Sorafenib Resistance in Hepatocellular Carcinoma: The Relevance of Genetic Heterogeneity
Abstract
1. Introduction
1.1. Hepatocellular Carcinoma: Obstacles in Conventional Treatment
1.2. Molecular Therapy with Sorafenib
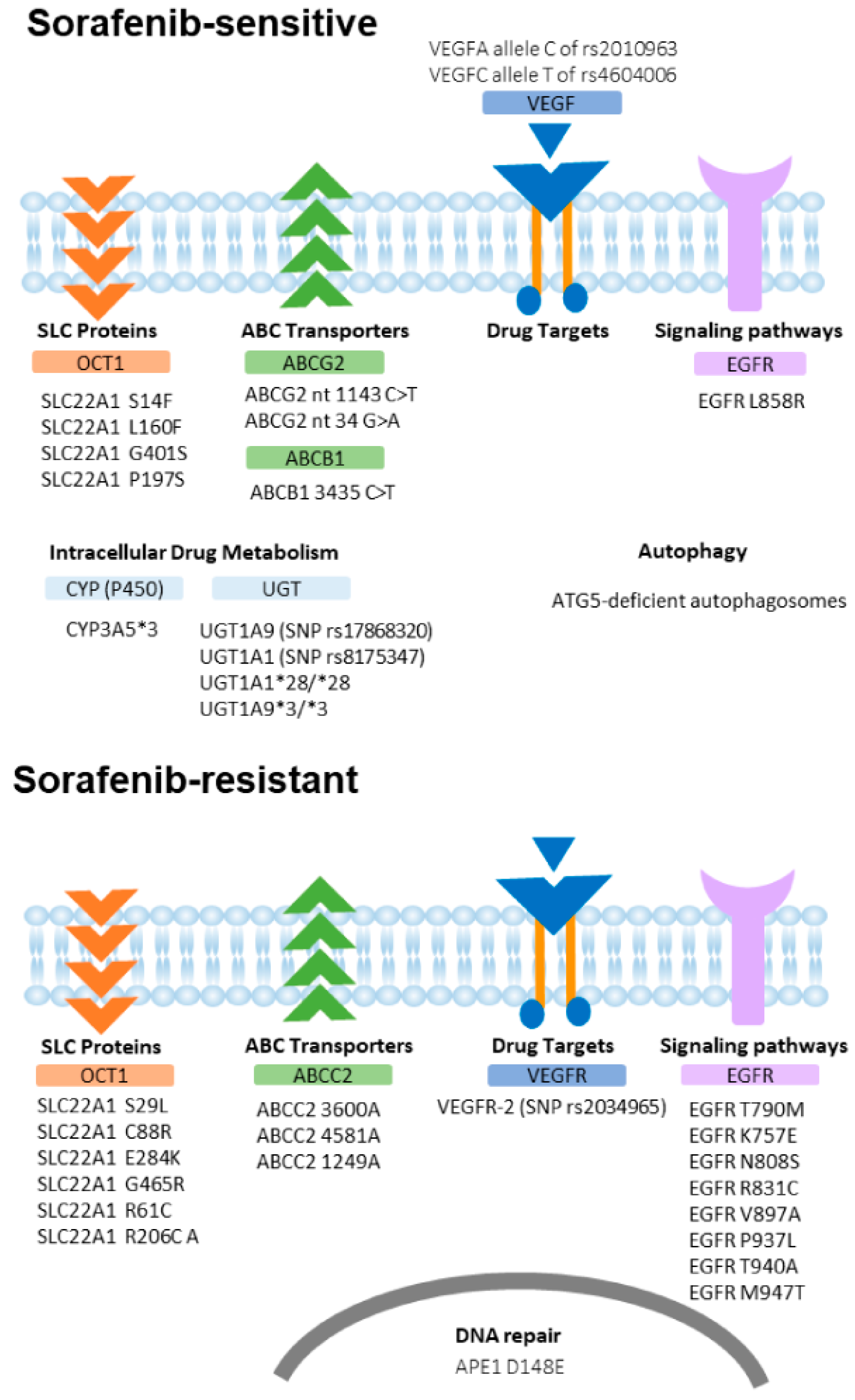
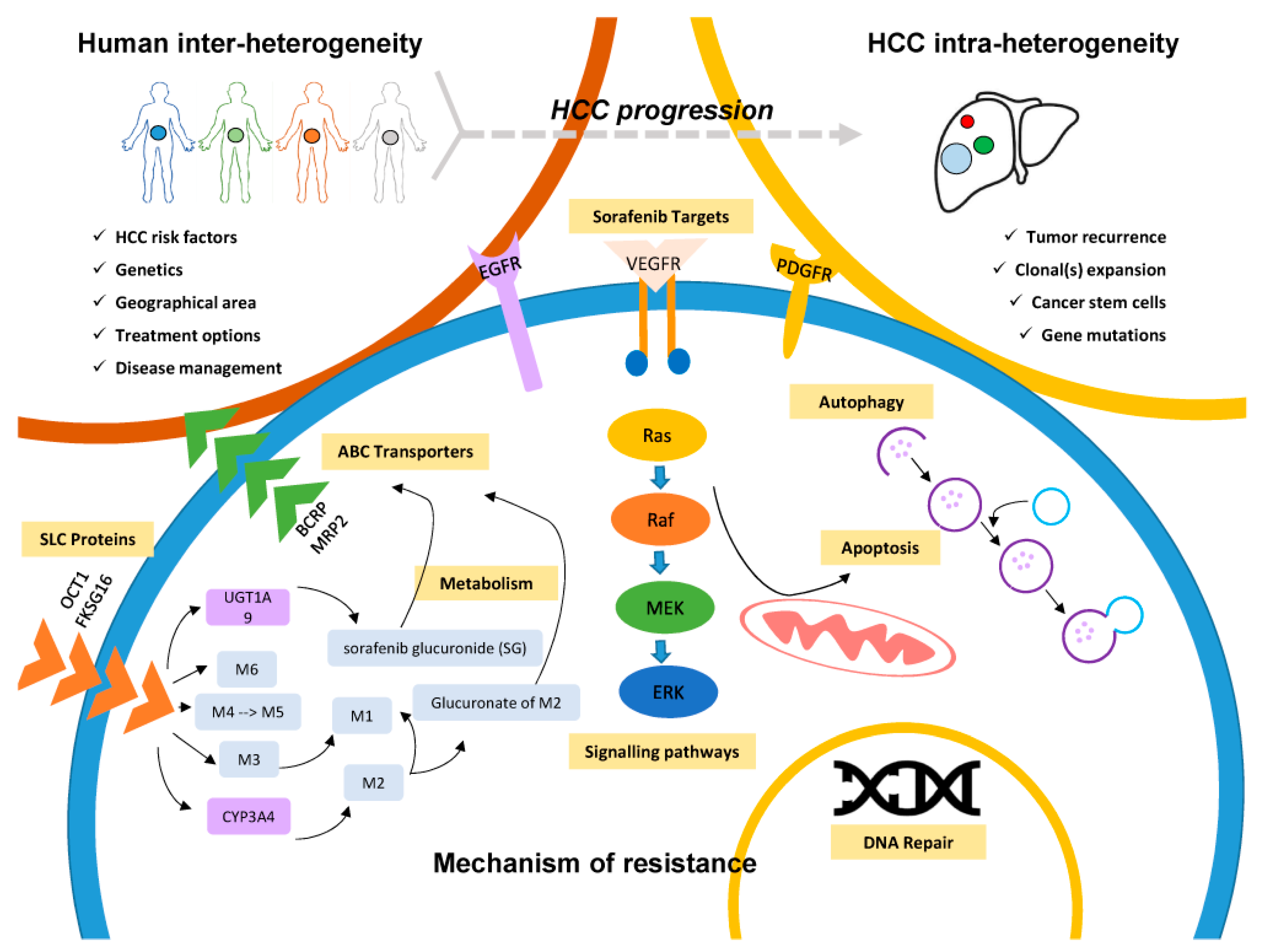
2. Mechanisms of Sorafenib Resistance and HCC Heterogeneity
2.1. Drug Efflux: ABC Transporters Family
2.2. Drug Uptake: The SLC Family
2.3. Alteration of Molecular Targets
2.4. Alteration of Signaling Pathways
2.5. Intracellular Drug Metabolism
2.6. DNA Repair Pathway
2.7. Autophagy
2.8. Hepatic Cancer Stem Cells
3. Future Perspectives
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Ferlay, J.; Colombet, M.; Soerjomataram, I.; Mathers, C.; Parkin, D.M.; Piñeros, M.; Znaor, A.; Bray, F. Estimating the global cancer incidence and mortality in 2018: GLOBOCAN sources and methods. Int. J. Cancer 2018. [Google Scholar] [CrossRef] [PubMed]
- Global Burden of Disease Liver Cancer Collaboration; Akinyemiju, T.; Abera, S.; Ahmed, M.; Alam, N.; Alemayohu, M.A.; Allen, C.; Al-Raddadi, R.; Alvis-Guzman, N.; Amoako, Y.; et al. The burden of primary liver cancer and underlying etiologies from 1990 to 2015 at the global, regional, and national level: Results from the Global Burden of Disease Study 2015. JAMA Oncol. 2017, 3, 1683–1691. [Google Scholar] [CrossRef] [PubMed]
- Colombo, M.; Maisonneuve, P. Controlling liver cancer mortality on a global scale: Still a long way to go. J. Hepatol. 2017, 67, 216–217. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.W.; Talati, C.; Kim, R. Hepatocellular carcinoma (HCC): Beyond sorafenib—Chemotherapy. J. Gastrointest. Oncol. 2017, 8, 256–265. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.D.; Hainaut, P.; Gores, G.J.; Amadou, A.; Plymoth, A.; Roberts, L.R. A global view of hepatocellular carcinoma: Trends, risk, prevention and management. Nat. Rev. Gastroenterol. Hepatol. 2019, 16, 589–604. [Google Scholar] [CrossRef]
- European Association for the Study of the Liver. European Organisation for Research and Treatment of Cancer EASL–EORTC Clinical Practice Guidelines: Management of hepatocellular carcinoma. J. Hepatol. 2012, 56, 908–943. [Google Scholar] [CrossRef]
- Belghiti, J.; Fuks, D. Liver resection and transplantation in hepatocellular carcinoma. Liver Cancer 2012, 1, 71–82. [Google Scholar] [CrossRef]
- Lohitesh, K.; Chowdhury, R.; Mukherjee, S. Resistance a major hindrance to chemotherapy in hepatocellular carcinoma: An insight. Cancer Cell Int. 2018, 18, 44. [Google Scholar] [CrossRef]
- Bruix, J.; Sherman, M. Management of hepatocellular carcinoma: An update. Hepatology 2011, 53, 1020–1022. [Google Scholar] [CrossRef]
- Takayasu, K.; Arii, S.; Ikai, I.; Omata, M.; Okita, K.; Ichida, T.; Matsuyama, Y.; Nakanuma, Y.; Kojiro, M.; Makuuchi, M.; et al. Prospective cohort study of transarterial chemoembolization for unresectable hepatocellular carcinoma in 8510 patients. Gastroenterology 2006, 131, 461–469. [Google Scholar] [CrossRef]
- Llovet, J.M.; Real, M.I.; Montaña, X.; Planas, R.; Coll, S.; Aponte, J.; Ayuso, C.; Sala, M.; Muchart, J.; Solà, R.; et al. Arterial embolisation or chemoembolisation versus symptomatic treatment in patients with unresectable hepatocellular carcinoma: A randomised controlled trial. Lancet 2002, 359, 1734–1739. [Google Scholar] [CrossRef]
- Malagari, K.; Pomoni, M.; Kelekis, A.; Pomoni, A.; Dourakis, S.; Spyridopoulos, T.; Moschouris, H.; Emmanouil, E.; Rizos, S.; Kelekis, D. Prospective randomized comparison of chemoembolization with doxorubicin-eluting beads and bland embolization with beadblock for hepatocellular carcinoma. Cardiovasc. Intervent. Radiol. 2010, 33, 541–551. [Google Scholar] [CrossRef] [PubMed]
- Roxburgh, P.; Evans, T.R.J. Systemic therapy of hepatocellular carcinoma: Are we making progress? Adv. Ther. 2008, 25, 1089–1104. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Lin, Y.; Zhang, J.; Zhang, Y.; Li, Y.; Liu, Z.; Li, Q.; Luo, M.; Liang, R.; Ye, J. Molecular targeted and immune checkpoint therapy for advanced hepatocellular carcinoma. J. Exp. Clin. Cancer Res. 2019, 38, 447. [Google Scholar] [CrossRef]
- Llovet, J.M.; Ricci, S.; Mazzaferro, V.; Hilgard, P.; Gane, E.; Blanc, J.-F.; De Oliveira, A.C.; Santoro, A.; Raoul, J.-L.; Forner, A.; et al. Sorafenib in advanced hepatocellular carcinoma. N. Engl. J. Med. 2008, 359, 378–390. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Cao, Y.; Chen, C.; Zhang, X.; McNabola, A.; Wilkie, D.; Wilhelm, S.; Lynch, M.; Carter, C. Sorafenib blocks the RAF/MEK/ERK pathway, inhibits tumor angiogenesis, and induces tumor cell apoptosis in hepatocellular carcinoma model PLC/PRF/5. Cancer Res. 2006, 66, 11851–11858. [Google Scholar] [CrossRef]
