Identification of Skin Lesions by Snapshot Hyperspectral Imaging
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
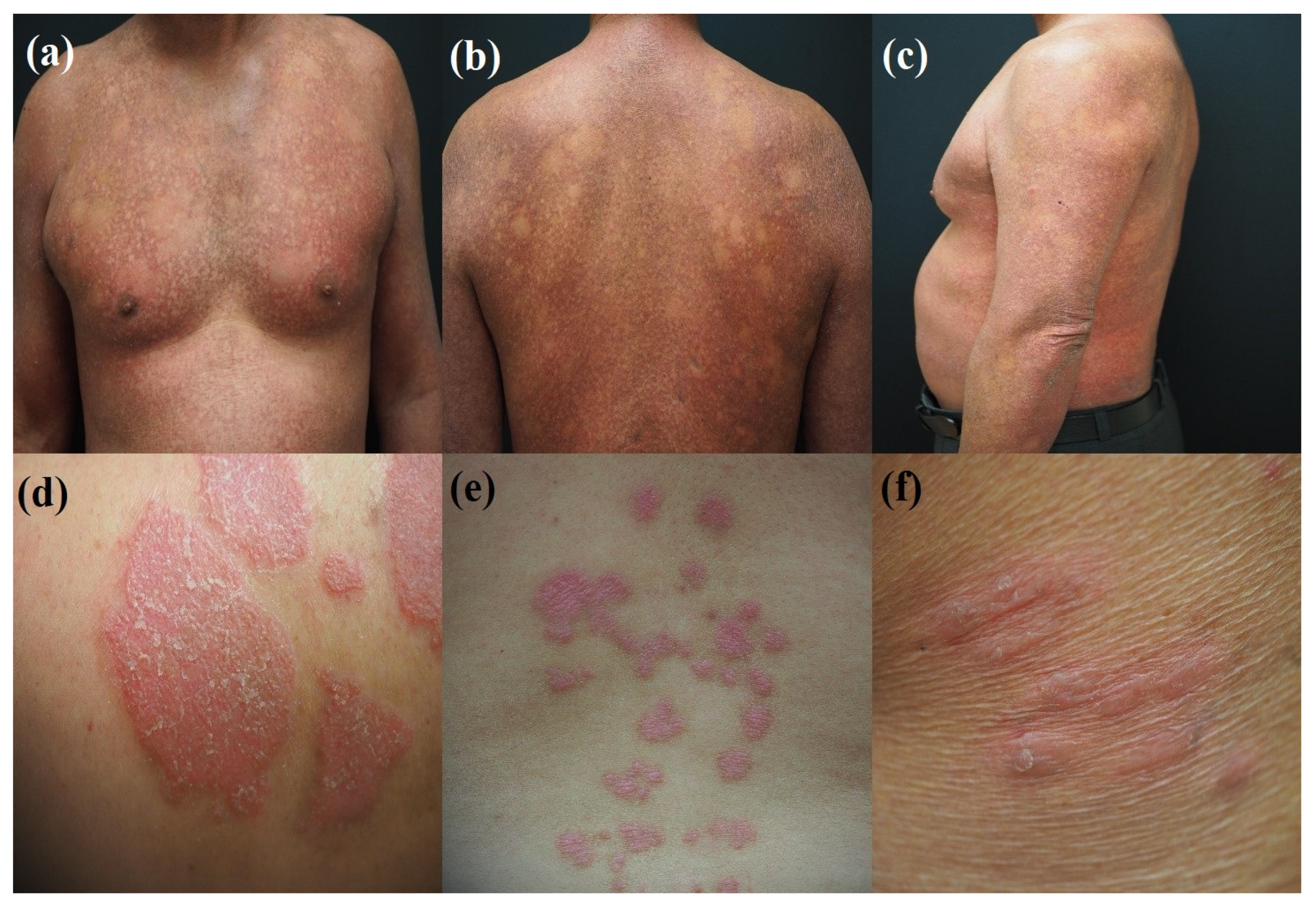
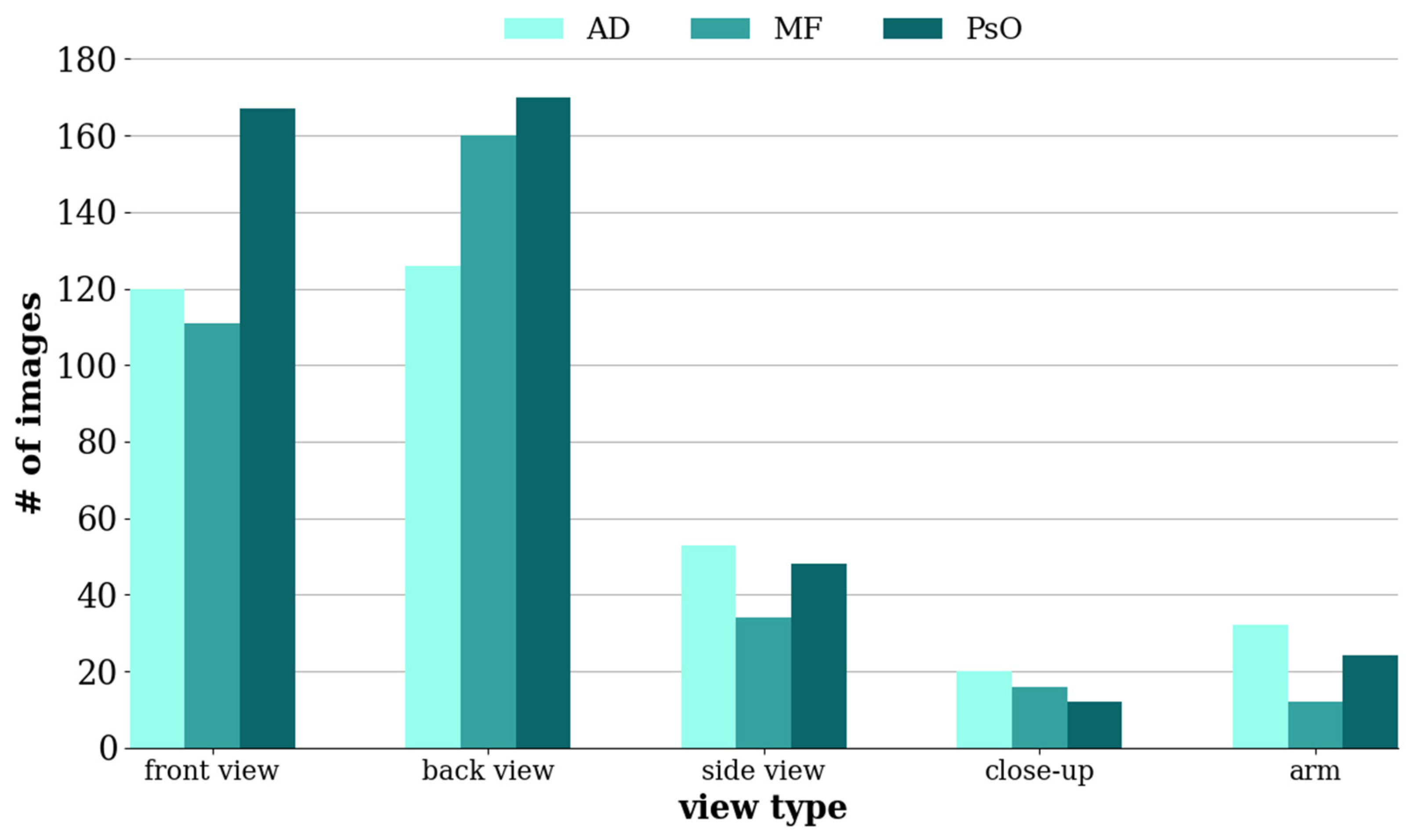
2.1. Sample Preparation
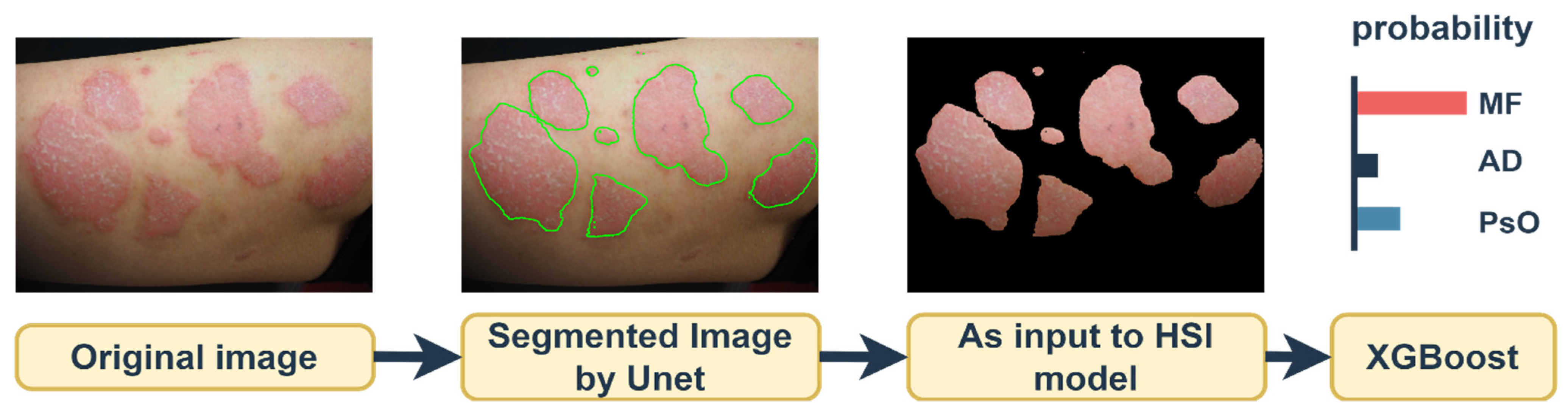
2.2. Segmentation Task
2.3. Model HSI-Feature Extraction Task
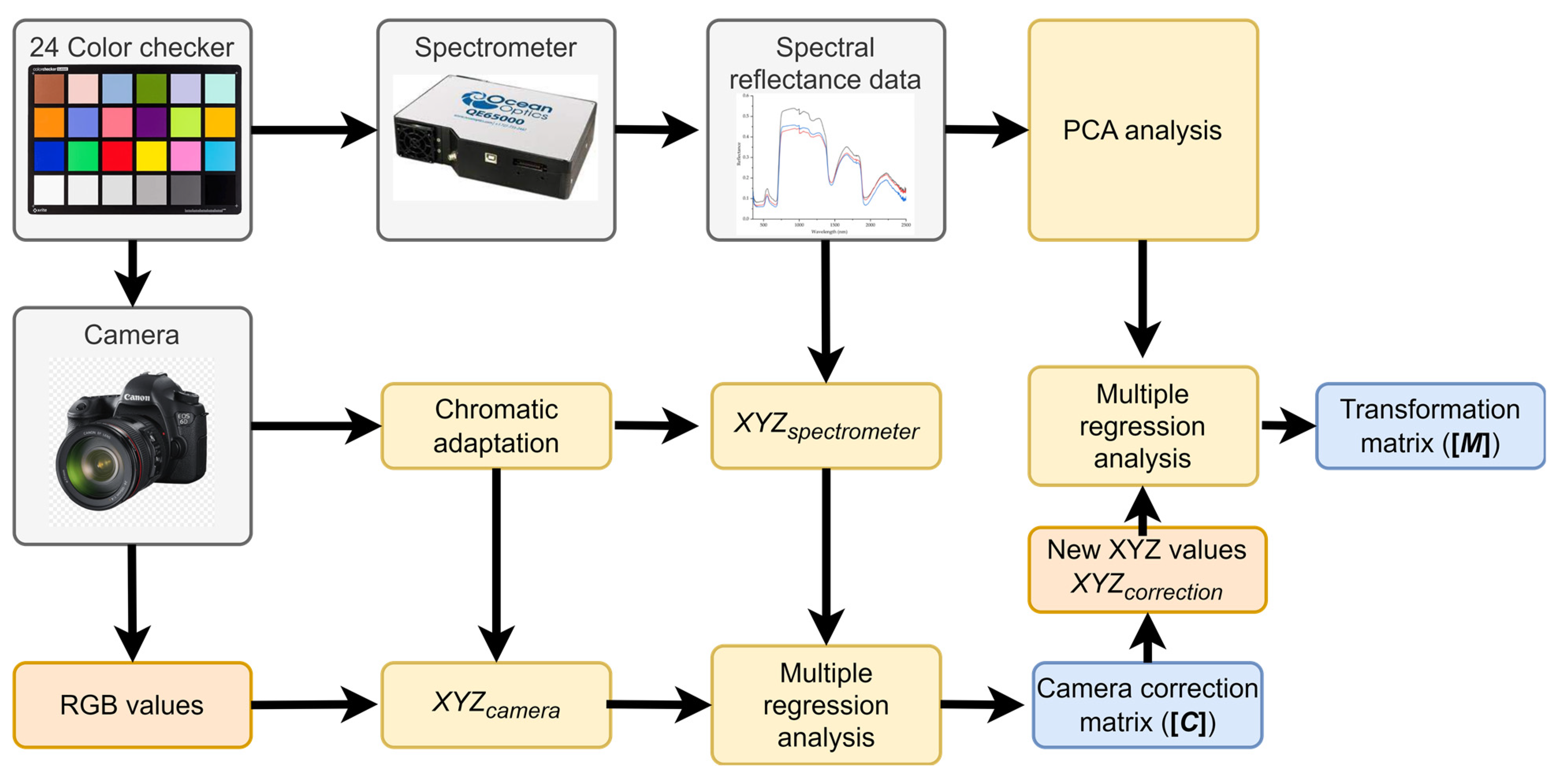
2.3.1. Spectrometer to XYZ Color Space Conversion
2.3.2. Non-Linear XYZ Correction
2.3.3. Correction Matrix C and the Calibration Camera and Spectrometer
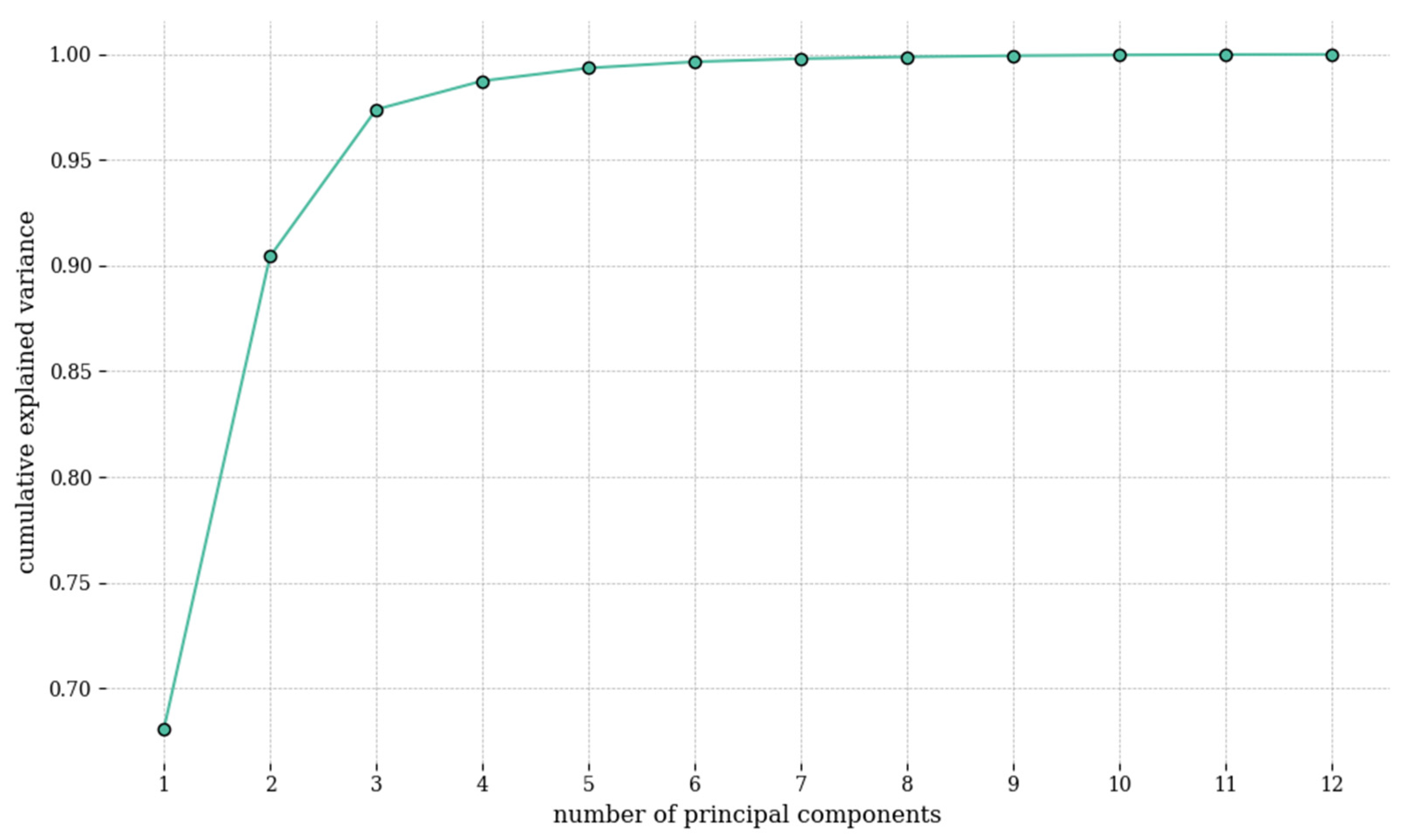
2.3.4. Principal Component Analysis of Reflectance Spectrum
2.3.5. The Hyperspectrum
2.4. Classification Task
2.5. Training Strategy and Performance Evaluation
3. Results and Discussions
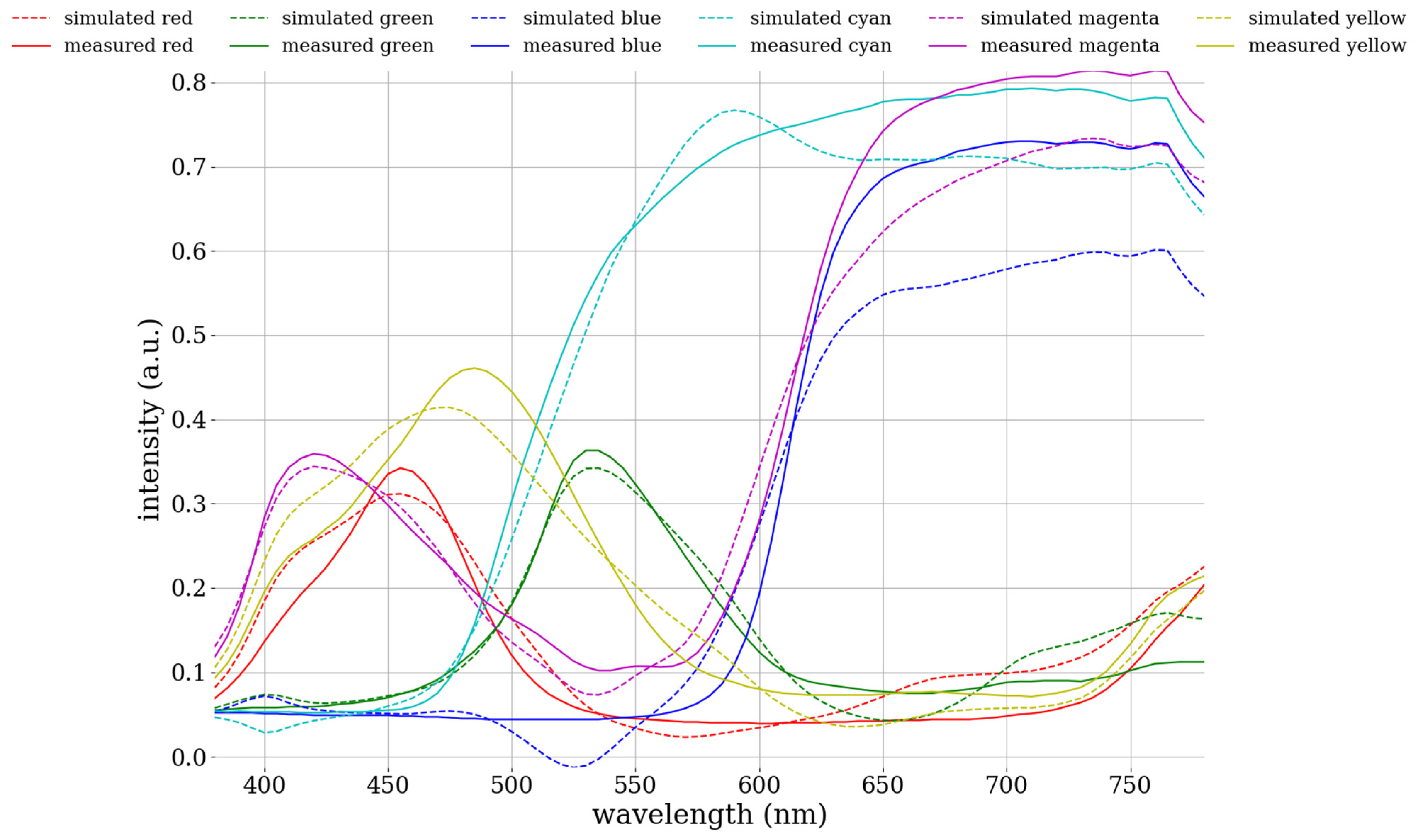
3.1. Evaluate the Performance of the HSI Model
3.2. Segmentation Model Results
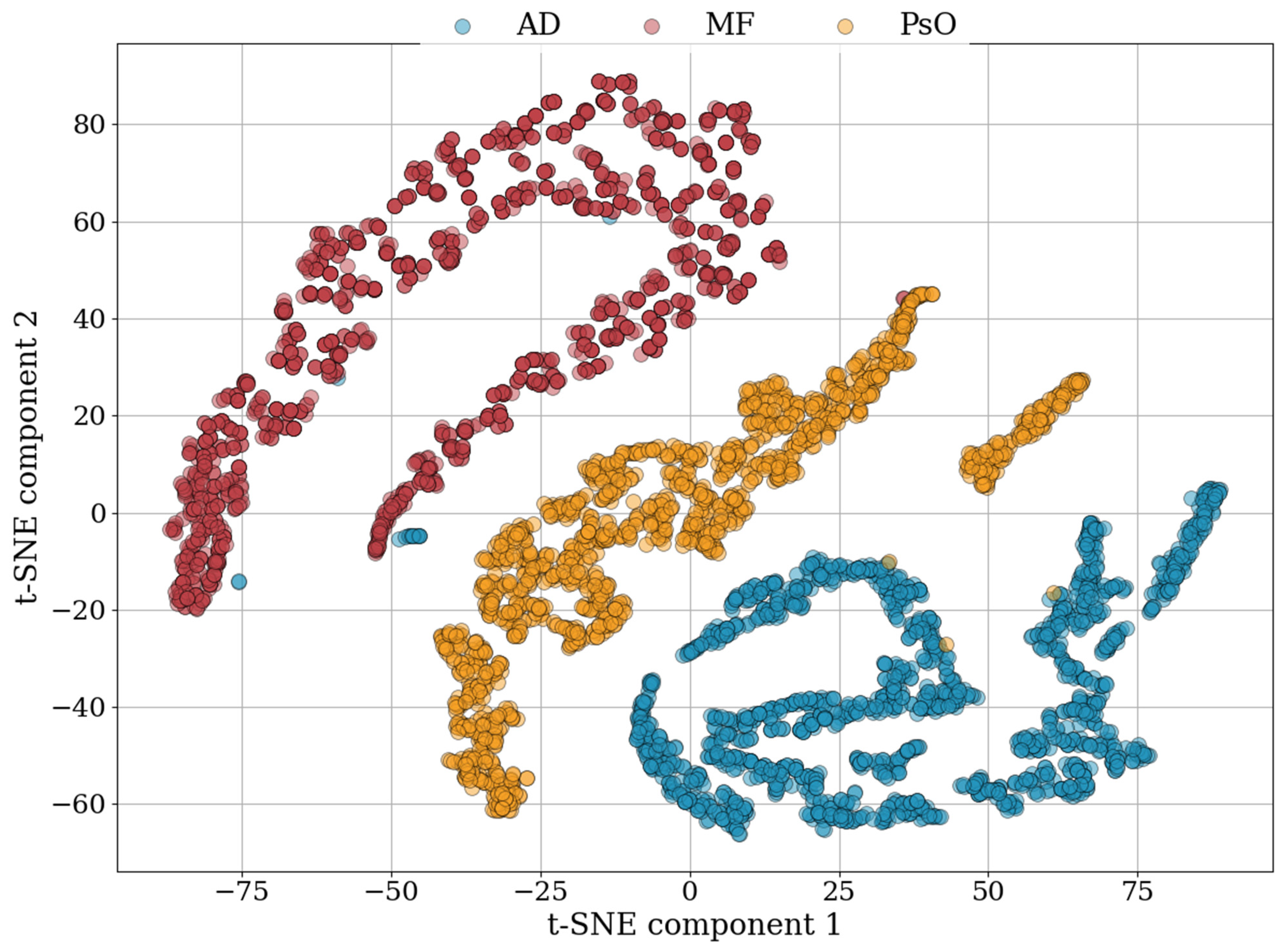
3.3. Classification Task Results
3.4. Comparison with Other Existing Studies
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Querfeld, C.; Zain, J.; Rosen, S.T. Primary cutaneous T-Cell lymphomas: Mycosis fungoides and Sezary syndrome. T-Cell NK-Cell Lymphomas Biol. Nov. Ther. 2019, 176, 225–248. [Google Scholar]
- Stoll, J.R.; Willner, J.; Oh, Y.; Pulitzer, M.; Moskowitz, A.; Horwitz, S.; Myskowski, P.; Noor, S.J. Primary cutaneous T-cell lymphomas other than mycosis fungoides and Sézary syndrome. Part I: Clinical and histologic features and diagnosis. J. Am. Acad. Dermatol. 2021, 85, 1073–1090. [Google Scholar] [CrossRef] [PubMed]
