Colorectal Cancer and Alcohol Consumption—Populations to Molecules
Abstract
1. Introduction
2. The Epidemiology of Alcohol and Colon Carcinogenesis
2.1. Alcohol and CRC
2.2. Alcohol and Polyps
3. The Metabolism of Ethanol
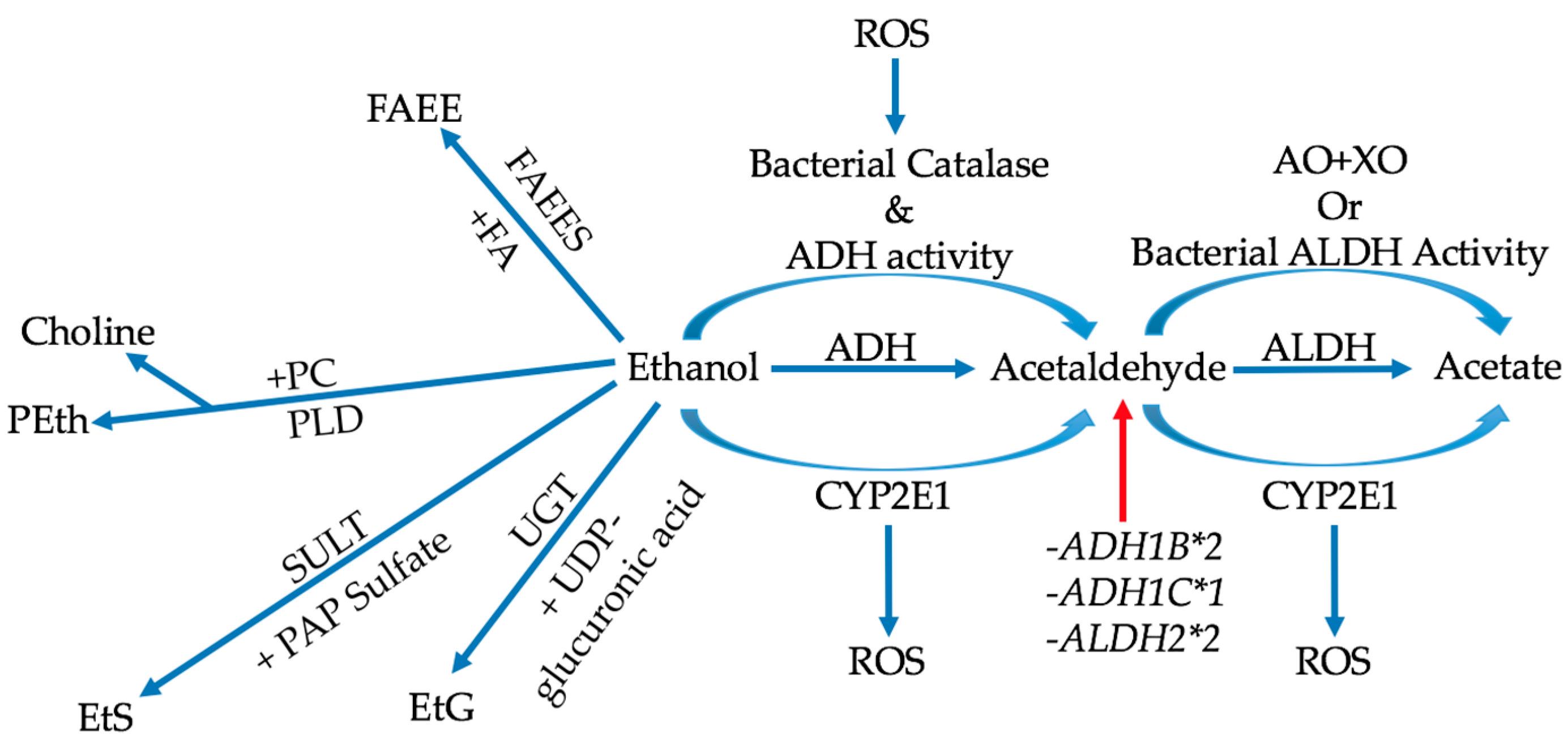
3.1. General Metabolism
3.2. Enzyme Polymorphisms and Bacterial Metabolism
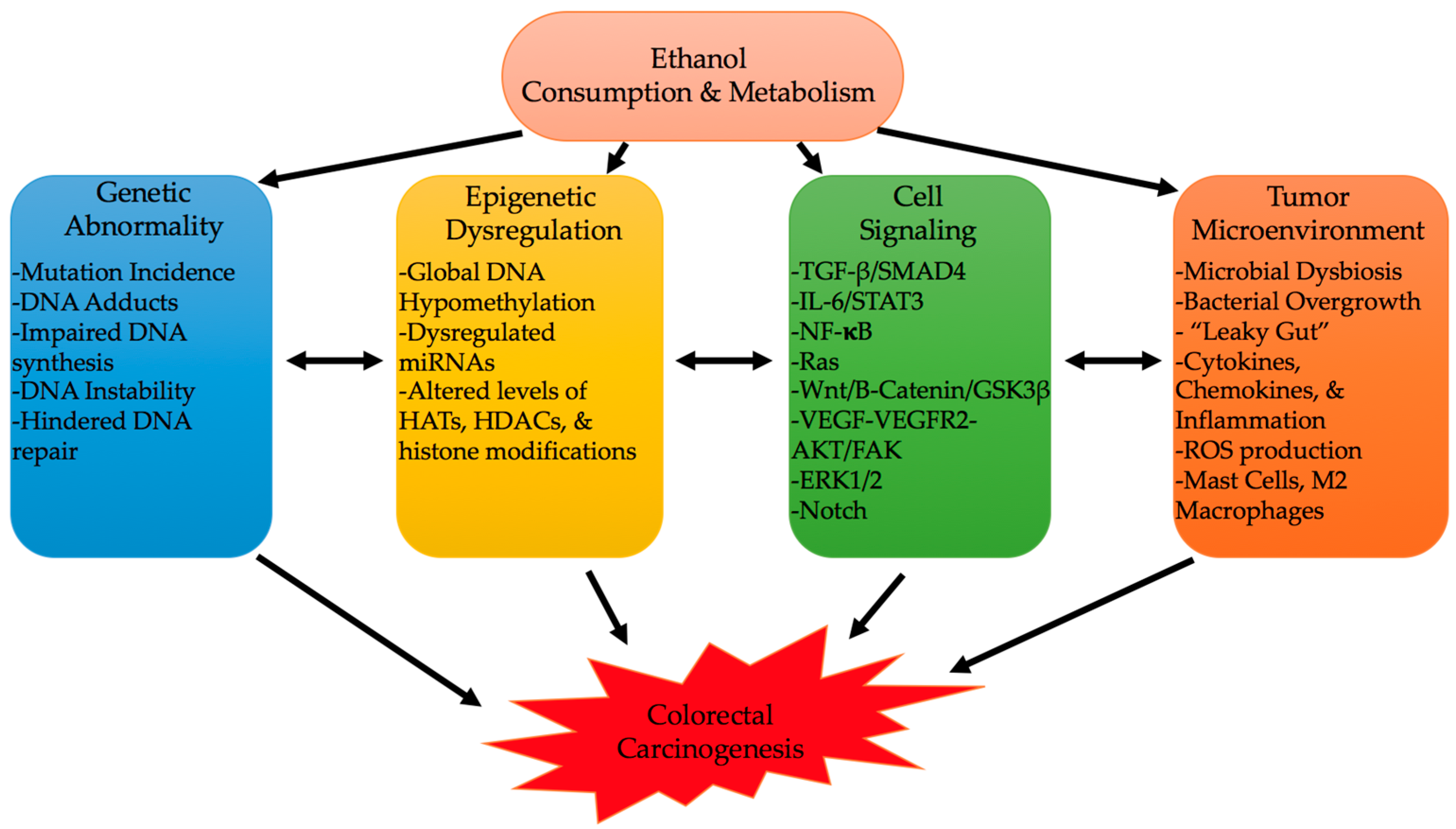
4. Genetic Stability, Epigenetic Modifications, and Mutation
4.1. Direct DNA Damage
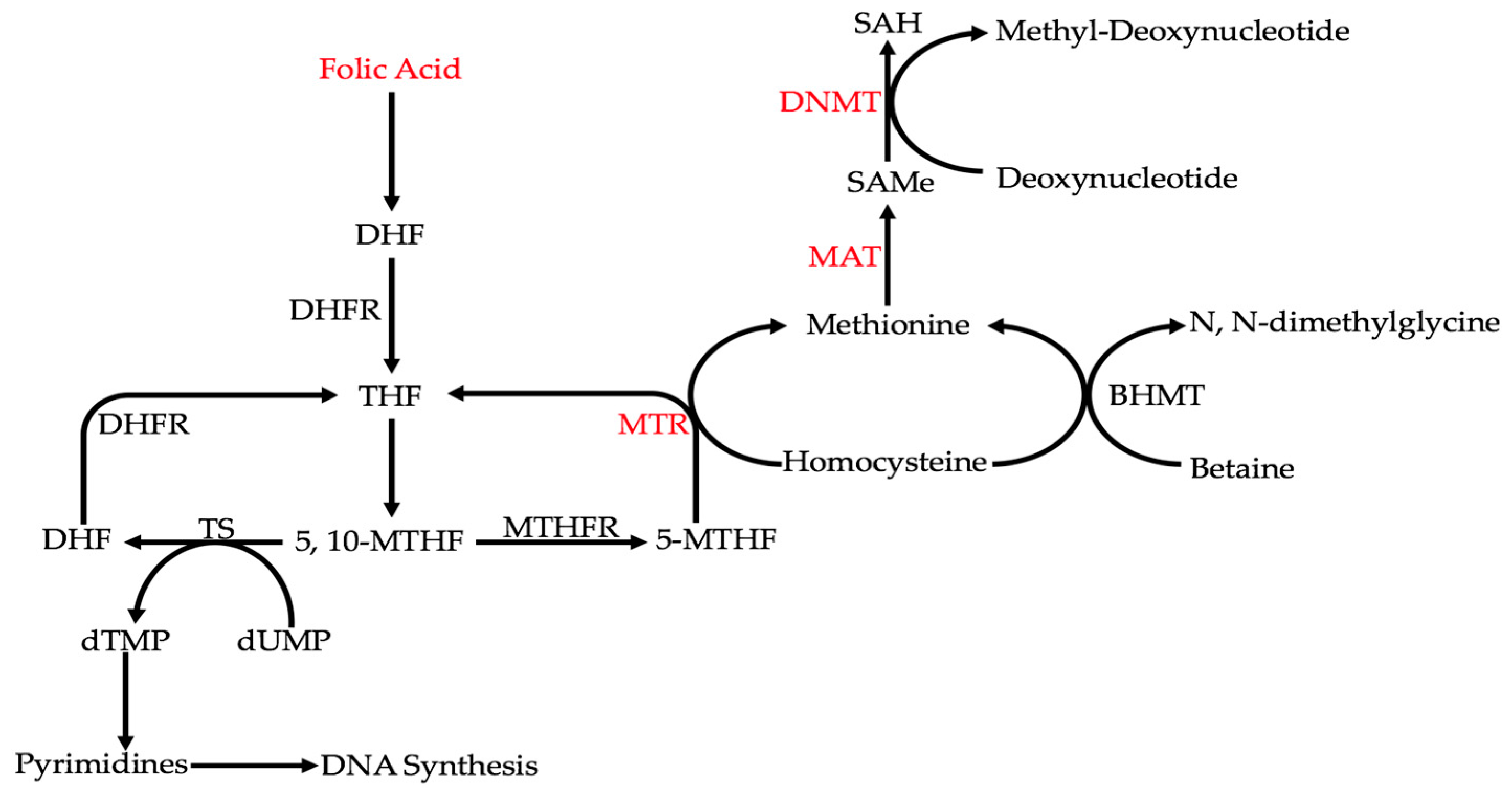
4.2. Indirect DNA Instability
5. Non-Coding RNAs, Cell Signaling, and Stemness
5.1. Non-Coding RNAs and Their Effects
5.2. Altered Histone Modifications
5.3. The Hedgehog Pathway
6. The Tumor Microenvironment
7. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Kolligs, F.T. Diagnostics and epidemiology of colorectal cancer. Visc. Med. 2016, 32, 158–164. [Google Scholar] [CrossRef] [PubMed]
- Roswall, N.; Weiderpass, E. Alcohol as a risk factor for cancer: Existing evidence in a global perspective. J. Prev. Med. Public Health 2015, 48, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Yabroff, K.R.; Warren, J.L.; Schrag, D.; Mariotto, A.; Meekins, A.; Topor, M.; Brown, M.L. Comparison of approaches for estimating incidence costs of care for colorectal cancer patients. Med. Care 2009, 47, S56–S63. [Google Scholar] [CrossRef] [PubMed]
- Bishehsari, F.; Mahdavinia, M.; Vacca, M.; Malekzadeh, R.; Mariani-Costantini, R. Epidemiological transition of colorectal cancer in developing countries: Environmental factors, molecular pathways, and opportunities for prevention. World J. Gastroenterol. 2014, 20, 6055–6072. [Google Scholar] [CrossRef] [PubMed]
- Bishehsari, F.; Magno, E.; Swanson, G.; Desai, V.; Voigt, R.M.; Forsyth, C.B.; Keshavarzian, A. Alcohol and gut-derived inflammation. Alcohol Res. Curr. Rev. 2017, 38, 163–171. [Google Scholar]
- Jemal, A.; Center, M.M.; DeSantis, C.; Ward, E.M. Global patterns of cancer incidence and mortality rates and trends. Cancer Epidemiol. Biomark. Prev. 2010, 19, 1893–1907. [Google Scholar] [CrossRef] [PubMed]
- Fedirko, V.; Tramacere, I.; Bagnardi, V.; Rota, M.; Scotti, L.; Islami, F.; Negri, E.; Straif, K.; Romieu, I.; La Vecchia, C.; et al. Alcohol drinking and colorectal cancer risk: An overall and dose-response meta-analysis of published studies. Ann. Oncol. 2011, 22, 1958–1972. [Google Scholar] [CrossRef]
- Seitz, H.K.; Becker, P. Alcohol metabolism and cancer risk. Alcohol Res. Health 2007, 30, 38–47. [Google Scholar] [PubMed]
- Shukla, S.D.; Lim, R.W. Epigenetic effects of ethanol on the liver and gastrointestinal system. Alcohol Res. Curr. Rev. 2013, 35, 47–55. [Google Scholar]
- Varela-Rey, M.; Woodhoo, A.; Martinez-Chantar, M.L.; Mato, J.M.; Lu, S.C. Alcohol, DNA methylation, and cancer. Alcohol Res. Curr. Rev. 2013, 35, 25–35. [Google Scholar]
- Dinis-Oliveira, R.J. Oxidative and non-oxidative metabolomics of ethanol. Curr. Drug Metab. 2016, 17, 327–335. [Google Scholar] [CrossRef] [PubMed]
- Haas, S.L.; Ye, W.; Lohr, J.M. Alcohol consumption and digestive tract cancer. Curr. Opin. Clin. Nutr. Metab. Care 2012, 15, 457–467. [Google Scholar] [CrossRef] [PubMed]
- Wimberly, A.L.; Forsyth, C.B.; Khan, M.W.; Pemberton, A.; Khazaie, K.; Keshavarzian, A. Ethanol-induced mast cell-mediated inflammation leads to increased susceptibility of intestinal tumorigenesis in the APC delta468 min mouse model of colon cancer. Alcohol. Clin. Exp. Res. 2013, 37, E199–E208. [Google Scholar] [CrossRef] [PubMed]
- Cai, S.; Li, Y.; Ding, Y.; Chen, K.; Jin, M. Alcohol drinking and the risk of colorectal cancer death: A meta-analysis. Eur. J. Cancer Prev. 2014, 23, 532–539. [Google Scholar] [CrossRef] [PubMed]
- Haggar, F.A.; Boushey, R.P. Colorectal cancer epidemiology: Incidence, mortality, survival, and risk factors. Clin. Colon Rectal Surg. 2009, 22, 191–197. [Google Scholar] [CrossRef] [PubMed]
- Yi, S.W.; Sull, J.W.; Linton, J.A.; Nam, C.M.; Ohrr, H. Alcohol consumption and digestive cancer mortality in Koreans: The Kangwha Cohort Study. J. Epidemiol. 2010, 20, 204–211. [Google Scholar] [CrossRef]
- Ibanez-Sanz, G.; Diez-Villanueva, A.; Alonso, M.H.; Rodriguez-Moranta, F.; Perez-Gomez, B.; Bustamante, M.; Martin, V.; Llorca, J.; Amiano, P.; Ardanaz, E.; et al. Risk model for colorectal cancer in Spanish population using environmental and genetic factors: Results from the MCC-Spain study. Sci. Rep. 2017, 7, 43263. [Google Scholar] [CrossRef] [PubMed]
