Lactate Dehydrogenases as Metabolic Links between Tumor and Stroma in the Tumor Microenvironment
Abstract
1. Introduction
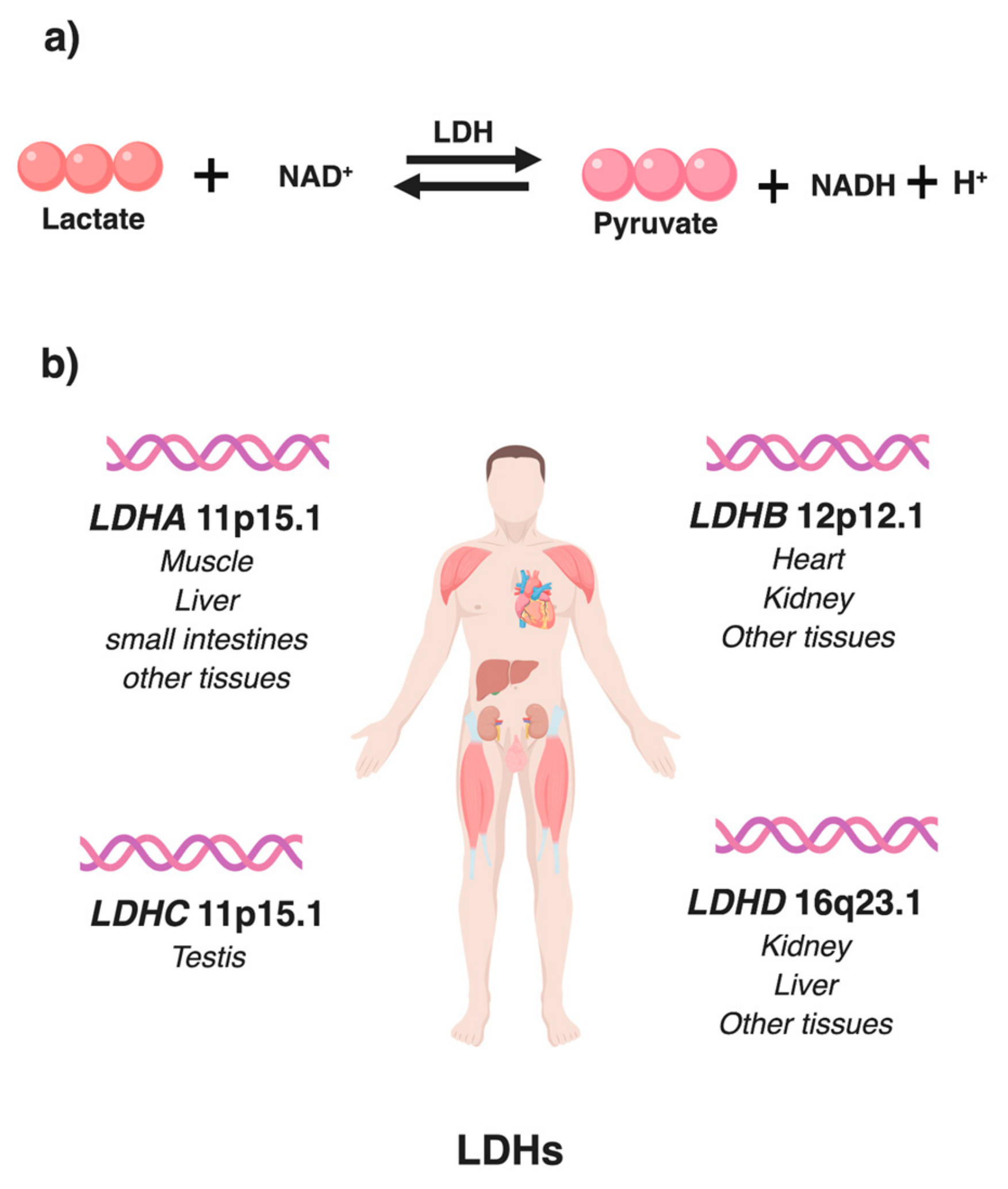
2. Lactate Dehydrogenase
3. LDHA in Cancers
3.1. HIF-1 and LDHA
3.2. c-Myc and LDHA
3.3. FOXM1 and LDHA
3.4. KLF4 and LDHA
3.5. Other Pathways
4. LDHB in Cancers
4.1. LDHB Promoter Hypermethylation
4.2. LDHB Regulation by Different Pathways
5. LDH-Mediated Metabolic Exchange between Cancer Cells and Stromal CAFs
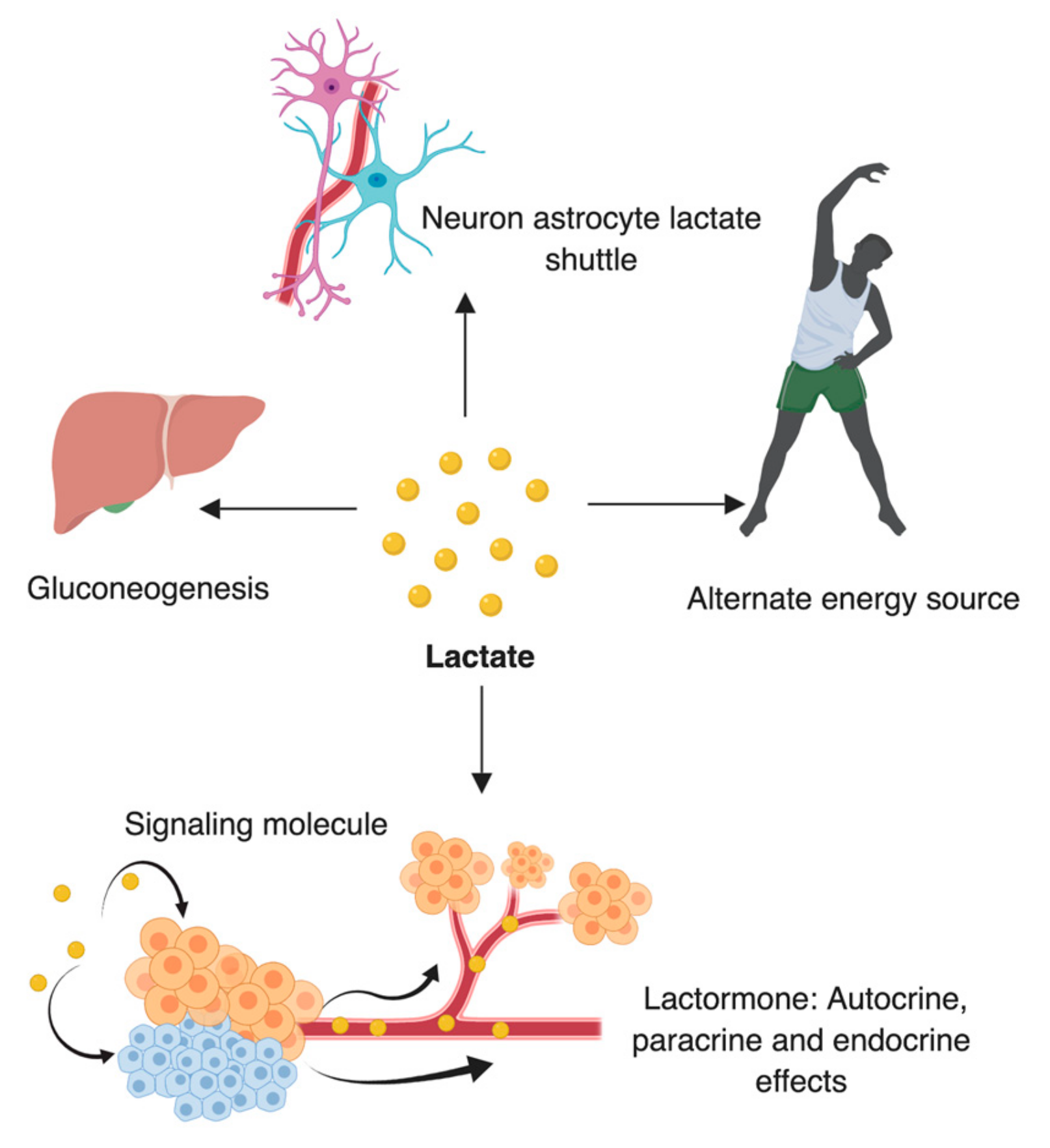
6. Lactate: A Substrate for LDH
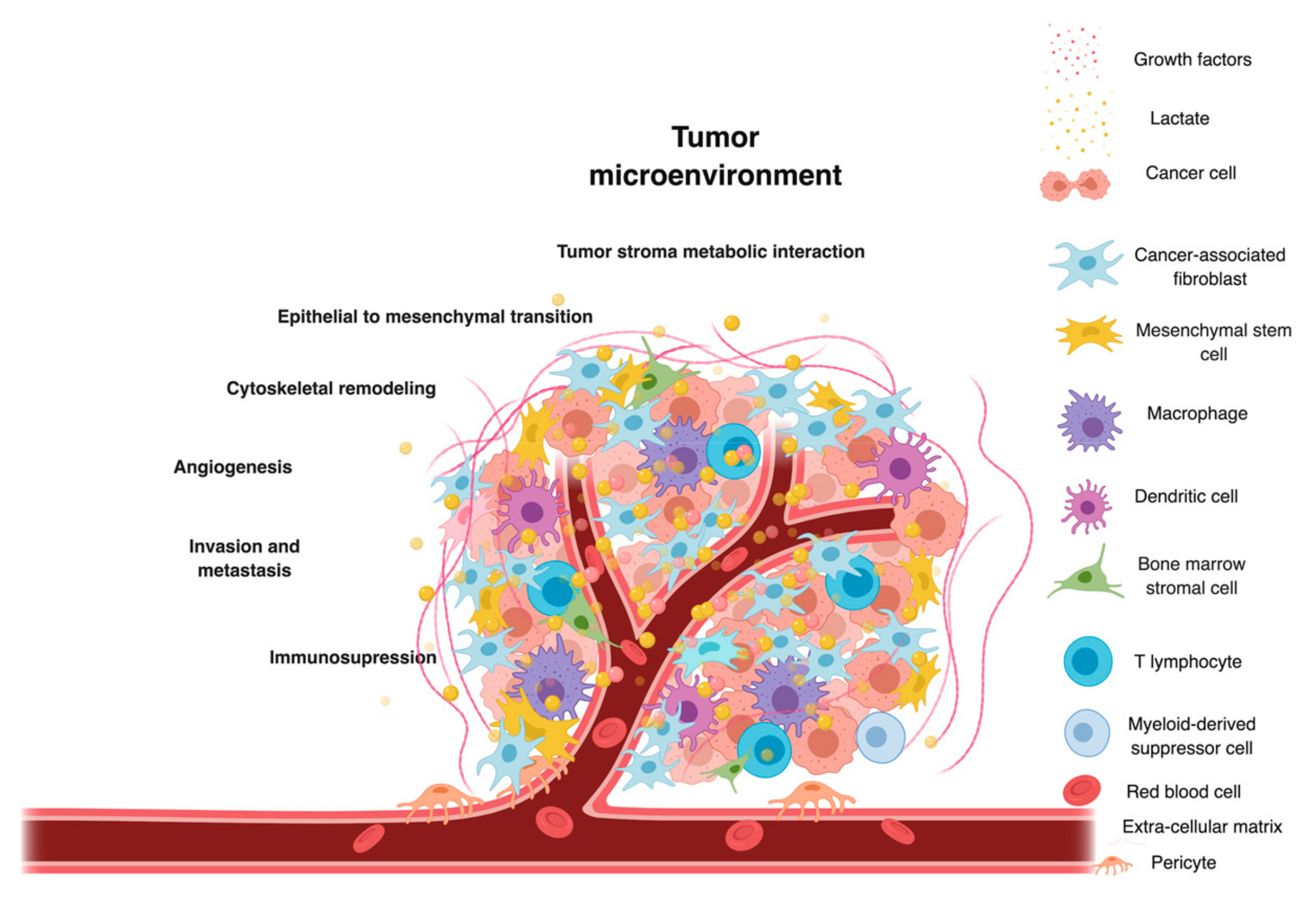
7. Role of Lactate in Regulating Immunometabolism in the TME
8. Serum LDH Profile in Cancers and Clinical Relevance
9. Future Directions in Therapeutic Designing
9.1. Small Molecule Inhibitors
9.2. Molecular Inhibition of LDHA and LDHB
9.3. Blocking Lactate Exchange between Tumor and Stroma
10. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Egeblad, M.; Nakasone, E.S.; Werb, Z. Tumors as organs: Complex tissues that interface with the entire organism. Dev. Cell 2010, 6, 884–901. [Google Scholar] [CrossRef]
- Lyssiotis, C.A.; Kimmelman, A.C. Metabolic Interactions in the Tumor Microenvironment. Trends Cell Biol. 2017, 11, 863–875. [Google Scholar] [CrossRef] [PubMed]
- Pavlova, N.N.; Thompson, C.B. The Emerging Hallmarks of Cancer Metabolism. Cell Metab. 2016, 1, 27–47. [Google Scholar] [CrossRef] [PubMed]
- Swierczynski, J.; Hebanowska, A.; Sledzinski, T. Role of abnormal lipid metabolism in development, progression, diagnosis and therapy of pancreatic cancer. World J. Gastroenterol. 2014, 20, 2279–3035. [Google Scholar] [CrossRef]
- Vander Heiden, M.G.; Cantley, L.C.; Thompson, C.B. Understanding the Warburg effect: The metabolic requirements of cell proliferation. Science 2009, 5930, 1029–1033. [Google Scholar] [CrossRef]
- Draoui, N.; Feron, O. Lactate shuttles at a glance: From physiological paradigms to anti-cancer treatments. Dis. Model Mech. 2011, 6, 727–732. [Google Scholar] [CrossRef]
- Valvona, C.J.; Fillmore, H.L.; Nunn, P.B.; Pilkington, G.J. The Regulation and Function of Lactate Dehydrogenase A: Therapeutic Potential in Brain Tumor. Brain Pathol. 2016, 26, 3–17. [Google Scholar] [CrossRef] [PubMed]
- Cui, J.; Shi, M.; Xie, D.; Wei, D.; Jia, Z.; Zheng, S.; Gao, Y.; Huang, S.; Xie, K. FOXM1 promotes the warburg effect and pancreatic cancer progression via transactivation of LDHA expression. Clin. Cancer Res. 2014, 10, 2595–2606. [Google Scholar] [CrossRef] [PubMed]
- Shi, M.; Cui, J.; Du, J.; Wei, D.; Jia, Z.; Zhang, J.; Zhu, Z.; Gao, Y.; Xie, K. A novel KLF4/LDHA signaling pathway regulates aerobic glycolysis in and progression of pancreatic cancer. Clin. Cancer Res. 2014, 16, 4370–4380. [Google Scholar] [CrossRef]
- Schlichtholz, B.; Turyn, J.; Goyke, E.; Biernacki, M.; Jaskiewicz, K.; Sledzinski, Z.; Swierczynski, J. Enhanced citrate synthase activity in human pancreatic cancer. Pancreas 2005, 30, 99–104. [Google Scholar] [CrossRef]
- Kurpińska, A.; Suraj, J.; Bonar, E.; Zakrzewska, A.; Stojak, M.; Sternak, M.; Jasztal, A.; Walczak, M. Proteomic characterization of early lung response to breast cancer metastasis in mice. Exp. Mol. Pathol. 2019, 18, 30482–30489. [Google Scholar] [CrossRef]
- Su, Y.; Yu, Q.H.; Wang, X.Y.; Yu, L.P.; Wang, Z.F.; Cao, Y.C.; Li, J.D. JMJD2A promotes the Warburg effect and nasopharyngeal carcinoma progression by transactivating LDHA expression. BMC Cancer 2017, 1, 477. [Google Scholar] [CrossRef] [PubMed]
- Jiang, W.; Zhou, F.; Li, N.; Li, Q.; Wang, L. FOXM1-LDHA signaling promoted gastric cancer glycolytic phenotype and progression. Int. J. Clin. Exp. Pathol. 2015, 6, 6756–6763. [Google Scholar]
- Koukourakis, M.I.; Kakouratos, C.; Kalamida, D.; Bampali, Z.; Mavropoulou, S.; Sivridis, E.; Giatromanolaki, A. Hypoxia-inducible proteins HIF1α and lactate dehydrogenase LDH5, key markers of anaerobic metabolism, relate with stem cell markers and poor post-radiotherapy outcome in bladder cancer. Int. J. Radiat. Biol. 2016, 92, 353–363. [Google Scholar] [CrossRef]
- Giatromanolaki, A.; Sivridis, E.; Gatter, K.C.; Turley, H.; Harris, A.L.; Koukourakis, M.I. Tumour and Angiogenesis Research Group. Lactate dehydrogenase 5 (LDH-5) expression in endometrial cancer relates to the activated VEGF/VEGFR2(KDR) pathway and prognosis. Gynecol. Oncol. 2006, 103, 912–918. [Google Scholar] [CrossRef]
- Markert, C.L.; Shaklee, J.B.; Whitt, G.S. Evolution of a gene. Multiple genes for LDH isozymes provide a model of the evolution of gene structure, function and regulation. Science 1975, 189, 102–114. [Google Scholar] [CrossRef]
- Flick, M.J.; Konieczny, S.F. Identification of putative mammalian D-lactate dehydrogenase enzymes. Biochem. Biophys. Res. Commun. 2002, 295, 910–916. [Google Scholar] [CrossRef]
- Ewaschuk, J.B.; Naylor, J.M.; Zello, G.A. D-lactate in human and ruminant metabolism. J. Nutr. 2005, 135, 1619–1625. [Google Scholar] [CrossRef] [PubMed]
- Kong, L.; Du, W.; Cui, Z.; Wang, L.; Yang, Z.; Zhang, H.; Lin, D. Expression of lactate dehydrogenase C in MDA-MB-231 cells and its role in tumor invasion and migration. Mol. Med. Rep. 2016, 13, 3533–3538. [Google Scholar] [CrossRef] [PubMed]
