Theranostics of Primary Prostate Cancer: Beyond PSMA and GRP-R
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Patient Characteristics
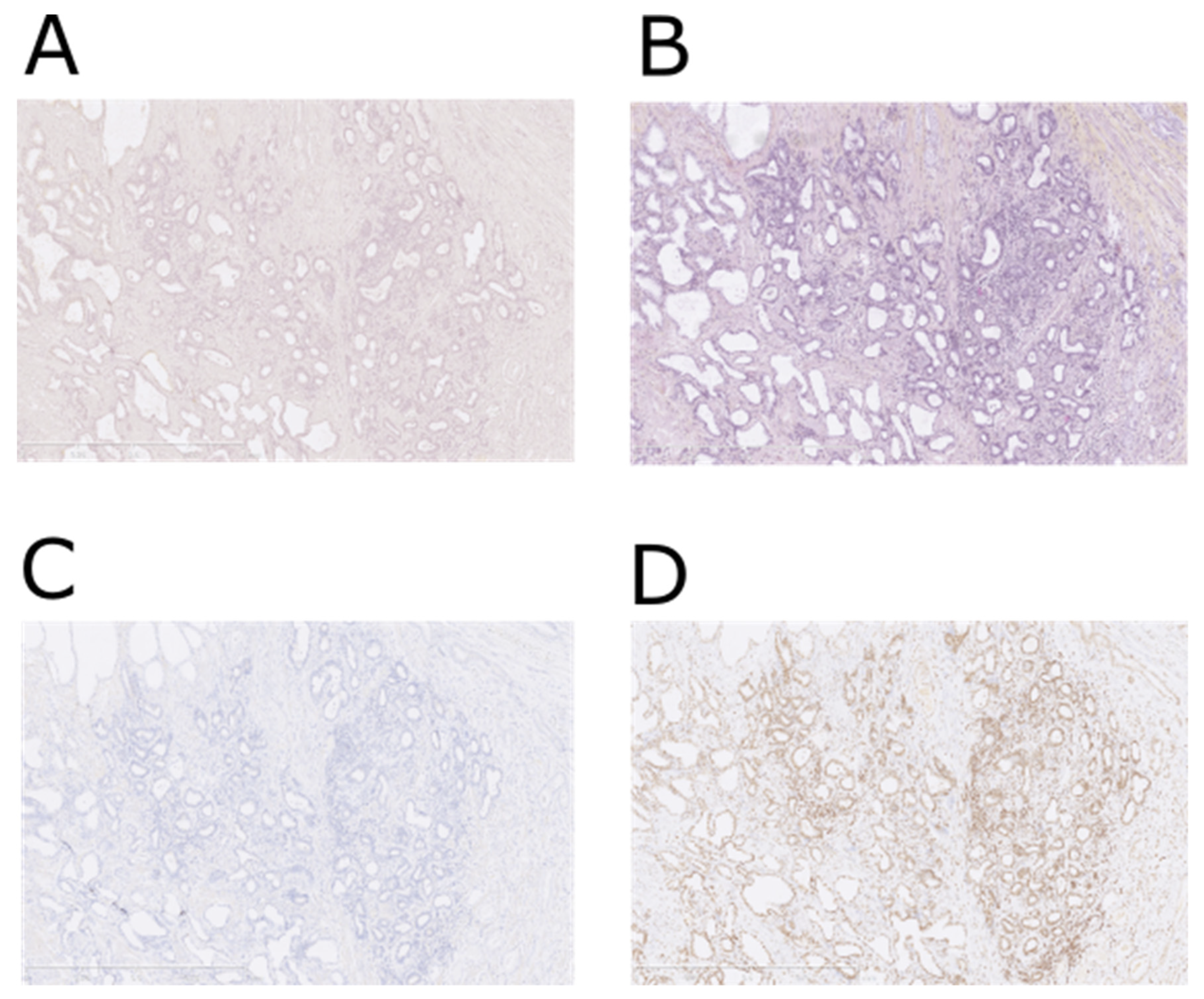
2.2. NTS1—Immunohistochemistry
2.3. Radiosynthesis and Quality Control of Radioligands
2.4. High-Resolution Microimaging
2.4.1. Binding Assay
2.4.2. Tissue Microimaging
2.5. Data Analysis
2.6. Statistical Analysis
3. Results
3.1. Study 1: Prospective NTS1 IHC Study
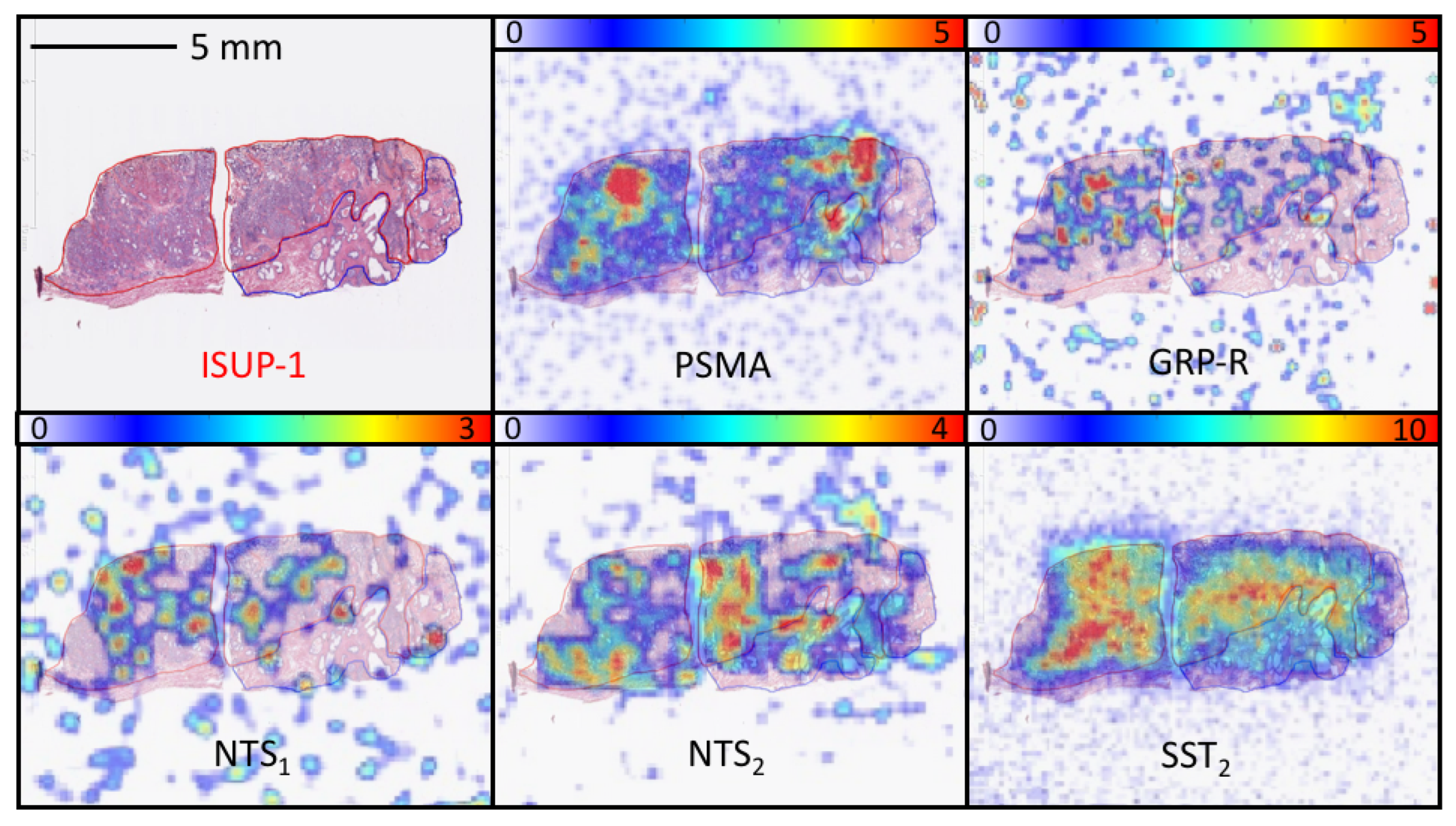
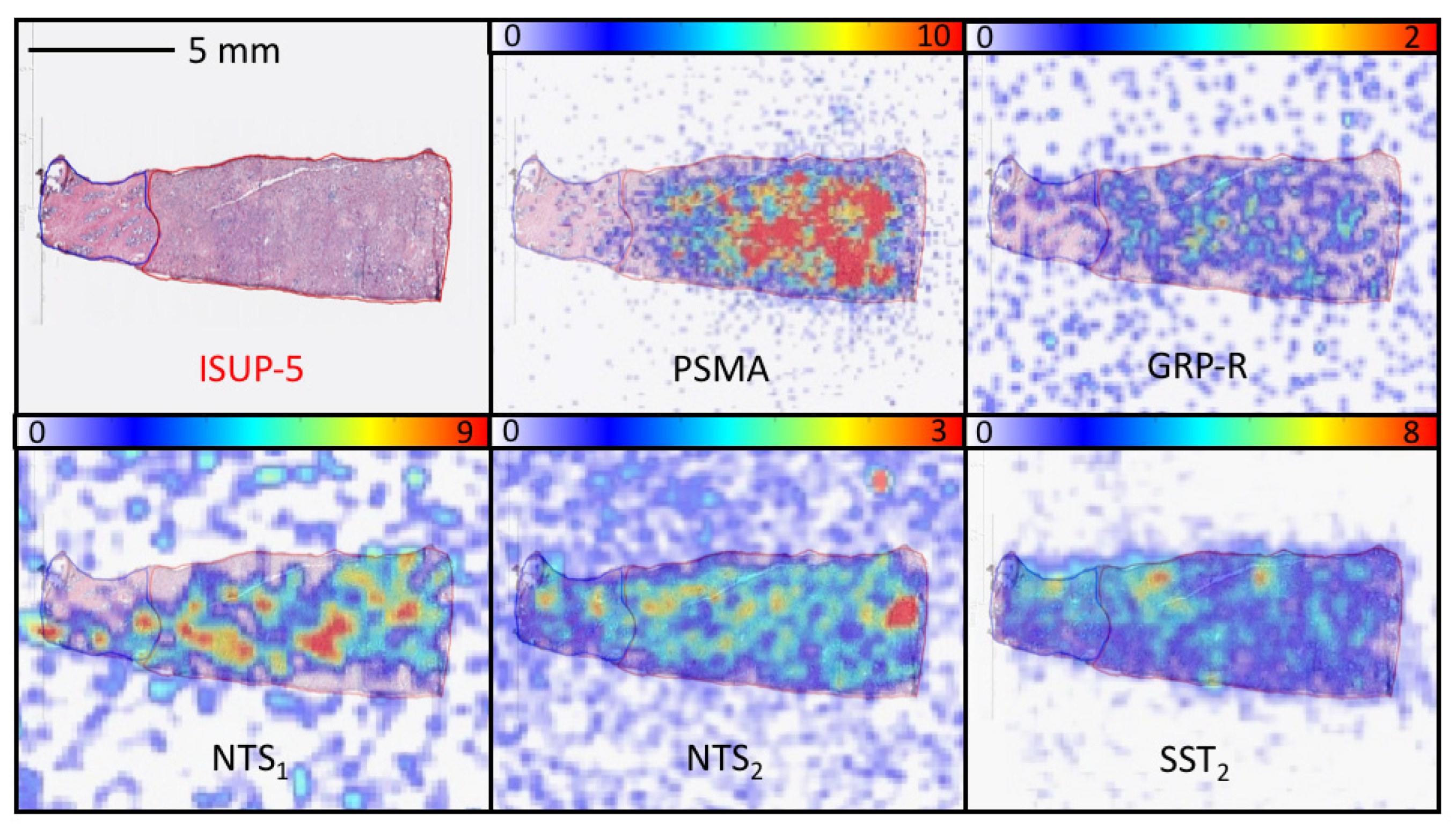
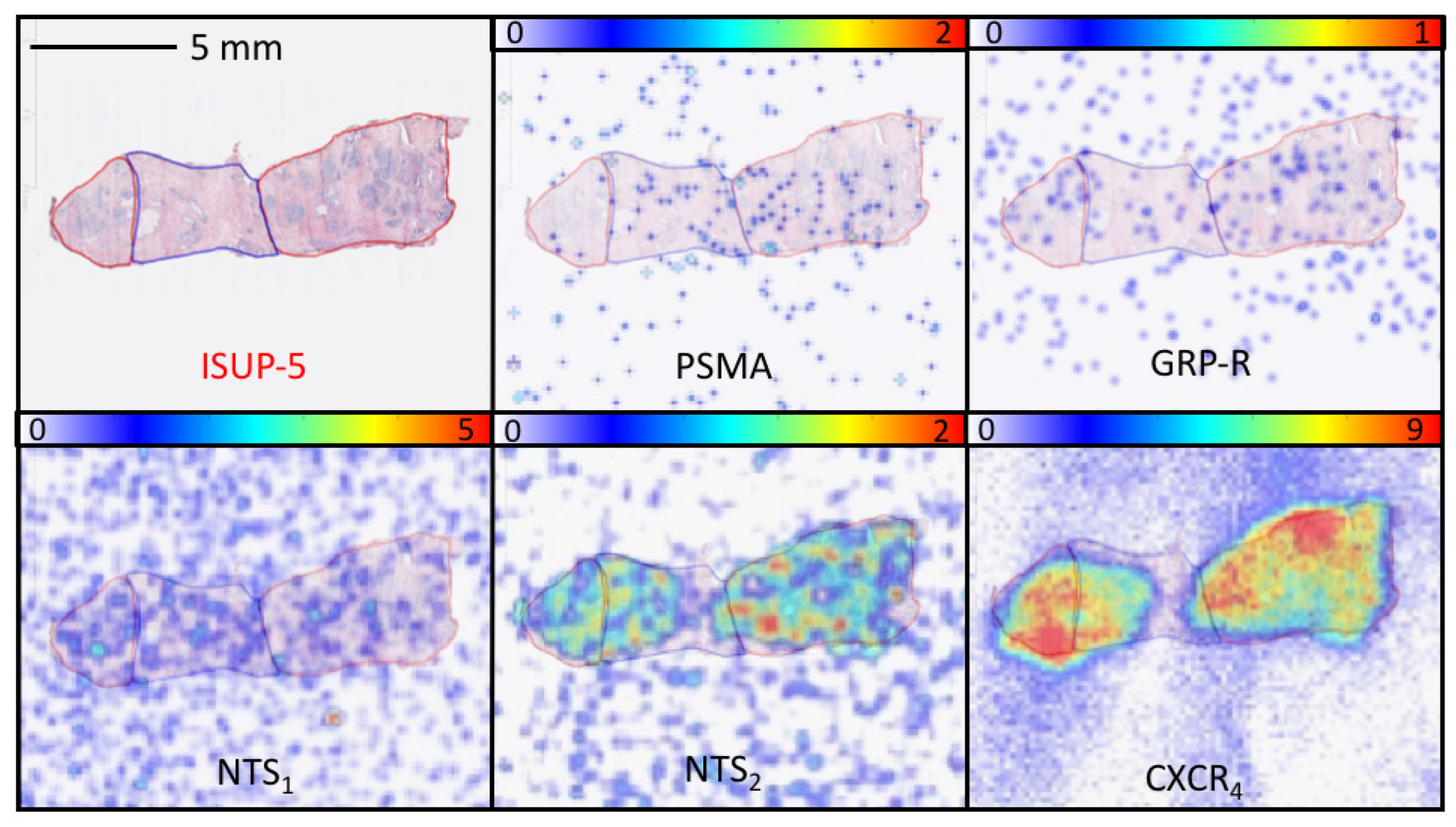
3.2. Study 2: Retrospective Study of the Expression of PSMA, GRP-R, NTS1, NTS2, SST2 and CXCR4 on Samples of Primary Prostate Cancer
3.3. Radiopharmaceuticals
3.4. Quantitative Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Wagle, N.S.; Jemal, A. Cancer statistics, 2023. CA Cancer J. Clin. 2023, 73, 17–48. [Google Scholar] [CrossRef] [PubMed]
- Fütterer, J.J.; Briganti, A.; De Visschere, P.; Emberton, M.; Giannarini, G.; Kirkham, A.; Taneja, S.S.; Thoeny, H.; Villeirs, G.; Villers, A. Can Clinically Significant Prostate Cancer Be Detected with Multiparametric Magnetic Resonance Imaging? A Systematic Review of the Literature. Eur. Urol. 2015, 68, 1045–1053. [Google Scholar] [CrossRef] [PubMed]
- Grenabo Bergdahl, A.; Wilderäng, U.; Aus, G.; Carlsson, S.; Damber, J.-E.; Franlund, M.; Geterud, K.; Khatami, A.; Socratous, A.; Stranne, J.; et al. Role of Magnetic Resonance Imaging in Prostate Cancer Screening: A Pilot Study Within the Göteborg Randomised Screening Trial. Eur. Urol. 2016, 70, 566–573. [Google Scholar] [CrossRef] [PubMed]
- Siddiqui, M.M.; Rais-Bahrami, S.; Turkbey, B.; George, A.K.; Rothwax, J.; Shakir, N.; Okoro, C.; Raskolnikov, D.; Parnes, H.L.; Linehan, W.M.; et al. Comparison of MR/Ultrasound Fusion–Guided Biopsy With Ultrasound-Guided Biopsy for the Diagnosis of Prostate Cancer. JAMA 2015, 313, 390. [Google Scholar] [CrossRef]
