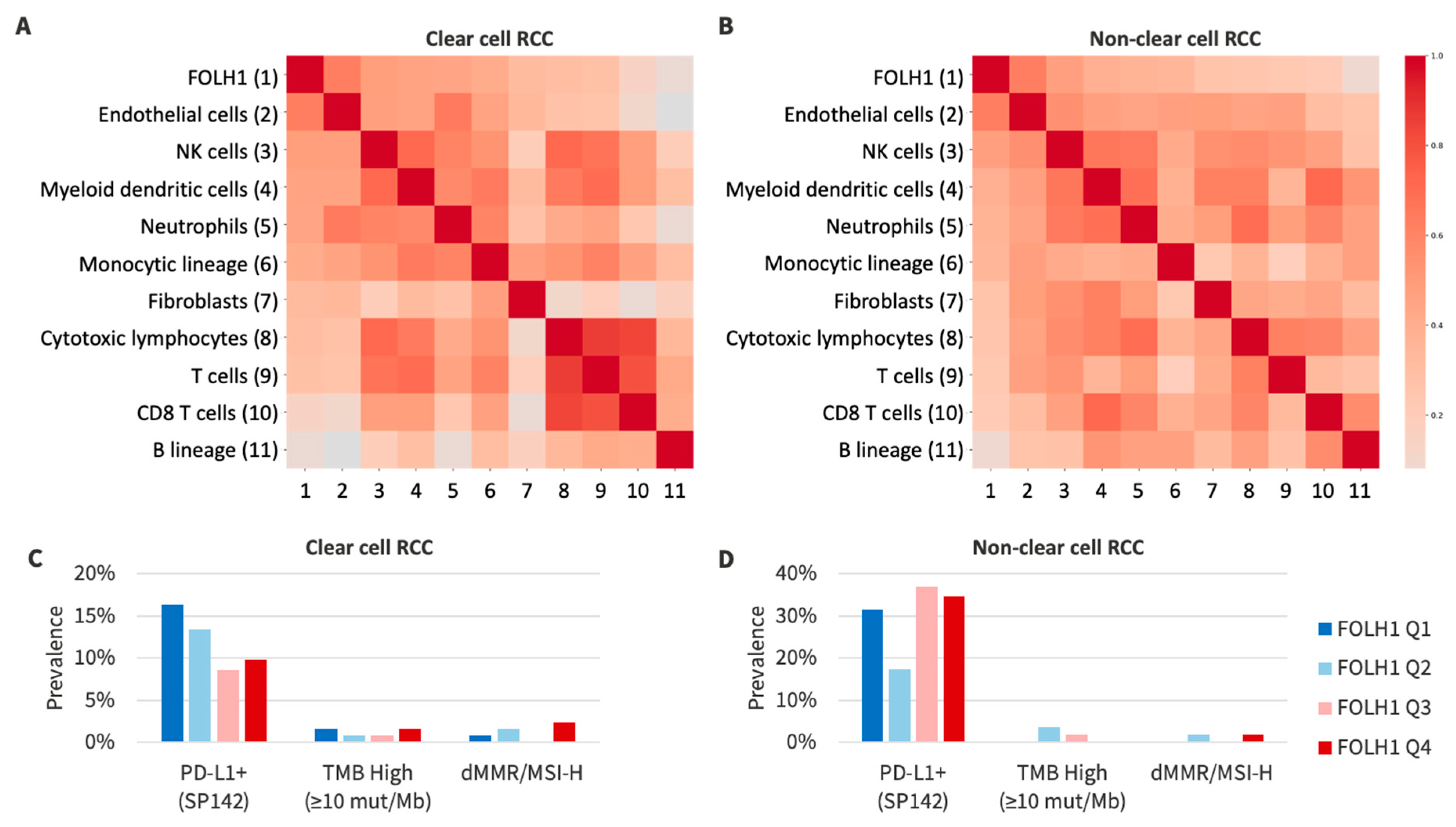
3.5. RNA Signatures and Tumor Microenvironment in FOLH1 Expression Groups
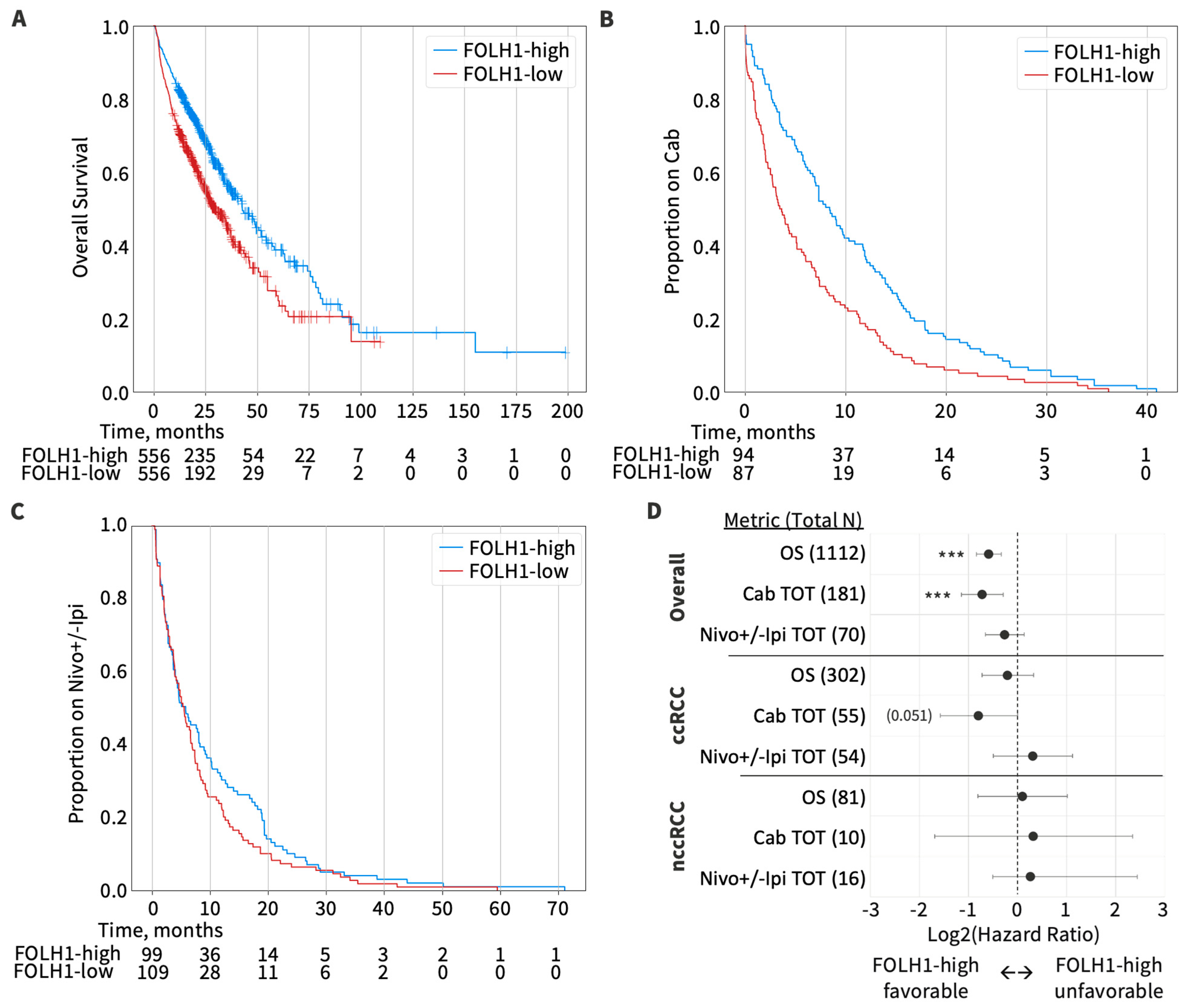
3.6. Overall Survival by FOLH1 Expression
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bahadoram, S.; Davoodi, M.; Hassanzadeh, S.; Bahadoram, M.; Barahman, M.; Mafakher, L. Renal cell carcinoma: An overview of the epidemiology, diagnosis, and treatment. G. Ital. Nefrol. 2022, 39, 2022. [Google Scholar]
- Padala, S.A.; Barsouk, A.; Thandra, K.C.; Saginala, K.; Mohammed, A.; Vakiti, A.; Rawla, P.; Barsouk, A. Epidemiology of Renal Cell Carcinoma. World J. Oncol. 2020, 11, 79–87. [Google Scholar] [CrossRef] [PubMed]
- Graves, A.; Hessamodini, H.; Wong, G.; Lim, W.H. Metastatic renal cell carcinoma: Update on epidemiology, genetics, and therapeutic modalities. Immunotargets Ther. 2013, 2, 73–90. [Google Scholar] [CrossRef] [PubMed]
- Heidenreich, A.; Ravery, V. Preoperative imaging in renal cell cancer. World J. Urol. 2004, 22, 307–315. [Google Scholar] [CrossRef] [PubMed]
- Klatte, T.; Rossi, S.H.; Stewart, G.D. Prognostic factors and prognostic models for renal cell carcinoma: A literature review. World J. Urol. 2018, 36, 1943–1952. [Google Scholar] [CrossRef] [PubMed]
- Kinoshita, Y.; Kuratsukuri, K.; Landas, S.; Imaida, K.; Rovito, P.M., Jr.; Wang, C.Y.; Haas, G.P. Expression of prostate-specific membrane antigen in normal and malignant human tissues. World J. Surg. 2006, 30, 628–636. [Google Scholar] [CrossRef]
- Al-Ahmadie, H.A.; Olgac, S.; Gregor, P.D.; Tickoo, S.K.; Fine, S.W.; Kondagunta, G.V.; Scher, H.I.; Morris, M.J.; Russo, P.; Motzer, R.J.; et al. Expression of prostate-specific membrane antigen in renal cortical tumors. Mod. Pathol. 2008, 21, 727–732. [Google Scholar] [CrossRef] [PubMed]
- Vig, S.V.L.; Zan, E.; Kang, S.K. Imaging for Metastatic Renal Cell Carcinoma. Urol. Clin. N. Am. 2020, 47, 281–291. [Google Scholar] [CrossRef]
- Liu, Y. The Place of FDG PET/CT in Renal Cell Carcinoma: Value and Limitations. Front. Oncol. 2016, 6, 201. [Google Scholar] [CrossRef]
- Cho, S.Y.; Szabo, Z. Molecular imaging of urogenital diseases. Semin. Nucl. Med. 2014, 44, 93–109. [Google Scholar] [CrossRef]
