Circulating Cell-Free DNA Profiling Predicts the Therapeutic Outcome in Advanced Hepatocellular Carcinoma Patients Treated with Combination Immunotherapy
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Patients and Study Design
2.2. Extraction and Quantitative Measurement of cfDNA
2.3. Library Preparation, Hybridization Capture and Sequencing of cfDNA
2.4. Variant Analysis
2.5. Statistical Analysis
3. Results
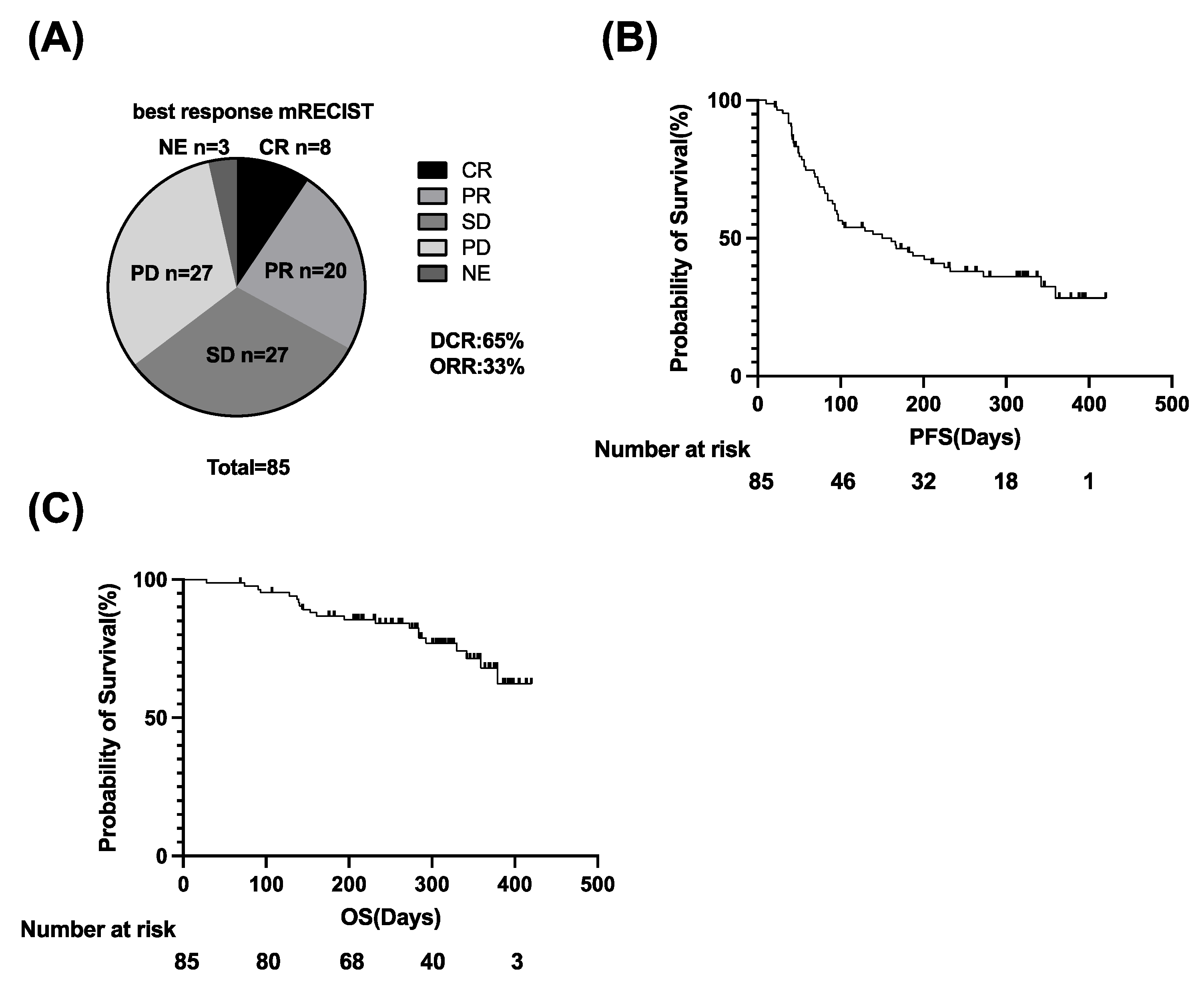
3.1. Clinical Outcomes of Atezo/Bev Therapy in u-HCC Patients
3.2. Patients with High Plasma cfDNA Levels Show a Significantly Lower ORR and Shorter PFS and OS Than Those with Low cfDNA Levels
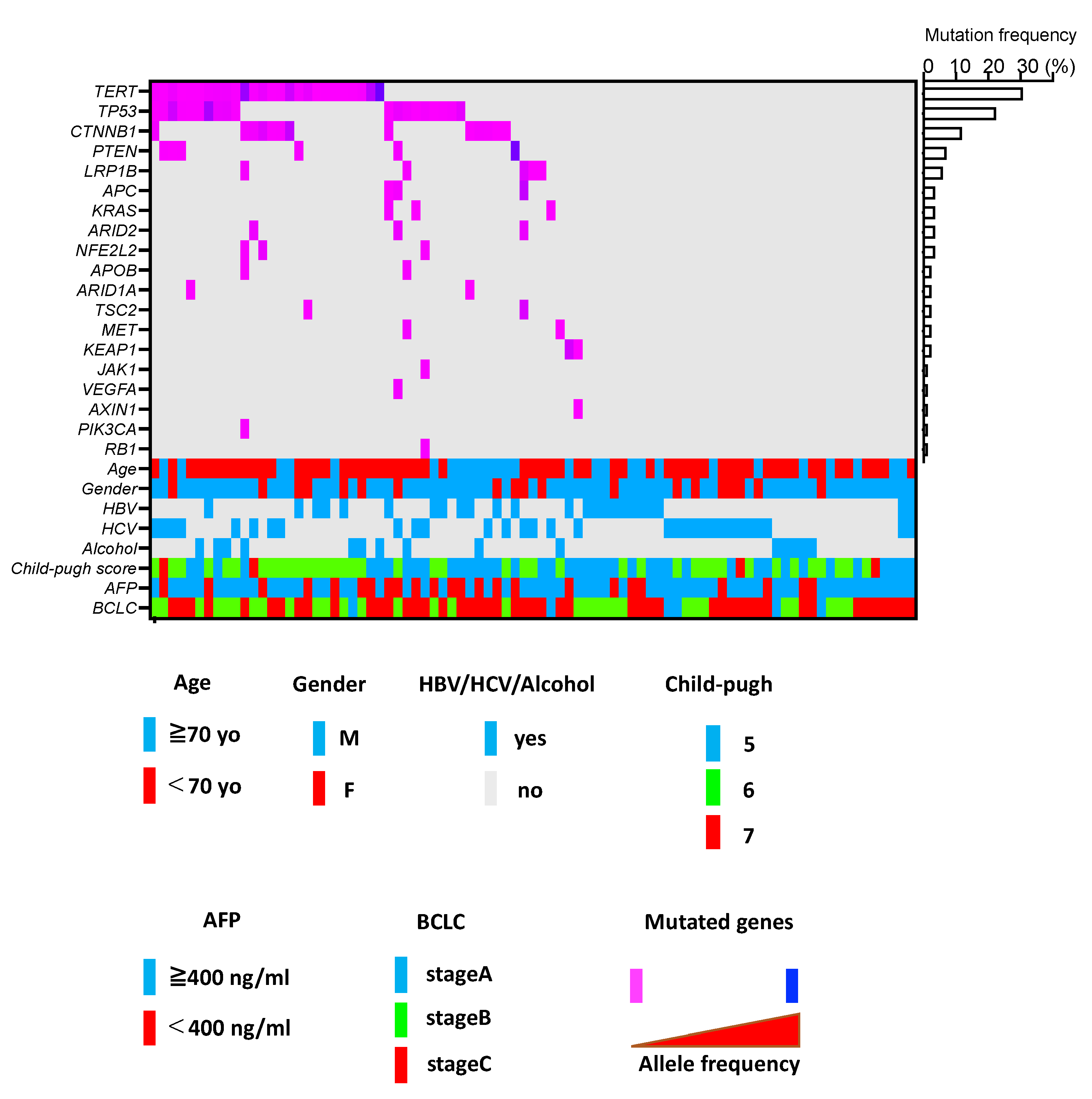
3.3. ctDNA Profiling in u-HCC Patients Treated with Atezo/Bev
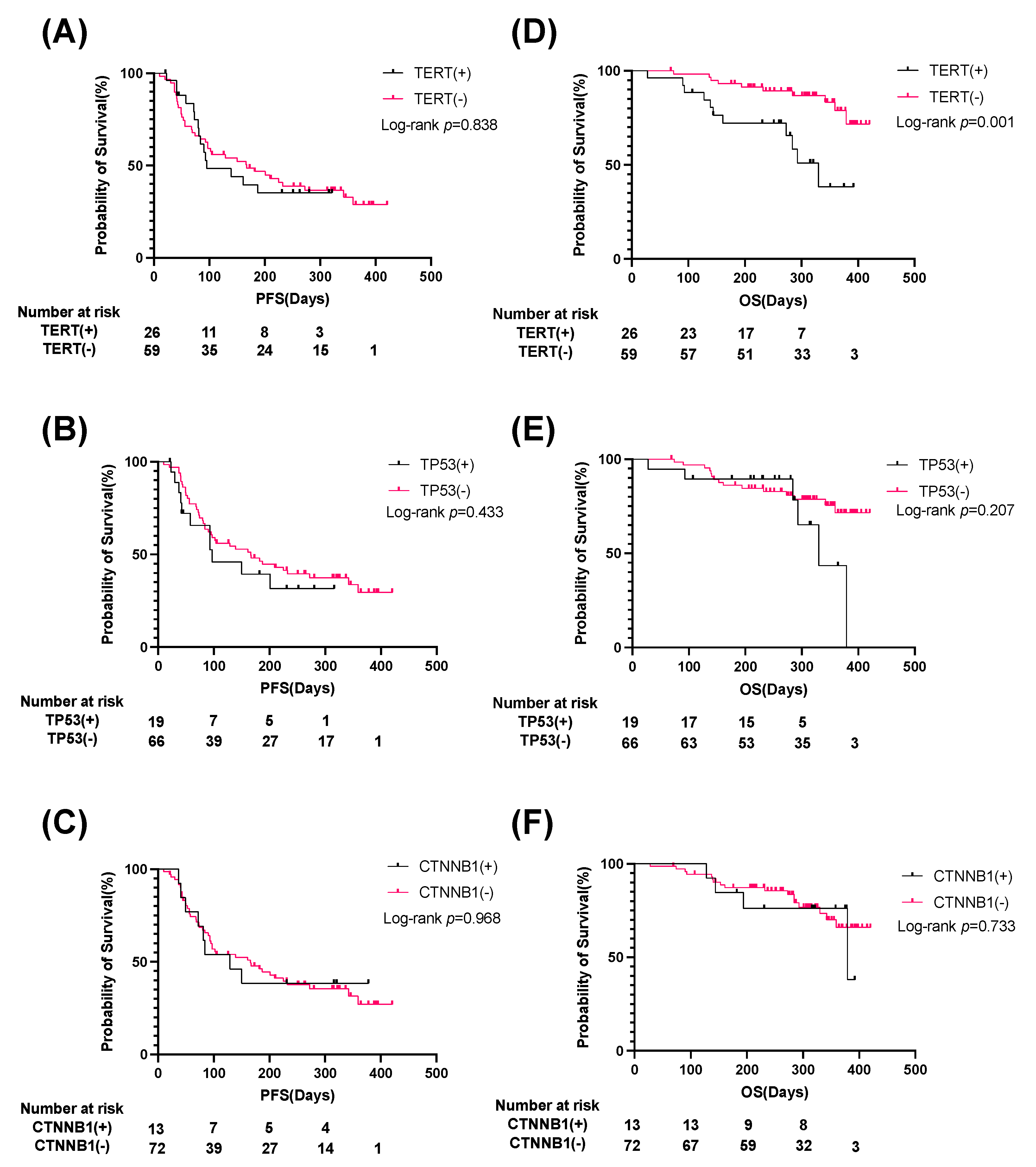
3.4. Patients with TERT Promoter ctDNA Show Significantly Shorter OS Than Those without
3.5. TERT ctDNA Mutation and AFP Level Can Be Used to Stratify u-HCC Patients Treated with Combination Immunotherapy Based on Prognosis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Myojin, Y.; Kodama, T.; Sakamori, R.; Maesaka, K.; Matsumae, T.; Sawai, Y.; Imai, Y.; Ohkawa, K.; Miyazaki, M.; Tanaka, S.; et al. Interleukin-6 Is a Circulating Prognostic Biomarker for Hepatocellular Carcinoma Patients Treated with Combined Immunotherapy. Cancers 2022, 14, 883. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.S.; Qin, S.; Ikeda, M.; Galle, P.R.; Ducreux, M.; Kim, T.-Y.; Kudo, M.; Breder, V.; Merle, P.; Kaseb, A.O.; et al. Atezolizumab plus Bevacizumab in Unresectable Hepatocellular Carcinoma. N. Engl. J. Med. 2020, 382, 1894–1905. [Google Scholar] [CrossRef] [PubMed]
- Cheng, A.L.; Qin, S.; Ikeda, M.; Galle, P.R.; Ducreux, M.; Kim, T.Y.; Kudo, M.; Breder, V.; Merle, P.; Kaseb, A.O.; et al. Updated efficacy and safety data from IMbrave150: Atezolizumab plus bevacizumab vs. sorafenib for unresectable hepatocellular carcinoma. J. Hepatol. 2021, 76, 862–873. [Google Scholar]
