Patient-Derived Tumor Organoids for Drug Repositioning in Cancer Care: A Promising Approach in the Era of Tailored Treatment
Abstract
Simple Summary
Abstract
1. Introduction
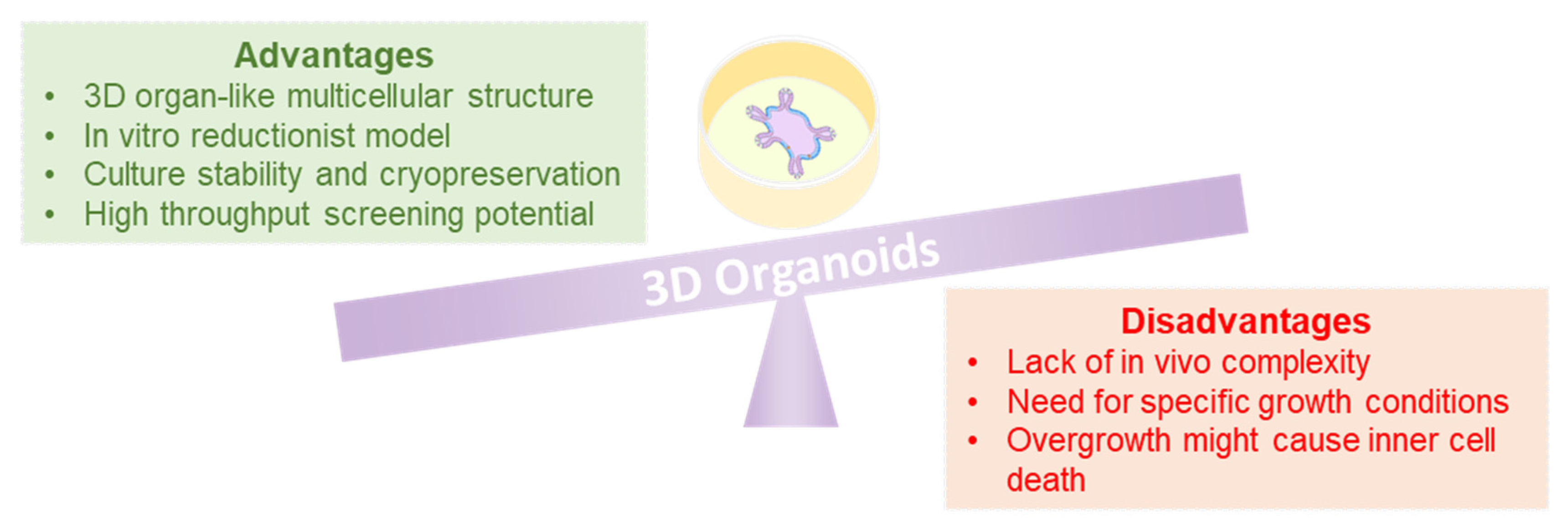
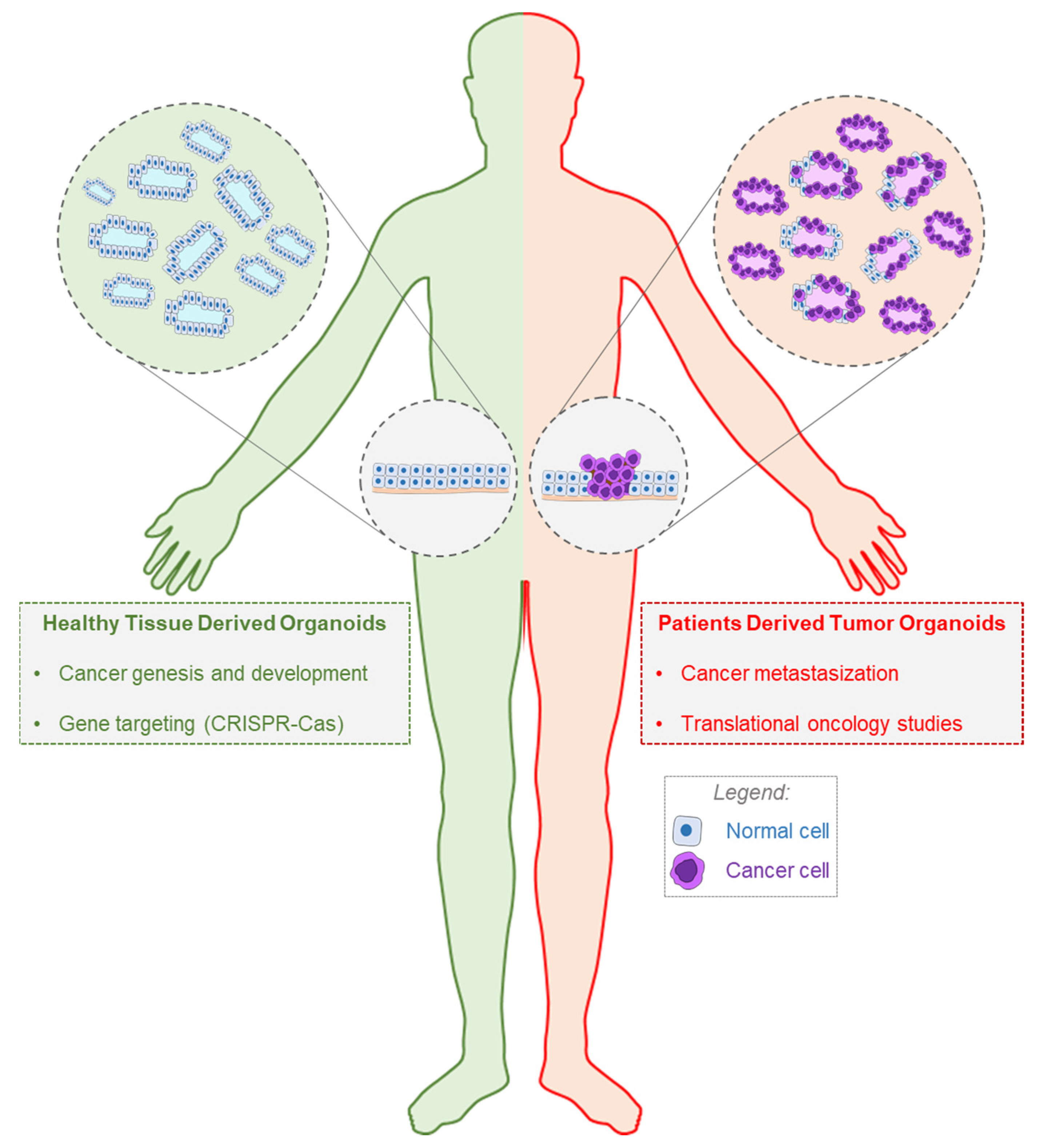
2. The Importance of Organoids in Oncology
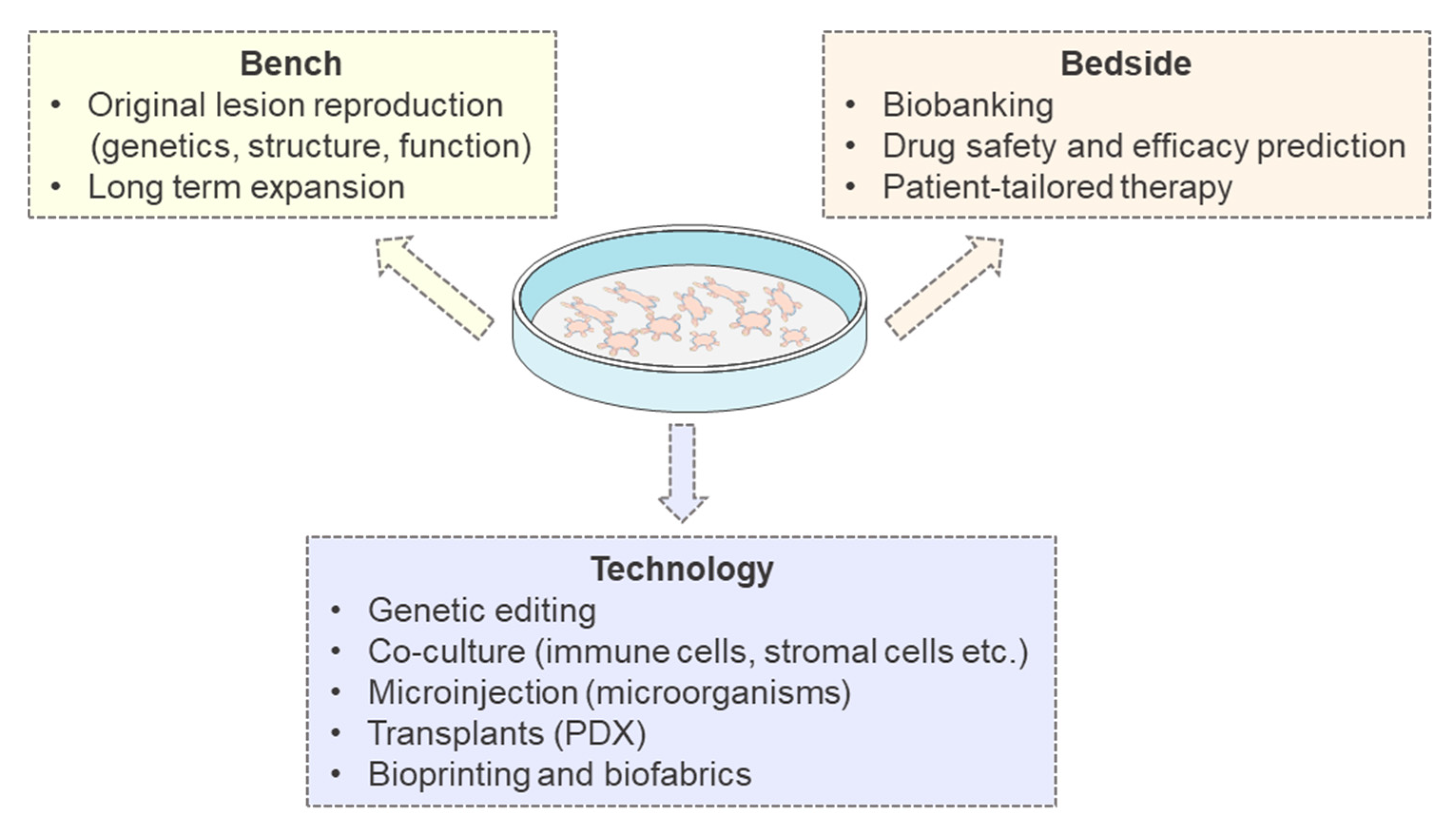
3. History and Applications of PDTOs in Translational Oncology
4. Current Ongoing Clinical Studies Using PTDOs in Cancer Care
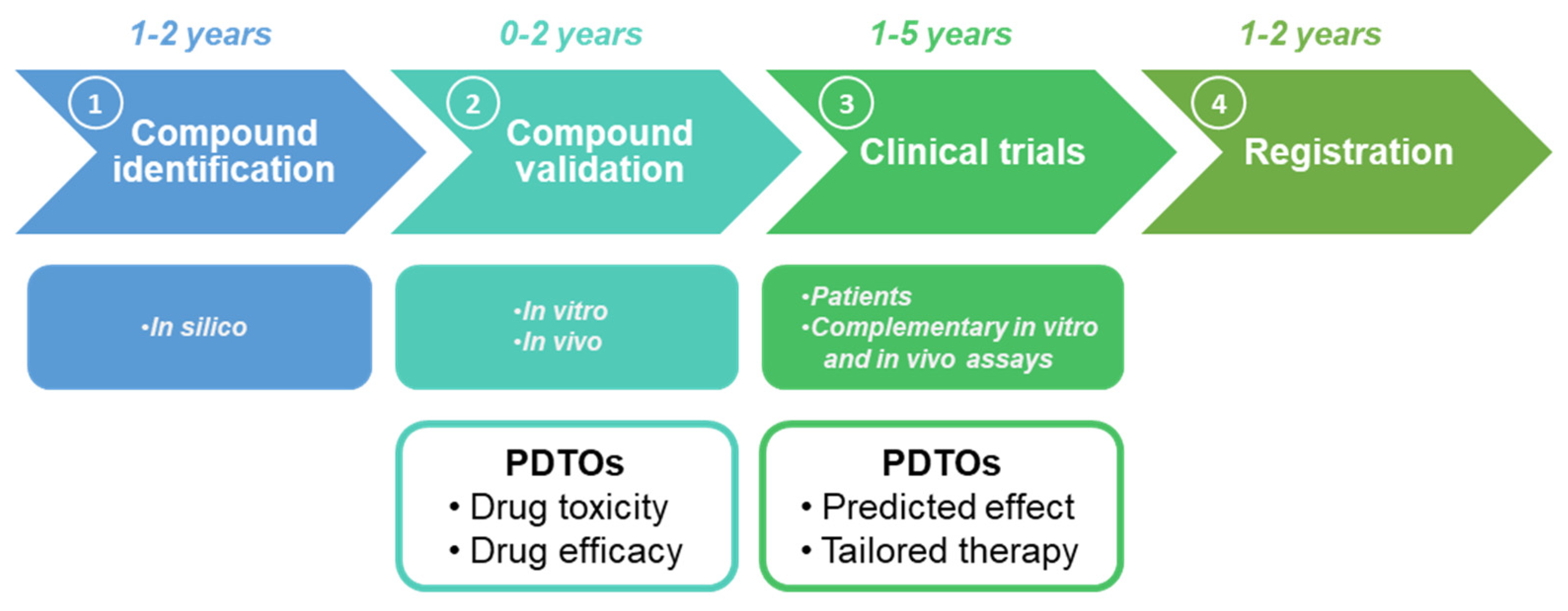
5. PDTOs and Drug Repositioning: A New Hope in Cancer Care
6. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Welch, D.R. Tumor Heterogeneity—A ’Contemporary Concept Founded on Historical Insights and Predictions. Cancer Res. 2016, 76, 4–6. [Google Scholar] [CrossRef] [PubMed]
- Foo, J.; Michor, F. Evolution of acquired resistance to anti-cancer therapy. J. Theor. Biol. 2014, 355, 10–20. [Google Scholar] [CrossRef] [PubMed]
- Walens, A.; Lin, J.; Damrauer, J.S.; McKinney, B.; Lupo, R.; Newcomb, R.; Fox, D.B.; Mabe, N.W.; Gresham, J.; Sheng, Z.; et al. Adaptation and selection shape clonal evolution of tumors during residual disease and recurrence. Nat. Commun. 2020, 11, 5017. [Google Scholar] [CrossRef] [PubMed]
- Granat, L.M.; Kambhampati, O.; Klosek, S.; Niedzwecki, B.; Parsa, K.; Zhang, D. The promises and challenges of patient-derived tumor organoids in drug development and precision oncology. Anim. Model. Exp. Med. 2019, 2, 150–161. [Google Scholar] [CrossRef]
- Nguyen, R.; Bae, S.D.W.; Zhou, G.; Read, S.A.; Ahlenstiel, G.; George, J.; Qiao, L. Application of organoids in translational research of human diseases with a particular focus on gastrointestinal cancers. Biochim. Biophys. Acta-Rev. Cancer 2020, 1873, 188350. [Google Scholar] [CrossRef]
- Lee, S.-J.; Lee, H.-A. Trends in the development of human stem cell-based non-animal drug testing models. Korean J. Physiol. Pharmacol. 2020, 24, 441–452. [Google Scholar] [CrossRef]
- Marshall, L.J.; Triunfol, M.; Seidle, T. Patient-Derived Xenograft vs. Organoids: A Preliminary Analysis of Cancer Research Output, Funding and Human Health Impact in 2014–2019. Animals 2020, 10, 1923. [Google Scholar] [CrossRef]
- Fan, H.; Demirci, U.; Chen, P. Emerging organoid models: Leaping forward in cancer research. J. Hematol. Oncol. 2019, 12, 142. [Google Scholar] [CrossRef]
- Xu, H.; Lyu, X.; Yi, M.; Zhao, W.; Song, Y.; Wu, K. Organoid technology and applications in cancer research. J. Hematol. Oncol. 2018, 11, 116. [Google Scholar] [CrossRef]
- Studer, L.; Vera, E.; Cornacchia, D. Programming and Reprogramming Cellular Age in the Era of Induced Pluripotency. Cell Stem Cell 2015, 16, 591–600. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Koo, B.-K.; Knoblich, J.A. Human organoids: Model systems for human biology and medicine. Nat. Rev. Mol. Cell Biol. 2020, 21, 571–584. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Yamanaka, S. Induction of Pluripotent Stem Cells from Mouse Embryonic and Adult Fibroblast Cultures by Defined Factors. Cell 2006, 126, 663–676. [Google Scholar] [CrossRef] [PubMed]
- Fujii, M.; Sato, T. Somatic cell-derived organoids as prototypes of human epithelial tissues and diseases. Nat. Mater. 2020. [Google Scholar] [CrossRef] [PubMed]
- McCauley, H.A.; Wells, J.M. Pluripotent stem cell-derived organoids: Using principles of developmental biology to grow human tissues in a dish. Development 2017, 144, 958–962. [Google Scholar] [CrossRef]
- Bartfeld, S.; Clevers, H. Stem cell-derived organoids and their application for medical research and patient treatment. J. Mol. Med. 2017, 95, 729–738. [Google Scholar] [CrossRef]
- Simian, M.; Bissell, M.J. Organoids: A historical perspective of thinking in three dimensions. J. Cell Biol. 2017, 216, 31–40. [Google Scholar] [CrossRef]
- Dontu, G. In vitro propagation and transcriptional profiling of human mammary stem/progenitor cells. Genes Dev. 2003, 17, 1253–1270. [Google Scholar] [CrossRef]
- Sato, T.; Vries, R.G.; Snippert, H.J.; van de Wetering, M.; Barker, N.; Stange, D.E.; van Es, J.H.; Abo, A.; Kujala, P.; Peters, P.J.; et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 2009, 459, 262–265. [Google Scholar] [CrossRef]
- Kaushik, G.; Ponnusamy, M.P.; Batra, S.K. Concise Review: Current Status of Three-Dimensional Organoids as Preclinical Models. Stem Cells 2018, 36, 1329–1340. [Google Scholar] [CrossRef]
- Perkhofer, L.; Frappart, P.-O.; Müller, M.; Kleger, A. Importance of organoids for personalized medicine. Per. Med. 2018, 15, 461–465. [Google Scholar] [CrossRef] [PubMed]
- Dekkers, J.F.; Wiegerinck, C.L.; de Jonge, H.R.; Bronsveld, I.; Janssens, H.M.; de Winter-de Groot, K.M.; Brandsma, A.M.; de Jong, N.W.M.; Bijvelds, M.J.C.; Scholte, B.J.; et al. A functional CFTR assay using primary cystic fibrosis intestinal organoids. Nat. Med. 2013, 19, 939–945. [Google Scholar] [CrossRef] [PubMed]
- Dekkers, J.F.; Berkers, G.; Kruisselbrink, E.; Vonk, A.; de Jonge, H.R.; Janssens, H.M.; Bronsveld, I.; van de Graaf, E.A.; Nieuwenhuis, E.E.S.; Houwen, R.H.J.; et al. Characterizing responses to CFTR-modulating drugs using rectal organoids derived from subjects with cystic fibrosis. Sci. Transl. Med. 2016, 8, 344ra84. [Google Scholar] [CrossRef] [PubMed]
- Boj, S.F.; Vonk, A.M.; Statia, M.; Su, J.; Vries, R.R.G.; Beekman, J.M.; Clevers, H. Forskolin-induced Swelling in Intestinal Organoids: An In Vitro Assay for Assessing Drug Response in Cystic Fibrosis Patients. J. Vis. Exp. 2017, 120, e55159. [Google Scholar] [CrossRef] [PubMed]
- Kondo, J.; Inoue, M. Application of Cancer Organoid Model for Drug Screening and Personalized Therapy. Cells 2019, 8, 470. [Google Scholar] [CrossRef]