- Maurer, T.; Eiber, M.; Schwaiger, M.; Gschwend, J.E. Current use of PSMA–PET in prostate cancer management. Nat. Rev. Urol. 2016, 13, 226–235. [Google Scholar] [CrossRef]
- Hofman, M.S.; Lawrentschuk, N.; Francis, R.J.; Tang, C.; Vela, I.; Thomas, P.; Rutherford, N.; Martin, J.M.; Frydenberg, M.; Shakher, R.; et al. Prostate-specific membrane antigen PET-CT in patients with high-risk prostate cancer before curative-intent surgery or radiotherapy (proPSMA): A prospective, randomised, multicentre study. Lancet 2020, 395, 1208–1216. [Google Scholar] [CrossRef]
- Morgat, C.; Schollhammer, R.; MacGrogan, G.; Barthe, N.; Vélasco, V.; Vimont, D.; Cazeau, A.-L.; Fernandez, P.; Hindié, E. Comparison of the binding of the gastrin-releasing peptide receptor (GRP-R) antagonist 68Ga-RM2 and 18F-FDG in breast cancer samples. PLoS ONE 2019, 14, e0210905. [Google Scholar] [CrossRef]
- Schollhammer, R.; Gallerande, H.D.C.; Yacoub, M.; Ranty, M.-L.Q.; Barthe, N.; Vimont, D.; Hindié, E.; Fernandez, P.; Morgat, C. Comparison of the radiolabeled PSMA-inhibitor 111In-PSMA-617 and the radiolabeled GRP-R antagonist 111In-RM2 in primary prostate cancer samples. EJNMMI Res. 2019, 9, 52. [Google Scholar] [CrossRef]
- Schollhammer, R.; Robert, G.; Asselineau, J.; Yacoub, M.; Vimont, D.; Balamoutoff, N.; Bladou, F.; Bénard, A.; Hindié, E.; de Clermont Gallerande, E.; et al. Comparison of 68Ga-PSMA-617 PET/CT and 68Ga-RM2 PET/CT in patients with localized prostate cancer candidate for radical prostatectomy: A prospective, single arm, single center, phase II study. J. Nucl. Med. 2023, 64, 379–385. [Google Scholar] [CrossRef]
- Morgat, C.; Chastel, A.; Molinie, V.; Schollhammer, R.; Macgrogan, G.; Vélasco, V.; Malavaud, B.; Fernandez, P.; Hindié, E. Neurotensin Receptor-1 Expression in Human Prostate Cancer: A Pilot Study on Primary Tumors and Lymph Node Metastases. Int. J. Mol. Sci. 2019, 20, 1721. [Google Scholar] [CrossRef]