- Klarich, D.S.; Brasser, S.M.; Hong, M.Y. Moderate alcohol consumption and colorectal cancer risk. Alcohol. Clin. Exp. Res. 2015, 39, 1280–1291. [Google Scholar] [CrossRef] [PubMed]
- Yun, J.W.; Son, M.J.; Abdelmegeed, M.A.; Banerjee, A.; Morgan, T.R.; Yoo, S.H.; Song, B.J. Binge alcohol promotes hypoxic liver injury through a CYP2E1-HIF-1alpha-dependent apoptosis pathway in mice and humans. Free Radic. Biol. Med. 2014, 77, 183–194. [Google Scholar] [CrossRef] [PubMed]
- Walter, V.; Jansen, L.; Ulrich, A.; Roth, W.; Blaker, H.; Chang-Claude, J.; Hoffmeister, M.; Brenner, H. Alcohol consumption and survival of colorectal cancer patients: A population-based study from germany. Am. J. Clin. Nutr. 2016, 103, 1497–1506. [Google Scholar] [CrossRef] [PubMed]
- Yang, B.; Gapstur, S.M.; Newton, C.C.; Jacobs, E.J.; Campbell, P.T. Alcohol intake and mortality among survivors of colorectal cancer: The cancer prevention study II nutrition cohort. Cancer 2017, 123, 2006–2013. [Google Scholar] [CrossRef] [PubMed]
- Pelser, C.; Arem, H.; Pfeiffer, R.M.; Elena, J.W.; Alfano, C.M.; Hollenbeck, A.R.; Park, Y. Prediagnostic lifestyle factors and survival after colon and rectal cancer diagnosis in the national institutes of health (NIH)-AARP diet and health study. Cancer 2014, 120, 1540–1547. [Google Scholar] [CrossRef] [PubMed]
- Ahrendt, S.A.; Hu, Y.; Buta, M.; McDermott, M.P.; Benoit, N.; Yang, S.C.; Wu, L.; Sidransky, D. P53 mutations and survival in stage I non-small-cell lung cancer: Results of a prospective study. J. Natl. Cancer Inst. 2003, 95, 961–970. [Google Scholar] [CrossRef] [PubMed]
- Wu, W.J.; Pruett, S.B. Ethanol decreases host resistance to pulmonary metastases in a mouse model: Role of natural killer cells and the ethanol-induced stress response. Int. J. Cancer 1999, 82, 886–892. [Google Scholar] [CrossRef]
- Paull, D.E.; Updyke, G.M.; Baumann, M.A.; Chin, H.W.; Little, A.G.; Adebonojo, S.A. Alcohol abuse predicts progression of disease and death in patients with lung cancer. Ann. Thorac. Surg. 2005, 80, 1033–1039. [Google Scholar] [CrossRef] [PubMed]
- Cho, E.; Lee, J.E.; Rimm, E.B.; Fuchs, C.S.; Giovannucci, E.L. Alcohol consumption and the risk of colon cancer by family history of colorectal cancer. Am. J. Clin. Nutr. 2012, 95, 413–419. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Zhu, Y.; Wang, P.P.; West, R.; Buehler, S.; Sun, Z.; Squires, J.; Roebothan, B.; McLaughlin, J.R.; Campbell, P.T.; et al. Interaction between alcohol drinking and obesity in relation to colorectal cancer risk: A case-control study in Newfoundland and Labrador, Canada. BMC Public Health 2012, 12, 94. [Google Scholar] [CrossRef] [PubMed]
- Owusu, D.; Quinn, M.; Wang, K.S. Alcohol consumption, depression, insomnia and colorectal cancer screening: Racial differences. Int. J. High Risk Behav. Addict. 2015, 4, e23424. [Google Scholar] [CrossRef]
- Gonzales, M.; Nelson, H.; Rhyne, R.L.; Stone, S.N.; Hoffman, R.M. Surveillance of colorectal cancer screening in New Mexico hispanics and non-hispanic whites. J. Community Health 2012, 37, 1279–1288. [Google Scholar] [CrossRef] [PubMed]
- Fagunwa, I.O.; Loughrey, M.B.; Coleman, H.G. Alcohol, smoking and the risk of premalignant and malignant colorectal neoplasms. Best Pract. Res. Clin. Gastroenterol. 2017, 31, 561–568. [Google Scholar] [CrossRef] [PubMed]
- Gao, C.M.; Takezaki, T.; Wu, J.Z.; Zhang, X.M.; Cao, H.X.; Ding, J.H.; Liu, Y.T.; Li, S.P.; Cao, J.; Matsuo, K.; et al. Polymorphisms of alcohol dehydrogenase 2 and aldehyde dehydrogenase 2 and colorectal cancer risk in Chinese males. World J. Gastroenterol. 2008, 14, 5078–5083. [Google Scholar] [CrossRef] [PubMed]
- Peng, G.S.; Chen, Y.C.; Tsao, T.P.; Wang, M.F.; Yin, S.J. Pharmacokinetic and pharmacodynamic basis for partial protection against alcoholism in Asians, heterozygous for the variant ALDH2*2 gene allele. Pharmacogenet. Genom. 2007, 17, 845–855. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.F.; Wang, J.; Yu, S.J.; Song, J.; Ji, M.Y.; Zhang, J.X.; Cao, Z.; Wang, J.; Dong, W.G. Meta-analysis of the ADH1B and ALDH2 polymorphisms and the risk of colorectal cancer in East Asians. Intern. Med. 2013, 52, 2693–2699. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Chen, C.; Wang, L.; Liao, Q.; Xu, L.; Huang, Y.; Zhang, C.; Ye, H.; Xu, X.; Ye, M.; Duan, S. Association between six genetic polymorphisms and colorectal cancer: A meta-analysis. Genet. Test. Mol. Biomark. 2014, 18, 187–195. [Google Scholar] [CrossRef] [PubMed]
- Chang, J.S.; Hsiao, J.R.; Chen, C.H. ALDH2 polymorphism and alcohol-related cancers in Asians: A public health perspective. J. Biomed. Sci. 2017, 24, 19. [Google Scholar] [CrossRef] [PubMed]