- Lehmann, R.; Lee, C.M.; Shugart, E.C.; Benedetti, M.; Charo, R.A.; Gartner, Z.; Hogan, B.; Knoblich, J.; Nelson, C.M.; Wilson, K.M. Human organoids: A new dimension in cell biology. Mol. Biol. Cell 2019, 30, 1129–1137. [Google Scholar] [CrossRef]
- Jackson, E.L.; Lu, H. Three-dimensional models for studying development and disease: Moving on from organisms to organs-on-a-chip and organoids. Integr. Biol. 2016, 8, 672–683. [Google Scholar] [CrossRef]
- Kretzschmar, K.; Clevers, H. Organoids: Modeling Development and the Stem Cell Niche in a Dish. Dev. Cell 2016, 38, 590–600. [Google Scholar] [CrossRef]
- Mizutani, T.; Clevers, H. Primary Intestinal Epithelial Organoid Culture; Humana: New York, NY, USA, 2020; pp. 185–200. [Google Scholar]
- Wu, Y.; Zhao, Y.; Huan, L.; Zhao, J.; Zhou, Y.; Xu, L.; Hu, Z.; Liu, Y.; Chen, Z.; Wang, L.; et al. An LTR Retrotransposon-Derived Long Noncoding RNA lncMER52A Promotes Hepatocellular Carcinoma Progression by Binding p120-Catenin. Cancer Res. 2020, 80, 976–987. [Google Scholar] [CrossRef]
- Artegiani, B.; Hendriks, D.; Beumer, J.; Kok, R.; Zheng, X.; Joore, I.; Chuva de Sousa Lopes, S.; van Zon, J.; Tans, S.; Clevers, H. Fast and efficient generation of knock-in human organoids using homology-independent CRISPR–Cas9 precision genome editing. Nat. Cell Biol. 2020, 22, 321–331. [Google Scholar] [CrossRef]
- Nagle, P.W.; Plukker, J.T.M.; Muijs, C.T.; van Luijk, P.; Coppes, R.P. Patient-derived tumor organoids for prediction of cancer treatment response. Semin. Cancer Biol. 2018, 53, 258–264. [Google Scholar] [CrossRef] [PubMed]
- Driehuis, E.; Kretzschmar, K.; Clevers, H. Establishment of patient-derived cancer organoids for drug-screening applications. Nat. Protoc. 2020, 15, 3380–3409. [Google Scholar] [CrossRef] [PubMed]
- Drost, J.; van Jaarsveld, R.H.; Ponsioen, B.; Zimberlin, C.; van Boxtel, R.; Buijs, A.; Sachs, N.; Overmeer, R.M.; Offerhaus, G.J.; Begthel, H.; et al. Sequential cancer mutations in cultured human intestinal stem cells. Nature 2015, 521, 43–47. [Google Scholar] [CrossRef]
- Matano, M.; Date, S.; Shimokawa, M.; Takano, A.; Fujii, M.; Ohta, Y.; Watanabe, T.; Kanai, T.; Sato, T. Modeling colorectal cancer using CRISPR-Cas9–mediated engineering of human intestinal organoids. Nat. Med. 2015, 21, 256–262. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Barral, A.; Costales-Carrera, A.; Buira, S.P.; Jung, P.; Ferrer-Mayorga, G.; Larriba, M.J.; Bustamante-Madrid, P.; Domínguez, O.; Real, F.X.; Guerra-Pastrián, L.; et al. Vitamin D differentially regulates colon stem cells in patient-derived normal and tumor organoids. FEBS J. 2020, 287, 53–72. [Google Scholar] [CrossRef] [PubMed]
- Costales-Carrera, A.; Fernández-Barral, A.; Bustamante-Madrid, P.; Domínguez, O.; Guerra-Pastrián, L.; Cantero, R.; del Peso, L.; Burgos, A.; Barbáchano, A.; Muñoz, A. Comparative Study of Organoids from Patient-Derived Normal and Tumor Colon and Rectal Tissue. Cancers 2020, 12, 2302. [Google Scholar] [CrossRef] [PubMed]
- Shinozawa, T.; Kimura, M.; Cai, Y.; Saiki, N.; Yoneyama, Y.; Ouchi, R.; Koike, H.; Maezawa, M.; Zhang, R.-R.; Dunn, A.; et al. High-Fidelity Drug-Induced Liver Injury Screen Using Human Pluripotent Stem Cell–Derived Organoids. Gastroenterology 2020. [Google Scholar] [CrossRef]
- Ooft, S.N.; Weeber, F.; Dijkstra, K.K.; McLean, C.M.; Kaing, S.; van Werkhoven, E.; Schipper, L.; Hoes, L.; Vis, D.J.; van de Haar, J.; et al. Patient-derived organoids can predict response to chemotherapy in metastatic colorectal cancer patients. Sci. Transl. Med. 2019, 11, eaay2574. [Google Scholar] [CrossRef]
- Bruun, J.; Kryeziu, K.; Eide, P.W.; Moosavi, S.H.; Eilertsen, I.A.; Langerud, J.; Røsok, B.; Totland, M.Z.; Brunsell, T.H.; Pellinen, T.; et al. Patient-Derived Organoids from Multiple Colorectal Cancer Liver Metastases Reveal Moderate Intra-patient Pharmacotranscriptomic Heterogeneity. Clin. Cancer Res. 2020, 26, 4107–4119. [Google Scholar] [CrossRef]
- Yang, H.; Sun, L.; Liu, M.; Mao, Y. Patient-derived organoids: A promising model for personalized cancer treatment. Gastroenterol. Rep. 2018, 6, 243–245. [Google Scholar] [CrossRef]
- Eiraku, M.; Watanabe, K.; Matsuo-Takasaki, M.; Kawada, M.; Yonemura, S.; Matsumura, M.; Wataya, T.; Nishiyama, A.; Muguruma, K.; Sasai, Y. Self-Organized Formation of Polarized Cortical Tissues from ESCs and Its Active Manipulation by Extrinsic Signals. Cell Stem Cell 2008, 3, 519–532. [Google Scholar] [CrossRef] [PubMed]
- Eiraku, M.; Takata, N.; Ishibashi, H.; Kawada, M.; Sakakura, E.; Okuda, S.; Sekiguchi, K.; Adachi, T.; Sasai, Y. Self-organizing optic-cup morphogenesis in three-dimensional culture. Nature 2011, 472, 51–56. [Google Scholar] [CrossRef] [PubMed]
- Sato, T.; Stange, D.E.; Ferrante, M.; Vries, R.G.J.; van Es, J.H.; van den Brink, S.; van Houdt, W.J.; Pronk, A.; van Gorp, J.; Siersema, P.D.; et al. Long-term Expansion of Epithelial Organoids From Human Colon, Adenoma, Adenocarcinoma, and Barrett’s Epithelium. Gastroenterology 2011, 141, 1762–1772. [Google Scholar] [CrossRef] [PubMed]
- Jung, P.; Sato, T.; Merlos-Suárez, A.; Barriga, F.M.; Iglesias, M.; Rossell, D.; Auer, H.; Gallardo, M.; Blasco, M.A.; Sancho, E.; et al. Isolation and in vitro expansion of human colonic stem cells. Nat. Med. 2011, 17, 1225–1227. [Google Scholar] [CrossRef] [PubMed]
- Lancaster, M.A.; Renner, M.; Martin, C.-A.; Wenzel, D.; Bicknell, L.S.; Hurles, M.E.; Homfray, T.; Penninger, J.M.; Jackson, A.P.; Knoblich, J.A. Cerebral organoids model human brain development and microcephaly. Nature 2013, 501, 373–379. [Google Scholar] [CrossRef] [PubMed]