- Ahn, T.; Roberts, M.J.; Abduljabar, A.; Joshi, A.; Perera, M.; Rhee, H.; Wood, S.; Vela, I. A Review of Prostate-Specific Membrane Antigen (PSMA) Positron Emission Tomography (PET) in Renal Cell Carcinoma (RCC). Mol. Imaging Biol. 2019, 21, 799–807. [Google Scholar] [CrossRef] [PubMed]
- Rhee, H.; Blazak, J.; Tham, C.M.; Ng, K.L.; Shepherd, B.; Lawson, M.; Preston, J.; Vela, I.; Thomas, P.; Wood, S. Pilot study: Use of gallium-68 PSMA PET for detection of metastatic lesions in patients with renal tumour. EJNMMI Res. 2016, 6, 76. [Google Scholar] [CrossRef] [PubMed]
- Martin, F.C.; Dorff, T.B.; Tran, B. The new era of prostate-specific membrane antigen-directed immunotherapies and beyond in advanced prostate cancer: A review. Ther. Adv. Med. Oncol. 2023, 15, 17588359231170474. [Google Scholar] [CrossRef]
- Sartor, O.; de Bono, J.; Chi, K.N.; Fizazi, K.; Herrmann, K.; Rahbar, K.; Tagawa, S.T.; Nordquist, L.T.; Vaishampayan, N.; El-Haddad, G.; et al. Lutetium-177-PSMA-617 for Metastatic Castration-Resistant Prostate Cancer. N. Engl. J. Med. 2021, 385, 1091–1103. [Google Scholar] [CrossRef] [PubMed]
- Baum, R.P.; Kulkarni, H.R.; Schuchardt, C.; Singh, A.; Wirtz, M.; Wiessalla, S.; Schottelius, M.; Mueller, D.; Klette, I.; Wester, H.-J. 177Lu-Labeled Prostate-Specific Membrane Antigen Radioligand Therapy of Metastatic Castration-Resistant Prostate Cancer: Safety and Efficacy. J. Nucl. Med. 2016, 57, 1006–1013. [Google Scholar] [CrossRef] [PubMed]
- Urso, L.; Castello, A.; Rocca, G.C.; Lancia, F.; Panareo, S.; Cittanti, C.; Uccelli, L.; Florimonte, L.; Castellani, M.; Ippolito, C.; et al. Role of PSMA-ligands imaging in Renal Cell Carcinoma management: Current status and future perspectives. J. Cancer Res. Clin. Oncol. 2022, 148, 1299–1311. [Google Scholar] [CrossRef] [PubMed]
- McDermott, D.F.; Huseni, M.A.; Atkins, M.B.; Motzer, R.J.; Rini, B.I.; Escudier, B.; Fong, L.; Joseph, R.W.; Pal, S.K.; Reeves, J.A.; et al. Clinical activity and molecular correlates of response to atezolizumab alone or in combination with bevacizumab versus sunitinib in renal cell carcinoma. Nat. Med. 2018, 24, 749–757. [Google Scholar] [CrossRef] [PubMed]
- Becht, E.; Giraldo, N.A.; Lacroix, L.; Buttard, B.; Elarouci, N.; Petitprez, F.; Selves, J.; Laurent-Puig, P.; Sautes-Fridman, C.; Fridman, W.H.; et al. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol. 2016, 17, 218. [Google Scholar] [CrossRef]
