The Diagnostic and Prognostic Value of a Liquid Biopsy for Esophageal Cancer: A Systematic Review and Meta-Analysis
Simple Summary
Abstract
1. Introduction
2. Results
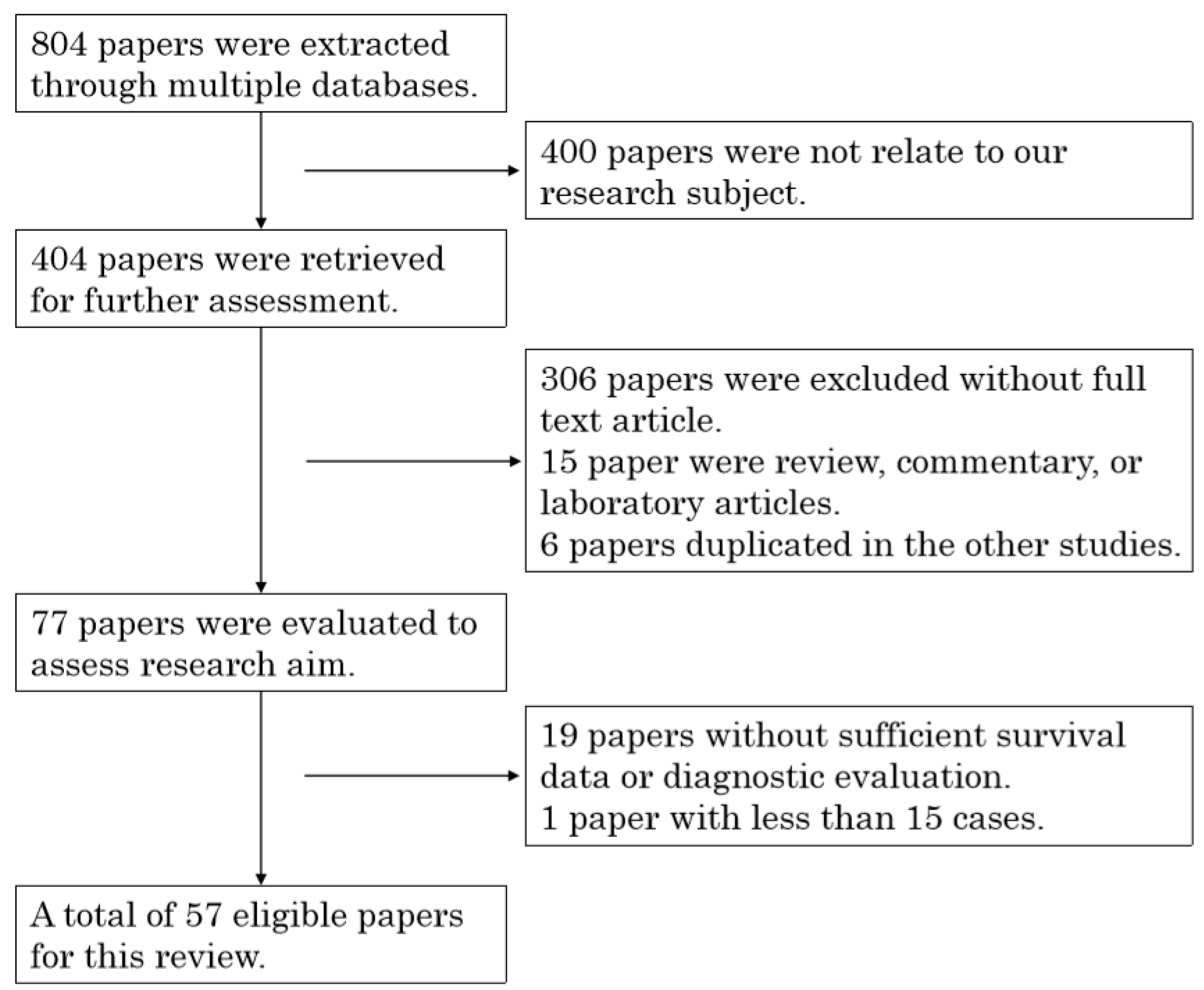
2.1. Literature Extraction
2.2. Summary of Different Technologies for Liquid Biopsy
2.3. Summary of Platforms for the Cytometric Isolation of CTCs
2.3.1. Immuno-Magnetic Technology
2.3.2. Filtration Technology
2.3.3. Microfluidic Technology
2.4. Summary of Platforms for the Non-Cytometric Isolation of CTCs
2.4.1. ELISA
2.4.2. RT-PCR
2.4.3. Non-Coding RNAs
2.4.4. Circulating Tumor DNA
2.4.5. LC–MS
2.5. Prognostic Value of CTC Identification
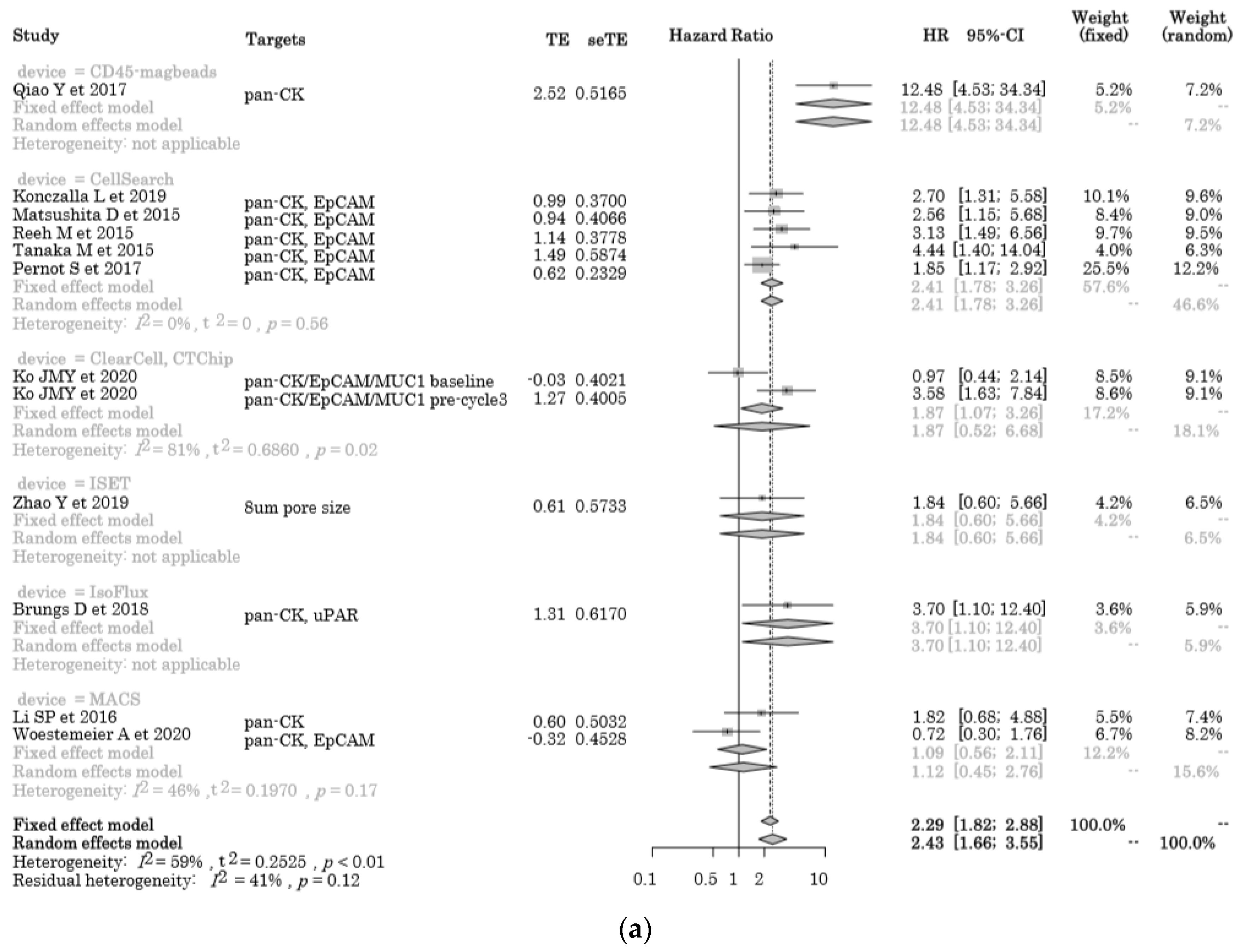
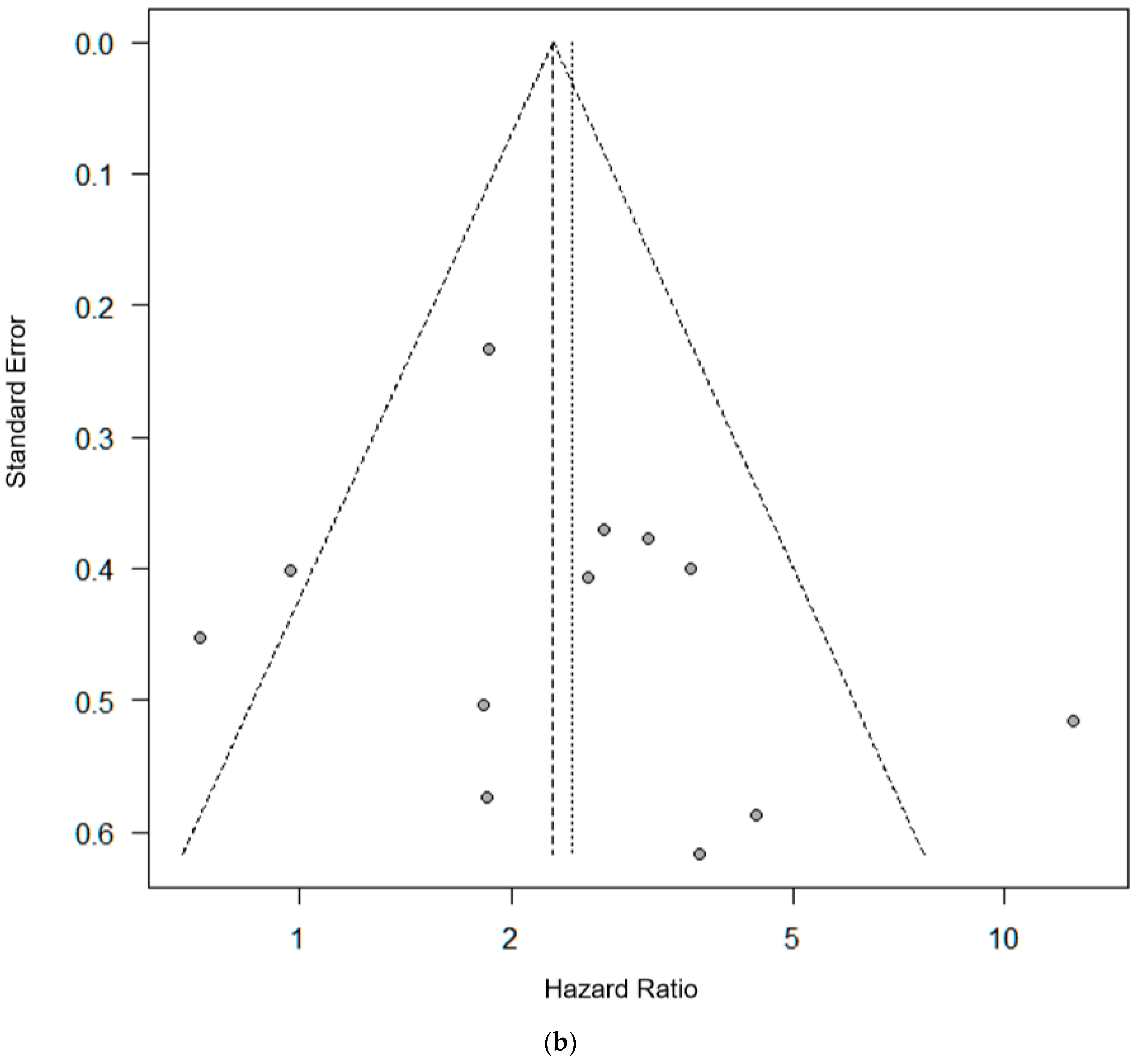
2.5.1. OS in the Cytometric Assay
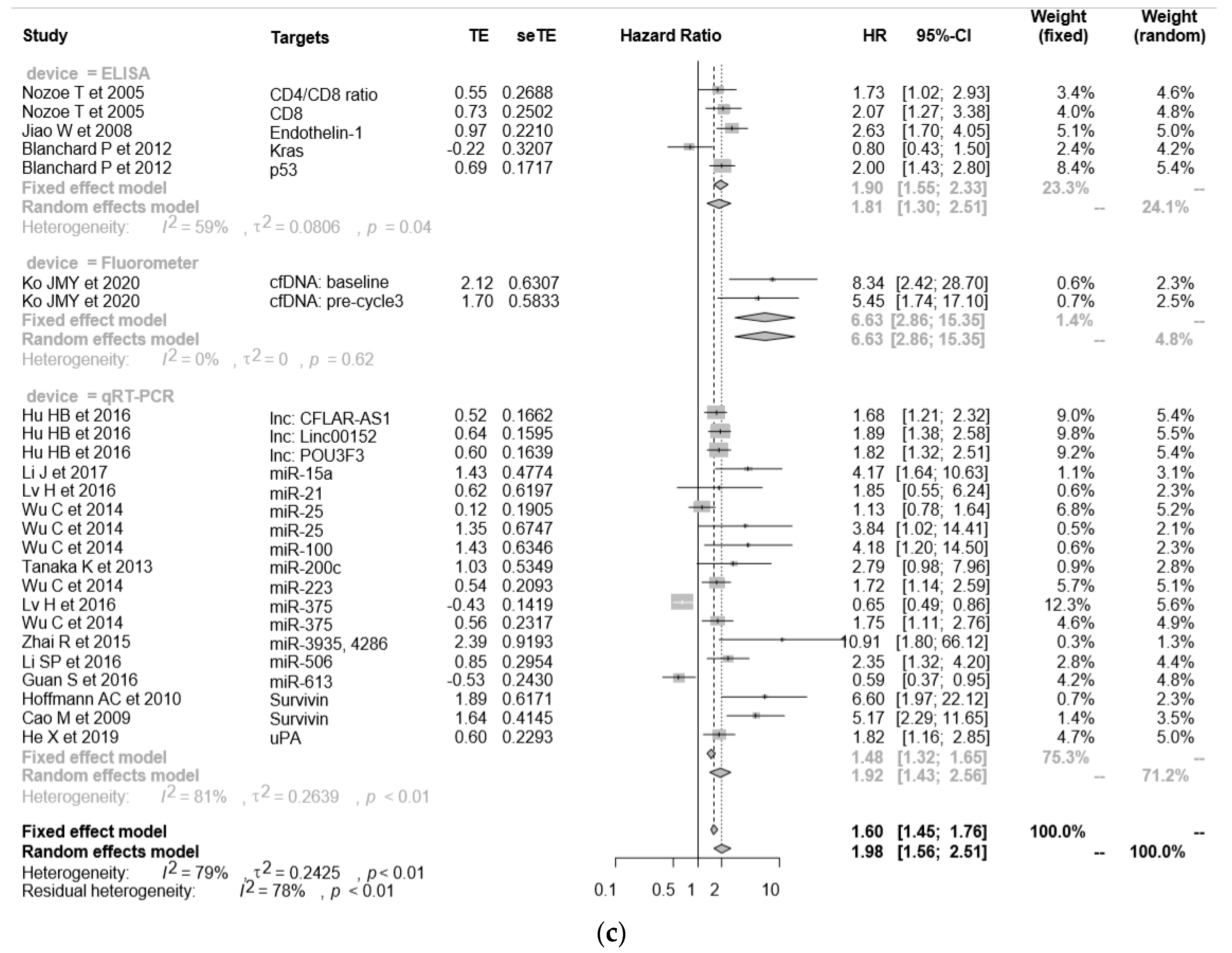
2.5.2. OS in the Non-Cytometric Assay
2.6. Progression-Free Survival (PFS) in the Cytometric Assay
2.7. PFS in the Non-Cytometric Assay
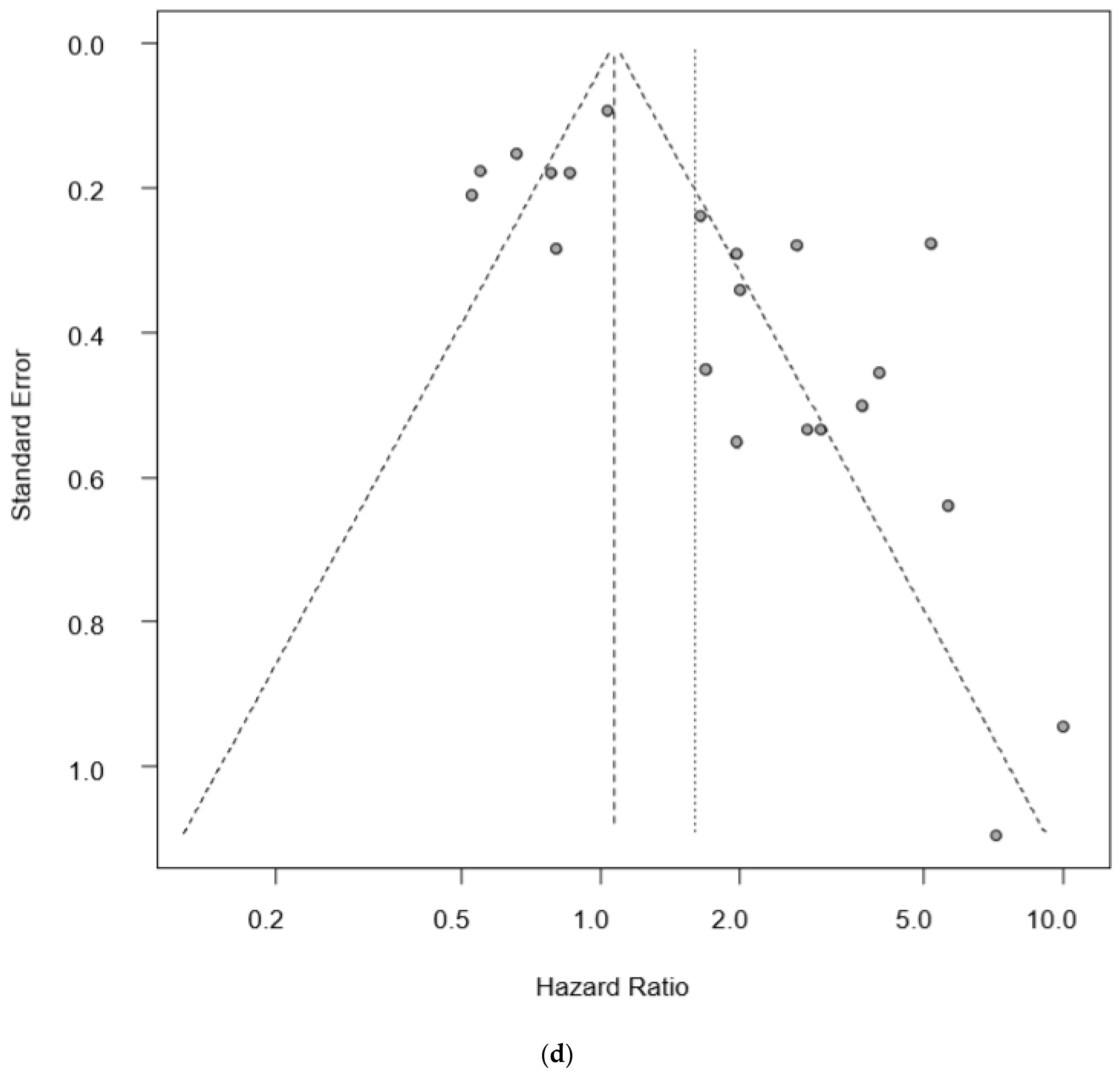
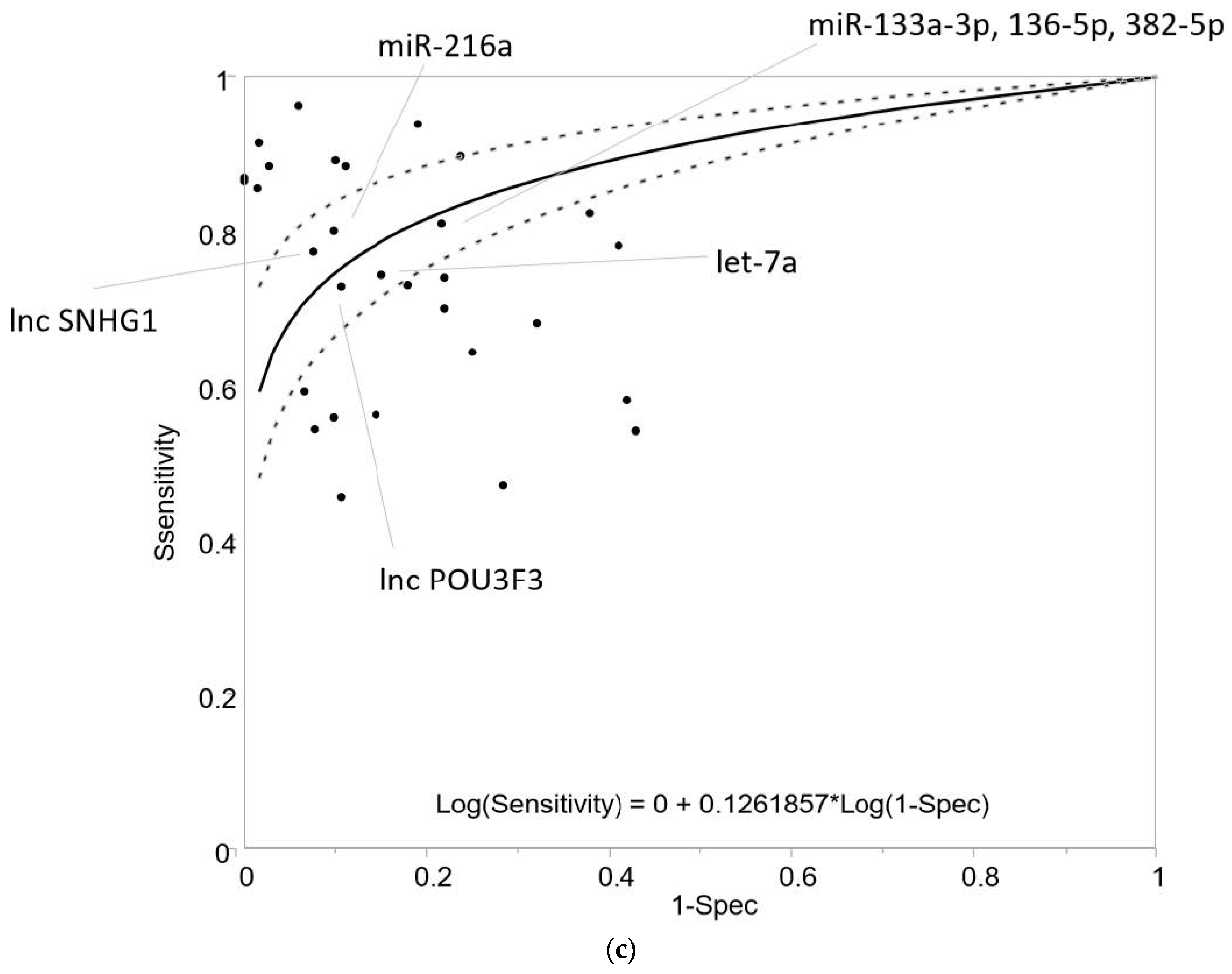
2.8. Early Diagnostic Value of “Liquid Biopsy”
3. Discussion
4. Materials and Methods
4.1. Literature Search Strategy
4.2. Inclusion and Exclusion Criteria
4.3. Data Extraction
4.4. Statistical Methods
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| CTC | Circulating tumor cell; |
| EC | Esophageal cancer; |
| EAC | Esophageal adenocarcinoma; |
| ESCC | Esophageal squamous cell carcinoma; |
| CT | Computed tomography; |
| PET | Positron emission tomography; |
| CEA | Carcinoembryonic antigen; |
| SCC | Squamous cell carcinoma antigen; |
| F-IHC | Fluorescence immunohistochemical staining; |
| PCR | Polymerase chain reaction; |
| qRT-PCR | Quantitative reverse transcription PCR; |
| NGS | Next-generation sequencing; |
| miRNA | Micro RNA; |
| FDA | Food and Drug Administration; |
| ISET | Isolation by size of epithelial tumor cells; |
| MACS | Magnetic cell separation system; |
| HR | Hazard ratio; |
| CI | Confidence interval; |
| ROC | Receiver operating characteristic; |
| AUC | Area under curve; |
| ELISA | Enzyme-linked immunosorbent assay; |
| LC–MS | Liquid chromatography–mass spectrometry; |
| CK | Cytokeratin; |
| EMT | Epithelial–mesenchymal transition; |
| EpCAM | Epithelial cell adhesion molecule; |
| DAPI | Dye 4, 6-diamidino-2-phenylidole dihydrochloride; |
| CTM | Circulating tumor microemboli; |
| FAST | Fluid-assisted separation technique; |
| ccfDNA | Circulating cell free DNA; |
| ctDNA | Circulating tumor DNA; |
| NMR | Nuclear magnetic resonance. |
References
- Arnold, M.; Soerjomataram, I.; Ferlay, J.; Forman, D. Global incidence of oesophageal cancer by histological subtype in 2012. Gut 2014, 64, 381–387. [Google Scholar] [CrossRef]
- Rice, T.W.; Apperson-Hansen, C.; DiPaola, L.M.; Semple, M.E.; Lerut, T.E.M.R.; Orringer, M.B.; Chen, L.-Q.; Hofstetter, W.L.; Smithers, B.M.; Rusch, V.W.; et al. Worldwide Esophageal Cancer Collaboration: Clinical staging data. Dis. Esophagus 2016, 29, 707–714. [Google Scholar] [CrossRef] [PubMed]
- Global Burden of Disease Cancer Collaboration; Fitzmaurice, C.; Allen, C.; Barber, R.M.; Barregard, L.; Bhutta, Z.A.; Brenner, H.; Dicker, D.J.; Chimed-Orchir, O.; Dandona, R.; et al. Global, Regional, and National Cancer Incidence, Mortality, Years of Life Lost, Years Lived With Disability, and Disability-Adjusted Life-years for 32 Cancer Groups, 1990 to 2015. JAMA Oncol. 2017, 3, 524–548. [Google Scholar] [CrossRef]
- Kosugi, S.-I.; Nishimaki, T.; Kanda, T.; Nakagawa, S.; Ohashi, M.; Hatakeyama, K. Clinical Significance of Serum Carcinoembryonic Antigen, Carbohydrate Antigen 19-9, and Squamous Cell Carcinoma Antigen Levels in Esophageal Cancer Patients. World J. Surg. 2004, 28, 680–685. [Google Scholar] [CrossRef] [PubMed]
- Ashworth, T.R. A case of cancer in which cells similar to those in the tumors were seen in the blood after death. Aust. Med. J. 1869, 14, 146–149. [Google Scholar]
- Allard, W.J. Tumor Cells Circulate in the Peripheral Blood of All Major Carcinomas but not in Healthy Subjects or Patients with Nonmalignant Diseases. Clin. Cancer Res. 2004, 10, 6897–6904. [Google Scholar] [CrossRef]
- Nakashima, S.; Natsugoe, S.; Matsumoto, M.; Miyazono, F.; Nakajo, A.; Uchikura, K.; Tokuda, K.; Ishigami, S.; Baba, M.; Takao, S.; et al. Clinical significance of circulating tumor cells in blood by molecular detection and tumor markers in esophageal cancer. Surgery 2003, 133, 162–169. [Google Scholar] [CrossRef] [PubMed]
- Kaganoi, J.; Shimada, Y.; Kano, M.; Okumura, T.; Watanabe, G.; Imamura, M. Detection of circulating oesophageal squamous cancer cells in peripheral blood and its impact on prognosis. Br. J. Surg. 2004, 91, 1055–1060. [Google Scholar] [CrossRef]
- Liu, Z.; Jiang, M.; Yan, F.; Xu, L.; Zhong, S.-L.; Ju, H. Multipoint quantification of multimarker genes in peripheral blood and micrometastasis characteristic in peri-operative esophageal cancer patients. Cancer Lett. 2008, 261, 46–54. [Google Scholar] [CrossRef]
- Cristofanilli, M.; Budd, G.T.; Ellis, M.J.; Stopeck, A.; Matera, J.; Miller, M.C.; Reuben, J.M.; Doyle, G.V.; Allard, W.J.; Terstappen, L.W.; et al. Circulating Tumor Cells, Disease Progression, and Survival in Metastatic Breast Cancer. N. Engl. J. Med. 2004, 351, 781–791. [Google Scholar] [CrossRef]
- Cohen, S.J.; Punt, C.J.A.; Iannotti, N.; Saidman, B.H.; Sabbath, K.D.; Gabrail, N.Y.; Picus, J.; Morse, M.; Mitchell, E.; Miller, M.C.; et al. Relationship of Circulating Tumor Cells to Tumor Response, Progression-Free Survival, and Overall Survival in Patients With Metastatic Colorectal Cancer. J. Clin. Oncol. 2008, 26, 3213–3221. [Google Scholar] [CrossRef] [PubMed]
