Germline Genetics of Prostate Cancer: Prevalence of Risk Variants and Clinical Implications for Disease Management
Abstract
Simple Summary
Abstract
1. Introduction
2. Germline Genetic Variations Associated with Prostate Cancer Risk
2.1. DNA Repair Genes
2.2. Mismatch Repair Genes
2.3. HOXB13
2.4. Health Disparities in Genetic Studies
3. Recommendation for Germline Genetic Testing in Prostate Cancer
Hereditary Genetic Testing Panels
4. Germline Markers Relevant for Prostate Cancer Specific Therapeutic Recommendations
Companion Diagnostics
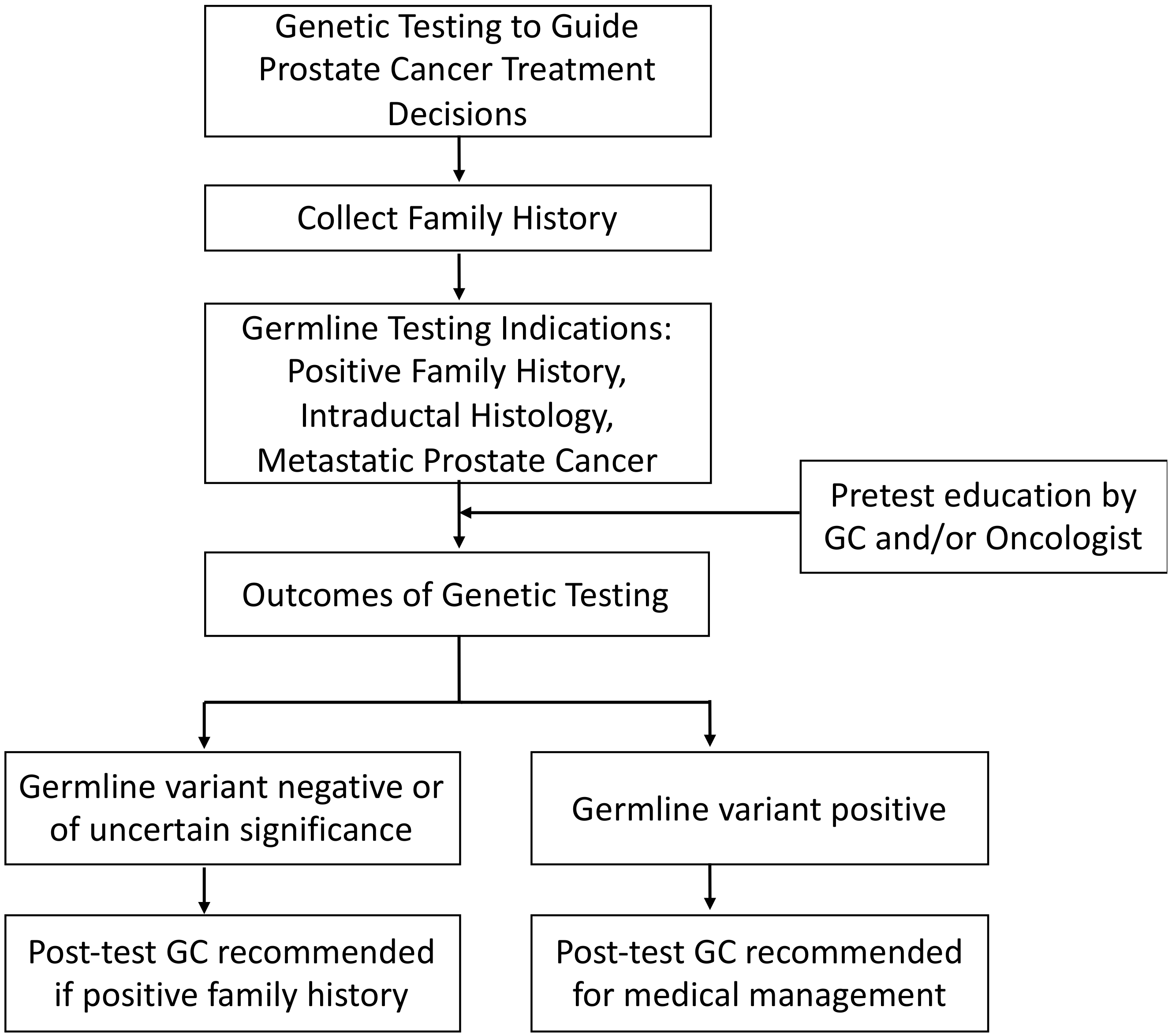
5. Genetic Counseling
6. Challenges and Barriers to Prostate Cancer Genetic Testing
7. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer Statistics, 2021. CA Cancer J. Clin. 2021, 71, 7–33. [Google Scholar] [CrossRef]
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2021. [Google Scholar] [CrossRef]
- Rawla, P. Epidemiology of Prostate Cancer. World J. Oncol. 2019, 10, 63–89. [Google Scholar] [CrossRef] [PubMed]
- Yang, R.; Zhou, Y.; Wang, Y.; Du, C.; Wu, Y. Trends in cancer incidence and mortality rates in the United States from 1975 to 2016. Ann. Transl. Med. 2020, 8, 1671. [Google Scholar] [CrossRef] [PubMed]
- Teo, M.Y.; Rathkopf, D.E.; Kantoff, P. Treatment of Advanced Prostate Cancer. Annu. Rev. Med. 2019, 70, 479–499. [Google Scholar] [CrossRef] [PubMed]
- Cancel-Tassin, G.; Cussenot, O. Genetic susceptibility to prostate cancer. BJU Int. 2005, 96, 1380–1385. [Google Scholar] [CrossRef]
- Ewing, C.M.; Ray, A.M.; Lange, E.M.; Zuhlke, K.A.; Robbins, C.M.; Tembe, W.D.; Wiley, K.E.; Isaacs, S.D.; Johng, D.; Wang, Y.; et al. Germline mutations in HOXB13 and prostate-cancer risk. N. Engl. J. Med. 2012, 366, 141–149. [Google Scholar] [CrossRef]
- PDQ Cancer Genetics Editorial Board. Genetics of Prostate Cancer (PDQ®): Health Professional Version. Bethesda, MD: National Cancer Institute. Available online: https://www.cancer.gov/types/prostate/hp/prostate-genetics-pdq (accessed on 15 January 2021).
- Conti, D.V.; Darst, B.F.; Moss, L.C.; Saunders, E.J.; Sheng, X.; Chou, A.; Schumacher, F.R.; Olama, A.A.A.; Benlloch, S.; Dadaev, T.; et al. Trans-ancestry genome-wide association meta-analysis of prostate cancer identifies new susceptibility loci and informs genetic risk prediction. Nat. Genet. 2021, 53, 65–75. [Google Scholar] [CrossRef]
- Al Olama, A.A.; Kote-Jarai, Z.; Berndt, S.I.; Conti, D.V.; Schumacher, F.; Han, Y.; Benlloch, S.; Hazelett, D.J.; Wang, Z.; Saunders, E.; et al. A meta-analysis of 87,040 individuals identifies 23 new susceptibility loci for prostate cancer. Nat. Genet. 2014, 46, 1103–1109. [Google Scholar] [CrossRef]
- Zheng, S.L.; Sun, J.; Wiklund, F.; Smith, S.; Stattin, P.; Li, G.; Adami, H.O.; Hsu, F.C.; Zhu, Y.; Balter, K.; et al. Cumulative association of five genetic variants with prostate cancer. N. Engl. J. Med. 2008, 358, 910–919. [Google Scholar] [CrossRef]
