Computational Detection of Extraprostatic Extension of Prostate Cancer on Multiparametric MRI Using Deep Learning
Abstract
:Simple Summary
Abstract
1. Introduction
| First Author (Year) | Evaluation Granularity | Radiologist Input Required | Patient Number | Method | AUC |
|---|---|---|---|---|---|
| Hou (2021) [7] | Per index lesion | Tumor segmentation | 849 | Radiologists CNN | 0.63–0.74 0.73–0.81 |
| Cuocolo (2021) [17] | Per index lesion | Tumor segmentation | 193 | Radiologists Radiomics + SVM | 81–83% acc 0.73–0.80 74–79% acc |
| Eurboonyanun (2021) [27] | Per index | Measure TCL | 95 | Logistic regression w/ | |
| lesion | absolute TCL (euclidean) | 0.80 | |||
| actual TCL (curvilinear) | 0.74 | ||||
| Losnegard (2020) [28] | Per index lesion | Tumor segmentation | 228 | Radiologists Radiomics + Random forest | 0.75 0.74 |
| Park (2020) [29] | Per patient | Measure TCL | 301 | Radiologists using MRI-based EPE grade, ESUR score, Likert scale, TCL | 0.77–0.81 0.79–0.81 0.78–0.79 0.78–0.85 |
| Xu (2020) [19] | Per lesion (all those MRI visible) | Tumor segmentation | 95 | Radiomics + Regression algorithm | 0.87 |
| Shiradkar (2020) [18] | Per index lesion | Tumor and periprostatic fat segmentation | 45 | Radiomics + SVM | 0.88 |
| Mehralivand (2019) [30] | Per index lesion | Measure TCL | 553 | Logistic regression w/ MRI-based EPE grade + clinical features | 0.77 0.81 |
| Ma (2019) [31] | Per index lesion | Tumor segmentation | 210 | Radiologists Radiomics + Regression algorithm | 0.60–0.70 0.88 |
| Stanzione (2019) [20] | Per index lesion | Tumor segmentation | 39 | Radiomics + Bayesian Network | 0.88 |
| Krishna (2017) [32] | Per lesion (all those MRI visible) | Tumor segmentation | 149 | Radiologists Logistic regression w/ PI-RADS scores, tumor size, TCL, ADC entropy | 0.61–0.67, 0.61–0.72, 0.73, 0.69, 0.76 |
2. Materials and Methods
2.1. Dataset
2.1.1. Population Characteristics
2.1.2. Image Acquisition
2.1.3. Labels
2.2. Data Pre-Processing
2.2.1. Histopathology Pre-Processing
- Registration: Each digital histopathology image was aligned with its corresponding T2w MR image using the automated affine and deformable registration method RAPSODI [33]. This enabled accurate mapping of pixel-level cancer and extraprostatic extension labels from digital histopathology images onto MRI. For details on this process, refer to [26,33].
- Smoothing: Images were smoothed with a Gaussian filter with mm to avoid downsampling artifacts.
- Resampling: The Gaussian smoothed images were downsampled to an X-Y size of pixels, resulting in an in-plane pixel size of mm2.
- Intensity normalization: Each RGB channel of the resulting digital histopathology images was Z-score normalized.
2.2.2. MRI Pre-Processing
- Affine Registration: The T2w images and ADC images were manually registered using an affine transformation driven by the prostate segmentations on both modalities.
- Resampling: The T2w images, ADC images, prostate masks and cancer labels were projected and resampled on the corresponding histopathology images, resulting in images of pixels, with pixel size of mm2.
- Intensity standardization: We followed the procedure by Nyul et al. [34]. Using the training dataset, we learned a set of intensity histogram landmarks for T2w and ADC sequences independently. Then, we transformed the image histograms to align with the learned mean histogram of each MRI sequence. The histogram average learned in the training set was also used to align the cases in the test set. This histogram alignment intensity standardization method helps ensure similar MRI intensity distribution for all patients irrespective of scanners and scanning protocols.
