A Novel Four-Gene Score to Predict Pathologically Complete (R0) Resection and Survival in Pancreatic Cancer
Abstract
:Simple Summary
Abstract
1. Introduction
2. Results
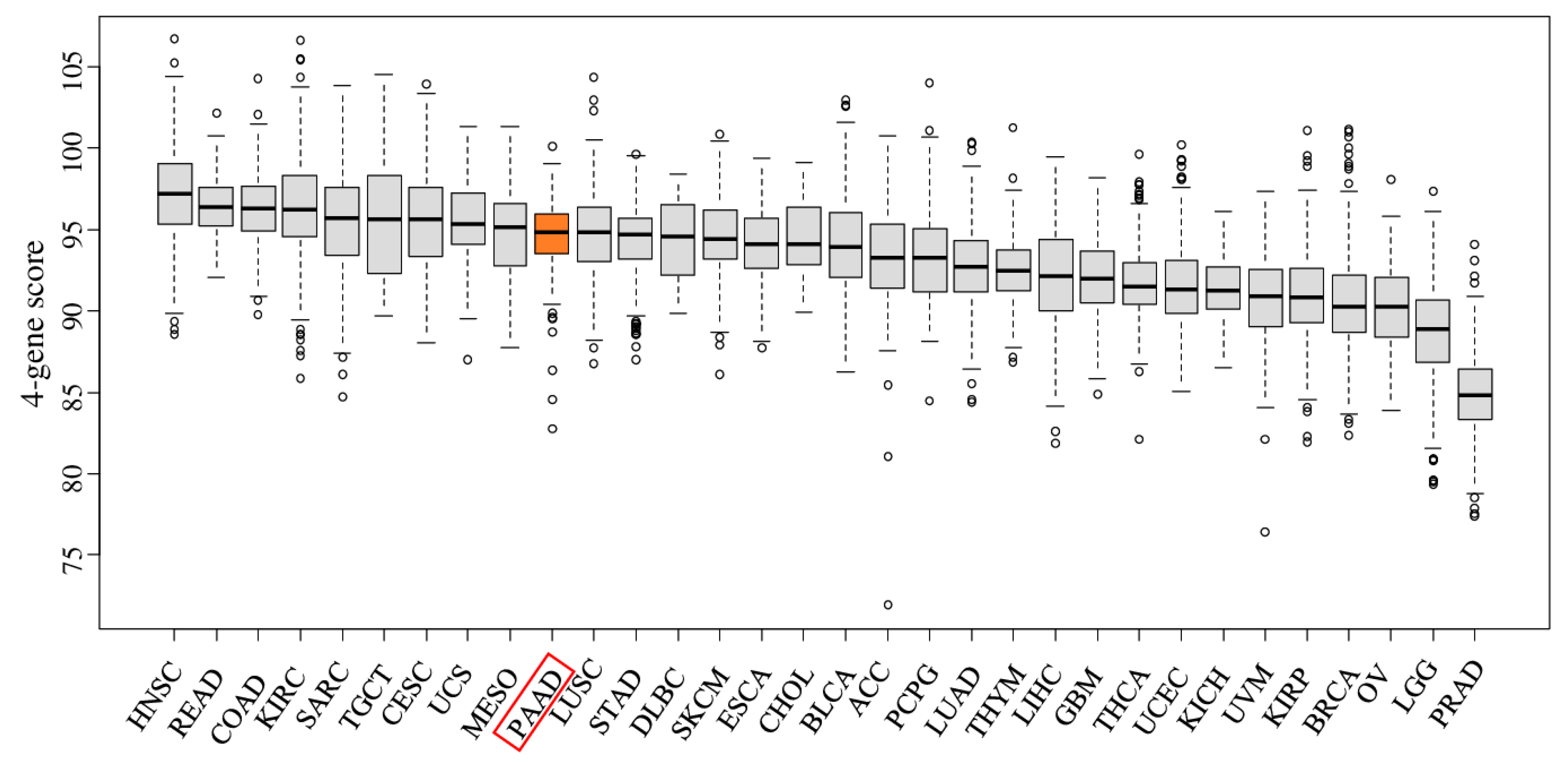
2.1. The Four-Gene Score Is Elevated in Pancreatic Cancer Tumors
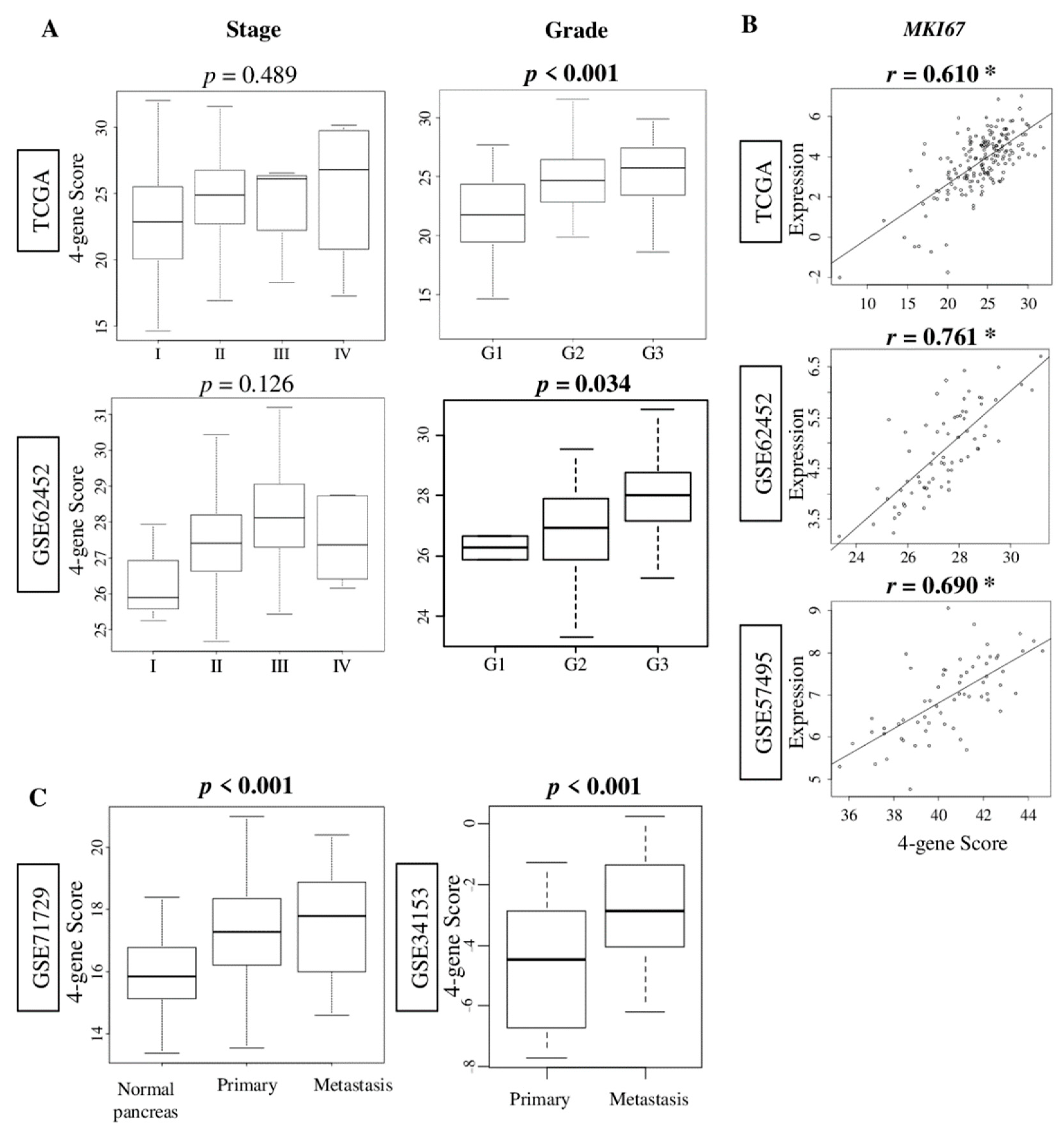
2.2. The Four-Gene Score Correlates with Pathological Grade and MKI67 Expression, and Was Elevated in Metastatic Pancreatic Cancer
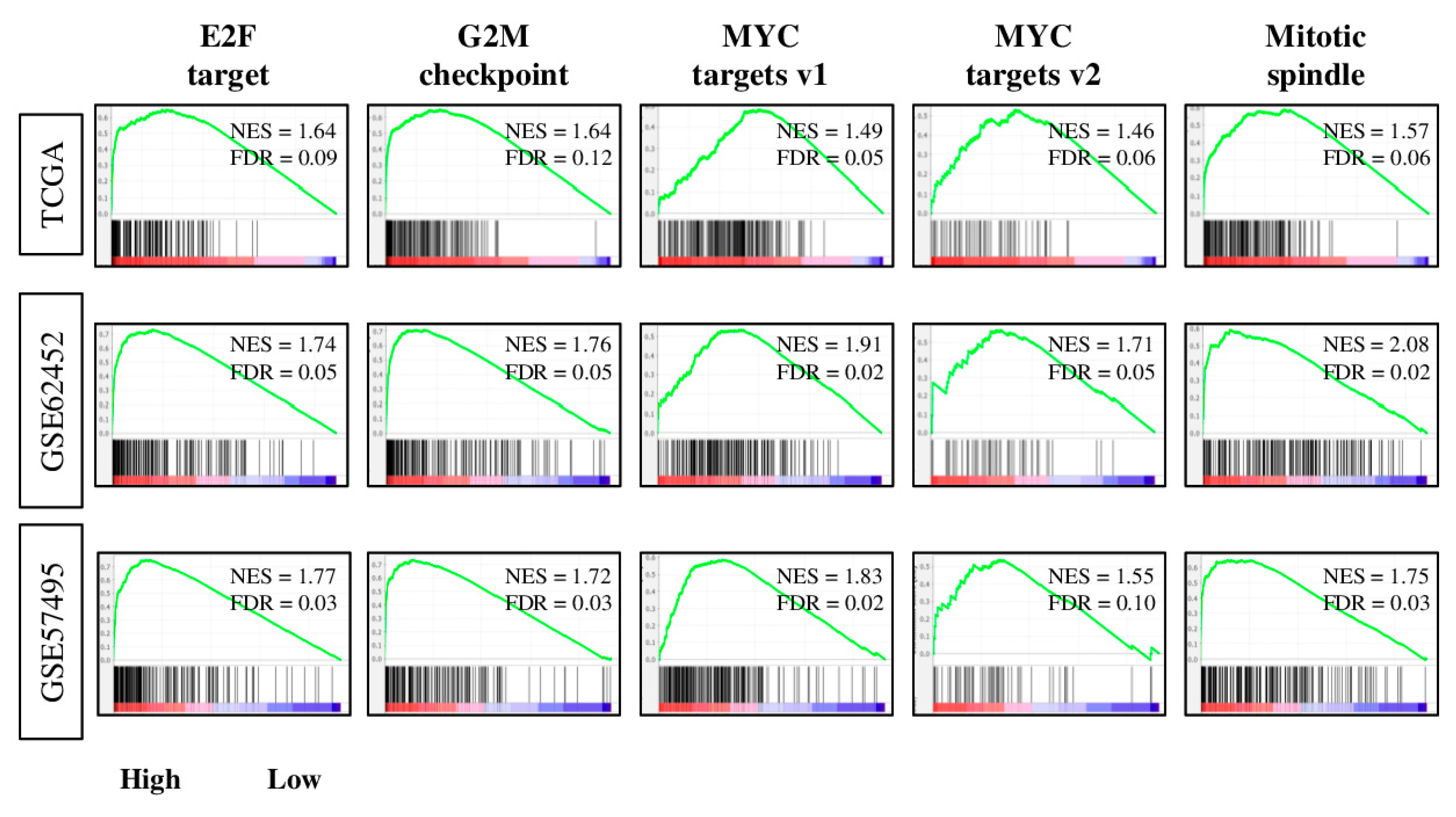
2.3. Tumors with High Four-Gene Score Have Enriched Expression of Cell Proiferation-Related and Pro-Cancerous Gene Sets
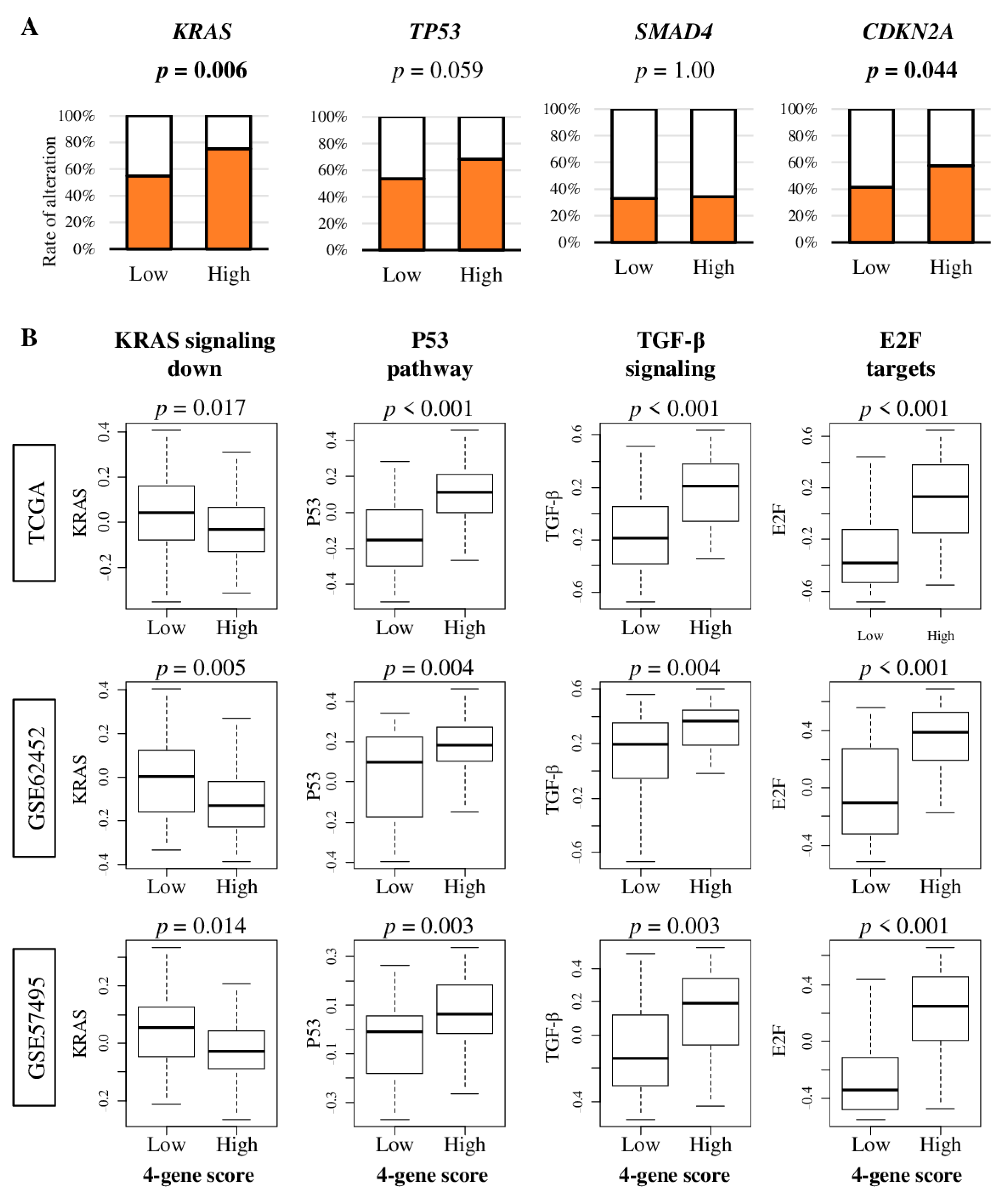
2.4. Pancreatic Cancer with High Four-Gene Sore Have Higher KRAS and CDKN2A Gene Alterations, and Enhanced Signaling of KRAS, p53, TGF-β, and E2F Target Pathways
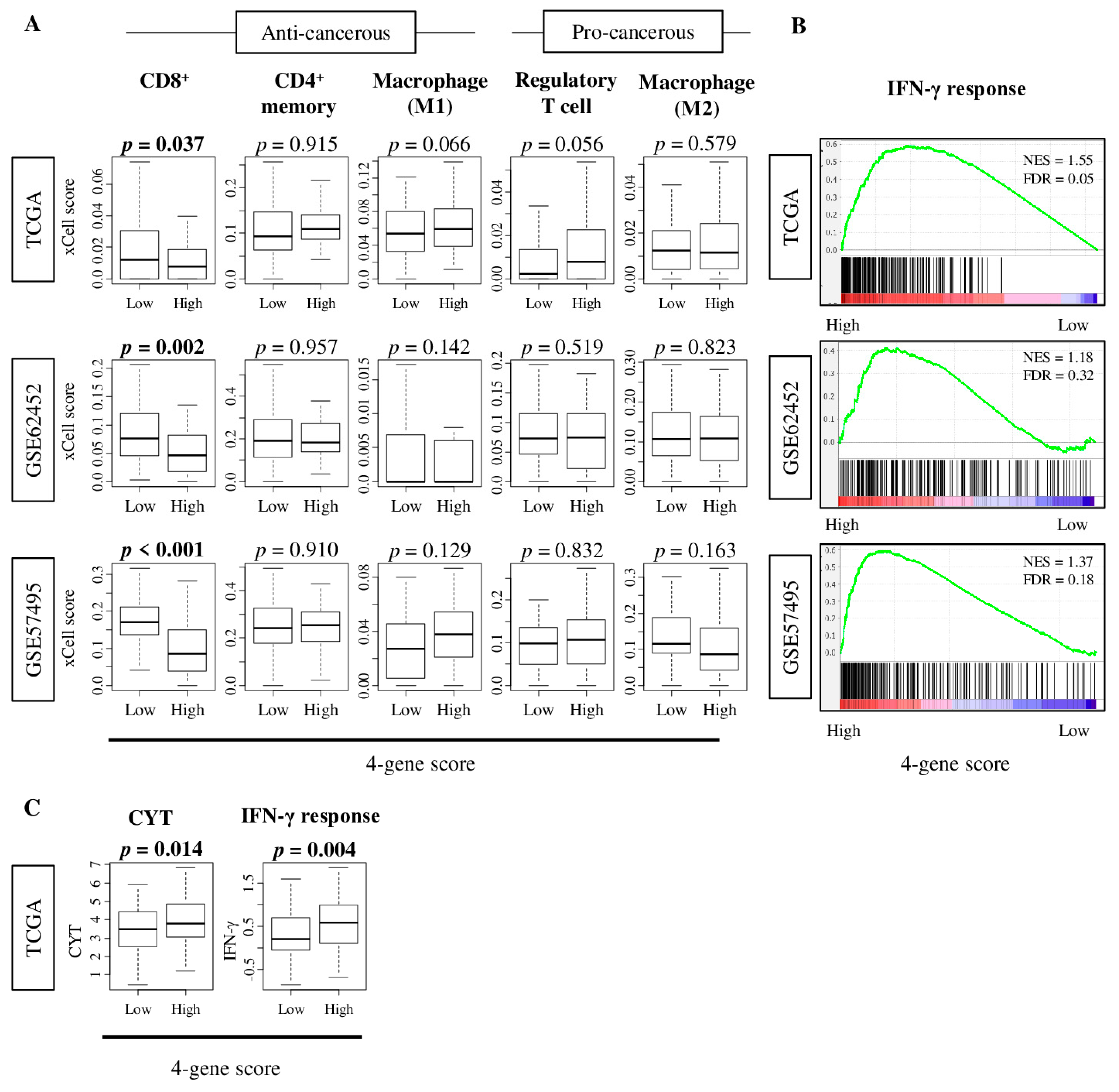
2.5. Tumors with High Four-Gene Score Have High Levels of Interferon -Γ Response and Cytolytic Activity
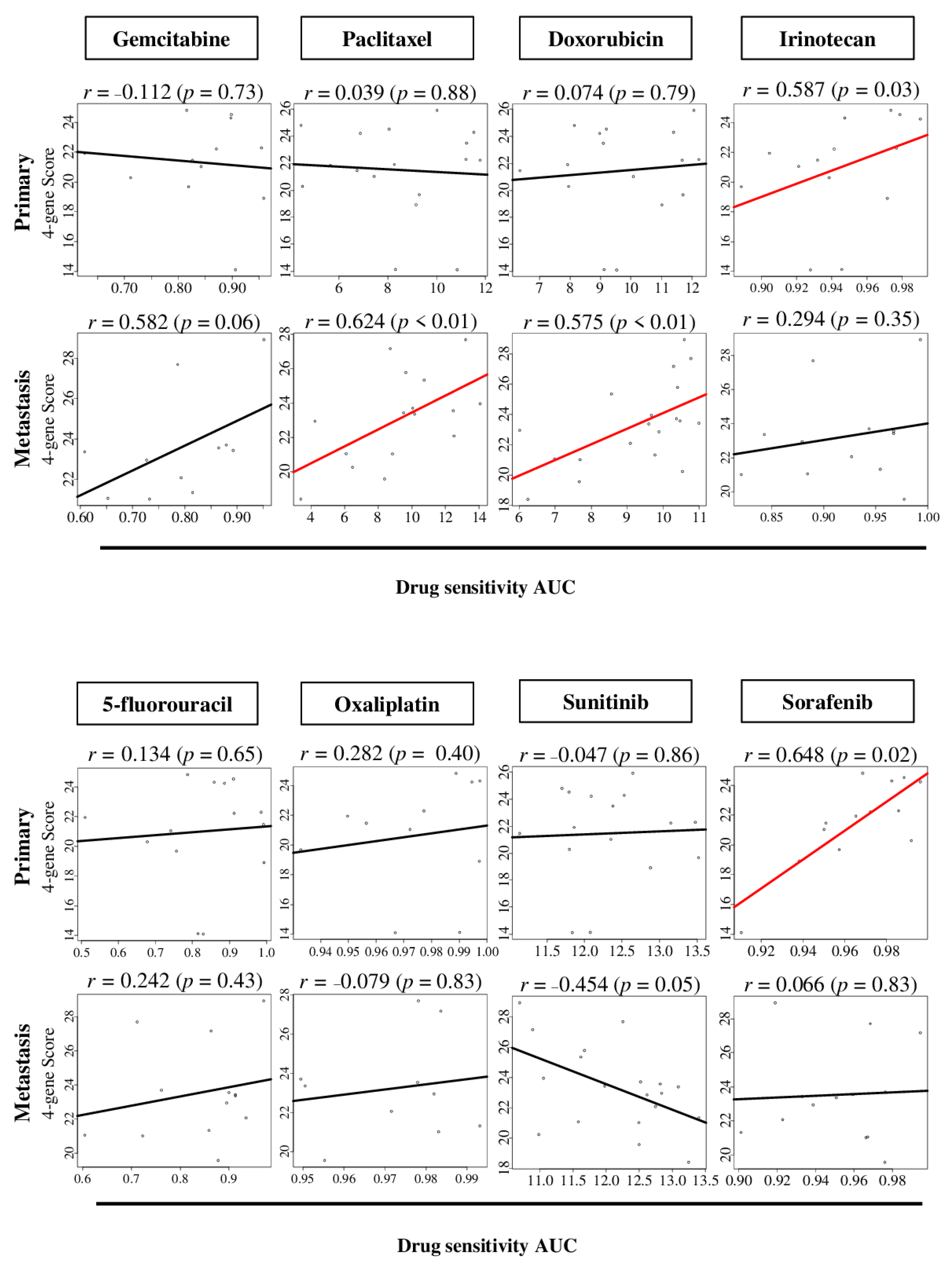
2.6. The Four-Gene Score Level Associates with Drug Sensitivity to Sorafenib in Primary Pancreatic Cancer, and Paclitaxel and Doxorubicin in Metastatic Pancreatic Cancer
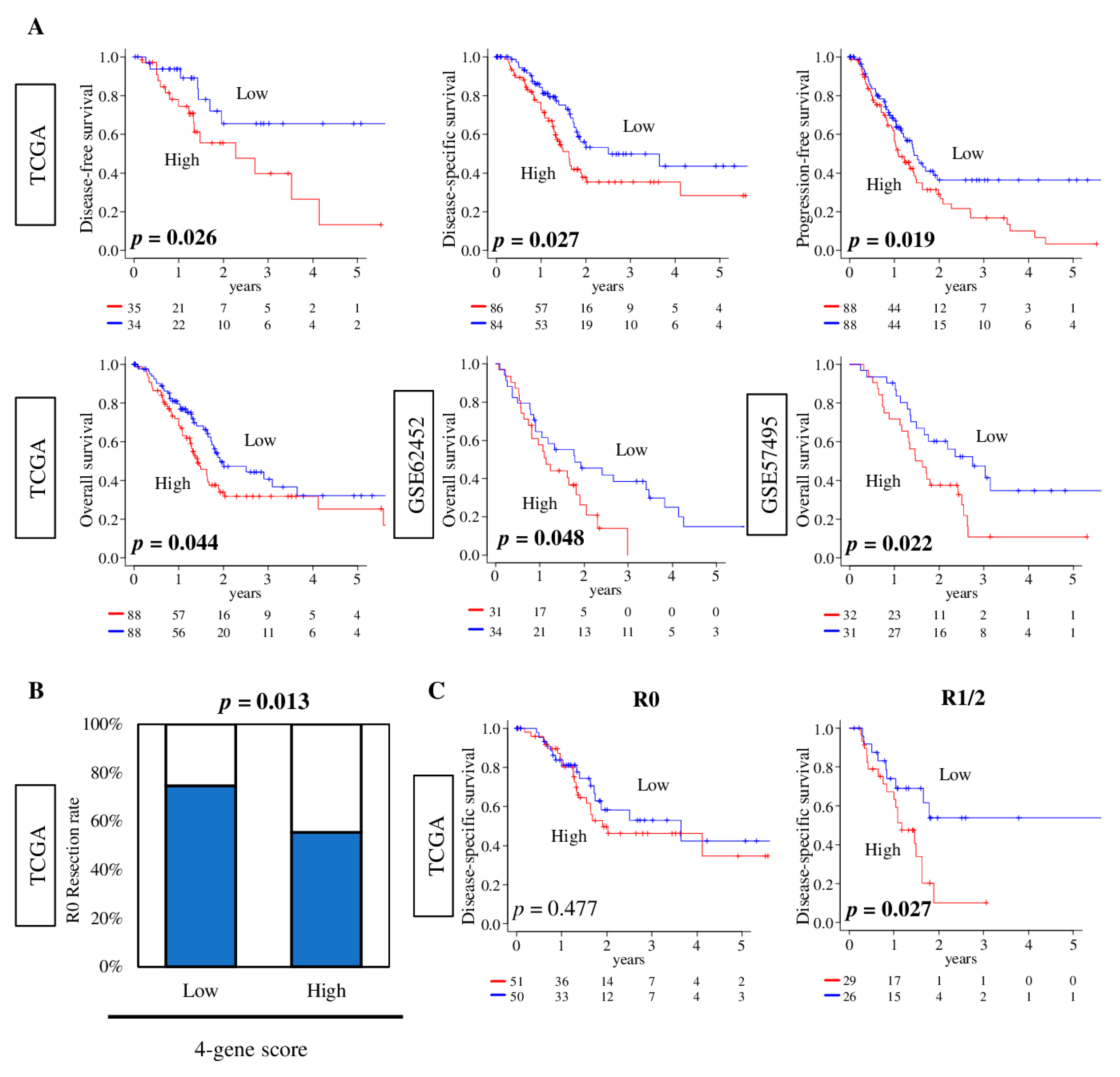
2.7. Pancreatic Cancer with a High Four-Gene Score Was Significantly Associated with Worse Survival and Pathologically Margin-Positive Resection
3. Discussion
4. Materials and Methods
4.1. Pancreatic Cancer Cohorts and Their Data
4.2. Gene Set Expression Analyses
4.3. Cell Composition Fraction Estimation
4.4. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Rawla, P.; Sunkara, T.; Gaduputi, V. Epidemiology of Pancreatic Cancer: Global Trends, Etiology and Risk Factors. World J. Oncol. 2019, 10, 10–27. [Google Scholar] [CrossRef] [PubMed]
- Collisson, E.A.; Bailey, P.; Chang, D.K.; Biankin, A.V. Molecular subtypes of pancreatic cancer. Nat. Rev. Gastroenterol. Hepatol. 2019, 16, 207–220. [Google Scholar] [CrossRef] [PubMed]
- Katz, M.H.; Pisters, P.W.; Evans, D.B.; Sun, C.C.; Lee, J.E.; Fleming, J.B.; Vauthey, J.N.; Abdalla, E.K.; Crane, C.H.; Wolff, R.A.; et al. Borderline resectable pancreatic cancer: The importance of this emerging stage of disease. J. Am. Coll. Surg. 2008, 206, 833–846; discussion 846–838. [Google Scholar] [CrossRef] [Green Version]
- Rau, B.M.; Moritz, K.; Schuschan, S.; Alsfasser, G.; Prall, F.; Klar, E. R1 resection in pancreatic cancer has significant impact on long-term outcome in standardized pathology modified for routine use. Surgery 2012, 152, S103–S111. [Google Scholar] [CrossRef] [PubMed]
