Tumor BRCA Testing in Epithelial Ovarian Cancers: Past and Future—Five-Years’ Single-Institution Experience of 762 Consecutive Patients
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Population
2.2. Next Generation Sequencing-Based BRCA Tumor Testing
2.3. Multiplex Ligation-Dependent Probe Amplification (MLPA)
2.4. Statistical Analysis
3. Results
3.1. Tumor BRCA Testing Results
3.2. Tumor BRCA1/2 Status According to Clinicopathological Characteristics
3.3. The “New Workflow” of Tumor BRCA Testing
3.4. Comparison between Tumor and Blood BRCA Test Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- AstraZeneca. Lynparza Summary of Product Characteristics. 2014. Available online: http://www.ema.europa.eu/docs/en_GB/document_library/EPAR_Product_Information/human/003726/WC500180151.pdf (accessed on 25 January 2022).
- Tesaro. Zejula Summary of Product Characteristics. 2017. Available online: http://www.ema.europa.eu/docs/en_GB/document_library/EPAR_Product_Information/human/004249/WC500239289.pdf (accessed on 25 January 2022).
- Clovis Oncology UK. Rubraca Summary of Product Characteristics. 2018. Available online: https://www.ema.europa.eu/documents/product-information/rubraca-epar-product-information_en.pdf (accessed on 25 January 2022).
- Tan, D.S.P.; Rothermundt, C.; Thomas, K.; Bancroft, E.; Eeles, R.; Shanley, S.; Ardern-Jones, A.; Norman, A.; Kaye, S.B.; Gore, M.E. “BRCAness” syndrome in ovarian cancer: A case-control study describing the clinical features and outcome of patients with epithelial ovarian cancer associated with BRCA1 and BRCA2 mutations. J. Clin. Oncol. 2008, 26, 5530–5536. [Google Scholar] [CrossRef] [PubMed]
- Pujade-Lauraine, E.; Ledermann, J.A.; Selle, F.; Gebski, V.; Penson, R.T.; Oza, A.M.; Korach, J.; Huzarski, T.; Poveda, A.; Pignata, S.; et al. Olaparib tablets as maintenance therapy in patients with platinum-sensitive, relapsed ovarian cancer and a BRCA1/2 mutation (SOLO2/ENGOT-Ov21): A double-blind, randomised, placebo-controlled, phase 3 trial. Lancet Oncol. 2017, 18, 1274–1284. [Google Scholar] [CrossRef] [Green Version]
- Oaknin, A.; Oza, A.M.; Lorusso, D.; Aghajanian, C.; Dean, A.; Colombo, N.; Weberpals, J.I.; Clamp, A.R.; Scambia, G.; Leary, A.; et al. Maintenance treatment with rucaparib for recurrent ovarian carcinoma in ARIEL3, a randomized phase 3 trial: The effects of best response to last platinum-based regimen and disease at baseline on efficacy and safety. Cancer Med. 2021, 10, 7162–7173. [Google Scholar] [CrossRef] [PubMed]
- Mirza, M.R.; Monk, B.J.; Herrstedt, J.; Oza, A.M.; Mahner, S.; Redondo, A.; Fabbro, M.; Ledermann, J.A.; Lorusso, D.; Vergote, I.; et al. Niraparib Maintenance Therapy in Platinum-Sensitive, Recurrent Ovarian Cancer. N. Engl. J. Med. 2016, 375, 2154–2164. [Google Scholar] [CrossRef]
- Banerjee, S.; Moore, K.N.; Colombo, N.; Scambia, G.; Kim, B.G.; Oaknin, A.; Friedlander, M.; Lisyanskaya, A.; Floquet, A.; Leary, A.; et al. Maintenance olaparib for patients with newly diagnosed advanced ovarian cancer and a BRCA mutation (SOLO1/GOG 3004): 5-year follow-up of a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet Oncol. 2021, 21, S1470–S2045. [Google Scholar] [CrossRef]
- González-Martín, A.; Pothuri, B.; Vergote, I.; DePont Christensen, R.; Graybill, W.; Mirza, M.R.; McCormick, C.; Lorusso, D.; Hoskins, P.; Freyer, G.; et al. Niraparib in Patients with Newly Diagnosed Advanced Ovarian Cancer. N. Engl. J. Med. 2019, 381, 2391–2402. [Google Scholar] [CrossRef] [Green Version]
- Coleman, R.L.; Fleming, G.F.; Brady, M.F.; Swisher, E.M.; Steffensen, K.D.; Friedlander, M.; Okamoto, A.; Moore, K.N.; Efrat Ben-Baruch, N.; Werner, T.L.; et al. Veliparib with First-Line Chemotherapy and as Maintenance Therapy in Ovarian Cancer. N. Engl. J. Med. 2019, 381, 2403–2415. [Google Scholar] [CrossRef]
- Konstantinopoulos, P.A.; Norquist, B.; Lacchetti, C.; Armstrong, D.; Grisham, R.N.; Goodfellow, P.J.; Kohn, E.C.; Levine, D.A.; Liu, J.F.; Lu, K.H.; et al. Germline and Somatic Tumor Testing in Epithelial Ovarian Cancer: ASCO Guideline. J. Clin. Oncol. 2020, 38, 1222–1245. [Google Scholar] [CrossRef]
- NCCN. 2020. Available online: https://crain-platform-genomeweb-prod.s3.amazonaws.com/s3fs-public/files_copied/nccn_nov_2020_brca_guidelines.pdf (accessed on 25 January 2022).