- Morichetti, D.; Mazzucchelli, R.; Santinelli, A.; Stramazzotti, D.; Lopez-Beltran, A.; Scarpelli, M.; Bono, A.; Cheng, L.; Montironi, R. Immunohistochemical Expression and Localization of Somatostatin Receptor Subtypes in Prostate Cancer with Neuroendocrine Differentiation. Int. J. Immunopathol. Pharmacol. 2010, 23, 511–522. [Google Scholar] [CrossRef] [PubMed]
- Dubrovska, A.; Elliott, J.; Salamone, R.J.; Telegeev, G.D.; Stakhovsky, A.E.; Schepotin, I.B.; Yan, F.; Wang, Y.; Bouchez, L.C.; Kularatne, S.A.; et al. CXCR4 Expression in Prostate Cancer Progenitor Cells. PLoS ONE 2012, 7, e31226. [Google Scholar] [CrossRef] [PubMed]
- Morgat, C.; Bonnefoi, H.; MacGrogan, G.; Brouste, V.; Vélasco, V.; Sévenet, N.; Fernandez, P.; Debled, M.; Hindié, E. Expression of Gastrin-Releasing Peptide Receptor in Breast Cancer and Its Association with Pathologic, Biologic, and Clinical Parameters: A Study of 1,432 Primary Tumors. J. Nucl. Med. 2017, 58, 1401–1407. [Google Scholar] [CrossRef] [PubMed]
- Benešová, M.; Schäfer, M.; Bauder-Wüst, U.; Afshar-Oromieh, A.; Kratochwil, C.; Mier, W.; Haberkorn, U.; Kopka, K.; Eder, M. Preclinical Evaluation of a Tailor-Made DOTA-Conjugated PSMA Inhibitor with Optimized Linker Moiety for Imaging and Endoradiotherapy of Prostate Cancer. J. Nucl. Med. 2015, 56, 914–920. [Google Scholar] [CrossRef]
- Chastel, A.; Vimont, D.; Claverol, S.; Zerna, M.; Bodin, S.; Berndt, M.; Chaignepain, S.; Hindié, E.; Morgat, C. 68Ga-Radiolabeling and Pharmacological Characterization of a Kit-Based Formulation of the Gastrin-Releasing Peptide Receptor (GRP-R) Antagonist RM2 for Convenient Preparation of [68Ga]Ga-RM2. Pharmaceutics 2021, 13, 1160. [Google Scholar] [CrossRef]
- Fanelli, R.; Chastel, A.; Previti, S.; Hindié, E.; Vimont, D.; Zanotti-Fregonara, P.; Fernandez, P.; Garrigue, P.; Lamare, F.; Schollhammer, R.; et al. Silicon-Containing Neurotensin Analogues as Radiopharmaceuticals for NTS1 -Positive Tumors Imaging. Bioconjug. Chem. 2020, 31, 2339–2349. [Google Scholar] [CrossRef]