- Qian, X.; Nguyen, H.N.; Song, M.M.; Hadiono, C.; Ogden, S.C.; Hammack, C.; Yao, B.; Hamersky, G.R.; Jacob, F.; Zhong, C.; et al. Brain-Region-Specific Organoids Using Mini-bioreactors for Modeling ZIKV Exposure. Cell 2016, 165, 1238–1254. [Google Scholar] [CrossRef] [PubMed]
- Renner, M.; Lancaster, M.A.; Bian, S.; Choi, H.; Ku, T.; Peer, A.; Chung, K.; Knoblich, J.A. Self-organized developmental patterning and differentiation in cerebral organoids. EMBO J. 2017, 36, 1316–1329. [Google Scholar] [CrossRef]
- Giandomenico, S.L.; Mierau, S.B.; Gibbons, G.M.; Wenger, L.M.D.; Masullo, L.; Sit, T.; Sutcliffe, M.; Boulanger, J.; Tripodi, M.; Derivery, E.; et al. Cerebral organoids at the air–liquid interface generate diverse nerve tracts with functional output. Nat. Neurosci. 2019, 22, 669–679. [Google Scholar] [CrossRef]
- Velasco, S.; Kedaigle, A.J.; Simmons, S.K.; Nash, A.; Rocha, M.; Quadrato, G.; Paulsen, B.; Nguyen, L.; Adiconis, X.; Regev, A.; et al. Individual brain organoids reproducibly form cell diversity of the human cerebral cortex. Nature 2019, 570, 523–527. [Google Scholar] [CrossRef]
- Karthaus, W.R.; Iaquinta, P.J.; Drost, J.; Gracanin, A.; van Boxtel, R.; Wongvipat, J.; Dowling, C.M.; Gao, D.; Begthel, H.; Sachs, N.; et al. Identification of Multipotent Luminal Progenitor Cells in Human Prostate Organoid Cultures. Cell 2014, 159, 163–175. [Google Scholar] [CrossRef]
- Chua, C.W.; Shibata, M.; Lei, M.; Toivanen, R.; Barlow, L.J.; Bergren, S.K.; Badani, K.K.; McKiernan, J.M.; Benson, M.C.; Hibshoosh, H.; et al. Single luminal epithelial progenitors can generate prostate organoids in culture. Nat. Cell Biol. 2014, 16, 951–961. [Google Scholar] [CrossRef] [PubMed]
- Boj, S.F.; Hwang, C.-I.; Baker, L.A.; Chio, I.I.C.; Engle, D.D.; Corbo, V.; Jager, M.; Ponz-Sarvise, M.; Tiriac, H.; Spector, M.S.; et al. Organoid Models of Human and Mouse Ductal Pancreatic Cancer. Cell 2015, 160, 324–338. [Google Scholar] [CrossRef] [PubMed]
- Kessler, M.; Hoffmann, K.; Brinkmann, V.; Thieck, O.; Jackisch, S.; Toelle, B.; Berger, H.; Mollenkopf, H.-J.; Mangler, M.; Sehouli, J.; et al. The Notch and Wnt pathways regulate stemness and differentiation in human fallopian tube organoids. Nat. Commun. 2015, 6, 8989. [Google Scholar] [CrossRef]
- Bartfeld, S.; Bayram, T.; van de Wetering, M.; Huch, M.; Begthel, H.; Kujala, P.; Vries, R.; Peters, P.J.; Clevers, H. In Vitro Expansion of Human Gastric Epithelial Stem Cells and Their Responses to Bacterial Infection. Gastroenterology 2015, 148, 126–136.e6. [Google Scholar] [CrossRef]
- Hubert, C.G.; Rivera, M.; Spangler, L.C.; Wu, Q.; Mack, S.C.; Prager, B.C.; Couce, M.; McLendon, R.E.; Sloan, A.E.; Rich, J.N. A Three-Dimensional Organoid Culture System Derived from Human Glioblastomas Recapitulates the Hypoxic Gradients and Cancer Stem Cell Heterogeneity of Tumors Found In Vivo. Cancer Res. 2016, 76, 2465–2477. [Google Scholar] [CrossRef]
- Turco, M.Y.; Gardner, L.; Hughes, J.; Cindrova-Davies, T.; Gomez, M.J.; Farrell, L.; Hollinshead, M.; Marsh, S.G.E.; Brosens, J.J.; Critchley, H.O.; et al. Long-term, hormone-responsive organoid cultures of human endometrium in a chemically defined medium. Nat. Cell Biol. 2017, 19, 568–577. [Google Scholar] [CrossRef] [PubMed]
- Broutier, L.; Mastrogiovanni, G.; Verstegen, M.M.; Francies, H.E.; Gavarró, L.M.; Bradshaw, C.R.; Allen, G.E.; Arnes-Benito, R.; Sidorova, O.; Gaspersz, M.P.; et al. Human primary liver cancer–derived organoid cultures for disease modeling and drug screening. Nat. Med. 2017, 23, 1424–1435. [Google Scholar] [CrossRef]
- Boretto, M.; Maenhoudt, N.; Luo, X.; Hennes, A.; Boeckx, B.; Bui, B.; Heremans, R.; Perneel, L.; Kobayashi, H.; Van Zundert, I.; et al. Patient-derived organoids from endometrial disease capture clinical heterogeneity and are amenable to drug screening. Nat. Cell Biol. 2019, 21, 1041–1051. [Google Scholar] [CrossRef] [PubMed]
- Schutgens, F.; Rookmaaker, M.B.; Margaritis, T.; Rios, A.; Ammerlaan, C.; Jansen, J.; Gijzen, L.; Vormann, M.; Vonk, A.; Viveen, M.; et al. Tubuloids derived from human adult kidney and urine for personalized disease modeling. Nat. Biotechnol. 2019, 37, 303–313. [Google Scholar] [CrossRef]
- Sachs, N.; Papaspyropoulos, A.; Zomer-van Ommen, D.D.; Heo, I.; Böttinger, L.; Klay, D.; Weeber, F.; Huelsz-Prince, G.; Iakobachvili, N.; Amatngalim, G.D.; et al. Long-term expanding human airway organoids for disease modeling. EMBO J. 2019, 38, e100300. [Google Scholar] [CrossRef]
- Dekkers, J.F.; Alieva, M.; Wellens, L.M.; Ariese, H.C.R.; Jamieson, P.R.; Vonk, A.M.; Amatngalim, G.D.; Hu, H.; Oost, K.C.; Snippert, H.J.G.; et al. High-resolution 3D imaging of fixed and cleared organoids. Nat. Protoc. 2019, 14, 1756–1771. [Google Scholar] [CrossRef] [PubMed]
- Walsh, A.J.; Cook, R.S.; Sanders, M.E.; Aurisicchio, L.; Ciliberto, G.; Arteaga, C.L.; Skala, M.C. Quantitative Optical Imaging of Primary Tumor Organoid Metabolism Predicts Drug Response in Breast Cancer. Cancer Res. 2014, 74, 5184–5194. [Google Scholar] [CrossRef] [PubMed]
- Grün, D.; Lyubimova, A.; Kester, L.; Wiebrands, K.; Basak, O.; Sasaki, N.; Clevers, H.; van Oudenaarden, A. Single-cell messenger RNA sequencing reveals rare intestinal cell types. Nature 2015, 525, 251–255. [Google Scholar] [CrossRef] [PubMed]
- Boers, S.N.; Delden, J.J.; Clevers, H.; Bredenoord, A.L. Organoid biobanking: Identifying the ethics. EMBO Rep. 2016, 17, 938–941. [Google Scholar] [CrossRef] [PubMed]
- Clinton, J.; McWilliams-Koeppen, P. Initiation, Expansion, and Cryopreservation of Human Primary Tissue-Derived Normal and Diseased Organoids in Embedded Three-Dimensional Culture. Curr. Protoc. Cell Biol. 2019, 82, e66. [Google Scholar] [CrossRef]