- Colombo, N.; Sessa, C.; du Bois, A.; Ledermann, J.; McCluggage, W.G.; McNeish, I.; Morice, P.; Pignata, S.; Ray-Coquard, I.; Vergote, I.; et al. ESMO-ESGO consensus conference recommendations on ovarian cancer: Pathology and molecular biology, early and advanced stages, borderline tumours and recurrent disease†. Ann. Oncol. 2019, 30, 672–705. [Google Scholar] [CrossRef] [Green Version]
- Pinto, C.; Bella, M.A.; Capoluongo, E.; Carrera, P.; Clemente, C.; Colombo, N.; Cortesi, L.; De Rosa, G.; Fenizia, F.; Genuardi, M.; et al. Recommendations for the implementation of BRCA testing in the care and treatment pathways of ovarian cancer patients. Future Oncol. 2016, 18, 2071–2075. [Google Scholar] [CrossRef] [Green Version]
- Capoluongo, E.; Ellison, G.; López-Guerrero, J.A.; Penault-Llorca, F.; Ligtenberg, M.J.L.; Banerjee, S.; Singer, C.; Friedman, E.; Markiefka, B.; Schirmacher, P.; et al. Guidance Statement on BRCA1/2 Tumor Testing in Ovarian Cancer Patients. Semin. Oncol. 2017, 44, 187–197. [Google Scholar] [CrossRef]
- Gori, S.; Barberis, M.; Bella, M.A.; Buttitta, F.; Capoluongo, E.; Carrera, P.; Colombo, N.; Cortesi, L.; Genuardi, M.; Gion, M.; et al. Recommendations for the implementation of BRCA testing in ovarian cancer patients and their relatives. Crit. Rev. Oncol. Hematol. 2019, 140, 67–72. [Google Scholar] [CrossRef]
- Guidelines for BRCA Test Implementation 2021. Raccomandazioni per L’Implementazione del Test BRCA Predittivo e Preventivo nei Tumori della Mammella, Dell’Ovaio, del Pancreas e della Prostata. A Cura del Panel Multidisciplinare Intersocietario: AIOM, AIRO, AISP, ANISC, AURO, Fondazione AIOM, SIAPEC-IAP, SIBIOC, SICO, SIF, SIGE, SIGU, SIU, SIURO, UROP, aBRCAdabra Onlus: Maggio 2021. Available online: https://www.aiom.it/wp-content/uploads/2021/07/2021_Racc_testBRCA_predittivo-preventivo.pdf (accessed on 25 January 2022).
- Palacios, J.; de la Hoya, M.; Bellosillo, B.; de Juan, I.; Matías-Guiu, X.; Lázaro, C.; Palanca, S.; Osorio, A.; Rojo, F.; Rosa-Rosa, J.M.; et al. Mutational Screening of BRCA1/2 Genes as a Predictive Factor for Therapeutic Response in Epithelial Ovarian Cancer: A Consensus Guide from the Spanish Society of Pathology (SEAP-IAP) and the Spanish Society of Human Genetics (AEGH). Virchows Arch. 2020, 476, 195–207. [Google Scholar] [CrossRef] [Green Version]
- Rivera, D.; Paudice, M.; Gismondi, V.; Anselmi, G.; Vellone, V.G.; Varesco, L.; Ligurian BRCA Working Group. Implementing NGS-based BRCA tumour tissue testing in FFPE ovarian carcinoma specimens: Hints from a real-life experience within the framework of expert recommendations. J. Clin. Pathol. 2021, 74, 596–603. [Google Scholar] [CrossRef]
- Fumagalli, C.; Tomao, F.; Betella, I.; Rappa, A.; Calvello, M.; Bonanni, B.; Bernard, L.; Peccatori, F.; Colombo, N.; Viale, G.; et al. Tumor BRCA Test for Patients with Epithelial Ovarian Cancer: The Role of Molecular Pathology in the Era of PARP Inhibitor Therapy. Cancers 2019, 11, 1641. [Google Scholar] [CrossRef] [Green Version]
- Akaev, I.; Rahimi, S.; Onifade, O.; Gardner, F.J.E.; Castells-Rufas, D.; Jones, E.; Acharige, S.; Yeoh, C.C. Tumour Versus Germline BRCA Testing in Ovarian Cancer: A Single-Site Institution Experience in the United Kingdom. Diagnostics 2021, 11, 547. [Google Scholar] [CrossRef]
- Vos, J.R.; Fakkert, I.E.; de Hullu, J.A.; van Altena, A.M.; Sie, A.S.; Ouchene, H.; Willems, R.W.; Nagtegaal, I.D.; Jongmans, M.C.J.; Mensenkamp, A.R.; et al. Universal Tumor DNA BRCA1/2 Testing of Ovarian Cancer: Prescreening PARPi Treatment and Genetic Predisposition. J. Natl. Cancer Inst. 2020, 112, 161–169. [Google Scholar] [CrossRef]
- Marchetti, C.; Minucci, A.; D’Indinosante, M.; Ergasti, R.; Arcieri, M.; Capoluongo, E.D.; Pietragalla, A.; Caricato, C.; Scambia, G.; Fagotti, A. Feasibility of tumor testing for BRCA status in high-grade serous ovarian cancer using fresh-frozen tissue based approach. Gynecol. Oncol. 2020, 158, 740–746. [Google Scholar] [CrossRef]
- FIGO Staging. 2014. Available online: https://www.sgo.org/wp-content/uploads/2012/09/FIGO-Ovarian-Cancer-Staging_1.10.14.pdf (accessed on 25 January 2022).
- ENIGMA BRCA1/2 Gene Variant Classification Criteria. Available online: https://enigmaconsortium.org/wp-content/uploads/2020/08/ENIGMA_Rules_2017-06-29-v2_5_1.pdf (accessed on 25 January 2022).
- BRCA Exchange Database. Available online: https://brcaexchange.org/ (accessed on 25 January 2022).
- ClinVar Database. Available online: https://www.ncbi.nlm.nih.gov/clinvar/ (accessed on 25 January 2022).
