The Use of Microfluidic Technology for Cancer Applications and Liquid Biopsy
Abstract
1. Introduction
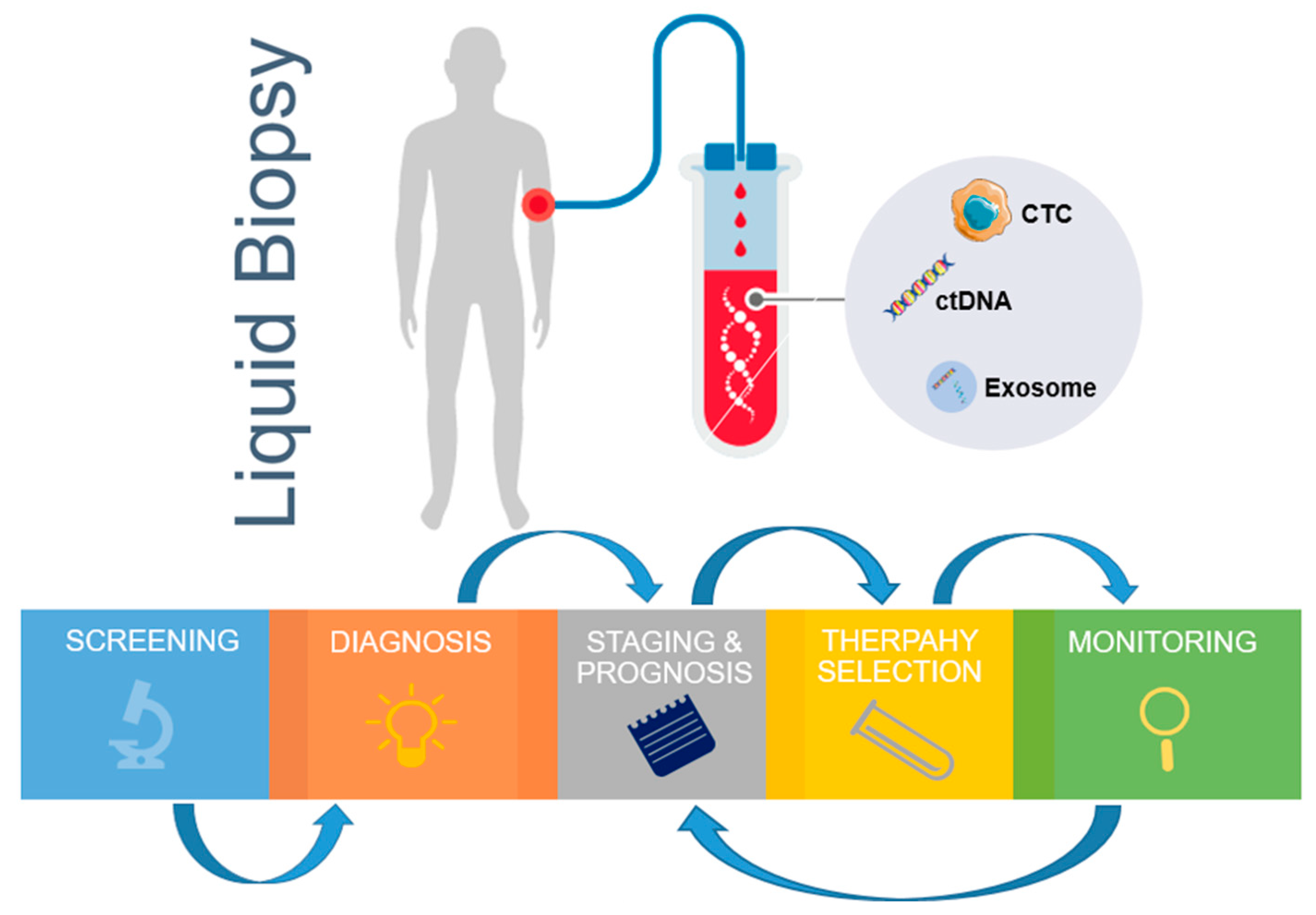
2. Circulating Biomarkers in Liquid Biopsy. Molecular, Biology, and Morphology Characteristics of CTC, ctDNA, and Exosomes
2.1. Circulating Tumor Cells (CTCs)
2.2. Monitoring of Disease
2.3. Epithelial to Mesenchymal Transition (EMT)
2.4. Biomarkers for Immunotherapy
2.5. Exosomes
2.6. ctDNA
3. The Role of Microfluidics in Liquid Biopsy
4. Circulating Tumor Cells Separation and Analysis on Microfluidic Chip
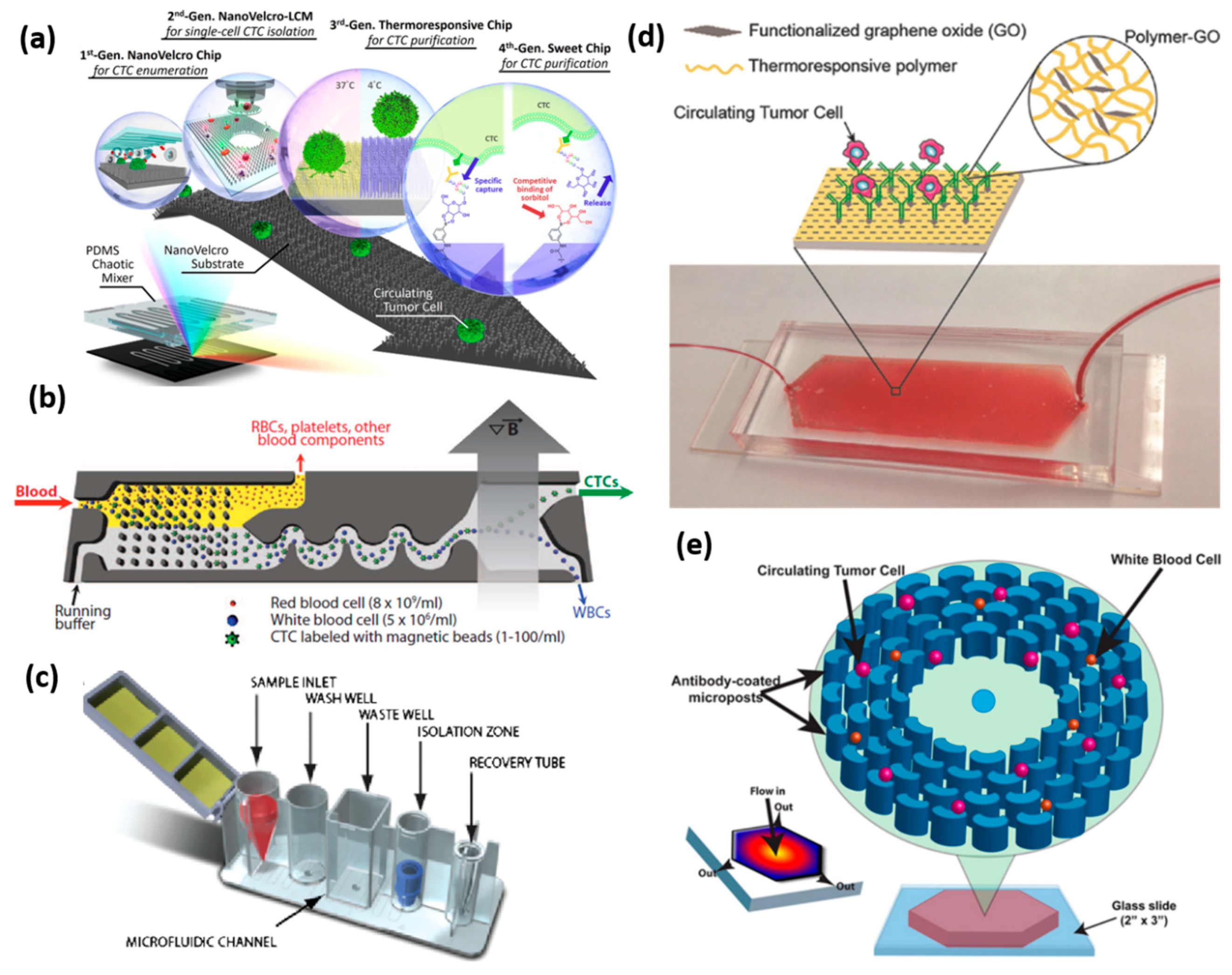
4.1. Postitive Selection of CTCs
4.1.1. Antibody-Coated Microstructure Separation
4.1.2. In Vivo Antibody-Coated Device Capture
4.1.3. Antibody-Coated Particles Capture Platform
4.2. Negative Seletion of CTCs
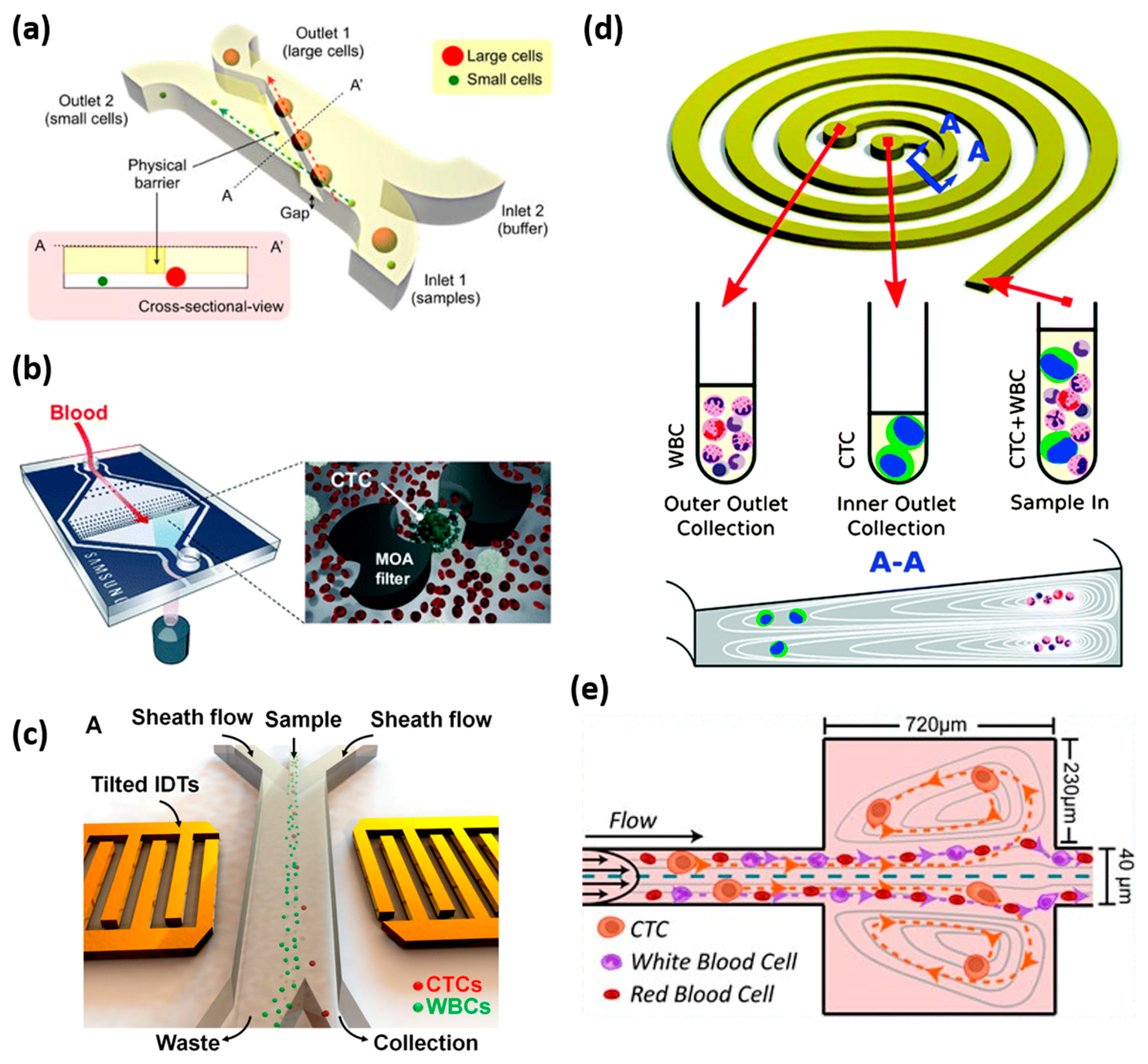
4.3. Label-Free Separation of CTCs
4.3.1. Size-Based Filtration
4.3.2. Density-Based Separation
4.3.3. Hydrodynamic-Based Separation
4.3.4. Acoustic Separation
4.3.5. Magnetophoresis Separation
4.3.6. Dielectrophoresis Separation
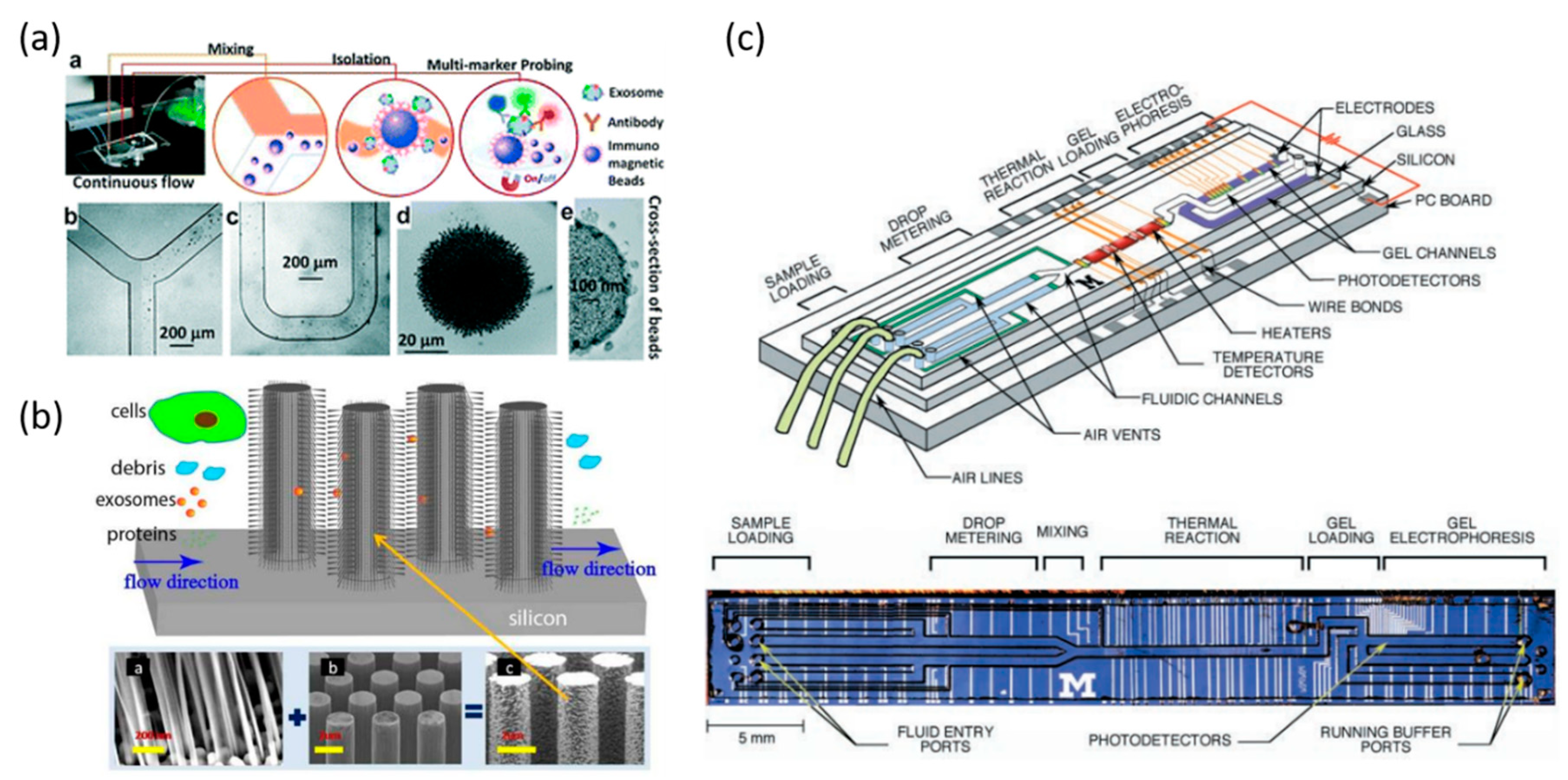
5. Exosome Separation and Analysis
5.1. Immuno-Affinity Based Separation and Analysis
5.2. Size-Based Exosomes Separation and Analysis
5.3. Other Technologies for Exosome Isolation
6. Circulating Tumor DNA Separation and Analysis
7. Microfluidic Techniques in CAR-T Cell Therapy
8. Challenges and Perspectives
Acknowledgments
Conflicts of Interest
References
- Lewis, J.M.; Heineck, D.P.; Heller, M.J. Detecting cancer biomarkers in blood: Challenges for new molecular diagnostic and point-of-care tests using cell-free nucleic acids. Expert Rev. Mol. Diagn. 2015, 15, 1187–1200. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.; Ma, J.; Zou, Z.; Jemal, A. Cancer statistics, 2014. CA Cancer J. Clin. 2014, 64, 9–29. [Google Scholar] [CrossRef] [PubMed]
- Etzioni, R.; Urban, N.; Ramsey, S.; McIntosh, M.; Schwartz, S.; Reid, B.; Radich, J.; Anderson, G.; Hartwell, L. The case for early detection. Nat. Rev. Cancer 2003, 3, 243–252. [Google Scholar] [CrossRef] [PubMed]
- Radhakrishna, S.; Gayathri, A.; Chegu, D. Needle core biopsy for breast lesions: An audit of 467 needle core biopsies. Indian J. Med. Paediatr. Oncol. 2013, 34, 252–256. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ilie, M.; Hofman, P. Pros: Can tissue biopsy be replaced by liquid biopsy? Transl. Lung Cancer Res. 2016, 5, 420–423. [Google Scholar] [CrossRef] [PubMed]
- Welch, H.G.; Black, W.C. Overdiagnosis in cancer. J. Natl. Cancer Inst. 2010, 102, 605–613. [Google Scholar] [CrossRef] [PubMed]
- Loughran, C.F.; Keeling, C.R. Seeding of tumour cells following breast biopsy: A literature review. Br. J. Radiol. 2011, 84, 869–874. [Google Scholar] [CrossRef] [PubMed]
- Yates, L.R.; Campbell, P.J. Evolution of the cancer genome. Nat. Rev. Genet. 2012, 13, 795–806. [Google Scholar] [CrossRef] [PubMed]
- Sacher, A.G.; Paweletz, C.; Dahlberg, S.E.; Alden, R.S.; O’Connell, A.; Feeney, N.; Mach, S.L.; Janne, P.A.; Oxnard, G.R. Prospective validation of rapid plasma genotyping for the detection of EGFR and KRAS mutations in advanced lung cancer. JAMA Oncol. 2016, 2, 1014–1022. [Google Scholar] [CrossRef] [PubMed]
- Hornberger, J.; Hirsch, F.R.; Li, Q.; Page, R.D. Outcome and economic implications of proteomic test-guided second- or third-line treatment for advanced non-small cell lung cancer: Extended analysis of the prose trial. Lung Cancer 2015, 88, 223–230. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, H.; Kulasinghe, A.; Perry, C.; Nelson, C.; Punyadeera, C. A liquid biopsy for head and neck cancers. Expert Rev. Mol. Diagn. 2016, 16, 165–172. [Google Scholar] [CrossRef] [PubMed]
- Gerlinger, M.; Rowan, A.J.; Horswell, S.; Math, M.; Larkin, J.; Endesfelder, D.; Gronroos, E.; Martinez, P.; Matthews, N.; Stewart, A.; et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 2012, 366, 883–892. [Google Scholar] [CrossRef] [PubMed]
- Crowley, E.; Di Nicolantonio, F.; Loupakis, F.; Bardelli, A. Liquid biopsy: Monitoring cancer-genetics in the blood. Nat. Rev. Clin. Oncol. 2013, 10, 472–484. [Google Scholar] [CrossRef] [PubMed]