- Bongaerts, B.W.; van den Brandt, P.A.; Goldbohm, R.A.; de Goeij, A.F.; Weijenberg, M.P. Alcohol consumption, type of alcoholic beverage and risk of colorectal cancer at specific subsites. Int. J. Cancer 2008, 123, 2411–2417. [Google Scholar] [CrossRef] [PubMed]
- Ferrari, P.; Jenab, M.; Norat, T.; Moskal, A.; Slimani, N.; Olsen, A.; Tjonneland, A.; Overvad, K.; Jensen, M.K.; Boutron-Ruault, M.C.; et al. Lifetime and baseline alcohol intake and risk of colon and rectal cancers in the European prospective investigation into cancer and nutrition (EPIC). Int. J. Cancer 2007, 121, 2065–2072. [Google Scholar] [CrossRef] [PubMed]
- Baron, J.A.; Sandler, R.S.; Haile, R.W.; Mandel, J.S.; Mott, L.A.; Greenberg, E.R. Folate intake, alcohol consumption, cigarette smoking, and risk of colorectal adenomas. J. Natl. Cancer Inst. 1998, 90, 57–62. [Google Scholar] [CrossRef] [PubMed]
- Giacosa, A.; Frascio, F.; Munizzi, F. Epidemiology of colorectal polyps. Tech. Coloproctol. 2004, 8, s243–s247. [Google Scholar] [CrossRef] [PubMed]
- Honjo, S.; Kono, S.; Shinchi, K.; Imanishi, K.; Hirohata, T. Cigarette smoking, alcohol use and adenomatous polyps of the sigmoid colon. Jpn. J. Cancer Res. Gann 1992, 83, 806–811. [Google Scholar] [CrossRef] [PubMed]
- Cope, G.F.; Wyatt, J.I.; Pinder, I.F.; Lee, P.N.; Heatley, R.V.; Kelleher, J. Alcohol consumption in patients with colorectal adenomatous polyps. Gut 1991, 32, 70–72. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zhu, J.Z.; Wang, Y.M.; Zhou, Q.Y.; Zhu, K.F.; Yu, C.H.; Li, Y.M. Systematic review with meta-analysis: Alcohol consumption and the risk of colorectal adenoma. Aliment. Pharmacol. Ther. 2014, 40, 325–337. [Google Scholar] [CrossRef] [PubMed]
- Bailie, L.; Loughrey, M.B.; Coleman, H.G. Lifestyle risk factors for serrated colorectal polyps: A systematic review and meta-analysis. Gastroenterology 2017, 152, 92–104. [Google Scholar] [CrossRef] [PubMed]
- Makkar, R.; Pai, R.K.; Burke, C.A. Sessile serrated polyps: Cancer risk and appropriate surveillance. Clevel. Clin. J. Med. 2012, 79, 865–871. [Google Scholar] [CrossRef] [PubMed]
- Bardou, M.; Montembault, S.; Giraud, V.; Balian, A.; Borotto, E.; Houdayer, C.; Capron, F.; Chaput, J.C.; Naveau, S. Excessive alcohol consumption favours high risk polyp or colorectal cancer occurrence among patients with adenomas: A case control study. Gut 2002, 50, 38–42. [Google Scholar] [CrossRef] [PubMed]
- Ma, A.T.; Therrien, A.; Giard, J.M.; von Renteln, D.; Bouin, M. Alcoholic liver disease is a strong predictor of colorectal polyps in liver transplant recipients. Endosc. Int. Open 2017, 5, E918–E923. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Boutron, M.C.; Faivre, J.; Dop, M.C.; Quipourt, V.; Senesse, P. Tobacco, alcohol, and colorectal tumors: A multistep process. Am. J. Epidemiol. 1995, 141, 1038–1046. [Google Scholar] [PubMed]
- Steinmetz, J.; Spyckerelle, Y.; Gueguen, R.; Dupre, C. Alcohol, tobacco and colorectal adenomas and cancer. Case-control study in a population with positive fecal occult blood tests. Presse Med. (Paris Fr. 1983) 2007, 36, 1174–1182. [Google Scholar] [CrossRef] [PubMed]
- Hassan, C.; Pickhardt, P.J.; Marmo, R.; Choi, J.R. Impact of lifestyle factors on colorectal polyp detection in the screening setting. Dis. Colon Rectum 2010, 53, 1328–1333. [Google Scholar] [CrossRef] [PubMed]
- Setshedi, M.; Wands, J.R.; Monte, S.M. Acetaldehyde adducts in alcoholic liver disease. Oxid. Med. Cell. Longev. 2010, 3, 178–185. [Google Scholar] [CrossRef] [PubMed]
- Tsuruya, A.; Kuwahara, A.; Saito, Y.; Yamaguchi, H.; Tsubo, T.; Suga, S.; Inai, M.; Aoki, Y.; Takahashi, S.; Tsutsumi, E.; et al. Ecophysiological consequences of alcoholism on human gut microbiota: Implications for ethanol-related pathogenesis of colon cancer. Sci. Rep. 2016, 6, 27923. [Google Scholar] [CrossRef] [PubMed]
- Na, H.K.; Lee, J.Y. Molecular basis of alcohol-related gastric and colon cancer. Int. J. Mol. Sci. 2017, 18, 1116. [Google Scholar]
- Seitz, H.K.; Stickel, F. Molecular mechanisms of alcohol-mediated carcinogenesis. Nat. Rev. Cancer 2007, 7, 599–612. [Google Scholar] [CrossRef] [PubMed]
- Seitz, H.K.; Stickel, F. Acetaldehyde as an underestimated risk factor for cancer development: Role of genetics in ethanol metabolism. Genes Nutr. 2010, 5, 121–128. [Google Scholar] [CrossRef] [PubMed]
- Kruman, I.I.; Henderson, G.I.; Bergeson, S.E. DNA damage and neurotoxicity of chronic alcohol abuse. Exp. Biol. Med. 2012, 237, 740–747. [Google Scholar] [CrossRef] [PubMed]
