Do GNAQ and GNA11 Differentially Affect Inflammation and HLA Expression in Uveal Melanoma?
Abstract
1. Introduction
2. Results
2.1. mRNA and Patient Distribution
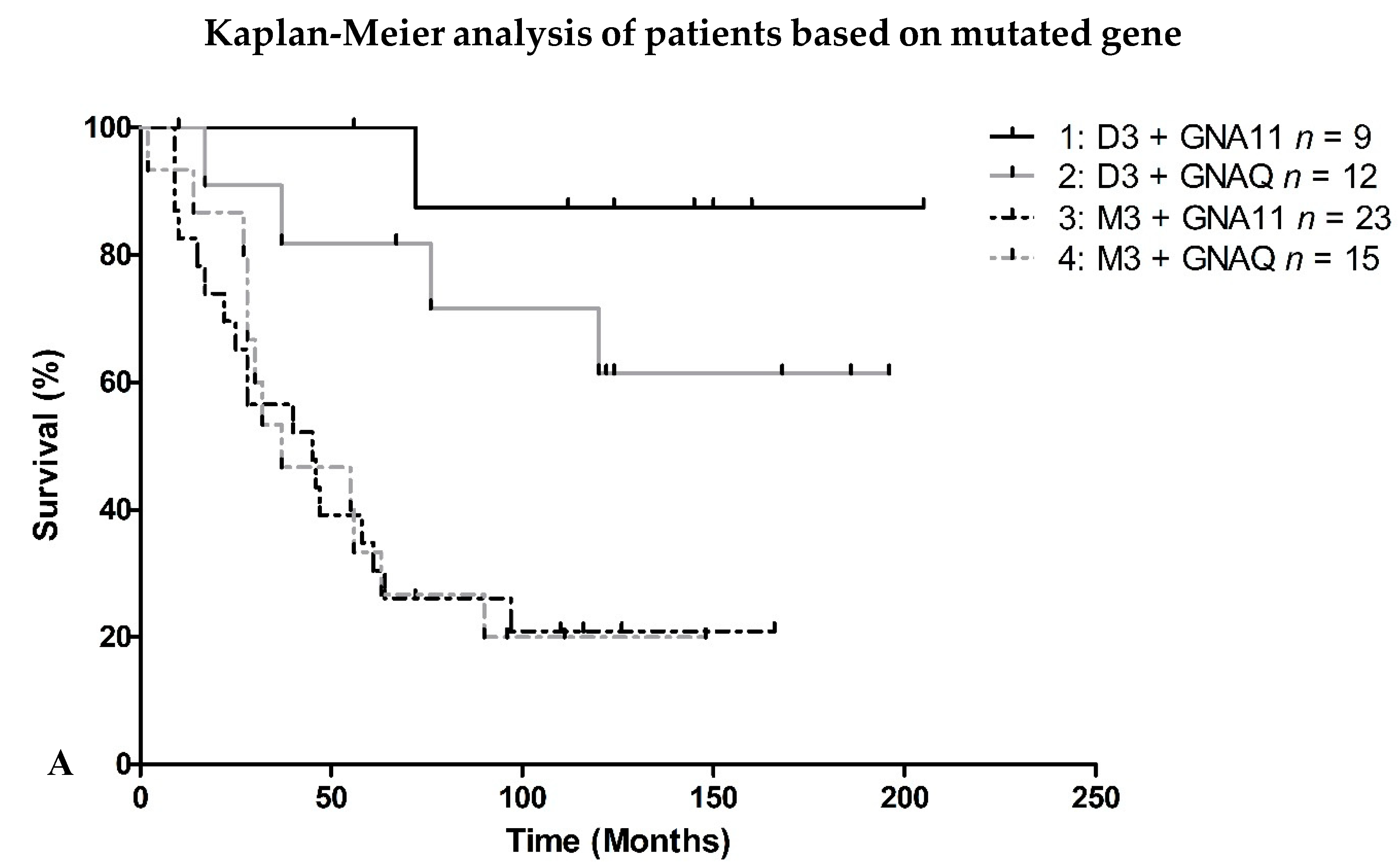
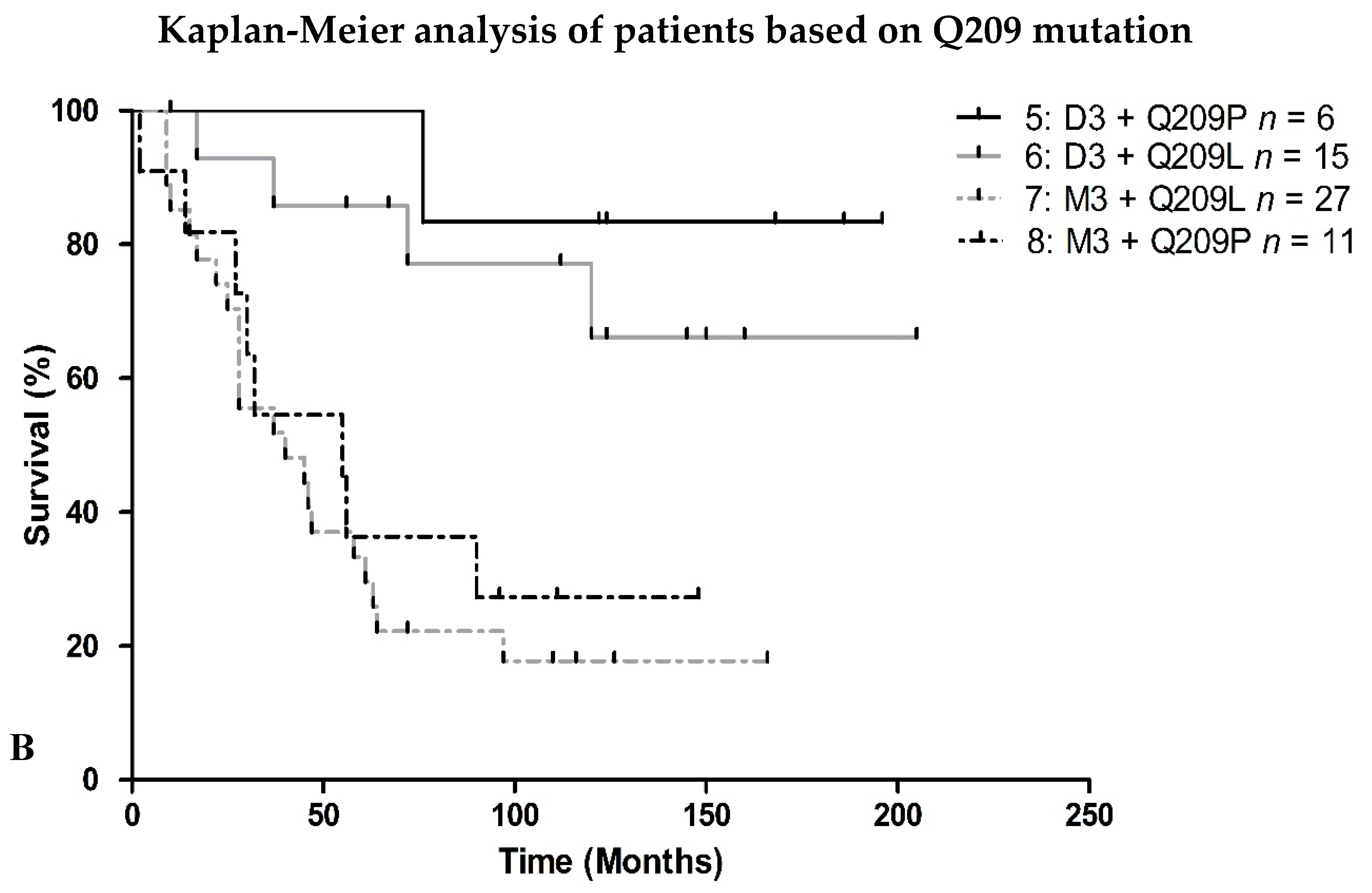
2.2. Survival
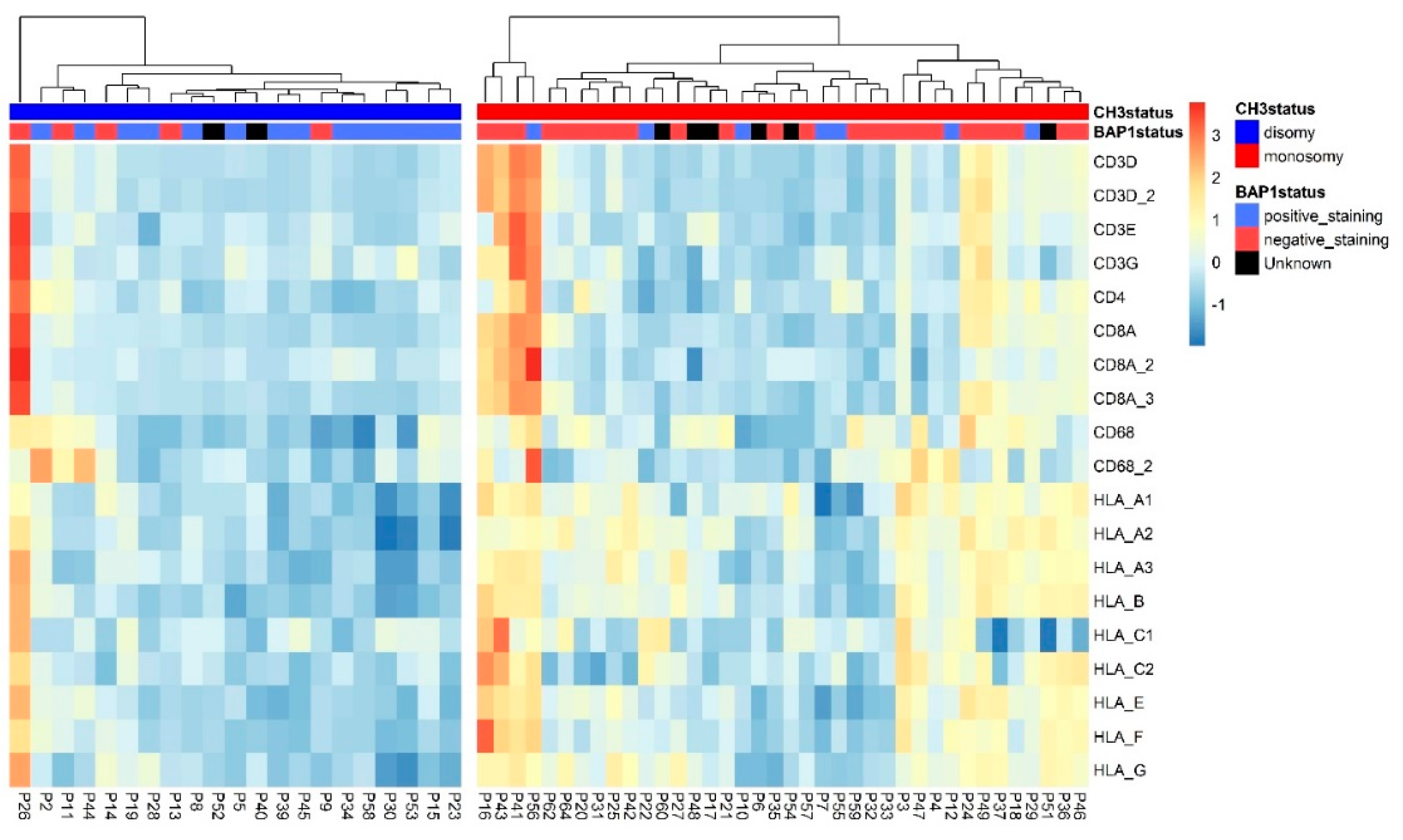
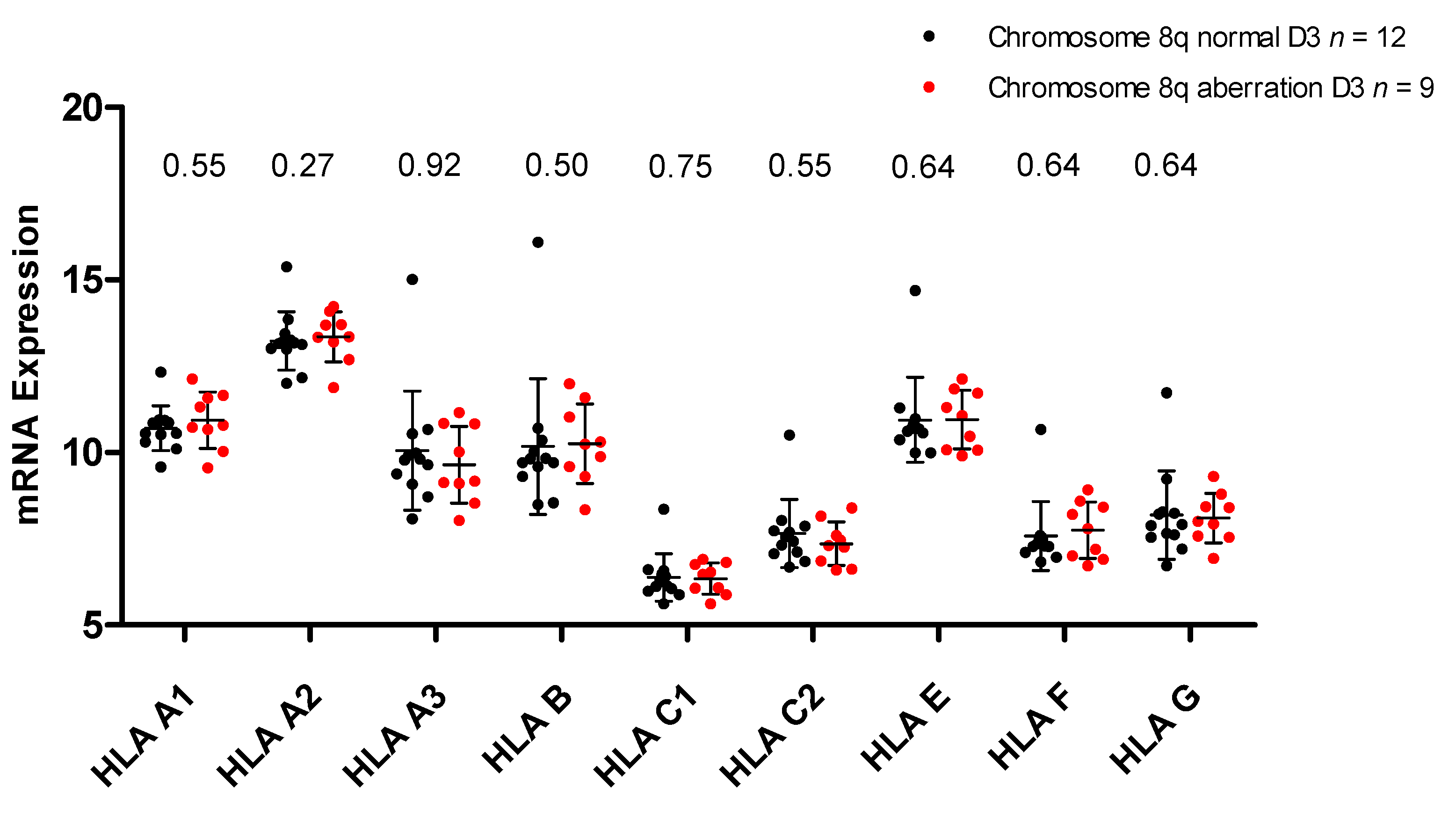
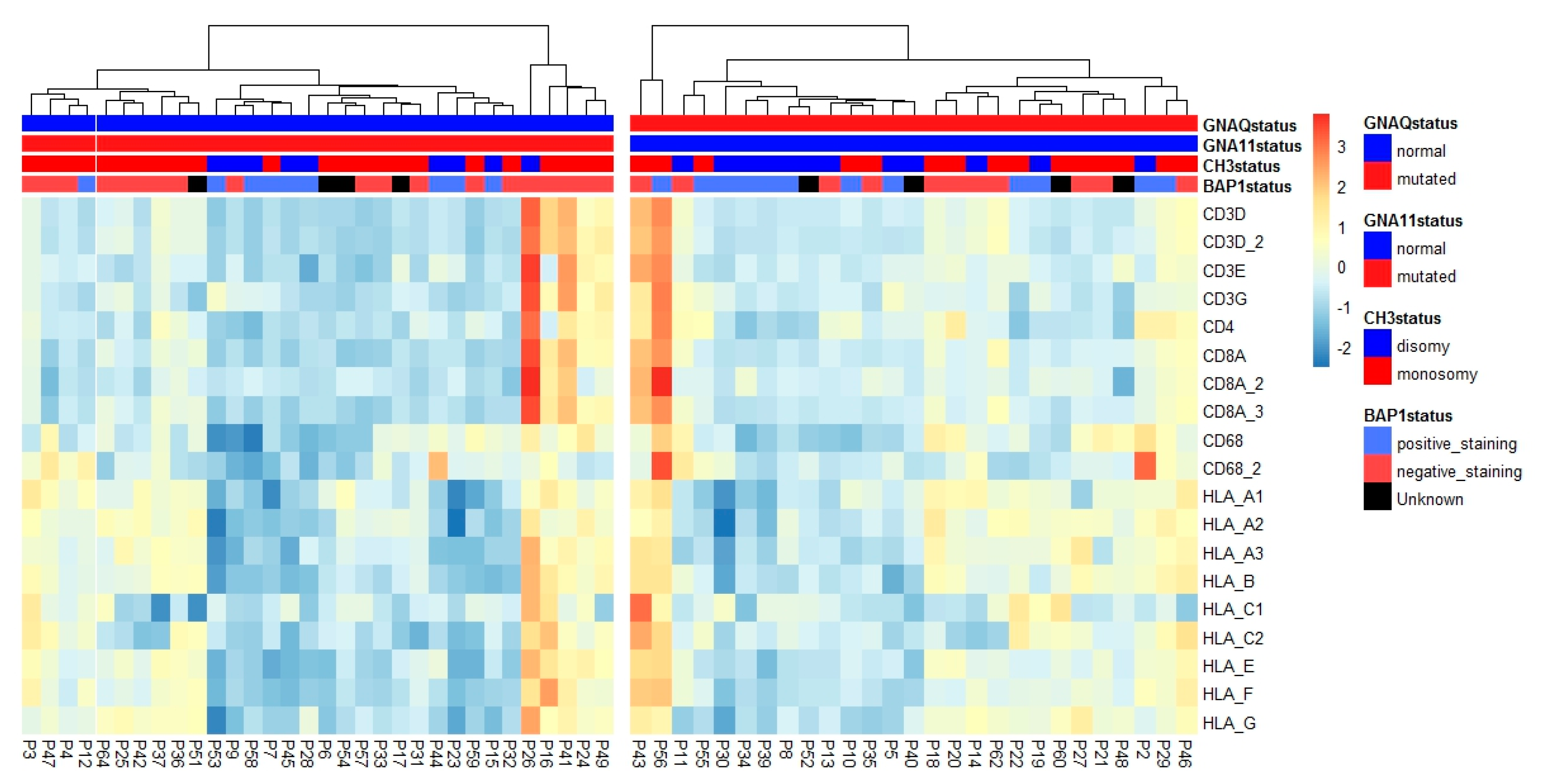
2.3. HLA Expression in Relation to Chromosome 8q Status
3. Discussion
4. Materials and Methods
4.1. Cases
4.2. Chromosome Analysis
4.3. GNAQ and GNA11 Mutations
4.4. mRNA Analysis Status
4.5. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Virgili, G.; Gatta, G.; Ciccolallo, L.; Capocaccia, R.; Biggeri, A.; Crocetti, E.; Lutz, J.M.; Paci, E.; EUROCARE Working Group. Incidence of Uveal Melanoma in Europe. Ophthalmology 2007, 114, 2309–2315. [Google Scholar] [CrossRef] [PubMed]
- Kujala, E.; Mäkitie, T.; Kivelä, T. Very Long-Term Prognosis of Patients with Malignant Uveal Melanoma. Investig. Ophthalmol. Vis. Sci. 2003, 44, 4651–4659. [Google Scholar] [CrossRef] [PubMed]
- Van Raamsdonk, C.D.; Bezrookove, V.; Green, G.; Bauer, J.; Gaugler, L.; O’Brien, J.M.; Simpson, E.M.; Barsh, G.S.; Bastian, B.C. Frequent somatic mutations of GNAQ in uveal melanoma and blue naevi. Nature 2009, 457, 599–602. [Google Scholar] [CrossRef] [PubMed]
- Van Raamsdonk, C.D.; Griewank, K.G.; Crosby, M.B.; Garrido, M.C.; Vemula, S.; Wiesner, T.; Obenauf, A.C.; Wackernagel, W.; Green, G.; Bouvier, N.; et al. Mutations in GNA11 in uveal melanoma. N. Engl. J. Med. 2010, 363, 2191–2199. [Google Scholar] [CrossRef] [PubMed]
- Vader, M.; Madigan, M.; Versluis, M.; Suleiman, H.; Gezgin, G.; Gruis, N.A.; Out Luiting, J.J.; Bergman, W.; Verdijk, R.M.; Jager, M.J.; et al. GNAQ and GNA11 mutations and downstream YAP activation in choroidal nevi. Br. J. Cancer 2017, 117, 884–887. [Google Scholar] [CrossRef] [PubMed]
- Harbour, J.W.; Roberson, E.D.O.; Anbunathan, H.; Onken, M.D.; Worley, L.A.; Bowcock, A.M. Recurrent mutations at codon 625 of the splicing factor SF3B1 in uveal melanoma. Nat. Genet. 2013, 45, 133–135. [Google Scholar] [CrossRef] [PubMed]
- Martin, M.; Maßhofer, L.; Temming, P.; Rahmann, S.; Metz, C.; Bornfeld, N.; van de Nes, J.; Klein-Hitpass, L.; Hinnebusch, A.G.; Horsthemke, B.; et al. Exome sequencing identifies recurrent somatic mutations in EIF1AX and SF3B1 in uveal melanoma with disomy 3. Nat. Genet. 2013, 45, 933–936. [Google Scholar] [CrossRef] [PubMed]
- Furney, S.J.; Pedersen, M.; Gentien, D.; Dumont, A.G.; Rapinat, A.; Desjardins, L.; Turajlic, S.; Piperno-Neumann, S.; de la Grange, P.; Roman-Roman, S.; et al. SF3B1 mutations are associated with alternative splicing in uveal melanoma. Cancer Discov. 2013, 3, 1122–1129. [Google Scholar] [CrossRef] [PubMed]
