Automated Workflow for Somatic and Germline Next Generation Sequencing Analysis in Routine Clinical Cancer Diagnostics
Abstract
1. Introduction
2. Results
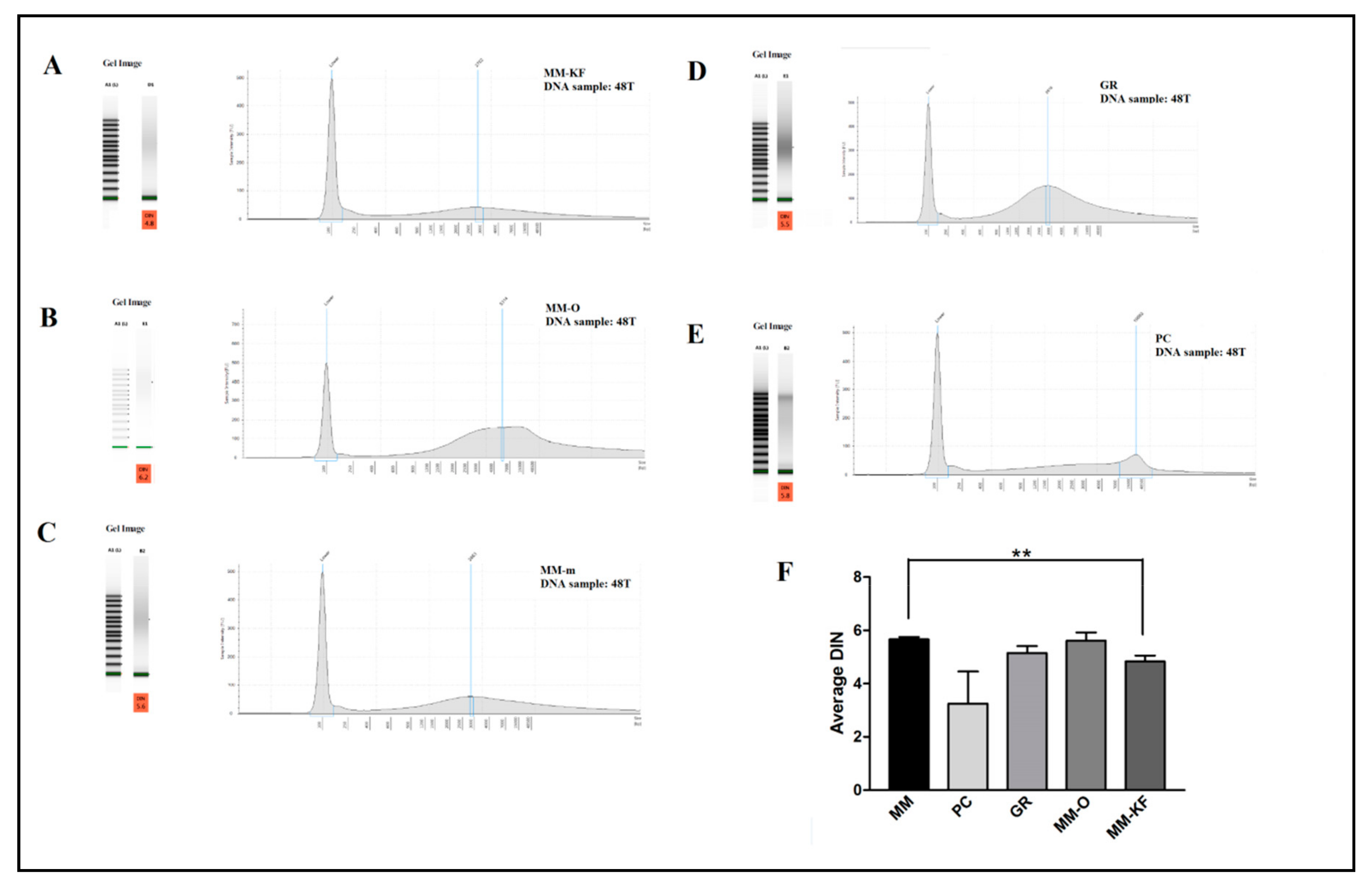
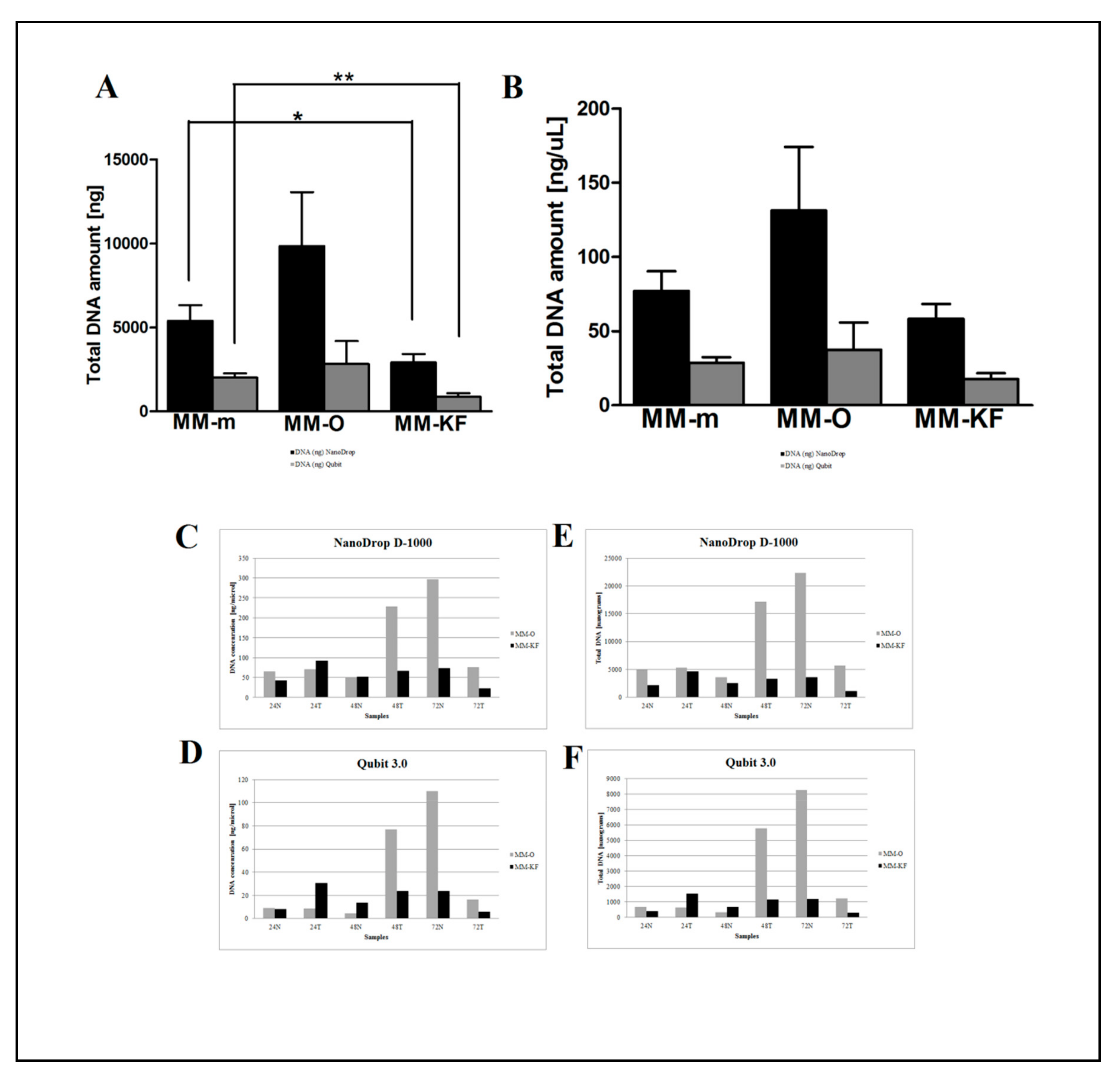
2.1. Manual vs. Automatic DNA from FFPE Tissue Extraction
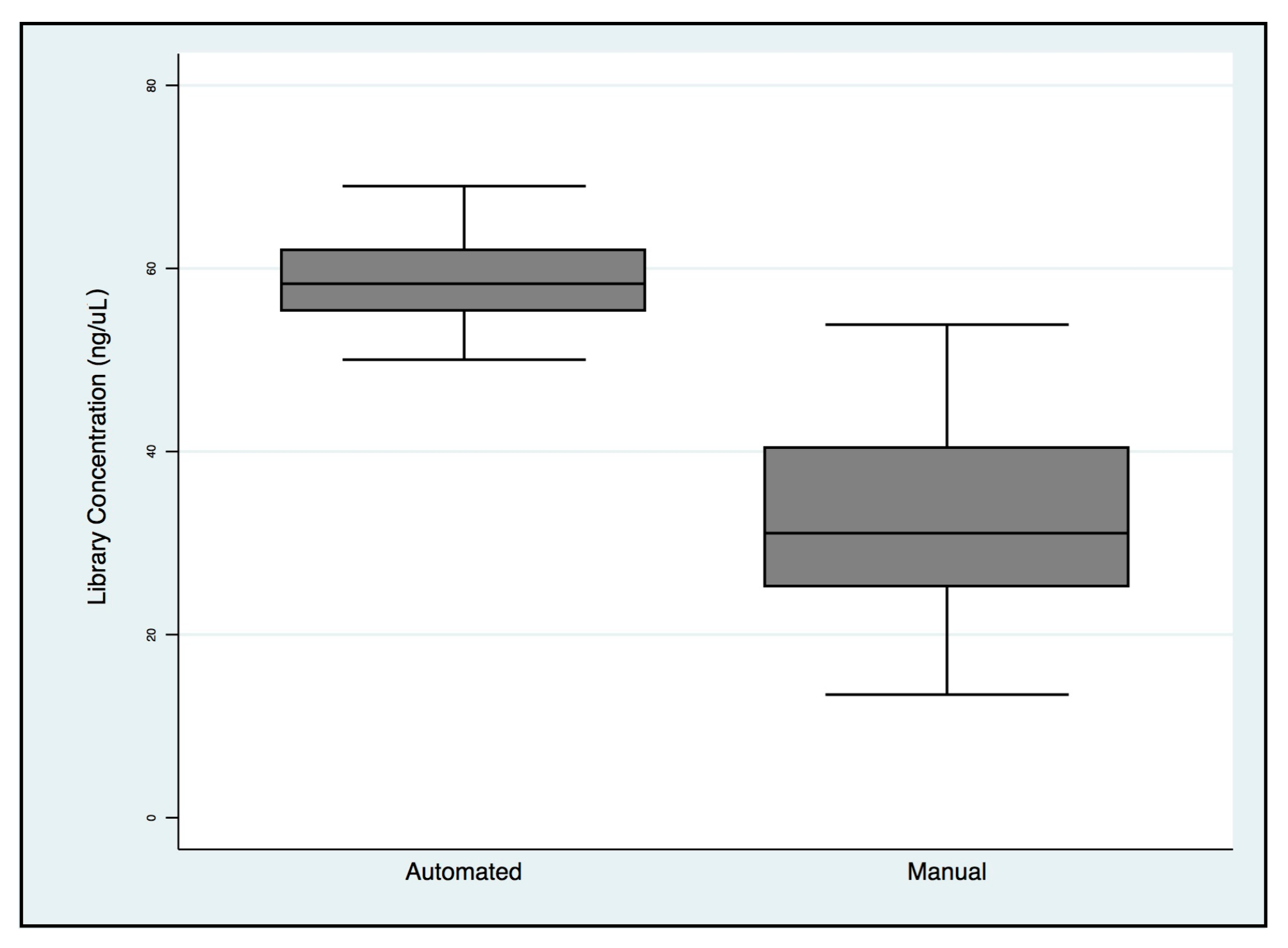
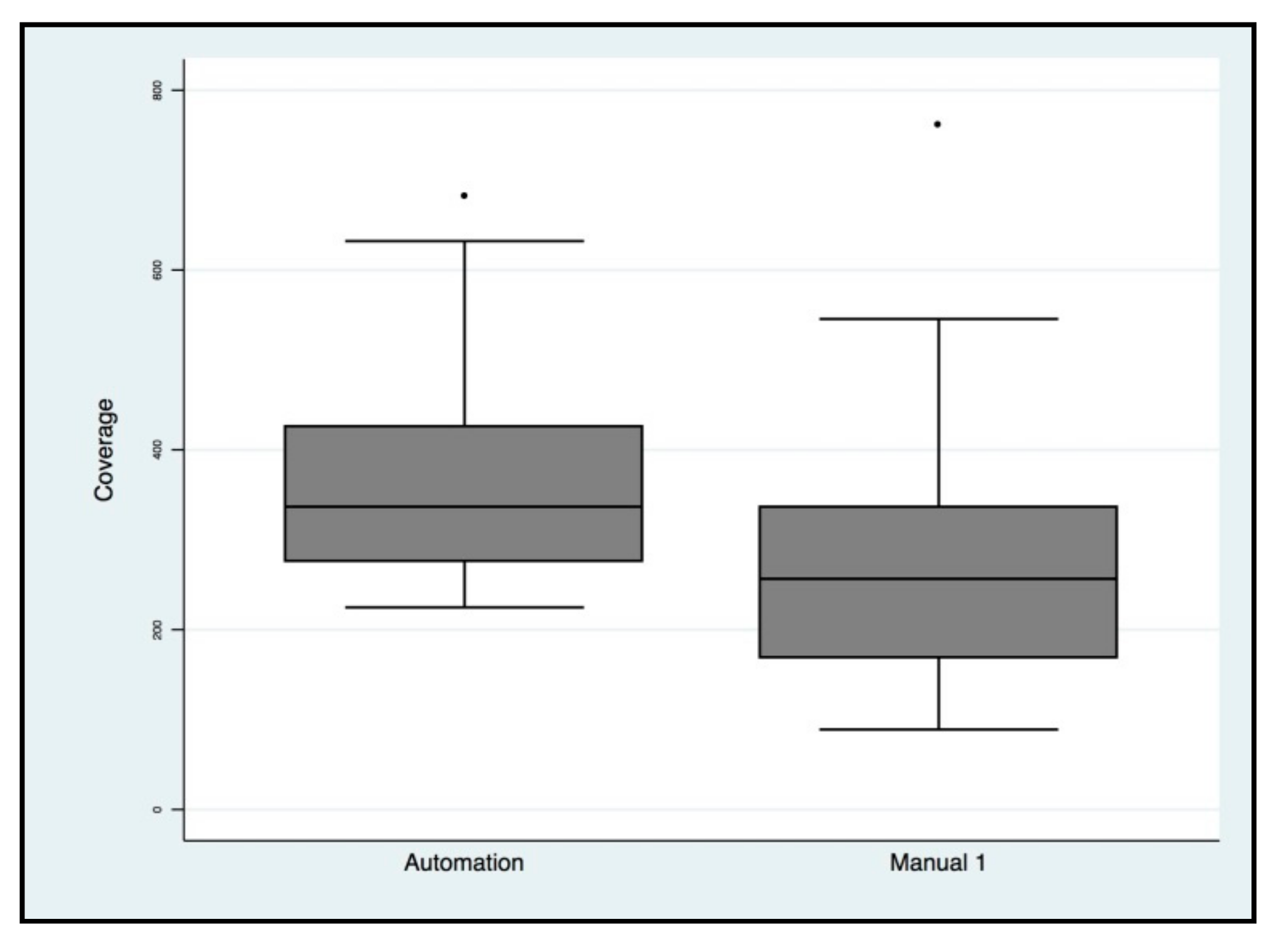
2.2. NGS Automatic Performance Workflow
3. Discussion
4. Materials and Methods
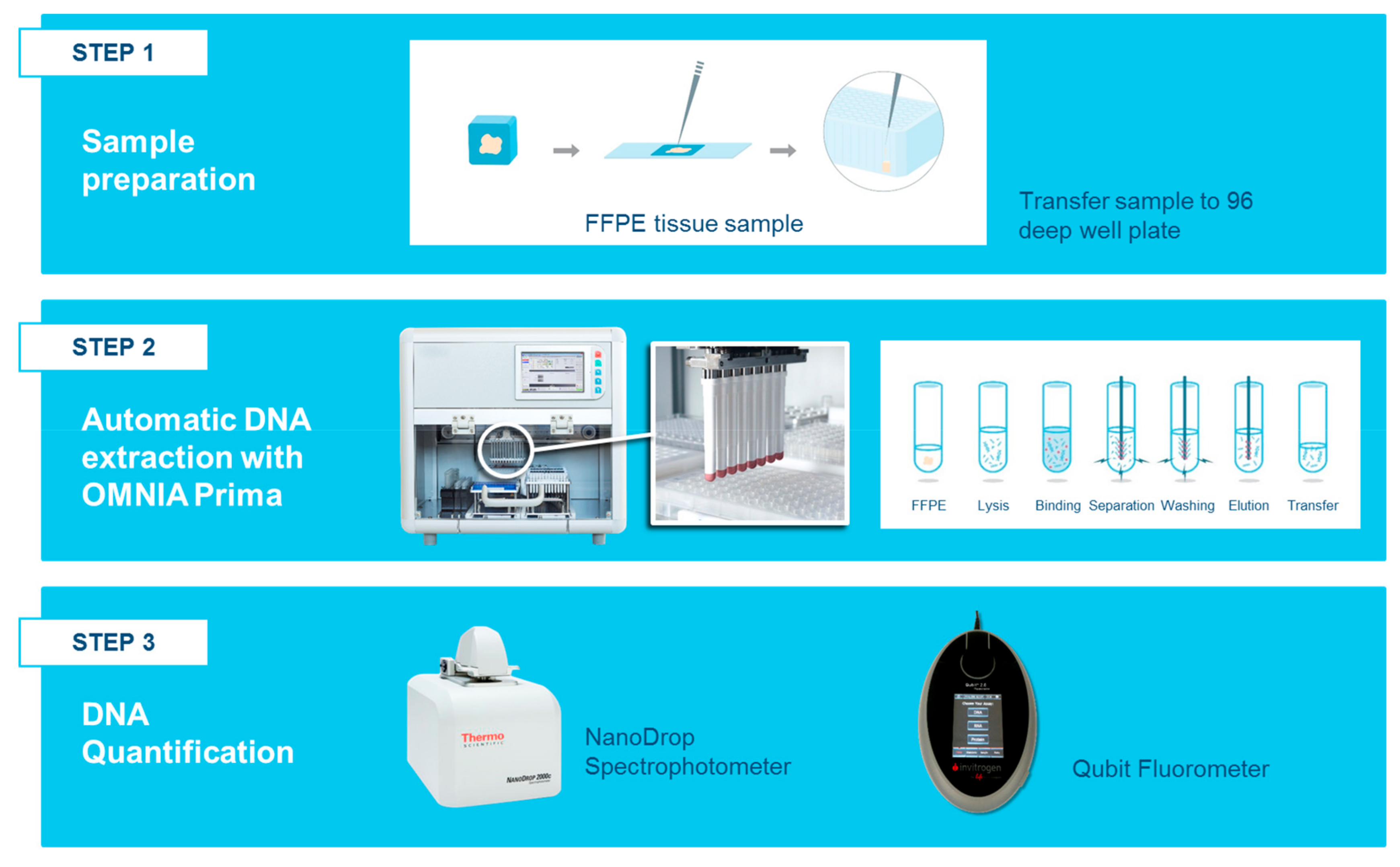
4.1. DNA Extraction Workflow
4.1.1. Biological Sample Collection and Macroscopic Evaluation
4.1.2. Manual DNA Isolation from FFPE Tissues
4.1.3. Automated DNA Isolation from FFPE Tissues
4.1.4. Quality and Quantity DNA Assessment
4.1.5. Assessment of PCR Amplifiable Fragment Length
4.1.6. Genomic DNA extraction
4.2. NGS Workflow
4.2.1. NGS Library Preparation, Quality and Quantity Assessment
4.2.2. NGS Analysis
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bonfiglio, S.; Vanni, I.; Rossella, V.; Truini, A.; Lazarevič, D.; Bello, M.G.D.; Alama, A.; Mora, M.; Rijavec, E.; Genova, C.; et al. Performance comparison of two commercial human whole-exome capture systems on formalin-fixed paraffin-embedded lung adenocarcinoma samples. BMC Cancer 2016, 16, 692. [Google Scholar] [CrossRef] [PubMed]
- Yohe, S.; Thyagarajan, B. Review of Clinical Next-Generation Sequencing. Arch. Pathol. Labor. Med. 2017, 141, 1544–1557. [Google Scholar] [CrossRef] [PubMed]
- Pikor, L.A.; Enfield, K.S.; Cameron, H.; Lam, W.L. DNA extraction from paraffin embedded material for genetic and epigenetic analyses. J. Vis. Exp. 2011, 49, e2763. [Google Scholar] [CrossRef] [PubMed]