- Katsuta, E.; Rashid, O.M.; Takabe, K. Fibroblasts as a Biological Marker for Curative Resection in Pancreatic Ductal Adenocarcinoma. Int. J. Mol. Sci. 2020, 21, 3890. [Google Scholar] [CrossRef]
- Demir, I.E.; Jäger, C.; Schlitter, A.M.; Konukiewitz, B.; Stecher, L.; Schorn, S.; Tieftrunk, E.; Scheufele, F.; Calavrezos, L.; Schirren, R.; et al. R0 Versus R1 Resection Matters after Pancreaticoduodenectomy, and Less after Distal or Total Pancreatectomy for Pancreatic Cancer. Ann. Surg. 2018, 268, 1058–1068. [Google Scholar] [CrossRef]
- Kelly, K.N.; Macedo, F.I.; Merchant, N.B. Neoadjuvant Therapy: When Should It Be Used for Pancreatic Cancer? Adv. Surg. 2020, 54, 49–68. [Google Scholar] [CrossRef]
- Hemmings, C.; Connor, S. Pathological assessment of tumour regression following neoadjuvant therapy in pancreatic carcinoma. Pathology 2020. [Google Scholar] [CrossRef]
- Rahib, L.; Smith, B.D.; Aizenberg, R.; Rosenzweig, A.B.; Fleshman, J.M.; Matrisian, L.M. Projecting cancer incidence and deaths to 2030: The unexpected burden of thyroid, liver, and pancreas cancers in the United States. Cancer Res. 2014, 74, 2913–2921. [Google Scholar] [CrossRef] [Green Version]
- Oshi, M.; Katsuta, E.; Yan, L.; Ebos, J.M.L.; Rashid, O.M.; Matsuyama, R.; Endo, I.; Takabe, K. A Novel 4-Gene Score to Predict Survival, Distant Metastasis and Response to Neoadjuvant Therapy in Breast Cancer. Cancers 2020, 12, 1148. [Google Scholar] [CrossRef]
- Wee, P.; Wang, Z. Epidermal Growth Factor Receptor Cell Proliferation Signaling Pathways. Cancers 2017, 9, 52. [Google Scholar] [CrossRef] [Green Version]
- Liu, J.; Lichtenberg, T.; Hoadley, K.A.; Poisson, L.M.; Lazar, A.J.; Cherniack, A.D.; Kovatich, A.J.; Benz, C.C.; Levine, D.A.; Lee, A.V.; et al. An Integrated TCGA Pan-Cancer Clinical Data Resource to Drive High-Quality Survival Outcome Analytics. Cell 2018, 173, 400–416.e411. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, S.; He, P.; Wang, J.; Schetter, A.; Tang, W.; Funamizu, N.; Yanaga, K.; Uwagawa, T.; Satoskar, A.R.; Gaedcke, J.; et al. A Novel MIF Signaling Pathway Drives the Malignant Character of Pancreatic Cancer by Targeting NR3C2. Cancer Res. 2016, 76, 3838–3850. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, D.T.; Davis-Yadley, A.H.; Huang, P.Y.; Husain, K.; Centeno, B.A.; Permuth-Wey, J.; Pimiento, J.M.; Malafa, M. Prognostic Fifteen-Gene Signature for Early Stage Pancreatic Ductal Adenocarcinoma. PLoS ONE 2015, 10, e0133562. [Google Scholar] [CrossRef] [PubMed]