- Cheng, A.-L.; Kang, Y.-K.; Chen, Z.; Tsao, C.-J.; Qin, S.; Kim, J.S.; Luo, R.; Feng, J.; Ye, S.; Yang, T.-S.; et al. Efficacy and safety of sorafenib in patients in the Asia-Pacific region with advanced hepatocellular carcinoma: A phase III randomised, double-blind, placebo-controlled trial. Lancet Oncol. 2009, 10, 25–34. [Google Scholar] [CrossRef]
- Chen, W.; Yang, J.; Zhang, Y.; Cai, H.; Chen, X.; Sun, D. Regorafenib reverses HGF-induced sorafenib resistance by inhibiting epithelial-mesenchymal transition in hepatocellular carcinoma. FEBS Open Bio 2019, 9, 335–347. [Google Scholar] [CrossRef]
- Al-Rajabi, R.; Patel, S.; Ketchum, N.S.; Jaime, N.A.; Lu, T.-W.; Pollock, B.H.; Mahalingam, D. Comparative dosing and efficacy of sorafenib in hepatocellular cancer patients with varying liver dysfunction. J. Gastrointest. Oncol. 2015, 6, 259–267. [Google Scholar] [CrossRef]
- Reiss, K.A.; Yu, S.; Mamtani, R.; Mehta, R.; D’Addeo, K.; Wileyto, E.P.; Taddei, T.H.; Kaplan, D.E. Starting dose of sorafenib for the treatment of hepatocellular carcinoma: A retrospective, multi-institutional study. J. Clin. Oncol. 2017, 35, 3575–3581. [Google Scholar] [CrossRef] [PubMed]
- Boyault, S.; Rickman, D.S.; De Reyniès, A.; Balabaud, C.; Rebouissou, S.; Jeannot, E.; Hérault, A.; Saric, J.; Belghiti, J.; Franco, D.; et al. Transcriptome classification of HCC is related to gene alterations and to new therapeutic targets. Hepatology 2007, 45, 42–52. [Google Scholar] [CrossRef] [PubMed]
- Calderaro, J.; Ziol, M.; Paradis, V.; Zucman-Rossi, J. Molecular and histological correlations in liver cancer. J. Hepatol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Marin, J.J.G.; Romero, M.R.; Briz, O. Molecular bases of liver cancer refractoriness to pharmacological treatment. Curr. Med. Chem. 2010, 17, 709–740. [Google Scholar] [CrossRef]
- Villanueva, A.; Llovet, J.M. Mutational landscape of HCC—the end of the beginning. Nat. Rev. Clin. Oncol. 2014, 11, 73–74. [Google Scholar] [CrossRef]
- Guichard, C.; Amaddeo, G.; Imbeaud, S.; Ladeiro, Y.; Pelletier, L.; Maad, I.B.; Calderaro, J.; Bioulac-Sage, P.; Letexier, M.; Degos, F.; et al. Integrated analysis of somatic mutations and focal copy-number changes identifies key genes and pathways in hepatocellular carcinoma. Nat. Genet. 2012, 44, 694–698. [Google Scholar] [CrossRef]
- Llovet, J.M. Focal gains of VEGFA: Candidate predictors of sorafenib response in hepatocellular carcinoma. Cancer Cell 2014, 25, 560–562. [Google Scholar] [CrossRef]
- Zucman-Rossi, J.; Villanueva, A.; Nault, J.-C.; Llovet, J.M. Genetic landscape and biomarkers of hepatocellular carcinoma. Gastroenterology 2015, 149, 1226–1239. [Google Scholar] [CrossRef] [PubMed]
- Yao, S.; Johnson, C.; Hu, Q.; Yan, L.; Liu, B.; Ambrosone, C.B.; Wang, J.; Liu, S. Differences in somatic mutation landscape of hepatocellular carcinoma in Asian American and European American populations. Oncotarget 2016, 7, 40491–40499. [Google Scholar] [CrossRef]
- Teufel, M.; Seidel, H.; Köchert, K.; Meinhardt, G.; Finn, R.S.; Llovet, J.M.; Bruix, J. Biomarkers associated with response to regorafenib in patients With hepatocellular carcinoma. Gastroenterology 2019, 156, 1731–1741. [Google Scholar] [CrossRef] [PubMed]
- Caruso, S.; Calatayud, A.-L.; Pilet, J.; La Bella, T.; Rekik, S.; Imbeaud, S.; Letouzé, E.; Meunier, L.; Bayard, Q.; Rohr-Udilova, N.; et al. Analysis of liver cancer cell lines identifies agents with likely efficacy against hepatocellular carcinoma and markers of response. Gastroenterology 2019. [Google Scholar] [CrossRef] [PubMed]
- Dean, M.; Hamon, Y.; Chimini, G. The Human ATP-Binding Cassette (ABC) transporter superfamily. J. Lipid Res. 2001, 42, 1007–1017. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Q.; Zhou, Y.; Lauschke, V.M. Ethnogeographic and inter-individual variability of human ABC transporters. Hum. Genet. 2020. [Google Scholar] [CrossRef] [PubMed]
- Lepper, E.R.; Nooter, K.; Verweij, J.; Acharya, M.R.; Figg, W.D.; Sparreboom, A. Mechanisms of resistance to anticancer drugs: The role of the polymorphic ABC transporters ABCB1 and ABCG2. Pharmacogenomics 2005, 6, 115–138. [Google Scholar] [CrossRef] [PubMed]
- Cascorbi, I. Role of pharmacogenetics of ATP-binding cassette transporters in the pharmacokinetics of drugs. Pharmacol. Ther. 2006, 112, 457–473. [Google Scholar] [CrossRef]
- Gao, J. Association of MDR1 gene polymorphisms with the risk of hepatocellular carcinoma in the Chinese Han population. Braz. J. Med. Biol. Res. 2013, 46, 311–317. [Google Scholar] [CrossRef][Green Version]
- Ren, Y.-Q.; Han, J.-Q.; Cao, J.-B.; Li, S.-X.; Fan, G.-R. Association of MDR1 gene polymorphisms with susceptibility to hepatocellular carcinoma in the Chinese population. Asian Pac. J. Cancer Prev. 2012, 13, 5451–5454. [Google Scholar] [CrossRef][Green Version]