- Hua, Y.; Liang, C.; Zhu, J.; Miao, C.; Yu, Y.; Xu, A.; Zhang, J.; Li, P.; Li, S.; Bao, M.; et al. Expression of lactate dehydrogenase C correlates with poor prognosis in renal cell carcinoma. Tumour Biol. 2017, 39, 1010428317695968. [Google Scholar] [CrossRef] [PubMed]
- Monroe, G.R.; van Eerde, A.M.; Tessadori, F.; Duran, K.J.; Savelberg, S.M.C.; van Alfen, J.C.; Terhal, P.A.; van der Crabben, S.N.; Lichtenbelt, K.D.; Fuchs, S.A.; et al. Identification of human D lactate dehydrogenase deficiency. Nat. Commun. 2019, 10, 1477. [Google Scholar] [CrossRef]
- Wang, Y.; Li, G.; Wan, F.; Dai, B.; Ye, D. Prognostic value of D-lactate dehydrogenase in patients with clear cell renal cell carcinoma. Oncol. Lett. 2018, 16, 866–874. [Google Scholar] [CrossRef]
- Ždralević, M.; Brand, A.; Di Ianni, L.; Dettmer, K.; Reinders, J.; Singer, K.; Peter, K.; Schnell, A.; Bruss, C.; Decking, S.M.; et al. Double genetic disruption of lactate dehydrogenases A and B is required to ablate the “Warburg effect” restricting tumor growth to oxidative metabolism. J. Biol. Chem. 2018, 293, 15947–15961. [Google Scholar]
- Read, J.A.; Winter, V.J.; Eszes, C.M.; Sessions, R.B.; Brady, R.L. Structural basis for altered activity of M- and H-isozyme forms of human lactate dehydrogenase. Proteins 2001, 43, 175–185. [Google Scholar] [CrossRef]
- Świderek, K.; Paneth, P. Binding ligands and cofactor to L-lactate dehydrogenase from human skeletal and heart muscles. J. Phys. Chem. B 2011, 115, 6366–6376. [Google Scholar] [CrossRef]
- Swiderek, K.; Paneth, P. Differences and similarities in binding of pyruvate and L-lactate in the active site of M4 and H4 isoforms of human lactate dehydrogenase. Arch. Biochem. Biophys. 2011, 505, 33–41. [Google Scholar] [CrossRef]
- Warburg, O. On the origin of cancer cells. Science 1956, 123, 309–314. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Chao, M.; Wu, H. Central role of lactate and proton in cancer cell resistance to glucose deprivation and its clinical translation. Signal Transduct. Target Ther. 2017, 2, 16047. [Google Scholar] [CrossRef]
- Cairns, R.A.; Harris, I.S.; Mak, T.W. Regulation of cancer cell metabolism. Nat. Rev. Cancer. 2011, 11, 85–95. [Google Scholar] [CrossRef]
- Schulze, A.; Harris, A.L. How cancer metabolism is tuned for proliferation and vulnerable to disruption. Nature 2012, 491, 364–373. [Google Scholar] [CrossRef] [PubMed]
- Jiang, F.; Ma, S.; Xue, Y.; Hou, J.; Zhang, Y. LDH-A promotes malignant progression via activation of epithelial-to-mesenchymal transition and conferring stemness in muscle-invasive bladder cancer. Biochem. Biophys. Res. Commun. 2016, 469, 985–992. [Google Scholar] [CrossRef] [PubMed]
- Arseneault, R.; Chien, A.; Newington, J.T.; Rappon, T.; Harris, R.; Cumming, R.C. Attenuation of LDHA expression in cancer cells leads to redox-dependent alterations in cytoskeletal structure and cell migration. Cancer Lett. 2013, 338, 255–266. [Google Scholar] [CrossRef]
- Valvona, C.J.; Fillmore, H.L. Oxamate, but Not Selective Targeting of LDH-A, Inhibits Medulloblastoma Cell Glycolysis, Growth and Motility. Brain Sci. 2018, 8, 56. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Yang, Z.; Chen, Z.; Chen, R.; Zhao, D.; Zhou, Y.; Qiao, L. Effects of the suppression of lactate dehydrogenase A on the growth and invasion of human gastric cancer cells. Oncol Rep. 2015, 33, 157–162. [Google Scholar] [CrossRef] [PubMed]
- Koh, M.Y.; Powis, G. Passing the baton: The HIF switch. Trends Biochem. Sci. 2012, 37, 364–372. [Google Scholar] [CrossRef]
- Semenza, G.L. Hypoxia-inducible factors in physiology and medicine. Cell 2012, 148, 399–408. [Google Scholar] [CrossRef]
- Augoff, K.; Hryniewicz-Jankowska, A.; Tabola, R. Lactate dehydrogenase 5: An old friend and a new hope in the war on cancer. Cancer Lett. 2015, 358, 1–7. [Google Scholar] [CrossRef]
- Semenza, G.L.; Jiang, B.H.; Leung, S.W.; Passantino, R.; Concordet, J.P.; Maire, P.; Giallongo, A. Hypoxia response elements in the aldolase, A.; enolase 1, and lactate dehydrogenase A gene promoters contain essential binding sites for hypoxia-inducible factor 1. J. Biol. Chem. 1996, 271, 32529–32537. [Google Scholar] [CrossRef]
- Koukourakis, M.I.; Giatromanolaki, A.; Sivridis, E.; Bougioukas, G.; Didilis, V.; Gatter, K.C.; Harris, A.L. Tumour and Angiogenesis Research Group. Lactate dehydrogenase-5 (LDH-5) overexpression in non-small-cell lung cancer tissues is linked to tumour hypoxia, angiogenic factor production and poor prognosis. Br. J. Cancer 2003, 89, 877–885. [Google Scholar] [CrossRef]
- Kolev, Y.; Uetake, H.; Takagi, Y.; Sugihara, K. Lactate dehydrogenase-5 (LDH-5) expression in human gastric cancer: Association with hypoxia-inducible factor (HIF-1alpha) pathway, angiogenic factors production and poor prognosis. Ann. Surg. Oncol. 2008, 15, 2336–2344. [Google Scholar] [CrossRef]
- Shim, H.; Dolde, C.; Lewis, B.C.; Wu, C.S.; Dang, G.; Jungmann, R.A.; Dalla-Favera, R.; Dang, C.V. c-Myc transactivation of LDH-A: Implications for tumor metabolism and growth. Proc. Natl. Acad. Sci. USA 1997, 94, 6658–6663. [Google Scholar] [CrossRef]
- Lewis, B.C.; Prescott, J.E.; Campbell, S.E.; Shim, H.; Orlowski, R.Z.; Dang, C.V. Tumor induction by the c-Myc target genes rcl and lactate dehydrogenase A. Cancer Res. 2000, 60, 6178–6183. [Google Scholar] [PubMed]