- Leiden Open Source Variation Database. Available online: https://databases.lovd.nl/shared/variants/BRCA1/unique (accessed on 25 January 2022).
- Turchiano, A.; Loconte, D.C.; De Nola, R.; Arezzo, F.; Chiarello, G.; Pantaleo, A.; Iacoviello, M.; Bagnulo, R.; De Luisi, A.; Perrelli, S.; et al. Beyond BRCA1/2: Homologous Recombination Repair Genetic Profile in a Large Cohort of Apulian Ovarian Cancers. Cancers 2022, 14, 365. [Google Scholar] [CrossRef]
- Fumagalli, C.; Guerini-Rocco, E.; Buttitta, F.; Iapicca, P.; You, W.; Mauri, M.; Felicioni, L.; Troncone, G.; Malapelle, U.; Scarpa, A.; et al. Reliability and reproducibility among different platforms for tumour BRCA testing in ovarian cancer: A study of the Italian NGS Network. J. Clin. Pathol. 2020, 74, 668–672. [Google Scholar] [CrossRef]
- Hodgson, D.R.; Brown, J.S.; Dearden, S.P.; Lai, Z.; Elks, C.E.; Milenkova, T.; Dougherty, B.A.; Lanchbury, J.S.; Perry, M.; Timms, K.M.; et al. Concordance of BRCA mutation detection in tumor versus blood, and frequency of bi-allelic loss of BRCA in tumors from patients in the phase III SOLO2 trial. Gynecol. Oncol. 2021, 163, 563–568. [Google Scholar] [CrossRef] [PubMed]
- Callens, C.; Vaur, D.; Soubeyran, I.; Rouleau, E.; Just, P.A.; Guillerm, E.; Golmard, L.; Goardon, N.; Sevenet, N.; Cabaret, O.; et al. Concordance Between Tumor and Germline BRCA Status in High-Grade Ovarian Carcinoma Patients in the Phase III PAOLA-1/ENGOT-ov25 Trial. J. Natl. Cancer Inst. 2021, 113, 917–923. [Google Scholar] [CrossRef] [PubMed]
- Soegaard, M.; Kjaer, S.K.; Cox, M.; Wozniak, E.; Høgdall, E.; Høgdall, C.; Blaakaer, J.; Jacobs, I.J.; Gayther, S.A.; Ramus, S.J. BRCA1 and BRCA2 mutation prevalence and clinical characteristics of a population-based series of ovarian cancer cases from Denmark. Clin. Cancer Res. 2008, 14, 3761–3767. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moslehi, R.; Chu, W.; Karlan, B.; Fishman, D.; Risch, H.; Fields, A.; Smotkin, D.; Ben-David, Y.; Rosenblatt, J.; Russo, D.; et al. BRCA1 and BRCA2 mutation analysis of 208 Ashkenazi Jewish women with ovarian cancer. Am. J. Hum. Genet. 2000, 66, 1259–1272. [Google Scholar] [CrossRef] [Green Version]
- Loizzi, V.; Cicinelli, E.; Santamaria, F.; Murgia, F.; Minicucci, V.; Resta, L.; Resta, N.; Natalicchio, M.I.; Ranieri, G.; Cormio, G. BRCAmut and “founder effect”: A prospective study in a single academic institution. Oncotarget 2018, 9, 22353–22358. [Google Scholar] [CrossRef] [Green Version]