- Intensity normalization: Finally, Z-score normalization was applied to the prostate regions of T2w and ADC images.
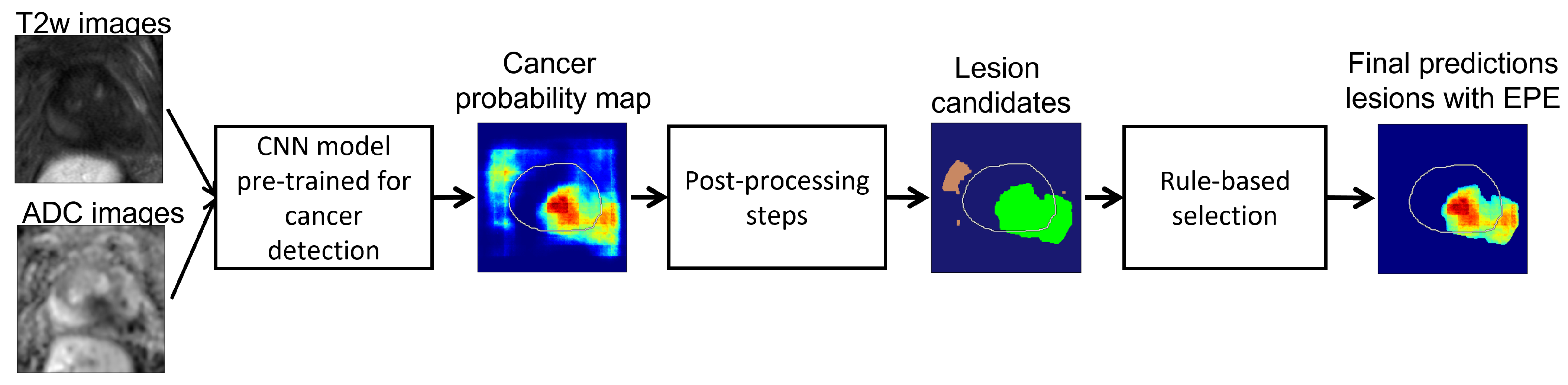
2.3. Proposed Approach
2.3.1. Step 1: Deep Learning Models for Cancer Detection
2.3.2. Step 2: Post-Processing Pipeline
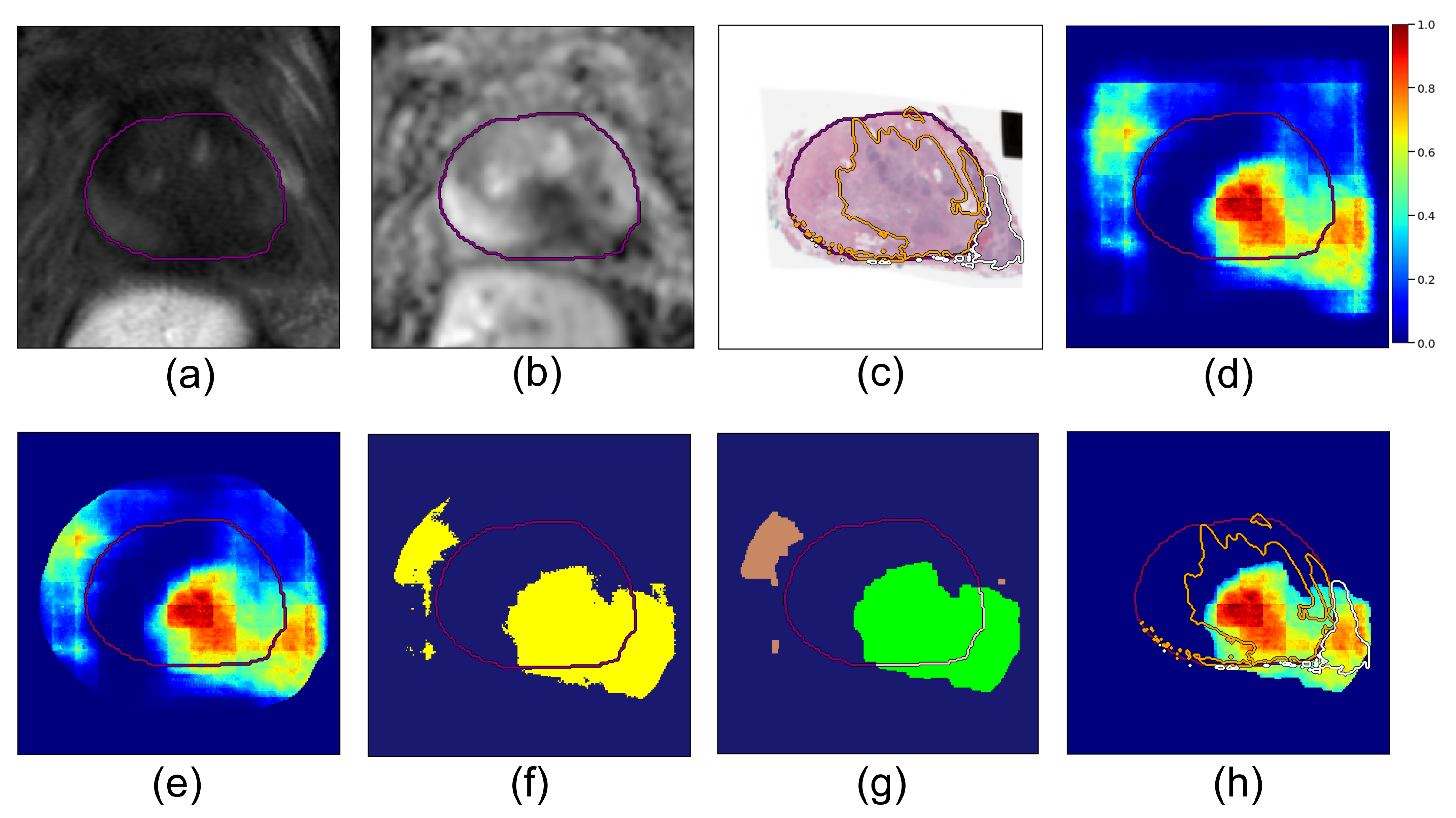
- Dilated prostate mask. The deep learning cancer predictions become less reliable the further we look outside the prostate, since other anatomical features may drive false positives. To prevent this, we applied a dilated prostate mask to the cancer probability map. Based on the diameter of the largest extraprostatic extension lesions in our cohort, we chose to dilate the original prostate mask using kernels of size pixels (corresponding to 1.86 cm × 1.86 cm):where represents the prostate segmentation mask (pixels have value 1 inside the prostate, value 0 outside) and is the resulting dilated prostate mask. We set all values in the probability map outside this region to zero:where is the cancer probability map output by the model (Equation (1)) (Figure 2d); ∗ denotes element-wise multiplication; the resulting is the 2D masked cancer probability map for a given slice (Figure 2e).
- Binary threshold. All pixels in the prediction map with probability greater than a fixed threshold, , were considered to be cancer, and the rest were set to zero; is a hyperparameter:and this results in a thresholded cancer probability map for the slice. We computed for all slices in a case, resulting in a volume for the patient, where z is the slice index.
- Connected components. Next, we computed all 3D connected components in the volume with connectivity value 26 using the python cc3d library [36]:Each component is a lesion candidate (Figure 2f). Note that the connected components function returns binary mask objects, i.e., each pixel in is either 0 or 1, and all the pixels with value 1 are connected.
- Logical rules: We used logical rules to prune these components and determine the final predictions for extraprostatic extension:
- Rule I: Component must predict cancer both inside:and outside the prostate capsulewhere is the prostate binary mask, i.e., for pixels inside the prostate and for pixels outside the prostate. If a component is either fully inside () or fully outside the prostate (), it is not a viable candidate for extraprostatic extension (Figure 2g, and brown components were discarded because they are fully outside the prostate and the green component was accepted since it crosses the prostate border).
- Rule II: For each viable lesion candidate, compute tumor–capsule contact line length (TCL) and compare with threshold . The overlap between a candidate and the prostate boundary defines a curvilinear segment (shown in pink in Figure 2g), and is the length of this segment.Lesion candidates with are discarded. Candidates with constitute our final predictions for cancer lesions with extraprostatic extension. Each final candidate is a binary mask; multiplying it element-wise with the probability map gives the probability map for cancer with extraprostatic extension (Figure 2h).We denote the final extraprostatic extension probability map for the entire case volume .