- Prescher, G.; Bornfeld, N.; Hirche, H.; Horsthemke, B.; Jockel, K.H.; Becher, R. Prognostic implications of monosomy 3 in uveal melanoma. Lancet 1996, 347, 1222–1225. [Google Scholar] [CrossRef]
- Ewens, K.G.; Kanetsky, P.A.; Richards-Yutz, J.; Purrazzella, J.; Shields, C.L.; Ganguly, T.; Ganguly, A. Chromosome 3 status combined with BAP1 and EIF1AX mutation profiles are associated with metastasis in uveal melanoma. Investig. Ophthalmol. Vis. Sci. 2014, 55, 5160–5167. [Google Scholar] [CrossRef]
- Koopmans, A.E.; Verdijk, R.M.; Brouwer, R.W.; van den Bosch, T.P.; van den Berg, M.M.; Vaarwater, J.; Kockx, C.E.; Paridaens, D.; Naus, N.C.; Nellist, M.; et al. Clinical significance of immunohistochemistry for detection of BAP1 mutations in uveal melanoma. Mod. Pathol. 2014, 27, 1321–1330. [Google Scholar] [CrossRef] [PubMed]
- Van Essen, T.H.; van Pelt, S.I.; Versluis, M.; Bronkhorst, I.H.; van Duinen, S.G.; Marinkovic, M.; Kroes, W.G.M.; Ruivenkamp, C.A.; Shukla, S.; de Klein, A.; et al. Prognostic parameters in uveal melanoma and their association with BAP1 expression. Br. J. Ophthalmol. 2014, 98, 1738–1743. [Google Scholar] [CrossRef]
- Robertson, A.G.; Shih, J.; Yau, C.; Gibb, E.A.; Oba, J.; Mungall, K.L.; Hess, J.M.; Uzunangelov, V.; Walter, V.; Danilova, L.; et al. Integrative analysis identifies four molecular and clinical subsets in uveal melanoma. Cancer Cell 2017, 32, 204–220. [Google Scholar] [CrossRef] [PubMed]
- Dogrusöz, M.; Jager, M.J. Genetic prognostication in uveal melanoma. Acta Ophthalmol. 2017, 96, 331–347. [Google Scholar] [CrossRef] [PubMed]
- Kaliki, S.; Shields, C.; Shields, J. Uveal melanoma: Estimating prognosis. Ind. J. Ophthalmol. 2015, 63, 93–102. [Google Scholar] [CrossRef] [PubMed]
- Versluis, M.; de Lange, M.J.; van Pelt, S.I.; Ruivenkamp, C.A.; Kroes, W.G.M.; Cao, J.; Jager, M.J.; Luyten, G.P.M.; van der Velden, P.A. Digital PCR validates 8q dosage as prognostic tool in uveal melanoma. PLoS ONE 2015, 10, e0116371. [Google Scholar] [CrossRef] [PubMed]
- Singh, N.; Singh, A.D.; Hide, W. Inferring an evolutionary tree of uveal melanoma from genomic copy number aberrations. Investig. Ophthalmol. Vis. Sci. 2015, 56, 6801–6809. [Google Scholar] [CrossRef]
- Decatur, C.L.; Ong, E.; Garg, N.; Anbunathan, H.; Bowcock, A.M.; Field, M.G.; Harbour, J.W. Driver mutations in uveal melanoma associations with gene expression profile and patient outcomes. JAMA Ophthalmol. 2016, 134, 728–734. [Google Scholar] [CrossRef]