- García-Díaz, N.; Piris, M.Á.; Ortiz-Romero, P.L.; Vaqué, J.P. Mycosis fungoides and sézary syndrome: An integrative review of the pathophysiology, molecular drivers, and targeted therapy. Cancers 2021, 13, 1931. [Google Scholar] [CrossRef] [PubMed]
- Dobos, G.; Lazaridou, I.; de Masson, A. Mycosis Fungoides and Sézary Syndrome: Microenvironment and Cancer Progression. Cancers 2023, 15, 746. [Google Scholar] [CrossRef] [PubMed]
- Hsiao, Y.-P.; Wang, H.-C.; Chen, S.-H.; Tsai, C.-H.; Yang, J.-H. Identified early stage mycosis fungoides from psoriasis and atopic dermatitis using non-invasive color contrast enhancement by LEDs lighting. Opt. Quantum Electron. 2015, 47, 1599–1611. [Google Scholar] [CrossRef]
- Fujimura, T.; Okuyama, R.; Ito, Y.; Aiba, S. Profiles of Foxp3+ regulatory T cells in eczematous dermatitis, psoriasis vulgaris and mycosis fungoides. Br. J. Dermatol. 2008, 158, 1256–1263. [Google Scholar] [CrossRef] [PubMed]
- Nashan, D.; Faulhaber, D.; Ständer, S.; Luger, T.; Stadler, R. Mycosis fungoides: A dermatological masquerader. Br. J. Dermatol. 2007, 156, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Cerroni, L. Mycosis fungoides-clinical and histopathologic features, differential diagnosis, and treatment. Semin. Cutan. Med. Surg. 2018, 37, 2–10. [Google Scholar] [CrossRef]
- Shamim, H.; Johnson, E.F.; Gibson, L.E.; Comfere, N. Mycosis fungoides with spongiosis: A potential diagnostic pitfall. J. Cutan. Pathol. 2019, 46, 645–652. [Google Scholar] [CrossRef]