- De Bono, J.S.; Scher, H.I.; Montgomery, R.B.; Parker, C.; Miller, M.C.; Tissing, H.; Doyle, G.; Terstappen, L.W.; Pienta, K.J.; Raghavan, D. Circulating Tumor Cells Predict Survival Benefit from Treatment in Metastatic Castration-Resistant Prostate Cancer. Clin. Cancer Res. 2008, 14, 6302–6309. [Google Scholar] [CrossRef] [PubMed]
- Matsushita, D.; Uenosono, Y.; Arigami, T.; Yanagita, S.; Nishizono, Y.; Hagihara, T.; Hirata, M.; Haraguchi, N.; Arima, H.; Kijima, Y.; et al. Clinical Significance of Circulating Tumor Cells in Peripheral Blood of Patients with Esophageal Squamous Cell Carcinoma. Ann. Surg. Oncol. 2015, 22, 3674–3680. [Google Scholar] [CrossRef]
- Vona, G.; Sabile, A.; Louha, M.; Sitruk, V.; Romana, S.; Schütze, K.; Capron, F.; Franco, D.; Pazzagli, M.; Vekemans, M.; et al. Isolation by Size of Epithelial Tumor Cells. Am. J. Pathol. 2000, 156, 57–63. [Google Scholar] [CrossRef]
- DeSitter, I.; Guerrouahen, B.S.; Benali-Furet, N.; Wechsler, J.; Jänne, P.A.; Kuang, Y.; Yanagita, M.; Wang, L.; Berkowitz, J.A.; Distel, R.J.; et al. A new device for rapid isolation by size and characterization of rare circulating tumor cells. Anticancer Res. 2011, 31, 427–441. [Google Scholar]
- Bobek, V.; Matkowski, R.; Gürlich, R.; Grabowski, K.; Szelachowska, J.; Lischke, R.; Schutzner, J.; Harustiak, T.; Pazdro, A.; Rzechonek, A.; et al. Cultivation of circulating tumor cells in esophageal cancer. Folia Histochem. Cytobiol. 2014, 52, 171–177. [Google Scholar] [CrossRef]
- Miltenyi, S.; Müller, W.; Weichel, W.; Radbruch, A. High gradient magnetic cell separation with MACS. Cytometry 1990, 11, 231–238. [Google Scholar] [CrossRef]
- Nagrath, S.; Sequist, L.V.; Maheswaran, S.; Bell, D.W.; Irimia, D.; Ulkus, L.; Smith, M.R.; Kwak, E.L.; Digumarthy, S.R.; Muzikansky, A.; et al. Isolation of rare circulating tumour cells in cancer patients by microchip technology. Nat. Cell Biol. 2007, 450, 1235–1239. [Google Scholar] [CrossRef]
- Harb, W.; Fan, A.; Tran, T.; Danila, D.C.; Keys, D.; Schwartz, M.; Ionescu-Zanetti, C. Mutational Analysis of Circulating Tumor Cells Using a Novel Microfluidic Collection Device and qPCR Assay. Transl. Oncol. 2013, 6, 528–538. [Google Scholar] [CrossRef] [PubMed]
- He, W.; Kularatne, S.A.; Kalli, K.R.; Prendergast, F.G.; Amato, R.J.; Klee, G.G.; Hartmann, L.C.; Low, P.S. Quantitation of circulating tumor cells in blood samples from ovarian and prostate cancer patients using tumor-specific fluorescent ligands. Int. J. Cancer 2008, 123, 1968–1973. [Google Scholar] [CrossRef]
- Zhu, Z.; Qiu, S.; Shao, K.; Hou, Y. Progress and challenges of sequencing and analyzing circulating tumor cells. Cell Biol. Toxicol. 2017, 34, 405–415. [Google Scholar] [CrossRef]
- Onidani, K.; Shoji, H.; Kakizaki, T.; Yoshimoto, S.; Okaya, S.; Miura, N.; Sekikawa, S.; Furuta, K.; Lim, C.T.; Shibahara, T.; et al. Monitoring of cancer patients via next-generation sequencing of patient-derived circulating tumor cells and tumor DNA. Cancer Sci. 2019, 110, 2590–2599. [Google Scholar] [CrossRef]
- Yu, J.; Leung, W.K.; Ebert, M.P.A.; Ng, E.K.W.; Go, M.Y.Y.; Wang, H.B.; Chung, S.C.S.; Malfertheiner, P.; Sung, J.J. Increased expression of survivin in gastric cancer patients and in first degree relatives. Br. J. Cancer 2002, 87, 91–97. [Google Scholar] [CrossRef]
- Hiraiwa, K.; Takeuchi, H.; Hasegawa, H.; Saikawa, Y.; Suda, K.; Ando, T.; Kumagai, K.; Irino, T.; Yoshikawa, T.; Matsuda, S.; et al. Clinical Significance of Circulating Tumor Cells in Blood from Patients with Gastrointestinal Cancers. Ann. Surg. Oncol. 2008, 15, 3092–3100. [Google Scholar] [CrossRef]
- Konczalla, L.; Ghadban, T.; Effenberger, K.E.; Wöstemeier, A.; Riethdorf, S.; Uzunoglu, F.G.; Izbicki, J.R.; Pantel, K.; Bockhorn, M.; Reeh, M. Prospective Comparison of the Prognostic Relevance of Circulating Tumor Cells in Blood and Disseminated Tumor Cells in Bone Marrow of a Single Patient’s Cohort with Esophageal Cancer. Ann. Surg. 2019. [Google Scholar] [CrossRef] [PubMed]
- Reeh, M.; Harms-Effenberger, K.; Koenig, A.M.; Riethdorf, S.; Eichstädt, D.; Vettorazzi, E.; Uzunoglu, F.G.; Vashist, Y.K.; Izbicki, J.R.; Pantel, K.; et al. Circulating Tumor Cells as a Biomarker for Preoperative Prognostic Staging in Patients with Esophageal Cancer. Ann. Surg. 2015, 261, 1124–1130. [Google Scholar] [CrossRef] [PubMed]
- Qiao, Y.; Li, J.; Shi, C.; Wang, W.; Qu, X.; Xiong, M.; Sun, Y.; Li, D.; Zhao, X.; Zhang, D. Prognostic value of circulating tumor cells in the peripheral blood of patients with esophageal squamous cell carcinoma. Onco Targets Ther. 2017, 10, 1363–1373. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Ma, G.; Zhao, P.; Fu, R.; Gao, L.; Jiang, X.; Hu, P.; Ren, T.; Wu, Y.; Wang, Z.; et al. Improvement of sensitive and specific detection of circulating tumor cells using negative enrichment and immunostaining-FISH. Clin. Chim. Acta 2018, 485, 95–102. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Li, J.; Wang, L.; Meng, P.; Zhao, J.; Han, P.; Xia, J.; Xu, J.; Wang, L.; Shen, F.; et al. Clinical significance of detecting circulating tumor cells in patients with esophageal squamous cell carcinoma by EpCAM-independent enrichment and immunostaining-fluorescence in situ hybridization. Mol. Med. Rep. 2019, 20, 1551–1560. [Google Scholar] [CrossRef]
- Baccelli, I.; Schneeweiss, A.; Riethdorf, S.; Stenzinger, A.; Schillert, A.; Vogel, V.; Klein, C.; Saini, M.; Bäuerle, T.; Wallwiener, M.; et al. Identification of a population of blood circulating tumor cells from breast cancer patients that initiates metastasis in a xenograft assay. Nat. Biotechnol. 2013, 31, 539–544. [Google Scholar] [CrossRef]