- Li, X.-F.; He, H.-B.; Zhu, Y.-S.; He, J.-K.; Ye, W.-W.; Chen, Y.-X.; Lou, L.-Q. Association between the c.3751G>a genetic variant of MDR1 and hepatocellular carcinoma risk in a Chinese Han population. Asian Pac. J. Cancer Prev. APJCP 2013, 14, 5361–5365. [Google Scholar] [CrossRef][Green Version]
- Sukowati, C.H.; Rosso, N.; Pascut, D.; Anfuso, B.; Torre, G.; Francalanci, P.; Crocè, L.S.; Tiribelli, C. Gene and functional up-regulation of the BCRP/ABCG2 transporter in hepatocellular carcinoma. BMC Gastroenterol. 2012, 12, 160. [Google Scholar] [CrossRef]
- Sun, Z.; Zhao, Z.; Li, G.; Dong, S.; Huang, Z.; Ye, L.; Liang, H.; Qu, J.; Ai, X.; Zhang, W.; et al. Relevance of two genes in the multidrug resistance of hepatocellular carcinoma: In vivo and clinical studies. Tumori J. 2010, 96, 90–96. [Google Scholar] [CrossRef]
- Luo, F.; Li, H.; Liang, J.; Jia, H.; Li, X.; Xiao, H.; He, X.; He, J.; Tian, Y.; Zhao, H. Downregulation of NPM reverses multidrug resistance in human hepatoma cells via inhibition of P-glycoprotein expression. Mol. Med. Rep. 2017, 15, 2360–2368. [Google Scholar] [CrossRef][Green Version]
- Hoffmann, K.; Franz, C.; Xiao, Z.; Mohr, E.; Serba, S.; Büchler, M.W.; Schemmer, P. Sorafenib modulates the gene expression of multi-drug resistance mediating ATP-binding cassette proteins in experimental hepatocellular carcinoma. Anticancer Res. 2010, 30, 4503–4508. [Google Scholar] [PubMed]
- Huang, W.-C.; Hsieh, Y.-L.; Hung, C.-M.; Chien, P.-H.; Chien, Y.-F.; Chen, L.-C.; Tu, C.-Y.; Chen, C.-H.; Hsu, S.-C.; Lin, Y.-M.; et al. BCRP/ABCG2 inhibition sensitizes hepatocellular carcinoma cells to sorafenib. PLoS ONE 2013, 8, e83627. [Google Scholar] [CrossRef] [PubMed]
- Tandia, M.; Mhiri, A.; Paule, B.; Saffroy, R.; Cailliez, V.; Noé, G.; Farinotti, R.; Bonhomme-Faivre, L. Correlation between clinical response to sorafenib in hepatocellular carcinoma treatment and polymorphisms of P-glycoprotein (ABCB1) and of breast cancer resistance protein (ABCG2): Monocentric study. Cancer Chemother. Pharmacol. 2017, 79, 759–766. [Google Scholar] [CrossRef] [PubMed]
- Boudou-Rouquette, P.; Narjoz, C.; Golmard, J.L.; Thomas-Schoemann, A.; Mir, O.; Taieb, F.; Durand, J.-P.; Coriat, R.; Dauphin, A.; Vidal, M.; et al. Early sorafenib-induced toxicity is associated with drug exposure and UGTIA9 genetic polymorphism in patients with solid tumors: A preliminary study. PLoS ONE 2012, 7, e42875. [Google Scholar] [CrossRef] [PubMed]
- Meier, Y.; Pauli-Magnus, C.; Zanger, U.M.; Klein, K.; Schaeffeler, E.; Nussler, A.K.; Nussler, N.; Eichelbaum, M.; Meier, P.J.; Stieger, B. Interindividual variability of canalicular ATP-binding-cassette (ABC)-transporter expression in human liver. Hepatol. Baltim. Md 2006, 44, 62–74. [Google Scholar] [CrossRef] [PubMed]
- Wei, D.; Zhang, H.; Peng, R.; Huang, C.; Bai, R. ABCC2 (1249G > A) polymorphism implicates altered transport activity for sorafenib. Xenobiotica 2017, 47, 1008–1014. [Google Scholar] [CrossRef]
- Wen, X.; Joy, M.S.; Aleksunes, L.M. In vitro transport activity and trafficking of MRP2/ABCC2 polymorphic variants. Pharm. Res. 2017, 34, 1637–1647. [Google Scholar] [CrossRef]
- Lian, G.; Yuan, J.; Gao, Y. In vitro transport ability of ABCC2 (G1249A) polymorphic variant towards anticancer drugs. OncoTargets Ther. 2020, 13, 1413–1419. [Google Scholar] [CrossRef]
- Lin, L.; Yee, S.W.; Kim, R.B.; Giacomini, K.M. SLC transporters as therapeutic targets: Emerging opportunities. Nat. Rev. Drug Discov. 2015, 14, 543–560. [Google Scholar] [CrossRef] [PubMed]
- Schaller, L.; Lauschke, V.M. The genetic landscape of the human solute carrier (SLC) transporter superfamily. Hum. Genet. 2019, 138, 1359–1377. [Google Scholar] [CrossRef] [PubMed]
- Seitz, T.; Stalmann, R.; Dalila, N.; Chen, J.; Pojar, S.; Dos Santos Pereira, J.N.; Krätzner, R.; Brockmöller, J.; Tzvetkov, M.V. Global genetic analyses reveal strong inter-ethnic variability in the loss of activity of the organic cation transporter OCT1. Genome Med. 2015, 7, 56. [Google Scholar] [CrossRef] [PubMed]
- Al-Abdulla, R.; Lozano, E.; Macias, R.I.R.; Monte, M.J.; Briz, O.; O’Rourke, C.J.; Serrano, M.A.; Banales, J.M.; Avila, M.A.; Martinez-Chantar, M.L.; et al. Epigenetic events involved in organic cation transporter 1-dependent impaired response of hepatocellular carcinoma to sorafenib. Br. J. Pharmacol. 2019, 176, 787–800. [Google Scholar] [CrossRef] [PubMed]