- Bardelli, A.; Pantel, K. Liquid biopsies, what we do not know (yet). Cancer Cell 2017, 31, 172–179. [Google Scholar] [CrossRef] [PubMed]
- He, M.; Zeng, Y. Microfluidic exosome analysis toward liquid biopsy for cancer. J. Lab. Autom. 2016, 21, 599–608. [Google Scholar] [CrossRef] [PubMed]
- Shields, C.W.T.; Reyes, C.D.; Lopez, G.P. Microfluidic cell sorting: A review of the advances in the separation of cells from debulking to rare cell isolation. Lab Chip 2015, 15, 1230–1249. [Google Scholar] [CrossRef] [PubMed]
- Alam, M.K.; Koomson, E.; Zou, H.; Yi, C.; Li, C.-W.; Xu, T.; Yang, M. Recent advances in microfluidic technology for manipulation and analysis of biological cells (2007–2017). Anal. Chim. Acta 2018. in perss. [Google Scholar] [CrossRef]
- Sun, Y.; Haglund, T.A.; Rogers, A.J.; Ghanim, A.F.; Sethu, P. Review: Microfluidics technologies for blood-based cancer liquid biopsies. Anal. Chim. Acta 2018, 1012, 10–29. [Google Scholar] [CrossRef] [PubMed]
- Hao, N.; Zhang, J.X.J. Microfluidic screening of circulating tumor biomarkers toward liquid biopsy. Sep. Purif. Rev. 2018, 47, 19–48. [Google Scholar] [CrossRef]
- Dive, C.; Brady, G. Snapshot: Circulating tumor cells. Cell 2017, 168, 742. [Google Scholar] [CrossRef] [PubMed]
- Allard, W.J.; Matera, J.; Miller, M.C.; Repollet, M.; Connelly, M.C.; Rao, C.; Tibbe, A.G.; Uhr, J.W.; Terstappen, L.W. Tumor cells circulate in the peripheral blood of all major carcinomas but not in healthy subjects or patients with nonmalignant diseases. Clin. Cancer Res. 2004, 10, 6897–6904. [Google Scholar] [CrossRef] [PubMed]
- Pantel, K.; Alix-Panabieres, C.; Riethdorf, S. Cancer micrometastases. Nat. Rev. Clin. Oncol. 2009, 6, 339–351. [Google Scholar] [CrossRef] [PubMed]
- Yu, M.; Bardia, A.; Wittner, B.S.; Stott, S.L.; Smas, M.E.; Ting, D.T.; Isakoff, S.J.; Ciciliano, J.C.; Wells, M.N.; Shah, A.M.; et al. Circulating breast tumor cells exhibit dynamic changes in epithelial and mesenchymal composition. Science 2013, 339, 580–584. [Google Scholar] [CrossRef] [PubMed]
- Gao, D.; Vela, I.; Sboner, A.; Iaquinta, P.J.; Karthaus, W.R.; Gopalan, A.; Dowling, C.; Wanjala, J.N.; Undvall, E.A.; Arora, V.K.; et al. Organoid cultures derived from patients with advanced prostate cancer. Cell 2014, 159, 176–187. [Google Scholar] [CrossRef] [PubMed]
- Hodgkinson, C.L.; Morrow, C.J.; Li, Y.; Metcalf, R.L.; Rothwell, D.G.; Trapani, F.; Polanski, R.; Burt, D.J.; Simpson, K.L.; Morris, K.; et al. Tumorigenicity and genetic profiling of circulating tumor cells in small-cell lung cancer. Nat. Med. 2014, 20, 897–903. [Google Scholar] [CrossRef] [PubMed]
- Khoo, B.L.; Lee, S.C.; Kumar, P.; Tan, T.Z.; Warkiani, M.E.; Ow, S.G.; Nandi, S.; Lim, C.T.; Thiery, J.P. Short-term expansion of breast circulating cancer cells predicts response to anti-cancer therapy. Oncotarget 2015, 6, 15578–15593. [Google Scholar] [CrossRef] [PubMed]
- Kasimir-Bauer, S.; Hoffmann, O.; Wallwiener, D.; Kimmig, R.; Fehm, T. Expression of stem cell and epithelial-mesenchymal transition markers in primary breast cancer patients with circulating tumor cells. Breast Cancer Res. 2012, 14, R15. [Google Scholar] [CrossRef] [PubMed]
- Tomaskovic-Crook, E.; Thompson, E.W.; Thiery, J.P. Epithelial to mesenchymal transition and breast cancer. Breast Cancer Res. 2009, 11, 213. [Google Scholar] [CrossRef] [PubMed]
- Thompson, E.W.; Nagaraj, S.H. Transition states that allow cancer to spread. Nature 2018, 556, 442–444. [Google Scholar] [CrossRef] [PubMed]
- Rao, C.G.; Chianese, D.; Doyle, G.V.; Miller, M.C.; Russell, T.; Sanders, R.A., Jr.; Terstappen, L.W. Expression of epithelial cell adhesion molecule in carcinoma cells present in blood and primary and metastatic tumors. Int. J. Oncol. 2005, 27, 49–57. [Google Scholar] [CrossRef] [PubMed]
- Joosse, S.A.; Pantel, K. Biologic challenges in the detection of circulating tumor cells. Cancer Res. 2013, 73, 8–11. [Google Scholar] [CrossRef] [PubMed]
- Barriere, G.; Fici, P.; Gallerani, G.; Fabbri, F.; Zoli, W.; Rigaud, M. Circulating tumor cells and epithelial, mesenchymal and stemness markers: Characterization of cell subpopulations. Ann. Transl. Med. 2014, 2, 109. [Google Scholar] [CrossRef] [PubMed]
