The Interplay between Long Noncoding RNAs and Proteins of the Epigenetic Machinery in Ovarian Cancer
Simple Summary
Abstract
1. Introduction
2. LncRNA Genes and Their Expression Patterns
3. LncRNAs and Their Association with Ovarian Cancer
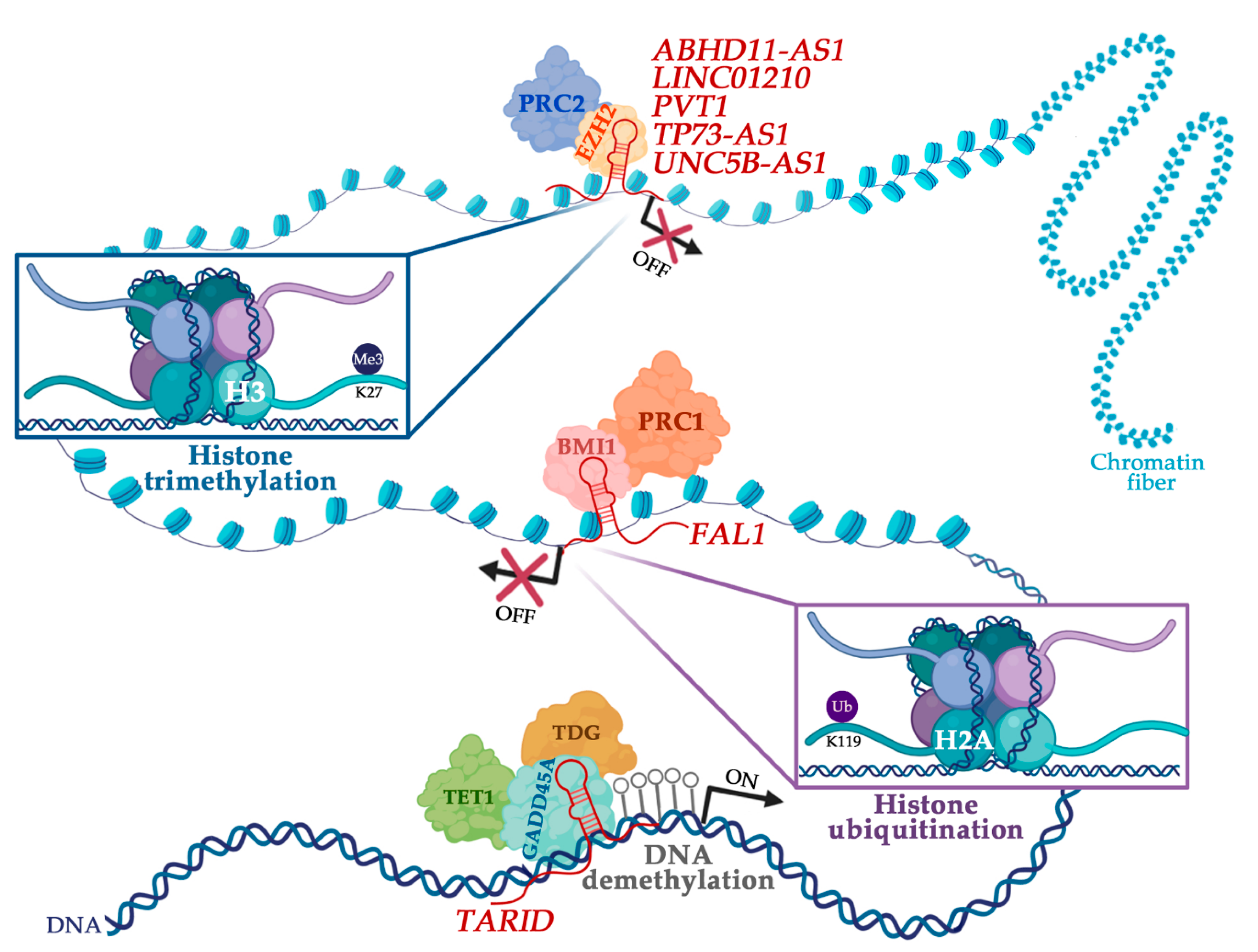
4. LncRNAs Involved in Epigenetic Mechanisms in Ovarian Cancer
| LncRNA | Regulatory Mechanism | Target | Source | Refs. |
|---|---|---|---|---|
| FAL1 (FALEC) | Stabilization of BMI1 protein and PRC1-mediated deposition of repressive histone marks | CDKN1A | LncTarD and EVLncRNAs databases | [84,85] |
| PVT1 | EZH2 recruitment | miR-214 | LncTarD database | [86] |
| TARID | Recruitment of GADD45A and TET-dependent DNA demethylation | TCF21 | LncTarD database | [87,88] |
| LINC01210 | EZH2 recruitment and PRC2-mediated deposition of repressive histone marks | KLF4 | Literature mining | [89] |
| TP73-AS1 | EZH2 recruitment and PRC2-mediated deposition of repressive histone marks | CDKN1A | Literature mining | [90] |
| UNC5B-AS1 | EZH2 recruitment and PRC2-mediated deposition of repressive histone marks | NDRG2 | Literature mining | [91] |
| ABHD11-AS1 | EZH2 recruitment and PRC2-mediated deposition of repressive histone marks | TIMP2 | Literature mining | [92] |
5. A Concise Guide to Exploring lncRNA Databases
6. Conclusions and Perspectives
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Timmermans, M.; Sonke, G.S.; Van de Vijver, K.K.; van der Aa, M.A.; Kruitwagen, R.F.P.M. No improvement in long-term survival for epithelial ovarian cancer patients: A population-based study between 1989 and 2014 in the Netherlands. Eur. J. Cancer 2018, 88, 31–37. [Google Scholar] [CrossRef] [PubMed]
- Sant, M.; Chirlaque Lopez, M.D.; Agresti, R.; Sánchez Pérez, M.J.; Holleczek, B.; Bielska-Lasota, M.; Dimitrova, N.; Innos, K.; Katalinic, A.; Langseth, H.; et al. Survival of women with cancers of breast and genital organs in Europe 1999–2007: Results of the EUROCARE-5 study. Eur. J. Cancer 2015, 51, 2191–2205. [Google Scholar] [CrossRef] [PubMed]
- Global Cancer Observatory GLOBOCAN 2018: Ovary Fact Sheet. Available online: https://gco.iarc.fr/today/data/factsheets/cancers/25-Ovary-fact-sheet.pdf (accessed on 24 February 2020).
