Elapid Snake Venom Analyses Show the Specificity of the Peptide Composition at the Level of Genera Naja and Notechis
Abstract
:1. Introduction
2. Results
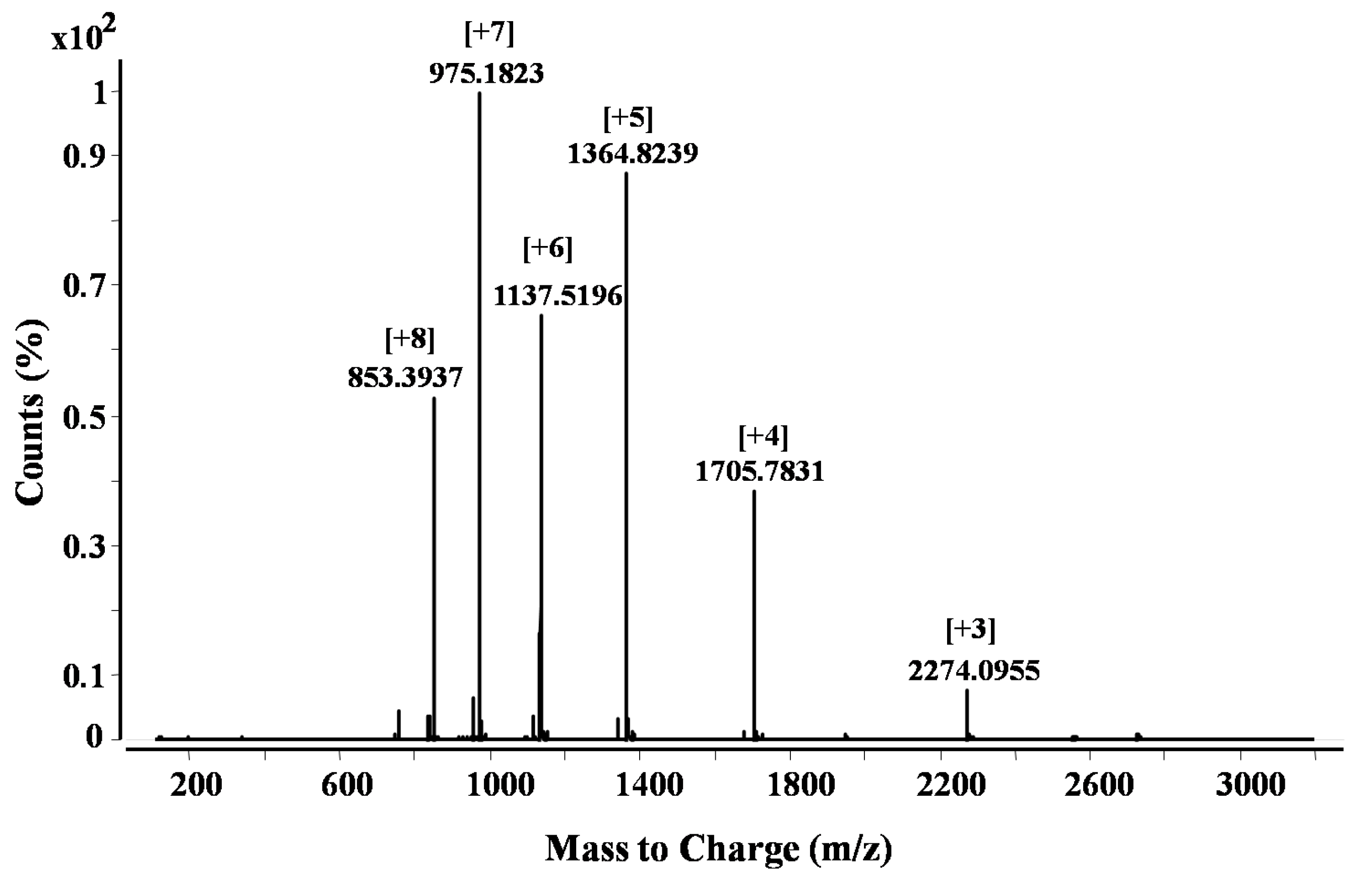
2.1. Purification and Identification of Peptides from N. m. mossambica Venom
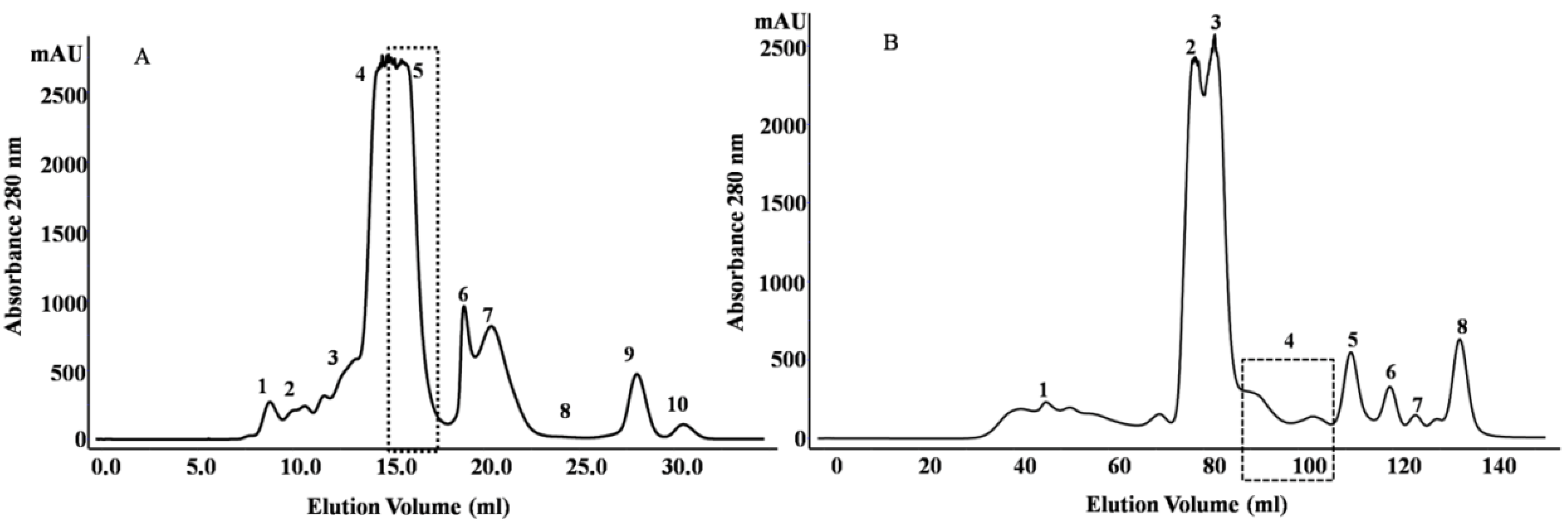
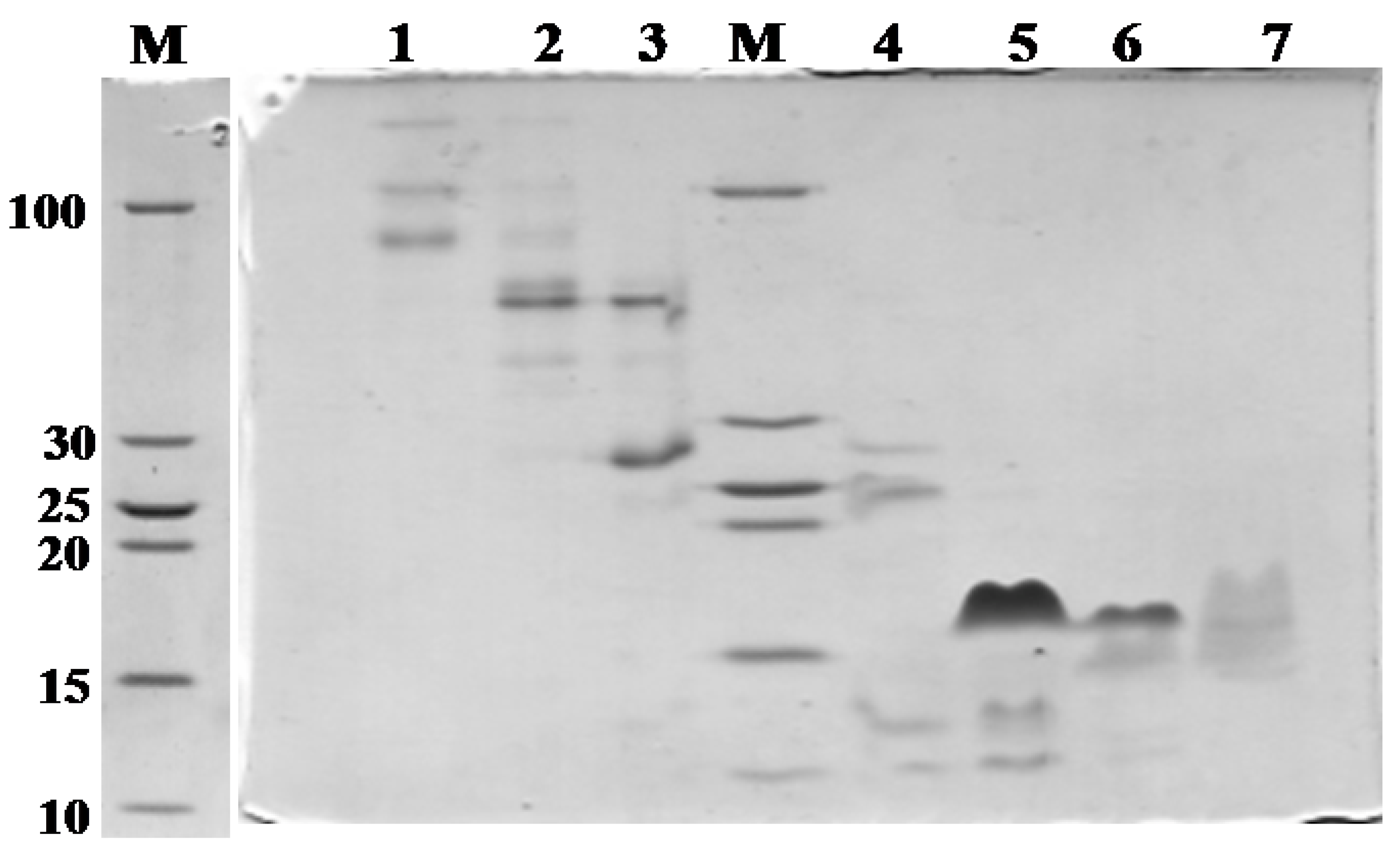
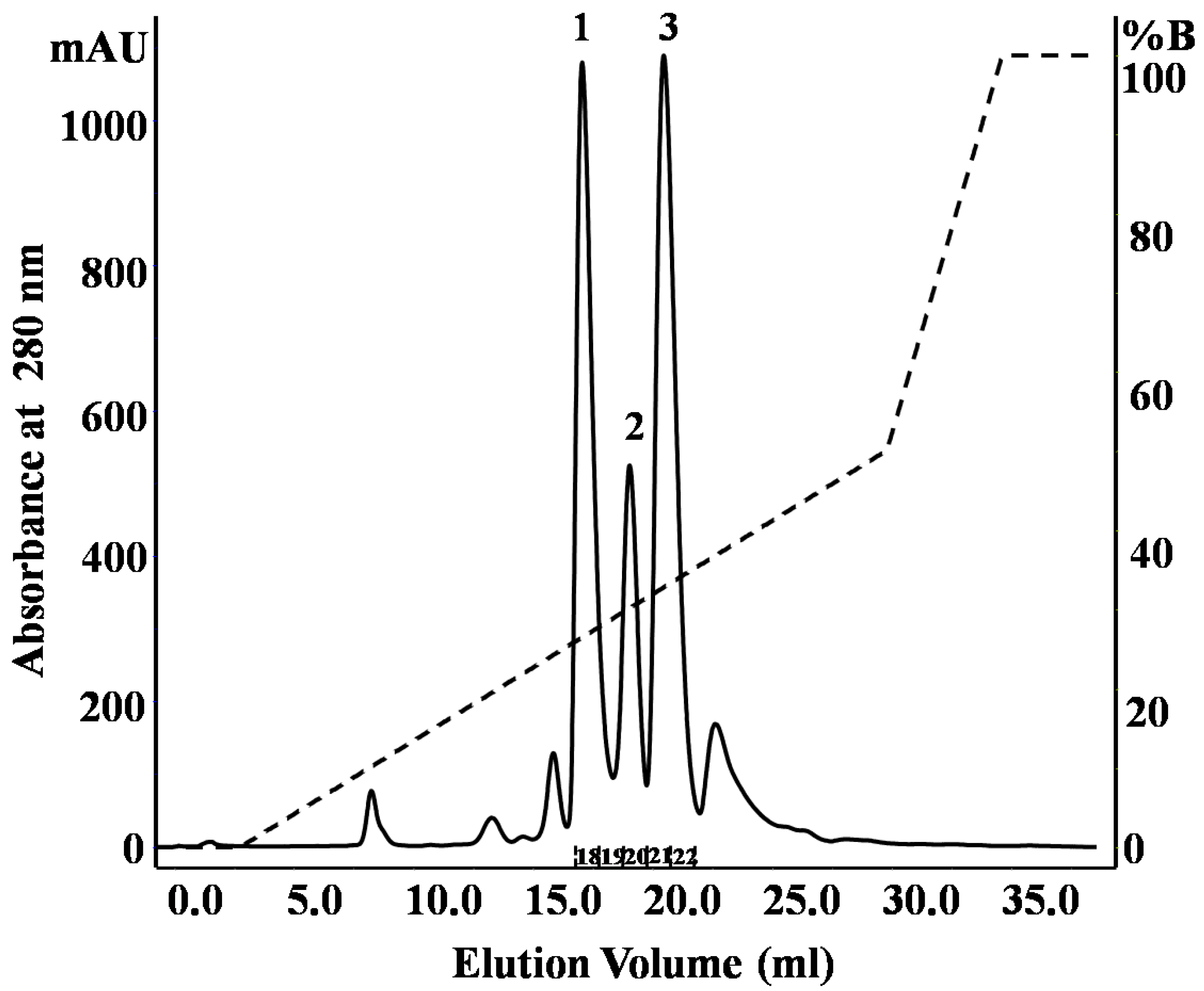