- Deneve, E.; Riethdorf, S.; Ramos, J.; Nocca, D.; Coffy, A.; Daures, J.P.; Maudelonde, T.; Fabre, J.M.; Pantel, K.; Alix-Panabieres, C. Capture of viable circulating tumor cells in the liver of colorectal cancer patients. Clin. Chem. 2013, 59, 1384–1392. [Google Scholar] [CrossRef] [PubMed]
- Hou, H.W.; Warkiani, M.E.; Khoo, B.L.; Li, Z.R.; Soo, R.A.; Tan, D.S.; Lim, W.T.; Han, J.; Bhagat, A.A.; Lim, C.T. Isolation and retrieval of circulating tumor cells using centrifugal forces. Sci. Rep. 2013, 3, 1259. [Google Scholar] [CrossRef] [PubMed]
- Karabacak, N.M.; Spuhler, P.S.; Fachin, F.; Lim, E.J.; Pai, V.; Ozkumur, E.; Martel, J.M.; Kojic, N.; Smith, K.; Chen, P.I.; et al. Microfluidic, marker-free isolation of circulating tumor cells from blood samples. Nat. Protoc. 2014, 9, 694–710. [Google Scholar] [CrossRef] [PubMed]
- Kulasinghe, A.; Perry, C.; Warkiani, M.E.; Blick, T.; Davies, A.; O’Byrne, K.; Thompson, E.W.; Nelson, C.C.; Vela, I.; Punyadeera, C. Short term ex-vivo expansion of circulating head and neck tumour cells. Oncotarget 2016, 7, 60101–60109. [Google Scholar] [CrossRef] [PubMed]
- Khoo, B.L.; Warkiani, M.E.; Tan, D.S.; Bhagat, A.A.; Irwin, D.; Lau, D.P.; Lim, A.S.; Lim, K.H.; Krisna, S.S.; Lim, W.T.; et al. Clinical validation of an ultra high-throughput spiral microfluidics for the detection and enrichment of viable circulating tumor cells. PLoS ONE 2014, 9, e99409. [Google Scholar] [CrossRef] [PubMed]
- Ferris, R.L.; Blumenschein, G., Jr.; Fayette, J.; Guigay, J.; Colevas, A.D.; Licitra, L.; Harrington, K.; Kasper, S.; Vokes, E.E.; Even, C.; et al. Nivolumab for recurrent squamous-cell carcinoma of the head and neck. N. Engl. J. Med. 2016, 375, 1856–1867. [Google Scholar] [CrossRef] [PubMed]
- Hellmann, M.D.; Ciuleanu, T.E.; Pluzanski, A.; Lee, J.S.; Otterson, G.A.; Audigier-Valette, C.; Minenza, E.; Linardou, H.; Burgers, S.; Salman, P.; et al. Nivolumab plus ipilimumab in lung cancer with a high tumor mutational burden. N. Engl. J. Med. 2018, 378, 2093–2104. [Google Scholar] [CrossRef] [PubMed]
- Wolchok, J.D.; Chiarion-Sileni, V.; Gonzalez, R.; Rutkowski, P.; Grob, J.J.; Cowey, C.L.; Lao, C.D.; Wagstaff, J.; Schadendorf, D.; Ferrucci, P.F.; et al. Overall survival with combined nivolumab and ipilimumab in advanced melanoma. N. Engl. J. Med. 2017, 377, 1345–1356. [Google Scholar] [CrossRef] [PubMed]
- Strati, A.; Koutsodontis, G.; Papaxoinis, G.; Angelidis, I.; Zavridou, M.; Economopoulou, P.; Kotsantis, I.; Avgeris, M.; Mazel, M.; Perisanidis, C.; et al. Prognostic significance of PD-L1 expression on circulating tumor cells in patients with head and neck squamous cell carcinoma. Ann. Oncol. 2017, 28, 1923–1933. [Google Scholar] [CrossRef] [PubMed]
- Kulasinghe, A.; Perry, C.; Kenny, L.; Warkiani, M.E.; Nelson, C.; Punyadeera, C. Pd-L1 expressing circulating tumour cells in head and neck cancers. BMC Cancer 2017, 17, 333. [Google Scholar] [CrossRef] [PubMed]
- Dhar, M.; Wong, J.; Che, J.; Matsumoto, M.; Grogan, T.; Elashoff, D.; Garon, E.B.; Goldman, J.W.; Sollier Christen, E.; Di Carlo, D.; et al. Evaluation of PD-L1 expression on vortex-isolated circulating tumor cells in metastatic lung cancer. Sci. Rep. 2018, 8, 2592. [Google Scholar] [CrossRef] [PubMed]
- Kulasinghe, A.; Kenny, L.; Punyadeera, C. Circulating tumour cell PD-L1 test for head and neck cancers. Oral Oncol. 2017, 75, 6–7. [Google Scholar] [CrossRef] [PubMed]
- Kallergi, G.; Vetsika, E.K.; Aggouraki, D.; Lagoudaki, E.; Koutsopoulos, A.; Koinis, F.; Katsarlinos, P.; Trypaki, M.; Messaritakis, I.; Stournaras, C.; et al. Evaluation of PD-L1/PD-1 on circulating tumor cells in patients with advanced non-small cell lung cancer. Ther. Adv. Med. Oncol. 2018, 10. [Google Scholar] [CrossRef] [PubMed]
- Jia, S.; Zhang, R.; Li, Z.; Li, J. Clinical and biological significance of circulating tumor cells, circulating tumor DNA, and exosomes as biomarkers in colorectal cancer. Oncotarget 2017, 8, 55632–55645. [Google Scholar] [CrossRef] [PubMed]
- Kalluri, R. The biology and function of exosomes in cancer. J. Clin. Investig. 2016, 126, 1208–1215. [Google Scholar] [CrossRef] [PubMed]