- Shim, H.; Chun, Y.S.; Lewis, B.C.; Dang, C.V. A unique glucose-dependent apoptotic pathway induced by c-Myc. Proc. Natl. Acad. Sci. USA 1998, 95, 1511–1516. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; Yu, Z.; Wang, J.; Zhou, L.; Zhang, J.; Yao, B.; Dou, J.; Qiu, Z.; Huang, C. Krüppel-Like Factor 4 Inhibits Pancreatic Cancer Epithelial-to-Mesenchymal Transition and Metastasis by Down-Regulating Caveolin-1 Expression. Cell. Physiol. Biochem. 2018, 46, 238–252. [Google Scholar] [CrossRef]
- Jin, L.; Chun, J.; Pan, C.; Alesi, G.N.; Li, D.; Magliocca, K.R.; Kang, Y.; Chen, Z.G.; Shin, D.M.; Khuri, F.R.; et al. Phosphorylation-mediated activation of LDHA promotes cancer cell invasion and tumour metastasis. Oncogene 2017, 36, 3797–3806. [Google Scholar] [CrossRef] [PubMed]
- Li, S.S.; Pan, Y.E.; Sharief, F.S.; Evans, M.J.; Lin, M.F.; Clinton, G.M.; Holbrook, J.J. Cancer-associated lactate dehydrogenase is a tyrosylphosphorylated form of human LDH-A, skeletal muscle isoenzyme. Cancer Investig. 1988, 6, 93–101. [Google Scholar] [CrossRef]
- Liu, J.; Chen, G.; Liu, Z.; Liu, S.; Cai, Z.; You, P.; Ke, Y.; Lai, L.; Huang, Y.; Gao, H.; et al. Aberrant FGFR Tyrosine Kinase Signaling Enhances the Warburg Effect by Reprogramming LDH Isoform Expression and Activity in Prostate Cancer. Cancer Res. 2018, 78, 4459–4470. [Google Scholar] [CrossRef] [PubMed]
- Dennison, J.B.; Molina, J.R.; Mitra, S.; González-Angulo, A.M.; Balko, J.M.; Kuba, M.G.; Sanders, M.E.; Pinto, J.A.; Gómez, H.L.; Arteaga, C.L.; et al. Lactate dehydrogenase B: A metabolic marker of response to neoadjuvant chemotherapy in breast cancer. Clin. Cancer Res. 2013, 19, 3703–3713. [Google Scholar] [CrossRef]
- Sun, W.; Zhang, X.; Ding, X.; Li, H.; Geng, M.; Xie, Z.; Wu, H.; Huang, M. Lactate dehydrogenase B is associated with the response to neoadjuvant chemotherapy in oral squamous cell carcinoma. PLoS ONE 2015, 10, e0125976. [Google Scholar] [CrossRef]
- Li, C.; Chen, Y.; Bai, P.; Wang, J.; Liu, Z.; Wang, T.; Cai, Q. LDHB may be a significant predictor of poor prognosis in osteosarcoma. Am. J. Transl. Res. 2016, 8, 4831–4843. [Google Scholar]
- Leiblich, A.; Cross, S.S.; Catto, J.W.; Phillips, J.T.; Leung, H.Y.; Hamdy, F.C.; Rehman, I. Lactate dehydrogenase-B is silenced by promoter hypermethylation in human prostate cancer. Oncogene 2006, 25, 2953–2960. [Google Scholar] [CrossRef] [PubMed]
- Maekawa, M.; Taniguchi, T.; Ishikawa, J.; Sugimura, H.; Sugano, K.; Kanno, T. Promoter hypermethylation in cancer silences LDHB, eliminating lactate dehydrogenase isoenzymes 1-4. Clin. Chem. 2003, 49, 1518–1520. [Google Scholar] [CrossRef]
- Cui, J.; Quan, M.; Jiang, W.; Hu, H.; Jiao, F.; Li, N.; Jin, Z.; Wang, L.; Wang, Y.; Wang, L. Suppressed expression of LDHB promotes pancreatic cancer progression via inducing glycolytic phenotype. Med. Oncol. 2015, 32, 143. [Google Scholar] [CrossRef]
- Brown, N.J.; Higham, S.E.; Perunovic, B.; Arafa, M.; Balasubramanian, S.; Rehman, I. Lactate dehydrogenase-B is silenced by promoter methylation in a high frequency of human breast cancers. PLoS ONE 2013, 8, e57697. [Google Scholar] [CrossRef]
- McCleland, M.L.; Adler, A.S.; Shang, Y.; Hunsaker, T.; Truong, T.; Peterson, D.; Torres, E.; Li, L.; Haley, B.; Stephan, J.P.; et al. An integrated genomic screen identifies LDHB as an essential gene for triple-negative breast cancer. Cancer Res. 2012, 72, 5812–5823. [Google Scholar] [CrossRef]
- Ding, S.J.; Li, Y.; Shao, X.X.; Zhou, H.; Zeng, R.; Tang, Z.Y.; Xia, Q.C. Proteome analysis of hepatocellular carcinoma cell strains, MHCC97-H and MHCC97-L, with different metastasis potentials. Proteomics 2004, 4, 982–994. [Google Scholar] [CrossRef] [PubMed]
- Chen, R.; Zhou, X.; Yu, Z.; Liu, J.; Huang, G. Low Expression of LDHB Correlates with Unfavorable Survival in Hepatocellular Carcinoma: Strobe-Compliant Article. Medicine 2015, 94, e1583. [Google Scholar] [CrossRef]
- Liao, A.C.; Li, C.F.; Shen, K.H.; Chien, L.H.; Huang, H.Y.; Wu, T.F. Loss of lactate dehydrogenase B subunit expression is correlated with tumour progression and independently predicts inferior disease-specific survival in urinary bladder urothelial carcinoma. Pathology 2011, 43, 707–712. [Google Scholar] [CrossRef]
- Zha, X.; Wang, F.; Wang, Y.; He, S.; Jing, Y.; Wu, X.; Zhang, H. Lactate dehydrogenase B is critical for hyperactive mTOR-mediated tumorigenesis. Cancer Res. 2011, 71, 13–18. [Google Scholar] [CrossRef]
- Brisson, L.; Bański, P.; Sboarina, M.; Dethier, C.; Danhier, P.; Fontenille, M.J.; Van Hée, V.F.; Vazeille, T.; Tardy, M.; Falces, J.; et al. Lactate Dehydrogenase B Controls Lysosome Activity and Autophagy in Cancer. Cancer Cell 2016, 30, 418–431. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.; Yan, H.; An, S.; Shen, M.; Jia, W.; Zhang, R.; Zhao, L.; Huang, G.; Liu, J. SIRT5-mediated deacetylation of LDHB promotes autophagy and tumorigenesis in colorectal cancer. Mol. Oncol. 2019, 13, 358–375. [Google Scholar] [CrossRef]
- Zhang, W.; Tong, D.; Liu, F.; Li, D.; Li, J.; Cheng, X.; Wang, Z. RPS7 inhibits colorectal cancer growth via decreasing HIF-1α-mediated glycolysis. Oncotarget 2016, 7, 5800–5814. [Google Scholar]