- Moffitt, R.A.; Marayati, R.; Flate, E.L.; Volmar, K.E.; Loeza, S.G.; Hoadley, K.A.; Rashid, N.U.; Williams, L.A.; Eaton, S.C.; Chung, A.H.; et al. Virtual microdissection identifies distinct tumor- and stroma-specific subtypes of pancreatic ductal adenocarcinoma. Nat. Genet. 2015, 47, 1168–1178. [Google Scholar] [CrossRef]
- Chaika, N.V.; Yu, F.; Purohit, V.; Mehla, K.; Lazenby, A.J.; DiMaio, D.; Anderson, J.M.; Yeh, J.J.; Johnson, K.R.; Hollingsworth, M.A.; et al. Differential expression of metabolic genes in tumor and stromal components of primary and metastatic loci in pancreatic adenocarcinoma. PLoS ONE 2012, 7, e32996. [Google Scholar] [CrossRef] [Green Version]
- Waddell, N.; Pajic, M.; Patch, A.M.; Chang, D.K.; Kassahn, K.S.; Bailey, P.; Johns, A.L.; Miller, D.; Nones, K.; Quek, K.; et al. Whole genomes redefine the mutational landscape of pancreatic cancer. Nature 2015, 518, 495–501. [Google Scholar] [CrossRef] [Green Version]
- Oshi, M.; Takahashi, H.; Tokumaru, Y.; Yan, L.; Rashid, O.M.; Nagahashi, M.; Matsuyama, R.; Endo, I.; Takabe, K. The E2F Pathway Score as a Predictive Biomarker of Response to Neoadjuvant Therapy in ER+/HER2- Breast Cancer. Cells 2020, 9, 1643. [Google Scholar] [CrossRef]
- Tokumaru, Y.; Oshi, M.; Katsuta, E.; Yan, L.; Satyananda, V.; Matsuhashi, N.; Futamura, M.; Akao, Y.; Yoshida, K.; Takabe, K. KRAS signaling enriched triple negative breast cancer is associated with favorable tumor immune microenvironment and better survival. Am. J. Cancer Res. 2020, 10, 897–907. [Google Scholar]
- Oshi, M.; Takahashi, H.; Tokumaru, Y.; Yan, L.; Rashid, O.M.; Matsuyama, R.; Endo, I.; Takabe, K. G2M Cell Cycle Pathway Score as a Prognostic Biomarker of Metastasis in Estrogen Receptor (ER)-Positive Breast Cancer. Int. J. Mol. Sci. 2020, 21, 2921. [Google Scholar] [CrossRef] [Green Version]
- Oshi, M.; Newman, S.; Tokumaru, Y.; Yan, L.; Matsuyama, R.; Endo, I.; Katz, M.H.G.; Takabe, K. High G2M Pathway Score Pancreatic Cancer is Associated with Worse Survival, Particularly after Margin-Positive (R1 or R2) Resection. Cancers 2020, 12, 2871. [Google Scholar] [CrossRef] [PubMed]