- van Santen, S.; Jansen, P.M.; Quint, K.D.; Vermeer, M.H.; Willemze, R. Plaque stage folliculotropic mycosis fungoides: Histopathologic features and prognostic factors in a series of 40 patients. J. Cutan. Pathol. 2020, 47, 241–250. [Google Scholar] [CrossRef]
- Hodak, E.; Geskin, L.; Guenova, E.; Ortiz-Romero, P.L.; Willemze, R.; Zheng, J.; Cowan, R.; Foss, F.; Mangas, C.; Querfeld, C. Real-Life barriers to diagnosis of early mycosis fungoides: An international expert panel discussion. Am. J. Clin. Dermatol. 2023, 24, 5–14. [Google Scholar] [CrossRef] [PubMed]
- Roelens, M.; de Masson, A.; Ram-Wolff, C.; Maki, G.; Cayuela, J.M.; Marie-Cardine, A.; Bensussan, A.; Toubert, A.; Bagot, M.; Moins-Teisserenc, H. Revisiting the initial diagnosis and blood staging of mycosis fungoides and Sézary syndrome with the KIR3DL2 marker. Br. J. Dermatol. 2020, 182, 1415–1422. [Google Scholar] [CrossRef] [PubMed]
- Horna, P.; Wang, S.A.; Wolniak, K.L.; Psarra, K.; Almeida, J.; Illingworth, A.J.; Johansson, U.; Craig, F.E.; Torres, R. Flow cytometric evaluation of peripheral blood for suspected Sézary syndrome or mycosis fungoides: International guidelines for assay characteristics. Cytom. Part B Clin. Cytom. 2021, 100, 142–155. [Google Scholar] [CrossRef] [PubMed]
- Lyapichev, K.A.; Bah, I.; Huen, A.; Duvic, M.; Routbort, M.J.; Wang, W.; Jorgensen, J.L.; Medeiros, L.J.; Vega, F.; Craig, F.E. Determination of immunophenotypic aberrancies provides better assessment of peripheral blood involvement by mycosis fungoides/Sézary syndrome than quantification of CD26− or CD7− CD4+ T-cells. Cytom. Part B Clin. Cytom. 2021, 100, 183–191. [Google Scholar] [CrossRef] [PubMed]