- Bodin, S.; Previti, S.; Jestin, E.; Vimont, D.; Ait-Arsa, I.; Lamare, F.; Rémond, E.; Hindié, E.; Cavelier, F.; Morgat, C. Design, Synthesis, and Biological Evaluation of the First Radio-Metalated Neurotensin Analogue Targeting Neurotensin Receptor 2. ACS Omega 2023, 8, 6994–7004. [Google Scholar] [CrossRef]
- Schottelius, M.; Šimeček, J.; Hoffmann, F.; Willibald, M.; Schwaiger, M.; Wester, H.-J. Twins in spirit—episode I: Comparative preclinical evaluation of [68Ga]DOTATATE and [68Ga]HA-DOTATATE. EJNMMI Res. 2015, 5, 22. [Google Scholar] [CrossRef]
- Poschenrieder, A.; Schottelius, M.; Schwaiger, M.; Kessler, H.; Wester, H.-J. The influence of different metal-chelate conjugates of pentixafor on the CXCR4 affinity. EJNMMI Res. 2016, 6, 36. [Google Scholar] [CrossRef]
- Reubi, J.C.; Kvols, L.K.; Waser, B.; Nagorney, D.M.; Heitz, P.U.; Charboneau, J.W.; Reading, C.C.; Moertel, C. Detection of somatostatin receptors in surgical and percutaneous needle biopsy samples of carcinoids and islet cell carcinomas. Cancer Res. 1990, 50, 5969–5977. [Google Scholar]
- Parent, E.E.; Schuster, D.M. Update on 18F-Fluciclovine PET for Prostate Cancer Imaging. J. Nucl. Med. 2018, 59, 733–739. [Google Scholar] [CrossRef] [PubMed]
- Nitsch, S.; Hakenberg, O.W.; Heuschkel, M.; Dräger, D.; Hildebrandt, G.; Krause, B.J.; Schwarzenböck, S.M. Evaluation of Prostate Cancer with 11C- and 18F-Choline PET/CT: Diagnosis and Initial Staging. J. Nucl. Med. 2016, 57, 38S–42S. [Google Scholar] [CrossRef] [PubMed]
- Touijer, K.A.; Michaud, L.; Alvarez, H.A.V.; Gopalan, A.; Kossatz, G.M.; Gonen, M.; Beattie, B.; Sandler, I.; Lyaschenko, S.; Eastham, J.A.; et al. Prospective Study of the Radiolabeled GRPR Antagonist BAY86-7548 for Positron Emission Tomography/Computed Tomography Imaging of Newly Diagnosed Prostate Cancer. Eur. Urol. Oncol. 2019, 2, 166–173. [Google Scholar] [CrossRef]
- He, T.; Wang, M.; Wang, H.; Tan, H.; Tang, Y.; Smith, E.; Wu, Z.; Liao, W.; Hu, S.; Li, Z. Evaluation of neurotensin receptor 1 as potential biomarker for prostate cancer theranostic use. Eur. J. Nucl. Med. Mol. Imaging 2019, 46, 2199–2207. [Google Scholar] [CrossRef]