- Maesaka, K.; Sakamori, R.; Yamada, R.; Doi, A.; Tahata, Y.; Miyazaki, M.; Ohkawa, K.; Mita, E.; Iio, S.; Nozaki, Y.; et al. Comparison of atezolizumab plus bevacizumab and lenvatinib in terms of efficacy and safety as primary systemic chemotherapy for hepatocellular carcinoma. Hepatol. Res. 2022, 52, 630–640. [Google Scholar] [CrossRef]
- Li, F.; Li, C.; Cai, X.; Xie, Z.; Zhou, L.; Cheng, B.; Zhong, R.; Xiong, S.; Li, J.; Chen, Z.; et al. The association between CD8+ tumor-infiltrating lymphocytes and the clinical outcome of cancer immunotherapy: A systematic review and meta-analysis. EClinicalMedicine 2021, 41, 101134. [Google Scholar] [CrossRef]
- Lu, S.; Stein, J.E.; Rimm, D.L.; Wang, D.W.; Bell, J.M.; Johnson, D.B.; Sosman, J.A.; Schalper, K.A.; Anders, R.A.; Wang, H.; et al. Comparison of Biomarker Modalities for Predicting Response to PD-1/PD-L1 Checkpoint Blockade: A Systematic Review and Meta-analysis. JAMA Oncol. 2019, 5, 1195–1204. [Google Scholar] [CrossRef]
- Ruiz de Galarreta, M.; Bresnahan, E.; Molina-Sanchez, P.; Lindblad, K.E.; Maier, B.; Sia, D.; Puigvehi, M. beta-Catenin Activation Promotes Immune Escape and Resistance to Anti-PD-1 Therapy in Hepatocellular Carcinoma. Cancer Discov. 2019, 9, 1124–1141. [Google Scholar] [CrossRef]
- Morita, M.; Nishida, N.; Sakai, K.; Aoki, T.; Chishina, H.; Takita, M.; Ida, H.; Hagiwara, S.; Minami, Y.; Ueshima, K.; et al. Immunological Microenvironment Predicts the Survival of the Patients with Hepatocellular Carcinoma Treated with Anti-PD-1 Antibody. Liver Cancer 2021, 10, 380–393. [Google Scholar] [CrossRef]
- Ray, S.K.; Mukherjee, S. Cell Free DNA as an Evolving Liquid Biopsy Biomarker for Initial Diagnosis and Therapeutic Nursing in Cancer- An Evolving Aspect in Medical Biotechnology. Curr. Pharm. Biotechnol. 2022, 23, 112–122. [Google Scholar] [CrossRef]
