Bone Morphogenetic Protein 4 Targeting Glioma Stem-Like Cells for Malignant Glioma Treatment: Latest Advances and Implications for Clinical Application
Abstract
1. Introduction
2. BMP4 Signal Pathways and Glioma Biology
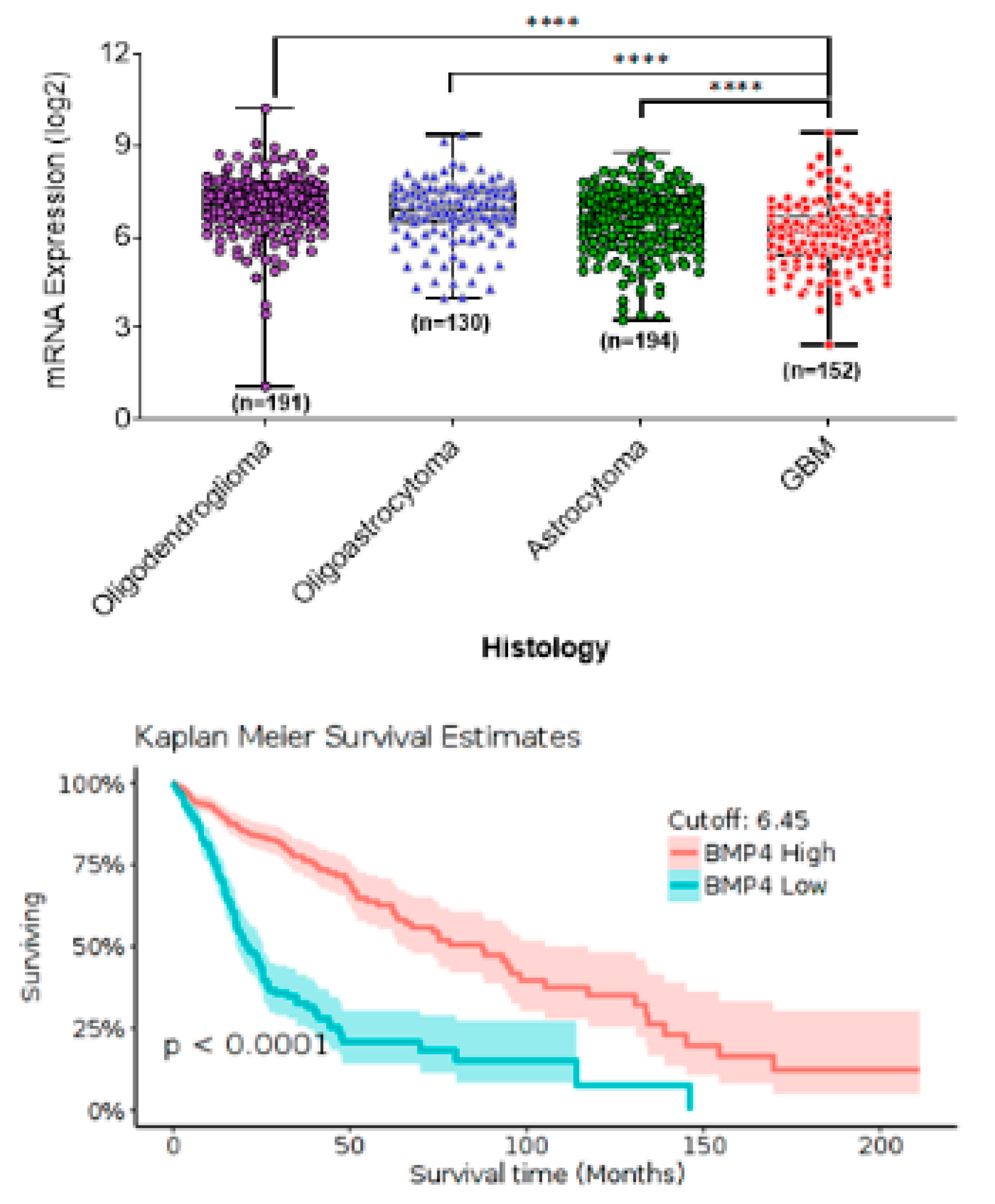
3. BMP4 Signaling in Adult High-Grade Gliomas
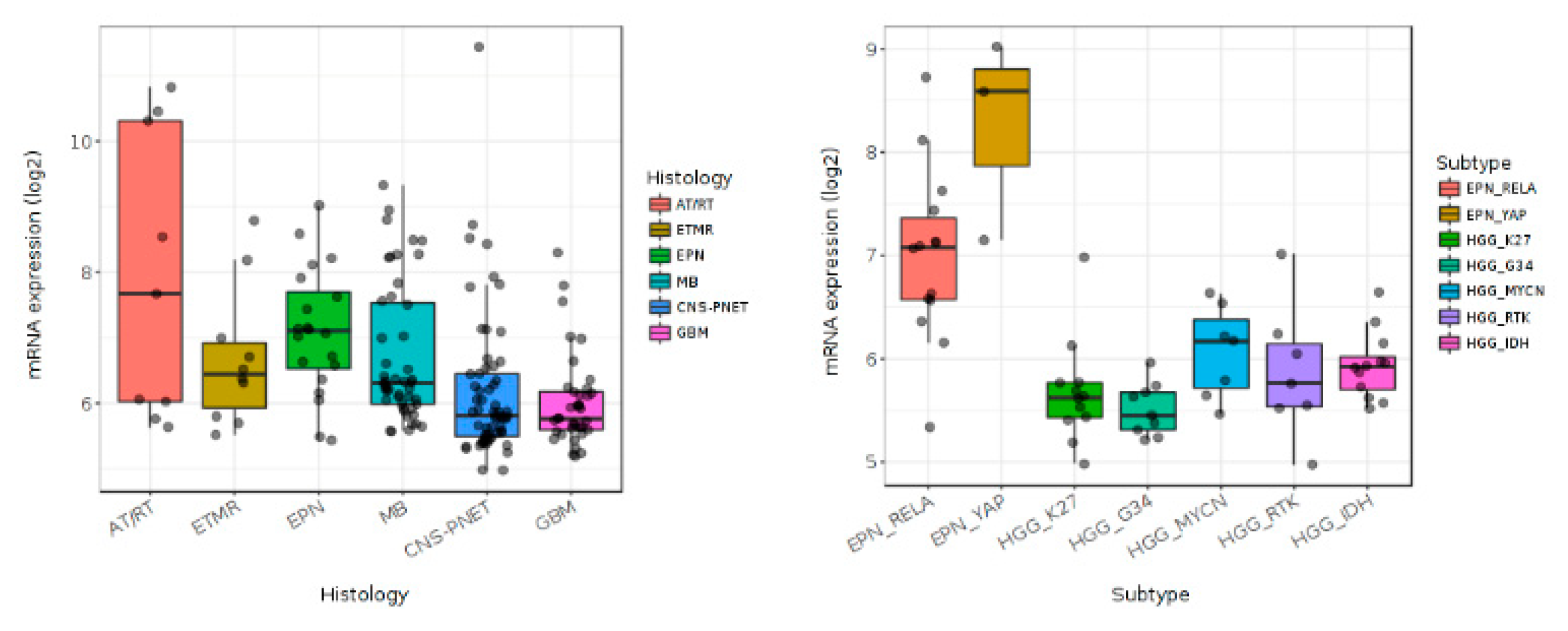
4. BMP4 Signaling in Pediatric High-Grade Gliomas
5. BMP4 Delivery Methods for Glioma Treatment
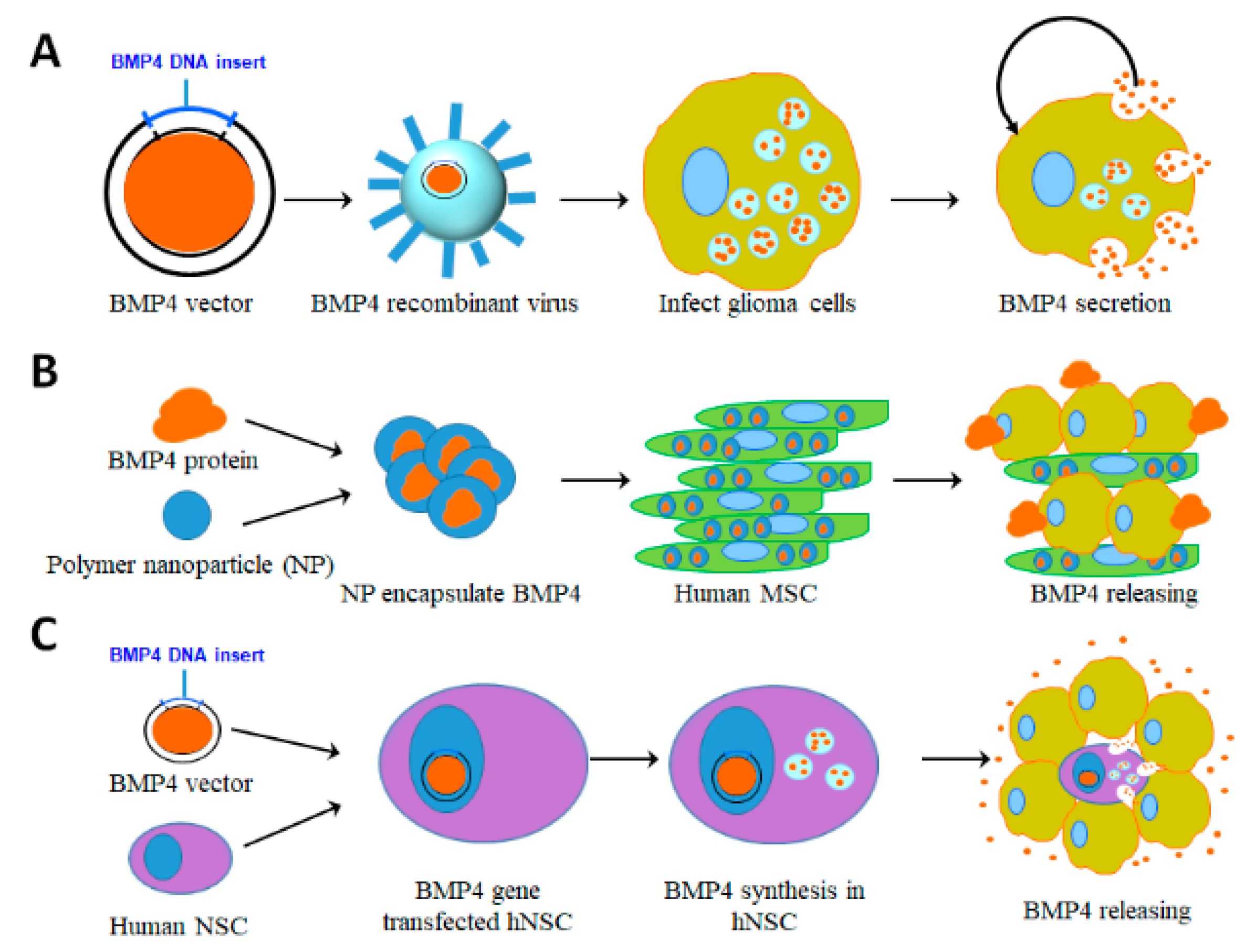
5.1. Viral Vector Based Delivery
5.2. Human Adipose-Derived Mesenchymal Stem Cell (hAMSC) Based Delivery
5.3. Human Neural Stem Cell (hNSC) Based Delivery
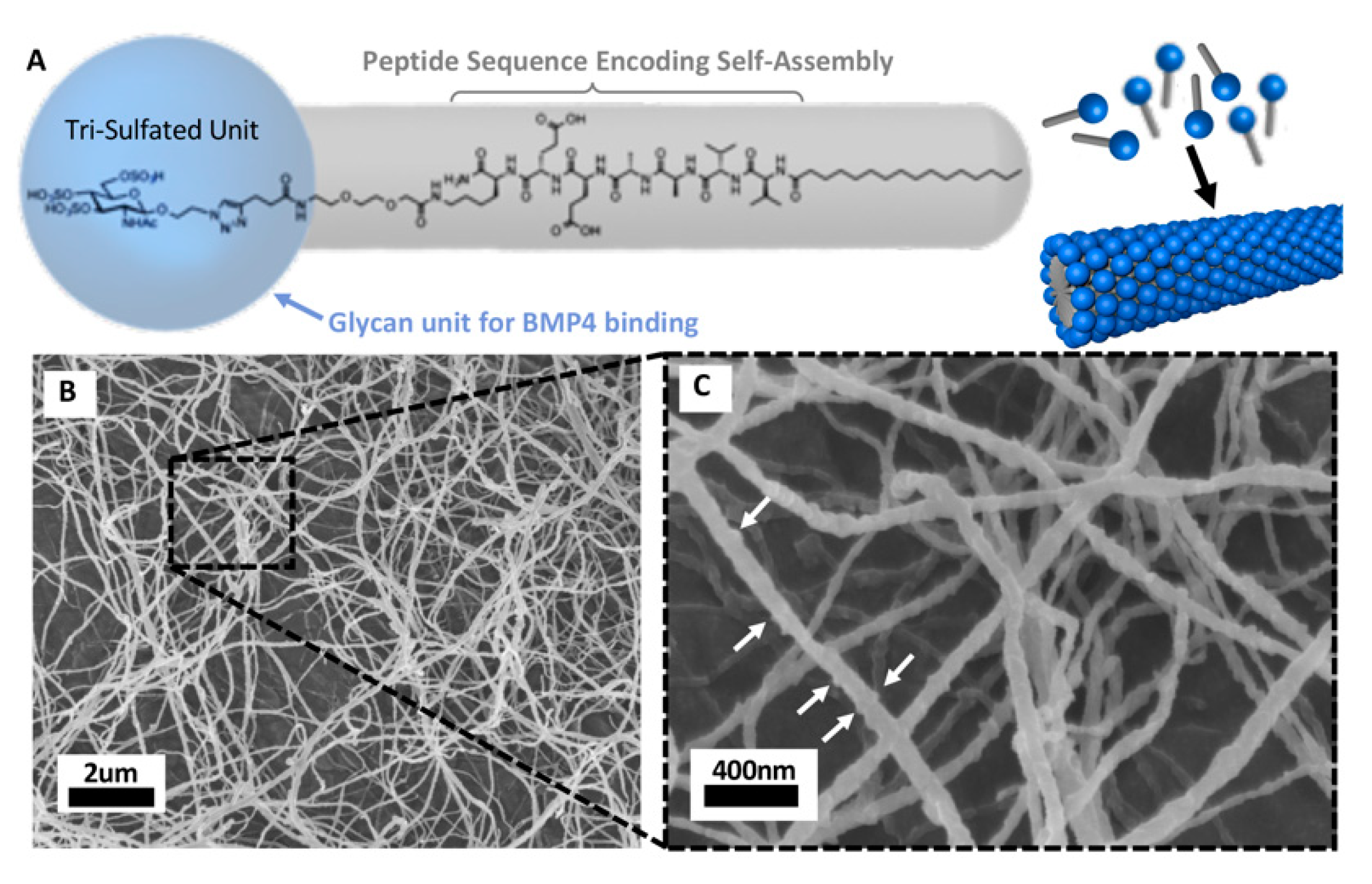
5.4. Biocompatible Nanomaterial Based Delivery
6. Future Prospects
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ostrom, Q.T.; Gittleman, H.; Liao, P.; Vecchione-Koval, T.; Wolinsky, Y.; Kruchko, C.; Barnholtz-Sloan, J.S. CBTRUS Statistical Report: Primary brain and other central nervous system tumors diagnosed in the United States in 2010–2014. Neuro-Oncology 2017, 19, v1–v88. [Google Scholar] [CrossRef]
- Aldape, K.; Brindle, K.M.; Chesler, L.; Chopra, R.; Gajjar, A.; Gilbert, M.R.; Gottardo, N.; Gutmann, D.H.; Hargrave, D.; Holland, E.C.; et al. Challenges to curing primary brain tumours. Nature reviews. Clin. Oncol. 2019, 16, 509–520. [Google Scholar] [CrossRef]
- Smith, J.S.; Jenkins, R.B. Genetic alterations in adult diffuse glioma: Occurrence, significance, and prognostic implications. Front. Biosci. 2000, 5, D213–D231. [Google Scholar] [CrossRef]
- Song, K.S.; Phi, J.H.; Cho, B.K.; Wang, K.C.; Lee, J.Y.; Kim, D.G.; Kim, I.H.; Ahn, H.S.; Park, S.H.; Kim, S.K. Long-term outcomes in children with glioblastoma. Journal of neurosurgery. Pediatrics 2010, 6, 145–149. [Google Scholar] [CrossRef]
- Johnson, B.E.; Mazor, T.; Hong, C.; Barnes, M.; Aihara, K.; McLean, C.Y.; Fouse, S.D.; Yamamoto, S.; Ueda, H.; Tatsuno, K.; et al. Mutational analysis reveals the origin and therapy-driven evolution of recurrent glioma. Science 2014, 343, 189–193. [Google Scholar] [CrossRef]
- Nishikawa, R. Pediatric and adult gliomas: How different are they? Neuro-Oncology 2010, 12, 1203–1204. [Google Scholar] [CrossRef]
- Sturm, D.; Bender, S.; Jones, D.T.; Lichter, P.; Grill, J.; Becher, O.; Hawkins, C.; Majewski, J.; Jones, C.; Costello, J.F.; et al. Paediatric and adult glioblastoma: Multiform (epi)genomic culprits emerge. Nature reviews. Cancer 2014, 14, 92–107. [Google Scholar] [CrossRef]
- Yan, H.; Parsons, D.W.; Jin, G.; McLendon, R.; Rasheed, B.A.; Yuan, W.; Kos, I.; Batinic-Haberle, I.; Jones, S.; Riggins, G.J.; et al. IDH1 and IDH2 mutations in gliomas. N. Engl. J. Med. 2009, 360, 765–773. [Google Scholar] [CrossRef]