- Melo, S.A.; Sugimoto, H.; O’Connell, J.T.; Kato, N.; Villanueva, A.; Vidal, A.; Qiu, L.; Vitkin, E.; Perelman, L.T.; Melo, C.A.; et al. Cancer exosomes perform cell-independent microrna biogenesis and promote tumorigenesis. Cancer Cell 2014, 26, 707–721. [Google Scholar] [CrossRef] [PubMed]
- Costa-Silva, B.; Aiello, N.M.; Ocean, A.J.; Singh, S.; Zhang, H.; Thakur, B.K.; Becker, A.; Hoshino, A.; Mark, M.T.; Molina, H.; et al. Pancreatic cancer exosomes initiate pre-metastatic niche formation in the liver. Nat. Cell Biol. 2015, 17, 816–826. [Google Scholar] [CrossRef] [PubMed]
- Zomer, A.; Maynard, C.; Verweij, F.J.; Kamermans, A.; Schafer, R.; Beerling, E.; Schiffelers, R.M.; de Wit, E.; Berenguer, J.; Ellenbroek, S.I.J.; et al. In vivo imaging reveals extracellular vesicle-mediated phenocopying of metastatic behavior. Cell 2015, 161, 1046–1057. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Yang, Y.; Zeng, Y.; He, M. A microfluidic exosearch chip for multiplexed exosome detection towards blood-based ovarian cancer diagnosis. Lab Chip 2016, 16, 489–496. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Skog, J.; Hsu, C.H.; Lessard, R.T.; Balaj, L.; Wurdinger, T.; Carter, B.S.; Breakefield, X.O.; Toner, M.; Irimia, D. Microfluidic isolation and transcriptome analysis of serum microvesicles. Lab Chip 2010, 10, 505–511. [Google Scholar] [CrossRef] [PubMed]
- Dudani, J.S.; Gossett, D.R.; Tse, H.T.; Lamm, R.J.; Kulkarni, R.P.; Carlo, D.D. Rapid inertial solution exchange for enrichment and flow cytometric detection of microvesicles. Biomicrofluidics 2015, 9, 014112. [Google Scholar] [CrossRef] [PubMed]
- Han, K.H.; Han, A.; Frazier, A.B. Microsystems for isolation and electrophysiological analysis of breast cancer cells from blood. Biosens. Bioelectron. 2006, 21, 1907–1914. [Google Scholar] [CrossRef] [PubMed]
- Bettegowda, C.; Sausen, M.; Leary, R.J.; Kinde, I.; Wang, Y.; Agrawal, N.; Bartlett, B.R.; Wang, H.; Luber, B.; Alani, R.M.; et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci. Transl. Med. 2014, 6, 224ra224. [Google Scholar] [CrossRef] [PubMed]
- Bellassai, N.; Spoto, G. Biosensors for liquid biopsy: Circulating nucleic acids to diagnose and treat cancer. Anal. Bioanal. Chem. 2016, 408, 7255–7264. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.J.; Brock, M.V.; Shih Ie, M.; Wang, T.H. Decoding circulating nucleic acids in human serum using microfluidic single molecule spectroscopy. J. Am. Chem. Soc. 2010, 132, 5793–5798. [Google Scholar] [CrossRef] [PubMed]
- Toepke, M.W.; Beebe, D.J. PDMS absorption of small molecules and consequences in microfluidic applications. Lab Chip 2006, 6, 1484–1486. [Google Scholar] [CrossRef] [PubMed]
- Whitesides, G.M. The origins and the future of microfluidics. Nature 2006, 442, 368–373. [Google Scholar] [CrossRef] [PubMed]
- Sequist, L.V.; Nagrath, S.; Toner, M.; Haber, D.A.; Lynch, T.J. The CTC-chip: An exciting new tool to detect circulating tumor cells in lung cancer patients. J. Thorac. Oncol. 2009, 4, 281–283. [Google Scholar] [CrossRef] [PubMed]
- Yoon, H.J.; Shanker, A.; Wang, Y.; Kozminsky, M.; Jin, Q.; Palanisamy, N.; Burness, M.L.; Azizi, E.; Simeone, D.M.; Wicha, M.S.; et al. Tunable thermal-sensitive polymer-graphene oxide composite for efficient capture and release of viable circulating tumor cells. Adv. Mater. 2016, 28, 4891–4897. [Google Scholar] [CrossRef] [PubMed]
- Murlidhar, V.; Zeinali, M.; Grabauskiene, S.; Ghannad-Rezaie, M.; Wicha, M.S.; Simeone, D.M.; Ramnath, N.; Reddy, R.M.; Nagrath, S. A radial flow microfluidic device for ultra-high-throughput affinity-based isolation of circulating tumor cells. Small 2014, 10, 4895–4904. [Google Scholar] [CrossRef] [PubMed]
- Jan, Y.J.; Chen, J.F.; Zhu, Y.; Lu, Y.T.; Chen, S.H.; Chung, H.; Smalley, M.; Huang, Y.W.; Dong, J.; Chen, L.C.; et al. Nanovelcro rare-cell assays for detection and characterization of circulating tumor cells. Adv. Drug Deliv. Rev. 2018, 125, 78–93. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Lu, Y.T.; Li, F.; Wu, K.; Hou, S.; Yu, J.; Shen, Q.; Wu, D.; Song, M.; OuYang, W.H.; et al. High-purity prostate circulating tumor cell isolation by a polymer nanofiber-embedded microchip for whole exome sequencing. Adv. Mater. 2013, 25, 2897–2902. [Google Scholar] [CrossRef] [PubMed]