| Algorithm 1 Steps for predicting lesions with extraprostatic extension |
|
2.4. Evaluation
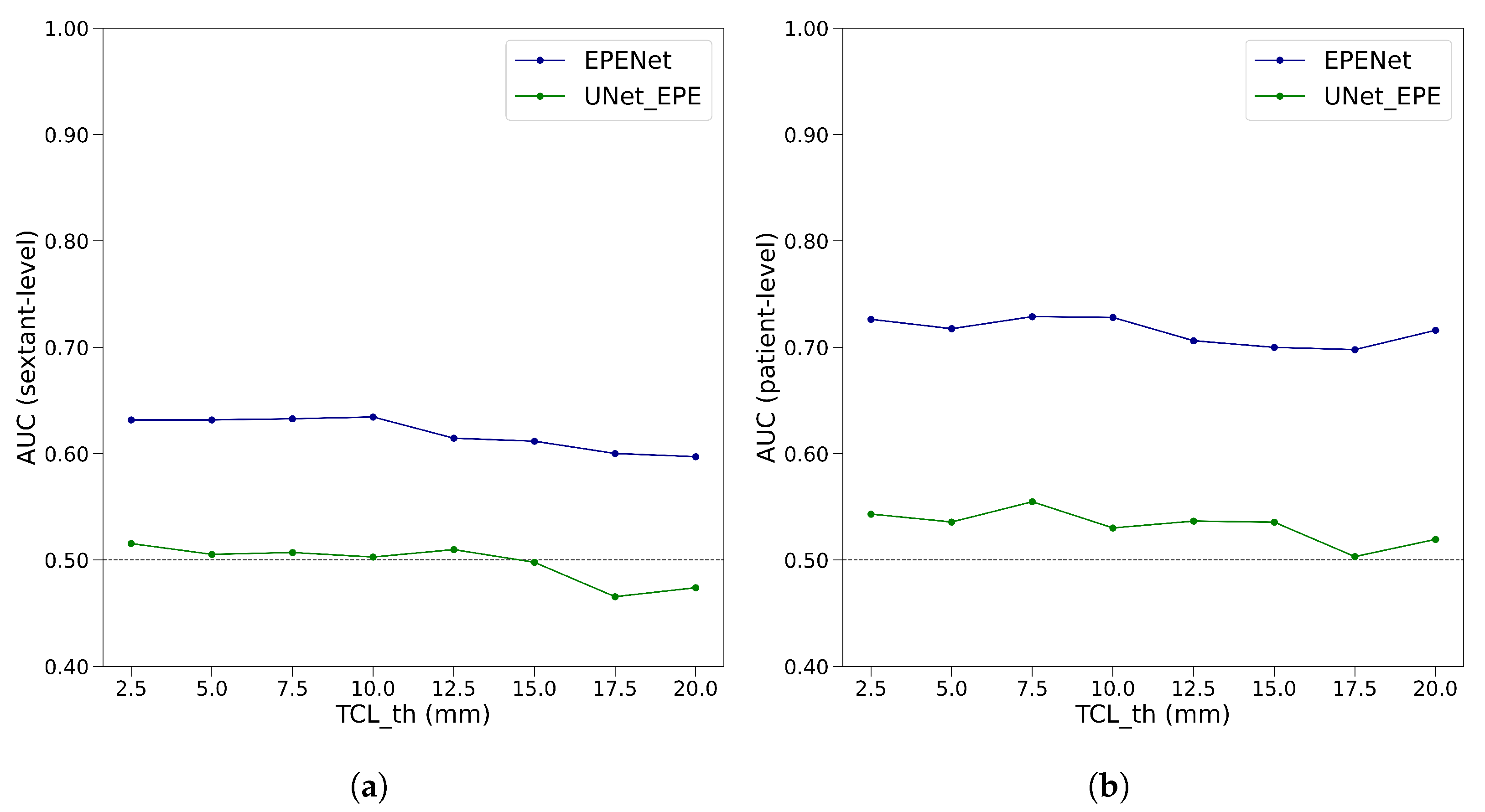
2.4.1. Patient-Level Evaluation
2.4.2. Sextant-Level Evaluation
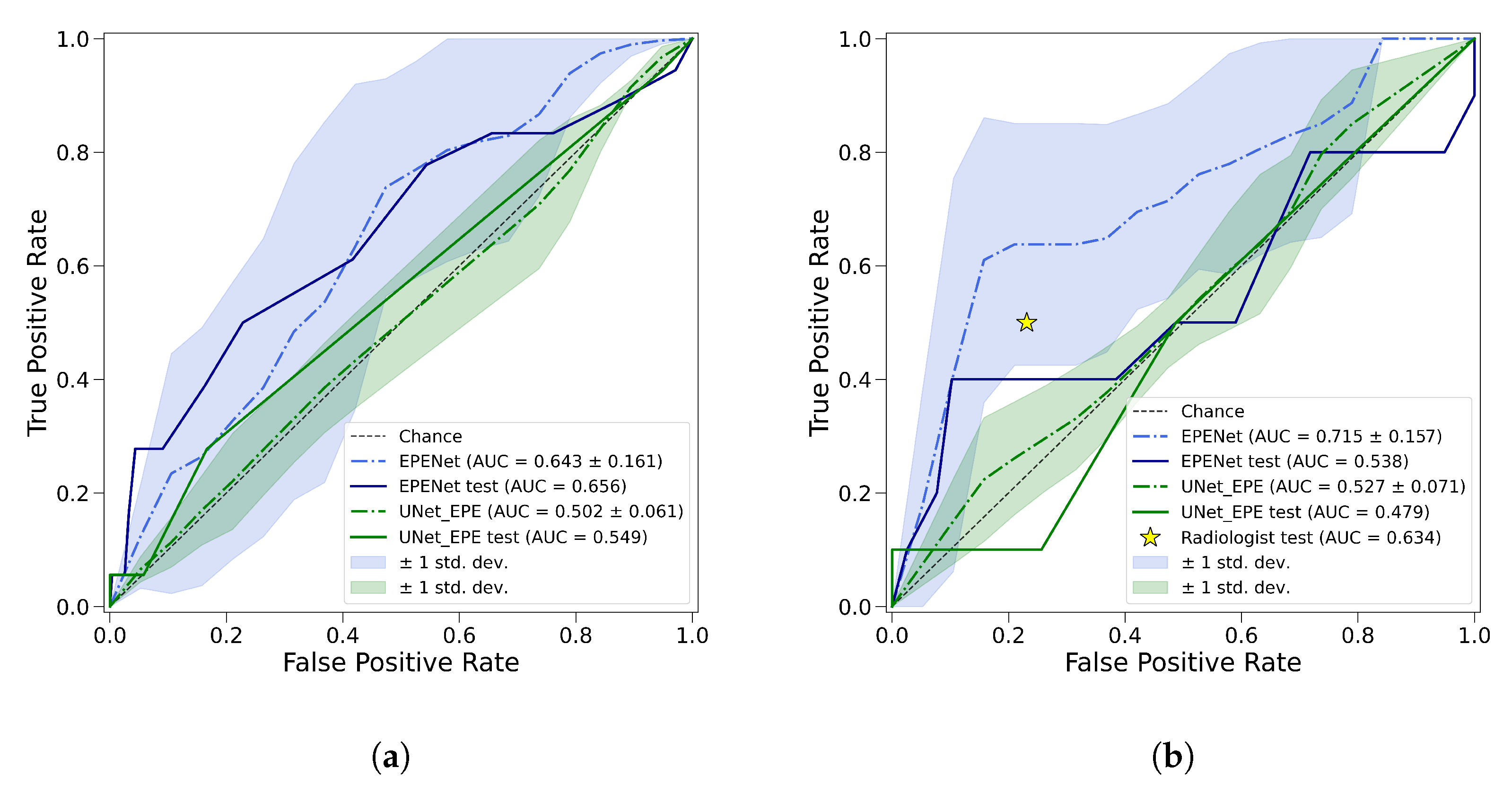
2.4.3. ROC Analysis
2.5. Experimental Design
Hyper-Parameter Optimization
3. Results
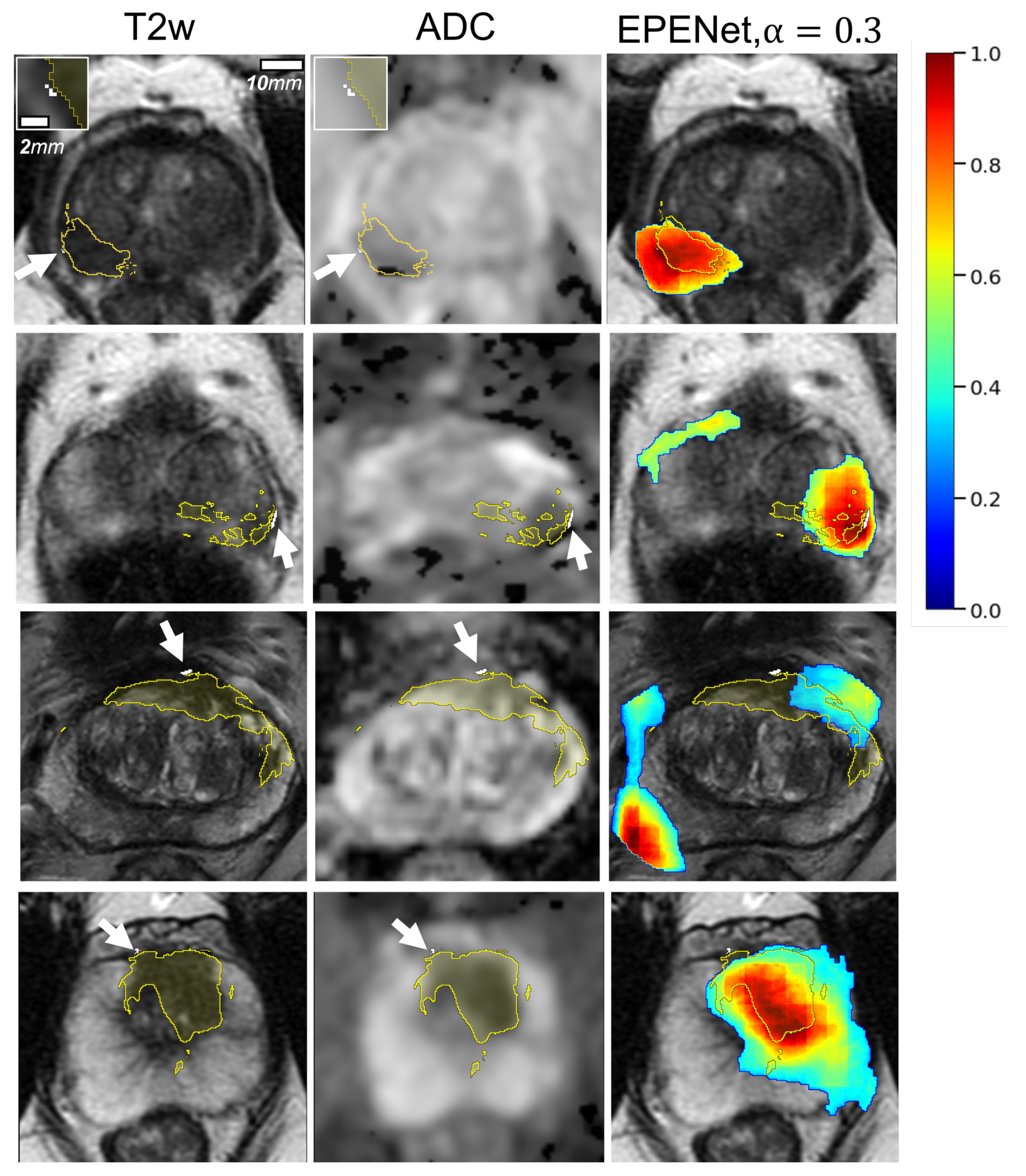
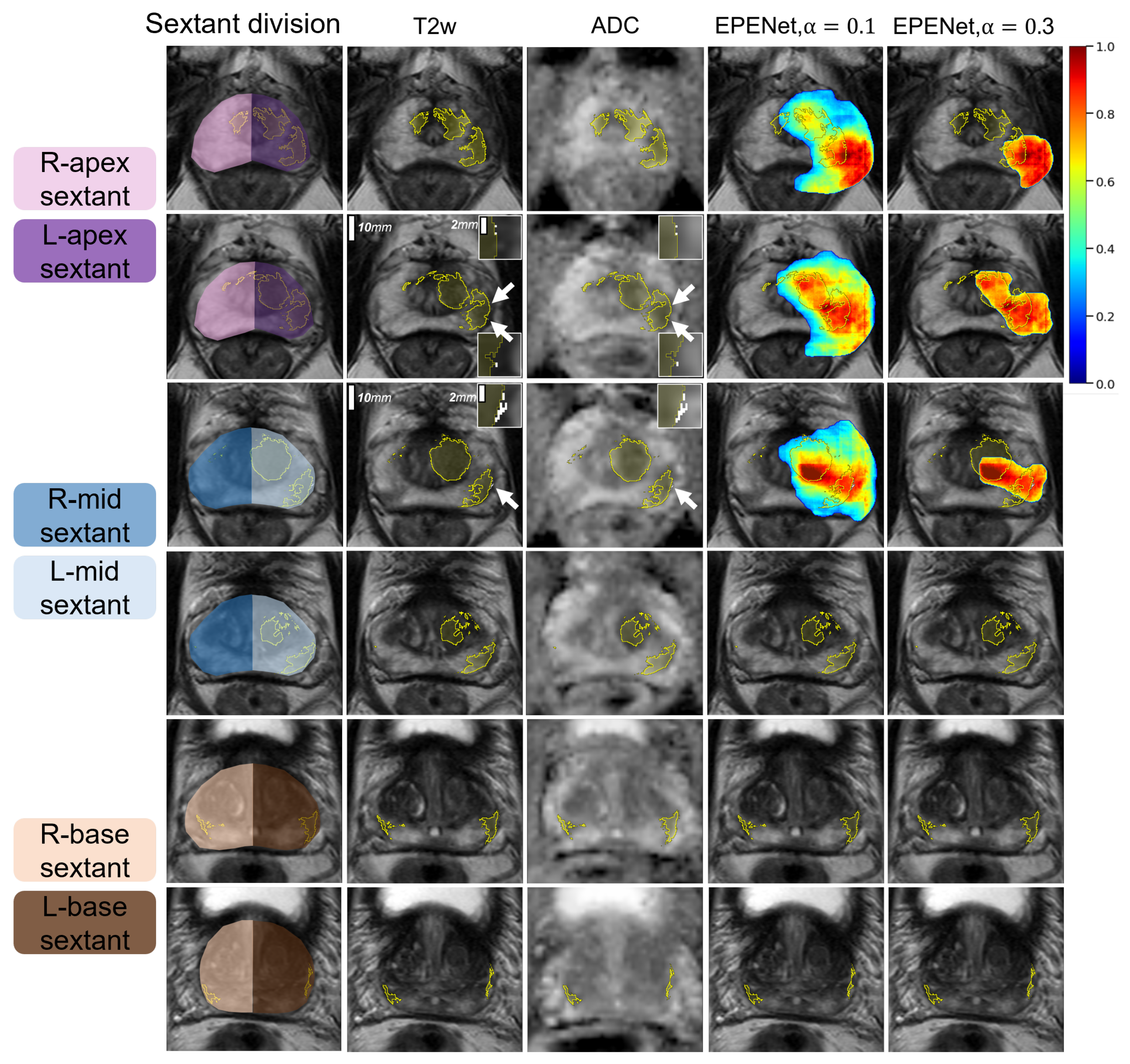
3.1. Qualitative Results
3.2. Quantitative Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ADC | Apparent Diffusion Coefficient |
| AUC | Area Under the Curve |
| CNN | Convolutional Neural Network |
| EPE | Extraprostatic Extension |
| MRI | Magnetic Resonance Imaging |
| PI-RADS | Prostate Imaging-Reporting and Data System |
| ROC | Receiver Operating Characteristic |
| SVM | Support Vector Machine |
| TCL | Tumor–capsule Contact Line Length (also known as Capsular Contact Length, CCL) |
Appendix A. Details of Deep Learning Models for Cancer Detection
Appendix B. Additional Visual Results
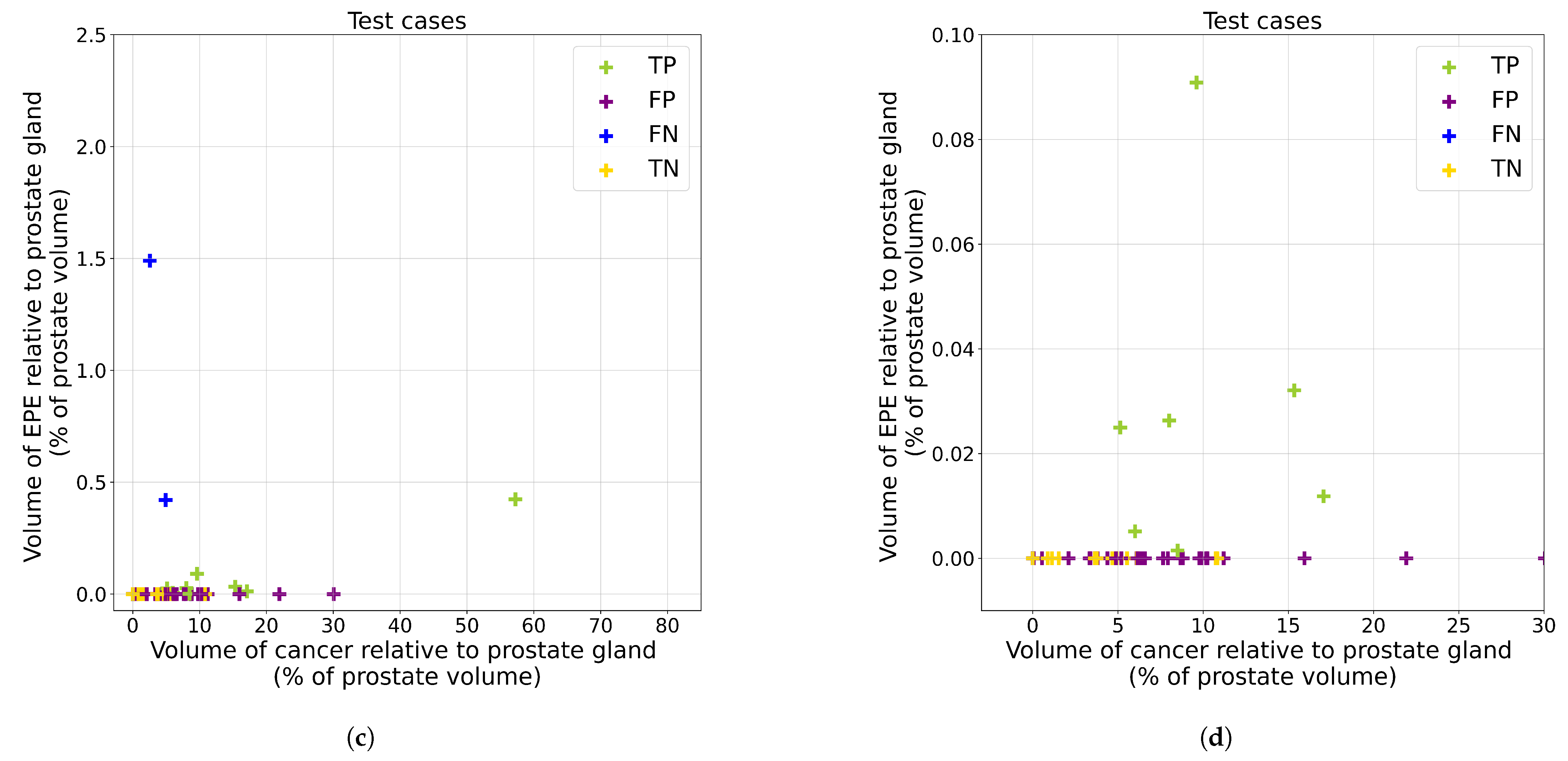
Appendix C. Analysis of False Positive Predictions

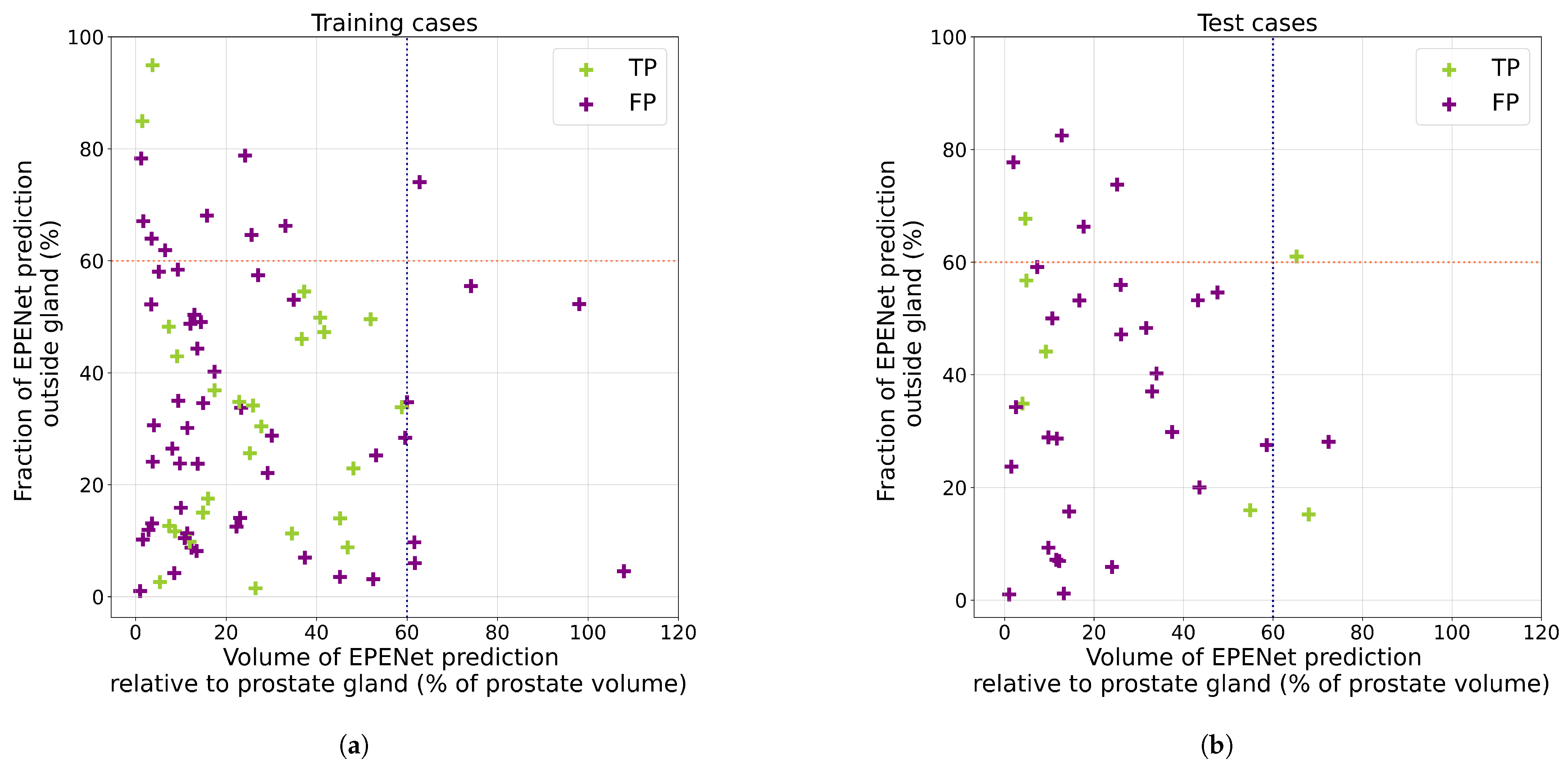
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef]
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer statistics, 2022. CA Cancer J. Clin. 2022, 72, 7–33. [Google Scholar] [CrossRef]
- Tourinho-Barbosa, R.; Srougi, V.; Nunes-Silva, I.; Baghdadi, M.; Rembeyo, G.; Eiffel, S.S.; Barret, E.; Rozet, F.; Galiano, M.; Cathelineau, X.; et al. Biochemical recurrence after radical prostatectomy: What does it mean? Int. Braz. J. 2018, 44, 14–21. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, X.; Kapoor, A.; Gu, Y.; Chow, M.; Xu, H.; Major, P.; Tang, D. Assessment of biochemical recurrence of prostate cancer (Review). Int. J. Oncol. 2019, 55, 1194–1212. [Google Scholar] [CrossRef] [Green Version]
- Park, C.K.; Chung, Y.S.; Choi, Y.D.; Ham, W.S.; Jang, W.S.; Cho, N.H. Revisiting extraprostatic extension based on invasion depth and number for new algorithm for substaging of pT3a prostate cancer. Sci. Rep. 2021, 11, 13952. [Google Scholar] [CrossRef] [PubMed]