- Marabelle, A.; Le, D.T.; Ascierto, P.A.; Di Giacomo, A.M.; De Jesus-Acosta, A.; Delord, J.-P.; Geva, R.; Gottfried, M.; Penel, N.; Hansen, A.R.; et al. Efficacy of Pembrolizumab in Patients With Noncolorectal High Microsatellite Instability/Mismatch Repair-Deficient Cancer: Results From the Phase II KEYNOTE-158 Study. J. Clin. Oncol. 2020, 38, 1–10. [Google Scholar] [CrossRef]
- Lopes, A.D.; Davis, W.L.; Rosenstraus, M.J.; Uveges, A.J.; Gilman, S.C. Immunohistochemical and pharmacokinetic characterization of the site-specific immunoconjugate CYT-356 derived from antiprostate monoclonal antibody 7E11-C5. Cancer Res. 1990, 50, 6423–6429. [Google Scholar]
- Silver, D.A.; Pellicer, I.; Fair, W.R.; Heston, W.D.; Cordon-Cardo, C. Prostate-specific membrane antigen expression in normal and malignant human tissues. Clin. Cancer Res. 1997, 3, 81–85. [Google Scholar] [PubMed]
- Chang, S.S.; Reuter, V.E.; Heston, W.D.; Bander, N.H.; Grauer, L.S.; Gaudin, P.B. Five different anti-prostate-specific membrane antigen (PSMA) antibodies confirm PSMA expression in tumor-associated neovasculature. Cancer Res. 1999, 59, 3192–3198. [Google Scholar] [PubMed]
- Chang, S.S.; Reuter, V.E.; Heston, W.D.; Gaudin, P.B. Metastatic renal cell carcinoma neovasculature expresses prostate-specific membrane antigen. Urology 2001, 57, 801–805. [Google Scholar] [CrossRef] [PubMed]
- de la Taille, A.; Katz, A.; Cao, Y.; McKiernan, J.; Buttyan, R.; Burchardt, M.; Burchardt, T.; Hayek, O.; Olsson, C.A.; Chopin, D.K.; et al. Blood-based RT-PCR assays of MN/CA9 or PSMA: Clinical application in renal cancer patients. Urology 2000, 56, 393–398. [Google Scholar] [CrossRef]
- de la Taille, A.; Cao, Y.; Sawczuk, I.S.; Nozemu, T.; D’Agati, V.; McKiernan, J.M.; Bagiella, E.; Buttyan, R.; Burchardt, M.; Olsson, C.A.; et al. Detection of prostate-specific membrane antigen expressing cells in blood obtained from renal cancer patients: A potential biomarker of vascular invasion. Cancer Detect. Prev. 2000, 24, 579–588. [Google Scholar]
- Baccala, A.; Sercia, L.; Li, J.; Heston, W.; Zhou, M. Expression of prostate-specific membrane antigen in tumor-associated neovasculature of renal neoplasms. Urology 2007, 70, 385–390. [Google Scholar] [CrossRef] [PubMed]