- Ke, Z.; Lin, M.; Chen, J.F.; Choi, J.S.; Zhang, Y.; Fong, A.; Liang, A.J.; Chen, S.F.; Li, Q.; Fang, W.; et al. Programming thermoresponsiveness of nanovelcro substrates enables effective purification of circulating tumor cells in lung cancer patients. ACS Nano 2015, 9, 62–70. [Google Scholar] [CrossRef] [PubMed]
- Mandair, D.; Vesely, C.; Ensell, L.; Lowe, H.; Spanswick, V.; Hartley, J.A.; Caplin, M.E.; Meyer, T. A comparison of cellcollector with cellsearch in patients with neuroendocrine tumours. Endocr. Relat. Cancer 2016, 23, L29–L32. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Theil, G.; Fischer, K.; Weber, E.; Medek, R.; Hoda, R.; Lucke, K.; Fornara, P. The use of a new cellcollector to isolate circulating tumor cells from the blood of patients with different stages of prostate cancer and clinical outcomes—A proof-of-concept study. PLoS ONE 2016, 11, e0158354. [Google Scholar] [CrossRef] [PubMed]
- Saucedo-Zeni, N.; Mewes, S.; Niestroj, R.; Gasiorowski, L.; Murawa, D.; Nowaczyk, P.; Tomasi, T.; Weber, E.; Dworacki, G.; Morgenthaler, N.G.; et al. A novel method for the in vivo isolation of circulating tumor cells from peripheral blood of cancer patients using a functionalized and structured medical wire. Int. J. Oncol. 2012, 41, 1241–1250. [Google Scholar] [PubMed]
- Hoshino, K.; Huang, Y.Y.; Lane, N.; Huebschman, M.; Uhr, J.W.; Frenkel, E.P.; Zhang, X. Microchip-based immunomagnetic detection of circulating tumor cells. Lab Chip 2011, 11, 3449–3457. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.Y.; Hoshino, K.; Chen, P.; Wu, C.H.; Lane, N.; Huebschman, M.; Liu, H.; Sokolov, K.; Uhr, J.W.; Frenkel, E.P.; et al. Immunomagnetic nanoscreening of circulating tumor cells with a motion controlled microfluidic system. Biomed. Microdevices 2013, 15, 673–681. [Google Scholar] [CrossRef] [PubMed]
- Ozkumur, E.; Shah, A.M.; Ciciliano, J.C.; Emmink, B.L.; Miyamoto, D.T.; Brachtel, E.; Yu, M.; Chen, P.I.; Morgan, B.; Trautwein, J.; et al. Inertial focusing for tumor antigen-dependent and -independent sorting of rare circulating tumor cells. Sci. Transl. Med. 2013, 5, 179ra147. [Google Scholar] [CrossRef] [PubMed]
- Adams, D.L.; Stefansson, S.; Haudenschild, C.; Martin, S.S.; Charpentier, M.; Chumsri, S.; Cristofanilli, M.; Tang, C.M.; Alpaugh, R.K. Cytometric characterization of circulating tumor cells captured by microfiltration and their correlation to the cellsearch® CTC test. Cytometry A 2015, 87, 137–144. [Google Scholar] [CrossRef] [PubMed]
- Raimondi, C.; Gradilone, A.; Naso, G.; Cortesi, E.; Gazzaniga, P. Clinical utility of circulating tumor cell counting through cellsearch®: The dilemma of a concept suspended in limbo. Onco Targets Ther. 2014, 7, 619–625. [Google Scholar] [PubMed]
- Bidard, F.C.; Proudhon, C.; Pierga, J.Y. Circulating tumor cells in breast cancer. Mol. Oncol. 2016, 10, 418–430. [Google Scholar] [CrossRef] [PubMed]
- Galletti, G.; Portella, L.; Tagawa, S.T.; Kirby, B.J.; Giannakakou, P.; Nanus, D.M. Circulating tumor cells in prostate cancer diagnosis and monitoring: An appraisal of clinical potential. Mol. Diagn. Ther. 2014, 18, 389–402. [Google Scholar] [CrossRef] [PubMed]
- Hardingham, J.E.; Grover, P.; Winter, M.; Hewett, P.J.; Price, T.J.; Thierry, B. Detection and clinical significance of circulating tumor cells in colorectal cancer—20 years of progress. Mol. Med. 2015, 21 (Suppl. 1), S25–S31. [Google Scholar]
- Alva, A.; Friedlander, T.; Clark, M.; Huebner, T.; Daignault, S.; Hussain, M.; Lee, C.; Hafez, K.; Hollenbeck, B.; Weizer, A.; et al. Circulating tumor cells as potential biomarkers in bladder cancer. J. Urol. 2015, 194, 790–798. [Google Scholar] [CrossRef] [PubMed]
- Harb, W.; Fan, A.; Tran, T.; Danila, D.C.; Keys, D.; Schwartz, M.; Ionescu-Zanetti, C. Mutational analysis of circulating tumor cells using a novel microfluidic collection device and QPCR assay. Transl. Oncol. 2013, 6, 528–538. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Lorencio, M.I.; Ramirez, P.; Saenz, L.; Martinez Sanchez, M.V.; De La Orden, V.; Mediero-Valeros, B.; Veganzones-De-Castro, S.; Baroja-Mazo, A.; Revilla Nuin, B.; Gonzalez, M.R.; et al. Comparison of two types of liquid biopsies in patients with hepatocellular carcinoma awaiting orthotopic liver transplantation. Transplant. Proc. 2015, 47, 2639–2642. [Google Scholar] [CrossRef] [PubMed]