- Von Felden, J.; Craig, A.J.; Garcia-Lezana, T.; Labgaa, I.; Haber, P.K.; D’Avola, D.; Asgharpour, A.; Dieterich, D.; Bonaccorso, A.; Torres-Martin, M.; et al. Mutations in circulating tumor DNA predict primary resistance to systemic therapies in advanced hepatocellular carcinoma. Oncogene 2021, 40, 140–151. [Google Scholar] [CrossRef] [PubMed]
- Wei, J.; Feng, J.; Weng, Y.; Xu, Z.; Jin, Y.; Wang, P.; Cui, X.; Ruan, P.; Luo, R.; Li, N.; et al. The Prognostic Value of ctDNA and bTMB on Immune Checkpoint Inhibitors in Human Cancer. Front. Oncol. 2021, 11, 706910. [Google Scholar] [CrossRef] [PubMed]
- Llovet, J.M.; Lencioni, R. mRECIST for HCC: Performance and novel refinements. J. Hepatol. 2020, 72, 288–306. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cancer Genome Atlas Research Network. Electronic address wbe, Cancer Genome Atlas Research N. Comprehensive and Integrative Genomic Characterization of Hepatocellular Carcinoma. Cell 2017, 169, 1327–1341.e23. [Google Scholar] [CrossRef] [Green Version]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M.; et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010, 20, 1297–1303. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [Green Version]
- Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 2011, 27, 2987–2993. [Google Scholar] [CrossRef] [Green Version]
- Wang, K.; Li, M.; Hakonarson, H. ANNOVAR: Functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010, 38, e164. [Google Scholar] [CrossRef]
- Totoki, Y.; Tatsuno, K.; Covington, K.R.; Ueda, H.; Creighton, C.J.; Kato, M.; Tsuji, S.; Donehower, L.A.; Slagle, B.L.; Nakamura, H.; et al. Trans-ancestry mutational landscape of hepatocellular carcinoma genomes. Nat. Genet. 2014, 46, 1267–1273. [Google Scholar] [CrossRef]