- Ferrer, I.; Armstrong, J.; Capellari, S.; Parchi, P.; Arzberger, T.; Bell, J.; Budka, H.; Strobel, T.; Giaccone, G.; Rossi, G.; et al. Effects of Formalin Fixation, Paraffin Embedding, and Time of Storage on DNA Preservation in Brain Tissue: A BrainNet Europe Study. Brain Pathol. 2007, 17, 297–303. [Google Scholar] [CrossRef]
- Patel, P.G.; Selvarajah, S.; Guérard, K.P.; Bartlett, J.M.S.; Lapointe, J.; Berman, D.M.; Okello, J.B.A.; Park, P.C. Reliability and performance of commercial RNA and DNA extraction kits for FFPE tissue cores. PLoS ONE 2017, 12, 0179732. [Google Scholar] [CrossRef]
- Guyard, A.; Boyez, A.; Pujals, A.; Robe, C.; Van Nhieu, J.T.; Allory, Y.; Moroch, J.; Georges, O.; Fournet, J.C.; Zafrani, E.S.; et al. DNA degrades during storage in formalin-fixed and paraffin-embedded tissue blocks. Virchows Arch. 2017, 471, 491–500. [Google Scholar] [CrossRef]
- Greer, C.E.; Wheeler, C.M.; Manos, M.M. Sample preparation and PCR amplification from paraffin-embedded tissues. Genome Res. 1994, 3, S113–S122. [Google Scholar] [CrossRef]
- Bonnet, E.; Moutet, M.L.; Baulard, C.; Bacq-Daian, D.; Sandron, F.; Mesrob, L.; Fin, B.; Delepine, M.; Palomares, M.A.; Jubin, C.; et al. Performance comparison of three DNA extraction kits on human whole-exome data from formalin-fixed paraffin-embedded normal and tumor samples. PLoS ONE 2018, 13, e0195471. [Google Scholar] [CrossRef]
- Janecka, A.; Adamczyk, A.; Gasińska, A. Comparison of eight commercially available kits for DNA extraction from formalin-fixed paraffin-embedded tissues. Anal. Biochem. 2015, 476, 8–10. [Google Scholar] [CrossRef]
- Callejas, S.; Álvarez, R.; Benguria, A.; Dopazo, A. AG-NGS: A powerful and user-friendly computing application for the semi-automated preparation of next-generation sequencing libraries using open liquid handling platforms. Biotechniques 2014, 56, 28–35. [Google Scholar] [CrossRef]