- Prignon, A.; Provost, C.; Alshoukr, F.; Wendum, D.; Couvelard, A.; Barbet, J.; Forgez, P.; Talbot, J.-N.; Gruaz-Guyon, A. Preclinical Evaluation of 68Ga-DOTA-NT-20.3: A Promising PET Imaging Probe To Discriminate Human Pancreatic Ductal Adenocarcinoma from Pancreatitis. Mol. Pharm. 2019, 16, 2776–2784. [Google Scholar] [CrossRef]
- Maschauer, S.; Greff, C.; Einsiedel, J.; Ott, J.; Tripal, P.; Hübner, H.; Gmeiner, P.; Prante, O. Improved radiosynthesis and preliminary in vivo evaluation of a 18F-labeled glycopeptide–peptoid hybrid for PET imaging of neurotensin receptor 2. Bioorg. Med. Chem. 2015, 23, 4026–4033. [Google Scholar] [CrossRef] [PubMed]
- Swift, S.L.; Burns, J.E.; Maitland, N.J. Altered Expression of Neurotensin Receptors Is Associated with the Differentiation State of Prostate Cancer. Cancer Res. 2010, 70, 347–356. [Google Scholar] [CrossRef] [PubMed]
- Reubi, J.C.; Waser, B.; Mäcke, H.; Rivier, J.E. Highly Increased 125I-JR11 Antagonist Binding In Vitro Reveals Novel Indications for sst 2 Targeting in Human Cancers. J. Nucl. Med. 2017, 58, 300–306. [Google Scholar] [CrossRef]
- Heidegger, I.; Fotakis, G.; Offermann, A.; Goveia, J.; Daum, S.; Salcher, S.; Noureen, A.; Timmer-Boscha, H.; Schäfer, G.; Walenkamp, A.; et al. Comprehensive characterization of the prostate tumor microenvironment identifies CXCR4/CXCL12 crosstalk as a novel antiangiogenic therapeutic target in prostate cancer. Mol. Cancer. 2022, 21, 132. [Google Scholar] [CrossRef]
- Available online: https://www.nccn.org/guidelines/guidelines-detail?category=1&id=1459 (accessed on 16 February 2023).
- Liu, C.; Liu, T.; Zhang, N.; Liu, Y.; Li, N.; Du, P.; Yang, Y.; Liu, M.; Gong, K.; Yang, X.; et al. 68Ga-PSMA-617 PET/CT: A promising new technique for predicting risk stratification and metastatic risk of prostate cancer patients. Eur. J. Nucl. Med. Mol. Imaging 2018, 45, 1852–1861. [Google Scholar] [CrossRef]