- Oshi, M.; Newman, S.; Tokumaru, Y.; Yan, L.; Matsuyama, R.; Endo, I.; Nagahashi, M.; Takabe, K. Intra-Tumoral Angiogenesis Is Associated with Inflammation, Immune Reaction and Metastatic Recurrence in Breast Cancer. Int. J. Mol. Sci. 2020, 21, 6708. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, H.; Asaoka, M.; Yan, L.; Rashid, O.M.; Oshi, M.; Ishikawa, T.; Nagahashi, M.; Takabe, K. Biologically Aggressive Phenotype and Anti-cancer Immunity Counterbalance in Breast Cancer with High Mutation Rate. Sci. Rep. 2020, 10, 1852. [Google Scholar] [CrossRef] [Green Version]
- Aran, D.; Hu, Z.; Butte, A.J. xCell: Digitally portraying the tissue cellular heterogeneity landscape. Genome Biol. 2017, 18, 220. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shi, W.; Jiang, T.; Nuciforo, P.; Hatzis, C.; Holmes, E.; Harbeck, N.; Sotiriou, C.; Peña, L.; Loi, S.; Rosa, D.D.; et al. Pathway level alterations rather than mutations in single genes predict response to HER2-targeted therapies in the neo-ALTTO trial. Ann. Oncol. Off. J. Eur. Soc. Med. Oncol. 2019, 30, 1018. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Sun, Z.; Zimmermann, M.T.; Bugrim, A.; Kocher, J.P. Predict drug sensitivity of cancer cells with pathway activity inference. BMC Med. Genom. 2019, 12, 15. [Google Scholar] [CrossRef] [Green Version]
- Khatri, P.; Sirota, M.; Butte, A.J. Ten years of pathway analysis: Current approaches and outstanding challenges. PLoS Comput. Biol. 2012, 8, e1002375. [Google Scholar] [CrossRef]
- Su, J.; Yoon, B.J.; Dougherty, E.R. Accurate and reliable cancer classification based on probabilistic inference of pathway activity. PLoS ONE 2009, 4, e8161. [Google Scholar] [CrossRef]
- Lee, E.; Chuang, H.Y.; Kim, J.W.; Ideker, T.; Lee, D. Inferring pathway activity toward precise disease classification. PLoS Comput. Biol. 2008, 4, e1000217. [Google Scholar] [CrossRef]
- Chang, Y.T.; Chang, M.C.; Wei, S.C.; Tien, Y.W.; Hsu, C.; Liang, P.C.; Tsao, P.N.; Jan, I.S.; Wong, J.M. Serum vascular endothelial growth factor/soluble vascular endothelial growth factor receptor 1 ratio is an independent prognostic marker in pancreatic cancer. Pancreas 2008, 37, 145–150. [Google Scholar] [CrossRef] [Green Version]
- Zhang, C.; Sun, J.B.; Liu, D.C.; Cui, Y.Q.; Liu, S.; Sun, H.C. Preliminary research on the pathological role of cathepsin-B in subcutaneous heteroplastic pancreatic carcinoma in nude mice. Chin. Med. J. 2009, 122, 2489–2496. [Google Scholar] [PubMed]
- Carmeliet, P.; Moons, L.; Luttun, A.; Vincenti, V.; Compernolle, V.; De Mol, M.; Wu, Y.; Bono, F.; Devy, L.; Beck, H.; et al. Synergism between vascular endothelial growth factor and placental growth factor contributes to angiogenesis and plasma extravasation in pathological conditions. Nat. Med. 2001, 7, 575–583. [Google Scholar] [CrossRef] [PubMed]