- Niederkorn, J. Ocular immune privilege and ocular melanoma: Parallel universes or immunological plagiarism? Front. Immunol. 2012, 3, 148. [Google Scholar] [CrossRef]
- Maat, W.; Ly, L.V.; Jordanova, E.S.; de Wolff-Rouendaal, D.; Schalij-Delfos, N.E.; Jager, M.J. Monosomy of Chromosome 3 and an inflammatory phenotype occur together in uveal melanoma. Investig. Ophthalmol. Vis. Sci. 2008, 49, 505–510. [Google Scholar] [CrossRef]
- Blom, D.J.; Luyten, G.P.M.; Mooy, C.; Kerkvliet, S.; Zwinderman, A.H.; Jager, M.J. Human leukocyte antigen class I expression. Marker of poor prognosis in uveal melanoma. Investig. Ophthalmol. Vis. Sci. 1997, 38, 1865–1872. [Google Scholar]
- Ericsson, C.; Seregard, S.; Bartolazzi, A.; Levitskaya, E.; Ferrone, S.; Kiessling, R.; Larsson, O. Association of HLA class I and class II antigen expression and mortality in uveal melanoma. Investig. Ophthalmol. Vis. Sci. 2001, 42, 2153–2156. [Google Scholar]
- Jager, M.J.; Hurks, H.M.; Levitskaya, J.; Kiessling, R. HLA expression in uveal melanoma: There is no rule without some exception. Hum. Immunol. 2002, 83, 444–451. [Google Scholar] [CrossRef]
- Van Essen, T.H.; van Pelt, S.I.; Bronkhorst, I.H.G.; Versluis, M.; Némati, F.; Laurent, C.; Luyten, G.P.M.; van Hall, T.; van den Elsen, P.J.; van der Velden, P.A.; et al. Upregulation of HLA expression in primary uveal melanoma by infiltrating leukocytes. PLoS ONE 2016, 11, e0164292. [Google Scholar] [CrossRef] [PubMed]
- Loumagne, L.; Baudhuin, J.; Favier, B.; Montespan, F.; Carosella, E.D.; Rouas-Freiss, N. In vivo evidence that secretion of HLA-G by immunogenic tumor cells allows their evasion from immunosurveillance. Int. J. Cancer 2014, 135, 2107–2117. [Google Scholar] [CrossRef] [PubMed]
- Lesport, E.; Baudhuin, J.; LeMaoult, J.; Sousa, S.; Doliger, C.; Carosella, E.D.; Favier, B. Human melanoma cell secreting human leukocyte antigen–G5 inhibit natural killer cell cytotoxicity by impairing lytic granules polarization toward target cell. Hum. Immunol. 2009, 70, 1000–1005. [Google Scholar] [CrossRef]
- Hurks, H.M.H.; Valter, M.M.; Wison, L.; Hilgert, I.; van den Elsen, J.; Jager, M.J. Uveal Melanoma: No expression of HLA-G. Investig. Ophthlalmol. Vis. Sci. 2001, 42, 3081–3084. [Google Scholar] [PubMed]
- Gezgin, G.; Dogrusöz, M.; van Essen, T.H.; Kroes, W.G.M.; Luyten, G.P.M.; van der Velden, P.A.; Walter, V.; Verdijk, R.M.; van Hall, T.; van der Burg, S.H.; et al. Genetic Evolution of uveal melanoma guides the development of an inflammatory microenvironment. Cancer Immunol. Immunother. 2017, 66, 904–912. [Google Scholar] [CrossRef]