- Sollier, E.; Go, D.E.; Che, J.; Gossett, D.R.; O’Byrne, S.; Weaver, W.M.; Kummer, N.; Rettig, M.; Goldman, J.; Nickols, N.; et al. Size-selective collection of circulating tumor cells using Vortex technology. Lab Chip 2014, 14, 63–77. [Google Scholar] [CrossRef]
- Zhao, Y.; Zhao, S.; Chen, Y.; Dong, X.; Peng, C.; Sun, Q.; Shan, L.; Wang, Z.; Zhao, X. Isolation of circulating tumor cells in patients undergoing surgery for esophageal cancer and a specific confirmation method. Oncol. Lett. 2019, 4, 3817–3825. [Google Scholar] [CrossRef]
- Ohnaga, T.; Shimada, Y.; Takata, K.; Obata, T.; Okumura, T.; Nagata, T.; Kishi, H.; Muraguchi, A.; Tsukada, K. Capture of esophageal and breast cancer cells with polymeric microfluidic devices for CTC isolation. Mol. Clin. Oncol. 2016, 4, 599–602. [Google Scholar] [CrossRef]
- Chudasama, D.Y.; Freydina, D.V.; Freidin, M.B.; Leung, M.; Fernandez, A.M.; Rice, A.; Nicholson, A.G.; Karteris, E.; Anikin, V.; Lim, E. Inertia based microfluidic capture and characterisation of circulating tumour cells for the diagnosis of lung cancer. Ann. Transl. Med. 2016, 4, 480. [Google Scholar] [CrossRef]
- Karabacak, N.M.; Spuhler, P.S.; Fachin, F.; Lim, E.J.; Pai, V.; Ozkumur, E.; Martel, J.M.; Kojic, N.; Smith, B.A.H.; Chen, P.-I.; et al. Microfluidic, marker-free isolation of circulating tumor cells from blood samples. Nat. Protoc. 2014, 9, 694–710. [Google Scholar] [CrossRef]
- Earhart, C.M.; Hughes, C.E.; Gaster, R.S.; Ooi, C.C.; Wilson, R.J.; Zhou, L.Y.; Humke, E.W.; Xu, L.; Wong, D.J.; Willingham, S.B.; et al. Isolation and mutational analysis of circulating tumor cells from lung cancer patients with magnetic sifters and biochips. Lab Chip 2014, 14, 78–88. [Google Scholar] [CrossRef]
- Choi, M.K.; Kim, G.H.; I, H.; Park, S.J.; Lee, M.W.; Lee, B.E.; Park, D.Y.; Cho, Y.-K. Circulating tumor cells detected using fluid-assisted separation technique in esophageal squamous cell carcinoma. J. Gastroenterol. Hepatol. 2018, 34, 552–560. [Google Scholar] [CrossRef]
- Ismail, M.M.; James, C.; Webbe, G. The enzyme linked immunosorbent assay (ELISA) for the determination of circulating antigen and antibody in Schistosoma haematobium-infected baboons. Trans. R. Soc. Trop. Med. Hyg. 1981, 75, 542–548. [Google Scholar] [CrossRef]
- Nozoe, T.; Maehara, Y.; Sugimachi, K. Preoperative sorting of circulating T lymphocytes in patients with esophageal squamous cell carcinoma: Its prognostic significance. World J. Gastroenterol. 2005, 11, 6689–6693. [Google Scholar] [CrossRef]
- Kimura, H.; Kato, H.; Tanaka, N.; Inose, T.; Faried, A.; Sohda, M.; Nakajima, M.; Fukai, Y.; Miyazaki, T.; Masuda, N.; et al. Preoperative serum vascular endothelial growth factor-C (VEGF-C) levels predict recurrence in patients with esophageal cancer. Anticancer Res. 2008, 28, 165–169. [Google Scholar]
- Curran, M.A.; Montalvo, W.; Yagita, H.; Allison, J.P. PD-1 and CTLA-4 combination blockade expands infiltrating T cells and reduces regulatory T and myeloid cells within B16 melanoma tumors. Proc. Natl. Acad. Sci. USA 2010, 107, 4275–4280. [Google Scholar] [CrossRef]
- Setoyama, T.; Natsugoe, S.; Okumura, H.; Matsumoto, M.; Uchikado, Y.; Ishigami, S.; Owaki, T.; Takao, S.; Aikou, T. Carcinoembryonic Antigen Messenger RNA Expression in Blood Predicts Recurrence in Esophageal Cancer. Clin. Cancer Res. 2006, 12, 5972–5977. [Google Scholar] [CrossRef]
- Srinivasula, S.M.; Ashwell, J.D. IAPs: What’s in a Name? Mol. Cell 2008, 30, 123–135. [Google Scholar] [CrossRef]
- Hoffmann, A.-C.; Vallböhmer, D.; Grimminger, P.P.; Metzger, R.; Prenzel, K.L.; Hoelscher, A.H.; Brabender, J. Preoperative survivin mRNA detection in peripheral blood is an independent predictor of outcome in esophageal carcinoma. Pharmacogenomics 2010, 11, 341–347. [Google Scholar] [CrossRef]
- Lambrechts, A.; Bosma, A.; Klaver, S.; Top, B.; Perebolte, L.; Veer, L.J.V.T.; Rodenhuis, S. Comparison of immunocytochemistry, reverse transcriptase polymerase chain reaction, and nucleic acid sequence–based amplification for the detection of circulating breast cancer cells. Breast Cancer Res. Treat. 1999, 56, 217–229. [Google Scholar] [CrossRef]
- Mitchell, P.S.; Parkin, R.K.; Kroh, E.M.; Fritz, B.R.; Wyman, S.K.; Pogosova-Agadjanyan, E.L.; Peterson, A.; Noteboom, J.; O’Briant, K.C.; Allen, A.; et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. USA 2008, 105, 10513–10518. [Google Scholar] [CrossRef]
- Li, J.; Li, M.; Gao, F.; Ge, X. Serum microRNA-15a level acts as a potential diagnostic and prognostic biomarker for human esophageal squamous cell carcinoma. Cancer Biomark. 2017, 18, 11–17. [Google Scholar] [CrossRef]
- Lv, H.; He, Z.; Wang, H.; Du, T.; Pang, Z. Differential expression of miR-21 and miR-75 in esophageal carcinoma patients and its clinical implication. Am. J. Transl. Res. 2016, 8, 3288–3298. [Google Scholar]
- Komatsu, S.; Ichikawa, D.; Kawaguchi, T.; Miyamae, M.; Okajima, W.; Ohashi, T.; Imamura, T.; Kiuchi, J.; Konishi, H.; Shiozaki, A.; et al. Circulating miR-21 as an independent predictive biomarker for chemoresistance in esophageal squamous cell carcinoma. Am. J. Cancer Res. 2016, 6, 1511–1523. [Google Scholar]
- Wu, C.; Li, M.; Hu, C.; Duan, H. Clinical significance of serum miR-223, miR-25 and miR-375 in patients with esophageal squamous cell carcinoma. Mol. Biol. Rep. 2014, 41, 1257–1266. [Google Scholar] [CrossRef]
- Gu, J.; Zhang, J.; Zheng, L.; Ajani, J.A.; Wu, X.; Ye, Y. Serum miR-331-3p predicts tumor recurrence in esophageal adenocarcinoma. Sci. Rep. 2018, 8, 14006. [Google Scholar] [CrossRef]