- Capper, D.; Jones, D.T.W.; Sill, M.; Hovestadt, V.; Schrimpf, D.; Sturm, D.; Koelsche, C.; Sahm, F.; Chavez, L.; Reuss, D.E.; et al. DNA methylation-based classification of central nervous system tumours. Nature 2018, 555, 469–474. [Google Scholar] [CrossRef]
- Patel, A.P.; Tirosh, I.; Trombetta, J.J.; Shalek, A.K.; Gillespie, S.M.; Wakimoto, H.; Cahill, D.P.; Nahed, B.V.; Curry, W.T.; Martuza, R.L.; et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 2014, 344, 1396–1401. [Google Scholar] [CrossRef]
- Eskilsson, E.; Rosland, G.V.; Solecki, G.; Wang, Q.; Harter, P.N.; Graziani, G.; Verhaak, R.G.W.; Winkler, F.; Bjerkvig, R.; Miletic, H. EGFR heterogeneity and implications for therapeutic intervention in glioblastoma. Neuro-Oncology 2018, 20, 743–752. [Google Scholar] [CrossRef]
- Mackay, A.; Burford, A.; Carvalho, D.; Izquierdo, E.; Fazal-Salom, J.; Taylor, K.R.; Bjerke, L.; Clarke, M.; Vinci, M.; Nandhabalan, M.; et al. Integrated Molecular Meta-Analysis of 1,000 Pediatric High-Grade and Diffuse Intrinsic Pontine Glioma. Cancer Cell 2017, 32, 520–537.e525. [Google Scholar] [CrossRef]
- Wu, G.; Broniscer, A.; McEachron, T.A.; Lu, C.; Paugh, B.S.; Becksfort, J.; Qu, C.; Ding, L.; Huether, R.; Parker, M.; et al. Somatic histone H3 alterations in pediatric diffuse intrinsic pontine gliomas and non-brainstem glioblastomas. Nat. Genet. 2012, 44, 251–253. [Google Scholar] [CrossRef]
- Schwartzentruber, J.; Korshunov, A.; Liu, X.Y.; Jones, D.T.; Pfaff, E.; Jacob, K.; Sturm, D.; Fontebasso, A.M.; Quang, D.A.; Tonjes, M.; et al. Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 2012, 482, 226–231. [Google Scholar] [CrossRef]
- Buczkowicz, P.; Hoeman, C.; Rakopoulos, P.; Pajovic, S.; Letourneau, L.; Dzamba, M.; Morrison, A.; Lewis, P.; Bouffet, E.; Bartels, U.; et al. Genomic analysis of diffuse intrinsic pontine gliomas identifies three molecular subgroups and recurrent activating ACVR1 mutations. Nat. Genet. 2014, 46, 451–456. [Google Scholar] [CrossRef]
- Fontebasso, A.M.; Papillon-Cavanagh, S.; Schwartzentruber, J.; Nikbakht, H.; Gerges, N.; Fiset, P.O.; Bechet, D.; Faury, D.; De Jay, N.; Ramkissoon, L.A.; et al. Recurrent somatic mutations in ACVR1 in pediatric midline high-grade astrocytoma. Nat. Genet. 2014, 46, 462–466. [Google Scholar] [CrossRef]
- Taylor, K.R.; Mackay, A.; Truffaux, N.; Butterfield, Y.; Morozova, O.; Philippe, C.; Castel, D.; Grasso, C.S.; Vinci, M.; Carvalho, D.; et al. Recurrent activating ACVR1 mutations in diffuse intrinsic pontine glioma. Nat. Genet. 2014, 46, 457–461. [Google Scholar] [CrossRef]
- Singh, S.K.; Clarke, I.D.; Hide, T.; Dirks, P.B. Cancer stem cells in nervous system tumors. Oncogene 2004, 23, 7267–7273. [Google Scholar] [CrossRef]
- Hemmati, H.D.; Nakano, I.; Lazareff, J.A.; Masterman-Smith, M.; Geschwind, D.H.; Bronner-Fraser, M.; Kornblum, H.I. Cancerous stem cells can arise from pediatric brain tumors. Proc. Natl. Acad. Sci. USA 2003, 100, 15178–15183. [Google Scholar] [CrossRef]