- Cheng, L.; MacLennan, G.T.; Bostwick, D.G. Urologic Surgical Pathology; Saunders: Philadelphia, PA, USA, 2019. [Google Scholar]
- Hou, Y.; Zhang, Y.H.; Bao, J.; Bao, M.L.; Yang, G.; Shi, H.B.; Song, Y.; Zhang, Y.D. Artificial intelligence is a promising prospect for the detection of prostate cancer extracapsular extension with mpMRI: A two-center comparative study. Eur. J. Nucl. Med. Mol. Imaging 2021, 48, 3805–3816. [Google Scholar] [CrossRef]
- Gandaglia, G.; Ploussard, G.; Valerio, M.; Mattei, A.; Fiori, C.; Roumiguié, M.; Fossati, N.; Stabile, A.; Beauval, J.B.; Malavaud, B.; et al. The Key Combined Value of Multiparametric Magnetic Resonance Imaging, and Magnetic Resonance Imaging–targeted and Concomitant Systematic Biopsies for the Prediction of Adverse Pathological Features in Prostate Cancer Patients Undergoing Radical Prostatectomy. Eur. Urol. 2020, 77, 733–741. [Google Scholar] [CrossRef]
- Diamand, R.; Ploussard, G.; Roumiguié, M.; Oderda, M.; Benamran, D.; Fiard, G.; Quackels, T.; Assenmacher, G.; Simone, G.; Van Damme, J.; et al. External Validation of a Multiparametric Magnetic Resonance Imaging–based Nomogram for the Prediction of Extracapsular Extension and Seminal Vesicle Invasion in Prostate Cancer Patients Undergoing Radical Prostatectomy. Eur. Urol. 2021, 79, 180–185. [Google Scholar] [CrossRef] [PubMed]
- Somford, D.; Hamoen, E.; Fütterer, J.; van Basten, J.; van de Kaa, C.H.; Vreuls, W.; van Oort, I.; Vergunst, H.; Kiemeney, L.; Barentsz, J.; et al. The Predictive Value of Endorectal 3 Tesla Multiparametric Magnetic Resonance Imaging for Extraprostatic Extension in Patients with Low, Intermediate and High Risk Prostate Cancer. J. Urol. 2013, 190, 1728–1734. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Wu, Y.; Guo, J.; Chen, H.; Weng, X.; Liu, X. Intrafascial nerve-sparing radical prostatectomy improves patients’ postoperative continence recovery and erectile function. Medicine 2018, 97, e11297. [Google Scholar] [CrossRef]
- Shieh, A.C.; Guler, E.; Ojili, V.; Paspulati, R.M.; Elliott, R.; Ramaiya, N.H.; Tirumani, S.H. Extraprostatic extension in prostate cancer: Primer for radiologists. Abdom. Radiol. 2020, 45, 4040–4051. [Google Scholar] [CrossRef]