- Li-Sheng Chen, S.; Ching-Yuan Fann, J.; Sipeky, C.; Yang, T.K.; Yueh-Hsia Chiu, S.; Ming-Fang Yen, A.; Laitinen, V.; Tammela, T.L.J.; Stenman, U.H.; Auvinen, A.; et al. Risk Prediction of Prostate Cancer with Single Nucleotide Polymorphisms and Prostate Specific Antigen. J. Urol. 2019, 201, 486–495. [Google Scholar] [CrossRef] [PubMed]
- Huynh-Le, M.P.; Fan, C.C.; Karunamuni, R.; Walsh, E.I.; Turner, E.L.; Lane, J.A.; Martin, R.M.; Neal, D.E.; Donovan, J.L.; Hamdy, F.C.; et al. A Genetic Risk Score to Personalize Prostate Cancer Screening, Applied to Population Data. Cancer Epidemiol. Biomark. Prev. 2020, 29, 1731–1738. [Google Scholar] [CrossRef] [PubMed]
- Nordstrom, T.; Aly, M.; Eklund, M.; Egevad, L.; Gronberg, H. A genetic score can identify men at high risk for prostate cancer among men with prostate-specific antigen of 1–3 ng/mL. Eur. Urol. 2014, 65, 1184–1190. [Google Scholar] [CrossRef]
- Black, M.H.; Li, S.; LaDuca, H.; Lo, M.T.; Chen, J.; Hoiness, R.; Gutierrez, S.; Tippin-Davis, B.; Lu, H.M.; Gielzak, M.; et al. Validation of a prostate cancer polygenic risk score. Prostate 2020, 80, 1314–1321. [Google Scholar] [CrossRef]
- Giglia-Mari, G.; Zotter, A.; Vermeulen, W. DNA damage response. Cold Spring Harb. Perspect. Biol. 2011, 3, a000745. [Google Scholar] [CrossRef]
- Pritchard, C.C.; Mateo, J.; Walsh, M.F.; De Sarkar, N.; Abida, W.; Beltran, H.; Garofalo, A.; Gulati, R.; Carreira, S.; Eeles, R.; et al. Inherited DNA-Repair Gene Mutations in Men with Metastatic Prostate Cancer. N. Engl. J. Med. 2016, 375, 443–453. [Google Scholar] [CrossRef] [PubMed]
- Darst, B.F.; Dadaev, T.; Saunders, E.; Sheng, X.; Wan, P.; Pooler, L.; Xia, L.Y.; Chanock, S.; Berndt, S.I.; Gapstur, S.M.; et al. Germline sequencing DNA repair genes in 5545 men with aggressive and non-aggressive prostate cancer. J. Natl. Cancer Inst. 2020. [Google Scholar] [CrossRef]
- Robinson, D.; Van Allen, E.M.; Wu, Y.M.; Schultz, N.; Lonigro, R.J.; Mosquera, J.M.; Montgomery, B.; Taplin, M.E.; Pritchard, C.C.; Attard, G.; et al. Integrative clinical genomics of advanced prostate cancer. Cell 2015, 161, 1215–1228. [Google Scholar] [CrossRef] [PubMed]
- Agalliu, I.; Karlins, E.; Kwon, E.M.; Iwasaki, L.M.; Diamond, A.; Ostrander, E.A.; Stanford, J.L. Rare germline mutations in the BRCA2 gene are associated with early-onset prostate cancer. Br. J. Cancer 2007, 97, 826–831. [Google Scholar] [CrossRef]
- Momozawa, Y.; Iwasaki, Y.; Hirata, M.; Liu, X.; Kamatani, Y.; Takahashi, A.; Sugano, K.; Yoshida, T.; Murakami, Y.; Matsuda, K.; et al. Germline Pathogenic Variants in 7636 Japanese Patients with Prostate Cancer and 12,366 Controls. J. Natl. Cancer Inst. 2020, 112, 369–376. [Google Scholar] [CrossRef] [PubMed]