- Bhaduri, A.; Di Lullo, E.; Jung, D.; Müller, S.; Crouch, E.E.; Espinosa, C.S.; Ozawa, T.; Alvarado, B.; Spatazza, J.; Cadwell, C.R.; et al. Outer Radial Glia-like Cancer Stem Cells Contribute to Heterogeneity of Glioblastoma. Cell Stem Cell 2020, 26, 48–63. [Google Scholar] [CrossRef]
- Das, S.; Srikanth, M.; Kessler, J.A. Cancer stem cells and glioma. Nat. Clin. Pract. Neurol. 2008, 4, 427–435. [Google Scholar] [CrossRef]
- Bao, S.; Wu, Q.; McLendon, R.E.; Hao, Y.; Shi, Q.; Hjelmeland, A.B.; Dewhirst, M.W.; Bigner, D.D.; Rich, J.N. Glioma stem cells promote radioresistance by preferential activation of the DNA damage response. Nature 2006, 444, 756–760. [Google Scholar] [CrossRef]
- Weng, Q.; Wang, J.; Wang, J.; He, D.; Cheng, Z.; Zhang, F.; Verma, R.; Xu, L.; Dong, X.; Liao, Y.; et al. Single-Cell Transcriptomics Uncovers Glial Progenitor Diversity and Cell Fate Determinants during Development and Gliomagenesis. Cell Stem Cell 2019, 24, 707–723. [Google Scholar] [CrossRef]
- Chen, J.; Li, Y.; Yu, T.S.; McKay, R.M.; Burns, D.K.; Kernie, S.G.; Parada, L.F. A restricted cell population propagates glioblastoma growth after chemotherapy. Nature 2012, 488, 522–526. [Google Scholar] [CrossRef]
- Hussein, D.; Punjaruk, W.; Storer, L.C.; Shaw, L.; Othman, R.; Peet, A.; Miller, S.; Bandopadhyay, G.; Heath, R.; Kumari, R.; et al. Pediatric brain tumor cancer stem cells: Cell cycle dynamics, DNA repair, and etoposide extrusion. Neuro-Oncology 2011, 13, 70–83. [Google Scholar] [CrossRef]
- Parada, L.F.; Dirks, P.B.; Wechsler-Reya, R.J. Brain Tumor Stem Cells Remain in Play. J. Clin. Oncol. 2017, 35, 2428–2431. [Google Scholar] [CrossRef]
- Yokoyama, Y.; Watanabe, T.; Tamura, Y.; Hashizume, Y.; Miyazono, K.; Ehata, S. Autocrine BMP-4 Signaling Is a Therapeutic Target in Colorectal Cancer. Cancer Res. 2017, 77, 4026–4038. [Google Scholar] [CrossRef]
- Chiou, J.; Su, C.-Y.; Jan, Y.-H.; Yang, C.-J.; Huang, M.-S.; Yu, Y.-L.; Hsiao, M. Decrease of FSTL1-BMP4-Smad signaling predicts poor prognosis in lung adenocarcinoma but not in squamous cell carcinoma. Sci. Rep. 2017, 7, 9830. [Google Scholar] [CrossRef]
- Park, W.Y.; Hong, B.J.; Lee, J.; Choi, C.; Kim, M.Y. H3K27 Demethylase JMJD3 Employs the NF-kappaB and BMP Signaling Pathways to Modulate the Tumor Microenvironment and Promote Melanoma Progression and Metastasis. Cancer Res. 2016, 76, 161–170. [Google Scholar] [CrossRef]
- Martinez, V.G.; Rubio, C.; Martinez-Fernandez, M.; Segovia, C.; Lopez-Calderon, F.; Garin, M.; Teijeira, A.; Munera-Maravilla, E.; Varas, A.; Sacedon, R.; et al. BMP4 induces M2 macrophage polarization and favors tumor progression in bladder cancer. Clin. Cancer Res. 2017, 23, 7388–7399. [Google Scholar] [CrossRef]