- de Bruin, E.C.; McGranahan, N.; Mitter, R.; Salm, M.; Wedge, D.C.; Yates, L.; Jamal-Hanjani, M.; Shafi, S.; Murugaesu, N.; Rowan, A.J.; et al. Spatial and temporal diversity in genomic instability processes defines lung cancer evolution. Science 2014, 346, 251–256. [Google Scholar] [CrossRef] [PubMed]
- Cahall, C.F.; Lilly, J.L.; Hirschowitz, E.A.; Berron, B.J. A quantitative perspective on surface marker selection for the isolation of functional tumor cells. Breast Cancer 2015, 9, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.S.; Sim, T.S.; Kim, Y.J.; Kim, S.S.; Jeong, H.; Park, J.M.; Moon, H.S.; Kim, S.I.; Gurel, O.; Lee, S.S.; et al. SSA-MOA: A novel CTC isolation platform using selective size amplification (SSA) and a multi-obstacle architecture (MOA) filter. Lab Chip 2012, 12, 2874–2880. [Google Scholar] [CrossRef] [PubMed]
- Chung, J.; Shao, H.; Reiner, T.; Issadore, D.; Weissleder, R.; Lee, H. Microfluidic cell sorter (MUFCS) for on-chip capture and analysis of single cells. Adv. Healthc. Mater. 2012, 1, 432–436. [Google Scholar] [CrossRef] [PubMed]
- Coumans, F.A.; van Dalum, G.; Beck, M.; Terstappen, L.W. Filtration parameters influencing circulating tumor cell enrichment from whole blood. PLoS ONE 2013, 8, e61774. [Google Scholar] [CrossRef] [PubMed]
- Au, S.H.; Edd, J.; Stoddard, A.E.; Wong, K.H.K.; Fachin, F.; Maheswaran, S.; Haber, D.A.; Stott, S.L.; Kapur, R.; Toner, M. Microfluidic isolation of circulating tumor cell clusters by size and asymmetry. Sci. Rep. 2017, 7, 2433. [Google Scholar] [CrossRef] [PubMed]
- Andersson, E.; Steven, K.; Guldberg, P. Size-based enrichment of exfoliated tumor cells in urine increases the sensitivity for DNA-based detection of bladder cancer. PLoS ONE 2014, 9, e94023. [Google Scholar] [CrossRef] [PubMed]
- Shen, Z.; Wu, A.; Chen, X. Current detection technologies for circulating tumor cells. Chem. Soc. Rev. 2017, 46, 2038–2056. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, R.; Gertler, R.; Friederichs, J.; Fuehrer, K.; Dahm, M.; Phelps, R.; Thorban, S.; Nekarda, H.; Siewert, J.R. Comparison of two density gradient centrifugation systems for the enrichment of disseminated tumor cells in blood. Cytometry 2002, 49, 150–158. [Google Scholar] [CrossRef] [PubMed]
- Sollier, E.; Go, D.E.; Che, J.; Gossett, D.R.; O’Byrne, S.; Weaver, W.M.; Kummer, N.; Rettig, M.; Goldman, J.; Nickols, N.; et al. Size-selective collection of circulating tumor cells using vortex technology. Lab Chip 2014, 14, 63–77. [Google Scholar] [CrossRef] [PubMed]
- Warkiani, M.E.; Khoo, B.L.; Wu, L.; Tay, A.K.; Bhagat, A.A.; Han, J.; Lim, C.T. Ultra-fast, label-free isolation of circulating tumor cells from blood using spiral microfluidics. Nat. Protoc. 2016, 11, 134–148. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Mao, Z.; Peng, Z.; Zhou, L.; Chen, Y.; Huang, P.H.; Truica, C.I.; Drabick, J.J.; El-Deiry, W.S.; Dao, M.; et al. Acoustic separation of circulating tumor cells. Proc. Natl. Acad. Sci. USA 2015, 112, 4970–4975. [Google Scholar] [CrossRef] [PubMed]
- Cushing, K.; Undvall, E.; Ceder, Y.; Lilja, H.; Laurell, T. Reducing wbc background in cancer cell separation products by negative acoustic contrast particle immuno-acoustophoresis. Anal. Chim. Acta 2018, 1000, 256–264. [Google Scholar] [CrossRef] [PubMed]
- Shields Iv, C.W.; Wang, J.L.; Ohiri, K.A.; Essoyan, E.D.; Yellen, B.B.; Armstrong, A.J.; Lopez, G.P. Magnetic separation of acoustically focused cancer cells from blood for magnetographic templating and analysis. Lab Chip 2016, 16, 3833–3844. [Google Scholar] [CrossRef] [PubMed]
- Warkiani, M.E.; Guan, G.; Luan, K.B.; Lee, W.C.; Bhagat, A.A.; Chaudhuri, P.K.; Tan, D.S.; Lim, W.T.; Lee, S.C.; Chen, P.C.; et al. Slanted spiral microfluidics for the ultra-fast, label-free isolation of circulating tumor cells. Lab Chip 2014, 14, 128–137. [Google Scholar] [CrossRef] [PubMed]
- Che, J.; Yu, V.; Dhar, M.; Renier, C.; Matsumoto, M.; Heirich, K.; Garon, E.B.; Goldman, J.; Rao, J.; Sledge, G.W.; et al. Classification of large circulating tumor cells isolated with ultra-high throughput microfluidic vortex technology. Oncotarget 2016, 7, 12748–12760. [Google Scholar] [CrossRef] [PubMed]