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Blagden, S.P. Harnessing Pandemonium: The Clinical Implications of Tumor Heterogeneity in Ovarian Cancer. Front. Oncol. 2015, 5, 149. [Google Scholar] [CrossRef]
- McCluggage, W.G. Morphological subtypes of ovarian carcinoma: A review with emphasis on new developments and pathogenesis. Pathology 2011, 43, 420–432. [Google Scholar] [CrossRef]
- Lisio, M.A.; Fu, L.; Goyeneche, A.; Gao, Z.H.; Telleria, C. High-grade serous ovarian cancer: Basic sciences, clinical and therapeutic standpoints. Int. J. Mol. Sci. 2019, 20, 952. [Google Scholar] [CrossRef]
- Matulonis, U.A.; Sood, A.K.; Fallowfield, L.; Howitt, B.E.; Sehouli, J.; Karlan, B.Y. Ovarian cancer. Nat. Rev. Dis. Prim. 2016, 2, 16061. [Google Scholar] [CrossRef]
- Menon, U.; Karpinskyj, C.; Gentry-Maharaj, A. Ovarian Cancer Prevention and Screening. Obstet. Gynecol. 2018, 131, 909–927. [Google Scholar] [CrossRef]
- Goff, B.A.; Mandel, L.; Muntz, H.G.; Melancon, C.H. Ovarian carcinoma diagnosis. Cancer 2000, 89, 2068–2075. [Google Scholar] [CrossRef]
- Tan, D.S.; Agarwal, R.; Kaye, S.B. Mechanisms of transcoelomic metastasis in ovarian cancer. Lancet Oncol. 2006, 7, 925–934. [Google Scholar] [CrossRef]
- Cancer Research UK Ovarian Cancer Survival Statistics. Available online: https://www.cancerresearchuk.org/health-professional/cancer-statistics/statistics-by-cancer-type/ovarian-cancer/survival (accessed on 24 February 2020).
- Moore, K.; Colombo, N.; Scambia, G.; Kim, B.-G.; Oaknin, A.; Friedlander, M.; Lisyanskaya, A.; Floquet, A.; Leary, A.; Sonke, G.S.; et al. Maintenance Olaparib in Patients with Newly Diagnosed Advanced Ovarian Cancer. N. Engl. J. Med. 2018, 379, 2495–2505. [Google Scholar] [CrossRef] [PubMed]
- Boyd, L.R.; Muggia, F.M. Carboplatin/Paclitaxel Induction in Ovarian Cancer: The Finer Points. Oncology 2018, 32, 418–420, 422–424. [Google Scholar] [PubMed]
- Piovesan, A.; Antonaros, F.; Vitale, L.; Strippoli, P.; Pelleri, M.C.; Caracausi, M. Human protein-coding genes and gene feature statistics in 2019. BMC Res. Notes 2019, 12, 315. [Google Scholar] [CrossRef] [PubMed]
- Dunham, I.; Kundaje, A.; Aldred, S.F.; Collins, P.J.; Davis, C.A.; Doyle, F.; Epstein, C.B.; Frietze, S.; Harrow, J.; Kaul, R.; et al. An integrated encyclopedia of DNA elements in the human genome. Nature 2012, 489, 57–74. [Google Scholar] [CrossRef]
- Rasool, M.; Malik, A.; Zahid, S.; Basit Ashraf, M.A.; Qazi, M.H.; Asif, M.; Zaheer, A.; Arshad, M.; Raza, A.; Jamal, M.S. Non-coding RNAs in cancer diagnosis and therapy. NonCoding RNA Res. 2016, 1, 69–76. [Google Scholar] [CrossRef]
- Derrien, T.; Johnson, R.; Bussotti, G.; Tanzer, A.; Djebali, S.; Tilgner, H.; Guernec, G.; Martin, D.; Merkel, A.; Knowles, D.G.; et al. The GENCODE v7 catalog of human long noncoding RNAs: Analysis of their gene structure, evolution, and expression. Genome Res. 2012, 22, 1775–1789. [Google Scholar] [CrossRef]
- Rasmussen, T.P. Parallels between artificial reprogramming and the biogenesis of cancer stem cells: Involvement of lncRNAs. Semin. Cancer Biol. 2019, 57, 36–44. [Google Scholar] [CrossRef]
- Fu, D.; Shi, Y.; Liu, J.-B.; Wu, T.-M.; Jia, C.-Y.; Yang, H.-Q.; Zhang, D.-D.; Yang, X.-L.; Wang, H.-M.; Ma, Y.-S. Targeting Long Non-coding RNA to Therapeutically Regulate Gene Expression in Cancer. Mol. Ther. Nucleic Acids 2020, 21, 712–724. [Google Scholar] [CrossRef]
- De Oliveira, J.C.; Oliveira, L.C.; Mathias, C.; Pedroso, G.A.; Lemos, D.S.; Salviano-Silva, A.; Jucoski, T.S.; Lobo-Alves, S.C.; Zambalde, E.P.; Cipolla, G.A.; et al. Long non-coding RNAs in cancer: Another layer of complexity. J. Gene Med. 2019, 21, 1–18. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Zhang, X.-Z.; Liu, H.; Chen, S.-R. Mechanisms of Long Non-Coding RNAs in Cancers and Their Dynamic Regulations. Cancers 2020, 12, 1245. [Google Scholar] [CrossRef] [PubMed]
- Hanly, D.J.; Esteller, M.; Berdasco, M. Interplay between long non-coding RNAs and epigenetic machinery: Emerging targets in cancer? Philos. Trans. R. Soc. B Biol. Sci. 2018, 373, 20170074. [Google Scholar] [CrossRef] [PubMed]
- Kong, L.; Zhang, C. LncRNA DLX6-AS1 aggravates the development of ovarian cancer via modulating FHL2 by sponging miR-195-5p. Cancer Cell Int. 2020, 20, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Ding, F.; Zheng, G. LncRNA TMPO-AS1 promotes LCN2 transcriptional activity and exerts oncogenic functions in ovarian cancer. FASEB J. 2020, 34, 11382–11394. [Google Scholar] [CrossRef]
- Elsayed, A.M.; Amero, P.; Salama, S.A.; Abdelaziz, A.H.; Lopez-Berestein, G.; Rodriguez-Aguayo, C. Back to the Future: Rethinking the Great Potential of lncRNAS for Optimizing Chemotherapeutic Response in Ovarian Cancer. Cancers 2020, 12, 2406. [Google Scholar] [CrossRef]