- Pennington, K.P.; Walsh, T.; Harrell, M.I.; Lee, M.K.; Pennil, C.C.; Rendi, M.H.; Thornton, A.; Norquist, B.M.; Casadei, S.; Nord, A.S.; et al. Germline and somatic mutations in homologous recombination genes predict platinum response and survival in ovarian, fallopian tube, and peritoneal carcinomas. Clin. Cancer Res. 2014, 20, 764–775. [Google Scholar] [CrossRef] [Green Version]
- Hennessy, B.T.; Timms, K.M.; Carey, M.S.; Gutin, A.; Meyer, L.A.; Flake, D.D., 2nd; Abkevich, V.; Potter, J.; Pruss, D.; Glenn, P.; et al. Somatic mutations in BRCA1 and BRCA2 could expand the number of patients that benefit from poly (ADP ribose) polymerase inhibitors in ovarian cancer. J. Clin. Oncol. 2010, 28, 3570–3576. [Google Scholar] [CrossRef] [Green Version]
- Thompson, D.; Easton, D.; Breast Cancer Linkage Consortium. Variation in BRCA1 cancer risks by mutation position. Cancer Epidemiol. Biomark. Prev. 2002, 11, 329–336. [Google Scholar]
- Gayther, S.A.; Mangion, J.; Russell, P.; Seal, S.; Barfoot, R.; Ponder, B.A.; Stratton, M.R.; Easton, D. Variation of risks of breast and ovarian cancer associated with different germline mutations of the BRCA2 gene. Nat. Genet. 1997, 15, 103–105. [Google Scholar] [CrossRef]
- Rebbeck, T.R.; Mitra, N.; Wan, F.; Sinilnikova, O.M.; Healey, S.; McGuffog, L.; Mazoyer, S.; Chenevix-Trench, G.; Easton, D.F.; Antoniou, A.C.; et al. Association of type and location of BRCA1 and BRCA2 mutations with risk of breast and ovarian cancer. JAMA 2015, 313, 1347–1361. [Google Scholar] [CrossRef] [Green Version]
- Solsky, I.; Chen, J.; Rebbeck, T.R. Precision prophylaxis: Identifying the optimal timing for risk-reducing salpingo-oophorectomy based on type of BRCA1 and BRCA2 cluster region mutations. Gynecol. Oncol. 2020, 156, 363–376. [Google Scholar] [CrossRef]
- Hollis, R.L.; Churchman, M.; Gourley, C. Distinct implications of different BRCA mutations: Efficacy of cytotoxic chemotherapy, PARP inhibition and clinical outcome in ovarian cancer. OncoTargets Ther. 2017, 10, 2539–2551. [Google Scholar] [CrossRef] [Green Version]
- Dimitrova, D.; Ruscito, I.; Olek, S.; Richter, R.; Hellwag, A.; Türbachova, I.; Woopen, H.; Baron, U.; Braicu, E.I.; Sehouli, J. Germline mutations of BRCA1 gene exon 11 are not associated with platinum response neither with survival advantage in patients with primary ovarian cancer: Understanding the clinical importance of one of the biggest human exons. A study of the Tumor Bank Ovarian Cancer (TOC) Consortium. Tumor Biol. 2016, 37, 12329–12337. [Google Scholar] [CrossRef]
- Wang, Y.; Bernhardy, A.J.; Cruz, C.; Krais, J.J.; Nacson, J.; Nicolas, E.; Peri, S.; van der Gulden, H.; van der Heijden, I.; O’Brien, S.W.; et al. The BRCA1-Δ11q Alternative Splice Isoform Bypasses Germline Mutations and Promotes Therapeutic Resistance to PARP Inhibition and Cisplatin. Cancer Res. 2016, 76, 2778–2790. [Google Scholar] [CrossRef] [Green Version]