- Zhang, L.; Sun, H.; Zhao, F.; Lu, P.; Ge, C.; Li, H.; Hou, H.; Yan, M.; Chen, T.; Jiang, G.; et al. BMP4 administration induces differentiation of CD133+ hepatic cancer stem cells, blocking their contributions to hepatocellular carcinoma. Cancer Res. 2012, 72, 4276–4285. [Google Scholar] [CrossRef]
- Yacqub-Usman, K.; Duong, C.V.; Clayton, R.N.; Farrell, W.E. Epigenomic silencing of the BMP-4 gene in pituitary adenomas: A potential target for epidrug-induced re-expression. Endocrinology 2012, 153, 3603–3612. [Google Scholar] [CrossRef]
- Zhao, X.; Sun, Q.; Dou, C.; Chen, Q.; Liu, B. BMP4 inhibits glioblastoma invasion by promoting E-cadherin and claudin expression. Front. Biosci. 2019, 24, 1060–1070. [Google Scholar]
- Jin, X.; Nie, E.; Zhou, X.; Zeng, A.; Yu, T.; Zhi, T.; Jiang, K.; Wang, Y.; Zhang, J.; You, Y. Fstl1 Promotes Glioma Growth Through the BMP4/Smad1/5/8 Signaling Pathway. Cell. Physiol. Biochem. 2017, 44, 1616–1628. [Google Scholar] [CrossRef]
- Liu, S.; Yin, F.; Fan, W.; Wang, S.; Guo, X.R.; Zhang, J.N.; Tian, Z.M.; Fan, M. Over-expression of BMPR-IB reduces the malignancy of glioblastoma cells by upregulation of p21 and p27Kip1. J. Exp. Clin. Cancer Res. 2012, 31, 52. [Google Scholar] [CrossRef]
- Liu, B.; Chen, Q.; Tian, D.; Wu, L.; Dong, H.; Wang, J.; Ji, B.; Zhu, X.; Cai, Q.; Wang, L.; et al. BMP4 reverses multidrug resistance through modulation of BCL-2 and GDNF in glioblastoma. Brain Res. 2013, 1507, 115–124. [Google Scholar] [CrossRef]
- Raja, E.; Komuro, A.; Tanabe, R.; Sakai, S.; Ino, Y.; Saito, N.; Todo, T.; Morikawa, M.; Aburatani, H.; Koinuma, D.; et al. Bone morphogenetic protein signaling mediated by ALK-2 and DLX2 regulates apoptosis in glioma-initiating cells. Oncogene 2017, 36, 4963–4974. [Google Scholar] [CrossRef]
- Lee, S.S.; Fyrner, T.; Chen, F.; Álvarez, Z.; Sleep, E.; Chun, D.S.; Weiner, J.A.; Cook, R.W.; Freshman, R.D.; Schallmo, M.S.; et al. Sulfated glycopeptide nanostructures for multipotent protein activation. Nat. Nanotechnol. 2017, 12, 821–829. [Google Scholar] [CrossRef]
- Bond, A.M.; Peng, C.Y.; Meyers, E.A.; McGuire, T.; Ewaleifoh, O.; Kessler, J.A. BMP signaling regulates the tempo of adult hippocampal progenitor maturation at multiple stages of the lineage. Stem Cells 2014, 32, 2201–2214. [Google Scholar] [CrossRef]
- Bond, A.M.; Bhalala, O.G.; Kessler, J.A. The dynamic role of bone morphogenetic proteins in neural stem cell fate and maturation. Dev. Neurobiol. 2012, 72, 1068–1084. [Google Scholar] [CrossRef]
- Gross, R.E.; Mehler, M.F.; Mabie, P.C.; Zang, Z.; Santschi, L.; Kessler, J.A. Bone morphogenetic proteins promote astroglial lineage commitment by mammalian subventricular zone progenitor cells. Neuron 1996, 17, 595–606. [Google Scholar] [CrossRef]