- Silva, A.; Bullock, M.; Calin, G. The clinical relevance of long non-coding RNAs in cancer. Cancers 2015, 7, 2169–2182. [Google Scholar] [CrossRef]
- Bhan, A.; Mandal, S.S. LncRNA HOTAIR: A master regulator of chromatin dynamics and cancer. Biochim. Biophys. Acta Rev. Cancer 2015, 1856, 151–164. [Google Scholar] [CrossRef]
- Wang, H.; Fu, Z.; Dai, C.; Cao, J.; Liu, X.; Xu, J.; Lv, M.; Gu, Y.; Zhang, J.; Hua, X.; et al. LncRNAs expression profiling in normal ovary, benign ovarian cyst and malignant epithelial ovarian cancer. Sci. Rep. 2016, 6, 38983. [Google Scholar] [CrossRef]
- Abildgaard, C.; Do Canto, L.M.; Steffensen, K.D.; Rogatto, S.R. Long Non-coding RNAs Involved in Resistance to Chemotherapy in Ovarian Cancer. Front. Oncol. 2020, 9, 1–17. [Google Scholar] [CrossRef]
- Xu, L.; Wu, Y.; Che, X.; Zhao, J.; Wang, F.; Wang, P.; Qu, X.; Liu, Y.P.; Li, Z. Cox-LASSO Analysis Reveals a Ten-lncRNA Signature to Predict Outcomes in Patients with High-Grade Serous Ovarian Cancer. DNA Cell Biol. 2019, 38, 1519–1528. [Google Scholar] [CrossRef]
- Teschendorff, A.E.; Lee, S.H.; Jones, A.; Fiegl, H.; Kalwa, M.; Wagner, W.; Chindera, K.; Evans, I.; Dubeau, L.; Orjalo, A.; et al. HOTAIR and its surrogate DNA methylation signature indicate carboplatin resistance in ovarian cancer. Genome Med. 2015, 7, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Lu, A.; Chen, L. LncRNAs in ovarian cancer. Clin. Chim. Acta 2019, 490, 17–27. [Google Scholar] [CrossRef] [PubMed]
- Zhan, L.; Li, J.; Wei, B. Long non-coding RNAs in ovarian cancer. J. Exp. Clin. Cancer Res. 2018, 37, 120. [Google Scholar] [CrossRef] [PubMed]
- Nam, J.W.; Choi, S.W.; You, B.H. Incredible RNA: Dual Functions of Coding and Noncoding. Mol. Cells 2016, 39, 367–374. [Google Scholar] [CrossRef] [PubMed]
- Wu, P.; Mo, Y.; Peng, M.; Tang, T.; Zhong, Y.; Deng, X.; Xiong, F.; Guo, C.; Wu, X.; Li, Y.; et al. Emerging role of tumor-related functional peptides encoded by lncRNA and circRNA. Mol. Cancer 2020, 19, 22. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Liu, C. Coding or Noncoding, the Converging Concepts of RNAs. Front. Genet. 2019, 10, 1–10. [Google Scholar] [CrossRef]
- Iyer, M.K.; Niknafs, Y.S.; Malik, R.; Singhal, U.; Sahu, A.; Hosono, Y.; Barrette, T.R.; Prensner, J.R.; Evans, J.R.; Zhao, S.; et al. The landscape of long noncoding RNAs in the human transcriptome. Nat. Genet. 2015, 47, 199–208. [Google Scholar] [CrossRef]
- Djebali, S.; Davis, C.A.; Merkel, A.; Dobin, A.; Lassmann, T.; Mortazavi, A.; Tanzer, A.; Lagarde, J.; Lin, W.; Schlesinger, F.; et al. Landscape of transcription in human cells. Nature 2012, 489, 101–108. [Google Scholar] [CrossRef]
- Dhanoa, J.K.; Sethi, R.S.; Verma, R.; Arora, J.S.; Mukhopadhyay, C.S. Long non-coding RNA: Its evolutionary relics and biological implications in mammals: A review. J. Anim. Sci. Technol. 2018, 60, 1–10. [Google Scholar] [CrossRef]
- Quinn, J.J.; Chang, H.Y. Unique features of long non-coding RNA biogenesis and function. Nat. Rev. Genet. 2016, 17, 47–62. [Google Scholar] [CrossRef]
- Ulitsky, I. Evolution to the rescue: Using comparative genomics to understand long non-coding RNAs. Nat. Rev. Genet. 2016, 17, 601–614. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.C.; Chang, H.Y. Molecular Mechanisms of Long Noncoding RNAs. Mol. Cell 2011, 43, 904–914. [Google Scholar] [CrossRef] [PubMed]
- Cabili, M.N.; Dunagin, M.C.; McClanahan, P.D.; Biaesch, A.; Padovan-Merhar, O.; Regev, A.; Rinn, J.L.; Raj, A. Localization and abundance analysis of human lncRNAs at single-cell and single-molecule resolution. Genome Biol. 2015, 16, 20. [Google Scholar] [CrossRef] [PubMed]
- Morlando, M.; Fatica, A. Alteration of epigenetic regulation by long noncoding RNAs in cancer. Int. J. Mol. Sci. 2018, 19, 570. [Google Scholar] [CrossRef]
- Huang, M.-S.; Zhu, T.; Li, L.; Xie, P.; Li, X.; Zhou, H.-H.; Liu, Z.-Q. LncRNAs and CircRNAs from the same gene: Masterpieces of RNA splicing. Cancer Lett. 2018, 415, 49–57. [Google Scholar] [CrossRef]
- Kong, Y.; Hsieh, C.H.; Alonso, L.C. ANRIL: A lncRNA at the CDKN2A/B locus with roles in cancer and metabolic disease. Front. Endocrinol. 2018, 9, 1–13. [Google Scholar] [CrossRef]
- Burd, C.E.; Jeck, W.R.; Liu, Y.; Sanoff, H.K.; Wang, Z.; Sharpless, N.E. Expression of Linear and Novel Circular Forms of an INK4/ARF-Associated Non-Coding RNA Correlates with Atherosclerosis Risk. PLoS Genet. 2010, 6, e1001233. [Google Scholar] [CrossRef]
- Xu, Q.; Deng, B.; Li, M.; Chen, Y.; Zhuan, L. CircRNA-UBAP2 promotes the proliferation and inhibits apoptosis of ovarian cancer though miR-382-5p/PRPF8 axis. J. Ovarian Res. 2020, 13, 1–10. [Google Scholar] [CrossRef]