- Yin, X.; Mead, B.E.; Safaee, H.; Langer, R.; Karp, J.M.; Levy, O. Engineering Stem Cell Organoids. Cell Stem Cell 2016, 18, 25–38. [Google Scholar] [CrossRef]
- Huch, M.; Knoblich, J.A.; Lutolf, M.P.; Martinez-Arias, A. The hope and the hype of organoid research. Development 2017, 144, 938–941. [Google Scholar] [CrossRef]
- Fujii, E.; Kato, A.; Suzuki, M. Patient-derived xenograft (PDX) models: Characteristics and points to consider for the process of establishment. J. Toxicol. Pathol. 2020, 33, 153–160. [Google Scholar] [CrossRef]
- Willemse, J.; Lieshout, R.; van der Laan, L.J.W.; Verstegen, M.M.A. From organoids to organs: Bioengineering liver grafts from hepatic stem cells and matrix. Best Pract. Res. Clin. Gastroenterol. 2017, 31, 151–159. [Google Scholar] [CrossRef]
- Brandenberg, N.; Hoehnel, S.; Kuttler, F.; Homicsko, K.; Ceroni, C.; Ringel, T.; Gjorevski, N.; Schwank, G.; Coukos, G.; Turcatti, G.; et al. High-throughput automated organoid culture via stem-cell aggregation in microcavity arrays. Nat. Biomed. Eng. 2020, 4, 863–874. [Google Scholar] [CrossRef]
- Grebenyuk, S.; Ranga, A. Engineering Organoid Vascularization. Front. Bioeng. Biotechnol. 2019, 7, 39. [Google Scholar] [CrossRef] [PubMed]
- Nikolaev, M.; Mitrofanova, O.; Broguiere, N.; Geraldo, S.; Dutta, D.; Tabata, Y.; Elci, B.; Brandenberg, N.; Kolotuev, I.; Gjorevski, N.; et al. Homeostatic mini-intestines through scaffold-guided organoid morphogenesis. Nature 2020, 585, 574–578. [Google Scholar] [CrossRef] [PubMed]
- Lawlor, K.T.; Vanslambrouck, J.M.; Higgins, J.W.; Chambon, A.; Bishard, K.; Arndt, D.; Er, P.X.; Wilson, S.B.; Howden, S.E.; Tan, K.S.; et al. Cellular extrusion bioprinting improves kidney organoid reproducibility and conformation. Nat. Mater. 2020. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.; Holtzinger, A.; Jagan, I.; BeGora, M.; Lohse, I.; Ngai, N.; Nostro, C.; Wang, R.; Muthuswamy, L.B.; Crawford, H.C.; et al. Ductal pancreatic cancer modeling and drug screening using human pluripotent stem cell– and patient-derived tumor organoids. Nat. Med. 2015, 21, 1364–1371. [Google Scholar] [CrossRef] [PubMed]
- Roerink, S.F.; Sasaki, N.; Lee-Six, H.; Young, M.D.; Alexandrov, L.B.; Behjati, S.; Mitchell, T.J.; Grossmann, S.; Lightfoot, H.; Egan, D.A.; et al. Intra-tumour diversification in colorectal cancer at the single-cell level. Nature 2018, 556, 457–462. [Google Scholar] [CrossRef] [PubMed]
- Pasch, C.A.; Favreau, P.F.; Yueh, A.E.; Babiarz, C.P.; Gillette, A.A.; Sharick, J.T.; Karim, M.R.; Nickel, K.P.; DeZeeuw, A.K.; Sprackling, C.M.; et al. Patient-Derived Cancer Organoid Cultures to Predict Sensitivity to Chemotherapy and Radiation. Clin. Cancer Res. 2019, 25, 5376–5387. [Google Scholar] [CrossRef]
- Nanki, Y.; Chiyoda, T.; Hirasawa, A.; Ookubo, A.; Itoh, M.; Ueno, M.; Akahane, T.; Kameyama, K.; Yamagami, W.; Kataoka, F.; et al. Patient-derived ovarian cancer organoids capture the genomic profiles of primary tumours applicable for drug sensitivity and resistance testing. Sci. Rep. 2020, 10, 12581. [Google Scholar] [CrossRef]
- Pleguezuelos-Manzano, C.; Puschhof, J.; Rosendahl Huber, A.; van Hoeck, A.; Wood, H.M.; Nomburg, J.; Gurjao, C.; Manders, F.; Dalmasso, G.; Stege, P.B.; et al. Mutational signature in colorectal cancer caused by genotoxic pks+ E. coli. Nature 2020, 580, 269–273. [Google Scholar] [CrossRef]
- Tsai, Y.-H.; Czerwinski, M.; Wu, A.; Dame, M.K.; Attili, D.; Hill, E.; Colacino, J.A.; Nowacki, L.M.; Shroyer, N.F.; Higgins, P.D.R.; et al. A Method for Cryogenic Preservation of Human Biopsy Specimens and Subsequent Organoid Culture. Cell. Mol. Gastroenterol. Hepatol. 2018, 6, 218–222.e7. [Google Scholar] [CrossRef]
- Bleijs, M.; Wetering, M.; Clevers, H.; Drost, J. Xenograft and organoid model systems in cancer research. EMBO J. 2019, 38, e101654. [Google Scholar] [CrossRef]
- Gao, D.; Vela, I.; Sboner, A.; Iaquinta, P.J.; Karthaus, W.R.; Gopalan, A.; Dowling, C.; Wanjala, J.N.; Undvall, E.A.; Arora, V.K.; et al. Organoid Cultures Derived from Patients with Advanced Prostate Cancer. Cell 2014, 159, 176–187. [Google Scholar] [CrossRef]
- Van de Wetering, M.; Francies, H.E.; Francis, J.M.; Bounova, G.; Iorio, F.; Pronk, A.; van Houdt, W.; van Gorp, J.; Taylor-Weiner, A.; Kester, L.; et al. Prospective Derivation of a Living Organoid Biobank of Colorectal Cancer Patients. Cell 2015, 161, 933–945. [Google Scholar] [CrossRef] [PubMed]
- Weeber, F.; van de Wetering, M.; Hoogstraat, M.; Dijkstra, K.K.; Krijgsman, O.; Kuilman, T.; Gadellaa-van Hooijdonk, C.G.M.; van der Velden, D.L.; Peeper, D.S.; Cuppen, E.P.J.G.; et al. Preserved genetic diversity in organoids cultured from biopsies of human colorectal cancer metastases. Proc. Natl. Acad. Sci. USA 2015, 112, 13308–13311. [Google Scholar] [CrossRef] [PubMed]
- Fujii, M.; Shimokawa, M.; Date, S.; Takano, A.; Matano, M.; Nanki, K.; Ohta, Y.; Toshimitsu, K.; Nakazato, Y.; Kawasaki, K.; et al. A Colorectal Tumor Organoid Library Demonstrates Progressive Loss of Niche Factor Requirements during Tumorigenesis. Cell Stem Cell 2016, 18, 827–838. [Google Scholar] [CrossRef] [PubMed]
- Seino, T.; Kawasaki, S.; Shimokawa, M.; Tamagawa, H.; Toshimitsu, K.; Fujii, M.; Ohta, Y.; Matano, M.; Nanki, K.; Kawasaki, K.; et al. Human Pancreatic Tumor Organoids Reveal Loss of Stem Cell Niche Factor Dependence during Disease Progression. Cell Stem Cell 2018, 22, 454–467.e6. [Google Scholar] [CrossRef] [PubMed]
- Sachs, N.; de Ligt, J.; Kopper, O.; Gogola, E.; Bounova, G.; Weeber, F.; Balgobind, A.V.; Wind, K.; Gracanin, A.; Begthel, H.; et al. A Living Biobank of Breast Cancer Organoids Captures Disease Heterogeneity. Cell 2018, 172, 373–386.e10. [Google Scholar] [CrossRef]