- Allen, P.B.; McCook-Veal, A.A.; Switchenko, J.M.; Paulino, D.M.; Niyogusaba, T.; Baird, K.M.; Tarabadkar, E.S.; Lechowicz, M.J. Staging lymph nodes and blood at diagnosis in mycosis fungoides identifies patients at increased risk of progression to advanced stage: A retrospective cohort study. Cancer 2023, 129, 541–550. [Google Scholar] [CrossRef] [PubMed]
- Guitart, J. Sézary syndrome and mycosis fungoides flow cytometric evaluation: The clinicians’ perspective. Cytom. Part B Clin. Cytom. 2021, 100, 129–131. [Google Scholar] [CrossRef] [PubMed]
- Vermeer, M.H.; Moins-Teisserenc, H.; Bagot, M.; Quaglino, P.; Whittaker, S. Flow cytometry for the assessment of blood tumour burden in cutaneous T-cell lymphoma: Towards a standardized approach. Br. J. Dermatol. 2022, 187, 21–28. [Google Scholar] [CrossRef] [PubMed]
- Illingworth, A.; Johansson, U.; Huang, S.; Horna, P.; Wang, S.A.; Almeida, J.; Wolniak, K.L.; Psarra, K.; Torres, R.; Craig, F.E. International guidelines for the flow cytometric evaluation of peripheral blood for suspected Sézary syndrome or mycosis fungoides: Assay development/optimization, validation, and ongoing quality monitors. Cytom. Part B Clin. Cytom. 2021, 100, 156–182. [Google Scholar] [CrossRef]
- Hodak, E.; Sherman, S.; Papadavid, E.; Bagot, M.; Querfeld, C.; Quaglino, P.; Prince, H.; Ortiz-Romero, P.; Stadler, R.; Knobler, R. Should we be imaging lymph nodes at initial diagnosis of early-stage mycosis fungoides? Results from the PROspective Cutaneous Lymphoma International Prognostic Index (PROCLIPI) international study. Br. J. Dermatol. 2021, 184, 524–531. [Google Scholar] [CrossRef]