- Spatz, S.; Tolkach, Y.; Jung, K.; Stephan, C.; Busch, J.; Ralla, B.; Rabien, A.; Feldmann, G.; Brossart, P.; Bundschuh, R.A.; et al. Comprehensive Evaluation of Prostate Specific Membrane Antigen Expression in the Vasculature of Renal Tumors: Implications for Imaging Studies and Prognostic Role. J. Urol. 2018, 199, 370–377. [Google Scholar] [CrossRef] [PubMed]
- Rowe, S.P.; Gorin, M.A.; Hammers, H.J.; Pomper, M.G.; Allaf, M.E.; Javadi, M.S. Detection of 18F-FDG PET/CT Occult Lesions With 18F-DCFPyL PET/CT in a Patient With Metastatic Renal Cell Carcinoma. Clin. Nucl. Med. 2016, 41, 83–85. [Google Scholar] [CrossRef] [PubMed]
- Ferro, M.; Crocetto, F.; Barone, B.; del Giudice, F.; Maggi, M.; Lucarelli, G.; Busetto, G.M.; Autorino, R.; Marchioni, M.; Cantiello, F.; et al. Artificial intelligence and radiomics in evaluation of kidney lesions: A com-prehensive literature review. Ther. Adv. Urol. 2023, 15, 17562872231164803. [Google Scholar] [CrossRef]
- Rizzo, A.; Racca, M.; Dall’armellina, S.; Rescigno, P.; Banna, G.L.; Albano, D.; Dondi, F.; Bertagna, F.; Annunziata, S.; Treglia, G. The Emerging Role of PET/CT with PSMA-Targeting Radiopharmaceuticals in Clear Cell Renal Cancer: An Updated Systematic Review. Cancers 2023, 15, 355. [Google Scholar] [CrossRef]
- Udovicich, C.; Callahan, J.; Bressel, M.; Ong, W.L.; Perera, M.; Tran, B.; Azad, A.; Haran, S.; Moon, D.; Chander, S.; et al. Impact of Prostate-specific Membrane Antigen Positron Emission Tomography/Computed Tomography in the Management of Oligometastatic Renal Cell Carcinoma. Eur. Urol. Open Sci. 2022, 44, 60–68. [Google Scholar] [CrossRef]
- Yin, Y.; Campbell, S.P.; Markowski, M.C.; Pierorazio, P.M.; Pomper, M.G.; Allaf, M.E.; Rowe, S.P.; Gorin, M.A. Inconsistent Detection of Sites of Metastatic Non-Clear Cell Renal Cell Carcinoma with PSMA-Targeted [(18)F]DCFPyL PET/CT. Mol. Imaging Biol. 2019, 21, 567–573. [Google Scholar] [CrossRef] [PubMed]
- Crocetto, F.; Falcone, A.; Mirto, B.F.; Sicignano, E.; Pagano, G.; Dinacci, F.; Varriale, D.; Machiella, F.; Giampaglia, G.; Calogero, A.; et al. Unlocking Precision Medicine: Liquid Biopsy Advancements in Renal Cancer Detection and Monitoring. Int. J. Mol. Sci. 2024, 25, 3867. [Google Scholar] [CrossRef]