- Fraile-Bethencourt, E.; Díez-Gómez, B.; Velásquez-Zapata, V.; Acedo, A.; Sanz, D.J.; Velasco, E.A. Functional classification of DNA variants by hybrid minigenes: Identification of 30 spliceogenic variants of BRCA2 exons 17 and 18. PLoS Genet. 2017, 13, e1006691. [Google Scholar] [CrossRef]
- Jimenez-Sainz, J.; Jensen, R.B. Imprecise Medicine: BRCA2 Variants of Uncertain Significance (VUS), the Challenges and Benefits to Integrate a Functional Assay Workflow with Clinical Decision Rules. Genes 2021, 12, 780. [Google Scholar] [CrossRef]
- Yadegari, F.; Majidzadeh, K. In silico analysis for determining the deleterious nonsynonymous single nucleotide polymorphisms of BRCA genes. Mol. Biol. Res. Commun. 2019, 8, 141–150. [Google Scholar] [CrossRef]
- Hart, S.N.; Hoskin, T.; Shimelis, H.; Moore, R.M.; Feng, B.; Thomas, A.; Lindor, N.M.; Polley, E.C.; Goldgar, D.E.; Iversen, E.; et al. Comprehensive annotation of BRCA1 and BRCA2 missense variants by functionally validated sequence-based computational prediction models. Genet. Med. 2019, 21, 71–80. [Google Scholar] [CrossRef]
- Dines, J.N.; Shirts, B.H.; Slavin, T.P.; Walsh, T.; King, M.C.; Fowler, D.M.; Pritchard, C.C. Systematic misclassification of missense variants in BRCA1 and BRCA2 “coldspots”. Genet. Med. 2020, 22, 825–830. [Google Scholar] [CrossRef] [Green Version]
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; Grody, W.W.; Hegde, M.; Lyon, E.; Spector, E.; et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 2015, 17, 405–424. [Google Scholar] [CrossRef] [Green Version]
- Witjes, V.M.; van Bommel, M.H.D.; Ligtenberg, M.J.L.; Vos, J.R.; Mourits, M.J.E.; Ausems, M.G.E.M.; de Hullu, J.A.; Bosse, T.; Hoogerbrugge, N. Probability of detecting germline BRCA1/2 pathogenic variants in histological subtypes of ovarian carcinoma. A meta-analysis. Gynecol. Oncol. 2021, 64, 221–230. [Google Scholar] [CrossRef]
- Hauke, J.; Hahnen, E.; Schneider, S.; Reuss, A.; Richters, L.; Kommoss, S.; Heimbach, A.; Marmé, F.; Schmidt, S.; Prieske, K.; et al. Deleterious somatic variants in 473 consecutive individuals with ovarian cancer: Results of the observational AGO-TR1 study (NCT02222883). J. Med. Genet. 2019, 56, 574–580. [Google Scholar] [CrossRef]
- Fuh, K.; Mullen, M.; Blachut, B.; Stover, E.; Konstantinopoulos, P.; Liu, J.; Matulonis, U.; Khabele, D.; Mosammaparast, N.; Vindigni, A. Homologous recombination deficiency real-time clinical assays, ready or not? Gynecol. Oncol. 2020, 159, 877–886. [Google Scholar] [CrossRef] [PubMed]