- Falagario, U.G.; Jambor, I.; Ratnani, P.; Martini, A.; Treacy, P.J.; Wajswol, E.; Lantz, A.; Papastefanou, G.; Weil, R.; Phillip, D.; et al. Performance of prostate multiparametric MRI for prediction of prostate cancer extraprostatic extension according to NCCN risk categories: Implication for surgical planning. Ital. J. Urol. Nephrol. 2020, 72, 746–754. [Google Scholar]
- De Rooij, M.; Hamoen, E.H.; Witjes, J.A.; Barentsz, J.O.; Rovers, M.M. Accuracy of Magnetic Resonance Imaging for Local Staging of Prostate Cancer: A Diagnostic Meta-analysis. Eur. Urol. 2016, 70, 233–245. [Google Scholar] [CrossRef]
- Eberhardt, S.C. Local Staging of Prostate Cancer with MRI: A Need for Standardization. Radiology 2019, 290, 720–721. [Google Scholar] [CrossRef] [PubMed]
- Choyke, P.L. A Grading System for Extraprostatic Extension of Prostate Cancer That We Can All Agree Upon? Radiol. Imaging Cancer 2020, 2, e190088. [Google Scholar] [CrossRef] [PubMed]
- Cuocolo, R.; Stanzione, A.; Faletti, R.; Gatti, M.; Calleris, G.; Fornari, A.; Gentile, F.; Motta, A.; Dell’Aversana, S.; Creta, M.; et al. MRI index lesion radiomics and machine learning for detection of extraprostatic extension of disease: A multicenter study. Eur. Radiol. 2021, 31, 7575–7583. [Google Scholar] [CrossRef]
- Shiradkar, R.; Zuo, R.; Mahran, A.; Ponsky, L.; Tirumani, S.H.; Madabhushi, A. Radiomic features derived from periprostatic fat on pre-surgical T2w MRI predict extraprostatic extension of prostate cancer identified on post-surgical pathology: Preliminary results. In Proceedings of the Medical Imaging 2020: Computer-Aided Diagnosis, Houston, TX, USA, 15–20 February 2020; Hahn, H.K., Mazurowski, M.A., Eds.; SPIE: Bellingham, WA, USA, 2020. [Google Scholar] [CrossRef]
- Xu, L.; Zhang, G.; Zhao, L.; Mao, L.; Li, X.; Yan, W.; Xiao, Y.; Lei, J.; Sun, H.; Jin, Z. Radiomics Based on Multiparametric Magnetic Resonance Imaging to Predict Extraprostatic Extension of Prostate Cancer. Front. Oncol. 2020, 10, 940. [Google Scholar] [CrossRef]
- Stanzione, A.; Cuocolo, R.; Cocozza, S.; Romeo, V.; Persico, F.; Fusco, F.; Longo, N.; Brunetti, A.; Imbriaco, M. Detection of Extraprostatic Extension of Cancer on Biparametric MRI Combining Texture Analysis and Machine Learning: Preliminary Results. Acad. Radiol. 2019, 26, 1338–1344. [Google Scholar] [CrossRef]