- Brozovich, A.; Garmezy, B.; Pan, T.; Wang, L.; Farach-Carson, M.C.; Satcher, R.L. All bone metastases are not created equal: Revisiting treatment resistance in renal cell carcinoma. J. Bone Oncol. 2021, 31, 100399. [Google Scholar] [CrossRef]
- Duarte, C.; Hu, J.; Beuselinck, B.; Panian, J.; Weise, N.; Dizman, N.; Collier, K.A.; Rathi, N.; Li, H.; Elias, R.; et al. Metastatic renal cell carcinoma to the pancreas and other sites-a multicenter retrospective study. EClinicalMedicine 2023, 60, 102018. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Q.; Li, P.; Zhou, X.; Qiang, Y.; Fan, J.; Lin, Y.; Chen, Y.; Guo, J.; Wang, F.; Xue, H.; et al. Deficiency of the X-inactivation escaping gene KDM5C in clear cell renal cell carcinoma promotes tumorigenicity by reprogramming glycogen metabolism and inhibiting ferroptosis. Theranostics 2021, 11, 8674–8691. [Google Scholar] [CrossRef] [PubMed]
- Motzer, R.J.; Banchereau, R.; Hamidi, H.; Powles, T.; McDermott, D.; Atkins, M.B.; Escudier, B.; Liu, L.-F.; Leng, N.; Abbas, A.R.; et al. Molecular Subsets in Renal Cancer Determine Outcome to Checkpoint and Angiogenesis Blockade. Cancer Cell 2020, 38, 803–817.e4. [Google Scholar] [CrossRef]
- Santos, M.; Lanillos, J.; Caleiras, E.; Valdivia, C.; Roldan-Romero, J.M.; Laínez, N.; Puente, J.; Beuselinck, B.; Oudard, S.; Zucman-Rossi, J.; et al. PBRM1 and KDM5C cooperate to define high-angiogenesis tumors and increased antiangiogenic response in renal cancer. Am. J. Cancer Res. 2023, 13, 2116–2125. [Google Scholar]
- Sonni, I.; Caron, J.; Kishan, A.U.; Muthusamy, V.R.; Calais, J. PSMA Expression in the Neovasculature Associated With Rectal Adenocarcinoma: A Potential Stromal Target for Nuclear Theranostics. Clin. Nucl. Med. 2020, 45, e309–e310. [Google Scholar] [CrossRef]
- Wernicke, A.G.; Kim, S.; Liu, H.; Bander, N.H.; Pirog, E.C. Prostate-specific Membrane Antigen (PSMA) Expression in the Neovasculature of Gynecologic Malignancies: Implications for PSMA-targeted Therapy. Appl. Immunohistochem. Mol. Morphol. 2017, 25, 271–276. [Google Scholar] [CrossRef]