- Klevebring, D.; Gry, M.; Lindberg, J.; Eidefors, A.; Lundeberg, J. Automation of cDNA Synthesis and Labelling Improves Reproducibility. J. Biomed. Biotechnol. 2009, 2009, 396808. [Google Scholar] [CrossRef] [PubMed]
- Sedlackova, T.; Repiska, G.; Celec, P.; Szemes, T.; Minárik, G. Fragmentation of DNA affects the accuracy of the DNA quantitation by the commonly used methods. Boil. Proced. Online 2013, 15, 5. [Google Scholar] [CrossRef] [PubMed]
- Ludyga, N.; Grünwald, B.; Azimzadeh, O.; Englert, S.; Höfler, H.; Tapio, S.; Aubele, M. Nucleic acids from long-term preserved FFPE tissues are suitable for downstream analyses. Virchows Arch. 2012, 460, 131–140. [Google Scholar] [CrossRef] [PubMed]
- Senguven, B.; Baris, E.; Oygür, T.; Berktas, M. Comparison of Methods for the Extraction of DNA from Formalin-Fixed, Paraffin-Embedded Archival Tissues. Int. J. Med. Sci. 2014, 11, 494–499. [Google Scholar] [CrossRef] [PubMed]
- Ghatak, S.; Muthukumaran, R.B.; Nachimuthu, S.K. A simple method of genomic DNA extraction from human samples for PCR-RFLP analysis. J. Biomol. Tech. JBT 2013, 24, 224–231. [Google Scholar] [CrossRef]
- Gilbert, M.T.P.; Haselkorn, T.; Bunce, M.; Sanchez, J.J.; Lucas, S.B.; Jewell, L.D.; Van Marck, E.; Worobey, M. The Isolation of Nucleic Acids from Fixed, Paraffin-Embedded Tissues–Which Methods Are Useful When? PLoS ONE 2007, 2, e537. [Google Scholar] [CrossRef]
- Spencer, D.H.; Sehn, J.K.; Abel, H.J.; Watson, M.A.; Pfeifer, J.D.; Duncavage, E.J. Comparison of clinical targeted next-generation sequence data from formalin-fixed and fresh-frozen tissue specimens. J. Mol. Diagn. 2013, 15, 623–633. [Google Scholar] [CrossRef]