- Judmann, B.; Braun, D.; Wängler, B.; Schirmacher, R.; Fricker, G.; Wängler, C. Current state of radiolabeled heterobivalent peptidic ligands in tumor imaging and therapy. Pharmaceuticals 2020, 13, 173. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Wang, M.; Wang, H.; Zhang, T.; Wu, Z.; Sutton, M.V.; Popik, V.V.; Jiang, G.; Li, Z. Development of bispecific NT-PSMA heterodimer for prostate cancer imaging: A potenial approach to address tumor heterogeneity. Bioconjug. Chem. 2019, 30, 1314–1322. [Google Scholar] [CrossRef] [PubMed]
| PSMA | GRP-R | NTS1 | ||||
|---|---|---|---|---|---|---|
| Patient | ISUP Score | IRS | SUVmax | IRS | SUVmax | IRS |
| 1 | 1 | 6 | 2.8 | 6 | 4.8 | 6 |
| 2 | 2 | 9 | 4.5 | 3 | 5.1 | 6 |
| 3 | 2 | 9 | 4.7 | 8 | 6.3 | 0 |
| 4 | 2 | 0 | 5 | 4 | 5.3 | 12 |
| 5 | 2 | 1 | 3.4 | 1 | 7.5 | 12 |
| 6 | 3 | 12 | 6.8 | 4 | 8.3 | 1 |
| 7 | 3 | 2 | 3.6 | 8 | 8.9 | 12 |
| 8 | 4 | 9 | 2.8 | 6 | 2.4 | 3 |
| 9 | 4 | 9 | 8.5 | 6 | 2.8 | 2 |
| 10 | 5 | 12 | 13.3 | 4 | 7.5 | 2 |
| 11 | 5 | 12 | 5.9 | 4 | 7.2 | 4 |
| 12 | 5 | 12 | 12.5 | 2 | 2.8 | 8 |
| 13 | 5 | 6 | 7.1 | 1 | 9.1 | 4 |
| 14 | 5 | 12 | 3.7 | 4 | 10.5 | 12 |
| 15 | 5 | 12 | 7.8 | 4 | 9 | 6 |
| 16 | 5 | 12 | 20.4 | 2 | 3.7 | 1 |
| Patient | Age | ISUP | Gleason Score | PSA (ng/mL) | Clinical Tumoral Size: cT | Pathological Tumoral Size: pT | Metastatic Risk |
|---|---|---|---|---|---|---|---|
| 1 | 67 | 1 | 6 (3 + 3) | 3.7 | 1 | 2c | High |
| 2 | 65 | 1 | 6 (3 + 3) | 5.26 | 1 | 2c | High |
| 3 | 57 | 1 | 6 (3 + 3) | 4.38 | 1 | 2a | Low |
| 4 | 51 | 1 | 6 (3 + 3) | 3.7 | 2 | 2a | Low |
| 5 | 63 | 1 | 6 (3 + 3) | 10 | 1 | 2c | High |
| 6 | 48 | 1 | 6 (3 + 3) | 4.51 | 1 | 2c | High |
| 7 | 56 | 1 | 6 (3 + 3) | 4.4 | 2 | 2c | High |
| 8 | 55 | 1 | 6 (3 + 3) | 3.7 | 2 | 2c | High |
| 9 | 70 | 2 | 7 (3 + 4) | 10.5 | 1 | 3a | High |
| 10 | 67 | 2 | 7 (3 + 4) | 5.65 | 2 | 2c | High |
| 11 | 57 | 2 | 7 (3 + 4) | 6 | 1 | 3a | High |
| 12 | 66 | 2 | 7 (3 + 4) | 10 | 2 | 2c | High |
| 13 | 59 | 2 | 7 (3 + 4) | 13 | 2 | 2b | Intermediate |
| 14 | 66 | 2 | 7 (3 + 4) | 14 | 2 | 2c | High |
| 15 | 67 | 2 | 7 (3 + 4) | 14 | 1 | 3a | High |