- Vlachogiannis, G.; Hedayat, S.; Vatsiou, A.; Jamin, Y.; Fernández-Mateos, J.; Khan, K.; Lampis, A.; Eason, K.; Huntingford, I.; Burke, R.; et al. Patient-derived organoids model treatment response of metastatic gastrointestinal cancers. Science 2018, 359, 920–926. [Google Scholar] [CrossRef]
- Lee, S.H.; Hu, W.; Matulay, J.T.; Silva, M.V.; Owczarek, T.B.; Kim, K.; Chua, C.W.; Barlow, L.J.; Kandoth, C.; Williams, A.B.; et al. Tumor Evolution and Drug Response in Patient-Derived Organoid Models of Bladder Cancer. Cell 2018, 173, 515–528.e17. [Google Scholar] [CrossRef]
- Tiriac, H.; Belleau, P.; Engle, D.D.; Plenker, D.; Deschênes, A.; Somerville, T.D.D.; Froeling, F.E.M.; Burkhart, R.A.; Denroche, R.E.; Jang, G.-H.; et al. Organoid Profiling Identifies Common Responders to Chemotherapy in Pancreatic Cancer. Cancer Discov. 2018, 8, 1112–1129. [Google Scholar] [CrossRef]
- Yan, H.H.N.; Siu, H.C.; Law, S.; Ho, S.L.; Yue, S.S.K.; Tsui, W.Y.; Chan, D.; Chan, A.S.; Ma, S.; Lam, K.O.; et al. A Comprehensive Human Gastric Cancer Organoid Biobank Captures Tumor Subtype Heterogeneity and Enables Therapeutic Screening. Cell Stem Cell 2018, 23, 882–897.e11. [Google Scholar] [CrossRef]
- Beshiri, M.L.; Tice, C.M.; Tran, C.; Nguyen, H.M.; Sowalsky, A.G.; Agarwal, S.; Jansson, K.H.; Yang, Q.; McGowen, K.M.; Yin, J.; et al. A PDX/Organoid Biobank of Advanced Prostate Cancers Captures Genomic and Phenotypic Heterogeneity for Disease Modeling and Therapeutic Screening. Clin. Cancer Res. 2018, 24, 4332–4345. [Google Scholar] [CrossRef] [PubMed]
- Driehuis, E.; Kolders, S.; Spelier, S.; Lõhmussaar, K.; Willems, S.M.; Devriese, L.A.; de Bree, R.; de Ruiter, E.J.; Korving, J.; Begthel, H.; et al. Oral Mucosal Organoids as a Potential Platform for Personalized Cancer Therapy. Cancer Discov. 2019, 9, 852–871. [Google Scholar] [CrossRef] [PubMed]
- Mullenders, J.; de Jongh, E.; Brousali, A.; Roosen, M.; Blom, J.P.A.; Begthel, H.; Korving, J.; Jonges, T.; Kranenburg, O.; Meijer, R.; et al. Mouse and human urothelial cancer organoids: A tool for bladder cancer research. Proc. Natl. Acad. Sci. USA 2019, 116, 4567–4574. [Google Scholar] [CrossRef] [PubMed]
- Kopper, O.; de Witte, C.J.; Lõhmussaar, K.; Valle-Inclan, J.E.; Hami, N.; Kester, L.; Balgobind, A.V.; Korving, J.; Proost, N.; Begthel, H.; et al. An organoid platform for ovarian cancer captures intra- and interpatient heterogeneity. Nat. Med. 2019, 25, 838–849. [Google Scholar] [CrossRef]
- Yao, Y.; Xu, X.; Yang, L.; Zhu, J.; Wan, J.; Shen, L.; Xia, F.; Fu, G.; Deng, Y.; Pan, M.; et al. Patient-Derived Organoids Predict Chemoradiation Responses of Locally Advanced Rectal Cancer. Cell Stem Cell 2020, 26, 17–26.e6. [Google Scholar] [CrossRef]
- Jacob, F.; Salinas, R.D.; Zhang, D.Y.; Nguyen, P.T.T.; Schnoll, J.G.; Wong, S.Z.H.; Thokala, R.; Sheikh, S.; Saxena, D.; Prokop, S.; et al. A Patient-Derived Glioblastoma Organoid Model and Biobank Recapitulates Inter- and Intra-tumoral Heterogeneity. Cell 2020, 180, 188–204.e22. [Google Scholar] [CrossRef]
- Calandrini, C.; Schutgens, F.; Oka, R.; Margaritis, T.; Candelli, T.; Mathijsen, L.; Ammerlaan, C.; van Ineveld, R.L.; Derakhshan, S.; de Haan, S.; et al. An organoid biobank for childhood kidney cancers that captures disease and tissue heterogeneity. Nat. Commun. 2020, 11, 1310. [Google Scholar] [CrossRef]
- Lampis, A.; Carotenuto, P.; Vlachogiannis, G.; Cascione, L.; Hedayat, S.; Burke, R.; Clarke, P.; Bosma, E.; Simbolo, M.; Scarpa, A.; et al. MIR21 Drives Resistance to Heat Shock Protein 90 Inhibition in Cholangiocarcinoma. Gastroenterology 2018, 154, 1066–1079.e5. [Google Scholar] [CrossRef]
- Narasimhan, V.; Wright, J.A.; Churchill, M.; Wang, T.; Rosati, R.; Lannagan, T.R.M.; Vrbanac, L.; Richardson, A.B.; Kobayashi, H.; Price, T.; et al. Medium-throughput Drug Screening of Patient-derived Organoids from Colorectal Peritoneal Metastases to Direct Personalized Therapy. Clin. Cancer Res. 2020, 26, 3662–3670. [Google Scholar] [CrossRef]
- World Health Organization WHO. WHO Tral Search. Available online: https://apps.who.int/trialsearch/Default.aspx (accessed on 5 November 2020).
- NIH National Institutes of Health. Clinical Trials NIH. Available online: https://clinicaltrials.gov/ (accessed on 5 November 2020).
- Al-Bedeary, S.; Getta, H.; Al-Sharafi, D. The hallmarks of cancer and their therapeutic targeting in current use and clinical trials. Iraqi J. Hematol. 2020, 9, 1. [Google Scholar] [CrossRef]
- Dinić, J.; Efferth, T.; García-Sosa, A.T.; Grahovac, J.; Padrón, J.M.; Pajeva, I.; Rizzolio, F.; Saponara, S.; Spengler, G.; Tsakovska, I. Repurposing old drugs to fight multidrug resistant cancers. Drug Resist. Updat. 2020, 52, 100713. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Zhou, L.; Xie, N.; Nice, E.C.; Zhang, T.; Cui, Y.; Huang, C. Overcoming cancer therapeutic bottleneck by drug repurposing. Signal Transduct. Target. Ther. 2020, 5, 113. [Google Scholar] [CrossRef] [PubMed]
- Ashburn, T.T.; Thor, K.B. Drug repositioning: Identifying and developing new uses for existing drugs. Nat. Rev. Drug Discov. 2004, 3, 673–683. [Google Scholar] [CrossRef] [PubMed]
- Lin, Z.; Li, G.; Xu, X.; Mei, J. Does metformin improve the efficacy of standard epidermal growth factor receptor-tyrosine kinase inhibitor treatment for patients with advanced non-small-cell lung cancer? Interact. Cardiovasc. Thorac. Surg. 2020. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Su, Z.; Wang, R.; Ge, F.; Huo, Z.; Wen, Y.; Li, C.; Liang, H.; He, J.; Liang, W. Aspirin and risk of different cancers: An umbrella meta-analysis. Ann. Transl. Med. 2020, 8, 1333. [Google Scholar] [CrossRef] [PubMed]