- Stover, E.H.; Fuh, K.; Konstantinopoulos, P.A.; Matulonis, U.A.; Liu, J.F. Clinical assays for assessment of homologous recombination DNA repair deficiency. Gynecol. Oncol. 2020, 159, 887–898. [Google Scholar] [CrossRef]
- Miller, R.E.; Leary, A.; Scott, C.L.; Serra, V.; Lord, C.J.; Bowtell, D.; Chang, D.K.; Garsed, D.W.; Jonkers, J.; Ledermann, J.A.; et al. ESMO recommendations on predictive biomarker testing for homologous recombination deficiency and PARP inhibitor benefit in ovarian cancer. Ann. Oncol. 2020, 31, 1606–1622. [Google Scholar] [CrossRef]
- Castroviejo-Bermejo, M.; Cruz, C.; Llop-Guevara, A.; Gutiérrez-Enríquez, S.; Ducy, M.; Ibrahim, Y.H.; Gris-Oliver, A.; Pellegrino, B.; Bruna, A.; Guzmán, M.; et al. A RAD51 assay feasible in routine tumor samples calls PARP inhibitor response beyond BRCA mutation. EMBO Mol. Med. 2018, 10, e9172. [Google Scholar] [CrossRef]
- vanWijk, L.M.; Vermeulen, S.; Meijers, M.; van Diest, M.F.; Ter Haar, N.T.; de Jonge, M.M.; Solleveld-Westerink, N.; van Wezel, T.; van Gent, D.C.; Kroep, J.R.; et al. The RECAP Test Rapidly and Reliably Identifies Homologous Recombination-Deficient Ovarian Carcinomas. Cancers 2020, 12, 2805. [Google Scholar] [CrossRef]
- vanWijk, L.M.; Kramer, C.J.H.; Vermeulen, S.; Ter Haar, N.T.; de Jonge, M.M.; Kroep, J.R.; de Kroon, C.D.; Gaarenstroom, K.N.; Vrieling, H.; Bosse, T.; et al. The RAD51-FFPE Test; Calibration of a Functional Homologous Recombination Deficiency Test on Diagnostic Endometrial and Ovarian Tumor Blocks. Cancers 2021, 13, 2994. [Google Scholar] [CrossRef] [PubMed]
- Fumagalli, C.; Guerini-Rocco, E.; Rappa, A.; Vacirca, D.; Ranghiero, A.; Barberis, M. 09-P60. The First 500 Tumor BRCA Tests and Beyond: The Crucial Role of Molecular Pathology in the Clinical Management of Ovarian Cancer Patients. J. Mol. Diagn. 2020, 22, S1–S103. [Google Scholar] [CrossRef]
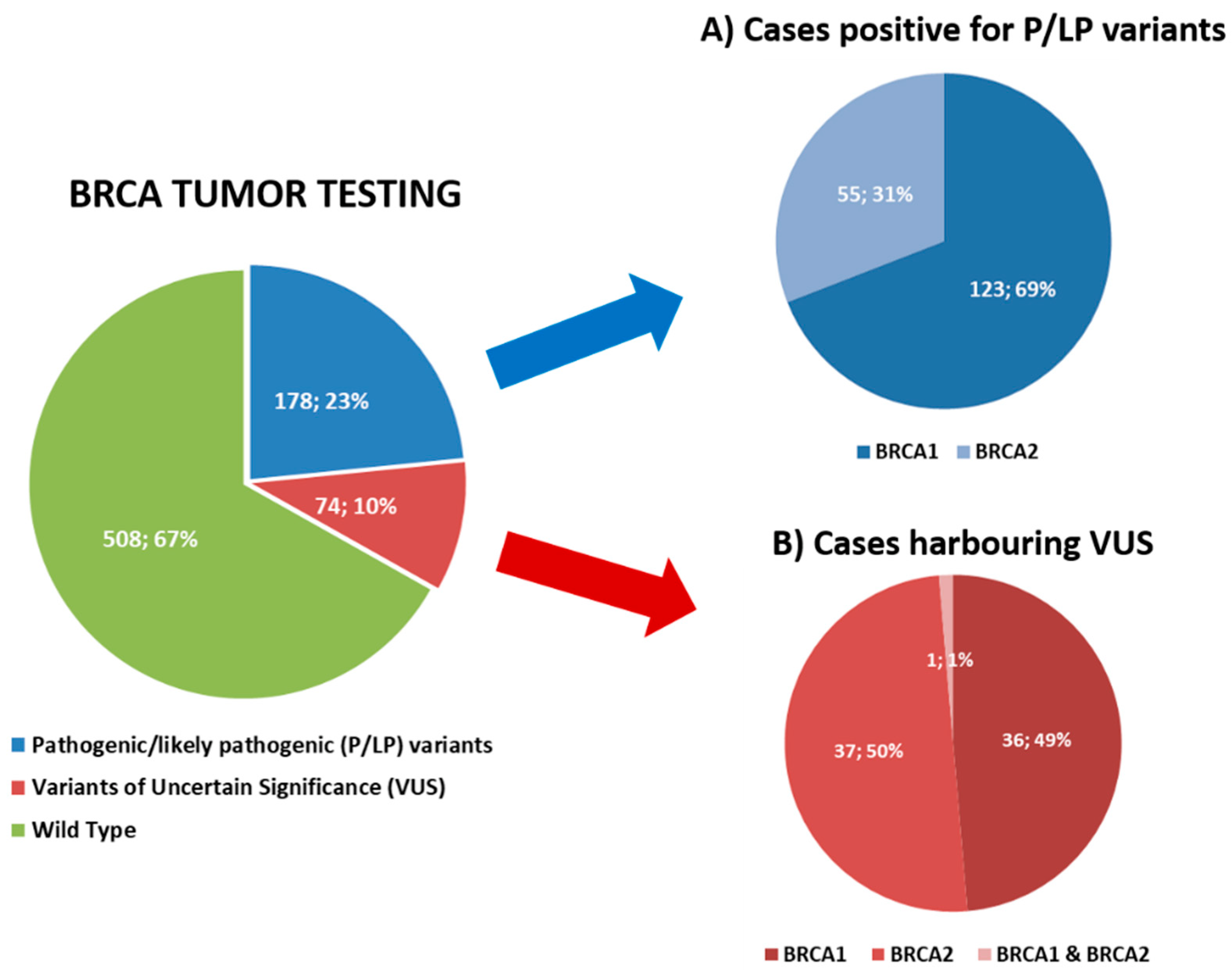
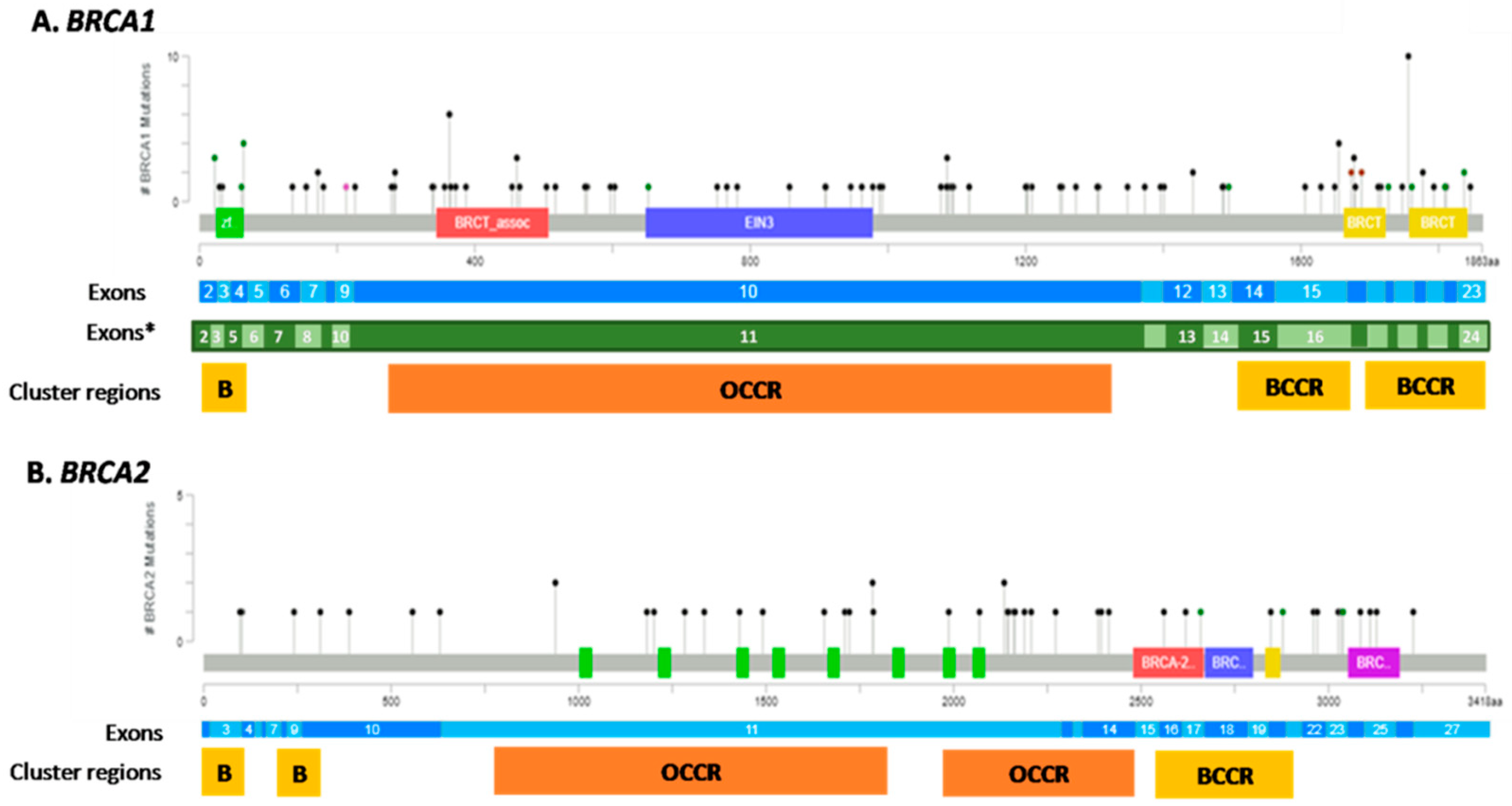
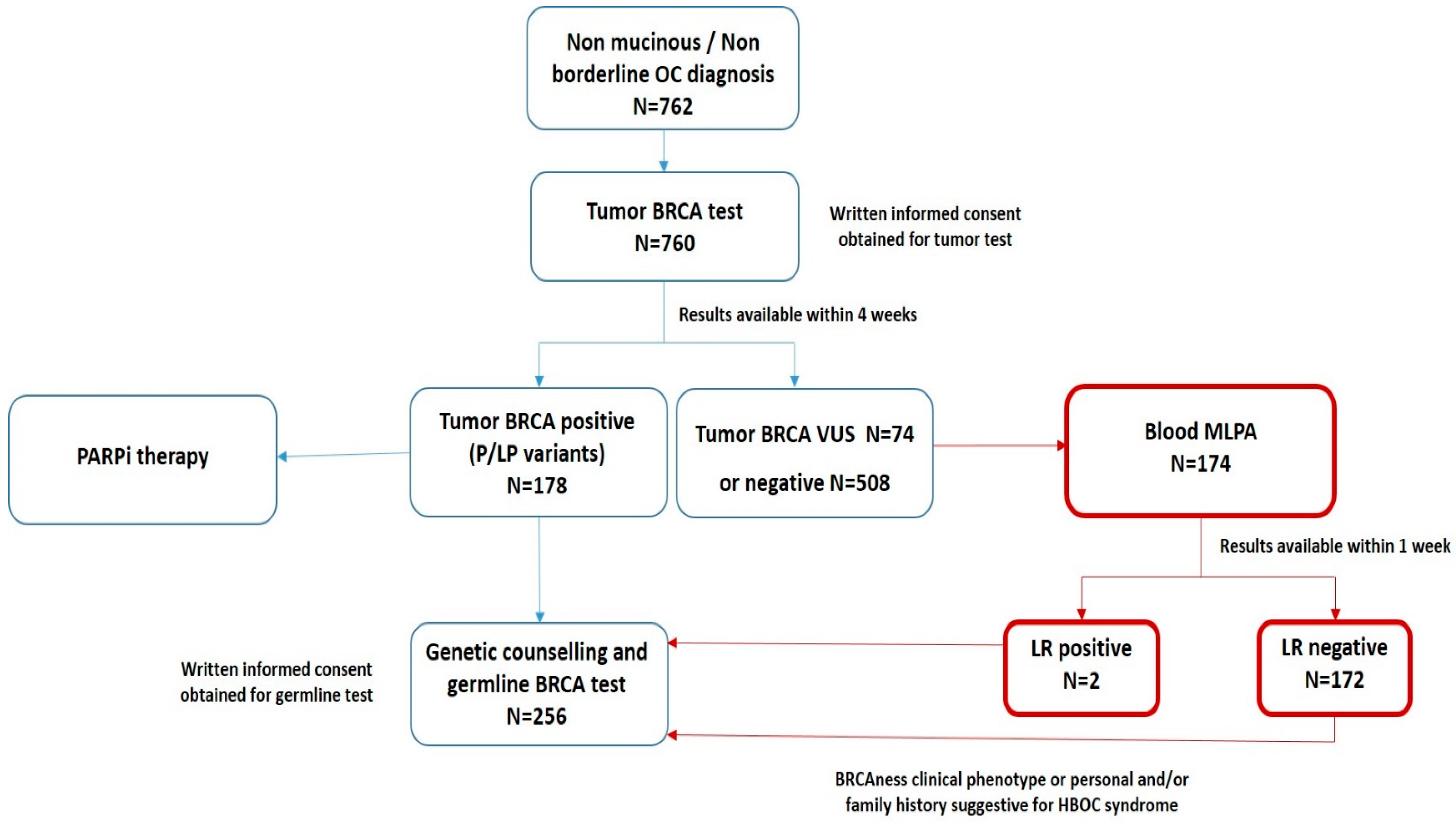
| Clinicopathological Characteristics | Study Cohort (n = 762) |
|---|---|
| Age (Median, range) | 61 (24–84) |
| Histological subtype | |
| High grade serous carcinoma | 636 (83.5%) |
| Low grade serous carcinoma | 17 (2.2%) |
| Clear cell carcinoma | 30 (3.9%) |
| Endometrioid carcinoma | 51 (6.7%) |
| Carcinoma—subtype not defined | 28 (3.7%) |
| Figo stage | |
| I (A-C) | 57 (7.5%) |
| II (A-B) | 61 (8%) |
| III (A-C) | 462 (60.6%) |
| IV (A-B) | 151 (19.8%) |
| NA | 31 (4.1%) |
| Tumor stage | |
| Early stage | 82 (10.8%) |
| Advanced stage | 649 (85.2%) |
| NA | 31 (4.1%) |
| Family history | |
| Positive | 427 (56%) |
| Negative | 240 (31.5%) |
| NA | 95 (12.5%) |