- Kim, J.H.; Kim, E.L.; Lee, Y.K.; Park, C.B.; Kim, B.W.; Wang, H.J.; Yoon, C.H.; Lee, S.J.; Yoon, G. Decreased lactate dehydrogenase B expression enhances claudin 1-mediated hepatoma cell invasiveness via mitochondrial defects. Exp. Cell Res. 2011, 317, 1108–1118. [Google Scholar] [CrossRef]
- Wu, G.; Yuan, S.; Chen, Z.; Chen, G.; Fan, Q.; Dong, H.; Ye, F.; Li, J.; Zhu, X. The KLF14 Transcription Factor Regulates Glycolysis by Downregulating LDHB in Colorectal Cancer. Int. J. Biol. Sci. 2019, 15, 628–635. [Google Scholar] [CrossRef]
- McCleland, M.L.; Adler, A.S.; Deming, L.; Cosino, E.; Lee, L.; Blackwood, E.M.; Solon, M.; Tao, J.; Li, L.; Shames, D.; et al. Lactate dehydrogenase B is required for the growth of KRAS-dependent lung adenocarcinomas. Clin. Cancer Res. 2013, 19, 773–784. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. The hallmarks of cancer. Cell 2000, 100, 57–70. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [PubMed]
- Tennant, D.A.; Durán, R.V.; Gottlieb, E. Targeting metabolic transformation for cancer therapy. Nat. Rev. Cancer 2010, 10, 267–277. [Google Scholar] [CrossRef]
- Kim, J. Regulation of Immune Cell Functions by Metabolic Reprogramming. J. Immunol. Res. 2018, 2018, 8605471. [Google Scholar] [CrossRef]
- Quail, D.F.; Joyce, J.A. Microenvironmental regulation of tumor progression and metastasis. Nat. Med. 2013, 19, 1423–1437. [Google Scholar] [CrossRef]
- Bussard, K.M.; Mutkus, L.; Stumpf, K.; Gomez-Manzano, C.; Marini, F.C. Tumor-associated stromal cells as key contributors to the tumor microenvironment. Breast Cancer Res. 2016, 18, 84. [Google Scholar] [CrossRef]
- LeBleu, V.S.; Kalluri, R. A peek into cancer-associated fibroblasts: Origins, functions and translational impact. Dis. Model. Mech. 2018, 11, dmm029447. [Google Scholar] [CrossRef]
- Schwörer, S.; Vardhana, S.A.; Thompson, C.B. Cancer Metabolism Drives a Stromal Regenerative Response. Cell Metab. 2019, 29, 576–591. [Google Scholar] [CrossRef]
- Tao, L.; Huang, G.; Song, H.; Chen, Y.; Chen, L. Cancer associated fibroblasts: An essential role in the tumor microenvironment. Oncol. Lett. 2017, 14, 2611–2620. [Google Scholar] [CrossRef]
- Shi, Y.; Du, L.; Lin, L.; Wang, Y. Tumour-associated mesenchymal stem/stromal cells: Emerging therapeutic targets. Nat. Rev. Drug Discov. 2017, 16, 35–52. [Google Scholar] [CrossRef]
- Rattigan, Y.I.; Patel, B.B.; Ackerstaff, E.; Sukenick, G.; Koutcher, J.A.; Glod, J.W.; Banerjee, D. Lactate is a mediator of metabolic cooperation between stromal carcinoma associated fibroblasts and glycolytic tumor cells in the tumor microenvironment. Exp. Cell Res. 2012, 318, 326–335. [Google Scholar] [CrossRef]
- Patel, B.B.; Ackerstaff, E.; Serganova, I.S.; Kerrigan, J.E.; Blasberg, R.G.; Koutcher, J.A.; Banerjee, D. Tumor stroma interaction is mediated by monocarboxylate metabolism. Exp. Cell Res. 2017, 352, 20–33. [Google Scholar] [CrossRef]
- Gouirand, V.; Guillaumond, F.; Vasseur, S. Influence of the Tumor Microenvironment on Cancer Cells Metabolic Reprogramming. Front Oncol. 2018, 8, 117. [Google Scholar] [CrossRef]
- Valkenburg, K.C.; de Groot, A.E.; Pienta, K.J. Targeting the tumour stroma to improve cancer therapy. Nat. Rev. Clin. Oncol. 2018, 15, 366–381. [Google Scholar] [CrossRef]
- Adeva-Andany, M.; López-Ojén, M.; Funcasta-Calderón, R.; Ameneiros-Rodríguez, E.; Donapetry-García, C.; Vila-Altesor, M.; Rodríguez-Seijas, J. Comprehensive review on lactate metabolism in human health. Mitochondrion 2014, 17, 76–100. [Google Scholar] [CrossRef]
- Brooks, G.A. The Science and Translation of Lactate Shuttle Theory. Cell Metab. 2018, 27, 757–785. [Google Scholar] [CrossRef] [PubMed]
- Consoli, A.; Nurjhan, N.; Reilly, J.J., Jr.; Bier, D.M.; Gerich, J.E. Contribution of liver and skeletal muscle to alanine and lactate metabolism in humans. Am. J. Physiol. 1990, 259, E677–E684. [Google Scholar] [CrossRef] [PubMed]
- Waterhouse, C.; Keilson, J. Cori cycle activity in man. J. Clin. Investig. 1969, 48, 2359–2366. [Google Scholar] [CrossRef] [PubMed]
- San-Millán, I.; Brooks, G.A. Reexamining cancer metabolism: Lactate production for carcinogenesis could be the purpose and explanation of the Warburg Effect. Carcinogenesis 2017, 38, 119–133. [Google Scholar] [CrossRef]
- Doherty, J.R.; Cleveland, J.L. Targeting lactate metabolism for cancer therapeutics. J. Clin. Investig. 2013, 123, 3685–3692. [Google Scholar] [CrossRef] [PubMed]
- Vlachostergios, P.J.; Oikonomou, K.G.; Gibilaro, E.; Apergis, G. Elevated lactic acid is a negative prognostic factor in metastatic lung cancer. Cancer Biomark. 2015, 15, 725–734. [Google Scholar] [CrossRef]
- Marchiq, I.; Pouysségur, J. Hypoxia, cancer metabolism and the therapeutic benefit of targeting lactate/H(+) symporters. J. Mol. Med. 2016, 94, 155–171. [Google Scholar] [CrossRef]
- Chang, C.H.; Pearce, E.L. Emerging concepts of T cell metabolism as a target of immunotherapy. Nat. Immunol. 2016, 17, 364–368. [Google Scholar] [CrossRef]
- Chang, C.H.; Qiu, J.; O’Sullivan, D.; Buck, M.D.; Noguchi, T.; Curtis, J.D.; Chen, Q.; Gindin, M.; Gubin, M.M.; van der Windt, G.J.; et al. Metabolic Competition in the Tumor Microenvironment Is a Driver of Cancer Progression. Cell 2015, 162, 1229–1241. [Google Scholar] [CrossRef]
- Buck, M.D.; O’Sullivan, D.; Pearce, E.L. T cell metabolism drives immunity. J. Exp. Med. 2015, 212, 1345–1360. [Google Scholar] [CrossRef]