- Lv, H.; Shi, D.; Fei, M.; Chen, Y.; Xie, F.; Wang, Z.; Wang, Y.; Hu, P. Association Between Statin Use and Prognosis of Breast Cancer: A Meta-Analysis of Cohort Studies. Front. Oncol. 2020, 10, 2113. [Google Scholar] [CrossRef]
- Caciolla, J.; Bisi, A.; Belluti, F.; Rampa, A.; Gobbi, S. Reconsidering Aromatase for Breast Cancer Treatment: New Roles for an Old Target. Molecules 2020, 25, 5351. [Google Scholar] [CrossRef]
- Chang, C.-H.; Lee, C.-H.; Ko, J.-C.; Chang, L.-Y.; Lee, M.-C.; Zhang, J.-F.; Wang, J.-Y.; Shih, J.-Y.; Yu, C.-J. Effect of β-Blocker in Treatment-Naïve Patients With Advanced Lung Adenocarcinoma Receiving First-Generation EGFR-TKIs. Front. Oncol. 2020, 10. [Google Scholar] [CrossRef]
- Manic, G.; Obrist, F.; Kroemer, G.; Vitale, I.; Galluzzi, L. Chloroquine and hydroxychloroquine for cancer therapy. Mol. Cell. Oncol. 2014, 1, e29911. [Google Scholar] [CrossRef]
- Gagelmann, N.; Kröger, N. The role of novel agents for consolidation after autologous transplantation in newly diagnosed multiple myeloma: A systematic review. Ann. Hematol. 2020. [Google Scholar] [CrossRef]
- Poortmans, P.M.; Guarneri, V.; Cardoso, M.-J. Cancer and COVID-19: What do we really know? Lancet 2020, 395, 1884–1885. [Google Scholar] [CrossRef]
- Zhou, Y.; Wang, F.; Tang, J.; Nussinov, R.; Cheng, F. Artificial intelligence in COVID-19 drug repurposing. Lancet Digit. Heal. 2020, 2, e667–e676. [Google Scholar] [CrossRef]
- Xue, H.; Li, J.; Xie, H.; Wang, Y. Review of Drug Repositioning Approaches and Resources. Int. J. Biol. Sci. 2018, 14, 1232–1244. [Google Scholar] [CrossRef] [PubMed]
- Ciallella, J.R.; Reaume, A.G. In vivo phenotypic screening: Clinical proof of concept for a drug repositioning approach. Drug Discov. Today Technol. 2017, 23, 45–52. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, G.F.; Pritchard, K. In Vitro Screening for Drug Repositioning. J. Biomol. Screen. 2015, 20, 167–179. [Google Scholar] [CrossRef] [PubMed]
- GNS, H.S.; GR, S.; Murahari, M.; Krishnamurthy, M. An update on Drug Repurposing: Re-written saga of the drug’s fate. Biomed. Pharmacother. 2019, 110, 700–716. [Google Scholar] [CrossRef]
- Hernandez, J.J.; Pryszlak, M.; Smith, L.; Yanchus, C.; Kurji, N.; Shahani, V.M.; Molinski, S.V. Giving Drugs a Second Chance: Overcoming Regulatory and Financial Hurdles in Repurposing Approved Drugs As Cancer Therapeutics. Front. Oncol. 2017, 7, 273. [Google Scholar] [CrossRef]
- Saiyed, M.M.; Ong, P.S.; Chew, L. Off-label drug use in oncology: A systematic review of literature. J. Clin. Pharm. Ther. 2017, 42, 251–258. [Google Scholar] [CrossRef]
- Herrero Fernandez, M.; Molina Villaverde, R.; Arroyo Yustos, M.; Navarro Expósito, F.; Lopez Gonzalez, J.L.; Luque Infantes, M.R.; Alvarez-Mon Soto, M. The Off-Label Use of Antineoplastics in Oncology Is Limited But Has Notable Scientific Support in a University Hospital Setting. Front. Pharmacol. 2019, 10. [Google Scholar] [CrossRef]
- Clevers, H. COVID-19: Organoids go viral. Nat. Rev. Mol. Cell Biol. 2020, 21, 355–356. [Google Scholar] [CrossRef]
- Xu, M.; Lee, E.M.; Wen, Z.; Cheng, Y.; Huang, W.-K.; Qian, X.; TCW, J.; Kouznetsova, J.; Ogden, S.C.; Hammack, C.; et al. Identification of small-molecule inhibitors of Zika virus infection and induced neural cell death via a drug repurposing screen. Nat. Med. 2016, 22, 1101–1107. [Google Scholar] [CrossRef] [PubMed]
- Della Corte, C.M.; Barra, G.; Ciaramella, V.; Di Liello, R.; Vicidomini, G.; Zappavigna, S.; Luce, A.; Abate, M.; Fiorelli, A.; Caraglia, M.; et al. Antitumor activity of dual blockade of PD-L1 and MEK in NSCLC patients derived three-dimensional spheroid cultures. J. Exp. Clin. Cancer Res. 2019, 38, 253. [Google Scholar] [CrossRef] [PubMed]
- Engel, R.M.; Chan, W.H.; Nickless, D.; Hlavca, S.; Richards, E.; Kerr, G.; Oliva, K.; McMurrick, P.J.; Jardé, T.; Abud, H.E. Patient-Derived Colorectal Cancer Organoids Upregulate Revival Stem Cell Marker Genes following Chemotherapeutic Treatment. J. Clin. Med. 2020, 9, 128. [Google Scholar] [CrossRef] [PubMed]
- Sudhan, D.R.; Guerrero-Zotano, A.; Won, H.; González Ericsson, P.; Servetto, A.; Huerta-Rosario, M.; Ye, D.; Lee, K.; Formisano, L.; Guo, Y.; et al. Hyperactivation of TORC1 Drives Resistance to the Pan-HER Tyrosine Kinase Inhibitor Neratinib in HER2-Mutant Cancers. Cancer Cell 2020, 37, 183–199.e5. [Google Scholar] [CrossRef]
- Yu, J.; Qin, B.; Moyer, A.M.; Nowsheen, S.; Liu, T.; Qin, S.; Zhuang, Y.; Liu, D.; Lu, S.W.; Kalari, K.R.; et al. DNA methyltransferase expression in triple-negative breast cancer predicts sensitivity to decitabine. J. Clin. Investig. 2018, 128, 2376–2388. [Google Scholar] [CrossRef]
- Kim, M.; Mun, H.; Sung, C.O.; Cho, E.J.; Jeon, H.-J.; Chun, S.-M.; Jung, D.J.; Shin, T.H.; Jeong, G.S.; Kim, D.K.; et al. Patient-derived lung cancer organoids as in vitro cancer models for therapeutic screening. Nat. Commun. 2019, 10, 3991. [Google Scholar] [CrossRef] [PubMed]
- Phan, N.; Hong, J.J.; Tofig, B.; Mapua, M.; Elashoff, D.; Moatamed, N.A.; Huang, J.; Memarzadeh, S.; Damoiseaux, R.; Soragni, A. A simple high-throughput approach identifies actionable drug sensitivities in patient-derived tumor organoids. Commun. Biol. 2019, 2, 78. [Google Scholar] [CrossRef]
- Pauli, C.; Hopkins, B.D.; Prandi, D.; Shaw, R.; Fedrizzi, T.; Sboner, A.; Sailer, V.; Augello, M.; Puca, L.; Rosati, R.; et al. Personalized In Vitro and In Vivo Cancer Models to Guide Precision Medicine. Cancer Discov. 2017, 7, 462–477. [Google Scholar] [CrossRef]
- Jabs, J.; Zickgraf, F.M.; Park, J.; Wagner, S.; Jiang, X.; Jechow, K.; Kleinheinz, K.; Toprak, U.H.; Schneider, M.A.; Meister, M.; et al. Screening drug effects in patient-derived cancer cells links organoid responses to genome alterations. Mol. Syst. Biol. 2017, 13, 955. [Google Scholar] [CrossRef]