- Calvani, J.; De Masson, A.; de Margerie-Mellon, C.; de Kerviler, É.; Ram-Wolff, C.; Gruber, A.; Meignin, V.; Brice, P.; Sadoux, A.; Mourah, S. Image-guided lymph node core-needle biopsy predicts survival in mycosis fungoides and Sézary syndrome. Br. J. Dermatol. 2021, 185, 419–427. [Google Scholar] [CrossRef]
- Jonak, C.; Tittes, J.; Brunner, P.M.; Guenova, E. Mycosis fungoides and Sézary syndrome. JDDG J. Der Dtsch. Dermatol. Ges. 2021, 19, 1307–1334. [Google Scholar] [CrossRef] [PubMed]
- McDonald, E.O.; Amanullah, A.A.; Park, P.S.U.; Song, W.; Werner, T.J.; Alavi, A.; Revheim, M.-E. The role of 18F-FDG PET/CT in primary cutaneous lymphoma: An educational review. Ann. Nucl. Med. 2023, 37, 328–348. [Google Scholar] [CrossRef] [PubMed]
- Gallamini, A.; Zwarthoed, C.; Borra, A. Positron emission tomography (PET) in oncology. Cancers 2014, 6, 1821–1889. [Google Scholar] [CrossRef] [PubMed]
- Salem, A.E.; Shah, H.R.; Covington, M.F.; Koppula, B.R.; Fine, G.C.; Wiggins, R.H.; Hoffman, J.M.; Morton, K.A. PET-CT in Clinical Adult Oncology: I. Hematologic Malignancies. Cancers 2022, 14, 5941. [Google Scholar] [CrossRef] [PubMed]
- Hsiao, Y.-P.; Chiu, C.-W.; Lu, C.-W.; Nguyen, H.T.; Tseng, Y.S.; Hsieh, S.-C.; Wang, H.-C. Identification of skin lesions by using single-step multiframe detector. J. Clin. Med. 2021, 10, 144. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wang, Y.; Cai, J.; Lee, T.K.; Miao, C.; Wang, Z.J. Ssd-kd: A self-supervised diverse knowledge distillation method for lightweight skin lesion classification using dermoscopic images. Med. Image Anal. 2023, 84, 102693. [Google Scholar] [CrossRef] [PubMed]