- Mitra, A.; Fisher, C.; Foster, C.S.; Jameson, C.; Barbachanno, Y.; Bartlett, J.; Bancroft, E.; Doherty, R.; Kote-Jarai, Z.; Peock, S.; et al. Prostate cancer in male BRCA1 and BRCA2 mutation carriers has a more aggressive phenotype. Br. J. Cancer 2008, 98, 502–507. [Google Scholar] [CrossRef] [PubMed]
- Agalliu, I.; Gern, R.; Leanza, S.; Burk, R.D. Associations of high-grade prostate cancer with BRCA1 and BRCA2 founder mutations. Clin. Cancer Res. 2009, 15, 1112–1120. [Google Scholar] [CrossRef]
- Isaacsson Velho, P.; Silberstein, J.L.; Markowski, M.C.; Luo, J.; Lotan, T.L.; Isaacs, W.B.; Antonarakis, E.S. Intraductal/ductal histology and lymphovascular invasion are associated with germline DNA-repair gene mutations in prostate cancer. Prostate 2018, 78, 401–407. [Google Scholar] [CrossRef]
- Nyberg, T.; Frost, D.; Barrowdale, D.; Evans, D.G.; Bancroft, E.; Adlard, J.; Ahmed, M.; Barwell, J.; Brady, A.F.; Brewer, C.; et al. Prostate Cancer Risks for Male BRCA1 and BRCA2 Mutation Carriers: A Prospective Cohort Study. Eur. Urol. 2020, 77, 24–35. [Google Scholar] [CrossRef] [PubMed]
- Leongamornlert, D.; Mahmud, N.; Tymrakiewicz, M.; Saunders, E.; Dadaev, T.; Castro, E.; Goh, C.; Govindasami, K.; Guy, M.; O’Brien, L.; et al. Germline BRCA1 mutations increase prostate cancer risk. Br. J. Cancer 2012, 106, 1697–1701. [Google Scholar] [CrossRef]
- Na, R.; Zheng, S.L.; Han, M.; Yu, H.; Jiang, D.; Shah, S.; Ewing, C.M.; Zhang, L.; Novakovic, K.; Petkewicz, J.; et al. Germline Mutations in ATM and BRCA1/2 Distinguish Risk for Lethal and Indolent Prostate Cancer and are Associated with Early Age at Death. Eur. Urol. 2017, 71, 740–747. [Google Scholar] [CrossRef]
- Castro, E.; Goh, C.; Olmos, D.; Saunders, E.; Leongamornlert, D.; Tymrakiewicz, M.; Mahmud, N.; Dadaev, T.; Govindasami, K.; Guy, M.; et al. Germline BRCA mutations are associated with higher risk of nodal involvement, distant metastasis, and poor survival outcomes in prostate cancer. J. Clin. Oncol. 2013, 31, 1748–1757. [Google Scholar] [CrossRef]
- Castro, E.; Goh, C.; Leongamornlert, D.; Saunders, E.; Tymrakiewicz, M.; Dadaev, T.; Govindasami, K.; Guy, M.; Ellis, S.; Frost, D.; et al. Effect of BRCA Mutations on Metastatic Relapse and Cause-specific Survival After Radical Treatment for Localised Prostate Cancer. Eur. Urol. 2015, 68, 186–193. [Google Scholar] [CrossRef]
- Tryggvadottir, L.; Vidarsdottir, L.; Thorgeirsson, T.; Jonasson, J.G.; Olafsdottir, E.J.; Olafsdottir, G.H.; Rafnar, T.; Thorlacius, S.; Jonsson, E.; Eyfjord, J.E.; et al. Prostate cancer progression and survival in BRCA2 mutation carriers. J. Natl. Cancer Inst. 2007, 99, 929–935. [Google Scholar] [CrossRef] [PubMed]
- Narod, S.A.; Neuhausen, S.; Vichodez, G.; Armel, S.; Lynch, H.T.; Ghadirian, P.; Cummings, S.; Olopade, O.; Stoppa-Lyonnet, D.; Couch, F.; et al. Rapid progression of prostate cancer in men with a BRCA2 mutation. Br. J. Cancer 2008, 99, 371–374. [Google Scholar] [CrossRef] [PubMed]