- Tolkach, Y.; Gevensleben, H.; Bundschuh, R.; Koyun, A.; Huber, D.; Kehrer, C.; Hecking, T.; Keyver-Paik, M.-D.; Kaiser, C.; Ahmadzadehfar, H.; et al. Prostate-specific membrane antigen in breast cancer: A comprehensive evaluation of expression and a case report of radionuclide therapy. Breast Cancer Res. Treat. 2018, 169, 447–455. [Google Scholar] [CrossRef] [PubMed]
- Sepe, P.; Ottini, A.; Pircher, C.C.; Franza, A.; Claps, M.; Guadalupi, V.; Verzoni, E.; Procopio, G. Characteristics and Treatment Challenges of Non-Clear Cell Renal Cell Carcinoma. Cancers 2021, 13, 3807. [Google Scholar] [CrossRef] [PubMed]
- Milowsky, M.I.; Nanus, D.M.; Kostakoglu, L.; Sheehan, C.E.; Vallabhajosula, S.; Goldsmith, S.J.; Ross, J.S.; Bander, N.H. Vascular targeted therapy with anti-prostate-specific membrane antigen monoclonal antibody J591 in advanced solid tumors. J. Clin. Oncol. 2007, 25, 540–547. [Google Scholar] [CrossRef]
- Zhang, J.; Schuchardt, C.; Chen, X.; Baum, R.P. Rapid Tumor Washout of 177Lu-PSMA Radioligand in Renal Cell Carcinoma. Clin. Nucl. Med. 2023, 48, 732–734. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Royz, E.; Pan, E.; Guer, M.; Elliott, A.; Siva, S.; Ravi, P.; McGregor, B.; Bagrodia, A.; Derweesh, I.; Barata, P.; et al. Characterization of FOLH1 Expression in Renal Cell Carcinoma. Cancers 2024, 16, 1855. https://doi.org/10.3390/cancers16101855
Royz E, Pan E, Guer M, Elliott A, Siva S, Ravi P, McGregor B, Bagrodia A, Derweesh I, Barata P, et al. Characterization of FOLH1 Expression in Renal Cell Carcinoma. Cancers. 2024; 16(10):1855. https://doi.org/10.3390/cancers16101855
Chicago/Turabian StyleRoyz, Eric, Elizabeth Pan, Melis Guer, Andrew Elliott, Shankar Siva, Praful Ravi, Bradley McGregor, Aditya Bagrodia, Ithaar Derweesh, Pedro Barata, and et al. 2024. "Characterization of FOLH1 Expression in Renal Cell Carcinoma" Cancers 16, no. 10: 1855. https://doi.org/10.3390/cancers16101855
APA StyleRoyz, E., Pan, E., Guer, M., Elliott, A., Siva, S., Ravi, P., McGregor, B., Bagrodia, A., Derweesh, I., Barata, P., Heath, E. I., Antonarakis, E. S., Darabi, S., Hoon, D. S. B., Mortazavi, A., Choueiri, T. K., Nabhan, C., Wei, S., & McKay, R. R. (2024). Characterization of FOLH1 Expression in Renal Cell Carcinoma. Cancers, 16(10), 1855. https://doi.org/10.3390/cancers16101855