- Sumithra, R.; Suhil, M.; Guru, D. Segmentation and classification of skin lesions for disease diagnosis. Procedia Comput. Sci. 2015, 45, 76–85. [Google Scholar] [CrossRef]
- Balaji, V.; Suganthi, S.; Rajadevi, R.; Kumar, V.K.; Balaji, B.S.; Pandiyan, S. Skin disease detection and segmentation using dynamic graph cut algorithm and classification through Naive Bayes classifier. Measurement 2020, 163, 107922. [Google Scholar] [CrossRef]
- Wu, H.; Chen, S.; Chen, G.; Wang, W.; Lei, B.; Wen, Z. FAT-Net: Feature adaptive transformers for automated skin lesion segmentation. Med. Image Anal. 2022, 76, 102327. [Google Scholar] [CrossRef]
- Ke, G.; Meng, Q.; Finley, T.; Wang, T.; Chen, W.; Ma, W.; Ye, Q.; Liu, T.-Y. Lightgbm: A highly efficient gradient boosting decision tree. Adv. Neural Inf. Process. Syst. 2017, 30, 3149–3157. [Google Scholar]
- Chan, I.L.; Cohen, S.; da Cunha, M.G.; Maluf, L.C. Characteristics and management of Asian skin. Int. J. Dermatol. 2019, 58, 131–143. [Google Scholar] [CrossRef] [PubMed]
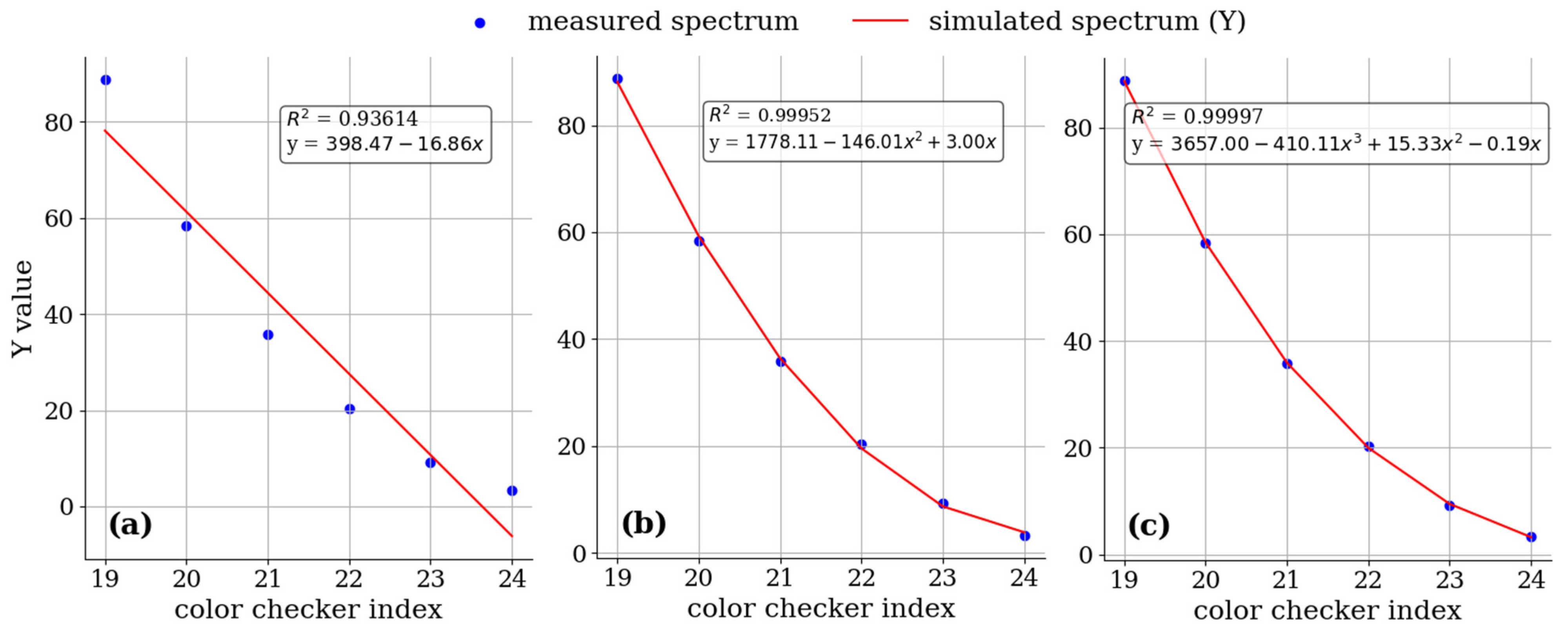
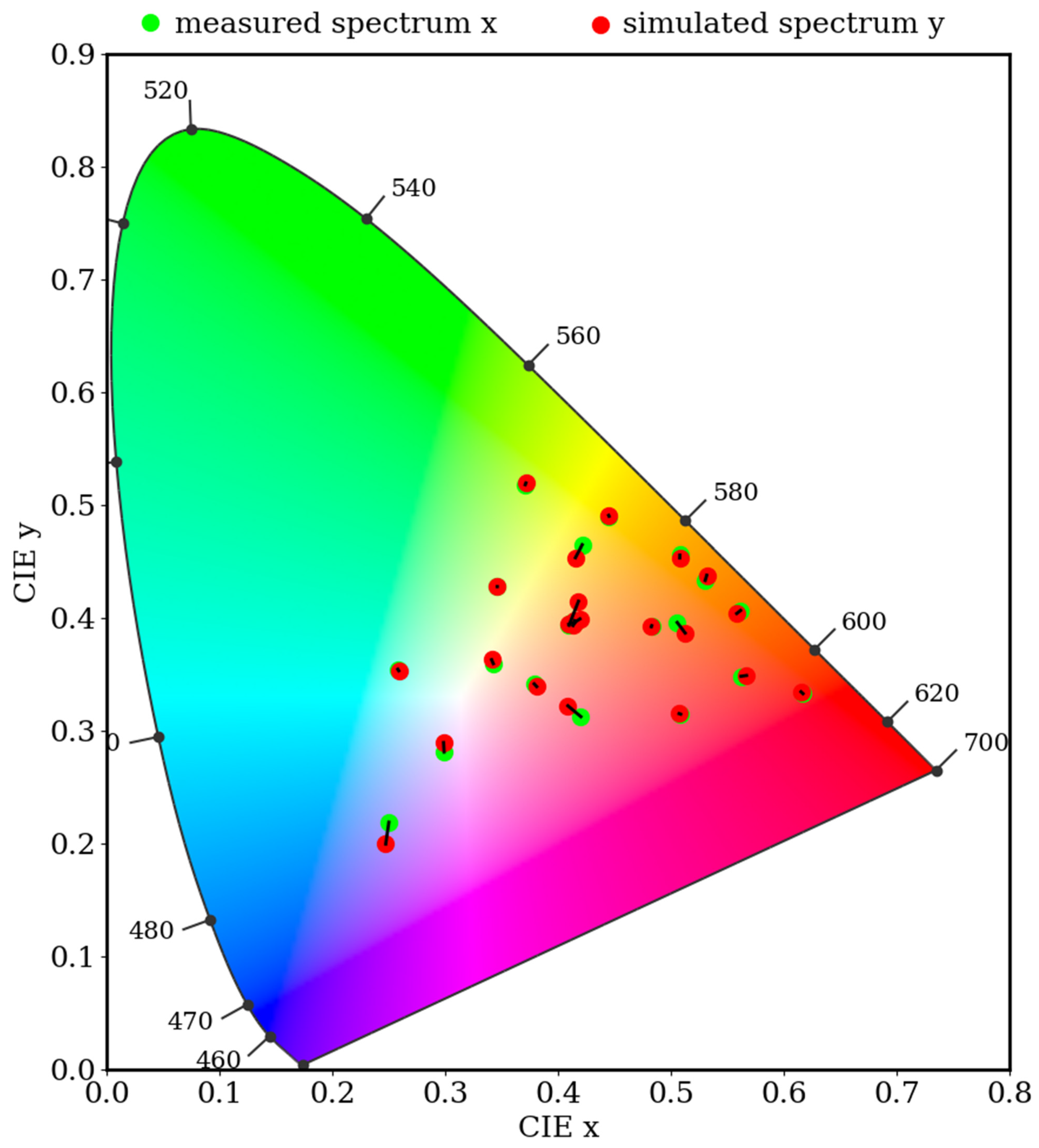
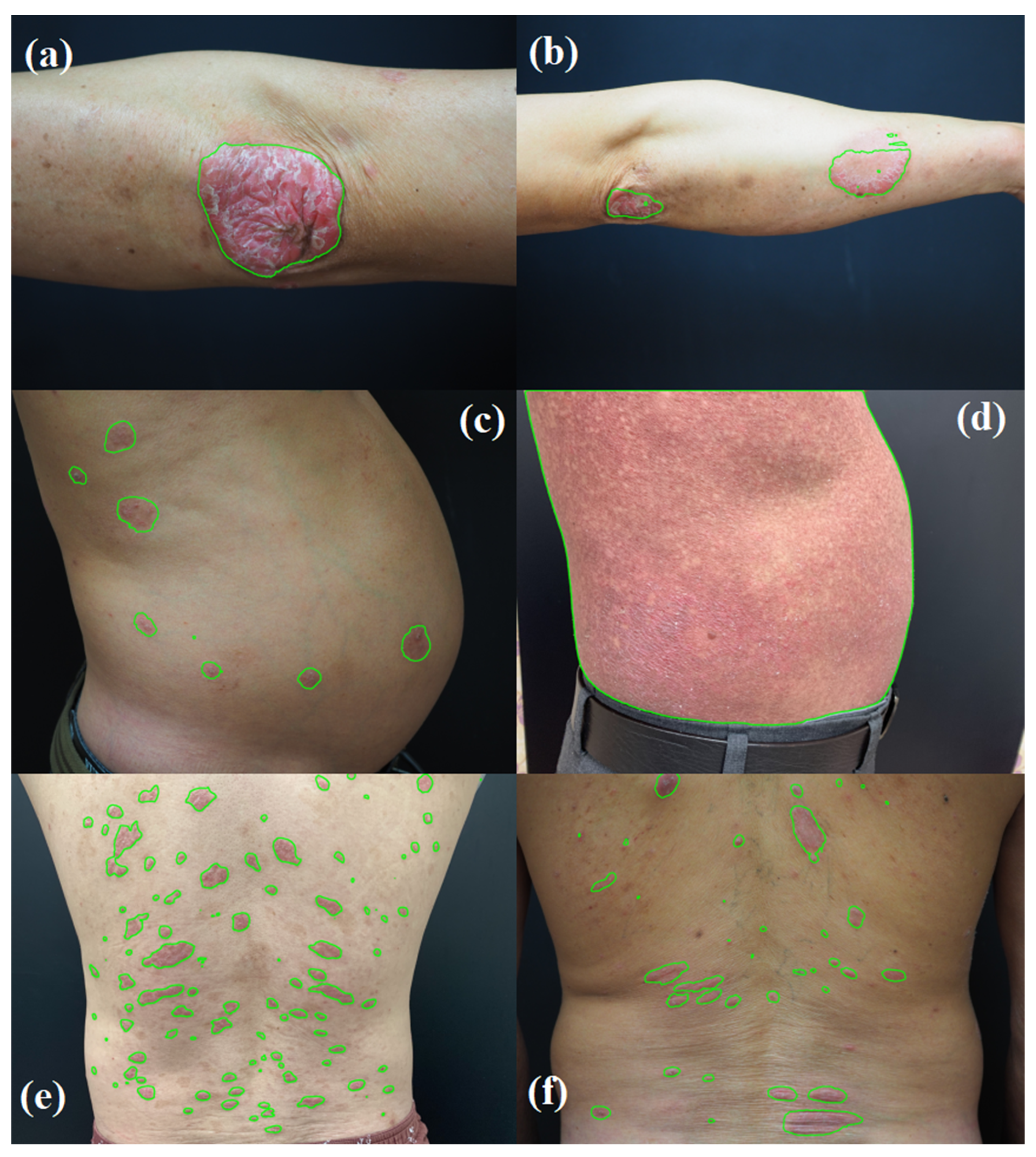
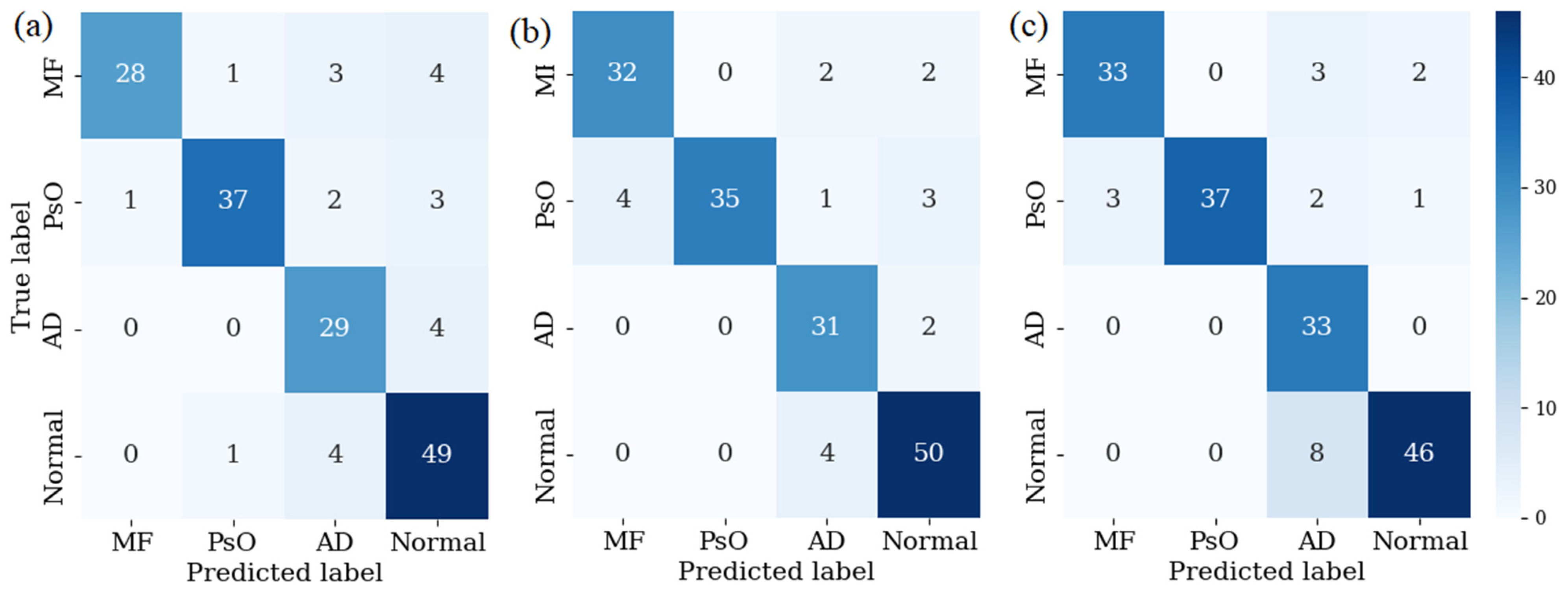
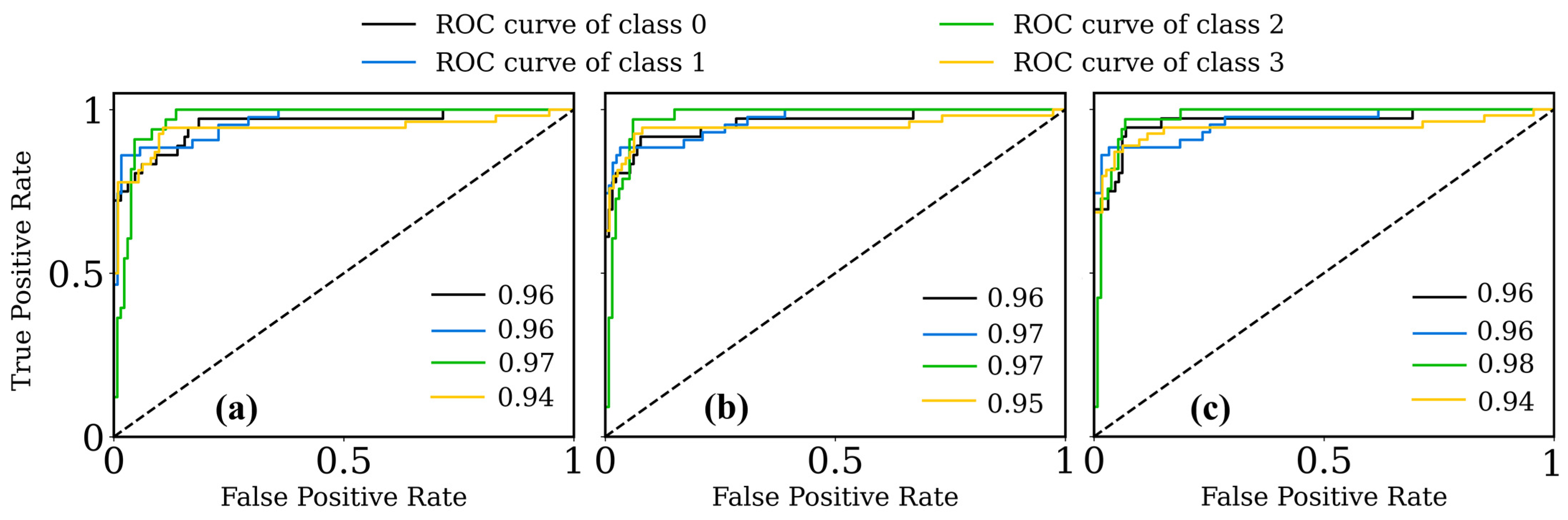
| Color No. | Measured | Simulated | L* (Measured) | a* (Measured) | b* (Measured) | L* (Simulated) | a* (Simulated) | b* (Simulated) | CIEDE 2000 |
|---|---|---|---|---|---|---|---|---|---|
| 1 | 38.94 | 24.35 | 36.89 | 38.81 | 24.92 | 38.23 | 0.52 | ||