- Tanaka, K.; Miyata, H.; Yamasaki, M.; Sugimura, K.; Takahashi, T.; Kurokawa, Y.; Nakajima, K.; Takiguchi, S.; Mori, M.; Doki, Y. Circulating miR-200c Levels Significantly Predict Response to Chemotherapy and Prognosis of Patients Undergoing Neoadjuvant Chemotherapy for Esophageal Cancer. Ann. Surg. Oncol. 2013, 20, 607–615. [Google Scholar] [CrossRef]
- Zhai, R.; Wei, Y.; Su, L.; Liu, G.; Kulke, M.H.; Wain, J.C.; Christiani, D.C. Whole-miRNome profiling identifies prognostic serum miRNAs in esophageal adenocarcinoma: The influence of Helicobacter pylori infection status. Carcinogenesis 2014, 36, 87–93. [Google Scholar] [CrossRef]
- Li, S.-P.; Su, H.-X.; Zhao, D.; Guan, Q.-L. Plasma miRNA-506 as a Prognostic Biomarker for Esophageal Squamous Cell Carcinoma. Med. Sci. Monit. 2016, 22, 2195–2201. [Google Scholar] [CrossRef][Green Version]
- Guan, S.; Wang, C.; Chen, X.; Liu, B.; Tan, B.; Liu, F.; Wang, D.; Han, L.; Wang, L.; Huang, X.; et al. MiR-613: A novel diagnostic and prognostic biomarker for patients with esophageal squamous cell carcinoma. Tumor Biol. 2015, 37, 4383–4391. [Google Scholar] [CrossRef]
- He, F.; Meng, W.-W.; Qu, Y.-H.; Zhou, M.-X.; He, J.; Lv, P.; Ming, L. Expression of circulating microRNA-20a and let-7a in esophageal squamous cell carcinoma. World J. Gastroenterol. 2015, 21, 4660–4665. [Google Scholar] [CrossRef]
- Cui, Y.; Xue, Y.; Dong, S.; Zhang, P. Plasma microRNA-9 as a diagnostic and prognostic biomarker in patients with esophageal squamous cell carcinoma. J. Int. Med. Res. 2017, 45, 1310–1317. [Google Scholar] [CrossRef]
- Shen, Y.; Ding, Y.; Ma, Q.; Zhao, L.; Guo, X.; Shao, Y.; Niu, C.; He, Y.; Zhang, F.; Zheng, D.; et al. Identification of Novel Circulating miRNA Biomarkers for the Diagnosis of Esophageal Squamous Cell Carcinoma and Squamous Dysplasia. Cancer Epidemiol. Biomark. Prev. 2019, 28, 1212–1220. [Google Scholar] [CrossRef]
- Chiam, K.; Wang, T.; Watson, D.I.; Mayne, G.C.; Irvine, T.S.; Bright, T.F.; Smith, L.; White, I.A.; Bowen, J.M.; Keefe, D.; et al. Circulating Serum Exosomal miRNAs As Potential Biomarkers for Esophageal Adenocarcinoma. J. Gastrointest. Surg. 2015, 19, 1208–1215. [Google Scholar] [CrossRef]
- Zheng, D.; Ding, Y.; Ma, Q.; Zhao, L.; Guo, X.; Shen, Y.; He, Y.; Wei, W.; Liu, F. Identification of Serum MicroRNAs as Novel Biomarkers in Esophageal Squamous Cell Carcinoma Using Feature Selection Algorithms. Front. Oncol. 2019, 8, 674. [Google Scholar] [CrossRef]
- Hirajima, S.; Komatsu, S.; Ichikawa, D.; Takeshita, H.; Konishi, H.; Shiozaki, A.; Morimura, R.; Tsujiura, M.; Nagata, H.; Kawaguchi, T.; et al. Clinical impact of circulating miR-18a in plasma of patients with oesophageal squamous cell carcinoma. Br. J. Cancer 2013, 108, 1822–1829. [Google Scholar] [CrossRef]
- Zhang, L.; Dong, B.; Ren, P.; Ye, H.; Shi, J.; Qin, J.; Wang, K.; Wang, P.; Zhang, J. Circulating plasma microRNAs in the detection of esophageal squamous cell carcinoma. Oncol. Lett. 2018, 16, 3303–3318. [Google Scholar] [CrossRef] [PubMed]
- Cheng, L.; Huang, Z.; Lv, H.; Wang, Y.; Sun, W.; Sun, X. Up-Regulation of MicroRNA-21 Indicates Poor Prognosis and Promotes Cell Proliferation in Esophageal Squamous Cell Carcinoma via Upregulation of lncRNA SNHG1. Cancer Manag. Res. 2020, 12, 1–14. [Google Scholar] [CrossRef]
- Ibuki, Y.; Nishiyama, Y.; Tsutani, Y.; Emi, M.; Hamai, Y.; Okada, M.; Tahara, H. Circulating microRNA/isomiRs as novel biomarkers of esophageal squamous cell carcinoma. PLoS ONE 2020, 15, e0231116. [Google Scholar] [CrossRef]
- Bus, P.; Kestens, C.; Kate, F.J.W.T.; Peters, W.; Drenth, J.P.H.; Roodhart, J.M.L.; Siersema, P.D.; Van Baal, J.W.P.M. Profiling of circulating microRNAs in patients with Barrett’s esophagus and esophageal adenocarcinoma. J. Gastroenterol. 2015, 51, 560–570. [Google Scholar] [CrossRef] [PubMed]
- Pavlov, K.; Kluiver, J.; Meijer, C.; Ek, W.B.-V.; Kruyt, F.A.E.; Karrenbeld, A.; Kleibeuker, J.H.; Peters, F.T.M.; Berg, A.V.D. Circulating miRNAs in patients with Barrett’s esophagus, high-grade dysplasia and esophageal adenocarcinoma. J. Gastrointest. Oncol. 2018, 9, 1150–1156. [Google Scholar] [CrossRef]
- Dong, S.; Yin, H.; Dong, C.; Sun, K.; Lv, P.; Meng, W.; Ming, L.; He, F. Predictive Value of Plasma MicroRNA-216a/b in the Diagnosis of Esophageal Squamous Cell Carcinoma. Dis. Mark. 2016, 2016, 1857067. [Google Scholar] [CrossRef]
- Huang, Z.; Zhang, L.; Zhu, D.; Shan, X.; Zhou, X.; Qi, L.-W.; Wu, L.; Zhu, J.; Cheng, W.; Zhang, H.; et al. A novel serum microRNA signature to screen esophageal squamous cell carcinoma. Cancer Med. 2016, 6, 109–119. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.; Wu, X.; Wang, J.; Lopez, J.; Zhou, W.; Yang, L.; Wang, S.E.; Raz, D.J.; Kim, J.Y. Circulating miRNA profile in esophageal adenocarcinoma. Am. J. Cancer Res. 2016, 6, 2713–2721. [Google Scholar]
- Perkel, J.M. Visiting “Noncodarnia”. Biotechniques 2013, 54, 301–304. [Google Scholar] [CrossRef]
- Qi, P.; Du, X. The long non-coding RNAs, a new cancer diagnostic and therapeutic gold mine. Mod. Pathol. 2012, 26, 155–165. [Google Scholar] [CrossRef] [PubMed]
- Tong-Xin, Y.; Wang, X.-W.; Zhou, X.-L.; Liu, Z.-H.; Yang, T.-X.; Shi, W.-H.; Xie, H.-W.; Lv, J.; Wu, Q.-Q.; Cao, X.-F. Identification of the long non-coding RNA POU3F3 in plasma as a novel biomarker for diagnosis of esophageal squamous cell carcinoma. Mol. Cancer 2015, 14, 3. [Google Scholar] [CrossRef] [PubMed]