- Heier, C.; Xie, H.; Zimmermann, R. Nonoxidative ethanol metabolism in humans-from biomarkers to bioactive lipids. IUBMB Life 2016, 68, 916–923. [Google Scholar] [CrossRef] [PubMed]
- Chiang, C.P.; Jao, S.W.; Lee, S.P.; Chen, P.C.; Chung, C.C.; Lee, S.L.; Nieh, S.; Yin, S.J. Expression pattern, ethanol-metabolizing activities, and cellular localization of alcohol and aldehyde dehydrogenases in human large bowel: Association of the functional polymorphisms of adh and aldh genes with hemorrhoids and colorectal cancer. Alcohol 2012, 46, 37–49. [Google Scholar] [CrossRef] [PubMed]
- Tillonen, J.; Kaihovaara, P.; Jousimies-Somer, H.; Heine, R.; Salaspuro, M. Role of catalase in in vitro acetaldehyde formation by human colonic contents. Alcohol. Clin. Exp. Res. 1998, 22, 1113–1119. [Google Scholar] [CrossRef] [PubMed]
- Tsuruya, A.; Kuwahara, A.; Saito, Y.; Yamaguchi, H.; Tenma, N.; Inai, M.; Takahashi, S.; Tsutsumi, E.; Suwa, Y.; Totsuka, Y.; et al. Major anaerobic bacteria responsible for the production of carcinogenic acetaldehyde from ethanol in the colon and rectum. Alcohol Alcohol. 2016, 51, 395–401. [Google Scholar] [CrossRef] [PubMed]
- Homann, N.; Tillonen, J.; Salaspuro, M. Microbially produced acetaldehyde from ethanol may increase the risk of colon cancer via folate deficiency. Int. J. Cancer 2000, 86, 169–173. [Google Scholar] [CrossRef]
- Visapaa, J.P.; Tillonen, J.; Salaspuro, M. Microbes and mucosa in the regulation of intracolonic acetaldehyde concentration during ethanol challenge. Alcohol Alcohol. 2002, 37, 322–326. [Google Scholar] [CrossRef] [PubMed]
- Elamin, E.E.; Masclee, A.A.; Dekker, J.; Jonkers, D.M. Ethanol metabolism and its effects on the intestinal epithelial barrier. Nutr. Rev. 2013, 71, 483–499. [Google Scholar] [CrossRef] [PubMed]
- Nosova, T.; Jokelainen, K.; Kaihovaara, P.; Heine, R.; Jousimies-Somer, H.; Salaspuro, M. Characteristics of aldehyde dehydrogenases of certain aerobic bacteria representing human colonic flora. Alcohol Alcohol. 1998, 33, 273–280. [Google Scholar] [CrossRef] [PubMed]
- Salaspuro, M. Microbial metabolism of ethanol and acetaldehyde and clinical consequences. Addict. Biol. 1997, 2, 35–46. [Google Scholar] [CrossRef] [PubMed]
- Moon, J.W.; Lee, S.K.; Lee, Y.W.; Lee, J.O.; Kim, N.; Lee, H.J.; Seo, J.S.; Kim, J.; Kim, H.S.; Park, S.H. Alcohol induces cell proliferation via hypermethylation of adhfe1 in colorectal cancer cells. BMC Cancer 2014, 14, 377. [Google Scholar] [CrossRef] [PubMed]
- Hutchison, J.; Cohen, Z.; Onyeagucha, B.C.; Funk, J.; Nelson, M.A. How micrornas influence both hereditary and inflammatory-mediated colon cancers. Cancer Genet. 2013, 206, 309–316. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Zhao, L.; Lei, L.; Lau, W.B.; Lau, B.; Yang, Q.; Le, X.; Yang, H.; Wang, C.; Luo, Z.; et al. Lncrnas: The bridge linking rna and colorectal cancer. Oncotarget 2017, 8, 12517–12532. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Bongaerts, B.W.; de Goeij, A.F.; de Vogel, S.; van den Brandt, P.A.; Goldbohm, R.A.; Weijenberg, M.P. Alcohol consumption and distinct molecular pathways to colorectal cancer. Br. J. Nutr. 2007, 97, 430–434. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Linhart, K.; Bartsch, H.; Seitz, H.K. The role of reactive oxygen species (ROS) and cytochrome P-450 2E1 in the generation of carcinogenic etheno-DNA adducts. Redox Biol. 2014, 3, 56–62. [Google Scholar] [CrossRef] [PubMed]
- Link, A.; Balaguer, F.; Shen, Y.; Lozano, J.J.; Leung, H.C.; Boland, C.R.; Goel, A. Curcumin modulates DNA methylation in colorectal cancer cells. PLoS ONE 2013, 8, e57709. [Google Scholar] [CrossRef] [PubMed]
- Hur, K.; Cejas, P.; Feliu, J.; Moreno-Rubio, J.; Burgos, E.; Boland, C.R.; Goel, A. Hypomethylation of long interspersed nuclear element-1 (LINE-1) leads to activation of proto-oncogenes in human colorectal cancer metastasis. Gut 2014, 63, 635–646. [Google Scholar] [CrossRef] [PubMed]
- Eussen, S.J.; Vollset, S.E.; Igland, J.; Meyer, K.; Fredriksen, A.; Ueland, P.M.; Jenab, M.; Slimani, N.; Boffetta, P.; Overvad, K.; et al. Plasma folate, related genetic variants, and colorectal cancer risk in EPIC. Cancer Epidemiol. Biomark. Prev. 2010, 19, 1328–1340. [Google Scholar] [CrossRef] [PubMed]
- McGlynn, A.P.; Wasson, G.R.; O’Reilly, S.L.; McNulty, H.; Downes, C.S.; Chang, C.K.; Hoey, L.; Molloy, A.M.; Ward, M.; Strain, J.J.; et al. Low colonocyte folate is associated with uracil misincorporation and global DNA hypomethylation in human colorectum. J. Nutr. 2013, 143, 27–33. [Google Scholar] [CrossRef] [PubMed]