- Sumathipala, Y.; Lay, N.; Turkbey, B. Prostate cancer detection from multi-institution multiparametric MRIs using deep convolutional neural networks. J. Med. Imaging 2018, 5, 044507. [Google Scholar] [CrossRef]
- Cao, R.; Bajgiran, A.M.; Mirak, S.A.; Shakeri, S.; Zhong, X.; Enzmann, D.; Raman, S.; Sung, K. Joint Prostate Cancer Detection and Gleason Score Prediction in mp-MRI via FocalNet. IEEE Trans. Med. Imaging 2019, 38, 2496–2506. [Google Scholar] [CrossRef] [Green Version]
- Sanyal, J.; Banerjee, I.; Hahn, L.; Rubin, D. An Automated Two-step Pipeline for Aggressive Prostate Lesion Detection from Multi-parametric MR Sequence. AMIA Jt. Summits Transl. Sci. Proc. 2020, 2020, 552–560. [Google Scholar]
- Wildeboer, R.R.; van Sloun, R.J.; Wijkstra, H.; Mischi, M. Artificial intelligence in multiparametric prostate cancer imaging with focus on deep-learning methods. Comput. Methods Programs Biomed. 2020, 189, 105316. [Google Scholar] [CrossRef]
- Seetharaman, A.; Bhattacharya, I.; Chen, L.C.; Kunder, C.A.; Shao, W.; Soerensen, S.J.C.; Wang, J.B.; Teslovich, N.C.; Fan, R.E.; Ghanouni, P.; et al. Automated detection of aggressive and indolent prostate cancer on magnetic resonance imaging. Med. Phys. 2021, 48, 2960–2972. [Google Scholar] [CrossRef]
- Bhattacharya, I.; Seetharaman, A.; Kunder, C.; Shao, W.; Chen, L.C.; Soerensen, S.J.; Wang, J.B.; Teslovich, N.C.; Fan, R.E.; Ghanouni, P.; et al. Selective identification and localization of indolent and aggressive prostate cancers via CorrSigNIA: An MRI-pathology correlation and deep learning framework. Med. Image Anal. 2022, 75, 102288. [Google Scholar] [CrossRef]
- Eurboonyanun, K.; Pisuchpen, N.; O’Shea, A.; Lahoud, R.M.; Atre, I.D.; Harisinghani, M. The absolute tumor-capsule contact length in the diagnosis of extraprostatic extension of prostate cancer. Abdom. Radiol. 2021, 46, 4014–4024. [Google Scholar] [CrossRef]