- Herraez, E.; Lozano, E.; Macias, R.I.R.; Vaquero, J.; Bujanda, L.; Banales, J.M.; Marin, J.J.G.; Briz, O. Expression of SLC22A1 variants may affect the response of hepatocellular carcinoma and cholangiocarcinoma to sorafenib. Hepatology 2013, 58, 1065–1073. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Neul, C.; Schaeffeler, E.; Frisch, F.; Winter, S.; Schwab, M.; Koepsell, H.; Hu, S.; Laufer, S.; Baker, S.D.; et al. Sorafenib activity and disposition in liver cancer does not depend on organic cation transporter 1. Clin. Pharmacol. Ther. 2020, 107, 227–237. [Google Scholar] [CrossRef]
- Geier, A.; Macias, R.I.R.; Bettinger, D.; Weiss, J.; Bantel, H.; Jahn, D.; Al-Abdulla, R.; Marin, J.J.G. The lack of the organic cation transporter OCT1 at the plasma membrane of tumor cells precludes a positive response to sorafenib in patients with hepatocellular carcinoma. Oncotarget 2017, 8, 15846–15857. [Google Scholar] [CrossRef]
- Martinez-Becerra, P.; Vaquero, J.; Romero, M.R.; Lozano, E.; Anadon, C.; Macias, R.I.R.; Serrano, M.A.; Grañé-Boladeras, N.; Muñoz-Bellvis, L.; Alvarez, L.; et al. No correlation between the expression of FXR and genes involved in multidrug resistance phenotype of primary liver tumors. Mol. Pharm. 2012, 9, 1693–1704. [Google Scholar] [CrossRef]
- Grimm, D.; Lieb, J.; Weyer, V.; Vollmar, J.; Darstein, F.; Lautem, A.; Hoppe-Lotichius, M.; Koch, S.; Schad, A.; Schattenberg, J.M.; et al. Organic cation transporter 1 (OCT1) mRNA expression in hepatocellular carcinoma as a biomarker for sorafenib treatment. BMC Cancer 2016, 16, 94. [Google Scholar] [CrossRef]
- Schaeffeler, E.; Hellerbrand, C.; Nies, A.T.; Winter, S.; Kruck, S.; Hofmann, U.; Van der Kuip, H.; Zanger, U.M.; Koepsell, H.; Schwab, M. DNA methylation is associated with downregulation of the organic cation transporter OCT1 (SLC22A1) in human hepatocellular carcinoma. Genome Med. 2011, 3, 82. [Google Scholar] [CrossRef]
- Arimany-Nardi, C.; Minuesa, G.; Keller, T.; Erkizia, I.; Koepsell, H.; Martinez-Picado, J.; Pastor-Anglada, M. Role of human organic cation transporter 1 (hOCT1) polymorphisms in lamivudine (3TC) uptake and drug-drug interactions. Front. Pharmacol. 2016, 7, 175. [Google Scholar] [CrossRef]
- Nies, A.T.; Koepsell, H.; Winter, S.; Burk, O.; Klein, K.; Kerb, R.; Zanger, U.M.; Keppler, D.; Schwab, M.; Schaeffeler, E. Expression of organic cation transporters OCT1 (SLC22A1) and OCT3 (SLC22A3) is affected by genetic factors and cholestasis in human liver. Hepatology 2009, 50, 1227–1240. [Google Scholar] [CrossRef] [PubMed]
- Swift, B.; Nebot, N.; Lee, J.K.; Han, T.; Proctor, W.R.; Thakker, D.R.; Lang, D.; Radtke, M.; Gnoth, M.J.; Brouwer, K.L.R. Sorafenib hepatobiliary disposition: Mechanisms of hepatic uptake and disposition of generated metabolites. Drug Metab. Dispos. 2013, 41, 1179–1186. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Zheng, B.; Meng, S.; Xu, Y.; Guo, J.; Chen, L.-J.; Xiao, J.; Zhang, W.; Tan, Z.-R.; Tang, J.; et al. Increased expression of SLC46A3 to oppose the progression of hepatocellular carcinoma and its effect on sorafenib therapy. Biomed. Pharmacother. 2019, 114, 108864. [Google Scholar] [CrossRef] [PubMed]
- Negri, F.V.; Dal Bello, B.; Porta, C.; Campanini, N.; Rossi, S.; Tinelli, C.; Poggi, G.; Missale, G.; Fanello, S.; Salvagni, S.; et al. Expression of pERK and VEGFR-2 in advanced hepatocellular carcinoma and resistance to sorafenib treatment. Liver Int. 2015, 35, 2001–2008. [Google Scholar] [CrossRef] [PubMed]
- Personeni, N.; Rimassa, L.; Pressiani, T.; Destro, A.; Ligorio, C.; Tronconi, M.C.; Bozzarelli, S.; Carnaghi, C.; Di Tommaso, L.; Giordano, L.; et al. Molecular determinants of outcome in sorafenib-treated patients with hepatocellular carcinoma. J. Cancer Res. Clin. Oncol. 2013, 139, 1179–1187. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Ma, X.-P.; Shi, Z.; Zhang, P.; Ding, D.-L.; Huang, H.-X.; Saiyin, H.G.; Chen, T.-Y.; Lu, P.-X.; Wang, N.-J.; et al. Epidermal growth factor receptor pathway polymorphisms and the prognosis of hepatocellular carcinoma. Am. J. Cancer Res. 2015, 5, 396–410. [Google Scholar] [PubMed]
- Iavarone, M.; Lampertico, P.; Iannuzzi, F.; Manenti, E.; Donato, M.F.; Arosio, E.; Bertolini, F.; Primignani, M.; Sangiovanni, A.; Colombo, M. Increased expression of vascular endothelial growth factor in small hepatocellular carcinoma. J. Viral Hepat. 2007, 14, 133–139. [Google Scholar] [CrossRef]
- Sheen, I.-S.; Jeng, K.-S.; Shih, S.-C.; Kao, C.-R.; Chang, W.-H.; Wang, H.-Y.; Wang, P.-C.; Wang, T.-E.; Shyung, L.-R.; Chen, C.-Z. Clinical significance of the expression of isoform 165 vascular endothelial growth factor mRNA in noncancerous liver remnants of patients with hepatocellular carcinoma. World J. Gastroenterol. 2005, 11, 187–192. [Google Scholar] [CrossRef]
- Chesnokov, M.S.; Khesina, P.A.; Shavochkina, D.A.; Kustova, I.F.; Dyakov, L.M.; Morozova, O.V.; Mugue, N.S.; Kudashkin, N.E.; Moroz, E.A.; Patyutko, Y.I.; et al. Shift in VEGFA isoform balance towards more angiogenic variants is associated with tumor stage and differentiation of human hepatocellular carcinoma. PeerJ 2018, 6, e4915. [Google Scholar] [CrossRef]