| 16 | 66 | 2 | 7 (3 + 4) | 10.4 | 0 | 3a | High |
| 17 | 67 | 2 | 7 (3 + 4) | 12.5 | 1 | 3a | High |
| 18 | 55 | 2 | 7 (3 + 4) | 13 | 1 | 3a | High |
| 19 | 49 | 2 | 7 (3 + 4) | 14.28 | 2 | 3b | High |
| 20 | 64 | 3 | 7 (4 + 3) | 8 | 1 | 3a | High |
| 21 | 60 | 3 | 7 (4 + 3) | 5.67 | 1 | 3b | High |
| 22 | 66 | 3 | 7 (4 + 3) | 4.28 | 2 | 3a | High |
| 23 | 58 | 3 | 7 (4 + 3) | 7.6 | 2 | 3a | High |
| 24 | 71 | 3 | 7 (4 + 3) | 6.4 | 2 | 3a | High |
| 25 | 67 | 3 | 7 (4 + 3) | 7.6 | 2 | 2c | High |
| 26 | 63 | 3 | 7 (4 + 3) | 28 | 2 | nd | High |
| 27 | 63 | 3 | 7 (4 + 3) | 25.6 | 3 | 3b | High |
| 28 | 68 | 3 | 7 (4 + 3) | 19 | 2 | 3a | High |
| 29 | 53 | 3 | 7 (4 + 3) | 20 | 2 | 3a | High |
| 30 | 75 | 4 | 8 (4 + 4) | 6 | 3 | 1b | High |
| 31 * | 71 | 4 | 8 (4 + 4) | 285 | 4 | nd | High |
| 32 | 63 | 4 | 8 (4 + 4) | 7 | 2 | 3a | High |
| 33 | 70 | 4 | 8 (4 + 4) | 3.9 | 2 | 3a | High |
| 34 | 70 | 4 | 8 (4 + 4) | 9.95 | 1 | 2c | High |
| 35 | 74 | 4 | 8 (5 + 3) | nd | nd | nd | High |
| 36 | 66 | 4 | 8 (4 + 4) | 44 | 2 | 3a | High |
| 37 | 59 | 4 | 8 (4 + 4) | 14 | 2 | 4 | High |
| 38 | 73 | 5 | 9 (4 + 5) | 10 | nd | 3b | High |
| 39 | 72 | 5 | 9 (4 + 5) | 20 | 3 | 3b | High |
| 40 | 63 | 5 | 9 (4 + 5) | 27 | 3 | 3b | High |
| 41 | 54 | 5 | 9 (4 + 5) | 30 | 3 | 3a | High |
| 42 | 60 | 5 | 9 (4 + 5) | 12.6 | 2 | 3a | High |
| 43 | 66 | 5 | 9 (4 + 5) | 4.4 | 2 | 2a | High |
| 44 | 63 | 5 | 9 (5 + 4) | 5 | 2 | 3a | High |
| 45 | 70 | 5 | 9 (4 + 5) | 24.5 | 2 | 3b | High |
| 46 | 56 | 5 | 9 (4 + 5) | 26 | 3 | 3a | High |
| ISUP | PSMA | GRP-R | NTS1 | SST2 | NTS2 | CXCR4 |
|---|---|---|---|---|---|---|
| 1 |  | |||||
| 78.8% ±10.0 (8) | 44.7% ±51.3 (8) | 38.3% ±33.6 (6) | 43.5% ±33.0 (8) | 29.4% ±37.5 (5) | 62.4% ±21.0 (4) | |
| 2 |  | |||||
| 81.0% ±15.4 (11) | 8.2% ±18.3 (11) | 10.0% ±20.0 (4) | 34.6% ±28.6 (9) | 37.1% ±29.6 (7) | 36.4% ±3.1 (2) | |
| 3 |  | |||||
| 89.0% ±9.8 (8) | 27.5% ±29.7 (9) | 64.7% ±68.7 (6) | 77.6% ±147 (9) | 36.4% ±38.9 (9) | 0% ±0 (0) | |
| 4 |  | |||||