- Fischer, C.; Mazzone, M.; Jonckx, B.; Carmeliet, P. FLT1 and its ligands VEGFB and PlGF: Drug targets for anti-angiogenic therapy? Nat. Rev. Cancer 2008, 8, 942–956. [Google Scholar] [CrossRef]
- Sabbaghian, M.S.; Rothberger, G.; Alongi, A.P.; Gagner, J.P.; Goldberg, J.D.; Rolnitzky, L.; Chiriboga, L.; Hajdu, C.H.; Zagzag, D.; Basch, R.; et al. Levels of elevated circulating endothelial cell decline after tumor resection in patients with pancreatic ductal adenocarcinoma. Anticancer Res. 2010, 30, 2911–2917. [Google Scholar] [PubMed]
- Asano, E.; Hasegawa, H.; Hyodo, T.; Ito, S.; Maeda, M.; Chen, D.; Takahashi, M.; Hamaguchi, M.; Senga, T. SHCBP1 is required for midbody organization and cytokinesis completion. Cell Cycle 2014, 13, 2744–2751. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Asano, E.; Hasegawa, H.; Hyodo, T.; Ito, S.; Maeda, M.; Takahashi, M.; Hamaguchi, M.; Senga, T. The Aurora-B-mediated phosphorylation of SHCBP1 regulates cytokinetic furrow ingression. J. Cell Sci. 2013, 126, 3263–3270. [Google Scholar] [CrossRef] [Green Version]
- Peng, C.; Zhao, H.; Chen, W.; Song, Y.; Wang, X.; Li, J.; Qiao, Y.; Wu, D.; Ma, S.; Wang, X.; et al. Identification of SHCBP1 as a novel downstream target gene of SS18-SSX1 and its functional analysis in progression of synovial sarcoma. Oncotarget 2016, 7, 66822–66834. [Google Scholar] [CrossRef] [Green Version]
- Itoh, S.; Lemay, S.; Osawa, M.; Che, W.; Duan, Y.; Tompkins, A.; Brookes, P.S.; Sheu, S.S.; Abe, J. Mitochondrial Dok-4 recruits Src kinase and regulates NF-kappaB activation in endothelial cells. J. Biol. Chem. 2005, 280, 26383–26396. [Google Scholar] [CrossRef] [Green Version]
- Bedirian, A.; Baldwin, C.; Abe, J.; Takano, T.; Lemay, S. Pleckstrin homology and phosphotyrosine-binding domain-dependent membrane association and tyrosine phosphorylation of Dok-4, an inhibitory adapter molecule expressed in epithelial cells. J. Biol. Chem. 2004, 279, 19335–19349. [Google Scholar] [CrossRef] [Green Version]
- Gray, S.G.; Al-Sarraf, N.; Baird, A.M.; Gately, K.; McGovern, E.; O’Byrne, K.J. Transcriptional regulation of IRS5/DOK4 expression in non-small-cell lung cancer cells. Clin. Lung Cancer 2008, 9, 367–374. [Google Scholar] [CrossRef]
- Zhang, L.; Li, R.; Hu, K.; Dai, Y.; Pang, Y.; Jiao, Y.; Liu, Y.; Cui, L.; Shi, J.; Cheng, Z.; et al. Prognostic role of DOK family adapters in acute myeloid leukemia. Cancer Gene Ther. 2019, 26, 305–312. [Google Scholar] [CrossRef] [PubMed]