- Wang, J.; Wu, A.; Yang, B.; Zhu, X.; Teng, Y.; Ai, Z. Profiling and bioinformatics analyses reveal differential circular RNA expression in ovarian cancer. Gene 2020, 724, 144150. [Google Scholar] [CrossRef]
- Zhao, X.; Cai, Y.; Xu, J. Circular RNAs: Biogenesis, Mechanism, and Function in Human Cancers. Int. J. Mol. Sci. 2019, 20, 3926. [Google Scholar] [CrossRef]
- Arnaiz, E.; Sole, C.; Manterola, L.; Iparraguirre, L.; Otaegui, D.; Lawrie, C.H. CircRNAs and cancer: Biomarkers and master regulators. Semin. Cancer Biol. 2019, 58, 90–99. [Google Scholar] [CrossRef] [PubMed]
- Vitiello, M.; Tuccoli, A.; Poliseno, L. Long non-coding RNAs in cancer: Implications for personalized therapy. Cell. Oncol. 2015, 38, 17–28. [Google Scholar] [CrossRef] [PubMed]
- Haddadi, N.; Lin, Y.; Travis, G.; Simpson, A.M.; McGowan, E.M.; Nassif, N.T. PTEN/PTENP1: “Regulating the regulator of RTK-dependent PI3K/Akt signalling”, new targets for cancer therapy. Mol. Cancer 2018, 17, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Sood, A.K.; Dang, C.V.; Zhang, L. The role of long noncoding RNAs in cancer: The dark matter matters. Curr. Opin. Genet. Dev. 2018, 48, 8–15. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.-N.; Xia, S.-Q.; Zhang, Y.-Y.; Zheng, J.-H.; Li, W. Circular RNAs: A novel type of biomarker and genetic tools in cancer. Oncotarget 2017, 8, 64551–64563. [Google Scholar] [CrossRef]
- Tran, A.M.; Chalbatani, G.M.; Berland, L.; Cruz De los Santos, M.; Raj, P.; Jalali, S.A.; Gharagouzloo, E.; Ivan, C.; Dragomir, M.P.; Calin, G.A. A New World of Biomarkers and Therapeutics for Female Reproductive System and Breast Cancers: Circular RNAs. Front. Cell Dev. Biol. 2020, 8, 1–17. [Google Scholar] [CrossRef]
- Poliseno, L. Pseudogenes: Newly Discovered Players in Human Cancer. Sci. Signal. 2012, 5, re5. [Google Scholar] [CrossRef]
- Fang, L.; Wang, H.; Li, P. Systematic analysis reveals a lncRNA-mRNA co-expression network associated with platinum resistance in high-grade serous ovarian cancer. Investig. New Drugs 2018, 36, 187–194. [Google Scholar] [CrossRef]
- Song, J.; Zhang, W.; Wang, S.; Liu, K.; Song, F.; Ran, L. A panel of 7 prognosis-related long non-coding RNAs to improve platinum-based chemoresistance prediction in ovarian cancer. Int. J. Oncol. 2018, 53, 866–876. [Google Scholar] [CrossRef]
- Liu, R.; Zeng, Y.; Zhou, C.F.; Wang, Y.; Li, X.; Liu, Z.Q.; Chen, X.P.; Zhang, W.; Zhou, H.H. Long noncoding RNA expression signature to predict platinum-based chemotherapeutic sensitivity of ovarian cancer patients. Sci. Rep. 2017, 7, 1–10. [Google Scholar] [CrossRef]
- Wang, L.; Hu, Y.; Xiang, X.; Qu, K.; Teng, Y. Identification of long non-coding RNA signature for paclitaxel-resistant patients with advanced ovarian cancer. Oncotarget 2017, 8, 64191–64202. [Google Scholar] [CrossRef] [PubMed]
- Yang, K.; Hou, Y.; Li, A.; Li, Z.; Wang, W.; Xie, H.; Rong, Z.; Lou, G.; Li, K. Identification of a six-lncRNA signature associated with recurrence of ovarian cancer. Sci. Rep. 2017, 7, 1–10. [Google Scholar] [CrossRef]
- Luo, P.; Liu, X.F.; Wang, Y.C.; Li, N.D.; Liao, S.J.; Yu, M.X.; Liang, C.Z.; Tu, J.C. Prognostic value of abnormally expressed lncRNAs in ovarian carcinoma: A systematic review and meta-analysis. Oncotarget 2017, 8, 23927–23936. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wang, H.; Song, T.; Zou, Y.; Jiang, J.; Fang, L.; Li, P. HOTAIR is a potential target for the treatment of cisplatin-resistant ovarian cancer. Mol. Med. Rep. 2015, 12, 2211–2216. [Google Scholar] [CrossRef] [PubMed]
- Fu, L.L.; Li, C.J.; Xu, Y.; Li, L.Y.; Zhou, X.; Li, D.D.; Chen, S.X.; Wang, F.G.; Zhang, X.Y.; Zheng, L.W. Role of IncRNAs as novel biomarkers and therapeutic targets in ovarian cancer. Crit. Rev. Eukaryot. Gene Expr. 2017, 27, 183–195. [Google Scholar] [CrossRef]
- Li, J.; Huang, H.; Li, Y.; Li, L.; Hou, W.; You, Z. Decreased expression of long non-coding RNA GAS5 promotes cell proliferation, migration and invasion, and indicates a poor prognosis in ovarian cancer. Oncol. Rep. 2016, 36, 3241–3250. [Google Scholar] [CrossRef]
- Guo, R.; Qin, Y. LEMD1-AS1 Suppresses Ovarian Cancer Progression Through Regulating miR-183-5p/TP53 Axis. Oncol. Targets. Ther. 2020, 13, 7387–7398. [Google Scholar] [CrossRef]
- Chang, H.; Zhang, X.; Li, B.; Meng, X. MAGI2-AS3 suppresses MYC signaling to inhibit cell proliferation and migration in ovarian cancer through targeting miR-525-5p/MXD1 axis. Cancer Med. 2020, 1–10. [Google Scholar] [CrossRef]
- Liu, S.; Lei, H.; Luo, F.; Li, Y.; Xie, L. The effect of lncRNA HOTAIR on chemoresistance of ovarian cancer through regulation of HOXA7. Biol. Chem. 2018, 399, 485–497. [Google Scholar] [CrossRef]