- Losnegård, A.; Reisæter, L.A.R.; Halvorsen, O.J.; Jurek, J.; Assmus, J.; Arnes, J.B.; Honoré, A.; Monssen, J.A.; Andersen, E.; Haldorsen, I.S.; et al. Magnetic resonance radiomics for prediction of extraprostatic extension in non-favorable intermediate- and high-risk prostate cancer patients. Acta Radiol. 2020, 61, 1570–1579. [Google Scholar] [CrossRef] [PubMed]
- Park, K.J.; hyun Kim, M.; Kim, J.K. Extraprostatic Tumor Extension: Comparison of Preoperative Multiparametric MRI Criteria and Histopathologic Correlation after Radical Prostatectomy. Radiology 2020, 296, 87–95. [Google Scholar] [CrossRef] [PubMed]
- Mehralivand, S.; Shih, J.H.; Harmon, S.; Smith, C.; Bloom, J.; Czarniecki, M.; Gold, S.; Hale, G.; Rayn, K.; Merino, M.J.; et al. A Grading System for the Assessment of Risk of Extraprostatic Extension of Prostate Cancer at Multiparametric MRI. Radiology 2019, 290, 709–719. [Google Scholar] [CrossRef]
- Ma, S.; Xie, H.; Wang, H.; Han, C.; Yang, J.; Lin, Z.; Li, Y.; He, Q.; Wang, R.; Cui, Y.; et al. MRI-Based Radiomics Signature for the Preoperative Prediction of Extracapsular Extension of Prostate Cancer. J. Magn. Reson. Imaging 2019, 50, 1914–1925. [Google Scholar] [CrossRef] [PubMed]
- Krishna, S.; Lim, C.S.; McInnes, M.D.; Flood, T.A.; Shabana, W.M.; Lim, R.S.; Schieda, N. Evaluation of MRI for diagnosis of extraprostatic extension in prostate cancer. J. Magn. Reson. Imaging 2017, 47, 176–185. [Google Scholar] [CrossRef]
- Rusu, M.; Shao, W.; Kunder, C.A.; Wang, J.B.; Soerensen, S.J.C.; Teslovich, N.C.; Sood, R.R.; Chen, L.C.; Fan, R.E.; Ghanouni, P.; et al. Registration of presurgical MRI and histopathology images from radical prostatectomy via RAPSODI. Med. Phys. 2020, 47, 4177–4188. [Google Scholar] [CrossRef] [PubMed]
- Nyul, L.; Udupa, J.; Zhang, X. New variants of a method of MRI scale standardization. IEEE Trans. Med. Imaging 2000, 19, 143–150. [Google Scholar] [CrossRef] [PubMed]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. In Lecture Notes in Computer Science; Springer International Publishing: Berlin/Heidelberg, Germany, 2015; pp. 234–241. [Google Scholar] [CrossRef] [Green Version]
- Silversmith, W. cc3d: Connected Components on Multilabel 3D & 2D Images. 2021. Available online: https://zenodo.org/record/5719536#.YpWNCv5BxPY (accessed on 1 May 2022).