- Markby, D.W.; Onrust, R.; Bourne, H.R. Separate GTP Binding and GTPase Activating Domains of Gα Subunit. Science 1993, 262, 1895–1901. [Google Scholar] [CrossRef]
- Bauer, J.; Kiliç, E.; Vaarwater, J.; Bastian, B.C.; Garbe, C.; de Klein, A. Oncogenix GNAQ mutations are not correlated with disease-free survival in uveal melanoma. Br. J. Cancer 2009, 101, 813–815. [Google Scholar] [CrossRef] [PubMed]
- Chua, V.; Lapadula, D.; Randolph, C.; Benovic, J.L.; Wedegaertner, P.B.; Aplin, A.E. Dysregulated GPCR signaling and therapeutic options in uveal melanoma. Mol. Cancer Res. 2017, 15, 501–506. [Google Scholar] [CrossRef] [PubMed]
- Bronkhorst, I.H.G.; Vu, T.H.K.; Jordanova, E.S.; Luyten, G.P.M.; van der Burg, S.H.; Jager, M.J. Different subsets of tumor-infiltrating lymphocytes correlate with macrophage influx and monosomy 3 in uveal melanoma. Investig. Ophthalmol. Vis. Sci. 2012, 53, 5370–5378. [Google Scholar] [CrossRef] [PubMed]
- Amaro, A.; Gangemi, R.; Piaggio, F.; Angelini, G.; Barisione, G.; Ferrini, S.; Pfeffer, U. The biology of uveal melanoma. Cancer Metastasis Rev. 2017, 36, 109–140. [Google Scholar] [CrossRef] [PubMed]
- O’Hayre, M.; Degese, M.S.; Gutkind, J.S. Novel insights into G protein and G-protein coupled receptor signalling in cancer. Curr. Opin. Cell Biol. 2014, 27, 126–135. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Li, J.; Zhu, M.; Fletscher, J.A.; Hodi, F.S. Protein Kinase C inhibitor AEB071 targets ocular melanoma harboring GNAQ mutations via effects on the PKC/Erk1/2 and PKC/NF-KB pathways. Mol. Cancer Ther. 2012, 11, 1905–1914. [Google Scholar] [CrossRef] [PubMed]
- Souri, Z.; Wierenga, A.P.A.; van Weeghel, C.; van der Velden, P.A.; Kroes, W.G.M.; Luyten, G.P.M.; van der Burg, S.; Jochemsen, A.G.; Jager, M.J. Loss of BAP1 Is Associated with Upregulation of the NfkB Pathway and Increased HLA Class I Expression in Uveal Melanoma. Cancers 2019, 11, 1102. [Google Scholar] [CrossRef]
- Vaqué, J.P.; Dorsam, R.T.; Feng, X.; Iglesias-Bartolome, R.; Forsthoefel, D.J.; Chen, Q.; Debant, A.; Seeger, M.A.; Ksander, B.R.; Teramoto, H.; et al. Genome-wide RNAi screen reveals a Trio-regulated Rho GTPase circuitry transducing mitogenic signals initiated by G protein-coupled receptors. Mol. Cell 2013, 49, 94–108. [Google Scholar] [CrossRef]
| Characteristics | D3 GNA11 (n = 9) | D3 GNAQ (n = 12) | M3 GNA11 (n = 23) | M3 GNAQ (n = 15) | Disomy 3 Tumors, GNA11 versus GNAQ p-Value | Monosomy 3 Tumors, GNA11 versus GNAQ p-Value | Monosomy 3 versus Disomy 3 p-Value |
|---|---|---|---|---|---|---|---|
| Gender n (%) | 0.70 1 | 0.85 1 | 0.21 1 | ||||
| Female | 3 (33) | 5 (42) | 13 (57) | 8 (53) | |||