- Xi, G.; Best, B.; Mania-Farnell, B.; James, C.D.; Tomita, T. Therapeutic Potential for Bone Morphogenetic Protein 4 in Human Malignant Glioma. Neoplasi 2017, 19, 261–270. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, D.; Taylor, K.R.; Olaciregui, N.G.; Molinari, V.; Clarke, M.; Mackay, A.; Ruddle, R.; Henley, A.; Valenti, M.; Hayes, A.; et al. ALK2 inhibitors display beneficial effects in preclinical models of ACVR1 mutant diffuse intrinsic pontine glioma. Commun. Biol. 2019, 2, 156. [Google Scholar] [CrossRef] [PubMed]
- Taylor, K.R.; Vinci, M.; Bullock, A.N.; Jones, C. ACVR1 mutations in DIPG: Lessons learned from FOP. Cancer Res. 2014, 74, 4565–4570. [Google Scholar] [CrossRef]
- Bowman, R.L.; Wang, Q.; Carro, A.; Verhaak, R.G.; Squatrito, M. GlioVis data portal for visualization and analysis of brain tumor expression datasets. Neuro-Oncology 2017, 19, 139–141. [Google Scholar] [CrossRef]
- Bao, Z.; Zhang, C.; Yan, W.; Liu, Y.; Li, M.; Zhang, W.; Jiang, T. BMP4, a strong better prognosis predictor, has a subtype preference and cell development association in gliomas. J. Transl. Med. 2013, 11, 100. [Google Scholar] [CrossRef]
- Wu, Q.; Yao, J. BMP4, a new prognostic factor for glioma. World J. Surg. Oncol. 2013, 11, 264. [Google Scholar] [CrossRef]
- Rich, J.N.; Bao, S. Chemotherapy and cancer stem cells. Cell Stem Cell 2007, 1, 353–355. [Google Scholar] [CrossRef]
- Rich, J.N. Cancer stem cells in radiation resistance. Cancer Res. 2007, 67, 8980–8984. [Google Scholar] [CrossRef]
- Stupp, R.; Hegi, M.E. Targeting brain-tumor stem cells. Nat. Biotechnol. 2007, 25, 193–194. [Google Scholar] [CrossRef]
- Piccirillo, S.G.; Reynolds, B.A.; Zanetti, N.; Lamorte, G.; Binda, E.; Broggi, G.; Brem, H.; Olivi, A.; Dimeco, F.; Vescovi, A.L. Bone morphogenetic proteins inhibit the tumorigenic potential of human brain tumour-initiating cells. Nature 2006, 444, 761–765. [Google Scholar] [CrossRef] [PubMed]
- Sachdeva, R.; Wu, M.; Johnson, K.; Kim, H.; Celebre, A.; Shahzad, U.; Graham, M.S.; Kessler, J.A.; Chuang, J.H.; Karamchandani, J.; et al. BMP signaling mediates glioma stem cell quiescence and confers treatment resistance in glioblastoma. Sci. Rep. 2019, 9, 14569. [Google Scholar] [CrossRef] [PubMed]
- Hughes, J.H.; Ewy, J.M.; Chen, J.; Wong, S.Y.; Tharp, K.M.; Stahl, A.; Kumar, S. Transcriptomic analysis reveals that BMP4 sensitizes glioblastoma tumor-initiating cells to mechanical cues. Matrix Biol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Verhaak, R.G.; Hoadley, K.A.; Purdom, E.; Wang, V.; Qi, Y.; Wilkerson, M.D.; Miller, C.R.; Ding, L.; Golub, T.; Mesirov, J.P.; et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 2010, 17, 98–110. [Google Scholar] [CrossRef] [PubMed]