| P/LP (n = 179 *) BRCA1 (n = 124 *), BRCA2 (n = 55) | VUS (n = 75 **) BRCA1 (n = 37 **), BRCA2 (n = 38 **) | p-Value | |
|---|---|---|---|
| VAF | |||
| (Median, range) | 74% (5–99%) | 47% (5–98%) | <0.0001 # |
| BRCA1 | 76.5% (6–99%) | 44.5% (9–93%) | |
| BRCA2 | 63% (5–93%) | 48% (5–98%) | |
| MUTATION TYPE | |||
| (N, %) | <0.0001 # | ||
| Truncating | 147 (82.1%) | 4 (5.3%) | |
| BRCA1 | 101 (81.5%) | 2 (5.4%) | |
| BRCA2 | 46 (83.6%) | 2 (5.3%) | |
| Missense | 18 (10.1%) | 37 (49.3%) | |
| BRCA1 | 101 (81.5%) | 13 (35.1%) | |
| BRCA2 | 46 (83.6%) | 24 (63.2%) | |
| Splice site | 7 (3.9%) | 1 (1.3%) | |
| BRCA1 | 2 (1.6%) | 1 (2.7%) | |
| BRCA2 | 5 (9.1%) | 0 | |
| Intronic | 3 (1.7%) | 28 (37.3%) | |
| BRCA1 | 2 (1.6%) | 17 (45.9%) | |
| BRCA2 | 1 (1.8%) | 11 (28.9%) | |
| Inframe | 4 (2.2%) | 5 (6.7%) | |
| BRCA1 | 4 (3.2%) | 4 (10.8%) | |
| BRCA2 | 0 | 1 (2.6%) |
| P/LP (n = 178) | VUS (n = 74) | WT (n = 508) | p-Value | |
|---|---|---|---|---|
| Age | 59 (37–81) | 59 (39–79) | 62 (24–84) | |
| (Median, range) | ||||
| Histological subtype | 0.002 * | |||
| High grade serous carcinoma (n = 635) | 168 (26.5%;944%) | 60 (9.4%;81.1%) | 407 (64.1%;80.1%) | |
| Low grade serous carcinoma (n = 17) | 0 | 2 (11.8%;2.7%) | 15 (88.2%;3%) | |
| Clear cell carcinoma (n = 30) | 0 | 4 (13.3%;5.4%) | 26 (86.7%;5.1%) | |
| Endometrioid carcinoma (n = 50) | 6 (12%;3.4%) | 2 (4%;2.7%) | 42 (84%;8.3%) | |
| Carcinoma—subtype not defined (n = 28) | 4 (14.3%; 2.2%) | 6 (21.4%; 8.1%) | 18 (64.3%;3.5%) | |
| Tumor status | 0.79 | |||
| Early (n = 81) | 17 (21%; 9.6%) | 8 (9.9%; 10.8%) | 56 (69.1%; 11%) | |
| Advanced (n = 649) | 152 (23.4%; 85.4%) | 62 (9.6%; 83.8%) | 435 (67%; 85.6%) | |
| NA (n = 30) | 9 (30%; 5.1%) | 4 (13.3%; 5.4%) | 17 (56.6%; 3.3%) | |
| FIGO stage | 0.11 | |||
| I (A-C) (n = 56) | 11 (19.6%; 6.2%) | 3 (5.4%; 4.1%) | 42 (75%; 8.3%) | |
| II (A-C) (n = 61) | 7 (11.5%; 3.9%) | 9 (14.8%; 12.2%) | 45 (73.8%; 8.9%) | |
| III (A-C) (n = 462) | 115 (24.9%; 64.6%) | 38 (8.2%; 51.4%) | 309 (66.9%; 60.8%) | |
| IV (A-B) (n = 151) | 36 (23.8%; 20.2%) | 20 (13.2%; 27%) | 95 (62.9%; 18.7%) | |
| NA (n = 30) | 9 (30%; 5.1%) | 4 (13.3%; 5.4%) | 17 (56.7%; 3.3%) | |
| Family history | 0.007 * | |||
| Positive (n = 427) | 118 (27.6%; 66.3%) | 44 (10.3%; 59.5%) | 265 (62.1%; 52.2%) | |
| Negative (n = 239) | 49 (20.5%; 27.5%) | 22 (9.2%; 29.7%) | 168 (70.3%; 33.1%) | |
| NA (n = 94) | 11 (11.7%; 6.2%) | 8 (8.5%; 10.8%) | 75 (79.8%; 14.8%) |
| B | P/LP (n = 100) | VUS (n = 6) | WT (n = 150) | |
|---|---|---|---|---|
| T | ||||
| P/LP (n = 132) | ||||
| 100 (76%) # | - | 32 (24%) | ||
| VUS (n = 16) | - | 6 (37.5%) # | 10 (62.5%) | |
| WT (n = 108) | - | - | 108 (100%) # | |