| Male | 6 (66) | 7 (58) | 10 (44) | 7 (47) | |||
| Survival n (%) | 0.11 1 | 0.86 1 | 0.001 1 | ||||
| Alive | 7 (78) | 3 (25) | 3 (13) | 2 (13) | |||
| Cause of Death: | |||||||
| Melanoma-related | 1 (11) | 4 (33) | 18 (78) | 12 (80) | |||
| Different cause | 1 (11) | 3 (25) | 1 (4) | 1 (7) | |||
| Unknown | 0 (0) | 2 (17) | 1 (4) | 0 (0) | |||
| Affected eye n (%) | 0.70 1 | 0.55 1 | 0.28 1 | ||||
| Right eye | 3 (33) | 5 (42) | 13 (57) | 7 (47) | |||
| Left eye | 6 (66) | 7 (58) | 10 (44) | 8 (53) | |||
| Age at enucleation | 0.14 2 | 0.65 2 | 0.012 2 | ||||
| Mean | 47.24 | 59.11 | 63.95 | 65.91 | |||
| ±St. dev. | ±17.82 | ±16.75 | ±15.28 | ±10.67 | |||
| Metastases n (%) | 0.24 1 | 0.90 1 | <0.001 1 | ||||
| No | 8 (89) | 8 (67) | 5 (22) | 3 (20) | |||
| Yes | 1 (11) | 4 (33) | 18 (78) | 12 (80) | |||
| Cell type n (%) | 0.43 1 | 0.78 1 | 0.04 1 | ||||
| Spindle | 5 (56) | 6 (50) | 4 (17) | 4 (26) | |||
| Mixed | 3 (33) | 6 (50) | 15 (65) | 9 (60) | |||
| Epithelioid | 1 (11) | 0 (0) | 4 (17) | 2 (13) | |||
| Basal diameter | 0.20 2 | 0.08 2 | 0.18 2 | ||||
| Mean (mm) | 12.00 | 13.42 | 13.13 | 15.67 | |||
| ±St. dev. | ±1.80 | ±3.09 | ±3.29 | ±4.62 | |||
| Prominence | 0.81 2 | 0.75 2 | 0.30 2 | ||||
| Mean (mm) | 7.00 | 7.29 | 7.85 | 8.13 | |||
| ±St. dev. | ±2.18 | ±3.39 | ±2.89 | ±2.59 | |||
| Mitotic count | 0.07 2 | 0.13 2 | 0.76 2 | ||||
| Mean 3 | 9.00 | 4.58 | 5.52 | 9.29 | |||
| ±St. Dev. | ±5.83 | ±3.77 | ±3.40 | ±8.33 | |||
| Ciliary body involvement n (%) | 0.72 1 | 0.58 1 | 0.009 1 | ||||
| No | 8 (89) | 10 (83) | 11 (48) | 8 (57) | |||
| Yes | 1 (11) | 2 (17) | 12 (52) | 6 (43) | |||
| Extra-ocular extension | - | 0.12 1 | 0.67 1 | ||||
| <5 mm | 0 (0) | 1 (50) | 4 (75) | 0 (0) | |||
| >5 mm | 0 (0) | 1 (50) | 1 (25) | 1 (100) | |||
| Chr. 8q status n (%) | 0.10 1 | 0.34 1 | <0.001 1 | ||||
| Normal | 7 (78) | 5 (42) | 4 (17) | 1 (7) | |||
| Aberrant | 2 (22) | 7 (58) | 19 (83) | 14 (93) | |||
| Chr. 6p status n (%) | 0.37 1 | 0.57 1 | <0.001 1 | ||||
| Normal | 2 (22) | 1 (8) | 20 (87) | 12 (80) | |||
| Aberrant | 7 (78) | 11 (92) | 3 (13) | 3 (20) |
| Characteristics | D3 Q209L (n = 15) | D3 Q209P (n = 6) | M3 Q209L (n = 27) | M3 Q209P (n = 11) | Disomy 3 Tumors Q209L versus Q209P p-Value | Monosomy 3 Tumors Q209L versus Q209P p-Value | Monosomy 3 versus Disomy 3 p-Value |
|---|---|---|---|---|---|---|---|
| Gender n (%) | 0.48 1 | 0.96 1 | 0.21 1 | ||||
| Female | 5 (33) | 3 (50) | 15 (56) | 6 (55) | |||
| Male | 10 (66) | 3 (50) | 12 (44) | 5 (45) | |||
| Age at enucleation | 0.92 2 | 0.70 2 | 0.15 2 | ||||
| Mean | 53.78 | 54.64 | 64.19 | 66.01 | |||