- Neftel, C.; Laffy, J.; Filbin, M.G.; Hara, T.; Shore, M.E.; Rahme, G.J.; Richman, A.R.; Silverbush, D.; Shaw, M.L.; Hebert, C.M.; et al. An Integrative Model of Cellular States, Plasticity, and Genetics for Glioblastoma. Cell 2019, 178, 835–849. [Google Scholar] [CrossRef] [PubMed]
- Gorovets, D.; Kannan, K.; Shen, R.; Kastenhuber, E.R.; Islamdoust, N.; Campos, C.; Pentsova, E.; Heguy, A.; Jhanwar, S.C.; Mellinghoff, I.K.; et al. IDH mutation and neuroglial developmental features define clinically distinct subclasses of lower grade diffuse astrocytic glioma. Clin. Cancer Res. 2012, 18, 2490–2501. [Google Scholar] [CrossRef]
- Phillips, H.S.; Kharbanda, S.; Chen, R.; Forrest, W.F.; Soriano, R.H.; Wu, T.D.; Misra, A.; Nigro, J.M.; Colman, H.; Soroceanu, L.; et al. Molecular subclasses of high-grade glioma predict prognosis, delineate a pattern of disease progression, and resemble stages in neurogenesis. Cancer Cell 2006, 9, 157–173. [Google Scholar] [CrossRef]
- Sturm, D.; Orr, B.A.; Toprak, U.H.; Hovestadt, V.; Jones, D.T.W.; Capper, D.; Sill, M.; Buchhalter, I.; Northcott, P.A.; Leis, I.; et al. New Brain Tumor Entities Emerge from Molecular Classification of CNS-PNETs. Cell 2016, 164, 1060–1072. [Google Scholar] [CrossRef]
- Louis, D.N.; Perry, A.; Reifenberger, G.; von Deimling, A.; Figarella-Branger, D.; Cavenee, W.K.; Ohgaki, H.; Wiestler, O.D.; Kleihues, P.; Ellison, D.W. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: A summary. Acta Neuropathol. 2016, 131, 803–820. [Google Scholar] [CrossRef]
- Jones, C.; Baker, S.J. Unique genetic and epigenetic mechanisms driving paediatric diffuse high-grade glioma. Nature reviews. Cancer 2014, 14. [Google Scholar] [CrossRef]
- Fukuda, T.; Scott, G.; Komatsu, Y.; Araya, R.; Kawano, M.; Ray, M.K.; Yamada, M.; Mishina, Y. Generation of a mouse with conditionally activated signaling through the BMP receptor, ALK2. Genesis 2006, 44, 159–167. [Google Scholar] [CrossRef] [PubMed]
- Caren, H.; Beck, S.; Pollard, S.M. Differentiation therapy for glioblastoma—Too many obstacles? Mol. Cell. Oncol. 2016, 3, e1124174. [Google Scholar] [CrossRef] [PubMed]
- Caren, H.; Stricker, S.H.; Bulstrode, H.; Gagrica, S.; Johnstone, E.; Bartlett, T.E.; Feber, A.; Wilson, G.; Teschendorff, A.E.; Bertone, P.; et al. Glioblastoma Stem Cells Respond to Differentiation Cues but Fail to Undergo Commitment and Terminal Cell-Cycle Arrest. Stem Cell Rep. 2015, 5, 829–842. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Liu, Y.; Zhang, J.; Yu, D.; Li, A.; Song, H.; Zhang, W.; Davis, D.; Gilbert, M.R.; Liu, F.; et al. Autocrine BMP4 Signaling Enhances Tumor Aggressiveness via Promoting Wnt/beta-Catenin Signaling in IDH1-mutant Gliomas. Transl. Oncol. 2019, 13, 125–134. [Google Scholar] [CrossRef] [PubMed]