- Bai, L.; Wang, A.; Zhang, Y.; Xu, X.; Zhang, X. Knockdown of MALAT1 enhances chemosensitivity of ovarian cancer cells to cisplatin through inhibiting the Notch1 signaling pathway. Exp. Cell Res. 2018, 366, 161–171. [Google Scholar] [CrossRef]
- Wang, H.; Liu, M.; Fang, L.; Jiang, J.; Zhang, Z.; Kuang, Y.; Wang, B.; Shang, X.; Han, P.; Li, Y.; et al. The cisplatin-induced lncRNA PANDAR dictates the chemoresistance of ovarian cancer via regulating SFRS2-mediated p53 phosphorylation. Cell Death Dis. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Ai, H.; Fan, X.; Chen, S.; Wang, Y.; Liu, L. Knockdown of long non-coding RNA HOTAIR reverses cisplatin resistance of ovarian cancer cells through inhibiting miR-138-5p-regulated EZH2 and SIRT1. Biol. Res. 2020, 53, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Blagden, S.; Abdel Mouti, M.; Chettle, J. Ancient and modern: Hints of a core post-transcriptional network driving chemotherapy resistance in ovarian cancer. Wiley Interdiscip. Rev. RNA 2018, 9, e1432. [Google Scholar] [CrossRef] [PubMed]
- Liu, E.; Liu, Z.; Zhou, Y.; Mi, R.; Wang, D. Overexpression of long non-coding RNA PVT1 in ovarian cancer cells promotes cisplatin resistance by regulating apoptotic pathways. Int. J. Clin. Exp. Med. 2015, 8, 20565–20572. [Google Scholar] [PubMed]
- Hosseini, E.S.; Meryet-Figuiere, M.; Sabzalipoor, H.; Kashani, H.H.; Nikzad, H.; Asemi, Z. Dysregulated expression of long noncoding RNAs in gynecologic cancers. Mol. Cancer 2017, 16, 1–13. [Google Scholar] [CrossRef]
- Wang, F.; Zhou, J.; Xie, X.; Hu, J.; Chen, L.; Hu, Q.; Guo, H.; Yu, C. Involvement of SRPK1 in cisplatin resistance related to long non-coding RNA UCA1 in human ovarian cancer cells. Neoplasma 2015, 62, 432–438. [Google Scholar] [CrossRef]
- Adriaens, C.; Standaert, L.; Barra, J.; Latil, M.; Verfaillie, A.; Kalev, P.; Boeckx, B.; Wijnhoven, P.W.G.; Radaelli, E.; Vermi, W.; et al. P53 induces formation of NEAT1 lncRNA-containing paraspeckles that modulate replication stress response and chemosensitivity. Nat. Med. 2016, 22, 861–868. [Google Scholar] [CrossRef]
- Cheng, Y.; He, C.; Wang, M.; Ma, X.; Mo, F.; Yang, S.; Han, J.; Wei, X. Targeting epigenetic regulators for cancer therapy: Mechanisms and advances in clinical trials. Signal. Transduct. Target. Ther. 2019, 4, 1–39. [Google Scholar] [CrossRef]
- Biswas, S.; Rao, C.M. Epigenetic tools (The Writers, The Readers and The Erasers) and their implications in cancer therapy. Eur. J. Pharmacol. 2018, 837, 8–24. [Google Scholar] [CrossRef]
- Beckedorff, F.C.; Amaral, M.S.; Deocesano-Pereira, C.; Verjovski-Almeida, S. Long non-coding RNAs and their implications in cancer epigenetics. Biosci. Rep. 2013, 33, 667–675. [Google Scholar] [CrossRef]
- Fang, Y.; Fullwood, M.J. Roles, Functions, and Mechanisms of Long Non-coding RNAs in Cancer. Genom. Proteom. Bioinform. 2016, 14, 42–54. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Feng, Y.; Zhang, D.; Zhao, S.D.; Hu, Z.; Greshock, J.; Zhang, Y.; Yang, L.; Zhong, X.; Wang, L.P.; et al. A Functional Genomic Approach Identifies FAL1 as an Oncogenic Long Noncoding RNA that Associates with BMI1 and Represses p21 Expression in Cancer. Cancer Cell 2014, 26, 344–357. [Google Scholar] [CrossRef] [PubMed]
- Activity, E.O. The focally amplified lncRNA FAL1 exhibits oncogenic activity. Cancer Discov. 2014, 4, 1253. [Google Scholar] [CrossRef]
- Chen, Y.; Du, H.; Bao, L.; Liu, W. LncRNA PVT1 promotes ovarian cancer progression by silencing miR-214. Cancer Biol. Med. 2018, 15, 238–250. [Google Scholar] [CrossRef]
- Delás, M.J.; Hannon, G.J. lncRNAs in development and disease: From functions to mechanisms. Open Biol. 2017, 7, 1–10. [Google Scholar] [CrossRef]
- Arab, K.; Park, Y.J.; Lindroth, A.M.; Schäfer, A.; Oakes, C.; Weichenhan, D.; Lukanova, A.; Lundin, E.; Risch, A.; Meister, M.; et al. Long noncoding RNA TARID directs demethylation and activation of the tumor suppressor TCF21 via GADD45A. Mol. Cell 2014, 55, 604–614. [Google Scholar] [CrossRef]
- Zhang, C.; Liu, J.; Zhang, Y.; Luo, C.; Zhu, T.; Zhang, R.; Yao, R. LINC01210 accelerates proliferation, invasion and migration in ovarian cancer through epigenetically downregulating KLF4. Biomed. Pharmacother. 2019, 119, 109431. [Google Scholar] [CrossRef]
- Li, Y.; Jiao, Y.; Hao, J.; Xing, H.; Li, C. Long noncoding RNA TP73-AS1 accelerates the epithelial ovarian cancer via epigenetically repressing p21. Am. J. Transl. Res. 2019, 11, 2447–2454. [Google Scholar]
- Wang, H.; Su, H.; Tan, Y. UNC5B-AS1 promoted ovarian cancer progression by regulating the H3K27me on NDRG2 via EZH2. Cell Biol. Int. 2020, 44, 1028–1036. [Google Scholar] [CrossRef]