| ±St. dev. | ± 19.21 | ±15.39 | ±14.20 | ±12.23 | |||
| Affected eye n (%) | 0.48 1 | 0.39 1 | 0.28 1 | ||||
| Right eye | 5 (33) | 3 (50) | 13 (48) | 7 (64) | |||
| Left eye | 10 (66) | 3 (50) | 14 (52) | 4 (36) | |||
| Survival n (%) | 0.88 1 | 0.75 1 | 0.001 1 | ||||
| Alive | 7 (47) | 3 (50) | 3 (11) | 2 (18) | |||
| Cause of Death: | |||||||
| Melanoma-related | 4 (27) | 1 (17) | 22 (81) | 8 (73) | |||
| Different cause | 3 (20) | 1 (17) | 1 (4) | 1 (9) | |||
| Unknown | 1 (7) | 1 (17) | 1 (4) | 0 (0) | |||
| Metastases n (%) | 0.63 1 | 0.55 1 | <0.001 1 | ||||
| No | 11 (73) | 5 (83) | 5 (19) | 3 (27) | |||
| Yes | 4 (27) | 1 (17) | 22 (81) | 8 (73) | |||
| Basal diameter | 0.34 2 | 0.27 2 | 0.30 2 | ||||
| Mean (mm) | 13.2 | 11.83 | 13.56 | 15.55 | |||
| ±St. dev. | ±2.6 | ±2.9 | ±3.3 | ±5.3 | |||
| Prominence | 0.67 2 | 0.56 2 | 0.86 2 | ||||
| Mean (mm) | 6.97 | 7.67 | 7.80 | 8.36 | |||
| ±St. dev. | ±2.7 | ±3.5 | ±2.8 | ±2.7 | |||
| Mitotic count | 0.29 2 | 0.22 2 | 0.96 2 | ||||
| Mean 3 | 7.20 | 4.67 | 5.81 | 9.64 | |||
| ±St. dev. | ±5.38 | ±4.41 | ±3.37 | ±9.39 | |||
| Ciliary body involvement n (%) | 0.24 1 | 0.13 1 | 0.011 1 | ||||
| No | 12 (80) | 6 (100) | 10 (37) | 8 (73) | |||
| Yes | 3 (20) | 0 (0) | 17 (63) | 3 (27) | |||
| Cell type | 0.78 1 | 0.70 1 | 0.043 1 | ||||
| Spindle | 8 (53) | 3 (50) | 5 (19) | 3 (27) | |||
| Mixed | 6 (40) | 3 (50) | 17 (63) | 7 (64) | |||
| Epithelioid | 1 (7) | 0 (0) | 5 (19) | 1 (9) | |||
| Extra-ocular extension | - | 0.12 1 | 0.67 1 | ||||
| <5 mm | 1 (50) | 0 (0) | 4 (80) | 0 (0) | |||
| >5 mm | 1 (50) | 0 (0) | 1 (20) | 1 (100) | |||
| Chr. 8q status n (%) | 0.16 1 | 0.64 1 | <0.001 1 | ||||
| Normal | 10 (66) | 2 (33) | 4 (15) | 1 (9) | |||
| Aberrant | 5 (33) | 4 (67) | 23 (85) | 10 (91) | |||
| Chr. 6 status n (%) | 0.24 1 | 0.80 1 | <0.001 1 | ||||
| Normal | 3 (20) | 0 (0) | 23 (85) | 9 (82) | |||
| Aberrant | 12 (80) | 6 (100) | 4 (15) | 2 (18) |
| mRNA Probe | Illumina | D3 GNA11 n = 9 Mean ± STD | D3 GNAQ n = 12 Mean ± STD | M3 GNA11 n = 23 Mean ± STD | M3 GNAQ n = 15 Mean ± STD | M3 GNA11 versus M3 GNAQ p-Value | D3 GNA11 versus D3 GNAQ p-Value | All M3 versus All D3 p-Value |
|---|---|---|---|---|---|---|---|---|
| HLA_A | ILMN 1671054 | 10.69 ±0.74 | 10.89 ±0.73 | 11.85 ±0.84 | 11.67 ±0.68 | 0.30 | 0.34 | <0.001 |
| HLA_A | ILMN 2203950 | 13.16 ±0.96 | 13.38 ±0.64 | 14.26 ±0.53 | 14.13 ±0.49 | 0.42 | 0.13 | <0.001 |
| HLA_A | ILMN 2186806 | 9.87 ±2.08 | 9.89 ±0.89 | 11.31 ±1.07 | 11.13 ±1.22 | 0.79 | 0.27 | <0.001 |
| HLA_B | ILMN 1778401 | 10.25 ±2.24 | 10.18 ±1.09 | 11.98 ±1.40 | 12.05 ±1.10 | 0.95 | 0.21 | <0.001 |