- Horwitz, E.; Stein, I.; Andreozzi, M.; Nemeth, J.; Shoham, A.; Pappo, O.; Schweitzer, N.; Tornillo, L.; Kanarek, N.; Quagliata, L.; et al. Human and mouse VEGFA-amplified hepatocellular carcinomas are highly sensitive to sorafenib treatment. Cancer Discov. 2014, 4, 730–743. [Google Scholar] [CrossRef]
- Scartozzi, M.; Faloppi, L.; Svegliati Baroni, G.; Loretelli, C.; Piscaglia, F.; Iavarone, M.; Toniutto, P.; Fava, G.; De Minicis, S.; Mandolesi, A.; et al. VEGF and VEGFR genotyping in the prediction of clinical outcome for HCC patients receiving sorafenib: The ALICE-1 study. Int. J. Cancer 2014, 135, 1247–1256. [Google Scholar] [CrossRef] [PubMed]
- Pinyol, R.; Sia, D.; Llovet, J.M. Immune Exclusion-Wnt/CTNNB1 class predicts resistance to immunotherapies in HCC. Clin. Cancer Res. 2019, 25, 2021–2023. [Google Scholar] [CrossRef] [PubMed]
- Harding, J.J.; Nandakumar, S.; Armenia, J.; Khalil, D.N.; Albano, M.; Ly, M.; Shia, J.; Hechtman, J.F.; Kundra, R.; El Dika, I.; et al. Prospective genotyping of hepatocellular carcinoma: Clinical implications of next-generation sequencing for matching patients to targeted and immune therapies. Clin. Cancer Res. 2019, 25, 2116–2126. [Google Scholar] [CrossRef]
- Sereno, M.; Moreno, V.; Moreno Rubio, J.; Gómez-Raposo, C.; García Sánchez, S.; Hernández Jusdado, R.; Falagan, S.; Zambrana Tébar, F.; Casado Sáenz, E. A significant response to sorafenib in a woman with advanced lung adenocarcinoma and a BRAF non-V600 mutation. Anti-Cancer Drugs 2015, 26, 1004–1007. [Google Scholar] [CrossRef] [PubMed]
- Casadei Gardini, A.; Chiadini, E.; Faloppi, L.; Marisi, G.; Delmonte, A.; Scartozzi, M.; Loretelli, C.; Lucchesi, A.; Oboldi, D.; Dubini, A.; et al. Efficacy of sorafenib in BRAF-mutated non-small-cell lung cancer (NSCLC) and no response in synchronous BRAF wild type-hepatocellular carcinoma: A case report. BMC Cancer 2016, 16, 429. [Google Scholar] [CrossRef] [PubMed]
- Llovet, J.M.; Bruix, J. Molecular targeted therapies in hepatocellular carcinoma. Hepatology 2008, 48, 1312–1327. [Google Scholar] [CrossRef]
- Zhu, Y.-J.; Zheng, B.; Wang, H.-Y.; Chen, L. New knowledge of the mechanisms of sorafenib resistance in liver cancer. Acta Pharmacol. Sin. 2017, 38, 614–622. [Google Scholar] [CrossRef]
- Ito, Y.; Takeda, T.; Sakon, M.; Tsujimoto, M.; Higashiyama, S.; Noda, K.; Miyoshi, E.; Monden, M.; Matsuura, N. Expression and clinical significance of erb-B receptor family in hepatocellular carcinoma. Br. J. Cancer 2001, 84, 1377–1383. [Google Scholar] [CrossRef]
- Wang, H.; Jiang, H.; Zhou, M.; Xu, Z.; Liu, S.; Shi, B.; Yao, X.; Yao, M.; Gu, J.; Li, Z. Epidermal growth factor receptor vIII enhances tumorigenicity and resistance to 5-fluorouracil in human hepatocellular carcinoma. Cancer Lett. 2009, 279, 30–38. [Google Scholar] [CrossRef]
- Blivet-Van Eggelpoël, M.-J.; Chettouh, H.; Fartoux, L.; Aoudjehane, L.; Barbu, V.; Rey, C.; Priam, S.; Housset, C.; Rosmorduc, O.; Desbois-Mouthon, C. Epidermal growth factor receptor and HER-3 restrict cell response to sorafenib in hepatocellular carcinoma cells. J. Hepatol. 2012, 57, 108–115. [Google Scholar] [CrossRef]
- Hoffmann, K.; Xiao, Z.; Franz, C.; Mohr, E.; Serba, S.; Büchler, M.W.; Schemmer, P. Involvement of the epidermal growth factor receptor in the modulation of multidrug resistance in human hepatocellular carcinoma cells in vitro. Cancer Cell Int. 2011, 11, 40. [Google Scholar] [CrossRef] [PubMed]
- Ezzoukhry, Z.; Louandre, C.; Trécherel, E.; Godin, C.; Chauffert, B.; Dupont, S.; Diouf, M.; Barbare, J.-C.; Mazière, J.-C.; Galmiche, A. EGFR activation is a potential determinant of primary resistance of hepatocellular carcinoma cells to sorafenib. Int. J. Cancer 2012, 131, 2961–2969. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Zhou, X.; Shen, H.; Wang, D.; Wang, Y. Phosphorylated ERK is a potential predictor of sensitivity to sorafenib when treating hepatocellular carcinoma: Evidence from an in vitrostudy. BMC Med. 2009, 7, 41. [Google Scholar] [CrossRef]
- Han, C.; Liao, X.; Qin, W.; Yu, L.; Liu, X.; Chen, G.; Liu, Z.; Lu, S.; Chen, Z.; Su, H.; et al. EGFR and SYNE2 are associated with p21 expression and SYNE2 variants predict post-operative clinical outcomes in HBV-related hepatocellular carcinoma. Sci. Rep. 2016, 6, 31237. [Google Scholar] [CrossRef] [PubMed]
- Sueangoen, N.; Tantiwetrueangdet, A.; Panvichian, R. HCC-derived EGFR mutants are functioning, EGF-dependent, and erlotinib-resistant. Cell Biosci. 2020, 10, 41. [Google Scholar] [CrossRef] [PubMed]