- Shibata, D.; Martin, W.J.; Arnheim, N. Analysis of DNA sequences in forty-year-old paraffin-embedded thin-tissue sections: A bridge between molecular biology and classical histology. Cancer Res. 1988, 48, 4564–4566. [Google Scholar]
- Srinivasan, M.; Sedmak, D.; Jewell, S. Effect of Fixatives and Tissue Processing on the Content and Integrity of Nucleic Acids. Am. J. Pathol. 2002, 161, 1961–1971. [Google Scholar] [CrossRef]
- Ben-Ezra, J.; Johnson, D.A.; Rossi, J.; Cook, N.; Wu, A. Effect of fixation on the amplification of nucleic acids from paraffin-embedded material by the polymerase chain reaction. J. Histochem. Cytochem. 1991, 39, 351–354. [Google Scholar] [CrossRef]
- Bonin, S.; Petrera, F.; Niccolini, B.; Stanta, G. PCR analysis in archival postmortem tissues. Mol. Pathol. 2003, 56, 184–186. [Google Scholar] [CrossRef] [PubMed]
- Simbolo, M.; Gottardi, M.; Corbo, V.; Fassan, M.; Mafficini, A.; Malpeli, G.; Lawlor, R.T.; Scarpa, A. DNA Qualification Workflow for Next Generation Sequencing of Histopathological Samples. PLoS ONE 2013, 8, e62692. [Google Scholar] [CrossRef] [PubMed]
- Russell, D.W. Isolation of High-molecular-weight DNA from Mammalian Cells Using Proteinase K and Phenol. Cold Spring Harb. Protoc. 2006, 2006, 1. [Google Scholar]
- Trombetta, D.; Graziano, P.; Scarpa, A.; Sparaneo, A.; Rossi, G.; Rossi, A.; Di Maio, M.; Antonello, D.; Mafficini, A.; Fabrizio, F.P.; et al. Frequent NRG1 fusions in Caucasian pulmonary mucinous adenocarcinoma predicted by Phospho-ErbB3 expression. Oncotarget 2018, 9, 9661–9671. [Google Scholar] [CrossRef]