- Mirtavoos-Mahyari, H.; Ghafouri-Fard, S.; Khosravi, A.; Motevaseli, E.; Esfahani-Monfared, Z.; Seifi, S.; Salimi, B.; Oskooei, V.K.; Ghadami, M.; Modarressi, M.H. Circulating free DNA concentration as a marker of disease recurrence and metastatic potential in lung cancer. Clin. Transl. Med. 2019, 8, 14. [Google Scholar] [CrossRef]
- Tissot, C.; Toffart, A.-C.; Villar, S.; Souquet, P.-J.; Merle, P.; Moro-Sibilot, D.; Pérol, M.; Zavadil, J.; Brambilla, C.; Olivier, M.; et al. Circulating free DNA concentration is an independent prognostic biomarker in lung cancer. Eur. Respir. J. 2015, 46, 1773–1780. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Gao, Y.; Vafaei, S.; Gu, X.; Zhong, X. The Prognostic Value of Plasma Cell-Free DNA Concentration in the Prostate Cancer: A Systematic Review and Meta-Analysis. Front. Oncol. 2021, 11, 599602. [Google Scholar] [CrossRef] [PubMed]
- Chen, E.; Cario, C.L.; Leong, L.; Lopez, K.; Márquez, C.P.; Chu, C.; Li, P.S.; Oropeza, E.; Tenggara, I.; Cowan, J.; et al. Cell-free DNA concentration and fragment size as a biomarker for prostate cancer. Sci. Rep. 2021, 11, 5040. [Google Scholar] [CrossRef] [PubMed]
- Garcia, D.F.; Hills, A.; Page, K.; Hastings, R.K.; Toghill, B.; Goddard, K.S.; Ion, C.; Ogle, O.; Boydell, A.R.; Gleason, K.; et al. Plasma cell-free DNA (cfDNA) as a predictive and prognostic marker in patients with metastatic breast cancer. Breast Cancer Res. 2019, 21, 149. [Google Scholar] [CrossRef]
- Su, Y.; Wang, L.; Jiang, C.; Yue, Z.; Fan, H.; Hong, H.; Duan, C.; Jin, M.; Zhang, D.; Qiu, L.; et al. Increased plasma concentration of cell-free DNA precedes disease recurrence in children with high-risk neuroblastoma. BMC Cancer 2020, 20, 102. [Google Scholar] [CrossRef] [Green Version]