- Fischer, K.; Hoffmann, P.; Voelkl, S.; Meidenbauer, N.; Ammer, J.; Edinger, M.; Gottfried, E.; Schwarz, S.; Rothe, G.; Hoves, S.; et al. Inhibitory effect of tumor cell-derived lactic acid on human T cells. Blood 2007, 109, 3812–3819. [Google Scholar] [CrossRef]
- Singer, K.; Gottfried, E.; Kreutz, M.; Mackensen, A. Suppression of T-cell responses by tumor metabolites. Cancer Immunol. Immunother. 2011, 60, 425–431. [Google Scholar] [CrossRef] [PubMed]
- Gottfried, E.; Kunz-Schughart, L.A.; Ebner, S.; Mueller-Klieser, W.; Hoves, S.; Andreesen, R.; Mackensen, A.; Kreutz, M. Tumor-derived lactic acid modulates dendritic cell activation and antigen expression. Blood 2006, 107, 2013–2021. [Google Scholar] [CrossRef]
- Angelin, A.; Gil-de-Gómez, L.; Dahiya, S.; Jiao, J.; Guo, L.; Levine, M.H.; Wang, Z.; Quinn, W.J., 3rd; Kopinski, P.K.; Wang, L.; et al. Foxp3 Reprograms T Cell Metabolism to Function in Low-Glucose, High-Lactate Environments. Cell Metab. 2017, 25, 1282–1293. [Google Scholar] [CrossRef]
- Huang, S.C.; Smith, A.M.; Everts, B.; Colonna, M.; Pearce, E.L.; Schilling, J.D.; Pearce, E.J. Metabolic Reprogramming Mediated by the mTORC2-IRF4 Signaling Axis Is Essential for Macrophage Alternative Activation. Immunity 2016, 45, 817–830. [Google Scholar] [CrossRef]
- Katanov, C.; Lerrer, S.; Liubomirski, Y.; Leider-Trejo, L.; Meshel, T.; Bar, J.; Feniger-Barish, R.; Kamer, I.; Soria-Artzi, G.; Kahani, H.; et al. Regulation of the inflammatory profile of stromal cells in human breast cancer: Prominent roles for TNF-α and the NF-κB pathway. Stem Cell Res. Ther. 2015, 6, 87. [Google Scholar] [CrossRef]
- Végran, F.; Boidot, R.; Michiels, C.; Sonveaux, P.; Feron, O. Lactate influx through the endothelial cell monocarboxylate transporter MCT1 supports an NF-κB/IL-8 pathway that drives tumor angiogenesis. Cancer Res. 2011, 71, 2550–2560. [Google Scholar] [CrossRef]
- O’Sullivan, D.; Sanin, D.E.; Pearce, E.J.; Pearce, E.L. Metabolic interventions in the immune response to cancer. Nat. Rev. Immunol. 2019, 19, 324–335. [Google Scholar] [CrossRef] [PubMed]
- Koukourakis, M.I.; Giatromanolaki, A.; Sivridis, E.; Gatter, K.C.; Trarbach, T.; Folprecht, G.; Shi, M.M.; Lebwohl, D.; Jalava, T.; Laurent, D.; et al. Prognostic and predictive role of lactate dehydrogenase 5 expression in colorectal cancer patients treated with PTK787/ZK 222584 (vatalanib) antiangiogenic therapy. Clin. Cancer Res. 2011, 17, 4892–4900. [Google Scholar] [CrossRef] [PubMed]
- Tas, F.; Aydiner, A.; Demir, C.; Topuz, E. Serum lactate dehydrogenase levels at presentation predict outcome of patients with limited-stage small-cell lung cancer. Am. J. Clin. Oncol. 2001, 24, 376–378. [Google Scholar] [CrossRef] [PubMed]
- Turen, S.; Ozyar, E.; Altundag, K.; Gullu, I.; Atahan, I.L. Serum lactate dehydrogenase level is a prognostic factor in patients with locoregionally advanced nasopharyngeal carcinoma treated with chemoradiotherapy. Cancer Investig. 2007, 25, 315–321. [Google Scholar] [CrossRef]
- Halabi, S.; Small, E.J.; Kantoff, P.W.; Kattan, M.W.; Kaplan, E.B.; Dawson, N.A.; Levine, E.G.; Blumenstein, B.A.; Vogelzang, N.J. Prognostic model for predicting survival in men with hormone-refractory metastatic prostate cancer. J. Clin. Oncol. 2003, 21, 1232–1237. [Google Scholar] [CrossRef]
- Brown, J.E.; Cook, R.J.; Lipton, A.; Coleman, R.E. Serum lactate dehydrogenase is prognostic for survival in patients with bone metastases from breast cancer: A retrospective analysis in bisphosphonate-treated patients. Clin. Cancer Res. 2012, 18, 6348–6355. [Google Scholar] [CrossRef]
- Petrelli, F.; Cabiddu, M.; Coinu, A.; Borgonovo, K.; Ghilardi, M.; Lonati, V.; Barni, S. Prognostic role of lactate dehydrogenase in solid tumors: A systematic review and meta-analysis of 76 studies. Acta Oncol. 2015, 54, 961–970. [Google Scholar] [CrossRef]
- Li, G.; Wang, Z.; Xu, J.; Wu, H.; Cai, S.; He, Y. The prognostic value of lactate dehydrogenase levels in colorectal cancer: A meta-analysis. BMC Cancer 2016, 16, 249. [Google Scholar] [CrossRef]
- Zhang, Y.; Xu, T.; Wang, Y.; Zhang, H.; Zhao, Y.; Yang, X.; Cao, Y.; Yang, W.; Niu, H. Prognostic Role of Lactate Dehydrogenase Expression in Urologic Cancers: A Systematic Review and Meta-Analysis. Oncol. Res. Treat. 2016, 39, 592–604. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Wei, S.; Su, L.; Lv, W.; Hong, J. Prognostic significance of pretreated serum lactate dehydrogenase level in nasopharyngeal carcinoma among Chinese population: A meta-analysis. Medicine 2016, 95, e4494. [Google Scholar] [CrossRef]
- Yuan, Z.Y.; Gao, S.G.; Mu, J.W.; Xue, Q.; Mao, Y.S.; Wang, D.L.; Zhao, J.; Gao, Y.S.; Huang, J.F.; He, J. Prognostic value of preoperative serum lactate dehydrogenase in thymic carcinoma. J. Thorac. Dis. 2016, 8, 2464–2472. [Google Scholar] [CrossRef]
- Marais, L.C.; Bertie, J.; Rodseth, R.; Sartorius, B.; Ferreira, N. Pre-treatment serum lactate dehydrogenase and alkaline phosphatase as predictors of metastases in extremity osteosarcoma. J. Bone Oncol. 2015, 4, 80–84. [Google Scholar] [CrossRef]