- Lathia, C.; Lettieri, J.; Cihon, F.; Gallentine, M.; Radtke, M.; Sundaresan, P. Lack of effect of ketoconazole-mediated CYP3A inhibition on sorafenib clinical pharmacokinetics. Cancer Chemother. Pharmacol. 2006, 57, 685–692. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.G.; Wang, Z.H.; Dong, W.; He, X.D.; Liu, F.C.; Liu, H. Specific CYP450 genotypes in the Chinese population affect sorafenib toxicity in HBV/HCV-associated hepatocellular carcinoma patients. Biomed. Environ. Sci. BES 2018, 31, 586–595. [Google Scholar] [CrossRef]
- Zanger, U.M.; Schwab, M. Cytochrome P450 enzymes in drug metabolism: Regulation of gene expression, enzyme activities, and impact of genetic variation. Pharmacol. Ther. 2013, 138, 103–141. [Google Scholar] [CrossRef]
- Bins, S.; Lenting, A.; El Bouazzaoui, S.; Van Doorn, L.; Oomen-de Hoop, E.; Eskens, F.A.; Van Schaik, R.H.; Mathijssen, R.H. Polymorphisms in SLCO1B1 and UGT1A1 are associated with sorafenib-induced toxicity. Pharmacogenomics 2016, 17, 1483–1490. [Google Scholar] [CrossRef]
- Peer, C.J.; Sissung, T.M.; Kim, A.; Jain, L.; Woo, S.; Gardner, E.R.; Kirkland, C.T.; Troutman, S.M.; English, B.C.; Richardson, E.D.; et al. Sorafenib is an inhibitor of UGT1A1 but is metabolized by UGT1A9: Implications of genetic variants on pharmacokinetics and hyperbilirubinemia. Clin. Cancer Res. 2012, 18, 2099–2107. [Google Scholar] [CrossRef]
- Cox, J.; Weinman, S. Mechanisms of doxorubicin resistance in hepatocellular carcinoma. Hepatic Oncol. 2016, 3, 57–59. [Google Scholar] [CrossRef] [PubMed]
- Tell, G.; Quadrifoglio, F.; Tiribelli, C.; Kelley, M.R. The many functions of APE1/Ref-1: Not only a DNA repair enzyme. Antioxid. Redox Signal. 2009, 11, 601–620. [Google Scholar] [CrossRef]
- Pascut, D.; Sukowati, C.H.C.; Antoniali, G.; Mangiapane, G.; Burra, S.; Mascaretti, L.G.; Buonocore, M.R.; Crocè, L.S.; Tiribelli, C.; Tell, G. Serum AP-endonuclease 1 (sAPE1) as novel biomarker for hepatocellular carcinoma. Oncotarget 2019, 10, 383–394. [Google Scholar] [CrossRef] [PubMed]
- Di Maso, V.; Avellini, C.; Crocè, L.S.; Rosso, N.; Quadrifoglio, F.; Cesaratto, L.; Codarin, E.; Bedogni, G.; Beltrami, C.A.; Tell, G.; et al. Subcellular localization of APE1/Ref-1 in human hepatocellular carcinoma: Possible prognostic significance. Mol. Med. 2007, 13, 89–96. [Google Scholar] [CrossRef] [PubMed]
- Chattopadhyay, R.; Das, S.; Maiti, A.K.; Boldogh, I.; Xie, J.; Hazra, T.K.; Kohno, K.; Mitra, S.; Bhakat, K.K. Regulatory role of human AP-endonuclease (APE1/Ref-1) in YB-1-mediated activation of the multidrug resistance gene MDR1. Mol. Cell. Biol. 2008, 28, 7066–7080. [Google Scholar] [CrossRef] [PubMed]
- Sengupta, S.; Mantha, A.K.; Mitra, S.; Bhakat, K.K. Human AP endonuclease (APE1/Ref-1) and its acetylation regulate YB-1-p300 recruitment and RNA polymerase II loading in the drug-induced activation of multidrug resistance gene MDR1. Oncogene 2011, 30, 482–493. [Google Scholar] [CrossRef]
- Zeng, X.; Liu, S.; Yu, H.; Ji, L.; Li, L.; Huang, J.; Bai, H.; Qiu, X. DNA repair capacity, DNA-strand break repair gene polymorphisms, and the incidence of hepatocellular carcinoma in southwestern Guangxi of China. DNA Cell Biol. 2012, 31, 1384–1391. [Google Scholar] [CrossRef]
- Lirussi, L.; Antoniali, G.; D’Ambrosio, C.; Scaloni, A.; Nilsen, H.; Tell, G. APE1 polymorphic variants cause persistent genomic stress and affect cancer cell proliferation. Oncotarget 2016, 7, 26293–26306. [Google Scholar] [CrossRef]
- Cao, L.; Cheng, H.; Jiang, Q.; Li, H.; Wu, Z. APEX1 is a novel diagnostic and prognostic biomarker for hepatocellular carcinoma. Aging 2020, 12, 4573–4591. [Google Scholar] [CrossRef]
- Alekseeva, I.V.; Davletgildeeva, A.T.; Arkova, O.V.; Kuznetsov, N.A.; Fedorova, O.S. The impact of single-nucleotide polymorphisms of human apurinic/apyrimidinic endonuclease 1 on specific DNA binding and catalysis. Biochimie 2019, 163, 73–83. [Google Scholar] [CrossRef]
- Yang, Z.; Zhao, J. Effect of APE1 and XRCC1 gene polymorphism on susceptibility to hepatocellular carcinoma and sensitivity to cisplatin. Int. J. Clin. Exp. Med. 2015, 8, 9931–9936. [Google Scholar] [PubMed]
- Mattar, M.-A.M.; Zekri, A.-R.N.; Hussein, N.; Morsy, H.; Esmat, G.; Amin, M.A. Polymorphisms of base-excision repair genes and the hepatocarcinogenesis. Gene 2018, 675, 62–68. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, S.; Takehara, T.; Hikita, H.; Kodama, T.; Tsunematsu, H.; Miyagi, T.; Hosui, A.; Ishida, H.; Tatsumi, T.; Kanto, T.; et al. Inhibition of autophagy potentiates the antitumor effect of the multikinase inhibitor sorafenib in hepatocellular carcinoma. Int. J. Cancer 2012, 131, 548–557. [Google Scholar] [CrossRef] [PubMed]