- Hiraoka, M.; Kagawa, Y. Genetic polymorphisms and folate status. Congenit. Anom. 2017, 57, 142–149. [Google Scholar] [CrossRef] [PubMed]
- Schernhammer, E.S.; Giovannucci, E.; Baba, Y.; Fuchs, C.S.; Ogino, S. B vitamins, methionine and alcohol intake and risk of colon cancer in relation to BRAF mutation and CpG island methylator phenotype (CIMP). PLoS ONE 2011, 6, e21102. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.H.; Wang, L.A.; Li, Z.; Peng, Y.; Cun, Y.P.; Dai, N.; Cheng, Y.; Xiao, H.; Xiong, Y.L.; Wang, D. APE1 polymorphisms are associated with colorectal cancer susceptibility in Chinese Hans. World J. Gastroenterol. 2014, 20, 8700–8708. [Google Scholar] [CrossRef] [PubMed]
- Sinha, R.; Hussain, S.; Mehrotra, R.; Kumar, R.S.; Kumar, K.; Pande, P.; Doval, D.C.; Basir, S.F.; Bharadwaj, M. Kras gene mutation and RASSF1A, FHIT and MGMT gene promoter hypermethylation: Indicators of tumor staging and metastasis in adenocarcinomatous sporadic colorectal cancer in Indian population. PLoS ONE 2013, 8, e60142. [Google Scholar] [CrossRef] [PubMed]
- Hansen, R.D.; Sorensen, M.; Tjonneland, A.; Overvad, K.; Wallin, H.; Raaschou-Nielsen, O.; Vogel, U. XPA A23G, XPC Lys939Gln, XPD Lys751Gln and XPD Asp312Asn polymorphisms, interactions with smoking, alcohol and dietary factors, and risk of colorectal cancer. Mutat. Res. 2007, 619, 68–80. [Google Scholar] [CrossRef] [PubMed]
- Monzo, M.; Moreno, I.; Navarro, A.; Ibeas, R.; Artells, R.; Gel, B.; Martinez, F.; Moreno, J.; Hernandez, R.; Navarro-Vigo, M. Single nucleotide polymorphisms in nucleotide excision repair genes XPA, XPD, XPG and ERCC1 in advanced colorectal cancer patients treated with first-line oxaliplatin/fluoropyrimidine. Oncology 2007, 72, 364–370. [Google Scholar] [CrossRef] [PubMed]
- Gil, J.; Ramsey, D.; Stembalska, A.; Karpinski, P.; Pesz, K.A.; Laczmanska, I.; Leszczynski, P.; Grzebieniak, Z.; Sasiadek, M.M. The C/A polymorphism in intron 11 of the XPC gene plays a crucial role in the modulation of an individual’s susceptibility to sporadic colorectal cancer. Mol. Biol. Rep. 2012, 39, 527–534. [Google Scholar] [CrossRef] [PubMed]
- Scharer, O.D. Nucleotide excision repair in eukaryotes. Cold Spring Harb. Perspect. Biol. 2013, 5, a012609. [Google Scholar] [CrossRef] [PubMed]
- Brenerman, B.M.; Illuzzi, J.L.; Wilson, D.M., 3rd. Base excision repair capacity in informing healthspan. Carcinogenesis 2014, 35, 2643–2652. [Google Scholar] [CrossRef] [PubMed]
- Slyskova, J.; Naccarati, A.; Pardini, B.; Polakova, V.; Vodickova, L.; Smerhovsky, Z.; Levy, M.; Lipska, L.; Liska, V.; Vodicka, P. Differences in nucleotide excision repair capacity between newly diagnosed colorectal cancer patients and healthy controls. Mutagenesis 2012, 27, 225–232. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Yu, Y.; Nangia-Makker, P.; Farhana, L.; Rajendra, S.G.; Levi, E.; Majumdar, A.P. miR-21 and miR-145 cooperation in regulation of colon cancer stem cells. Mol. Cancer 2015, 14, 98. [Google Scholar] [CrossRef] [PubMed]
- Chi, Y.; Zhou, D. Micrornas in colorectal carcinoma—From pathogenesis to therapy. J. Exp. Clin. Cancer Res. 2016, 35, 43. [Google Scholar] [CrossRef] [PubMed]
- Lamichhane, T.N.; Leung, C.A.; Douti, L.Y.; Jay, S.M. Ethanol induces enhanced vascularization bioactivity of endothelial cell-derived extracellular vesicles via regulation of microRNAs and long non-coding RNAs. Sci. Rep. 2017, 7, 13794. [Google Scholar] [CrossRef] [PubMed]
- Almeida, A.L.; Bernardes, M.V.; Feitosa, M.R.; Peria, F.M.; Tirapelli, D.P.; Rocha, J.J.; Feres, O. Serological under expression of microRNA-21, microRNA-34a and microRNA-126 in colorectal cancer. Acta Cir. Bras. 2016, 31, 13–18. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Bu, P.; Ai, Y.; Srinivasan, T.; Chen, H.J.; Xiang, K.; Lipkin, S.M.; Shen, X. A long non-coding RNA targets microRNA miR-34a to regulate colon cancer stem cell asymmetric division. eLife 2016, 5, e14620. [Google Scholar] [CrossRef] [PubMed]
- Sun, C.; Wang, F.J.; Zhang, H.G.; Xu, X.Z.; Jia, R.C.; Yao, L.; Qiao, P.F. miR-34a mediates oxaliplatin resistance of colorectal cancer cells by inhibiting macroautophagy via transforming growth factor-beta/smad4 pathway. World J. Gastroenterol. 2017, 23, 1816–1827. [Google Scholar] [CrossRef] [PubMed]