- Zhang, J.P.; Wang, H.B.; Lin, Y.H.; Xu, J.; Wang, J.; Wang, K.; Liu, W.L. Lactate Dehydrogenase Is an Important Prognostic Indicator for Hepatocellular Carcinoma after Partial Hepatectomy. Transl. Oncol. 2015, 8, 497–503. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Yang, Q.; Wang, H.; Wang, Z.; Zuo, D.; Cai, Z.; Hua, Y. Prognostic significance of serum lactate dehydrogenase levels in Ewing’s sarcoma: A meta-analysis. Mol. Clin. Oncol. 2016, 5, 832–838. [Google Scholar] [CrossRef]
- Yu, S.L.; Xu, L.T.; Qi, Q.; Geng, Y.W.; Chen, H.; Meng, Z.Q.; Wang, P.; Chen, Z. Serum lactate dehydrogenase predicts prognosis and correlates with systemic inflammatory response in patients with advanced pancreatic cancer after gemcitabine-based chemotherapy. Sci. Rep. 2017, 7, 45194. [Google Scholar] [CrossRef]
- Jia, Z.; Zhang, J.; Wang, Z.; Wang, B.; Wang, L.; Cao, J.; Tao, Z.; Hu, X. An explorative analysis of the prognostic value of lactate dehydrogenase for survival and the chemotherapeutic response in patients with advanced triple-negative breast cancer. Oncotarget 2018, 9, 10714–10722. [Google Scholar] [CrossRef] [PubMed]
- Rani, R.; Kumar, V. Recent Update on Human Lactate Dehydrogenase Enzyme 5 (hLDH5) Inhibitors: A Promising Approach for Cancer Chemotherapy. J. Med. Chem. 2016, 59, 487–496. [Google Scholar] [CrossRef]
- Boudreau, A.; Purkey, H.E.; Hitz, A.; Robarge, K.; Peterson, D.; Labadie, S.; Kwong, M.; Hong, R.; Gao, M.; Del Nagro, C.; et al. Metabolic plasticity underpins innate and acquired resistance to LDHA inhibition. Nat. Chem. Biol. 2016, 12, 779–786. [Google Scholar] [CrossRef]
- Ward, R.A.; Brassington, C.; Breeze, A.L.; Caputo, A.; Critchlow, S.; Davies, G.; Goodwin, L.; Hassall, G.; Greenwood, R.; Holdgate, G.A.; et al. Design and synthesis of novel lactate dehydrogenase A inhibitors by fragment-based lead generation. J. Med. Chem. 2012, 55, 3285–3306. [Google Scholar] [CrossRef]
- Shi, Y.; Pinto, B.M. Human lactate dehydrogenase a inhibitor: A molecular dynamics investigation. PLoS ONE 2014, 9, e86365. [Google Scholar] [CrossRef]
- Sun, R.; Li, X.; Li, Y.; Zhang, X.; Li, X.; Li, X.; Shi, Z.; Bao, J. Screening of novel inhibitors targeting lactate dehydrogenase A via four molecular docking strategies and dynamics simulations. J. Mol. Model. 2015, 21, 133. [Google Scholar] [CrossRef]
- Tuccinardi, T.; Poli, G.; Corchia, I.; Granchi, C.; Lapillo, M.; Macchia, M.; Minutolo, F.; Ortore, G.; Martinelli, A. A Virtual Screening Study for Lactate Dehydrogenase 5 Inhibitors by Using a Pharmacophore-based Approach. Mol. Inform. 2016, 35, 434–439. [Google Scholar] [CrossRef]
- Buonfiglio, R.; Ferraro, M.; Falchi, F.; Cavalli, A.; Masetti, M.; Recanatini, M. Collecting and assessing human lactate dehydrogenase-A conformations for structure-based virtual screening. J. Chem. Inf. Model. 2013, 53, 2792–2797. [Google Scholar] [CrossRef]
- Le, A.; Cooper, C.R.; Gouw, A.M.; Dinavahi, R.; Maitra, A.; Deck, L.M.; Royer, R.E.; Vander Jagt, D.L.; Semenza, G.L.; Dang, C.V. Inhibition of lactate dehydrogenase A induces oxidative stress and inhibits tumor progression. Proc. Natl. Acad. Sci. USA 2010, 107, 2037–2042. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Chen, X.; Yu, Y.; Li, J.; Hu, Q.; Xue, C.; Chen, J.; Shen, S.; Luo, Y.; Ren, F.; et al. LDHA is a direct target of miR-30d-5p and contributes to aggressive progression of gallbladder carcinoma. Mol. Carcinog. 2018, 57, 772–783. [Google Scholar] [CrossRef]
- Xian, Z.Y.; Liu, J.M.; Chen, Q.K.; Chen, H.Z.; Ye, C.J.; Xue, J.; Yang, H.Q.; Li, J.L.; Liu, X.F.; Kuang, S.J. Inhibition of LDHA suppresses tumor progression in prostate cancer. Tumour Biol. 2015, 36, 8093–8100. [Google Scholar] [CrossRef]
- Yu, Y.; Deck, J.A.; Hunsaker, L.A.; Deck, L.M.; Royer, R.E.; Goldberg, E.; Vander Jagt, D.L. Selective active site inhibitors of human lactate dehydrogenases A4, B4, and C4. Biochem. Pharmacol. 2001, 62, 81–89. [Google Scholar] [CrossRef]
- Vander Jagt, D.L.; Deck, L.M.; Royer, R.E. Gossypol: Prototype of inhibitors targeted to dinucleotide folds. Curr. Med. Chem. 2000, 7, 479–498. [Google Scholar] [CrossRef]
- Coyle, T.; Levante, S.; Shetler, M.; Winfield, J. In vitro and in vivo cytotoxicity of gossypol against central nervous system tumor cell lines. J. Neurooncol. 1994, 19, 25–35. [Google Scholar] [CrossRef]
- Shelley, M.D.; Hartley, L.; Fish, R.G.; Groundwater, P.; Morgan, J.J.; Mort, D.; Mason, M.; Evans, A. Stereo-specific cytotoxic effects of gossypol enantiomers and gossypolone in tumour cell lines. Cancer Lett. 1999, 135, 171–180. [Google Scholar] [CrossRef]
- Van Poznak, C.; Seidman, A.D.; Reidenberg, M.M.; Moasser, M.M.; Sklarin, N.; Van Zee, K.; Borgen, P.; Gollub, M.; Bacotti, D.; Yao, T.J.; et al. Oral gossypol in the treatment of patients with refractory metastatic breast cancer: A phase I/II clinical trial. Breast Cancer Res. Treat. 2001, 66, 239–248. [Google Scholar] [CrossRef]
- Flack, M.R.; Pyle, R.G.; Mullen, N.M.; Lorenzo, B.; Wu, Y.W.; Knazek, R.A.; Nisula, B.C.; Reidenberg, M.M. Oral gossypol in the treatment of metastatic adrenal cancer. J. Clin. Endocrinol. Metab. 1993, 76, 1019–1024. [Google Scholar]