| 2 | 67.93 | 32.09 | 49.08 | 67.95 | 32.03 | 48.94 | 0.05 | ||
| 3 | 49.45 | 0.07 | 9.87 | 49.34 | −0.66 | 9.68 | 1.01 | ||
| 4 | 42.33 | −4.05 | 39.45 | 42.59 | −3.97 | 38.03 | 0.57 | ||
| 5 | 55.89 | 16.17 | 11.32 | 55.88 | 16.12 | 11.33 | 0.04 | ||
| 6 | 69.10 | −19.39 | 31.48 | 69.10 | −19.46 | 31.45 | 0.05 | ||
| 7 | 65.32 | 46.33 | 82.11 | 65.37 | 46.43 | 81.81 | 0.14 | ||
| 8 | 39.09 | 8.89 | −10.49 | 39.23 | 9.36 | −10.40 | 0.49 | ||
| 9 | 55.11 | 58.96 | 46.87 | 55.06 | 59.03 | 47.19 | 0.14 | ||
| 10 | 31.22 | 24.81 | 6.34 | 31.78 | 25.36 | 6.05 | 0.57 | ||
| 11 | 71.87 | −5.83 | 76.56 | 71.87 | −5.85 | 76.51 | 0.02 | ||
| 12 | 74.87 | 34.26 | 90.13 | 74.84 | 34.16 | 90.25 | 0.08 | ||
| 13 | 27.87 | 11.77 | −23.10 | 27.60 | 11.34 | −23.26 | 0.46 | ||
| 14 | 53.97 | −27.31 | 50.50 | 53.95 | −27.22 | 50.74 | 0.11 | ||
| 15 | 45.76 | 66.20 | 51.52 | 45.76 | 65.96 | 51.01 | 0.17 | ||
| 16 | 83.76 | 23.22 | 100.15 | 83.76 | 23.26 | 100.18 | 0.02 | ||
| 17 | 54.10 | 58.28 | 24.01 | 54.11 | 58.30 | 24.01 | 0.01 | ||
| 18 | 48.03 | −23.70 | −0.08 | 48.08 | −23.35 | −0.02 | 0.19 | ||
| 19 | 95.47 | 15.52 | 46.28 | 95.48 | 15.52 | 46.28 | 0.00 | ||
| 20 | 80.98 | 13.58 | 40.13 | 80.95 | 13.62 | 40.11 | 0.04 | ||
| 21 | 66.39 | 11.35 | 34.00 | 66.52 | 11.47 | 34.18 | 0.14 | ||
| 22 | 52.19 | 9.40 | 28.14 | 51.90 | 9.29 | 28.26 | 0.31 | ||
| 23 | 36.45 | 6.84 | 21.33 | 36.46 | 6.79 | 21.44 | 0.09 | ||
| 24 | 21.36 | 4.87 | 14.76 | 21.05 | 3.68 | 15.29 | 1.55 | ||
| average | 0.28 |
| k-Fold | Sensitivity | Specificity | F1-Score | ROC-AUC |
|---|---|---|---|---|