- Heidarizadi, A.; Salimi, M.; Mozdarani, H. Study of DOK4 gene expression and promoter methylation in sporadic breast cancer. Neoplasma 2020, 67, 916–921. [Google Scholar] [CrossRef] [PubMed]
- Hannafon, B.N.; Sebastiani, P.; de las Morenas, A.; Lu, J.; Rosenberg, C.L. Expression of microRNA and their gene targets are dysregulated in preinvasive breast cancer. Breast Cancer Res. BCR 2011, 13, R24. [Google Scholar] [CrossRef] [Green Version]
- Babbitt, S.E.; Sutherland, M.C.; San Francisco, B.; Mendez, D.L.; Kranz, R.G. Mitochondrial cytochrome c biogenesis: No longer an enigma. Trends Biochem. Sci. 2015, 40, 446–455. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van Rahden, V.A.; Rau, I.; Fuchs, S.; Kosyna, F.K.; de Almeida, H.L., Jr.; Fryssira, H.; Isidor, B.; Jauch, A.; Joubert, M.; Lachmeijer, A.M.; et al. Clinical spectrum of females with HCCS mutation: From no clinical signs to a neonatal lethal form of the microphthalmia with linear skin defects (MLS) syndrome. Orphanet J. Rare Dis. 2014, 9, 53. [Google Scholar] [CrossRef] [Green Version]
- Wimplinger, I.; Shaw, G.M.; Kutsche, K. HCCS loss-of-function missense mutation in a female with bilateral microphthalmia and sclerocornea: A novel gene for severe ocular malformations? Mol. Vis. 2007, 13, 1475–1482. [Google Scholar]
- Lee, J.S.; Ruppin, E. Multiomics Prediction of Response Rates to Therapies to Inhibit Programmed Cell Death 1 and Programmed Cell Death 1 Ligand 1. JAMA Oncol. 2019, 5, 1614–1618. [Google Scholar] [CrossRef]
- Vonderheide, R.H.; Bayne, L.J. Inflammatory networks and immune surveillance of pancreatic carcinoma. Curr. Opin. Immunol. 2013, 25, 200–205. [Google Scholar] [CrossRef] [Green Version]
- Farhood, B.; Najafi, M.; Mortezaee, K. CD8(+) cytotoxic T lymphocytes in cancer immunotherapy: A review. J. Cell. Physiol. 2019, 234, 8509–8521. [Google Scholar] [CrossRef]
- Orhan, A.; Vogelsang, R.P.; Andersen, M.B.; Madsen, M.T.; Hölmich, E.R.; Raskov, H.; Gögenur, I. The prognostic value of tumour-infiltrating lymphocytes in pancreatic cancer: A systematic review and meta-analysis. Eur. J. Cancer 2020, 132, 71–84. [Google Scholar] [CrossRef]
- Zhang, J.; Wang, Y.F.; Wu, B.; Zhong, Z.X.; Wang, K.X.; Yang, L.Q.; Wang, Y.Q.; Li, Y.Q.; Gao, J.; Li, Z.S. Intraepithelial Attack Rather than Intratumorally Infiltration of CD8+T Lymphocytes is a Favorable Prognostic Indicator in Pancreatic Ductal Adenocarcinoma. Curr. Mol. Med. 2017, 17, 689–698. [Google Scholar] [CrossRef] [PubMed]