- Pan, J.; Lu, C.; Jun, W.; Wu, Y.; Shi, X.; Ding, Y. The up-regulation of P62 levels is associated with resistance of sorafenib in hepatocarcinoma cells. Int. J. Clin. Exp. Pathol. 2019, 12, 2622–2630. [Google Scholar] [PubMed]
- Sun, T.; Liu, H.; Ming, L. Multiple roles of autophagy in the sorafenib resistance of hepatocellular carcinoma. Cell. Physiol. Biochem. 2017, 44, 716–727. [Google Scholar] [CrossRef]
- Singh, R.; Cuervo, A.M. Autophagy in the cellular energetic balance. Cell Metab. 2011, 13, 495–504. [Google Scholar] [CrossRef]
- Zhang, K.; Chen, J.; Zhou, H.; Chen, Y.; Zhi, Y.; Zhang, B.; Chen, L.; Chu, X.; Wang, R.; Zhang, C. PU.1/microRNA-142-3p targets ATG5/ATG16L1 to inactivate autophagy and sensitize hepatocellular carcinoma cells to sorafenib. Cell Death Dis. 2018, 9, 312. [Google Scholar] [CrossRef]
- Kharaziha, P.; Chioureas, D.; Baltatzis, G.; Fonseca, P.; Rodriguez, P.; Gogvadze, V.; Lennartsson, L.; Björklund, A.-C.; Zhivotovsky, B.; Grandér, D.; et al. Sorafenib-induced defective autophagy promotes cell death by necroptosis. Oncotarget 2015, 6, 37066–37082. [Google Scholar] [CrossRef]
- Li, N.; Fan, X.; Wang, X.; Deng, H.; Zhang, K.; Zhang, X.; Han, Q.; Lv, Y.; Liu, Z. Autophagy-related 5 gene rs510432 polymorphism is associated with hepatocellular carcinoma in patients with chronic hepatitis B virus infection. Immunol. Invest. 2019, 48, 378–391. [Google Scholar] [CrossRef]
- Li, N.; Fan, X.; Wang, X.; Zhang, X.; Zhang, K.; Han, Q.; Lv, Y.; Liu, Z. Genetic association of polymorphisms at the intergenic region between PRDM1 and ATG5 with hepatitis B virus infection in Han Chinese patients. J. Med. Virol. 2019. [Google Scholar] [CrossRef]
- Shen, M.; Lin, L. Functional variants of autophagy-related genes are associated with the development of hepatocellular carcinoma. Life Sci. 2019, 235, 116675. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Dang, H.; Wang, X.W. The significance of intertumor and intratumor heterogeneity in liver cancer. Exp. Mol. Med. 2018, 50, e416. [Google Scholar] [CrossRef] [PubMed]
- Sukowati, C.H.C. Heterogeneity of Hepatic Cancer Stem Cells. Adv. Exp. Med. Biol. 2019, 1139, 59–81. [Google Scholar] [CrossRef] [PubMed]
- Sukowati, C.H.C.; Anfuso, B.; Torre, G.; Francalanci, P.; Crocè, L.S.; Tiribelli, C. The expression of CD90/Thy-1 in hepatocellular carcinoma: An in vivo and in vitro study. PloS One 2013, 8, e76830. [Google Scholar] [CrossRef]
- Jia, Q.; Zhang, X.; Deng, T.; Gao, J. Positive correlation of Oct4 and ABCG2 to chemotherapeutic resistance in CD90(+)CD133(+) liver cancer stem cells. Cell. Reprogram. 2013, 15, 143–150. [Google Scholar] [CrossRef]
- Hu, C.; Li, H.; Li, J.; Zhu, Z.; Yin, S.; Hao, X.; Yao, M.; Zheng, S.; Gu, J. Analysis of ABCG2 expression and side population identifies intrinsic drug efflux in the HCC cell line MHCC-97L and its modulation by Akt signaling. Carcinogenesis 2008, 29, 2289–2297. [Google Scholar] [CrossRef]
- Bort, A.; Sánchez, B.G.; Mateos-Gómez, P.A.; Vara-Ciruelos, D.; Rodríguez-Henche, N.; Díaz-Laviada, I. Targeting AMP-activated kinase impacts hepatocellular cancer stem cells induced by long-term treatment with sorafenib. Mol. Oncol. 2019, 13, 1311–1331. [Google Scholar] [CrossRef]
- Kahraman, D.C.; Kahraman, T.; Cetin-Atalay, R. Targeting PI3K/Akt/mTOR pathway identifies differential expression and functional role of IL8 in liver cancer stem cell enrichment. Mol. Cancer Ther. 2019, 18, 2146–2157. [Google Scholar] [CrossRef]
- Qiu, Y.; Shan, W.; Yang, Y.; Jin, M.; Dai, Y.; Yang, H.; Jiao, R.; Xia, Y.; Liu, Q.; Ju, L.; et al. Reversal of sorafenib resistance in hepatocellular carcinoma: Epigenetically regulated disruption of 14-3-3η/hypoxia-inducible factor-1α. Cell Death Discov. 2019, 5, 120. [Google Scholar] [CrossRef]
- Huang, M.; Chen, C.; Geng, J.; Han, D.; Wang, T.; Xie, T.; Wang, L.; Wang, Y.; Wang, C.; Lei, Z.; et al. Targeting KDM1A attenuates Wnt/β-catenin signaling pathway to eliminate sorafenib-resistant stem-like cells in hepatocellular carcinoma. Cancer Lett. 2017, 398, 12–21. [Google Scholar] [CrossRef]
- Xin, H.-W.; Ambe, C.M.; Hari, D.M.; Wiegand, G.W.; Miller, T.C.; Chen, J.-Q.; Anderson, A.J.; Ray, S.; Mullinax, J.E.; Koizumi, T.; et al. Label-retaining liver cancer cells are relatively resistant to sorafenib. Gut 2013, 62, 1777–1786. [Google Scholar] [CrossRef]
- Kahraman, D.C.; Hanquet, G.; Jeanmart, L.; Lanners, S.; Šramel, P.; Boháč, A.; Cetin-Atalay, R. Quinoides and VEGFR2 TKIs influence the fate of hepatocellular carcinoma and its cancer stem cells. MedChemComm 2017, 8, 81–87. [Google Scholar] [CrossRef] [PubMed]
- Zhai, B.; Zhang, X.; Sun, B.; Cao, L.; Zhao, L.; Li, J.; Ge, N.; Chen, L.; Qian, H.; Yin, Z. MK2206 overcomes the resistance of human liver cancer stem cells to sorafenib by inhibition of pAkt and upregulation of pERK. Tumor Biol. 2016, 37, 8047–8055. [Google Scholar] [CrossRef] [PubMed]