| 94.7% ±4.6 (8) | 16.2% ±27.8 (8) | 39.1% ±47.2 (7) | 7.4% ±10.0 (3) | 37.3% ±32.6 (6) | 32.0% ±45.2 (2) | |
| 5 |  | |||||
| 73.6% ±25.7 (8) | 20.8% ±39.4 (8) | 13.3% ±23.0 (7) | 32.0% ±28.7 (9) | 54.6% ±35.6 (6) | 43.9% ±73.3 (4) | |
| Total |  | |||||
| 83.3% ±16.0 (43) | 22.5% ±34.7 (44) | 34.2% ±44.8 (30) | 43.9% ±75.3 (38) | 39.0% ±33.8 (33) | 46.8% ±43.9 (12) | |
| ISUP | GRP-R | NTS1 | SST2 | NTS2 | CXCR4 |
|---|---|---|---|---|---|
| 1 | 1 | 1 | 2 | 1 | 2 |
| 2 | 0 | 0 | 0 | 1 | 0 |
| 3 | 0 | 2 | 1 | 1 | 0 |
| 4 | 0 | 3 | 0 | 0 | 0 |
| 5 | 1 | 0 | 0 | 1 | 1 |
| Total | 2 | 6 | 3 | 4 | 3 |
| ISUP | PSMA | NTS1 | SST2 | NTS2 | CXCR4 |
|---|---|---|---|---|---|
| 1 | 7 | 2 | 4 | 2 | 4 |
| 2 | 11 | 1 | 6 | 5 | 2 |
| 3 | 8 | 3 | 3 | 5 | NA |
| 4 | 8 | 2 | 2 | 5 | NA |
| 5 | 9 | 0 | 4 | 3 | 2 |
| Total | 43 | 8 | 19 | 20 | 8 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Schollhammer, R.; Quintyn Ranty, M.-L.; de Clermont Gallerande, H.; Cavelier, F.; Valverde, I.E.; Vimont, D.; Hindié, E.; Morgat, C. Theranostics of Primary Prostate Cancer: Beyond PSMA and GRP-R. Cancers 2023, 15, 2345. https://doi.org/10.3390/cancers15082345
Schollhammer R, Quintyn Ranty M-L, de Clermont Gallerande H, Cavelier F, Valverde IE, Vimont D, Hindié E, Morgat C. Theranostics of Primary Prostate Cancer: Beyond PSMA and GRP-R. Cancers. 2023; 15(8):2345. https://doi.org/10.3390/cancers15082345
Chicago/Turabian StyleSchollhammer, Romain, Marie-Laure Quintyn Ranty, Henri de Clermont Gallerande, Florine Cavelier, Ibai E. Valverde, Delphine Vimont, Elif Hindié, and Clément Morgat. 2023. "Theranostics of Primary Prostate Cancer: Beyond PSMA and GRP-R" Cancers 15, no. 8: 2345. https://doi.org/10.3390/cancers15082345
APA StyleSchollhammer, R., Quintyn Ranty, M.-L., de Clermont Gallerande, H., Cavelier, F., Valverde, I. E., Vimont, D., Hindié, E., & Morgat, C. (2023). Theranostics of Primary Prostate Cancer: Beyond PSMA and GRP-R. Cancers, 15(8), 2345. https://doi.org/10.3390/cancers15082345