- Panagopoulou, M.; Karaglani, M.; Balgkouranidou, I.; Biziota, E.; Koukaki, T.; Karamitrousis, E.; Nena, E.; Tsamardinos, I.; Kolios, G.; Lianidou, E.; et al. Circulating cell-free DNA in breast cancer: Size profiling, levels, and methylation patterns lead to prognostic and predictive classifiers. Oncogene 2019, 38, 3387–3401. [Google Scholar] [CrossRef]
- Dziadziuszko, R.; Peters, S.; Mok, T.; Camidge, D.R.; Gadgeel, S.M.; Ou, S.-H.I.; Konopa, K.; Noé, J.; Nowicka, M.; Bordogna, W.; et al. Circulating Cell-free DNA as a Prognostic Biomarker in Patients with Advanced ALK+ Non-small Cell Lung Cancer in the Global Phase III ALEX Trial. Clin. Cancer Res. 2022, 28, 1800–1808. [Google Scholar] [CrossRef]
- Tran, N.H.; Kisiel, J.; Roberts, L.R. Using cell-free DNA for HCC surveillance and prognosis. JHEP Rep. 2021, 3, 100304. [Google Scholar] [CrossRef]
- Tokuhisa, Y.; Iizuka, N.; Sakaida, I.; Moribe, T.; Fujita, N.; Miura, T.; Tamatsukuri, S.; Ishitsuka, H.; Uchida, K.; Terai, S.; et al. Circulating cell-free DNA as a predictive marker for distant metastasis of hepatitis C virus-related hepatocellular carcinoma. Br. J. Cancer 2007, 97, 1399–1403. [Google Scholar] [CrossRef]
- Nakatsuka, T.; Nakagawa, H.; Hayata, Y.; Wake, T.; Yamada, T.; Kinoshita, M.N.; Nakagomi, R.; Sato, M.; Minami, T.; Uchino, K.; et al. Post-treatment cell-free DNA as a predictive biomarker in molecular-targeted therapy of hepatocellular carcinoma. J. Gastroenterol. 2021, 56, 456–469. [Google Scholar] [CrossRef]
- Fujimoto, A.; Furuta, M.; Totoki, Y.; Tsunoda, T.; Kato, M.; Shiraishi, Y.; Tanaka, H.; Taniguchi, H.; Kawakami, Y.; Ueno, M.; et al. Whole-genome mutational landscape and characterization of noncoding and structural mutations in liver cancer. Nat. Genet. 2016, 48, 500–509. [Google Scholar] [CrossRef] [PubMed]
- Schulze, K.; Imbeaud, S.; Letouzé, E.; Alexandrov, L.B.; Calderaro, J.; Rebouissou, S.; Couchy, G. Exome sequencing of hepatocellular carcinomas identifies new mutational signatures and potential therapeutic targets. Nat. Genet. 2015, 47, 505–511. [Google Scholar] [CrossRef] [PubMed]