- NIH. National Institutes of Health Office of Cancer Genomics (OCG) NIH. Available online: https://ocg.cancer.gov/programs/hcmi/overview (accessed on 5 November 2020).
- OcellO, B.V. Available online: https://ocello.nl/ (accessed on 29 November 2020).
- Crown Bioscience Inc. Available online: https://www.crownbio.com/ (accessed on 29 November 2020).
- Cellesce Ltd. Available online: https://cellesce.com/ (accessed on 29 November 2020).
| Author | Year | Tissue(s) | Cell Type | Stem Cell | Drug Screening | Biobanking | PDX | Relevant Assays/Treatments | Study Potential(s) | Reference |
|---|---|---|---|---|---|---|---|---|---|---|
| Eiraku | 2008 | Cortical neurons | N | E | na | na | na | na | na | [42] |
| Sato | 2009 | Small intestine | N | A | na | na | na | na | na | [19] |
| Eiraku | 2011 | Retinal epithelium | N | E | na | na | na | na | na | [43] |
| Jung | 2011 | Colon | N, T | A | na | na | na | na | na | [45] |
| Sato | 2011 | Colon | N, T | A | na | na | na | na | na | [44] |
| Lancaster | 2013 | Brain | N | E | na | na | na | na | na | [46] |
| Chua | 2014 | Prostate | N, T | A | na | na | na | na | Biobanking, Drug screening | [52] |
| Gao | 2014 | Prostate, Blood | M | A | PI3-kinase pathway inhibitors, Everolimus, BKM-120 | na | Y | na | Biobanking, Drug screening | [82] |
| Karthaus | 2014 | Prostate | N, T | A | na | na | Y | na | na | [51] |
| Bartfeld | 2015 | Stomach | N, T | A | na | na | na | H. pylori | Drug screening | [55] |
| Boj | 2015 | Pancreas | N, T | A | na | na | Y | na | na | [53] |
| Huang | 2015 | Pancreas | N, T | A | Histone methyltransferase EZH2 inhibitors | na | Y | Patient predictive screening | [75] | |
| Kessler | 2015 | Fallopian tube | N | A | na | na | na | na | na | [54] |
| van de Wetering | 2015 | Colon | N, T | A | 83 compound-library | Y | na | na | Drug screening | [83] |
| Weeber | 2015 | Colon | M | A | na | na | na | na | Drug screening | [84] |
| Fujii | 2016 | Colon | T | A | na | Y | Y | na | Drug screening | [85] |
| Hubert | 2016 | Brain | T | A | na | na | Y | na | Drug screening | [56] |
| Broutier | 2017 | Liver | T | A | Gemcitabine, Nutlin-3a, LGK974, ERK inhibitor (SCH772984) | na | Y | na | Drug screening | [58] |
| Turco | 2017 | Endometrium | N | A | na | na | na | na | Biobanking, Drug screening | [57] |
| Beshiri | 2018 | Prostate | T | A | Olaparib | Y | Y | na | na | [92] |
| Lampis | 2018 | Liver | T | A | 484 compound-library (new small-molecules) | Y | Y | na | na | [99] |
| Lee | 2018 | Bladder | T | A | 40 compound-library (ERK, MEK inhibitors) | Y | Y | na | Patient predictive screening | [89] |
| Roerink | 2018 | Colon | T | A | na | na | na | Single-cell sequencing | na | [76] |
| Sachs | 2018 | Breast | T, M | A | Afatinib, Gefitinib, Pictilisib, GDC-0068, AZD8055, Everolimus, Tamoxifen | Y | Y | Gene engineering CRISPR-cas9 | na | [87] |
| Seino | 2018 | Pancreas | T | A | na | Y | Y | Gene engineering CRISPR-cas9 | na | [86] |
| Tiriac | 2018 | Pancreas | T | A | Gemcitabine, Paclitaxel, Irinotecan, 5-Fluorouracil, Oxaliplatin | Y | na | na | Patient predictive screening | [90] |
| Vlachogiannis | 2018 | Stomach, Intestine | M | A | 55 compound-library (FDA approved) | Y | Y | na | Patient predictive screening | [88] |
| Yan | 2018 | Stomach | N, T, M | A | 37 compound-library (FDA approved and new) | Y | Y | na | na | [91] |
| Boretto | 2019 | Endometrium | N, T | A | Paclitaxel, 5-Fluorouracil, Carboplatin, Doxorubicin, Everolimus | Y | Y | na | na | [59] |
| Driehuis | 2019 | Oral mucosa | T | A | Nutlin-3, Niraparib, AZD4547, Everolimus, Vemurafenib, Alpelisib, Cisplatin, Carboplatin, Cetuximab | Y | Y | Herpes simplex virus, Human papillomavirus | Patient predictive screening | [93] |
| Kopper | 2019 | Ovary | N, T, M | A | Carboplatin, Paclitaxel, MK2206, AZD8055, Pictilisib, Alpelisib, Nutlin3a, Adavosertib, Gemcitabine, Niraparib | Y | Y | Gene engineering CRISPR-cas9 | na | [95] |
| Mullenders | 2019 | Bladder | T | A | Epirubicin, Mitomycin C, Gemcitabine, Vincristine, Doxorubicin, Cisplatin | Y | na | Gene engineering CRISPR-cas9 | Patient predictive screening | [94] |
| Ooft | 2019 | Colon | T, M | A | 5-Fluorouracil, Capecitabine, Irinotecan, Oxaliplatin | na | na | na | Patient predictive screening | [39] |
| Pasch | 2019 | Various | T | A | 5-Fluorouracil, Oxaliplatin, Ionizing radiation | na | na | na | Patient predictive screening | [77] |
| Sachs | 2019 | Lung, Broncho-alvolar lavage | N, T | A | Paclitaxel, Methotrexate, Crizotinib, Cisplatin, Nutlin-3a, Erlotinib, Alpelisib, Gefitinib | Y | Y | Respiratory syncytial virus | Drug screening | [61] |
| Schutgens | 2019 | Kidney, Urine | N, T | A | Cidofovir | na | na | BK virus, Organ-on-a-chip | Drug screening | [60] |
| Calandrini | 2020 | Kidney | N, T | A | Actinomycin D, Vincristine, Doxorubicin, Etoposide | Y | na | Single-cell sequencing | na | [98] |
| Jacob | 2020 | Brain | T | A | Ionizing radiation, Temozolomide, Gefitinib, Trametinib, Everolimus | Y | Y | CAR-T cells Co-culture | na | [97] |
| Nanki | 2020 | Ovary | T | A | 23 compound-library (FDA approved) | na | na | na | na | [78] |
| Narasimhan | 2020 | Colon | M | A | 87 compound pan-cancer library | na | Y | na | Patient predictive screening | [100] |
| Pleguezuelos-Manzano | 2020 | Colon | T | A | na | na | na | E. coli | na | [79] |
| Yao | 2020 | Rectum | T | A | Ionizing radiation, 5-Fluorouracil, and Irinotecan | Y | na | na | Patient predictive screening | [96] |
| Registry | Study ID | Origin | PDTOs Establishment | Drug Screening | Biobanking | Tumor Type | Enrollment | Location |