| Fraction number (SEC) | Observed m/z | Score/sequence coverage | Sequence | Inhibitory activity | Homology with peptide from | Peptide family |
|---|---|---|---|---|---|---|
| 5 | 975.1823 (M + H)7+ | 274/70% | LKCNQLIPPFWKTCPKGKNLCYKMTMRAAPMVPVKRGCIDVCPKSSLLIKYMCCNTNKCN | Bacterial subtilisin (StmPr1), chymotrypsin and 20S proteasome | P01467: Naja mossambica | Snake venom three-finger toxin (CTX M1) |
| 5, 6 | 6896.40 (M + H)+ | 71/48% | LKCNRLIPPFWKTCPEGKNLCYKMTMRLAPKVPVKRGCIDVCPKSSLLIKYMCCNTNKCN | P01470: Naja mossambica | Snake venom three-finger toxin (CTX M3) | |
| 5 | 6727.402 (M + H)+ | 119/93% | LKCNKLIPIAYKTCPEGKNLCYKMMLASKKMVPVKRGCINVCPKNSALVKYVCCSTDRCN | P01452: Naja mossambica | Snake venom three-finger toxin (CTX M4) | |
| 8 | 873.4511 (M + H)+ | 32 | ZQKFSPR | P85314: Bothrops moojeni | Zinc metalloproteinase | |