- Harding, J.J.; Nandakumar, S.; Armenia, J.; Khalil, D.N.; Albano, M.; Ly, M.; Shia, J.; Hechtman, J.F.; Kundra, R.; El Dika, I.; et al. Prospective Genotyping of Hepatocellular Carcinoma: Clinical Implications of Next-Generation Sequencing for Matching Patients to Targeted and Immune Therapies. Clin. Cancer Res. 2019, 25, 2116–2126. [Google Scholar] [CrossRef] [Green Version]
- Lee, W.S.; Yang, H.; Chon, H.J.; Kim, C. Combination of anti-angiogenic therapy and immune checkpoint blockade normalizes vascular-immune crosstalk to potentiate cancer immunity. Exp. Mol. Med. 2020, 52, 1475–1485. [Google Scholar] [CrossRef]
- Li, H.; Li, J.; Zhang, C.; Zhang, C.; Wang, H. TERT mutations correlate with higher TMB value and unique tumor microenvironment and may be a potential biomarker for anti-CTLA4 treatment. Cancer Med. 2020, 9, 7151–7160. [Google Scholar] [CrossRef]
- Nault, J.C.; Calderaro, J.; Di Tommaso, L.; Balabaud, C.; Zafrani, E.S.; Bioulac-Sage, P.; Roncalli, M.; Zucman-Rossi, J. Telomerase reverse transcriptase promoter mutation is an early somatic genetic alteration in the transformation of premalignant nodules in hepatocellular carcinoma on cirrhosis. Hepatology 2014, 60, 1983–1992. [Google Scholar] [CrossRef] [PubMed]
- Kaneko, S.; Tsuchiya, K.; Yasui, Y.; Inada, K.; Kirino, S.; Yamashita, K.; Osawa, L.; Hayakawa, Y.; Sekiguchi, S.; Higuchi, M.; et al. Strategy for advanced hepatocellular carcinoma based on liver function and portal vein tumor thrombosis. Hepatol. Res. 2020, 50, 1375–1385. [Google Scholar] [CrossRef] [PubMed]


| Factor | Unit | Value |
|---|---|---|
| Age | Years Old | 74 (65–80) |
| Gender | Male/Female | 66/19 |
| ECOG PS | 0/1 | 76/9 |
| Etiology | HBV/HCV/HBV + HCV/alcohol/others | 22/29/2/15/17 |