- Concolino, P.; Costella, A.; Minucci, A.; Scaglione, G.L.; Santonocito, C.; Salutari, V.; Scambia, G.; Zuppi, C.; Capoluongo, E. A preliminary Quality Control (QC) for next generation sequencing (NGS) library evaluation turns out to be a very useful tool for a rapid detection of BRCA1/2 deleterious mutations. Clin. Chim. Acta 2014, 437, 72–77. [Google Scholar] [CrossRef]
- Concolino, P.; Rizza, R.; Hackmann, K.; Minucci, A.; Scaglione, G.L.; De Bonis, M.; Costella, A.; Zuppi, C.; Schrock, E.; Capoluongo, E. Identification and Characterization of a New BRCA2 Rearrangement in an Italian Family with Hereditary Breast and Ovarian Cancer Syndrome. Mol. Diagn. Ther. 2017, 21, 539–545. [Google Scholar] [CrossRef]
- Minucci, A.; Scambia, G.; Santonocito, C.; Concolino, P.; Canu, G.; Mignone, F.; De Bonis, M. Clinical impact on ovarian cancer patients of massive parallel sequencing for BRCA mutation detection: The experience at Gemelli hospital and a literature review. Expert Rev. Mol. Diagn. 2015, 15, 1383–1403. [Google Scholar] [CrossRef]
- Scaglione, G.L.; Concolino, P.; De Bonis, M.; De Paolis, E.; Minucci, A.; Ferrandina, G.; Scambia, G.; Capoluongo, E. A Whole Germline BRCA2 Gene Deletion: How to Learn from CNV In Silico Analysis. Int. J. Mol. Sci. 2018, 19, 961. [Google Scholar] [CrossRef]
- Capoluongo, E.; Scambia, G.; Nabholtz, J.M. Main implications related to the switch to BRCA1/2 tumor testing in ovarian cancer patients: A proposal of a consensus. Oncotarget 2018, 9, 19463–19468. [Google Scholar] [CrossRef]
- Capoluongo, E.; Ellison, G.; López-Guerrero, J.A.; Penault-Llorca, F.; Ligtenberg, M.J.; Banerjee, S.; Büttner, R. Guidance Statement on BRCA1/2 Tumor Testing in Ovarian Cancer Patients. Semin. Oncol. 2017, 44, 187–197. [Google Scholar] [CrossRef]