- Zeng, X.Y.; Jiang, X.Y.; Yong, J.H.; Xie, H.; Yuan, J.; Zeng, D.; Dou, Y.Y.; Xiao, S.S. lncRNA ABHD11-AS1, regulated by the EGFR pathway, contributes to the ovarian cancer tumorigenesis by epigenetically suppressing TIMP2. Cancer Med. 2019, 8, 7074–7085. [Google Scholar] [CrossRef]
- Schuettengruber, B.; Bourbon, H.M.; Di Croce, L.; Cavalli, G. Genome Regulation by Polycomb and Trithorax: 70 Years and Counting. Cell 2017, 171, 34–57. [Google Scholar] [CrossRef] [PubMed]
- Zhu, S.; Zhao, D.; Yan, L.; Jiang, W.; Kim, J.S.; Gu, B.; Liu, Q.; Wang, R.; Xia, B.; Zhao, J.C.; et al. BMI1 regulates androgen receptor in prostate cancer independently of the polycomb repressive complex 1. Nat. Commun. 2018, 9, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Comet, I.; Riising, E.M.; Leblanc, B.; Helin, K. Maintaining cell identity: PRC2-mediated regulation of transcription and cancer. Nat. Rev. Cancer 2016, 16, 803–810. [Google Scholar] [CrossRef] [PubMed]
- Jones, B.A.; Varambally, S.; Arend, R.C. Histone Methyltransferase EZH2: A Therapeutic Target for Ovarian Cancer. Mol. Cancer Ther. 2018, 17, 591–602. [Google Scholar] [CrossRef] [PubMed]
- Penna, E.; Orso, F.; Taverna, D. miR-214 as a Key Hub that Controls Cancer Networks: Small Player, Multiple Functions. J. Investig. Dermatol. 2015, 135, 960–969. [Google Scholar] [CrossRef]
- Wang, B.; Shen, A.; Ouyang, X.; Zhao, G.; Du, Z.; Huo, W.; Zhang, T.; Wang, Y.; Yang, C.; Dong, P.; et al. KLF4 expression enhances the efficacy of chemotherapy drugs in ovarian cancer cells. Biochem. Biophys. Res. Commun. 2017, 484, 486–492. [Google Scholar] [CrossRef]
- Yi, X.; Guo, J.; Guo, J.; Sun, S.; Yang, P.; Wang, J.; Li, Y.; Xie, L.; Cai, J.; Wang, Z. EZH2-mediated epigenetic silencing of TIMP2 promotes ovarian cancer migration and invasion. Sci. Rep. 2017, 7, 1–16. [Google Scholar] [CrossRef]
- Yuan, C.; Liu, X.; Liu, X.; Yang, N.; Liu, Z.; Yan, S.; Shen, K.; Kong, B. The GADD45A (1506T>C) Polymorphism Is Associated with Ovarian Cancer Susceptibility and Prognosis. PLoS ONE 2015, 10, e0138692. [Google Scholar] [CrossRef]
- Arab, K.; Karaulanov, E.; Musheev, M.; Trnka, P.; Schäfer, A.; Grummt, I.; Niehrs, C. GADD45A binds R-loops and recruits TET1 to CpG island promoters. Nat. Genet. 2019, 51, 217–223. [Google Scholar] [CrossRef]
- Wu, X.; Zhang, Y. TET-mediated active DNA demethylation: Mechanism, function and beyond. Nat. Rev. Genet. 2017, 18, 517–534. [Google Scholar] [CrossRef]
- Nie, L.; Wu, H.J.; Hsu, J.M.; Chang, S.S.; LaBaff, A.M.; Li, C.W.; Wang, Y.; Hsu, J.L.; Hung, M.C. Long non-coding RNAs: Versatile master regulators of gene expression and crucial players in cancer. Am. J. Transl. Res. 2012, 4, 127–150. [Google Scholar] [PubMed]
- Davidovich, C.; Zheng, L.; Goodrich, K.J.; Cech, T.R. Promiscuous RNA binding by Polycomb repressive complex 2. Nat. Struct. Mol. Biol. 2013, 20, 1250–1257. [Google Scholar] [CrossRef] [PubMed]
- Fang, S.; Zhang, L.; Guo, J.; Niu, Y.; Wu, Y.; Li, H.; Zhao, L.; Li, X.; Teng, X.; Sun, X.; et al. NONCODEV5: A comprehensive annotation database for long non-coding RNAs. Nucleic Acids Res. 2018, 46, D308–D314. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, S.; Deb, A.; Maji, R.K.; Saha, S.; Ghosh, Z. LncRBase: An Enriched Resource for lncRNA Information. PLoS ONE 2014, 9, e108010. [Google Scholar] [CrossRef] [PubMed]
- Volders, P.-J.; Anckaert, J.; Verheggen, K.; Nuytens, J.; Martens, L.; Mestdagh, P.; Vandesompele, J. LNCipedia 5: Towards a reference set of human long non-coding RNAs. Nucleic Acids Res. 2019, 47, D135–D139. [Google Scholar] [CrossRef]
- Ma, L.; Cao, J.; Liu, L.; Du, Q.; Li, Z.; Zou, D.; Bajic, V.B.; Zhang, Z. LncBook: A curated knowledgebase of human long non-coding RNAs. Nucleic Acids Res. 2019, 47, D128–D134. [Google Scholar] [CrossRef]
- Zhou, B.; Zhao, H.; Yu, J.; Guo, C.; Dou, X.; Song, F.; Hu, G.; Cao, Z.; Qu, Y.; Yang, Y.; et al. EVLncRNAs: A manually curated database for long non-coding RNAs validated by low-throughput experiments. Nucleic Acids Res. 2018, 46, D100–D105. [Google Scholar] [CrossRef]
- Ma, L.; Li, A.; Zou, D.; Xu, X.; Xia, L.; Yu, J.; Bajic, V.B.; Zhang, Z. LncRNAWiki: Harnessing community knowledge in collaborative curation of human long non-coding RNAs. Nucleic Acids Res. 2015, 43, D187–D192. [Google Scholar] [CrossRef]
- LNCipedia. Available online: https://lncipedia.org (accessed on 27 April 2020).
- LncRNAWiki. Available online: http://lncrna.big.ac.cn (accessed on 30 April 2020).
- LncBook. Available online: http://bigd.big.ac.cn/lncbook (accessed on 20 May 2020).
- Li, J.H.; Liu, S.; Zhou, H.; Qu, L.H.; Yang, J.H. StarBase v2.0: Decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 2014, 42, 92–97. [Google Scholar] [CrossRef]
- ENCORI. Available online: http://starbase.sysu.edu.cn/ (accessed on 27 June 2020).