- Edwards, S.M.; Evans, D.G.; Hope, Q.; Norman, A.R.; Barbachano, Y.; Bullock, S.; Kote-Jarai, Z.; Meitz, J.; Falconer, A.; Osin, P.; et al. Prostate cancer in BRCA2 germline mutation carriers is associated with poorer prognosis. Br. J. Cancer 2010, 103, 918–924. [Google Scholar] [CrossRef] [PubMed]
- Taylor, R.A.; Fraser, M.; Livingstone, J.; Espiritu, S.M.; Thorne, H.; Huang, V.; Lo, W.; Shiah, Y.J.; Yamaguchi, T.N.; Sliwinski, A.; et al. Germline BRCA2 mutations drive prostate cancers with distinct evolutionary trajectories. Nat. Commun. 2017, 8, 13671. [Google Scholar] [CrossRef] [PubMed]
- Karlsson, Q.; Brook, M.N.; Dadaev, T.; Wakerell, S.; Saunders, E.J.; Muir, K.; Neal, D.E.; Giles, G.G.; MacInnis, R.J.; Thibodeau, S.N.; et al. Rare Germline Variants in ATM Predispose to Prostate Cancer: A PRACTICAL Consortium Study. Eur. Urol. Oncol. 2021. [Google Scholar] [CrossRef] [PubMed]
- Nicolosi, P.; Ledet, E.; Yang, S.; Michalski, S.; Freschi, B.; O’Leary, E.; Esplin, E.D.; Nussbaum, R.L.; Sartor, O. Prevalence of Germline Variants in Prostate Cancer and Implications for Current Genetic Testing Guidelines. JAMA Oncol. 2019, 5, 523–528. [Google Scholar] [CrossRef]
- Chung, J.H.; Dewal, N.; Sokol, E.; Mathew, P.; Whitehead, R.; Millis, S.Z.; Frampton, G.M.; Bratslavsky, G.; Pal, S.K.; Lee, R.J.; et al. Prospective Comprehensive Genomic Profiling of Primary and Metastatic Prostate Tumors. JCO Precis. Oncol. 2019, 3, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Sedhom, R.; Antonarakis, E.S. Clinical implications of mismatch repair deficiency in prostate cancer. Future Oncol. 2019, 15, 2395–2411. [Google Scholar] [CrossRef] [PubMed]
- Ryan, S.; Jenkins, M.A.; Win, A.K. Risk of prostate cancer in Lynch syndrome: A systematic review and meta-analysis. Cancer Epidemiol. Biomark. Prev. 2014, 23, 437–449. [Google Scholar] [CrossRef]
- Dominguez-Valentin, M.; Sampson, J.R.; Seppala, T.T.; Ten Broeke, S.W.; Plazzer, J.P.; Nakken, S.; Engel, C.; Aretz, S.; Jenkins, M.A.; Sunde, L.; et al. Cancer risks by gene, age, and gender in 6350 carriers of pathogenic mismatch repair variants: Findings from the Prospective Lynch Syndrome Database. Genet. Med. 2020, 22, 15–25. [Google Scholar] [CrossRef]
- Martin-Lopez, J.V.; Fishel, R. The mechanism of mismatch repair and the functional analysis of mismatch repair defects in Lynch syndrome. Fam. Cancer 2013, 12, 159–168. [Google Scholar] [CrossRef] [PubMed]
- Antonarakis, E.S.; Shaukat, F.; Isaacsson Velho, P.; Kaur, H.; Shenderov, E.; Pardoll, D.M.; Lotan, T.L. Clinical Features and Therapeutic Outcomes in Men with Advanced Prostate Cancer and DNA Mismatch Repair Gene Mutations. Eur. Urol. 2019, 75, 378–382. [Google Scholar] [CrossRef]
- Cai, Q.; Wang, X.; Li, X.; Gong, R.; Guo, X.; Tang, Y.; Yang, K.; Niu, Y.; Zhao, Y. Germline HOXB13 p.Gly84Glu mutation and cancer susceptibility: A pooled analysis of 25 epidemiological studies with 145,257 participates. Oncotarget 2015, 6, 42312–42321. [Google Scholar] [CrossRef]