| Sample | Extraction Method | NanoDrop (DNA, ng/µL) | Total DNA (ng) NanoDrop | 260/280 Ratio | Qubit (DNA, ng/µL) | Total DNA (ng) Qubit |
|---|---|---|---|---|---|---|
| 24N | MM | 35.9 | 2513 | 1.83 | 12.4 | 868 |
| 24T | MM | 126.4 | 8848 | 1.74 | 26 | 1820 |
| 48N | MM | 64.8 | 4536 | 1.74 | 30.4 | 2128 |
| 48T | MM | 103 | 7210 | 1.78 | 36.6 | 2562 |
| 72N | MM | 58.9 | 4123 | 1.73 | 29 | 2030 |
| 72T | MM | 73 | 5110 | 1.76 | 37.8 | 2646 |
| 24N | PC | 171 | 3420 | 1.99 | 46.8 | 936 |
| 24T | PC | 623.6 | 12472 | 1.94 | 12.92 | 258.4 |
| 48N | PC | 156 | 3120 | 1.98 | 224 | 4480 |
| 48T | PC | 325.4 | 6508 | 1.98 | 154 | 3080 |
| 72N | PC | 546.6 | 10932 | 1.95 | 56 | 1120 |
| 72T | PC | 106.8 | 2136 | 2.07 | 4.6 | 92 |
| 24N | GR | 305.2 | 9156 | 1.95 | 74.4 | 2232 |
| 24T | GR | 647.4 | 19422 | 1.91 | 60.4 | 1812 |
| 48N | GR | 211.6 | 6348 | 1.93 | 244 | 7320 |
| 48T | GR | 393.6 | 11808 | 1.92 | 117.6 | 3528 |
| 72N | GR | 565.6 | 16968 | 1.9 | 112.8 | 3384 |
| 72T | GR | 194 | 5820 | 1.97 | 37.6 | 1128 |
| Sample | DNA Extraction Method | DNA (ng/µL) Nanodrop | DNA (ng) Nanodrop | 260/280 Ratio | DNA (ng/µL) Qubit | DNA (ng) Qubit |
|---|---|---|---|---|---|---|
| 24N | MM-O | 65.6 | 4920 | 1.6 | 8.94 | 670.5 |
| 24T | MM-O | 71.2 | 5340 | 1.62 | 8.48 | 636 |
| 48N | MM-O | 48.7 | 3648.8 | 1.63 | 4.28 | 321 |
| 48T | MM-O | 229.2 | 17190 | 1.84 | 77.2 | 5790 |
| 72N | MM-O | 297.4 | 22305 | 1.8 | 110 | 8250 |
| 72T | MM-O | 75.7 | 56738 | 1.71 | 16.6 | 1245 |
| 24N | MM-KF | 43.1 | 2155 | 1.7 | 8.34 | 417 |
| 24T | MM-KF | 92.6 | 4630 | 1.75 | 30.8 | 1540 |
| 48N | MM-KF | 51.6 | 2580 | 1.67 | 13.5 | 675 |
| 48T | MM-KF | 66.7 | 3335 | 1.71 | 23.6 | 1180 |