- Gascoyne, P.R.; Shim, S. Isolation of circulating tumor cells by dielectrophoresis. Cancers 2014, 6, 545–579. [Google Scholar] [CrossRef] [PubMed]
- Liga, A.; Vliegenthart, A.D.; Oosthuyzen, W.; Dear, J.W.; Kersaudy-Kerhoas, M. Exosome isolation: A microfluidic road-map. Lab Chip 2015, 15, 2388–2394. [Google Scholar] [CrossRef] [PubMed]
- Im, H.; Shao, H.; Park, Y.I.; Peterson, V.M.; Castro, C.M.; Weissleder, R.; Lee, H. Label-free detection and molecular profiling of exosomes with a nano-plasmonic sensor. Nat. Biotechnol. 2014, 32, 490–495. [Google Scholar] [CrossRef] [PubMed]
- He, M.; Crow, J.; Roth, M.; Zeng, Y.; Godwin, A.K. Integrated immunoisolation and protein analysis of circulating exosomes using microfluidic technology. Lab Chip 2014, 14, 3773–3780. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Wu, H.J.; Fine, D.; Schmulen, J.; Hu, Y.; Godin, B.; Zhang, J.X.; Liu, X. Ciliated micropillars for the microfluidic-based isolation of nanoscale lipid vesicles. Lab Chip 2013, 13, 2879–2882. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.; Shao, H.; Weissleder, R.; Lee, H. Acoustic purification of extracellular microvesicles. ACS Nano 2015, 9, 2321–2327. [Google Scholar] [CrossRef] [PubMed]
- Yeo, J.C.; Kenry; Zhao, Z.; Zhang, P.; Wang, Z.; Lim, C.T. Label-free extraction of extracellular vesicles using centrifugal microfluidics. Biomicrofluidics 2018, 12, 024103. [Google Scholar] [CrossRef]
- Ebrahimi Warkiani, M.; Lou, C.-P.; Gong, H.-Q. Fabrication of multi-layer polymeric micro-sieve having narrow slot pores with conventional ultraviolet-lithography and micro-fabrication techniques. Biomicrofluidics 2011, 5, 036504. [Google Scholar] [CrossRef] [PubMed]
- Warkiani, M.E.; Lou, C.-P.; Liu, H.-B.; Gong, H.-Q. A high-flux isopore micro-fabricated membrane for effective concentration and recovering of waterborne pathogens. Biomed. Microdevices 2012, 14, 669–677. [Google Scholar] [CrossRef] [PubMed]
- Davies, R.T.; Kim, J.; Jang, S.C.; Choi, E.J.; Gho, Y.S.; Park, J. Microfluidic filtration system to isolate extracellular vesicles from blood. Lab Chip 2012, 12, 5202–5210. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.; Jo, W.; Heo, Y.; Kang, J.Y.; Kwak, R.; Park, J. Isolation of extracellular vesicle from blood plasma using electrophoretic migration through porous membrane. Sens. Actuators B Chem. 2016, 233, 289–297. [Google Scholar] [CrossRef]
- Contreras-Naranjo, J.C.; Wu, H.J.; Ugaz, V.M. Microfluidics for exosome isolation and analysis: Enabling liquid biopsy for personalized medicine. Lab Chip 2017, 17, 3558–3577. [Google Scholar] [CrossRef] [PubMed]
- Dawson, S.J.; Tsui, D.W.; Murtaza, M.; Biggs, H.; Rueda, O.M.; Chin, S.F.; Dunning, M.J.; Gale, D.; Forshew, T.; Mahler-Araujo, B.; et al. Analysis of circulating tumor DNA to monitor metastatic breast cancer. N. Engl. J. Med. 2013, 368, 1199–1209. [Google Scholar] [CrossRef] [PubMed]
- Lo, Y.M.; Corbetta, N.; Chamberlain, P.F.; Rai, V.; Sargent, I.L.; Redman, C.W.; Wainscoat, J.S. Presence of fetal DNA in maternal plasma and serum. Lancet 1997, 350, 485–487. [Google Scholar] [CrossRef]
- Reinholt, S.J.; Baeumner, A.J. Microfluidic isolation of nucleic acids. Angew. Chem. Int. Ed. Engl. 2014, 53, 13988–14001. [Google Scholar] [CrossRef] [PubMed]
- Burns, M.A.; Johnson, B.N.; Brahmasandra, S.N.; Handique, K.; Webster, J.R.; Krishnan, M.; Sammarco, T.S.; Man, P.M.; Jones, D.; Heldsinger, D.; et al. An integrated nanoliter DNA analysis device. Science 1998, 282, 484–487. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Zhou, J.; Sutherland, A.; Wei, W.; Shin, Y.S.; Xue, M.; Heath, J.R. Microfluidics-based single-cell functional proteomics for fundamental and applied biomedical applications. Annu. Rev. Anal. Chem. 2014, 7, 275–295. [Google Scholar] [CrossRef] [PubMed]
- Lagally, E.T.; Medintz, I.; Mathies, R.A. Single-molecule DNA amplification and analysis in an integrated microfluidic device. Anal. Chem. 2001, 73, 565–570. [Google Scholar] [CrossRef] [PubMed]