- Elliott, S.P.; Shinohara, K.; Logan, S.L.; Carroll, P.R. Sextant Prostate Biopsies Predict Side and Sextant Site of Extracapsular Extension of Prostate Cancer. J. Urol. 2002, 168, 105–109. [Google Scholar] [CrossRef]
- Mayes, J.M.; Mouraviev, V.; Sun, L.; Madden, J.F.; Polascik, T.J. Can the conventional sextant prostate biopsy reliably diagnose unilateral prostate cancer in low-risk, localized, prostate cancer? Prostate Cancer Prostatic Dis. 2008. [Google Scholar] [CrossRef]
- Kim, T.H.; Woo, S.; Han, S.; Suh, C.H.; Ghafoor, S.; Hricak, H.; Vargas, H.A. The Diagnostic Performance of the Length of Tumor Capsular Contact on MRI for Detecting Prostate Cancer Extraprostatic Extension: A Systematic Review and Meta-Analysis. Korean J. Radiol. 2020, 21, 684. [Google Scholar] [CrossRef]
- Kikinis, R.; Pieper, S.D.; Vosburgh, K.G. 3D Slicer: A Platform for Subject-Specific Image Analysis, Visualization, and Clinical Support. In Intraoperative Imaging and Image-Guided Therapy; Springer: New York, NY, USA, 2013; pp. 277–289. [Google Scholar] [CrossRef]
- Kumar, A.; Patel, V.R.; Panaiyadiyan, S.; Bhat, K.R.S.; Moschovas, M.C.; Nayak, B. Nerve-sparing robot-assisted radical prostatectomy: Current perspectives. Asian J. Urol. 2021, 8, 2–13. [Google Scholar] [CrossRef]
- Schelb, P.; Kohl, S.; Radtke, J.P.; Wiesenfarth, M.; Kickingereder, P.; Bickelhaupt, S.; Kuder, T.A.; Stenzinger, A.; Hohenfellner, M.; Schlemmer, H.P.; et al. Classification of Cancer at Prostate MRI: Deep Learning versus Clinical PI-RADS Assessment. Radiology 2019, 293, 607–617. [Google Scholar] [CrossRef]
- de Vente, C.; Vos, P.; Hosseinzadeh, M.; Pluim, J.; Veta, M. Deep Learning Regression for Prostate Cancer Detection and Grading in Bi-Parametric MRI. IEEE Trans. Biomed. Eng. 2021, 68, 374–383. [Google Scholar] [CrossRef] [PubMed]
- Xie, S.; Tu, Z. Holistically-Nested Edge Detection. arXiv 2015, arXiv:1504.06375. [Google Scholar]
- Bhattacharya, I.; Seetharaman, A.; Shao, W.; Sood, R.; Kunder, C.A.; Fan, R.E.; Soerensen, S.J.C.; Wang, J.B.; Ghanouni, P.; Teslovich, N.C.; et al. CorrSigNet: Learning CORRelated Prostate Cancer SIGnatures from Radiology and Pathology Images for Improved Computer Aided Diagnosis. In Proceedings of the MICCAI 2020 Medical Image Computing and Computer Assisted Intervention, Lima, Peru, 4–8 October 2020; Springer International Publishing: Cham, Switzerland, 2020; pp. 315–325. [Google Scholar] [CrossRef]
| Cohort | Train | Test |
|---|---|---|
| Patient number | 74 | 49 |
| Lesion count | 90 | 58 |
| Indolent | 9 | 10 |
| Aggressive | 81 | 48 |
| EPE (pathologically proven) | 29 | 10 |
| Lesion volume (mm3) | 1541.6 (714.7, 3418.6) | 1099.1 (743.2, 2544.7) |
| EPE volume (where applicable) | 8.6 (3.6, 44.6) | 10.6 (5.6, 36.3) |
| Number of Patients | 123 |
|---|---|
| T2w | |
| Repetition time (TR, range) (s) | |
| Echo time (TE, range) (ms) | |
| Pixel size (range) (mm) | |
| Distance between slices (mm) | |
| Matrix size | |
| Number of slices | |
| ADC | |
| b-values (s/mm2) | |
| Pixel size (range) (mm) | |
| Distance between slices (mm) | |
| Matrix size | |
| Number of slices |
| Mode | Threshold | Cohort | Sensitivity | Specificity | Sensitivity | Specificity |
|---|---|---|---|---|---|---|
| (Patient) | (Patient) | (Sextant) | (Sextant) | |||
| % | % | % | % | |||
| EPENet | 0.10 | cross-val | ||||
| test | 90.0 | 0.0 | 88.9 | 13.0 | ||
| EPENet | 0.15 | cross-val | ||||
| test | 90.0 | 0.0 | 83.3 | 23.9 | ||
| EPENet | 0.20 | cross-val | ||||
| test | 80.0 | 5.1 | 83.3 | 34.4 | ||
| EPENet | 0.25 | cross-val | ||||
| test | 80.0 | 12.8 | 77.8 | 45.7 | ||
| EPENet | 0.30 | cross-val | 95.0 ± 10.0 | 26.8 ± 8.8 | 64.4 ± 21.6 | 54.6 ± 8.1 |
| test | 80.0 | 28.2 | 61.1 | 58.3 | ||
| EPENet | 0.35 | cross-val | ||||
| test | 50.0 | 41.0 | 55.6 | 67.8 | ||
| EPENet | 0.40 | cross-val | ||||
| test | 50.0 | 51.3 | 50.0 | 77.2 | ||
| EPENet | 0.45 | cross-val | ||||
| test | 40.0 | 61.5 | 38.9 | 83.7 | ||
| EPENet | 0.50 | cross-val | ||||
| test | 40.0 | 74.4 | 27.8 | 90.9 | ||
| EPENet | 0.55 | cross-val | ||||
| test | 40.0 | 87.2 | 27.8 | 95.7 | ||
| EPENet | 0.60 | cross-val | ||||
| test | 40.0 | 89.7 | 16.7 | 96.7 | ||
| Radiologists | — | test | 50.0 | 76.9 | — | – |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Moroianu, Ş.L.; Bhattacharya, I.; Seetharaman, A.; Shao, W.; Kunder, C.A.; Sharma, A.; Ghanouni, P.; Fan, R.E.; Sonn, G.A.; Rusu, M. Computational Detection of Extraprostatic Extension of Prostate Cancer on Multiparametric MRI Using Deep Learning. Cancers 2022, 14, 2821. https://doi.org/10.3390/cancers14122821
Moroianu ŞL, Bhattacharya I, Seetharaman A, Shao W, Kunder CA, Sharma A, Ghanouni P, Fan RE, Sonn GA, Rusu M. Computational Detection of Extraprostatic Extension of Prostate Cancer on Multiparametric MRI Using Deep Learning. Cancers. 2022; 14(12):2821. https://doi.org/10.3390/cancers14122821
Chicago/Turabian StyleMoroianu, Ştefania L., Indrani Bhattacharya, Arun Seetharaman, Wei Shao, Christian A. Kunder, Avishkar Sharma, Pejman Ghanouni, Richard E. Fan, Geoffrey A. Sonn, and Mirabela Rusu. 2022. "Computational Detection of Extraprostatic Extension of Prostate Cancer on Multiparametric MRI Using Deep Learning" Cancers 14, no. 12: 2821. https://doi.org/10.3390/cancers14122821
APA StyleMoroianu, Ş. L., Bhattacharya, I., Seetharaman, A., Shao, W., Kunder, C. A., Sharma, A., Ghanouni, P., Fan, R. E., Sonn, G. A., & Rusu, M. (2022). Computational Detection of Extraprostatic Extension of Prostate Cancer on Multiparametric MRI Using Deep Learning. Cancers, 14(12), 2821. https://doi.org/10.3390/cancers14122821