| HLA_C | ILMN 2150787 | 6.57 ±0.74 | 6.22 ±0.41 | 6.52 ±0.64 | 6.53 ±0.75 | 0.46 | 0.21 | 0.38 |
| HLA_C | ILMN 1721113 | 7.63 ±1.18 | 7.46 ±0.53 | 8.20 ±1.21 | 8.35 ±1.13 | 0.70 | 0.80 | 0.013 |
| HLA_E | ILMN 1765258 | 11.10 ±1.52 | 10.84 ±0.59 | 11.83 ±1.11 | 11.73 ±0.85 | 0.74 | 0.75 | <0.001 |
| HLA_F | ILMN 1762861 | 7.65 ±1.18 | 7.65 ±0.70 | 8.64 ±1.17 | 8.46 ±0.97 | 0.68 | 0.59 | <0.001 |
| HLA_G | ILMN 1656670 | 8.25 ±1.48 | 8.08 ±0.64 | 9.05 ±0.76 | 8.88 ±0.70 | 0.39 | 0.70 | <0.001 |
| CD68 | ILMN 1714861 | 10.36 ±1.30 | 10.37 ±0.86 | 11.07 ±0.75 | 11.01 ±0.82 | 0.95 | 0.70 | 0.008 |
| CD3D | ILMN 2261416 | 7.07 ±1.70 | 6.66 ±0.40 | 7.28 ±1.12 | 7.28 ±1.16 | 0.95 | 0.92 | 0.03 |
| CD3D | ILMN 2325837 | 7.14 ±1.71 | 6.57 ±0.43 | 7.37 ±1.23 | 7.31 ±1.16 | 0.98 | 0.46 | 0.022 |
| CD3E | ILMN 1739794 | 6.56 ±0.50 | 6.38 ±0.07 | 6.51 ±0.28 | 6.57 ±0.31 | 0.55 | 0.50 | 0.19 |
| CD3G | ILMN 1717197 | 6.68 ±0.45 | 6.53 ±0.11 | 6.63 ±0.28 | 6.57 ±0.29 | 0.39 | 0.64 | 0.77 |
| CD8A | ILMN 1768482 | 7.30 ±2.06 | 6.78 ±0.45 | 7.45 ±1.23 | 7.54 ±1.29 | 0.86 | 0.86 | 0.06 |
| CD8A | ILMN 1760374 | 6.58 ±0.58 | 6.36 ±0.10 | 6.47 ±0.29 | 6.54 ±0.45 | 0.63 | 0.27 | 0.25 |
| CD8A | ILMN 2353732 | 7.22 ±2.11 | 6.65 ±0.43 | 7.42 ±1.25 | 7.46 ±1.31 | 0.98 | 0.80 | 0.022 |
| CD4 | ILMN 1727284 | 6.65 ±0.49 | 6.54 ±0.29 | 6.69 ±0.24 | 6.73 ±0.36 | 0.98 | 0.86 | 0.044 |
| mRNA Probe | Illumina | D3 Q209L n = 15 Mean ± STD | D3 Q209P n = 6 Mean ± STD | M3 Q209L n = 27 Mean ± STD | M3 Q209P n = 11 Mean ± STD | M3 Q209L versus M3 Q209P p-Value | D3 Q209L versus D3 Q209P p-Value | All M3 versus All D3 p-Value |
|---|---|---|---|---|---|---|---|---|
| HLA_A | ILMN 1671054 | 10.88 ±0.73 | 10.63 ±0.72 | 11.85 ±0.79 | 11.60 ±0.74 | 0.30 | 0.79 | <0.001 |
| HLA_A | ILMN 2203950 | 13.35 ±0.80 | 13.12 ±0.75 | 14.23 ±0.51 | 14.16 ±0.55 | 0.87 | 0.91 | <0.001 |
| HLA_A | ILMN 2186806 | 10.08 ±1.63 | 9.39 ±0.93 | 11.20 ±1.10 | 11.33 ±1.22 | 0.70 | 0.37 | <0.001 |
| HLA_B | ILMN 1778401 | 10.34 ±1.78 | 9.89 ±1.26 | 11.99 ±1.34 | 12.05 ±1.17 | 1.00 | 0.97 | <0.001 |
| HLA_C | ILMN 2150787 | 6.34 ±0.66 | 6.44 ±0.37 | 6.44 ±0.63 | 6.74 ±0.77 | 0.46 | 0.51 | 0.38 |
| HLA_C | ILMN 1721113 | 7.57 ±0.95 | 7.43 ±0.58 | 8.16 ±1.19 | 8.50 ±1.12 | 0.42 | 0.97 | 0.013 |
| HLA_E | ILMN 1765258 | 11.01 ±1.21 | 10.81 ±0.61 | 11.82 ±1.05 | 11.71 ±0.93 | 0.63 | 1.00 | <0.001 |
| HLA_F | ILMN 1762861 | 7.72 ±1.00 | 7.49 ±0.69 | 8.58 ±1.13 | 8.54 ±1.02 | 1.00 | 0.67 | <0.001 |
| HLA_G | ILMN 1656670 | 8.32 ±1.16 | 7.73 ±0.61 | 9.00 ±0.76 | 8.96 ±0.69 | 0.85 | 0.15 | <0.001 |