- Hu, H.-B.; Jie, H.-Y.; Zheng, X.-X. Three Circulating LncRNA Predict Early Progress of Esophageal Squamous Cell Carcinoma. Cell. Physiol. Biochem. 2016, 40, 117–125. [Google Scholar] [CrossRef] [PubMed]
- Mandel, P.; Metais, P. Les acides nucléiques du plasma sanguin chez l’homme. CR Seances Soc. Biol. Fil. 1948, 142, 241. [Google Scholar]
- Ko, J.M.Y.; Ng, H.Y.; Lam, K.-O.; Chiu, K.W.-H.; Kwong, D.L.W.; Lo, A.W.; Wong, J.C.; Lin, R.C.W.; Fong, H.C.H.; Li, J.Y.K.; et al. Liquid Biopsy Serial Monitoring of Treatment Responses and Relapse in Advanced Esophageal Squamous Cell Carcinoma. Cancers 2020, 12, 1352. [Google Scholar] [CrossRef]
- Forshew, T.; Murtaza, M.; Parkinson, C.; Gale, D.; Tsui, D.W.Y.; Kaper, F.; Dawson, S.-J.; Piskorz, A.M.; Jimenez-Linan, M.; Bentley, D.; et al. Noninvasive Identification and Monitoring of Cancer Mutations by Targeted Deep Sequencing of Plasma DNA. Sci. Transl. Med. 2012, 4, 136ra68. [Google Scholar] [CrossRef]
- Grölz, D.; Hauch, S.; Schlumpberger, M.; Guenther, K.; Voss, T.; Sprenger-Haussels, M.; Oelmüller, U. Liquid Biopsy Preservation Solutions for Standardized Pre-Analytical Workflows—Venous Whole Blood and Plasma. Curr. Pathobiol. Rep. 2018, 6, 275–286. [Google Scholar] [CrossRef]
- Zhang, J.; Bowers, J.; Liu, L.; Wei, S.; Gowda, G.A.N.; Hammoud, Z.; Raftery, D. Esophageal Cancer Metabolite Biomarkers Detected by LC-MS and NMR Methods. PLoS ONE 2012, 7, e30181. [Google Scholar] [CrossRef]
- Xu, J.; Chen, Y.; Zhang, R.; He, J.; Song, Y.; Wang, J.; Wang, H.; Wang, L.; Zhan, Q.; Abliz, Z. Global metabolomics reveals potential urinary biomarkers of esophageal squamous cell carcinoma for diagnosis and staging. Sci. Rep. 2016, 6, 35010. [Google Scholar] [CrossRef]
- Zhu, Z.-J.; Qi, Z.; Zhang, J.; Xue, W.-H.; Li, L.-F.; Shen, Z.-B.; Li, Z.-Y.; Yuan, Y.-L.; Wang, W.-B.; Zhao, J. Untargeted Metabolomics Analysis of Esophageal Squamous Cell Carcinoma Discovers Dysregulated Metabolic Pathways and Potential Diagnostic Biomarkers. J. Cancer 2020, 11, 3944–3954. [Google Scholar] [CrossRef]
- Aretz, I.; Meierhofer, D. Advantages and Pitfalls of Mass Spectrometry Based Metabolome Profiling in Systems Biology. Int. J. Mol. Sci. 2016, 17, 632. [Google Scholar] [CrossRef]
- Woestemeier, A.; Harms-Effenberger, K.; Karstens, K.-F.; Konczalla, L.; Ghadban, T.; Uzunoglu, F.G.; Izbicki, J.R.; Bockhorn, M.; Pantel, K.; ReehMichael, M. Clinical Relevance of Circulating Tumor Cells in Esophageal Cancer Detected by a Combined MACS Enrichment Method. Cancers 2020, 12, 718. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, M.; Takeuchi, H.; Osaki, Y.; Hiraiwa, K.; Nakamura, R.; Oyama, T.; Takahashi, T.; Wada, N.; Kawakubo, H.; Saikawa, Y.; et al. Prognostic significance of circulating tumor cells in patients with advanced esophageal cancer. Esophagus 2014, 12, 352–359. [Google Scholar] [CrossRef]
- Pernot, S.; Badoual, C.; Terme, M.; Castan, F.; Cazes, A.; Bouche, O.; Bennouna, J.; Francois, E.; Ghiringhelli, F.; De La Fouchardiere, C.; et al. Dynamic evaluation of circulating tumour cells in patients with advanced gastric and oesogastric junction adenocarcinoma: Prognostic value and early assessment of therapeutic effects. Eur. J. Cancer 2017, 79, 15–22. [Google Scholar] [CrossRef]
- Brungs, D.; Lynch, D.; Luk, A.W.; Minaei, E.; Ranson, M.; Aghmesheh, M.; Vine, K.L.; Carolan, M.; Jaber, M.; De Souza, P.; et al. Cryopreservation for delayed circulating tumor cell isolation is a valid strategy for prognostic association of circulating tumor cells in gastroesophageal cancer. World J. Gastroenterol. 2018, 24, 810–818. [Google Scholar] [CrossRef]
- Han, L.; Li, Y.-J.; Zhang, W.-D.; Song, P.-P.; Li, H.; Li, S. Clinical significance of tumor cells in the peripheral blood of patients with esophageal squamous cell carcinoma. Med. Baltim. 2019, 98, e13921. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Li, Y.; Yuan, D.; Peng, Y.; Qin, J. Practical value of identifying circulating tumor cells to evaluate esophageal squamous cell carcinoma staging and treatment efficacy. Thorac. Cancer 2018, 9, 956–966. [Google Scholar] [CrossRef] [PubMed]
- Jiao, W.; Xu, J.; Zheng, J.; Shen, Y.; Lin, L.; Li, J. Elevation of circulating big endothelin-1: An independent prognostic factor for tumor recurrence and survival in patients with esophageal squamous cell carcinoma. BMC Cancer 2008, 8, 334. [Google Scholar] [CrossRef] [PubMed]
- Blanchard, P.; Quero, L.; Pacault, V.; Schlageter, M.-H.; Baruch-Hennequin, V.; Hennequin, C. Prognostic significance of anti-p53 and anti-KRas circulating antibodies in esophageal cancer patients treated with chemoradiotherapy. BMC Cancer 2012, 12, 119. [Google Scholar] [CrossRef] [PubMed]
- Cao, M.; Yie, S.-M.; Wu, S.-M.; Chen, S.; Lou, B.; He, X.; Ye, S.-R.; Xie, K.; Rao, L.; Gao, E.; et al. Detection of survivin-expressing circulating cancer cells in the peripheral blood of patients with esophageal squamous cell carcinoma and its clinical significance. Clin. Exp. Metastasis 2009, 26, 751–758. [Google Scholar] [CrossRef]
- He, X.; Xu, X.; Zhu, G.; Ye, H. Circulating uPA as a potential prognostic biomarker for resectable esophageal squamous cell carcinoma. Med. Baltim. 2019, 98, e14717. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, K.; Yano, M.; Motoori, M.; Kishi, K.; Miyashiro, I.; Shingai, T.; Gotoh, K.; Noura, S.; Takahashi, H.; Ohue, M.; et al. CEA-Antigen and SCC-Antigen mRNA Expression in Peripheral Blood Predict Hematogenous Recurrence After Resection in Patients with Esophageal Cancer. Ann. Surg. Oncol. 2010, 17, 2779–2786. [Google Scholar] [CrossRef] [PubMed]