- Krattinger, R.; Bostrom, A.; Lee, S.M.L.; Thasler, W.E.; Schioth, H.B.; Kullak-Ublick, G.A.; Mwinyi, J. Chenodeoxycholic acid significantly impacts the expression of miRNAs and genes involved in lipid, bile acid and drug metabolism in human hepatocytes. Life Sci. 2016, 156, 47–56. [Google Scholar] [CrossRef] [PubMed]
- Lavery, C.A.; Kurowska-Stolarska, M.; Holmes, W.M.; Donnelly, I.; Caslake, M.; Collier, A.; Baker, A.H.; Miller, A.M. miR-34a(⁻/⁻) mice are susceptible to diet-induced obesity. Obesity 2016, 24, 1741–1751. [Google Scholar] [CrossRef] [PubMed]
- Salvoza, N.C.; Klinzing, D.C.; Gopez-Cervantes, J.; Baclig, M.O. Association of circulating serum miR-34a and miR-122 with dyslipidemia among patients with non-alcoholic fatty liver disease. PLoS ONE 2016, 11, e0153497. [Google Scholar] [CrossRef] [PubMed]
- Dippold, R.P.; Vadigepalli, R.; Gonye, G.E.; Patra, B.; Hoek, J.B. Chronic ethanol feeding alters miRNA expression dynamics during liver regeneration. Alcohol. Clin. Exp. Res. 2013, 37, E59–E69. [Google Scholar] [CrossRef] [PubMed]
- Barr, T.; Girke, T.; Sureshchandra, S.; Nguyen, C.; Grant, K.; Messaoudi, I. Alcohol consumption modulates host defense in rhesus macaques by altering gene expression in circulating leukocytes. J. Immunol. 2016, 196, 182–195. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Liu, H.; Jin, W.; Ding, Z.; Zheng, S.; Yu, Y. Tissue microRNA-21 expression predicted recurrence and poor survival in patients with colorectal cancer—A meta-analysis. Onco Targets Ther. 2016, 9, 2615–2624. [Google Scholar] [PubMed]
- Mima, K.; Nishihara, R.; Nowak, J.A.; Kim, S.A.; Song, M.; Inamura, K.; Sukawa, Y.; Masuda, A.; Yang, J.; Dou, R.; et al. MicroRNA miR21 and T cells in colorectal cancer. Cancer Immunol. Res. 2016, 4, 33–40. [Google Scholar] [CrossRef] [PubMed]
- Francis, H.; McDaniel, K.; Han, Y.; Liu, X.; Kennedy, L.; Yang, F.; McCarra, J.; Zhou, T.; Glaser, S.; Venter, J.; et al. Regulation of the extrinsic apoptotic pathway by microRNA-21 in alcoholic liver injury. J. Biol. Chem. 2014, 289, 27526–27539. [Google Scholar] [CrossRef] [PubMed]
- Yeligar, S.; Tsukamoto, H.; Kalra, V.K. Ethanol-induced expression of ET-1 and ET-BR in liver sinusoidal endothelial cells and human endothelial cells involves hypoxia-inducible factor-1alpha and microRNA-199. J. Immunol. 2009, 183, 5232–5243. [Google Scholar] [CrossRef] [PubMed]
- Nagel, R.; le Sage, C.; Diosdado, B.; van der Waal, M.; Oude Vrielink, J.A.; Bolijn, A.; Meijer, G.A.; Agami, R. Regulation of the adenomatous polyposis coli gene by the miR-135 family in colorectal cancer. Cancer Res. 2008, 68, 5795–5802. [Google Scholar] [CrossRef] [PubMed]
- Golubovskaya, V.M.; Sumbler, B.; Ho, B.; Yemma, M.; Cance, W.G. miR-138 and miR-135 directly target focal adhesion kinase, inhibit cell invasion, and increase sensitivity to chemotherapy in cancer cells. Anti-Cancer Agents Med. Chem. 2014, 14, 18–28. [Google Scholar] [CrossRef]
- Ma, Y.; Semba, S.; Khan, R.I.; Bochimoto, H.; Watanabe, T.; Fujiya, M.; Kohgo, Y.; Liu, Y.; Taniguchi, T. Focal adhesion kinase regulates intestinal epithelial barrier function via redistribution of tight junction. Biochim. Biophys. Acta 2013, 1832, 151–159. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, Y.; Peng, H.; Mastej, V.; Chen, W. Microrna regulation of endothelial junction proteins and clinical consequence. Mediat. Inflamm. 2016, 2016, 5078627. [Google Scholar] [CrossRef] [PubMed]
- Bi, Y.L.; Mi, P.Y.; Zhao, S.J.; Pan, H.M.; Li, H.J.; Liu, F.; Shao, L.R.; Zhang, H.F.; Zhang, P.; Jiang, S.L. Salinomycin exhibits anti-angiogenic activity against human glioma in vitro and in vivo by suppressing the VEGF-VEGFR2-AKT/FAK signaling axis. Int. J. Mol. Med. 2017, 39, 1255–1261. [Google Scholar] [CrossRef] [PubMed]
- Pal-Bhadra, M.; Bhadra, U.; Jackson, D.E.; Mamatha, L.; Park, P.H.; Shukla, S.D. Distinct methylation patterns in histone H3 at Lys-4 and Lys-9 correlate with up- & down-regulation of genes by ethanol in hepatocytes. Life Sci. 2007, 81, 979–987. [Google Scholar] [PubMed]
- Aroor, A.R.; James, T.T.; Jackson, D.E.; Shukla, S.D. Differential changes in map kinases, histone modifications, and liver injury in rats acutely treated with ethanol. Alcohol. Clin. Exp. Res. 2010, 34, 1543–1551. [Google Scholar] [CrossRef] [PubMed]
- Bardag-Gorce, F.; French, B.A.; Joyce, M.; Baires, M.; Montgomery, R.O.; Li, J.; French, S. Histone acetyltransferase p300 modulates gene expression in an epigenetic manner at high blood alcohol levels. Exp. Mol. Pathol. 2007, 82, 197–202. [Google Scholar] [CrossRef] [PubMed]