| Child-pugh | 5/6/7 | 41/40/4 |
| PT | % | 92 (82–102) |
| ALB | g/dL | 3.7 (3.2–3.9) |
| T-BIL | mg/dL | 0.7 (0.5–1.0) |
| ALBI score | −2.35 (−2.69–−2.02) | |
| ALT | U/L | 26 (17–35) |
| PLT | ×104/μL | 13.8 (11.2–17.6) |
| NLR | 2.4 (1.8–3.6) | |
| AFP | ng/mL | 11 (3–887) |
| DCP | mAU/mL | 333 (65–2614) |
| Prior systemic therapy | Yes/No | 37/48 |
| Extrahepatic metastasis | Yes/No | 38/47 |
| Macrovascular invasion | Yes/No | 15/70 |
| Maximal tumor size | cm | 2.3 (1.6–4.5) |
| Intrahepatic tumor number | ≥5/≤4 | 36/49 |
| BCLC stage | A/B/C | 6/31/48 |
| Observation period | Days | 286 (216–359) |
| Factor | Unit | Univariate Analysis | Multivariate Analysis | |||
|---|---|---|---|---|---|---|
| Hazard Ratio (95% CI) | p Value | Hazard Ratio (95% CI) | p Value | |||
| Age | Years Old | ≥70/<70 | 0.96 (0.39–2.39) | 0.939 | ||
| Gender | Male/Female | 0.56 (0.23–1.39) | 0.212 | |||
| ECOG PS | 0/1 | 0.59 (0.17–2.00) | 0.398 | |||
| Etiology | Viral/non-Viral | 1.41 (0.55–3.65) | 0.477 | |||
| PT | % | ≥90/<90 | 1.12 (0.43–2.90) | 0.817 | ||
| ALB | g/dL | ≥4.0/<4.0 | 0.95 (0.35–2.60) | 0.920 | ||
| T-BIL | mg/dL | ≥0.7/<0.7 | 2.26 (0.83–6.18) | 0.112 | ||
| ALBI score | ≥−2.27/<−2.27 | 1.30 (0.55–3.09) | 0.546 | |||
| ALT | U/L | ≥45/<45 | 0.62 (0.14–2.66) | 0.520 | ||
| PLT | ×104/μL | ≥15/<15 | 0.47 (0.16–1.39) | 0.170 | ||
| NLR | ≥3.0/<3.0 | 3.98 (1.62–9.78) | 0.003 | 2.42 (0.80–7.36) | 0.119 | |
| AFP | ng/mL | ≥400/<400 | 4.79 (1.99–11.54) | 0.001 | 4.90 (1.58–15.13) | 0.006 |
| DCP | mAU/mL | ≥200/<200 | 2.96 (1.06–8.25) | 0.038 | 1.72 (0.53–5.57) | 0.365 |
| Prior systemic therapy | Yes/No | 0.99 (0.42–2.34) | 0.984 | |||
| Extrahepatic metastasis | Yes/No | 0.99 (0.42–2.35) | 0.980 | |||
| Macrovascular invasion | Yes/No | 3.61 (1.49–8.76) | 0.005 | 1.85 (0.62–5.59) | 0.273 | |
| Intrahepatic tumor number | ≥5/≤4 | 2.29 (0.96–5.46) | 0.061 | |||