- Yin, X.-D.; Yuan, X.; Xue, J.-J.; Wang, R.; Zhang, Z.-R.; Tong, J.-D. Clinical significance of carcinoembryonic antigen-, cytokeratin 19-, or survivin-positive circulating tumor cells in the peripheral blood of esophageal squamous cell carcinoma patients treated with radiotherapy. Dis. Esophagus 2012, 25, 750–756. [Google Scholar] [CrossRef]
- Honma, H.; Kanda, T.; Ito, H.; Wakai, T.; Nakagawa, S.; Ohashi, M.; Koyama, Y.; Valera, V.A.; Akazawa, K.; Hatakeyama, K. Squamous cell carcinoma-antigen messenger RNA level in peripheral blood predicts recurrence after resection in patients with esophageal squamous cell carcinoma. Surgery 2006, 139, 678–685. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, C.-C.; Hsu, H.-S.; Chang, S.-C.; Chen, Y.-J. Circulating Cell-Free DNA Levels Could Predict Oncological Outcomes of Patients Undergoing Esophagectomy for Esophageal Squamous Cell Carcinoma. Int. J. Mol. Sci. 2016, 17, 2131. [Google Scholar] [CrossRef] [PubMed]
- Diakowska, D.; Nienartowicz, M.; Grabowski, K.; Rosińczuk, J.; Krzystek-Korpacka, M. Toll-like receptors TLR-2, TLR-4, TLR-7, and TLR-9 in tumor tissue and serum of the patients with esophageal squamous cell carcinoma and gastro-esophageal junction cancer. Adv. Clin. Exp. Med. 2018, 28, 515–522. [Google Scholar] [CrossRef]
- Liao, Y.; Xing, S.; Xu, B.; Liu, W.; Zhang, G. Evaluation of the circulating level of fibroblast activation protein α for diagnosis of esophageal squamous cell carcinoma. Oncotarget 2017, 8, 30050–30062. [Google Scholar] [CrossRef]
- Laird, P.W. The power and the promise of DNA methylation markers. Nat. Rev. Cancer 2003, 3, 253–266. [Google Scholar] [CrossRef]
- Duffy, M.J.; Napieralski, R.; Martens, J.; Span, P.N.; Spyratos, F.; Sweep, F.C.; Brünner, N.; Foekens, J.; Schmitt, M. Methylated genes as new cancer biomarkers. Eur. J. Cancer 2009, 45, 335–346. [Google Scholar] [CrossRef]
- Roy, D.; Tiirikainen, M. Diagnostic Power of DNA Methylation Classifiers for Early Detection of Cancer. Trends Cancer 2020, 6, 78–81. [Google Scholar] [CrossRef]
- Zhu, Y.; Zhang, H.; Chen, N.; Hao, J.; Jin, H.; Ma, X. Diagnostic value of various liquid biopsy methods for pancreatic cancer. Med. Baltim. 2020, 99, e18581. [Google Scholar] [CrossRef]
- Jin, X.; Cai, C.; Qiu, Y. Diagnostic Value of Circulating microRNAs in Hepatitis B Virus-Related Hepatocellular Carcinoma: A Systematic Review and Meta-Analysis. J. Cancer 2019, 10, 4754–4764. [Google Scholar] [CrossRef]
- Cheng, X.; Zhang, L.; Chen, Y.; Qing, C. Circulating cell-free DNA and circulating tumor cells, the “liquid biopsies” in ovarian cancer. J. Ovarian Res. 2017, 10, 75. [Google Scholar] [CrossRef] [PubMed]
- Danese, E.; Montagnana, M.; Lippi, G. Circulating molecular biomarkers for screening or early diagnosis of colorectal cancer: Which is ready for prime time? Ann. Transl. Med. 2019, 7, 610. [Google Scholar] [CrossRef] [PubMed]
- Umer, M.; Vaidyanathan, R.; Nguyen, N.-T.; Shiddiky, M.J.A. Circulating tumor microemboli: Progress in molecular understanding and enrichment technologies. Biotechnol. Adv. 2018, 36, 1367–1389. [Google Scholar] [CrossRef] [PubMed]
- Alix-Panabières, C.; Bartkowiak, K.; Pantel, K. Functional studies on circulating and disseminated tumor cells in carcinoma patients. Mol. Oncol. 2016, 10, 443–449. [Google Scholar] [CrossRef] [PubMed]
- Kanda, Y. Investigation of the freely available easy-to-use software ‘EZR’ for medical statistics. Bone Marrow Transplant. 2012, 48, 452–458. [Google Scholar] [CrossRef]

| Author Year, Country | Technology | Molecules | SS | CS | Age (Years) | Pathology | DR | BS (mL) | DR in Stage I–II | DR in Stage III–IV | DR in T 1–2 | DR in T 3–4 | OS/PFS | HR | 95% CIs (Hi) | 95% CIs (Low) | p-Value |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Zhao et al. [32], 2019, China | ISET | 8-um pore size | 55 | 20 | - | ESCC | 52.7% | 5 | 33.3% | 66.7% | 55.6% | 52.2% | OS | 1.84 | 0.60 | 5.66 | 0.290 |
| Han et al. [86], 2019, China | ISET | 8-um pore size | 60 | 0 | 62.2 | ESCC | 33.3% | 5 | 23.5% | 37.2% | 0.0% | 40.0% | PFS | 5.63 | 1.77 | 17.89 | 0.003 |
| Qiao et al. [27], 2017, China | CD45 magnetic beads | pan-CK | 59 | 25 | 61.5 | ESCC | 79.7% | 7.5 | 23.5% | 69.0% | 12.5% | 60.8% | OS | 12.48 | 8.24 | 34.34 | 0.037 |
| PFS | 6.52 | 1.24 | 34.34 | 0.027 | |||||||||||||
| Li et al. [54], 2016, China | MACS | pan-CK | 140 | 25 | 62.8 | ESCC | 44.3% | 5 | 31.0% | 43.5% | 36.4% | 51.4% | OS | 1.82 | 0.91 | 4.88 | 0.046 |
| PFS | 1.86 | 0.87 | 3.15 | 0.035 | |||||||||||||
| Woestemeier et al. [82], 2020, Germany | MACS | pan-CK, EpCAM | 90 | 0 | 63.7 | EC | 25.6% | 7.5 | 7.1% | 8.3% | 7.5% | 8.0% | OS | 0.72 | 0.30 | 1.76 | 0.474 |
| Konczalla et al. [25], 2019, Germany | CellSearch | pan-CK, EpCAM | 76 | 0 | - | EC | 19.7% | 7.5 | 17.9% | 21.6% | 10.3% | 25.5% | OS | 2.70 | 1.31 | 5.58 | 0.007 |
| PFS | 4.04 | 1.86 | 8.76 | 0.001 | |||||||||||||
| Matsushita et al. [13], 2015, Japan | CellSearch | pan-CK, EpCAM | 90 | - | 65 | ESCC | 27.8% | 7.5 | 0.0% | 28.4% | 25.0% | 28.0% | OS | 2.56 | 1.15 | 5.68 | 0.021 |
| PFS | 1.25 | 0.65 | 2.31 | 0.497 | |||||||||||||
| Reeh et al. [26], 2015, Germany | CellSearch | pan-CK, EpCAM | 100 | - | 66 | EC | 18.0% | 7.5 | 15.1% | 21.3% | 32.6% | 27.8% | OS | 3.13 | 1.49 | 6.56 | 0.003 |
| PFS | 5.06 | 2.23 | 11.48 | 0.001 | |||||||||||||
| Tanaka et al. [83], 2015, Japan | CellSearch | pan-CK, EpCAM | 38 | - | 63 | EC | 50.0% | 7.5 | 20.0% | 50.0% | 40.0% | 44.4% | OS | 4.44 | 1.40 | 14.04 | 0.011 |
| Pernot et al. [84], 2017, France | CellSearch | pan-CK, EpCAM | 106 | 0 | -. | EAC | 46.2% | 7.5 | - | - | - | - | OS | 1.85 | 1.17 | 2.92 | 0.010 |