| CD68 | ILMN 1714861 | 10.39 ±1.16 | 10.30 ±0.75 | 11.07 ±0.74 | 10.99 ±0.87 | 0.80 | 0.85 | 0.008 |
| CD3D | ILMN 2261416 | 6.92 ±1.32 | 6.61 ±0.45 | 7.23 ±1.06 | 7.41 ±1.29 | 0.85 | 0.51 | 0.033 |
| CD3D | ILMN 2325837 | 6.94 ±1.39 | 6.49 ±0.47 | 7.31 ±1.17 | 7.44 ±1.29 | 0.75 | 0.23 | 0.016 |
| CD3E | ILMN 1739794 | 6.49 ±0.39 | 6.37 ±0.09 | 6.51 ±0.26 | 6.60 ±0.35 | 0.56 | 0.41 | 0.19 |
| CD3G | ILMN 1717197 | 6.60 ±0.36 | 6.58 ±0.13 | 6.61 ±0.27 | 6.60 ±0.33 | 0.70 | 0.51 | 0.77 |
| CD8A | ILMN 1768482 | 7.10 ±1.59 | 6.75 ±0.58 | 7.39 ±1.17 | 7.71 ±1.43 | 0.56 | 0.67 | 0.061 |
| CD8A | ILMN 1760374 | 6.50 ±0.46 | 6.35 ±0.09 | 6.47 ±0.27 | 6.57 ±0.52 | 0.80 | 0.41 | 0.25 |
| CD8A | ILMN 2353732 | 7.00 ±1.63 | 6.62 ±0.55 | 7.35 ±1.19 | 7.64 ±1.44 | 0.52 | 0.46 | 0.019 |
| CD4 | ILMN 1727284 | 6.62 ±0.41 | 6.51 ±0.21 | 6.70 ±0.25 | 6.72 ±0.40 | 0.90 | 0.61 | 0.036 |
| Gene | Probe |
|---|---|
| HLA-A | ILMN_1671054 |
| HLA-A | ILMN_2203950 |
| HLA-A | ILMN_2186806 |
| HLA-B | ILMN_1778401 |
| HLA-C | ILMN_2150787 |
| HLA-C | ILMN_1721113 |
| HLA-E | ILMN_1765258 |
| HLA-F | ILMN_1762861 |
| HLA-G | ILMN_1656670 |
| CD68 | ILMN 1714861 |
| CD3D | ILMN 2261416 |
| CD3D | ILMN 2325837 |
| CD3E | ILMN 1739794 |
| CD3G | ILMN 1717197 |
| CD8A | ILMN 1768482 |
| CD8A | ILMN 1760374 |
| CD8A | ILMN 2353732 |
| CD4 | ILMN 1727284 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
van Weeghel, C.; Wierenga, A.P.A.; Versluis, M.; van Hall, T.; van der Velden, P.A.; Kroes, W.G.M.; Pfeffer, U.; Luyten, G.P.M.; Jager, M.J. Do GNAQ and GNA11 Differentially Affect Inflammation and HLA Expression in Uveal Melanoma? Cancers 2019, 11, 1127. https://doi.org/10.3390/cancers11081127
van Weeghel C, Wierenga APA, Versluis M, van Hall T, van der Velden PA, Kroes WGM, Pfeffer U, Luyten GPM, Jager MJ. Do GNAQ and GNA11 Differentially Affect Inflammation and HLA Expression in Uveal Melanoma? Cancers. 2019; 11(8):1127. https://doi.org/10.3390/cancers11081127
Chicago/Turabian Stylevan Weeghel, Christiaan, Annemijn P. A. Wierenga, Mieke Versluis, Thorbald van Hall, Pieter A. van der Velden, Wilma G. M. Kroes, Ulrich Pfeffer, Gregorius P. M. Luyten, and Martine J. Jager. 2019. "Do GNAQ and GNA11 Differentially Affect Inflammation and HLA Expression in Uveal Melanoma?" Cancers 11, no. 8: 1127. https://doi.org/10.3390/cancers11081127
APA Stylevan Weeghel, C., Wierenga, A. P. A., Versluis, M., van Hall, T., van der Velden, P. A., Kroes, W. G. M., Pfeffer, U., Luyten, G. P. M., & Jager, M. J. (2019). Do GNAQ and GNA11 Differentially Affect Inflammation and HLA Expression in Uveal Melanoma? Cancers, 11(8), 1127. https://doi.org/10.3390/cancers11081127