|---|---|---|---|---|---|---|---|---|
| Chinese CTR | ChiCTR1800016734 | S, B | Y | N | N | Stomach | 10 | China |
| Chinese CTR | ChiCTR1800017767 | S, B | Y | N | N | Ovary | 120 | China |
| Chinese CTR | ChiCTR2000034996 | S, B | Y | N | N | Esophagus | 50 | China |
| US NIH | NCT02436564 | S, B | Y | N | N | Liver, Pancreas | 75 | United Kingdom |
| US NIH | NCT02910895 | S, B | Y | N | N | Soft Tissue | 40 | Netherlands |
| US NIH | NCT03140592 | B | Y | N | N | Pancreas | 300 | USA |
| US NIH | NCT03952793 | Ext B (Metastasis) | Y | N | N | Prostate | 20 | France |
| US NIH | NCT03990675 | B | Y | N | N | Pancreas | 50 | Germany |
| US NIH | NCT04219137 | S, B | Y | N | N | Stomach, Esophagus | 120 | Canada |
| US NIH | NCT04342286 | S, B | Y | N | N | Kidney | 20 | China |
| US NIH | NCT04371198 | S, B | Y | N | N | Rectum | 20 | USA |
| US NIH | NCT04478877 | S | Y | N | N | Meningis | 30 | Hong Kong |
| Netherlands CTR | NL8956 | S, B | Y | N | N | Larynx | 20 | Netherlands |
| Netherlands CTR | NTR6150 | Blood Circ (Metastasis) | Y | N | N | Prostate | 46 | Netherlands |
| Netherlands CTR | NTR7286 | B | Y | N | N | Breast | 30 | Netherlands |
| Thailand CTR | TCTR20200827007 | B | Y | N | N | Pancreas | 28 | Thailand |
| Chinese CTR | ChiCTR1800018069 | S, B | Y | Y | N | Breast | 100 | China |
| Chinese CTR | ChiCTR1900023682 | S, B | Y | Y | N | Endometrium | 20 | China |
| Chinese CTR | ChiCTR1900027081 | S, B | Y | Y | N | Nasopharynx | 25 | China |
| Chinese CTR | ChiCTR1900028000 | B | Y | Y | N | Pancreas | 50 | China |
| Chinese CTR | ChiCTR2000028856 | S, B | Y | Y | N | Stomach | 59 | China |
| Chinese CTR | ChiCTR2000028889 | S, B | Y | Y | N | Stomach | 40 | China |
| Chinese CTR | ChiCTR2000029049 | S, B | Y | Y | N | Colon, Rectum | 50 | China |
| Chinese CTR | ChiCTR2000032765 | S, B | Y | Y | N | Bladder | 1500 | China |
| Chinese CTR | ChiCTR2000035441 | S, B | Y | Y | N | Breast | 30 | China |
| Chinese CTR | ChiCTR2000036347 | Ext B (Metastasis) | Y | Y | N | Prostate | 100 | China |
| Chinese CTR | ChiCTR2000037214 | B | Y | Y | N | Pancreas | 200 | China |
| Chinese CTR | ChiCTR2000037237 | Ext B (Metastasis) | Y | Y | N | Breast | 90 | China |
| Chinese CTR | ChiCTR-ONC-17011405 | S, Ext B (Metastasis) | Y | Y | N | Breast | 100 | China |
| Chinese CTR | ChiCTR-OOC-17012057 | B | Y | Y | N | Pancreas | 50 | China |
| Indian CTR | CTRI/2017/05/008512 | B | Y | Y | N | Pancreas | 20 | India |
| German CTR | DRKS00021088 | B | Y | Y | N | Pancreas | 118 | Germany |
| US NIH | NCT03283527 | B | Y | Y | N | Esophagus | 100 | Netherlands |
| US NIH | NCT03307538 | S, B | Y | Y | N | Bile duct | 6 | Netherlands |
| US NIH | NCT03429816 | B | Y | Y | N | Stomach, Esophagus | 40 | Germany |
| US NIH | NCT03453307 | S, B | Y | Y | N | Lung | 100 | China |
| US NIH | NCT03500068 | Ext B (Metastasis) | Y | Y | N | Pancreas | 30 | Netherlands |
| US NIH | NCT03544047 | S, B | Y | Y | N | Breast | 50 | China |
| US NIH | NCT03577808 | S, B | Y | Y | N | Rectum | 80 | China |
| US NIH | NCT03764553 | S, B | Y | Y | N | Esophagus | 310 | Netherlands |
| US NIH | NCT03890614 | B (Bone Marrow) | Y | Y | N | Myeloid cells | 40 | USA |
| US NIH | NCT03896958 | S, B | Y | Y | N | Various (solid) | 1000 | USA |
| US NIH | NCT03925233 | S, B | Y | Y | N | Breast | 300 | China |
| US NIH | NCT03979170 | S, B | Y | Y | N | Lung | 50 | Switzerland |
| US NIH | NCT04072445 | S, B | Y | Y | N | Bile duct | 28 | USA |
| US NIH | NCT04261192 | S, B | Y | Y | N | Head-neck | 98 | France |
| US NIH | NCT04278326 | S, B | Y | Y | N | Vagina, Cervix, Penis, Oropharynx | 50 | France |
| US NIH | NCT04279509 | B | Y | Y | N | Various (solid) | 35 | Singapore |
| US NIH | NCT04450706 | S, Ext B (Metastasis) | Y | Y | N | Breast | 15 | USA |
| US NIH | NCT04469556 | B | Y | Y | N | Pancreas | 150 | Canada |
| US NIH | NCT04526587 | S, B | Y | Y | N | Breast | 300 | USA |
| US NIH | NCT04555473 | B | Y | Y | N | Ovary | 48 | Italy |
| US NIH | NCT04561453 | S, B | Y | Y | N | Biliary tract | 20 | USA |
| US NIH | NCT04587128 | S, B | Y | Y | N | Colon, Rectum | 110 | USA |
| Netherlands CTR | NTR7521 | S, B | Y | Y | N | Colon | 150 | Netherlands |
| Chinese CTR | ChiCTR1800017855 | S, B | Y | Y | Y | Bones | 200 | China |
| Chinese CTR | ChiCTR1900024322 | S, B | Y | Y | Y | Nasopharynx | 160 | China |
| US NIH | NCT03544255 | S, B | Y | Y | Y | Pancreas | 50 | China |
| US NIH | NCT03655015 | S | Y | Y | Y | Lung | 50 | USA |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vivarelli, S.; Candido, S.; Caruso, G.; Falzone, L.; Libra, M. Patient-Derived Tumor Organoids for Drug Repositioning in Cancer Care: A Promising Approach in the Era of Tailored Treatment. Cancers 2020, 12, 3636. https://doi.org/10.3390/cancers12123636
Vivarelli S, Candido S, Caruso G, Falzone L, Libra M. Patient-Derived Tumor Organoids for Drug Repositioning in Cancer Care: A Promising Approach in the Era of Tailored Treatment. Cancers. 2020; 12(12):3636. https://doi.org/10.3390/cancers12123636
Chicago/Turabian StyleVivarelli, Silvia, Saverio Candido, Giuseppe Caruso, Luca Falzone, and Massimo Libra. 2020. "Patient-Derived Tumor Organoids for Drug Repositioning in Cancer Care: A Promising Approach in the Era of Tailored Treatment" Cancers 12, no. 12: 3636. https://doi.org/10.3390/cancers12123636
APA StyleVivarelli, S., Candido, S., Caruso, G., Falzone, L., & Libra, M. (2020). Patient-Derived Tumor Organoids for Drug Repositioning in Cancer Care: A Promising Approach in the Era of Tailored Treatment. Cancers, 12(12), 3636. https://doi.org/10.3390/cancers12123636