| PFS | 1.59 | 1.04 | 2.43 | 0.030 | |||||||||||||
| Zhang et al. [29], 2019, China | SE-iFISH | pan-CK/CEP8 | 63 | 50 | -. | ESCC | 74.6% | 7.5 | 21.1% | 32.0% | 20.0% | 26.4% | PFS | 3.92 | 0.91 | 16.95 | 0.047 |
| Brungs et al. [85], 2018, Australia | IsoFlux | pan-CK/uPAR | 43 | 0 | 64 | EAC | 46.5% | 7.5 | 27.8% | 60.0% | - | - | OS | 3.70 | 1.20 | 12.4 | 0.030 |
| Ko et al. [75], 2020, Korea | ClearCell, CTChip | pan-CK/EpCAM/MUC1 baseline | 57 | 19 | 63 | ESCC | 70.9% | 5 | - | - | - | -. | OS | 0.97 | 0.44 | 2.14 | 0.946 |
| PFS | 1.35 | 0.69 | 2.63 | 0.380 | |||||||||||||
| pan-CK/EpCAM/MUC1 pre-cycle3 | OS | 3.58 | 1.63 | 7.84 | 0.001 | ||||||||||||
| PFS | 3.68 | 1.73 | 7.81 | 0.001 | |||||||||||||
| Chen et al. [87], 2018, China | CanPatrol RNA-FISH | Vimentin/twist | 71 | 40 | 62.7 | ESCC | 64.8% | 5 | - | - | - | PFS | 0.46 | 0.12 | 1.68 | 0.237 |
| Author, Year, Country | Technology | Molecules | SS | CS | Age (Years) | Pathology | DR | BS (mL) | DR in Stage I–II | DR in Stage III–IV | DR in T 1–2 | DR in T 3–4 | OS/PFS | HR | 95% CIs (Hi) | 95% CIs (Low) | p-Value |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Ko et al. [75], 2020, Korea | Fluorometer | cfDNA: Baseline | 57 | 19 | 63.0 | ESCC | - | 5 | - | - | - | - | OS | 8.34 | 2.42 | 28.7 | 0.001 |
| PFS | 1.96 | 0.67 | 5.76 | 0.222 | |||||||||||||
| Fluorometer | cfDNA: Pre-cycle3 | 57 | 19 | 63.0 | ESCC | - | 5 | - | - | - | - | OS | 5.45 | 1.74 | 17.1 | 0.004 | |
| PFS | 1.68 | 0.7 | 4.06 | 0.249 | |||||||||||||
| Nozoe et al. [39], 2005, Japan | ELISA | CD4/CD8 ratio | 134 | - | 62.0 | ESCC | 35.80% | - | 38.10% | 32.00% | 40.30% | 31.90% | OS | 1.73 | 1.02 | 2.93 | 0.043 |
| PFS | 2.07 | 1.26 | 3.38 | 0.004 | |||||||||||||
| Jiao et al. [88], 2008, China | ELISA | Endothelin-1 | 108 | 82 | 64.5 | ESCC | - | -. | - | - | - | - | OS | 2.63 | 1.38 | 4.05 | 0.003 |
| Blanchard et al. [89], 2012, France | ELISA | Kras | 84 | - | 60.0 | EC | 22.60% | - | - | - | 7.70% | 25.40% | OS | 0.8 | 0.5 | 1.5 | 0.5 |
| PFS | 0.8 | 0.5 | 1.4 | 0.5 | |||||||||||||
| ELISA | p53 | 84 | - | 60.0 | EC | 28.60% | - | - | - | 7.70% | 32.40% | OS | 2 | 1.05 | 2.8 | 0.04 | |
| PFS | 2 | 1 | 3.9 | 0.04 | |||||||||||||
| Kimura et al. [40], 2008, Japan | ELISA | VEGF-C | 80 | 20 | 62.8 | EC | - | - | - | - | - | - | PFS | 5.6 | 1.6 | 19.6 | 0.007 |
| Hu et al. [73], 2016, China | qRT-PCR | lnc CFLAR-AS1 | 205 | 210 | 54.3 | ESCC | - | - | - | - | - | - | OS | 1.68 | 1.08 | 2.32 | N.D. |
| Linc00152 | 205 | 210 | 54.3 | ESCC | - | - | - | - | - | - | OS | 1.89 | 1.22 | 2.58 | N.D. | ||
| lnc POU3F3 | 205 | 210 | 54.3 | ESCC | - | - | - | - | - | - | OS | 1.82 | 1.17 | 2.51 | N.D. | ||
| Li et al. [47], 2017, China | qRT-PCR | miR-15a | 106 | 106 | 62.3 | EC | - | - | - | - | - | - | OS | 4.17 | 1.97 | 10.63 | 0.01 |
| PFS | 4.01 | 1.62 | 9.82 | 0.01 | |||||||||||||
| Lv et al. [48], 2016, China | qRT-PCR | miR-21 | 126 | 80 | 59.1 | ESCC | - | - | 48.40% | 60.90% | 34.50% | 62.00% | OS | 1.85 | 1.48 | 6.24 | 0.012 |
| miR-375 | 126 | 80 | 59.1 | ESCC | - | - | 66.10% | 34.40% | 45.50% | 39.40% | OS | 0.65 | 0.27 | 0.86 | 0.041 | ||
| Komatsu et al. [49], 2016, Japan | qRT-PCR | miR-21 | 37 | 20 | - | ESCC | 43.20% | 7 | - | - | - | - | PFS | 9.95 | 1.56 | 63.42 | 0.015 |
| Wu et al. [50], 2014, China | qRT-PCR | miR-25 | 194 | 94 | - | ESCC | - | 2 | 61.90% | 62.50% | - | - | OS | 1.13 | 0.78 | 1.64 | 0.526 |
| miR-25 | 63 | 63 | - | ESCC | - | 3 | - | - | - | - | OS | 3.84 | 1.02 | 14.41 | 0.046 | ||
| miR-100 | 63 | 63 | - | ESCC | - | 3 | - | - | - | - | OS | 4.18 | 1.21 | 14.5 | 0.024 | ||
| miR-223 | 194 | 94 | - | ESCC | - | 2 | 56.80% | 59.40% | - | - | OS | 1.72 | 1.14 | 2.59 | 0.01 | ||
| miR-375 | 194 | 94 | - | ESCC | - | 2 | 70.30% | 64.10% | - | - | OS | 1.75 | 1.11 | 2.76 | 0.016 | ||
| Gu et al. [51], 2018, China | qRT-PCR | miR-25-3p | 329 | - | 61.7 | EAC | - | - | - | - | - | - | PFS | 1.04 | 0.73 | 1.25 | 0.817 |
| miR-30c-5p | 329 | - | 61.7 | EAC | -. | - | - | - | - | - | PFS | 0.86 | 0.61 | 1.22 | 0.397 | ||
| miR-152-3p | 329 | - | 61.7 | EAC | - | - | - | - | - | - | PFS | 0.78 | 0.54 | 1.11 | 0.161 | ||
| miR-331-3p | 329 | - | 61.7 | EAC | - | - | - | - | - | - | PFS | 0.55 | 0.38 | 0.78 | 0.001 | ||
| Tanaka et al. [52], 2013, Japan | qRT-PCR | miR-200c | 64 | 27 | - | ESCC | - | 8 | 28.00% | 64.10% | 30.00% | 59.10% | OS | 2.79 | 1.11 | 7.96 | N.D. |
| PFS | 2.79 | 1.11 | 7.96 | 0.029 | |||||||||||||
| Zhai et al. [53], 2015, China | qRT-PCR | miR-3935, 4286 | 30 | 30 | 68.7 | EC | - | - | - | - | - | - | OS | 10.91 | 1.8 | 66.12 | 0.009 |
| Li et al. [54], 2016, China | qRT-PCR | miR-506 | 100 | 40 | 59.2 | ESCC | - | 5 | 30.40% | 51.90% | 35.40% | 48.10% | OS | 2.35 | 1.32 | 4.2 | 0.004 |
| PFS | 2.65 | 1.53 | 4.58 | 1 | |||||||||||||
| Guan et al. [55], 2016, China | qRT-PCR | miR-613 | 75 | 75 | 65.0 | ESCC | - | - | - | - | - | - | OS | 0.59 | 0.34 | 0.95 | 0.031 |
| PFS | 0.66 | 0.48 | 0.89 | 0.006 | |||||||||||||
| Setoyama et al. [42], 2006, Japan | qRT-PCR | CEA | 106 | 28 | 63.3 | EC | 36.80% | 5 | 19.70% | 60.00% | 18.30% | 56.50% | PFS | 0.53 | 0.32 | 0.8 | 0.002 |
| Tanaka et al. [92], 2010, Japan | qRT-PCR | CEA, SCC | 244 | - | - | ESCC | 16.80% | 8 | 15.30% | 18.60% | 17.00% | 16.70% | PFS | 1.65 | 1.03 | 2.63 | 0.037 |
| Yin et al. [93], 2012, China | qRT-PCR | CEA, survivin, CK19 | 72 | - | 63.0 | EC | 54.20% | - | - | - | -. | - | PFS | 3.68 | 1.38 | 9.84 | 0.008 |
| Honma et al. [94], 2006, Japan | qRT-PCR | SCC | 46 | 42 | 66.0 | ESCC | 30.40% | 3 | 16.70% | 39.30% | 21.10% | 37.00% | PFS | 3 | 1.05 | 8.54 | 0.04 |