- Olmsted, E.A.; Kaplan, F.S.; Shore, E.M. Bone morphogenetic protein-4 regulation in fibrodysplasia ossificans progressiva. Clin. Orthop. Relat. Rew. 2003, 408, 331–343. [Google Scholar] [CrossRef]
- Calpe, S.; Correia, A.C.; Sancho-Serra Mdel, C.; Krishnadath, K.K. Comparison of newly developed anti-bone morphogenetic protein 4 llama-derived antibodies with commercially available BMP4 inhibitors. Mabs 2016, 8, 678–688. [Google Scholar] [CrossRef]
- Glaser, D.L.; Economides, A.N.; Wang, L.; Liu, X.; Kimble, R.D.; Fandl, J.P.; Wilson, J.M.; Stahl, N.; Kaplan, F.S.; Shore, E.M. In vivo somatic cell gene transfer of an engineered Noggin mutein prevents BMP4-induced heterotopic ossification. J. Bone Jt. Surg. Am. 2003, 85, 2332–2342. [Google Scholar] [CrossRef]
- Duggal, R.; Geissinger, U.; Zhang, Q.; Aguilar, J.; Chen, N.G.; Binda, E.; Vescovi, A.L.; Szalay, A.A. Vaccinia virus expressing bone morphogenetic protein-4 in novel glioblastoma orthotopic models facilitates enhanced tumor regression and long-term survival. J. Transl. Med. 2013, 11, 155. [Google Scholar] [CrossRef]
- Choudhury, S.R.; Hudry, E.; Maguire, C.A.; Sena-Esteves, M.; Breakefield, X.O.; Grandi, P. Viral vectors for therapy of neurologic diseases. Neuropharmacology 2017, 120, 63–80. [Google Scholar] [CrossRef]
- Tyler, M.A.; Ulasov, I.V.; Sonabend, A.M.; Nandi, S.; Han, Y.; Marler, S.; Roth, J.; Lesniak, M.S. Neural stem cells target intracranial glioma to deliver an oncolytic adenovirus in vivo. Gene Ther. 2008, 16, 262–278. [Google Scholar] [CrossRef]
- Li, Q.; Wijesekera, O.; Salas, S.J.; Wang, J.Y.; Zhu, M.; Aprhys, C.; Chaichana, K.L.; Chesler, D.A.; Zhang, H.; Smith, C.L.; et al. Mesenchymal stem cells from human fat engineered to secrete BMP4 are nononcogenic, suppress brain cancer, and prolong survival. Clin. Cancer Res. 2014, 20, 2375–2387. [Google Scholar] [CrossRef] [PubMed]
- Spaeth, E.L.; Dembinski, J.L.; Sasser, A.K.; Watson, K.; Klopp, A.; Hall, B.; Andreeff, M.; Marini, F. Mesenchymal stem cell transition to tumor-associated fibroblasts contributes to fibrovascular network expansion and tumor progression. PLoS ONE 2009, 4, e4992. [Google Scholar] [CrossRef] [PubMed]
- Mangraviti, A.; Tzeng, S.Y.; Gullotti, D.; Kozielski, K.L.; Kim, J.E.; Seng, M.; Abbadi, S.; Schiapparelli, P.; Sarabia-Estrada, R.; Vescovi, A.; et al. Non-virally engineered human adipose mesenchymal stem cells produce BMP4, target brain tumors, and extend survival. Biomaterials 2016, 100, 53–66. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Yin, F.; Zhao, M.; Zhou, C.; Ren, J.; Huang, Q.; Zhao, Z.; Mitra, R.; Fan, W.; Fan, M. The homing and inhibiting effects of hNSCs-BMP4 on human glioma stem cells. Oncotarget 2016, 7, 17920–17931. [Google Scholar] [CrossRef]