| k3 | 85.61% | 95.25% | 86.26% | 0.9051 |
| k5 | 89.20% | 96.35% | 89.19% | 0.9270 |
| k7 | 90.72% | 96.76% | 90.08% | 0.9351 |
| Model | Number of Params | Size (MB) | IoU | AUC-ROC | Time Prediction per Image (second) |
|---|---|---|---|---|---|
| U-Net | 31M | 124 | 0.8003 | 0.8490 | 0.0741 |
| U-Net++ | 55M | 220 | 0.8977 | 0.9334 | 0.1182 |
| U-Net Attention | 35M | 140 | 0.8521 | 0.9097 | 0.0792 |
| DeepLabv3 | 41M | 164 | 0.8730 | 0.9597 | 0.0836 |
| Ours (3-folds) | 35M | 148 | 0.8521 | 0.9051 | 0.0810 |
| Ours (5-folds) | 35M | 148 | 0.8521 | 0.9270 | 0.0810 |
| Ours (7-folds) | 35M | 148 | 0.8521 | 0.9351 | 0.0810 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, H.-Y.; Nguyen, H.-T.; Lin, T.-L.; Saenprasarn, P.; Liu, P.-H.; Wang, H.-C. Identification of Skin Lesions by Snapshot Hyperspectral Imaging. Cancers 2024, 16, 217. https://doi.org/10.3390/cancers16010217
Huang H-Y, Nguyen H-T, Lin T-L, Saenprasarn P, Liu P-H, Wang H-C. Identification of Skin Lesions by Snapshot Hyperspectral Imaging. Cancers. 2024; 16(1):217. https://doi.org/10.3390/cancers16010217
Chicago/Turabian StyleHuang, Hung-Yi, Hong-Thai Nguyen, Teng-Li Lin, Penchun Saenprasarn, Ping-Hung Liu, and Hsiang-Chen Wang. 2024. "Identification of Skin Lesions by Snapshot Hyperspectral Imaging" Cancers 16, no. 1: 217. https://doi.org/10.3390/cancers16010217
APA StyleHuang, H.-Y., Nguyen, H.-T., Lin, T.-L., Saenprasarn, P., Liu, P.-H., & Wang, H.-C. (2024). Identification of Skin Lesions by Snapshot Hyperspectral Imaging. Cancers, 16(1), 217. https://doi.org/10.3390/cancers16010217