- Cheng, L.; Wang, P.; Tian, R.; Wang, S.; Guo, Q.; Luo, M.; Zhou, W.; Liu, G.; Jiang, H.; Jiang, Q. LncRNA2Target v2.0: A comprehensive database for target genes of lncRNAs in human and mouse. Nucleic Acids Res. 2019, 47, D140–D144. [Google Scholar] [CrossRef]
- Zhao, H.; Shi, J.; Zhang, Y.; Xie, A.; Yu, L.; Zhang, C.; Lei, J.; Xu, H.; Leng, Z.; Li, T.; et al. LncTarD: A manually-curated database of experimentally-supported functional lncRNA-target regulations in human diseases. Nucleic Acids Res. 2020, 48, D118–D126. [Google Scholar] [CrossRef] [PubMed]
- Teng, X.; Chen, X.; Xue, H.; Tang, Y.; Zhang, P.; Kang, Q.; Hao, Y.; Chen, R.; Zhao, Y.; He, S. NPInter v4.0: An integrated database of ncRNA interactions. Nucleic Acids Res. 2019, 48, D160–D165. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Li, X.; Gao, Y.; Guo, Q.; Wang, Y.; Fang, Y.; Ma, X.; Zhi, H.; Zhou, D.; Shen, W.; et al. LncACTdb 2.0: An updated database of experimentally supported ceRNA interactions curated from low-and high-throughput experiments. Nucleic Acids Res. 2019, 47, D121–D127. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Zhi, H.; Zhang, Y.; Liu, Y.; Zhang, J.; Gao, Y.; Guo, M.; Ning, S.; Li, X. miRSponge: A manually curated database for experimentally supported miRNA sponges and ceRNAs. Database 2015, 2015, bav098. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Bai, J.; Wu, A.; Wang, Y.; Zhang, J.; Wang, Z.; Li, Y.; Xu, J.; Li, X. Co-LncRNA: Investigating the lncRNA combinatorial effects in GO annotations and KEGG pathways based on human RNA-Seq data. Database 2015, 2015, bav082. [Google Scholar] [CrossRef]
- Zhi, H.; Li, X.; Wang, P.; Gao, Y.; Gao, B.; Zhou, D.; Zhang, Y.; Guo, M.; Yue, M.; Shen, W.; et al. Lnc2Meth: A manually curated database of regulatory relationships between long non-coding RNAs and DNA methylation associated with human disease. Nucleic Acids Res. 2018, 46, D133–D138. [Google Scholar] [CrossRef]
- Bao, Z.; Yang, Z.; Huang, Z.; Zhou, Y.; Cui, Q.; Dong, D. LncRNADisease 2.0: An updated database of long non-coding RNA-associated diseases. Nucleic Acids Res. 2019, 47, D1034–D1037. [Google Scholar] [CrossRef]
- Jia, K.; Gao, Y.; Shi, J.; Zhou, Y.; Zhou, Y.; Cui, Q. Annotation and curation of the causality information in LncRNADisease. Database 2020, 2020, baz150. [Google Scholar] [CrossRef]
- Gao, Y.; Wang, P.; Wang, Y.; Ma, X.; Zhi, H.; Zhou, D.; Li, X.; Fang, Y.; Shen, W.; Xu, Y.; et al. Lnc2Cancer v2.0: Updated database of experimentally supported long non-coding RNAs in human cancers. Nucleic Acids Res. 2019, 47, D1028–D1033. [Google Scholar] [CrossRef]
- Lnc2Cancer 3.0. Available online: http://www.bio-bigdata.net/lnc2cancer/ (accessed on 5 July 2020).
- Cui, T.; Zhang, L.; Huang, Y.; Yi, Y.; Tan, P.; Zhao, Y.; Hu, Y.; Xu, L.; Li, E.; Wang, D. MNDR v2.0: An updated resource of ncRNA-disease associations in mammals. Nucleic Acids Res. 2018, 46, D371–D374. [Google Scholar] [CrossRef]
- Zhang, W.; Yao, G.; Wang, J.; Yang, M.; Wang, J.; Zhang, H.; Li, W. ncRPheno: A comprehensive database platform for identification and validation of disease related noncoding RNAs. RNA Biol. 2020, 17, 943–955. [Google Scholar] [CrossRef] [PubMed]
- Ren, C.; An, G.; Zhao, C.; Ouyang, Z.; Bo, X.; Shu, W. Lnc2Catlas: An atlas of long noncoding RNAs associated with risk of cancers. Sci. Rep. 2018, 8, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Xia, S.; Feng, J.; Chen, K.; Ma, Y.; Gong, J.; Cai, F.; Jin, Y.; Gao, Y.; Xia, L.; Chang, H.; et al. CSCD: A database for cancer-specific circular RNAs. Nucleic Acids Res. 2018, 46, D925–D929. [Google Scholar] [CrossRef] [PubMed]
- Yao, D.; Zhang, L.; Zheng, M.; Sun, X.; Lu, Y.; Liu, P. Circ2Disease: A manually curated database of experimentally validated circRNAs in human disease. Sci. Rep. 2018, 8, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Fan, C.; Lei, X.; Fang, Z.; Jiang, Q.; Wu, F.-X. CircR2Disease: A manually curated database for experimentally supported circular RNAs associated with various diseases. Database 2018, 2018, bay044. [Google Scholar] [CrossRef] [PubMed]
- CircR2Disease. Available online: http://bioinfo.snnu.edu.cn/CircR2Disease/ (accessed on 26 June 2020).
- Vo, J.N.; Cieslik, M.; Zhang, Y.; Shukla, S.; Xiao, L.; Zhang, Y.; Wu, Y.M.; Dhanasekaran, S.M.; Engelke, C.G.; Cao, X.; et al. The Landscape of Circular RNA in Cancer. Cell 2019, 176, 869–881.e13. [Google Scholar] [CrossRef]
- Tay, Y.; Rinn, J.; Pandolfi, P.P. The multilayered complexity of ceRNA crosstalk and competition. Nature 2014, 505, 344–352. [Google Scholar] [CrossRef]
- Yang, C.; Wu, D.; Gao, L.; Liu, X.; Jin, Y.; Wang, D.; Wang, T.; Li, X. Competing endogenous RNA networks in human cancer: Hypothesis, validation, and perspectives. Oncotarget 2016, 7, 13479–13490. [Google Scholar] [CrossRef]
- Salmena, L.; Poliseno, L.; Tay, Y.; Kats, L.; Pandolfi, P.P. A ceRNA hypothesis: The rosetta stone of a hidden RNA language? Cell 2011, 146, 353–358. [Google Scholar] [CrossRef]
- Li, J.; Han, L.; Roebuck, P.; Diao, L.; Liu, L.; Yuan, Y.; Weinstein, J.N.; Liang, H. TANRIC: An interactive open platform to explore the function of IncRNAs in cancer. Cancer Res. 2015, 75, 3728–3737. [Google Scholar] [CrossRef]
- Anaya, J. OncoLnc: Linking TCGA survival data to mRNAs, miRNAs, and lncRNAs. PeerJ Comput. Sci. 2016, 2, e67. [Google Scholar] [CrossRef]
- Wang, J.; Ma, R.; Ma, W.; Chen, J.; Yang, J.; Xi, Y.; Cui, Q. LncDisease: A sequence based bioinformatics tool for predicting lncRNA-disease associations. Nucleic Acids Res. 2016, 44, e90. [Google Scholar] [CrossRef] [PubMed]
- Morel, D.; Jeffery, D.; Aspeslagh, S.; Almouzni, G.; Postel-Vinay, S. Combining epigenetic drugs with other therapies for solid tumours–past lessons and future promise. Nat. Rev. Clin. Oncol. 2020, 17, 91–107. [Google Scholar] [CrossRef] [PubMed]