- Choudhury, M.; Pandey, R.S.; Clemens, D.L.; Davis, J.W.; Lim, R.W.; Shukla, S.D. Knock down of GCN5 histone acetyltransferase by siRNA decreases ethanol-induced histone acetylation and affects differential expression of genes in human hepatoma cells. Alcohol 2011, 45, 311–324. [Google Scholar] [CrossRef] [PubMed]
- Kirpich, I.; Ghare, S.; Zhang, J.; Gobejishvili, L.; Kharebava, G.; Barve, S.J.; Barker, D.; Moghe, A.; McClain, C.J.; Barve, S. Binge alcohol-induced microvesicular liver steatosis and injury are associated with down-regulation of hepatic Hdac 1, 7, 9, 10, 11 and up-regulation of Hdac 3. Alcohol. Clin. Exp. Res. 2012, 36, 1578–1586. [Google Scholar] [CrossRef] [PubMed]
- Chan, I.S.; Guy, C.D.; Machado, M.V.; Wank, A.; Kadiyala, V.; Michelotti, G.; Choi, S.; Swiderska-Syn, M.; Karaca, G.; Pereira, T.A.; et al. Alcohol activates the hedgehog pathway and induces related procarcinogenic processes in the alcohol-preferring rat model of hepatocarcinogenesis. Alcohol. Clin. Exp. Res. 2014, 38, 787–800. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.W.; Lin, H.; Lu, Y.; Xia, W.; Gao, J.; Li, Z.S. Sonic hedgehog expression in a rat model of chronic pancreatitis. World J. Gastroenterol. 2014, 20, 4712–4717. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.X.; Yang, H.T.; Zdanowicz, M.; Sicklick, J.K.; Qi, Y.; Camp, T.J.; Diehl, A.M. Fetal alcohol exposure impairs hedgehog cholesterol modification and signaling. Lab. Invest. 2007, 87, 231–240. [Google Scholar] [CrossRef] [PubMed]
- Latchoumycandane, C.; Hanouneh, M.; Nagy, L.E.; McIntyre, T.M. Inflammatory paf receptor signaling initiates hedgehog signaling and kidney fibrogenesis during ethanol consumption. PLoS ONE 2015, 10, e0145691. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Wei, L.; Shen, A.; Cai, Q.; Xu, W.; Li, H.; Zhan, Y.; Hong, Z.; Peng, J. Hedyotis diffusa willd extract suppresses sonic hedgehog signaling leading to the inhibition of colorectal cancer angiogenesis. Int. J. Oncol. 2013, 42, 651–656. [Google Scholar] [CrossRef] [PubMed]
- Wei, L.; Lin, J.; Xu, W.; Cai, Q.; Shen, A.; Hong, Z.; Peng, J. Scutellaria barbata D. Don inhibits tumor angiogenesis via suppression of hedgehog pathway in a mouse model of colorectal cancer. Int. J. Mol. Sci. 2012, 13, 9419–9430. [Google Scholar] [CrossRef] [PubMed]
- Nicholas, N.S.; Apollonio, B.; Ramsay, A.G. Tumor microenvironment (TME)-driven immune suppression in B cell malignancy. Biochim. Biophys. Acta 2016, 1863, 471–482. [Google Scholar] [CrossRef] [PubMed]
- Moossavi, S.; Bishehsari, F. Inflammation in sporadic colorectal cancer. Arch. Iran. Med. 2012, 15, 166–170. [Google Scholar] [PubMed]
- Engen, P.A.; Green, S.J.; Voigt, R.M.; Forsyth, C.B.; Keshavarzian, A. The gastrointestinal microbiome: Alcohol effects on the composition of intestinal microbiota. Alcohol Res. 2015, 37, 223–236. [Google Scholar] [PubMed]
- Elamin, E.; Masclee, A.; Troost, F.; Dekker, J.; Jonkers, D. Activation of the epithelial-to-mesenchymal transition factor snail mediates acetaldehyde-induced intestinal epithelial barrier disruption. Alcohol. Clin. Exp. Res. 2014, 38, 344–353. [Google Scholar] [CrossRef] [PubMed]
- Davis, B.T.; Voigt, R.M.; Shaikh, M.; Forsyth, C.B.; Keshavarzian, A. CREB protein mediates alcohol-induced circadian disruption and intestinal permeability. Alcohol. Clin. Exp. Res. 2017, 41, 2007–2014. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.; Forsyth, C.B.; Farhadi, A.; Rangan, J.; Jakate, S.; Shaikh, M.; Banan, A.; Fields, J.Z.; Keshavarzian, A. Nitric oxide-mediated intestinal injury is required for alcohol-induced gut leakiness and liver damage. Alcohol. Clin. Exp. Res. 2009, 33, 1220–1230. [Google Scholar] [CrossRef] [PubMed]
- Forsyth, C.B.; Shaikh, M.; Bishehsari, F.; Swanson, G.; Voigt, R.M.; Dodiya, H.; Wilkinson, P.; Samelco, B.; Song, S.; Keshavarzian, A. Alcohol feeding in mice promotes colonic hyperpermeability and changes in colonic organoid stem cell fate. Alcohol. Clin. Exp. Res. 2017, 41, 2100–2113. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Wang, S.; Qi, Y.; Chen, L.; Frank, J.A.; Yang, X.H.; Zhang, Z.; Shi, X.; Luo, J. Role of MCP-1 in alcohol-induced aggressiveness of colorectal cancer cells. Mol. Carcinog. 2016, 55, 1002–1011. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Zhang, F.; Ren, H.; Luo, J.; Wang, S. Role of cytokines and chemokines in alcohol-induced tumor promotion. OncoTargets Ther. 2017, 10, 1665–1671. [Google Scholar] [CrossRef] [PubMed]