- Colombo, F.; Baldan, F.; Mazzucchelli, S.; Martin-Padura, I.; Marighetti, P.; Cattaneo, A.; Foglieni, B.; Spreafico, M.; Guerneri, S.; Baccarin, M.; et al. Evidence of distinct tumour-propagating cell populations with different properties in primary human hepatocellular carcinoma. PLoS ONE 2011, 6, e21369. [Google Scholar] [CrossRef] [PubMed]
- Gao, Q.; Wang, Z.-C.; Duan, M.; Lin, Y.-H.; Zhou, X.-Y.; Worthley, D.L.; Wang, X.-Y.; Niu, G.; Xia, Y.; Deng, M.; et al. Cell culture system for analysis of genetic heterogeneity within hepatocellular carcinomas and response to pharmacologic agents. Gastroenterology 2017, 152, 232–242. [Google Scholar] [CrossRef] [PubMed]
- Hagiwara, S.; Kudo, M.; Nagai, T.; Inoue, T.; Ueshima, K.; Nishida, N.; Watanabe, T.; Sakurai, T. Activation of JNK and high expression level of CD133 predict a poor response to sorafenib in hepatocellular carcinoma. Br. J. Cancer 2012, 106, 1997–2003. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.H.; Park, J.-W.; Kim, J.S.; Lee, S.-K.; Hong, E.K. Stem cell markers predict the response to sorafenib in patients with hepatocellular carcinoma. Gut Liver 2019, 13, 342–348. [Google Scholar] [CrossRef] [PubMed]
- Turner, N.C.; Reis-Filho, J.S. Genetic heterogeneity and cancer drug resistance. Lancet Oncol. 2012, 13, e178–e185. [Google Scholar] [CrossRef]
- Broutier, L.; Mastrogiovanni, G.; Verstegen, M.M.; Francies, H.E.; Gavarró, L.M.; Bradshaw, C.R.; Allen, G.E.; Arnes-Benito, R.; Sidorova, O.; Gaspersz, M.P.; et al. Human primary liver cancer–derived organoid cultures for disease modeling and drug screening. Nat. Med. 2017, 23, 1424–1435. [Google Scholar] [CrossRef]
- Wang, X.; Zhang, H.; Chen, X. Drug resistance and combating drug resistance in cancer. Cancer Drug Resist. 2019. [Google Scholar] [CrossRef]
| Factors | Mechanism Contributing to Resistance | Molecular Markers | Effect to Chemotherapy or Cellular Function |
|---|---|---|---|
| Drug Concentration | |||
| Drug efflux via ABC transporters | Overexpression of specific ABC transporters; genetic polymorphism | ABCB1; ABCC1; ABCC2; ABCC3; ABCG2 | Decreased intracellular drug concentration |
| Drug uptake via SLC transport proteins | Downregulation of SLC transport proteins; genetic polymorphism; localization (plasma membrane expression only) | SLC22A1; SLC22A3; SLC46A3 | Decreased intracellular drug concentration |
| Cellular Processes | |||
| Alteration of target molecule | Overexpression of drug target molecules; genetic polymorphism and mutation | Raf; VEGF | Promote angiogenesis; sustained activation of signaling pathway promoting cellular proliferation and growth |
| Alteration in signaling pathway | Overexpression and sustained activation of signaling pathway molecules; genetic polymorphism and mutation | EGFR Ras/Raf/MEK/ERK pathway) | Sustained activation of signaling pathway promoting cellular proliferation and growth |
| Intracellular drug metabolism | Genetic polymorphism in metabolizing enzymes | CYP3A5; CYP450; UGT1A9; UGT1A1 | Decrease in drug metabolism |
| DNA repair | Overexpression sustained activation of DNA repair proteins; genetic polymorphism | APE1 | Promote DNA repair therefore inhibiting apoptosis |
| Autophagy | Stress response that function as double-edged; Agonist or antagonist to treatment | ATGs | Promote cell survival and proliferation or otherwise |
| Intra- and Inter-Cellular Properties | |||
| Cancer Stem Cells | Presence of cancer stem cell properties poses survival advantage | CD133, CD90, EpCAM, ABCG2 | Failed therapy |
| Tumor genetic heterogeneity | Tumoral heterogeneity dictates differential sensitivity of cells to chemotherapy within a tissue; cells ability for clonal expansion allows survival of resistant cells | - | Failed therapy |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cabral, L.K.D.; Tiribelli, C.; Sukowati, C.H.C. Sorafenib Resistance in Hepatocellular Carcinoma: The Relevance of Genetic Heterogeneity. Cancers 2020, 12, 1576. https://doi.org/10.3390/cancers12061576
Cabral LKD, Tiribelli C, Sukowati CHC. Sorafenib Resistance in Hepatocellular Carcinoma: The Relevance of Genetic Heterogeneity. Cancers. 2020; 12(6):1576. https://doi.org/10.3390/cancers12061576
Chicago/Turabian StyleCabral, Loraine Kay D., Claudio Tiribelli, and Caecilia H. C. Sukowati. 2020. "Sorafenib Resistance in Hepatocellular Carcinoma: The Relevance of Genetic Heterogeneity" Cancers 12, no. 6: 1576. https://doi.org/10.3390/cancers12061576
APA StyleCabral, L. K. D., Tiribelli, C., & Sukowati, C. H. C. (2020). Sorafenib Resistance in Hepatocellular Carcinoma: The Relevance of Genetic Heterogeneity. Cancers, 12(6), 1576. https://doi.org/10.3390/cancers12061576