- Srikanth, M.; Das, S.; Berns, E.J.; Kim, J.; Stupp, S.I.; Kessler, J.A. Nanofiber-mediated inhibition of focal adhesion kinase sensitizes glioma stemlike cells to epidermal growth factor receptor inhibition. Neuro-Oncology 2013, 15, 319–329. [Google Scholar] [CrossRef]
- Webber, M.J.; Kessler, J.A.; Stupp, S.I. Emerging peptide nanomedicine to regenerate tissues and organs. J. Intern. Med. 2010, 267, 71–88. [Google Scholar] [CrossRef]
- Lee, S.S.; Hsu, E.L.; Mendoza, M.; Ghodasra, J.; Nickoli, M.S.; Ashtekar, A.; Polavarapu, M.; Babu, J.; Riaz, R.M.; Nicolas, J.D.; et al. Gel scaffolds of BMP-2-binding peptide amphiphile nanofibers for spinal arthrodesis. Adv. Healthc. Mater. 2015, 4, 131–141. [Google Scholar] [CrossRef]
- Lee, J.; Son, M.J.; Woolard, K.; Donin, N.M.; Li, A.; Cheng, C.H.; Kotliarova, S.; Kotliarov, Y.; Walling, J.; Ahn, S.; et al. Epigenetic-Mediated Dysfunction of the Bone Morphogenetic Protein Pathway Inhibits Differentiation of Glioblastoma-Initiating Cells. Cancer Cell 2008, 13, 69–80. [Google Scholar] [CrossRef]
- Yan, K.; Wu, Q.; Yan, D.H.; Lee, C.H.; Rahim, N.; Tritschler, I.; DeVecchio, J.; Kalady, M.F.; Hjelmeland, A.B.; Rich, J.N. Glioma cancer stem cells secrete Gremlin1 to promote their maintenance within the tumor hierarchy. Genes Dev. 2014, 28, 1085–1100. [Google Scholar] [CrossRef]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nayak, S.; Mahenthiran, A.; Yang, Y.; McClendon, M.; Mania-Farnell, B.; James, C.D.; Kessler, J.A.; Tomita, T.; Cheng, S.-Y.; Stupp, S.I.; et al. Bone Morphogenetic Protein 4 Targeting Glioma Stem-Like Cells for Malignant Glioma Treatment: Latest Advances and Implications for Clinical Application. Cancers 2020, 12, 516. https://doi.org/10.3390/cancers12020516
Nayak S, Mahenthiran A, Yang Y, McClendon M, Mania-Farnell B, James CD, Kessler JA, Tomita T, Cheng S-Y, Stupp SI, et al. Bone Morphogenetic Protein 4 Targeting Glioma Stem-Like Cells for Malignant Glioma Treatment: Latest Advances and Implications for Clinical Application. Cancers. 2020; 12(2):516. https://doi.org/10.3390/cancers12020516
Chicago/Turabian StyleNayak, Sonali, Ashorne Mahenthiran, Yongyong Yang, Mark McClendon, Barbara Mania-Farnell, Charles David James, John A. Kessler, Tadanori Tomita, Shi-Yuan Cheng, Samuel I. Stupp, and et al. 2020. "Bone Morphogenetic Protein 4 Targeting Glioma Stem-Like Cells for Malignant Glioma Treatment: Latest Advances and Implications for Clinical Application" Cancers 12, no. 2: 516. https://doi.org/10.3390/cancers12020516
APA StyleNayak, S., Mahenthiran, A., Yang, Y., McClendon, M., Mania-Farnell, B., James, C. D., Kessler, J. A., Tomita, T., Cheng, S.-Y., Stupp, S. I., & Xi, G. (2020). Bone Morphogenetic Protein 4 Targeting Glioma Stem-Like Cells for Malignant Glioma Treatment: Latest Advances and Implications for Clinical Application. Cancers, 12(2), 516. https://doi.org/10.3390/cancers12020516