- Balachandran, V.P.; Łuksza, M.; Zhao, J.N.; Makarov, V.; Moral, J.A.; Remark, R.; Herbst, B.; Askan, G.; Bhanot, U.; Senbabaoglu, Y.; et al. Identification of unique neoantigen qualities in long-term survivors of pancreatic cancer. Nature 2017, 551, 512–516. [Google Scholar] [CrossRef] [PubMed]
- Carstens, J.L.; Correa de Sampaio, P.; Yang, D.; Barua, S.; Wang, H.; Rao, A.; Allison, J.P.; LeBleu, V.S.; Kalluri, R. Spatial computation of intratumoral T cells correlates with survival of patients with pancreatic cancer. Nat. Commun. 2017, 8, 15095. [Google Scholar] [CrossRef] [PubMed]
- Alberts, S.R.; Schroeder, M.; Erlichman, C.; Steen, P.D.; Foster, N.R.; Moore, D.F., Jr.; Rowland, K.M., Jr.; Nair, S.; Tschetter, L.K.; Fitch, T.R. Gemcitabine and ISIS-2503 for patients with locally advanced or metastatic pancreatic adenocarcinoma: A North Central Cancer Treatment Group phase II trial. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2004, 22, 4944–4950. [Google Scholar] [CrossRef]
- Usón Junior, P.L.S.; Rother, E.T.; Maluf, F.C.; Bugano, D.D.G. Meta-analysis of Modified FOLFIRINOX Regimens for Patients with Metastatic Pancreatic Cancer. Clin. Colorectal Cancer 2018, 17, 187–197. [Google Scholar] [CrossRef]
- Ottaiano, A.; Capozzi, M.; De Divitiis, C.; De Stefano, A.; Botti, G.; Avallone, A.; Tafuto, S. Gemcitabine mono-therapy versus gemcitabine plus targeted therapy in advanced pancreatic cancer: A meta-analysis of randomized phase III trials. Acta Oncol. Stockh. Swed. 2017, 56, 377–383. [Google Scholar] [CrossRef] [Green Version]
- Archibugi, L.; Testoni, S.G.G.; Redegalli, M.; Petrone, M.C.; Reni, M.; Falconi, M.; Doglioni, C.; Capurso, G.; Arcidiacono, P.G. New era for pancreatic endoscopic ultrasound: From imaging to molecular pathology of pancreatic cancer. World J. Gastrointest. Oncol. 2019, 11, 933–945. [Google Scholar] [CrossRef]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef] [Green Version]
- Gao, J.; Aksoy, B.A.; Dogrusoz, U.; Dresdner, G.; Gross, B.; Sumer, S.O.; Sun, Y.; Jacobsen, A.; Sinha, R.; Larsson, E.; et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 2013, 6, pl1. [Google Scholar] [CrossRef] [Green Version]
- Liberzon, A.; Birger, C.; Thorvaldsdóttir, H.; Ghandi, M.; Mesirov, J.P.; Tamayo, P. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 2015, 1, 417–425. [Google Scholar] [CrossRef] [Green Version]