| Kaganoi et al. [8], 2004, Japan | qRT-PCR | SCC | 70 | 19 | - | ESCC | 32.80% | 10 | 18.80% | 63.60% | 13.60% | 65.40% | PFS | 7.15 | 1.25 | 61.1 | 0.038 |
| Hoffmann et al. [44], 2010, Germany | qRT-PCR | Survivin | 62 | - | - | EC | 77.00% | - | - | - | - | - | OS | 6.6 | 1.97 | 22.12 | 0.002 |
| Cao et al. [90], 2009, China | qRT-PCR | Survivin | 108 | 75 | 58.9 | ESCC | 47.20% | 2 | 20.90% | 64.60% | 36.50% | 57.10% | OS | 5.17 | 2.3 | 11.65 | 0.001 |
| PFS | 5.18 | 2.42 | 8.93 | 0.005 | |||||||||||||
| He et al. [91], 2019, China | qRT-PCR | uPA | 205 | - | - | ESCC | 25.90% | - | 22.90% | 28.40% | 16.70% | 29.70% | OS | 1.82 | 1.16 | 2.85 | 0.009 |
| PFS | 1.97 | 1.11 | 3.49 | 0.02 |
| Author, Year, Country | Technology | Molecules | SS | CS | Age (Years) | Pathology | AUC | Sensitivity | Specificity | 95% CIs (Hi) | 95% CIs (Low) |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Ko et al. [75], 2020, Korea | ClearCell CTChip | pan-CK/EpCAM/MUC1 baseline | 57 | 19 | 63 | ESCC | 0.681 | 0.455 | 0.895 | ||
| Hsieh et al. [95], 2016, Taiwan | qRT-PCR | ctDNA: Copy number | 81 | 95 | 60.4 | ESCC | 0.991 | 0.963 | 0.941 | 0.982 | 0.999 |
| Liao et al. [97], 2017, China | ELISA | FAPα | 151 | 194 | 62 | ESCC | 0.714 | 0.561 | 0.856 | ||
| Tong et al. [72], 2015, China | qRT-PCR | lnc POU3F3 | 147 | 23 | - | ESCC | 0.842 | 0.728 | 0.894 | 0.748 | 0.853 |
| Luoet al. [63], 2020, China | qRT-PCR | lnc SNHG1 | 60 | 60 | - | ESCC | 0.85 | 0.774 | 0.925 | ||
| Zhang et al. [78], 2012, China | LC–MS | Metabolites | 67 | 34 | - | EAC | 0.92 | 0.89 | 0.9 | ||
| Xuet al. [79], 2106, China | LC–MS | Metabolites | 62 | 62 | 62 | EC | 0.981 | 0.913 | 0.984 | ||
| Zhu et al. [80], 2020, China | LC–MS | Metabolites | 140 | 170 | 60 | ESCC | 0.965 | 0.883 | 0.889 | 0.936 | 0.993 |
| He et al. [56], 2015, China | qRT-PCR | let-7a | 70 | 40 | 60.5 | ESCC | 0.829 | 0.743 | 0.85 | 0.754 | 0.904 |
| Cui et al. [57], 2017, China | qRT-PCR | miR-9 | 131 | 131 | - | ESCC | 0.913 | 0.855 | 0.985 | 0.873 | 0.953 |
| Shen et al. [58], 2019, China | qRT-PCR | miR-16-5p, 197-5p, 451a, 92a-3p | 96 | 78 | 60.1 | ESCC | 0.856 | 0.896 | 0.763 | 0.794 | 0.905 |
| Zheng et al. [60], 2019, China | qRT-PCR | miR-16-5p, 451a, 574-5p | 23 | 23 | - | ESCC | 0.76 | 0.73 | 0.82 | ||
| Hirajima et al. [61], 2013, Japan | qRT-PCR | miR-18a | 106 | 54 | - | ESCC | 0.9449 | 0.868 | 1 | ||
| He et al. [56], 2015, China | qRT-PCR | miR-20a | 70 | 40 | 60.5 | ESCC | 0.767 | 0.643 | 0.75 | 0.677 | 0.857 |
| Zhang et al. [62], 2018, China | qRT-PCR | miR-21 | 125 | 125 | 63 | ESCC | 0.8 | 0.74 | 0.78 | ||
| Luo et al. [63], 2020, China | qRT-PCR | miR-21 | 60 | 60 | - | ESCC | 0.928 | 0.883 | 0.973 | ||
| Komatsu et al. [49], 2016, Japan | qRT-PCR | miR-21 | 37 | 20 | - | ESCC | 0.8154 | 0.542 | 0.923 | ||
| Zhang et al. [62], 2018, China | qRT-PCR | miR-25 | 125 | 125 | 63 | ESCC | 0.55 | 0.54 | 0.57 | ||
| Wu et al. [50], 2014, China | qRT-PCR | miR-25 | 194 | 94 | - | ESCC | 0.593 | 0.471 | 0.716 | ||
| Ibuki et al. [64], 2020, Japan | qRT-PCR | miR-30a-5p, 205-5p, 574-3p | 66 | 42 | - | ESCC | 0.95 | 0.938 | 0.81 | 0.91 | 1 |
| Zhang et al. [62], 2018, China | qRT-PCR | miR-100 | 125 | 125 | 63 | ESCC | 0.58 | 0.58 | 0.58 | ||
| Bus et al. [65] 2016, Netherlands | qRT-PCR | miR-133a-3p, 136-5p, 382-5p | 59 | 15 | 65.8 | EAC | 0.797 | 0.8095 | 0.7838 | ||
| Pavlov et al. [66], 2018, Netherlands | qRT-PCR | miR-199a-3p, 320e | 17 | 19 | 65.1 | EAC | 0.786 | 0.823 | 0.622 | ||
| Dong et al. [67], 2016, China | qRT-PCR | miR-216a | 120 | 51 | - | ESCC | 0.877 | 0.8 | 0.902 | ||
| Dong et al. [67], 2016, China | qRT-PCR | miR-216b | 120 | 51 | - | ESCC | 0.756 | 0.558 | 0.902 | ||
| Zhang et al. [62], 2018, China | qRT-PCR | miR-223 | 125 | 125 | 63 | ESCC | 0.73 | 0.68 | 0.68 | ||
| Zhang et al. [62], 2018, China | qRT-PCR | miR-375 | 125 | 125 | 63 | ESCC | 0.69 | 0.78 | 0.59 | ||
| Li et al. [54], 2016, China | qRT-PCR | miR-506 | 100 | 40 | 59.2 | ESCC | 0.835 | 0.8636 | 1 | ||
| Tong et al. [72], 2015, China | qRT-PCR | SCC | 147 | 23 | - | ESCC | 0.784 | 0.592 | 0.935 | 0.727 | 0.841 |
| Diakowska et al. [96], 2019, Poland | qRT-PCR | TLR-4 | 27 | 38 | - | EAC | 0.787 | 0.7 | 0.78 | 0.661 | 0.909 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Matsushita, D.; Arigami, T.; Okubo, K.; Sasaki, K.; Noda, M.; Kita, Y.; Mori, S.; Uenosono, Y.; Ohtsuka, T.; Natsugoe, S. The Diagnostic and Prognostic Value of a Liquid Biopsy for Esophageal Cancer: A Systematic Review and Meta-Analysis. Cancers 2020, 12, 3070. https://doi.org/10.3390/cancers12103070
Matsushita D, Arigami T, Okubo K, Sasaki K, Noda M, Kita Y, Mori S, Uenosono Y, Ohtsuka T, Natsugoe S. The Diagnostic and Prognostic Value of a Liquid Biopsy for Esophageal Cancer: A Systematic Review and Meta-Analysis. Cancers. 2020; 12(10):3070. https://doi.org/10.3390/cancers12103070
Chicago/Turabian StyleMatsushita, Daisuke, Takaaki Arigami, Keishi Okubo, Ken Sasaki, Masahiro Noda, Yoshiaki Kita, Shinichiro Mori, Yoshikazu Uenosono, Takao Ohtsuka, and Shoji Natsugoe. 2020. "The Diagnostic and Prognostic Value of a Liquid Biopsy for Esophageal Cancer: A Systematic Review and Meta-Analysis" Cancers 12, no. 10: 3070. https://doi.org/10.3390/cancers12103070
APA StyleMatsushita, D., Arigami, T., Okubo, K., Sasaki, K., Noda, M., Kita, Y., Mori, S., Uenosono, Y., Ohtsuka, T., & Natsugoe, S. (2020). The Diagnostic and Prognostic Value of a Liquid Biopsy for Esophageal Cancer: A Systematic Review and Meta-Analysis. Cancers, 12(10), 3070. https://doi.org/10.3390/cancers12103070