- Kote-Jarai, Z.; Mikropoulos, C.; Leongamornlert, D.A.; Dadaev, T.; Tymrakiewicz, M.; Saunders, E.J.; Jones, M.; Jugurnauth-Little, S.; Govindasami, K.; Guy, M.; et al. Prevalence of the HOXB13 G84E germline mutation in British men and correlation with prostate cancer risk, tumour characteristics and clinical outcomes. Ann. Oncol. 2015, 26, 756–761. [Google Scholar] [CrossRef]
- Wei, J.; Shi, Z.; Na, R.; Wang, C.H.; Resurreccion, W.K.; Zheng, S.L.; Hulick, P.J.; Cooney, K.A.; Helfand, B.T.; Isaacs, W.B.; et al. Germline HOXB13 G84E mutation carriers and risk to twenty common types of cancer: Results from the UK Biobank. Br. J. Cancer 2020, 123, 1356–1359. [Google Scholar] [CrossRef] [PubMed]
- Petrovics, G.; Price, D.K.; Lou, H.; Chen, Y.; Garland, L.; Bass, S.; Jones, K.; Kohaar, I.; Ali, A.; Ravindranath, L.; et al. Increased frequency of germline BRCA2 mutations associates with prostate cancer metastasis in a racially diverse patient population. Prostate Cancer Prostatic Dis. 2019, 22, 406–410. [Google Scholar] [CrossRef] [PubMed]
- Sartor, O.; Yang, S.; Ledet, E.; Moses, M.; Nicolosi, P. Inherited DNA-repair gene mutations in African American men with prostate cancer. Oncotarget 2020, 11, 440–442. [Google Scholar] [CrossRef] [PubMed][Green Version]
- National Comprehensive Cancer Network. NCCN Guidelines: Prostate Cancer; Version 2.2021; National Comprehensive Cancer Network: Plymouth Meeting, PA, USA, 2020. [Google Scholar]
- Giri, V.N.; Knudsen, K.E.; Kelly, W.K.; Cheng, H.H.; Cooney, K.A.; Cookson, M.S.; Dahut, W.; Weissman, S.; Soule, H.R.; Petrylak, D.P.; et al. Implementation of Germline Testing for Prostate Cancer: Philadelphia Prostate Cancer Consensus Conference 2019. J. Clin. Oncol. 2020, 38, 2798–2811. [Google Scholar] [CrossRef]
- Carlo, M.I.; Giri, V.N.; Paller, C.J.; Abida, W.; Alumkal, J.J.; Beer, T.M.; Beltran, H.; George, D.J.; Heath, E.I.; Higano, C.S.; et al. Evolving Intersection Between Inherited Cancer Genetics and Therapeutic Clinical Trials in Prostate Cancer: A White Paper From the Germline Genetics Working Group of the Prostate Cancer Clinical Trials Consortium. JCO Precis Oncol. 2018, 2018, 1–14. [Google Scholar] [CrossRef]
- Gillessen, S.; Attard, G.; Beer, T.M.; Beltran, H.; Bjartell, A.; Bossi, A.; Briganti, A.; Bristow, R.G.; Chi, K.N.; Clarke, N.; et al. Management of Patients with Advanced Prostate Cancer: Report of the Advanced Prostate Cancer Consensus Conference 2019. Eur. Urol. 2020, 77, 508–547. [Google Scholar] [CrossRef] [PubMed]
- Loeb, S.; Giri, V.N. Clinical Implications of Germline Testing in Newly Diagnosed Prostate Cancer. Eur. Urol. Oncol. 2021, 4, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Invitae Prostate Cancer Panel. Available online: https://www.invitae.com/en/physician/tests/01362/ (accessed on 12 February 2021).