| 72N | MM-KF | 73.1 | 3655 | 1.75 | 24 | 1200 |
| 72T | MM-KF | 22.8 | 1140 | 1.51 | 5.76 | 288 |
| Sample | Extraction Method | DNA (ng/µL) NanoDrop | DNA (ng) NanoDrop | 260/280 | DNA (ng/µL) Qubit | DNA (ng) Qubit |
|---|---|---|---|---|---|---|
| 24N | MM-O | 65.6 | 4920 | 1.6 | 8.94 | 670.5 |
| 24T | MM-O | 71.2 | 5340 | 1.62 | 8.48 | 636 |
| 48N | MM-O | 48.7 | 3648.8 | 1.63 | 4.28 | 321 |
| 48T | MM-O | 229.2 | 17190 | 1.84 | 77.2 | 5790 |
| 72N | MM-O | 297.4 | 22305 | 1.8 | 110 | 8250 |
| 72T | MM-O | 75.7 | 5673,8 | 1.71 | 16.6 | 1245 |
| 24N | MM-KF | 43.1 | 2155 | 1.7 | 8.34 | 417 |
| 24T | MM-KF | 92.6 | 4630 | 1.75 | 30.8 | 1540 |
| 48N | MM-KF | 51.6 | 2580 | 1.67 | 13.5 | 675 |
| 48T | MM-KF | 66.7 | 3335 | 1.71 | 23.6 | 1180 |
| 72N | MM-KF | 73.1 | 3655 | 1.75 | 24 | 1200 |
| 72T | MM-KF | 22.8 | 1140 | 1.51 | 5.76 | 288 |
| Sample | DNA Extraction Method | DIN Value | QC Score |
|---|---|---|---|
| 24N | MM | 5.5 | −0.04 |
| 24T | MM | 5.9 | −0.04 |
| 48N | MM | 5.4 | −0.04 |
| 48T | MM | 5.6 | 0.15 |
| 72N | MM | 5.7 | −0.04 |
| 72T | MM | 5.9 | −0.04 |
| 24N | FC | 6.1 | −0.03 |
| 24T | FC | nv | −0.04 |
| 48N | FC | 1.9 | −0.02 |
| 48T | FC | 5.8 | 0.01 |
| 72N | FC | 5.7 | 0.01 |
| 72T | FC | nv | −0.01 |
| 24N | GR | 5.3 | 0.12 |
| 24T | GR | 5.7 | −0.03 |
| 48N | GR | 4.2 | −0.02 |
| 48T | GR | 5.5 | −0.01 |
| 72N | GR | 5.7 | −0.01 |
| 72T | GR | 4.5 | −0.01 |
| 24N | MM-O | 5.6 | −0.04 |
| 24T | MM-O | 5.1 | −0.04 |
| 48N | MM-O | 4.4 | −0.04 |
| 48T | MM-O | 6.2 | −0.03 |
| 72N | MM-O | 6.2 | −0.04 |
| 72T | MM-O | 6.2 | −0.02 |
| 24N | MM-KF | 4.7 | 0.01 |
| 24T | MM-KF | 5.8 | 0.00 |
| 48N | MM-KF | 4.8 | 0.03 |
| 48T | MM-KF | 4.8 | 0.03 |
| 72N | MM-KF | 4.8 | 0.04 |