| BCLC stage | A,B/C | 0.46 (0.18–1.18) | 0.104 | |||
| cfDNA | ng/uL | ≥2.23/<2.23 | 2.99 (1.16–7.75) | 0.024 | 2.92 (0.98–8.71) | 0.054 |
| TERT | Yes/No | 3.93 (1.63–9.44) | 0.002 | 3.25 (1.14–9.28) | 0.028 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Matsumae, T.; Kodama, T.; Myojin, Y.; Maesaka, K.; Sakamori, R.; Takuwa, A.; Oku, K.; Motooka, D.; Sawai, Y.; Oshita, M.; et al. Circulating Cell-Free DNA Profiling Predicts the Therapeutic Outcome in Advanced Hepatocellular Carcinoma Patients Treated with Combination Immunotherapy. Cancers 2022, 14, 3367. https://doi.org/10.3390/cancers14143367
Matsumae T, Kodama T, Myojin Y, Maesaka K, Sakamori R, Takuwa A, Oku K, Motooka D, Sawai Y, Oshita M, et al. Circulating Cell-Free DNA Profiling Predicts the Therapeutic Outcome in Advanced Hepatocellular Carcinoma Patients Treated with Combination Immunotherapy. Cancers. 2022; 14(14):3367. https://doi.org/10.3390/cancers14143367
Chicago/Turabian StyleMatsumae, Takayuki, Takahiro Kodama, Yuta Myojin, Kazuki Maesaka, Ryotaro Sakamori, Ayako Takuwa, Keiko Oku, Daisuke Motooka, Yoshiyuki Sawai, Masahide Oshita, and et al. 2022. "Circulating Cell-Free DNA Profiling Predicts the Therapeutic Outcome in Advanced Hepatocellular Carcinoma Patients Treated with Combination Immunotherapy" Cancers 14, no. 14: 3367. https://doi.org/10.3390/cancers14143367
APA StyleMatsumae, T., Kodama, T., Myojin, Y., Maesaka, K., Sakamori, R., Takuwa, A., Oku, K., Motooka, D., Sawai, Y., Oshita, M., Nakabori, T., Ohkawa, K., Miyazaki, M., Tanaka, S., Mita, E., Tawara, S., Yakushijin, T., Nozaki, Y., Hagiwara, H., ... Takehara, T. (2022). Circulating Cell-Free DNA Profiling Predicts the Therapeutic Outcome in Advanced Hepatocellular Carcinoma Patients Treated with Combination Immunotherapy. Cancers, 14(14), 3367. https://doi.org/10.3390/cancers14143367