- Diehl, F.; Li, M.; Dressman, D.; He, Y.; Shen, D.; Szabo, S.; Diaz, L.A., Jr.; Goodman, S.N.; David, K.A.; Juhl, H.; et al. Detection and quantification of mutations in the plasma of patients with colorectal tumors. Proc. Natl. Acad. Sci. USA 2005, 102, 16368–16373. [Google Scholar] [CrossRef] [PubMed]
- Volik, S.; Alcaide, M.; Morin, R.D.; Collins, C. Cell-free DNA (cfDNA): Clinical significance and utility in cancer shaped by emerging technologies. Mol. Cancer Res. 2016, 14, 898–908. [Google Scholar] [CrossRef] [PubMed]
- Adriani, G.; Pavesi, A.; Tan, A.T.; Bertoletti, A.; Thiery, J.P.; Kamm, R.D. Microfluidic models for adoptive cell-mediated cancer immunotherapies. Drug Discov. Today 2016, 21, 1472–1478. [Google Scholar] [CrossRef] [PubMed]
- Bancroft, D. Draper Looks to Ultrasound and Geometry to Reduce Car-t Manufacturing Costs. Available online: https://www.biopharma-reporter.com/Article/2017/02/07/Draper-invents-CAR-T-manufacturing-solution-Ultrasonic-biochips (accessed on 29 July 2018).
- Warkiani, M.E.; Wu, L.; Tay, A.K.; Han, J. Large-volume microfluidic cell sorting for biomedical applications. Annu. Rev. Biomed. Eng. 2015, 17, 1–34. [Google Scholar] [CrossRef] [PubMed]
- FDA Grants Breakthrough Device Designation to Foundation Medicine Liquid Biopsy Assay. Available online: https://www.genomeweb.com/molecular-diagnostics/fda-grants-breakthrough-device-designation-foundation-medicine-liquid-biopsy#.W17gv5_-iUk (accessed on 29 July 2018).
- Cobas EGFR Mutation Test v2. Available online: https://www.fda.gov/drugs/informationondrugs/approveddrugs/ucm504540.htm (accessed on 29 July 2018).
- Myriad Receives FDA Approval of Bracanalysis CDx® as Companion Diagnostic for Lynparza™ (olaparib) in Patients with Metastatic Breast Cancer. Available online: http://investor.myriad.com/news-releases/news-release-details/myriad-receives-fda-approval-bracanalysis-cdxr-companion (accessed on 29 July 2018).
| Types of Liquid Biopsy Samples | Circulating Tumor Cells | ctDNA | Exosomes |
|---|---|---|---|
| Sample Sources | Peripheral Blood | Plasma or Serum | Plasma or Other Body Fluids |
| Separation or detection methods | Positive selection by antibody coated microstructure and microparticles | Digital PCR, next generation sequencing, allele-specific PCR | Antibody coated microchannel and microparticles |
| Negative selection by antibody coated microstructure and microparticles | Label-free methods: Size based nanowire capture, acoustic nanofilter | ||
| Label-free methods: size-based filtration, density-based separation, hydrodynamic, acoustic, magnetophoresis, de-electrophoretic | |||
| Strength | ● High specificity of tumor-derived | ● High sensitivity | ● Samples available in various body fluids |
| ● Approved by FDA clinical practice | ● Ongoing clinical trials for treatment suggestions | ● DNA, RNA, and protein could all be investigated | |
| ● DNA, RNA, and protein could all be investigated | ● Functional studies available | ||
| ● Functional studies available | |||
| Limitations | ● Loss of heterogeneity on non-label free isolation methods | ● No RNA or protein could be investigated | ● Loss of heterogeneity on non-label free isolation methods |
| ● Instability of tumor diagnosis at early stage | ● Tumor specific mutations are hard to distinguish due to redundant background noise signals | ● Absence of specific tumor derived markers |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kulasinghe, A.; Wu, H.; Punyadeera, C.; Warkiani, M.E. The Use of Microfluidic Technology for Cancer Applications and Liquid Biopsy. Micromachines 2018, 9, 397. https://doi.org/10.3390/mi9080397
Kulasinghe A, Wu H, Punyadeera C, Warkiani ME. The Use of Microfluidic Technology for Cancer Applications and Liquid Biopsy. Micromachines. 2018; 9(8):397. https://doi.org/10.3390/mi9080397
Chicago/Turabian StyleKulasinghe, Arutha, Hanjie Wu, Chamindie Punyadeera, and Majid Ebrahimi Warkiani. 2018. "The Use of Microfluidic Technology for Cancer Applications and Liquid Biopsy" Micromachines 9, no. 8: 397. https://doi.org/10.3390/mi9080397
APA StyleKulasinghe, A., Wu, H., Punyadeera, C., & Warkiani, M. E. (2018). The Use of Microfluidic Technology for Cancer Applications and Liquid Biopsy. Micromachines, 9(8), 397. https://doi.org/10.3390/mi9080397