| SOMATIC P/LP (n = 32; 24%) | GERMLINE P/LP (n = 100; 76%) | p-Value | |
|---|---|---|---|
| Age (Median, range) | 62.5 (40–79) | 56 (37–78) | 0.03 * |
| Histological subtype | |||
| High grade serous carcinoma (n = 125) | 28 (22.4%; 87.5%) | 97 (77.6%; 97%) | 0.04 * |
| Endometrioid carcinoma (n = 6) | 4 (66.7%; 12.5%) | 2 (33.3%; 2%) | |
| Carcinoma—subtype not defined (n = 1) | 0 | 1 (100%; 1%) | |
| FIGO stage | |||
| I (A-C) (n = 10) | 4 (40%;12.5%) | 6 (60%; 6%) | 0.05 * |
| II (A-B) (n = 12) | 7 (58.3%; 21.9%) | 5 (41.7%; 5%) | |
| III (A-C) (n = 82) | 12 (14.6%; 37.5%) | 70 (85.4%; 70%) | |
| IV (A-B) (n = 27) | 9 (33.3%; 28.1%) | 18 (66.7%; 18%) | |
| NA (n = 1) | 0 | 1 (100%; 1%) | |
| Tumor status | 0.2 | ||
| Early (n = 16) | 6 (37.5%; 18.8%) | 10 (62.5%; 10%) | |
| Advanced (n = 115) | 26 (22.6%; 81.3%) | 89 (77.4%; 89%) | |
| NA (n = 1) | 0 | 1 (100%; 1%) | |
| Family history | |||
| Positive (n = 95) | 17 (17.9%; 53.1%) | 78 (82.1%; 78%) | 0.01 * |
| Negative (n = 34) | 15 (44%; 46.9%) | 19 (56%; 19%) | |
| NA (n = 3) | 0 (0%) | 3 (100%; 3%) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fumagalli, C.; Betella, I.; Rappa, A.; di Giminiani, M.; Gaiano, M.; De Vitis, L.A.; Zambetti, B.; Vacirca, D.; Multinu, F.; Venetis, K.; et al. Tumor BRCA Testing in Epithelial Ovarian Cancers: Past and Future—Five-Years’ Single-Institution Experience of 762 Consecutive Patients. Cancers 2022, 14, 1638. https://doi.org/10.3390/cancers14071638
Fumagalli C, Betella I, Rappa A, di Giminiani M, Gaiano M, De Vitis LA, Zambetti B, Vacirca D, Multinu F, Venetis K, et al. Tumor BRCA Testing in Epithelial Ovarian Cancers: Past and Future—Five-Years’ Single-Institution Experience of 762 Consecutive Patients. Cancers. 2022; 14(7):1638. https://doi.org/10.3390/cancers14071638
Chicago/Turabian StyleFumagalli, Caterina, Ilaria Betella, Alessandra Rappa, Maria di Giminiani, Michela Gaiano, Luigi Antonio De Vitis, Benedetta Zambetti, Davide Vacirca, Francesco Multinu, Konstantinos Venetis, and et al. 2022. "Tumor BRCA Testing in Epithelial Ovarian Cancers: Past and Future—Five-Years’ Single-Institution Experience of 762 Consecutive Patients" Cancers 14, no. 7: 1638. https://doi.org/10.3390/cancers14071638
APA StyleFumagalli, C., Betella, I., Rappa, A., di Giminiani, M., Gaiano, M., De Vitis, L. A., Zambetti, B., Vacirca, D., Multinu, F., Venetis, K., Colombo, N., Barberis, M., & Guerini Rocco, E. (2022). Tumor BRCA Testing in Epithelial Ovarian Cancers: Past and Future—Five-Years’ Single-Institution Experience of 762 Consecutive Patients. Cancers, 14(7), 1638. https://doi.org/10.3390/cancers14071638