- Hammer, A.M.; Morris, N.L.; Earley, Z.M.; Choudhry, M.A. The first line of defense: The effects of alcohol on post-burn intestinal barrier, immune cells, and microbiome. Alcohol Res. Curr. Rev. 2015, 37, 209–222. [Google Scholar]
- Tsuchimoto, Y.; Asai, A.; Tsuda, Y.; Ito, I.; Nishiguchi, T.; Garcia, M.C.; Suzuki, S.; Kobayashi, M.; Higuchi, K.; Suzuki, F. M2b monocytes provoke bacterial pneumonia and gut bacteria-associated sepsis in alcoholics. J. Immunol. 2015, 195, 5169–5177. [Google Scholar] [CrossRef] [PubMed]
- Roelands, J.; Kuppen, P.J.K.; Vermeulen, L.; Maccalli, C.; Decock, J.; Wang, E.; Marincola, F.M.; Bedognetti, D.; Hendrickx, W. Immunogenomic classification of colorectal cancer and therapeutic implications. Int. J. Mol. Sci. 2017, 18. [Google Scholar] [CrossRef] [PubMed]
- Patel, S.; Behara, R.; Swanson, G.R.; Forsyth, C.B.; Voigt, R.M.; Keshavarzian, A. Alcohol and the intestine. Biomolecules 2015, 5, 2573–2588. [Google Scholar] [CrossRef] [PubMed]
- Bishehsari, F.; Saadalla, A.; Khazaie, K.; Engen, P.A.; Voigt, R.M.; Shetuni, B.B.; Forsyth, C.; Shaikh, M.; Vitaterna, M.H.; Turek, F.; et al. Light/dark shifting promotes alcohol-induced colon carcinogenesis: Possible role of intestinal inflammatory milieu and microbiota. Int. J. Mol. Sci. 2016, 17. [Google Scholar] [CrossRef] [PubMed]
- Abdelmegeed, M.A.; Banerjee, A.; Jang, S.; Yoo, S.H.; Yun, J.W.; Gonzalez, F.J.; Keshavarzian, A.; Song, B.J. CYP2E1 potentiates binge alcohol-induced gut leakiness, steatohepatitis, and apoptosis. Free Radic. Biol. Med. 2013, 65, 1238–1245. [Google Scholar] [CrossRef] [PubMed]
- Shukla, P.K.; Chaudhry, K.K.; Mir, H.; Gangwar, R.; Yadav, N.; Manda, B.; Meena, A.S.; Rao, R. Chronic ethanol feeding promotes azoxymethane and dextran sulfate sodium-induced colonic tumorigenesis potentially by enhancing mucosal inflammation. BMC Cancer 2016, 16, 189. [Google Scholar] [CrossRef] [PubMed]
- Voigt, R.M.; Forsyth, C.B.; Keshavarzian, A. Circadian disruption: Potential implications in inflammatory and metabolic diseases associated with alcohol. Alcohol Res. 2013, 35, 87–96. [Google Scholar] [PubMed]
- Summa, K.C.; Voigt, R.M.; Forsyth, C.B.; Shaikh, M.; Cavanaugh, K.; Tang, Y.; Vitaterna, M.H.; Song, S.; Turek, F.W.; Keshavarzian, A. Disruption of the circadian clock in mice increases intestinal permeability and promotes alcohol-induced hepatic pathology and inflammation. PLoS ONE 2013, 8, e67102. [Google Scholar] [CrossRef] [PubMed]
- Preuss, F.; Tang, Y.; Laposky, A.D.; Arble, D.; Keshavarzian, A.; Turek, F.W. Adverse effects of chronic circadian desynchronization in animals in a “challenging” environment. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2008, 295, R2034–R2040. [Google Scholar] [CrossRef] [PubMed]
- Morimoto, L.M.; Newcomb, P.A.; Ulrich, C.M.; Bostick, R.M.; Lais, C.J.; Potter, J.D. Risk factors for hyperplastic and adenomatous polyps: Evidence for malignant potential? Cancer Epidemiol. Biomark. Prev. 2002, 11, 1012–1018. [Google Scholar]
- Erhardt, J.G.; Kreichgauer, H.P.; Meisner, C.; Bode, J.C.; Bode, C. Alcohol, cigarette smoking, dietary factors and the risk of colorectal adenomas and hyperplastic polyps—A case control study. Eur. J. Nutr. 2002, 41, 35–43. [Google Scholar] [CrossRef] [PubMed]
- Giovannucci, E.; Rimm, E.B.; Ascherio, A.; Stampfer, M.J.; Colditz, G.A.; Willett, W.C. Alcohol, low-methionine-low-folate diets, and risk of colon cancer in men. J. Natl. Cancer Inst. 1995, 87, 265–273. [Google Scholar] [CrossRef] [PubMed]
- Kune, G.A.; Vitetta, L. Alcohol consumption and the etiology of colorectal cancer: A review of the scientific evidence from 1957 to 1991. Nutr. Cancer 1992, 18, 97–111. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rossi, M.; Jahanzaib Anwar, M.; Usman, A.; Keshavarzian, A.; Bishehsari, F. Colorectal Cancer and Alcohol Consumption—Populations to Molecules. Cancers 2018, 10, 38. https://doi.org/10.3390/cancers10020038
Rossi M, Jahanzaib Anwar M, Usman A, Keshavarzian A, Bishehsari F. Colorectal Cancer and Alcohol Consumption—Populations to Molecules. Cancers. 2018; 10(2):38. https://doi.org/10.3390/cancers10020038
Chicago/Turabian StyleRossi, Marco, Muhammad Jahanzaib Anwar, Ahmad Usman, Ali Keshavarzian, and Faraz Bishehsari. 2018. "Colorectal Cancer and Alcohol Consumption—Populations to Molecules" Cancers 10, no. 2: 38. https://doi.org/10.3390/cancers10020038
APA StyleRossi, M., Jahanzaib Anwar, M., Usman, A., Keshavarzian, A., & Bishehsari, F. (2018). Colorectal Cancer and Alcohol Consumption—Populations to Molecules. Cancers, 10(2), 38. https://doi.org/10.3390/cancers10020038