- Cossío, F.P.; Esteller, M.; Berdasco, M. Towards a more precise therapy in cancer: Exploring epigenetic complexity. Curr. Opin. Chem. Biol. 2020, 57, 41–49. [Google Scholar] [CrossRef] [PubMed]
- Salehi, S.; Taheri, M.N.; Azarpira, N.; Zare, A.; Behzad-Behbahani, A. State of the art technologies to explore long non-coding RNAs in cancer. J. Cell. Mol. Med. 2017, 21, 3120–3140. [Google Scholar] [CrossRef] [PubMed]
- Schmitt, A.M.; Chang, H.Y. Long Noncoding RNAs in Cancer Pathways. Cancer Cell 2016, 29, 452–463. [Google Scholar] [CrossRef]
- Gilbert, L.A.; Horlbeck, M.A.; Adamson, B.; Villalta, J.E.; Chen, Y.; Whitehead, E.H.; Guimaraes, C.; Panning, B.; Ploegh, H.L.; Bassik, M.C.; et al. Genome-Scale CRISPR-Mediated Control of Gene Repression and Activation. Cell 2014, 159, 647–661. [Google Scholar] [CrossRef]
| Database | Aim | Website | Refs. |
|---|---|---|---|
| lncRNA annotation | |||
| NONCODE v5.0 | Collects and annotates a comprehensive set of ncRNAs, especially lncRNAs. | http://www.noncode.org/ | [105] |
| LNCipedia 5.2 | Collects sequences and annotation data for a comprehensive set of lncRNA transcripts. | https://lncipedia.org | [107,111] |
| LncRNAWiki | Collects and integrates comprehensive information on human lncRNAs relying on collaborative curation. | http://lncrna.big.ac.cn | [110,112] |
| EVLncRNAs | Collects lncRNAs that were validated by low-throughput experiments. | http://biophy.dzu.edu.cn/EVLncRNAs | [109] |
| LncRBase | Provides information on a comprehensive set of lncRNAs, ranging from basic transcript features to additional details on their genomic context. | http://bicresources.jcbose.ac.in/zhumur/lncrbase | [106] |
| LncBook | Collects a comprehensive set of human lncRNAs and provides multi-omics data integration, functional annotation, and disease association for these transcripts. | http://bigd.big.ac.cn/lncbook | [108,113] |
| lncRNA regulatory functions (interactions, mechanisms, and effects) | |||
| ENCORI | Integrates multi-dimensional sequencing data to provide a comprehensive collection of RNA–RNA and protein–RNA interaction networks. | http://starbase.sysu.edu.cn/ | [114,115] |
| LncRNA2Target v2.0 | Collects lncRNA–target associations supported by high- and low-throughput methods. | http://123.59.132.21/lncrna2target/ | [116] |
| LncTarD | Collects and illustrates experimentally supported lncRNA–target regulations in human diseases. | http://bio-bigdata.hrbmu.edu.cn/LncTarD | [117] |
| NPInter v4.0 | Integrates literature reported evidence and high-throughput data processing to collect and illustrate functional interactions between ncRNAs and biomolecules. | http://bigdata.ibp.ac.cn/npinter | [118] |
| LncACTdb 2.0 | Collects and illustrates information on ceRNA relationships between different types of RNAs in diverse diseases. | http://www.bio-bigdata.net/LncACTdb/ | [119] |
| miRSponge | Collects experimentally validated miRNA sponges and ceRNA interactions. | http://bio-bigdata.hrbmu.edu.cn/miRSponge/ | [120] |
| Co-LncRNA | Allows the identification of lncRNA coexpressed protein-coding genes and the assessment of the effects of individual or multiple lncRNAs in GO annotations and KEGG pathways. | http://bio-bigdata.hrbmu.edu.cn/Co-LncRNA/ | [121] |
| Lnc2Meth | Integrates experimental and computational evidence to provide information on regulatory relationships between lncRNAs and DNA methylation in diverse human diseases. | http://bio-bigdata.hrbmu.edu.cn/Lnc2Meth/ | [122] |
| lncRNA–disease associations | |||
| LncRNADisease 2.0 | Integrates experimental and computational evidence to provide information on lncRNA/circRNA–disease associations, including causal relationships. | http://www.rnanut.net/lncrnadisease/ | [123,124] |
| Lnc2Cancer v3.0 | Collects and integrates experimentally supported associations between lncRNAs and cancer subtypes. | http://www.bio-bigdata.net/lnc2cancer/ | [125,126] |
| MNDR v2.0 | Collects and integrates experimentally supported and predicted ncRNA–disease associations in mammals. | http://www.rna-society.org/mndr/ | [127] |
| ncRPheno | Collects comprehensive ncRNA–disease association data and offers web applications to facilitate analysis and visualization of this data. | http://lilab2.sysu.edu.cn/ncrpheno | [128] |
| Lnc2Catlas | Integrates multiple methods and data sources to collect quantitative associations between lncRNAs and cancer. | http://lnc2catlas.bioinfotech.org/ | [129] |
| CSCD | Collects and integrates information on cancer-specific circRNAs. | http://gb.whu.edu.cn/CSCD | [130] |
| Circ2Disease | Collects information on experimentally supported circRNAs in human diseases. | http://bioinformatics.zju.edu.cn/Circ2Disease/index.html | [131] |
| CircR2Disease | Collects information on experimentally validated associations between circRNAs and diseases. | http://bioinfo.snnu.edu.cn/CircR2Disease/ | [132,133] |
| MiOncoCirc | Collects data on circRNA formation and abundance in cancer. | https://mioncocirc.github.io/ | [134] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Calanca, N.; Abildgaard, C.; Rainho, C.A.; Rogatto, S.R. The Interplay between Long Noncoding RNAs and Proteins of the Epigenetic Machinery in Ovarian Cancer. Cancers 2020, 12, 2701. https://doi.org/10.3390/cancers12092701
Calanca N, Abildgaard C, Rainho CA, Rogatto SR. The Interplay between Long Noncoding RNAs and Proteins of the Epigenetic Machinery in Ovarian Cancer. Cancers. 2020; 12(9):2701. https://doi.org/10.3390/cancers12092701
Chicago/Turabian StyleCalanca, Naiade, Cecilie Abildgaard, Cláudia Aparecida Rainho, and Silvia Regina Rogatto. 2020. "The Interplay between Long Noncoding RNAs and Proteins of the Epigenetic Machinery in Ovarian Cancer" Cancers 12, no. 9: 2701. https://doi.org/10.3390/cancers12092701
APA StyleCalanca, N., Abildgaard, C., Rainho, C. A., & Rogatto, S. R. (2020). The Interplay between Long Noncoding RNAs and Proteins of the Epigenetic Machinery in Ovarian Cancer. Cancers, 12(9), 2701. https://doi.org/10.3390/cancers12092701