- Prostate Cancer Comprehensive Panel. Fulgent. Available online: https://www.fulgentgenetics.com/comprehensivecancer-prostate (accessed on 12 February 2021).
- ProstateNext. Ambry Genetics. Available online: https://www.ambrygen.com/providers/genetic-testing/5/oncology/prostatenext (accessed on 12 February 2021).
- Hereditary Prostate Cancer Panel. GeneDx. Available online: https://www.genedx.com/wp-content/uploads/2017/08/201340_Hereditary-Prostate-Cancer-Panel.pdf (accessed on 12 February 2021).
- Prostate Cancer Panel. Prevention Genetics. Available online: https://www.preventiongenetics.com/testInfo?val=Prostate+Cancer+Panel (accessed on 12 February 2021).
- Myriad mRisk Hereditary Cancer. Myriad Genetics, Inc. Available online: https://myriadmyrisk.com/ (accessed on 12 February 2021).
- Color—Overview for Providers. Available online: https://www.color.com/providers/about-color (accessed on 12 February 2021).
- Mateo, J.; Carreira, S.; Sandhu, S.; Miranda, S.; Mossop, H.; Perez-Lopez, R.; Nava Rodrigues, D.; Robinson, D.; Omlin, A.; Tunariu, N.; et al. DNA-Repair Defects and Olaparib in Metastatic Prostate Cancer. N. Engl. J. Med. 2015, 373, 1697–1708. [Google Scholar] [CrossRef] [PubMed]
- Mateo, J.; Porta, N.; Bianchini, D.; McGovern, U.; Elliott, T.; Jones, R.; Syndikus, I.; Ralph, C.; Jain, S.; Varughese, M.; et al. Olaparib in patients with metastatic castration-resistant prostate cancer with DNA repair gene aberrations (TOPARP-B): A multicentre, open-label, randomised, phase 2 trial. Lancet Oncol. 2020, 21, 162–174. [Google Scholar] [CrossRef]
- De Bono, J.; Mateo, J.; Fizazi, K.; Saad, F.; Shore, N.; Sandhu, S.; Chi, K.N.; Sartor, O.; Agarwal, N.; Olmos, D.; et al. Olaparib for Metastatic Castration-Resistant Prostate Cancer. N. Engl. J. Med. 2020, 382, 2091–2102. [Google Scholar] [CrossRef] [PubMed]
- Abida, W.; Patnaik, A.; Campbell, D.; Shapiro, J.; Bryce, A.H.; McDermott, R.; Sautois, B.; Vogelzang, N.J.; Bambury, R.M.; Voog, E.; et al. Rucaparib in Men with Metastatic Castration-Resistant Prostate Cancer Harboring a BRCA1 or BRCA2 Gene Alteration. J. Clin. Oncol. 2020, 38, 3763–3772. [Google Scholar] [CrossRef]
- Abida, W.; Campbell, D.; Patnaik, A.; Shapiro, J.D.; Sautois, B.; Vogelzang, N.J.; Voog, E.G.; Bryce, A.H.; McDermott, R.; Ricci, F.; et al. Non-BRCA DNA Damage Repair Gene Alterations and Response to the PARP Inhibitor Rucaparib in Metastatic Castration-Resistant Prostate Cancer: Analysis from the Phase II TRITON2 Study. Clin. Cancer Res. 2020, 26, 2487–2496. [Google Scholar] [CrossRef] [PubMed]
- Smith, M.R.; Sandhu, S.; Kelly, W.K.; Scher, H.I.; Efstathiou, E.; Lara, P.N.; Yu, E.Y.; George, D.J.; Chi, K.N.; Saad, F.; et al. Pre-specified interim analysis of GALAHAD: A phase II study of niraparib in patients (pts) with metastatic castration-resistant prostate cancer (mCRPC) and biallelic DNA-repair gene defects (DRD). Ann. Oncol. 2019, 30 (Suppl. 5), V884–V885. [Google Scholar] [CrossRef]
- Le, D.T.; Durham, J.N.; Smith, K.N.; Wang, H.; Bartlett, B.R.; Aulakh, L.K.; Lu, S.; Kemberling, H.; Wilt, C.; Luber, B.S.; et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science 2017, 357, 409–413. [Google Scholar] [CrossRef]