- Bushunow, P.; Reidenberg, M.M.; Wasenko, J.; Winfield, J.; Lorenzo, B.; Lemke, S.; Himpler, B.; Corona, R.; Coyle, T. Gossypol treatment of recurrent adult malignant gliomas. J. Neurooncol. 1999, 43, 79–86. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Xiong, Y.; Qiao, T.; Li, X.; Jia, L.; Han, Y. Lactate dehydrogenase A: A key player in carcinogenesis and potential target in cancer therapy. Cancer Med. 2018, 7, 6124–6136. [Google Scholar] [CrossRef]
- Manerba, M.; Vettraino, M.; Fiume, L.; Di Stefano, G.; Sartini, A.; Giacomini, E.; Buonfiglio, R.; Roberti, M.; Recanatini, M. Galloflavin (CAS 568-80-9): A novel inhibitor of lactate dehydrogenase. Chem. Med. Chem. 2012, 7, 311–317. [Google Scholar] [CrossRef]
- Farabegoli, F.; Vettraino, M.; Manerba, M.; Fiume, L.; Roberti, M.; Di Stefano, G. Galloflavin, a new lactate dehydrogenase inhibitor, induces the death of human breast cancer cells with different glycolytic attitude by affecting distinct signaling pathways. Eur. J. Pharm. Sci. 2012, 47, 729–738. [Google Scholar] [CrossRef]
- Granchi, C.; Paterni, I.; Rani, R.; Minutolo, F. Small-molecule inhibitors of human LDH5. Future Med. Chem. 2013, 5, 1967–1991. [Google Scholar] [CrossRef] [PubMed]
- Granchi, C.; Roy, S.; Giacomelli, C.; Macchia, M.; Tuccinardi, T.; Martinelli, A.; Lanza, M.; Betti, L.; Giannaccini, G.; Lucacchini, A.; et al. Discovery of N-hydroxyindole-based inhibitors of human lactate dehydrogenase isoform A (LDH-A) as starvation agents against cancer cells. J. Med. Chem. 2011, 54, 1599–1612. [Google Scholar] [CrossRef] [PubMed]
- Granchi, C.; Roy, S.; De Simone, A.; Salvetti, I.; Tuccinardi, T.; Martinelli, A.; Macchia, M.; Lanza, M.; Betti, L.; Giannaccini, G.; et al. N-Hydroxyindole-based inhibitors of lactate dehydrogenase against cancer cell proliferation. Eur. J. Med. Chem. 2011, 46, 5398–5407. [Google Scholar] [CrossRef]
- Granchi, C.; Roy, S.; Mottinelli, M.; Nardini, E.; Campinoti, F.; Tuccinardi, T.; Lanza, M.; Betti, L.; Giannaccini, G.; Lucacchini, A.; et al. Synthesis of sulfonamide-containing N-hydroxyindole-2-carboxylates as inhibitors of human lactate dehydrogenase-isoform 5. Bioorg. Med. Chem. Lett. 2011, 21, 7331–7336. [Google Scholar] [CrossRef]
- Wittrup, A.; Lieberman, J. Knocking down disease: A progress report on siRNA therapeutics. Nat. Rev. Genet. 2015, 16, 543–552. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Xu, L.; Wu, Q.; Liu, M.; Tang, F.; Cai, Y.; Fan, W.; Huang, H.; Gu, X. Inhibition of LDHA Deliver Potential Anticancer Performance in Renal Cell Carcinoma. Urol. Int. 2017, 99, 237–244. [Google Scholar] [CrossRef]
- Li, J.; Zhu, S.; Tong, J.; Hao, H.; Yang, J.; Liu, Z.; Wang, Y. Suppression of lactate dehydrogenase A compromises tumor progression by downregulation of the Warburg effect in glioblastoma. Neuroreport 2016, 27, 110–115. [Google Scholar] [CrossRef]
- Wang, J.; Wang, H.; Liu, A.; Fang, C.; Hao, J.; Wang, Z. Lactate dehydrogenase A negatively regulated by miRNAs promotes aerobic glycolysis and is increased in colorectal cancer. Oncotarget 2015, 6, 19456–19468. [Google Scholar] [CrossRef] [PubMed]
- Sheng, S.L.; Liu, J.J.; Dai, Y.H.; Sun, X.G.; Xiong, X.P.; Huang, G. Knockdown of lactate dehydrogenase A suppresses tumor growth and metastasis of human hepatocellular carcinoma. FEBS J. 2012, 279, 3898–3910. [Google Scholar] [CrossRef] [PubMed]
- Dorneburg, C.; Fischer, M.; Barth, T.F.E.; Mueller-Klieser, W.; Hero, B.; Gecht, J.; Carter, D.R.; De Preter, K.; Mayer, B.; Christner, L.; et al. LDHA in neuroblastoma is associated with poor outcome and its depletion decreases neuroblastoma growth independent of aerobic glycolysis. Clin. Cancer Res. 2018, 24, 5772–5783. [Google Scholar] [CrossRef]
- Cui, X.G.; Han, Z.T.; He, S.H.; Wu, X.D.; Chen, T.R.; Shao, C.H.; Chen, D.L.; Su, N.; Chen, Y.M.; Wang, T.; et al. HIF1/2α mediates hypoxia-induced LDHA expression in human pancreatic cancer cells. Oncotarget 2017, 8, 24840–24852. [Google Scholar]
- Di, H.; Zhang, X.; Guo, Y.; Shi, Y.; Fang, C.; Yuan, Y.; Wang, J.; Shang, C.; Guo, W.; Li, C. Silencing LDHA inhibits proliferation, induces apoptosis and increases chemosensitivity to temozolomide in glioma cells. Oncol. Lett. 2018, 15, 5131–5136. [Google Scholar] [CrossRef]
- Fantin, V.R.; St-Pierre, J.; Leder, P. Attenuation of LDH-A expression uncovers a link between glycolysis, mitochondrial physiology, and tumor maintenance. Cancer Cell 2006, 9, 425–434. [Google Scholar] [CrossRef]
- Hao, J.; Graham, P.; Chang, L.; Ni, J.; Wasinger, V.; Beretov, J.; Deng, J.; Duan, W.; Bucci, J.; Malouf, D.; et al. Proteomic identification of the lactate dehydrogenase A in a radioresistant prostate cancer xenograft mouse model for improving radiotherapy. Oncotarget 2016, 7, 74269–74285. [Google Scholar] [CrossRef]
- Hua, G.; Liu, Y.; Li, X.; Xu, P.; Luo, Y. Targeting glucose metabolism in chondrosarcoma cells enhances the sensitivity to doxorubicin through the inhibition of lactate dehydrogenase-A. Oncol. Rep. 2014, 31, 2727–2734. [Google Scholar] [CrossRef] [PubMed]
- Zhou, M.; Zhao, Y.; Ding, Y.; Liu, H.; Liu, Z.; Fodstad, O.; Riker, A.I.; Kamarajugadda, S.; Lu, J.; Owen, L.B.; et al. Warburg effect in chemosensitivity: Targeting lactate dehydrogenase-A re-sensitizes taxol-resistant cancer cells to taxol. Mol. Cancer 2010, 9, 33. [Google Scholar] [CrossRef]