- Oshi, M.; Tokumaru, Y.; Asaoka, M.; Yan, L.; Satyananda, V.; Matsuyama, R.; Matsuhashi, N.; Futamura, M.; Ishikawa, T.; Yoshida, K.; et al. M1 Macrophage and M1/M2 ratio defined by transcriptomic signatures resemble only part of their conventional clinical characteristics in breast cancer. Sci. Rep. 2020, 10, 16554. [Google Scholar] [CrossRef] [PubMed]
- Oshi, M.; Asaoka, M.; Tokumaru, Y.; Yan, L.; Matsuyama, R.; Ishikawa, T.; Endo, I.; Takabe, K. CD8 T Cell Score as a Prognostic Biomarker for Triple Negative Breast Cancer. Int. J. Mol. Sci. 2020, 21, 6968. [Google Scholar] [CrossRef] [PubMed]
- Tokumaru, Y.; Oshi, M.; Katsuta, E.; Yan, L.; Huang, J.L.; Nagahashi, M.; Matsuhashi, N.; Futamura, M.; Yoshida, K.; Takabe, K. Intratumoral Adipocyte-High Breast Cancer Enrich for Metastatic and Inflammation-Related Pathways but Associated with Less Cancer Cell Proliferation. Int. J. Mol. Sci. 2020, 21, 5744. [Google Scholar] [CrossRef] [PubMed]
- Oshi, M.; Newman, S.; Murthy, V.; Tokumaru, Y.; Yan, L.; Matsuyama, R.; Endo, I.; Takabe, K. ITPKC as a Prognostic and Predictive Biomarker of Neoadjuvant Chemotherapy for Triple Negative Breast Cancer. Cancers 2020, 12, 2758. [Google Scholar] [CrossRef]
- Rooney, M.S.; Shukla, S.A.; Wu, C.J.; Getz, G.; Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 2015, 160, 48–61. [Google Scholar] [CrossRef] [Green Version]
- Thorsson, V.; Gibbs, D.L.; Brown, S.D.; Wolf, D.; Bortone, D.S.; Ou Yang, T.H.; Porta-Pardo, E.; Gao, G.F.; Plaisier, C.L.; Eddy, J.A.; et al. The Immune Landscape of Cancer. Immunity 2019, 51, 411–412. [Google Scholar] [CrossRef] [Green Version]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oshi, M.; Tokumaru, Y.; Patel, A.; Yan, L.; Matsuyama, R.; Endo, I.; Katz, M.H.G.; Takabe, K. A Novel Four-Gene Score to Predict Pathologically Complete (R0) Resection and Survival in Pancreatic Cancer. Cancers 2020, 12, 3635. https://doi.org/10.3390/cancers12123635
Oshi M, Tokumaru Y, Patel A, Yan L, Matsuyama R, Endo I, Katz MHG, Takabe K. A Novel Four-Gene Score to Predict Pathologically Complete (R0) Resection and Survival in Pancreatic Cancer. Cancers. 2020; 12(12):3635. https://doi.org/10.3390/cancers12123635
Chicago/Turabian StyleOshi, Masanori, Yoshihisa Tokumaru, Ankit Patel, Li Yan, Ryusei Matsuyama, Itaru Endo, Matthew H.G. Katz, and Kazuaki Takabe. 2020. "A Novel Four-Gene Score to Predict Pathologically Complete (R0) Resection and Survival in Pancreatic Cancer" Cancers 12, no. 12: 3635. https://doi.org/10.3390/cancers12123635