- Abida, W.; Cheng, M.L.; Armenia, J.; Middha, S.; Autio, K.A.; Vargas, H.A.; Rathkopf, D.; Morris, M.J.; Danila, D.C.; Slovin, S.F.; et al. Analysis of the Prevalence of Microsatellite Instability in Prostate Cancer and Response to Immune Checkpoint Blockade. JAMA Oncol. 2019, 5, 471–478. [Google Scholar] [CrossRef] [PubMed]
- Barata, P.; Agarwal, N.; Nussenzveig, R.; Gerendash, B.; Jaeger, E.; Hatton, W.; Ledet, E.; Lewis, B.; Layton, J.; Babiker, H.; et al. Clinical activity of pembrolizumab in metastatic prostate cancer with microsatellite instability high (MSI-H) detected by circulating tumor DNA. J. Immunother. Cancer 2020, 8, e001065. [Google Scholar] [CrossRef] [PubMed]
- FDA. List of Cleared or Approved Companion Diagnostic Devices (In Vitro and Imaging Tools). Available online: https://www.fda.gov/medical-devices/vitro-diagnostics/list-cleared-or-approved-companion-diagnostic-devices-vitro-and-imaging-tools (accessed on 1 March 2021).
- BRACAnalysis CDx. Myriad Genetics, Inc. Available online: https://myriad.com/managed-care/bracanalysis-cdx/ (accessed on 12 February 2021).
- FoundationOne CDx. Available online: https://www.foundationmedicine.com/test/foundationone-cdx (accessed on 12 February 2021).
- FoundationOne Liquid CDx. Available online: https://www.foundationmedicine.com/test/foundationone-liquid-cdx (accessed on 12 February 2021).
- Jacobs, M.F.; Milliron, K.J. Genetic counseling and previvorship in patients with urologic malignancies. Curr. Opin. Urol. 2019, 29, 371–377. [Google Scholar] [CrossRef]
- Neviere, Z.; De La Motte Rouge, T.; Floquet, A.; Johnson, A.; Berthet, P.; Joly, F. How and when to refer patients for oncogenetic counseling in the era of PARP inhibitors. Ther. Adv. Med. Oncol. 2020, 12, 1758835919897530. [Google Scholar] [CrossRef]
- Cheng, H.; Powers, J.; Schaffer, K.; Sartor, O. Practical Methods for Integrating Genetic Testing into Clinical Practice for Advanced Prostate Cancer. Am. Soc. Clin. Oncol. Educ. Book 2018, 38, 372–381. [Google Scholar] [CrossRef] [PubMed]
- Paller, C.J.; Antonarakis, E.S.; Beer, T.M.; Borno, H.T.; Carlo, M.I.; George, D.J.; Graff, J.N.; Gupta, S.; Heath, E.I.; Higano, C.S.; et al. Germline Genetic Testing in Advanced Prostate Cancer; Practices and Barriers: Survey Results from the Germline Genetics Working Group of the Prostate Cancer Clinical Trials Consortium. Clin. Genitourin Cancer 2019, 17, 275–282.e1. [Google Scholar] [CrossRef] [PubMed]
- Suri, Y.; Nandagopal, L.; Basu, A. Inadequacies in genetic testing referrals and counseling in prostate cancer. J. Clin. Oncol. 2020, 38 (Suppl. 29), 43. [Google Scholar] [CrossRef]
| Prevalence of Germline Variants (%) | |||||
|---|---|---|---|---|---|
| Gene | Established Association with PCa Risk | Pritchard et al. [17] | Nicolosi et al. [35] | Robinson et al. [19] | * Chung et al. [36] |
| Patients with mPC (n = 692) | Patients with PC (n = 3607) | Patients with mCRPC (n = 150) | Primary or Metastatic PC Tumor Samples (n = 3476) | ||
| ATM | yes | 1.6 | 2.0 | 0.67 | 1.9 |
| BRCA1 | yes | 0.9 | 1.3 | 0.33 | 0.35 |
| BRCA2 | yes | 5.4 | 4.7 | 5.3 | 5.7 |
| BRIP1 | - | 0.2 | 0.3 | - | - |
| CHEK2 | yes | 1.9 | 2.9 | - | 1.4 |
| FANCA | - | - | - | - | 0.68 |