- Giatromanolaki, A.; Koukourakis, M.I.; Koutsopoulos, A.; Mendrinos, S.; Sivridis, E. The metabolic interactions between tumor cells and tumor-associated stroma (TAS) in prostatic cancer. Cancer Biol. Ther. 2012, 13, 1284–1289. [Google Scholar] [CrossRef] [PubMed]
- Hirschhaeuser, F.; Sattler, U.G.; Mueller-Klieser, W. Lactate: A metabolic key player in cancer. Cancer Res. 2011, 71, 6921–6925. [Google Scholar] [CrossRef] [PubMed]
- Nenu, I.; Gafencu, G.A.; Popescu, T.; Kacso, G. Lactate—A new frontier in the immunology and therapy of prostate cancer. J. Cancer Res. Ther. 2017, 13, 406–411. [Google Scholar] [PubMed]
- Ippolito, L.; Morandi, A.; Giannoni, E.; Chiarugi, P. Lactate: A Metabolic Driver in the Tumour Landscape. Trends Biochem. Sci. 2019, 44, 153–166. [Google Scholar] [CrossRef] [PubMed]
| Cancer Type | Sample/Model Systems | Mechanisms | Expression | Reference |
|---|---|---|---|---|
| Pancreatic cancer | pancreatic tumors and cancer cell lines | increased pancreatic cancer cell growth and metastasis by FOXM1-LDHA signaling | LDHA overexpression | [8] |
| Pancreatic cancer | pancreatic tumors, in vitro and in orthotopic mouse models | increased pancreatic cancer cell growth and metastasis by KLF4-LDHA signaling | LDHA overexpression | [9] |
| Pancreatic cancer | pancreatic ductal carcinoma samples | increased pancreatic cancer cell growth by citrate synthase mediated glucose to lipid conversion | LDHA overexpression | [10] |
| Nasopharyngeal cancer | NPC tumor tissues and cell lines | increased cell proliferation, migration and invasion by JMJD2A-LDHA signaling | LDHA overexpression | [12] |
| Gastric cancer | GC cell lines | promoted cancer growth by glycolytic phenotype (increased lactate production and glucose utilization) by FOXM1-LDHA signaling | LDHA overexpression | [13] |
| Bladder cancer | invasive transitional cell bladder cancer tissues | poor local relapse free survival, LDHA Silencing lead to strong radio-sensitization. | LDH-5 over expression | [14] |
| Endometrial cancer | stage I endometrial adenocarcinoma cells | LDH-5 mediated expression of phosphorylated VEGFR2/KDR receptors pathway | LDH-5 over expression | [15] |
| Bladder cancer | BC cell lines, muscle-invasive bladder cancer samples | epithelial-to-mesenchymal transition | LDHA expression | [31] |
| Cancer Type | Sample/Model System | Mechanism | Expression | Reference |
|---|---|---|---|---|
| Prostate cancer | poorly metastatic and highly metastatic variant of human prostate cancer cell lines and primary cancer tissues | promoter hypermethylation | loss of LDHB expression | [51] |
| Gastric cancer cell lines and pancreatic cancer | four gastric cancer cell lines and one pancreatic cancer cell line | promoter hypermethylation | loss of LDHB expression, loss of LDH1-4 | [52] |
| Pancreatic cancer | pancreatic cancer tissues and cell lines | promoter hypermethylation | suppressed LDHB and even total loss | [53] |
| Breast cancer | breast cancer (adenocarcinoma) tissues and cell lines | promoter hypermethylation | Absent or decreased expression of LDH isoenzymes 1–4 | [54] |
| Breast cancer | basal-like/triple-negative breast cancers | glycolytic pathway | high LDHB expression | [55] |
| Hepatocellular carcinoma | Highly metastatic cell line | Metastasis potential | down-regulated LDHB expression | [56] |
| Hepatocellular carcinoma | HCC cancer tissues | unfavorable survival outcomes | Low LDHB expression | [57] |
| Urinary bladder urothelial carcinoma (UBUC) | UBUC cancer tissues | tumor progression and inferior disease-specific survival | Loss of LDHB expression | [58] |
| Colorectal cancer | colorectal cancer tissues, in vitro and in vivo model | increased autophagy, accelerated cancer growth mediated by SIRT5/LDHB pathway | Deacetylation of LDHB | [61] |
| Colorectal cancer | colorectal cancer tissues, CRC cell lines, | RPS7 mediated cell glycolysis inhibition, inhibits colorectal cancer growth | Suppressed LDHB expression | [62] |
| Hepatoma cancer | hepatoma cell lines | claudin-1-mediated high invasive activity | Decreased LDHB expression | [63] |
| Colorectal cancer | stage I-III CRC patient samples, in vitro | KLF14 regulates glycolysis | downregulating LDHB expression | [64] |
| Lung cancer | KRAS-dependent lung adenocarcinoma samples, vitro and in vivo | predictor of shorter survival in patients, KRAS-mutant lung tumors are more dependent on glycolysis for proliferation compared with KRAS wild-type lung tumors | high LDHB expression | [65] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mishra, D.; Banerjee, D. Lactate Dehydrogenases as Metabolic Links between Tumor and Stroma in the Tumor Microenvironment. Cancers 2019, 11, 750. https://doi.org/10.3390/cancers11060750
Mishra D, Banerjee D. Lactate Dehydrogenases as Metabolic Links between Tumor and Stroma in the Tumor Microenvironment. Cancers. 2019; 11(6):750. https://doi.org/10.3390/cancers11060750
Chicago/Turabian StyleMishra, Deepshikha, and Debabrata Banerjee. 2019. "Lactate Dehydrogenases as Metabolic Links between Tumor and Stroma in the Tumor Microenvironment" Cancers 11, no. 6: 750. https://doi.org/10.3390/cancers11060750
APA StyleMishra, D., & Banerjee, D. (2019). Lactate Dehydrogenases as Metabolic Links between Tumor and Stroma in the Tumor Microenvironment. Cancers, 11(6), 750. https://doi.org/10.3390/cancers11060750