| 72T | MM-KF | 4.1 | 0.03 |
| NGS Variables | Automated | Manual |
|---|---|---|
| Cluster passing filter | 92% | 85.9% |
| Q30 score | 85 | 80 |
| Error rate | 0.15% | 0.18% |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Muscarella, L.A.; Fabrizio, F.P.; De Bonis, M.; Mancini, M.T.; Balsamo, T.; Graziano, P.; Centra, F.; Sparaneo, A.; Trombetta, D.; Bonfitto, A.; et al. Automated Workflow for Somatic and Germline Next Generation Sequencing Analysis in Routine Clinical Cancer Diagnostics. Cancers 2019, 11, 1691. https://doi.org/10.3390/cancers11111691
Muscarella LA, Fabrizio FP, De Bonis M, Mancini MT, Balsamo T, Graziano P, Centra F, Sparaneo A, Trombetta D, Bonfitto A, et al. Automated Workflow for Somatic and Germline Next Generation Sequencing Analysis in Routine Clinical Cancer Diagnostics. Cancers. 2019; 11(11):1691. https://doi.org/10.3390/cancers11111691
Chicago/Turabian StyleMuscarella, Lucia Anna, Federico Pio Fabrizio, Maria De Bonis, Maria Teresa Mancini, Teresa Balsamo, Paolo Graziano, Flavia Centra, Angelo Sparaneo, Domenico Trombetta, Antonio Bonfitto, and et al. 2019. "Automated Workflow for Somatic and Germline Next Generation Sequencing Analysis in Routine Clinical Cancer Diagnostics" Cancers 11, no. 11: 1691. https://doi.org/10.3390/cancers11111691
APA StyleMuscarella, L. A., Fabrizio, F. P., De Bonis, M., Mancini, M. T., Balsamo, T., Graziano, P., Centra, F., Sparaneo, A., Trombetta, D., Bonfitto, A., Scagliusi, V., Larizza, P., Capoluongo, E. D., & Fazio, V. M. (2019). Automated Workflow for Somatic and Germline Next Generation Sequencing Analysis in Routine Clinical Cancer Diagnostics. Cancers, 11(11), 1691. https://doi.org/10.3390/cancers11111691