| NBN | - | 0.3 | 0.3 | - | - |
| PALB2 | - | 0.4 | 0.7 | - | - |
| RAD51B | - | 0.3 | - | - | - |
| RAD51C | - | 0.14 | 0.2 | - | - |
| RAD51D | - | 0.4 | 0.2 | - | - |
| Prostate Cancer Genetic Tests | |||
|---|---|---|---|
| Test Name | Company | Genes Tested | Type of Sample |
| Prostate Cancer Panel [52] | Invitae | ATM, BRCA1, BRCA2, CHEK2, EPCAM, HOXB13, MLH1, MSH2, MS6, NBN, PMS2, TP53 -Add-on panel (ATR, BRIP1, FANCA, GEN1, PALB2, RAD51C, RAD51D) | Whole Blood or Buccal Swab or Saliva |
| Prostate Cancer Comprehensive Panel [53] | Fulgent | ATM, BRCA1, BRCA2, CHEK2, EPCAM, HOXB13, MLH1, MSH2, MSH6, NBN, PMS2, TP53 | Whole Blood or Buccal Swab or Saliva |
| Prostate Next [54] | Ambry Genetics | ATM, BRCA1, BRCA2, CHEK2, EPCAM, HOXB13, MLH1, MSH2, MSH6, NBN, PALB2, PMS2, RAD51D, TP53 | Whole Blood |
| Hereditary Prostate Cancer Panel [55] | GeneDX | ATM, BRCA1, BRCA2, BRIP1, CHEK2, EPCAM, HOXB13, MLH1, MSH2, MSH6, NBN, PALB2, PMS2, RAD51C, RAD51D, TP53 | Whole Blood or Buccal Swab |
| Prostate Cancer Panel [56] | Prevention Genetics | ATM, BRCA1, BRCA2, BRIP1, CHEK2, EPCAM, HOXB13, MLH1, MSH2, MSH6, NBN, PALB2, PMS2, RAD51C, RAD51D, TP53 | Whole Blood |
| myRisk [57] | Myriad Genetics | APC, ATM, AXIN2, BARD1, BMPR1A, BRCA1, BRCA2, BRIP1, CDH1, CDK4, CDKN2A, CHEK2, EPCAM, GREM1, HOXB13 GALNT12, MLH1, MSH2, MSH3, MSH6, MUTYH, NBN, NTHL1, PALB2, PMS2, POLE, POLD1, PTEN, RAD51C, RAD51D, RNF43, RPS20, SMAD4, STK11, TP53 | Whole Blood or Saliva |
| Hereditary Cancer Panel [58] | Color | APC, ATM, BAP1, BARD1, BMPR1A, BRCA1, BRCA2, BRIP1, CDH1, CDK4, CDKN2A, CHEK2, EPCAM, GREM1, MITF, MLH1, MSH2, MSH6, MUTYH, NBN, PALB2, PMS2, POLE, POLD1, PTEN, RAD51C, RAD51D, SMAD4, STK11, TP53 | Saliva |
| Companion Diagnostics | |||
|---|---|---|---|
| Test Name | Company | Genes Tested | Type of Sample |
| * BRACAnalysis CDx [69] | Myriad Genetics | BRCA1, BRCA2 | Whole Blood |
| * FoundationOne CDx [70] | Foundation Medicine | BRCA1, BRCA2, ATM, BARD1, BRIP1, CDK12, CHEK1, CHEK2, FANCL, PALB2, RAD51B, RAD51C, RAD51D and RAD54L | Biopsy # |
| ** FoundationOne Liquid CDx [71] | Foundation Medicine | BRCA1, BRCA2, ATM | Whole blood |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Doan, D.K.; Schmidt, K.T.; Chau, C.H.; Figg, W.D. Germline Genetics of Prostate Cancer: Prevalence of Risk Variants and Clinical Implications for Disease Management. Cancers 2021, 13, 2154. https://doi.org/10.3390/cancers13092154
Doan DK, Schmidt KT, Chau CH, Figg WD. Germline Genetics of Prostate Cancer: Prevalence of Risk Variants and Clinical Implications for Disease Management. Cancers. 2021; 13(9):2154. https://doi.org/10.3390/cancers13092154
Chicago/Turabian StyleDoan, David K., Keith T. Schmidt, Cindy H. Chau, and William D. Figg. 2021. "Germline Genetics of Prostate Cancer: Prevalence of Risk Variants and Clinical Implications for Disease Management" Cancers 13, no. 9: 2154. https://doi.org/10.3390/cancers13092154
APA StyleDoan, D. K., Schmidt, K. T., Chau, C. H., & Figg, W. D. (2021). Germline Genetics of Prostate Cancer: Prevalence of Risk Variants and Clinical Implications for Disease Management. Cancers, 13(9), 2154. https://doi.org/10.3390/cancers13092154