| 8 | 901.4614 (M + H)+ | 30 | ZQRFSPR | Q072L5: Bothrops asper | Zinc metalloproteinase/disintegrin | |
| 8 | 779.3983 (M + H)+ | 22 | ZXWPRP | ACE | P0C7R6: Agkistrodon piscivorus piscivorus | BPP |
| 10 | 1214.6581 (M + H)+ | 51 | ZXWPRPQXPP | ACE | P0C7S6: Crotalus atrox | BPP |
| 10 | 1276.6380 (M + H)+ | 45 | ZQWPPGHHXPP | ACE | P0C7J9: Crotalus adamanteus | BPP |
| 10 | 1063.5611 (M + H)+ | 20 | TPPAGPDVGPR | Q27J49: Lachesis muta muta | Bradykinin inhibitor peptide | |
| 10 | 1230.6743 (M + H)+ | 18 | QXWPRPQXPP | ACE | P0C7S6: Crotalus atrox | BPP |
| 10 | 1277.6380 (M + H)+ | 26 | ZEWPPGHHXPP | ACE | Q27J49: Lachesis muta muta | BPP |
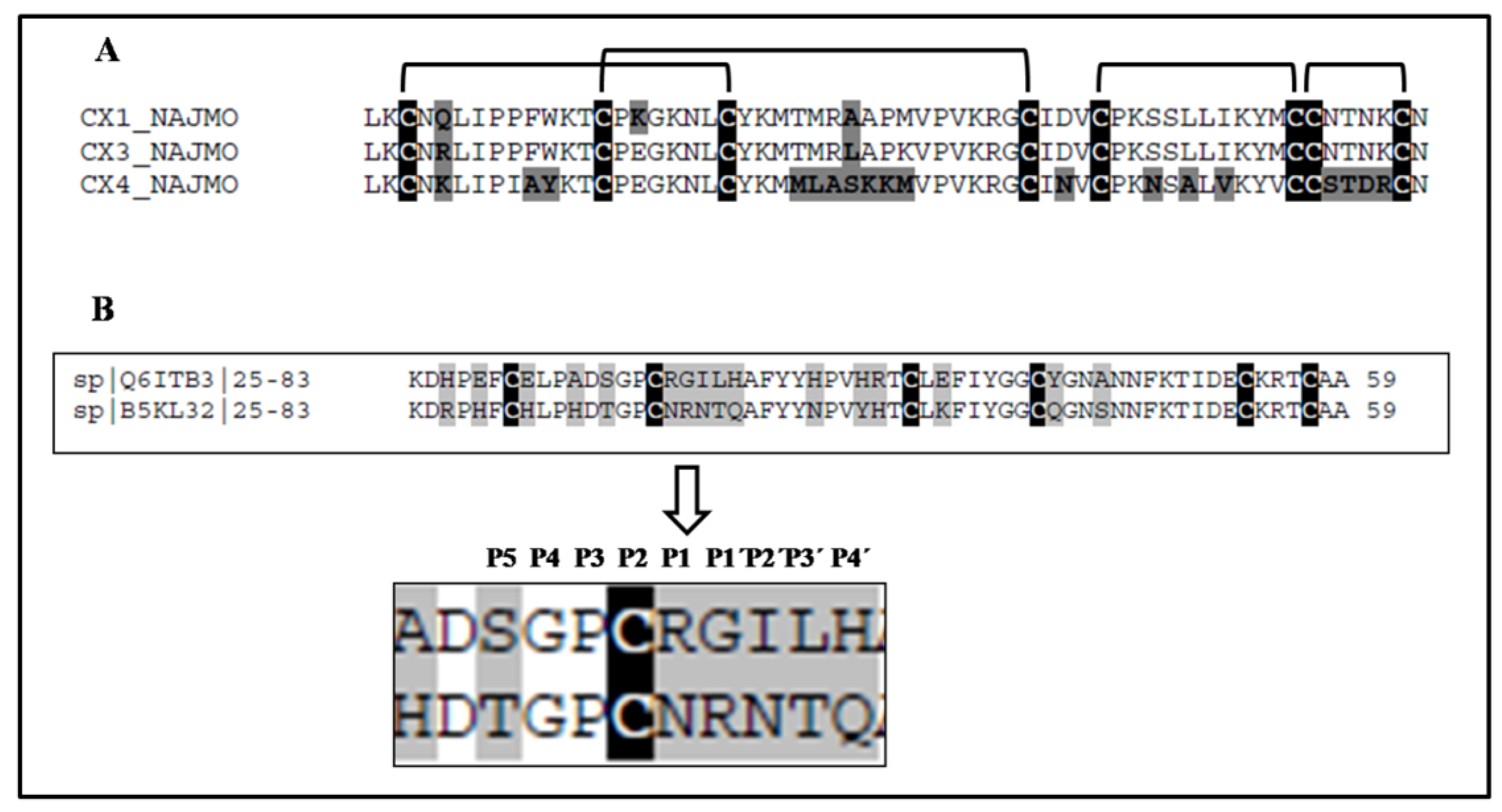
2.2. Purification and Identification of Peptides from N. scutatus (Kangaroo Island) Venom
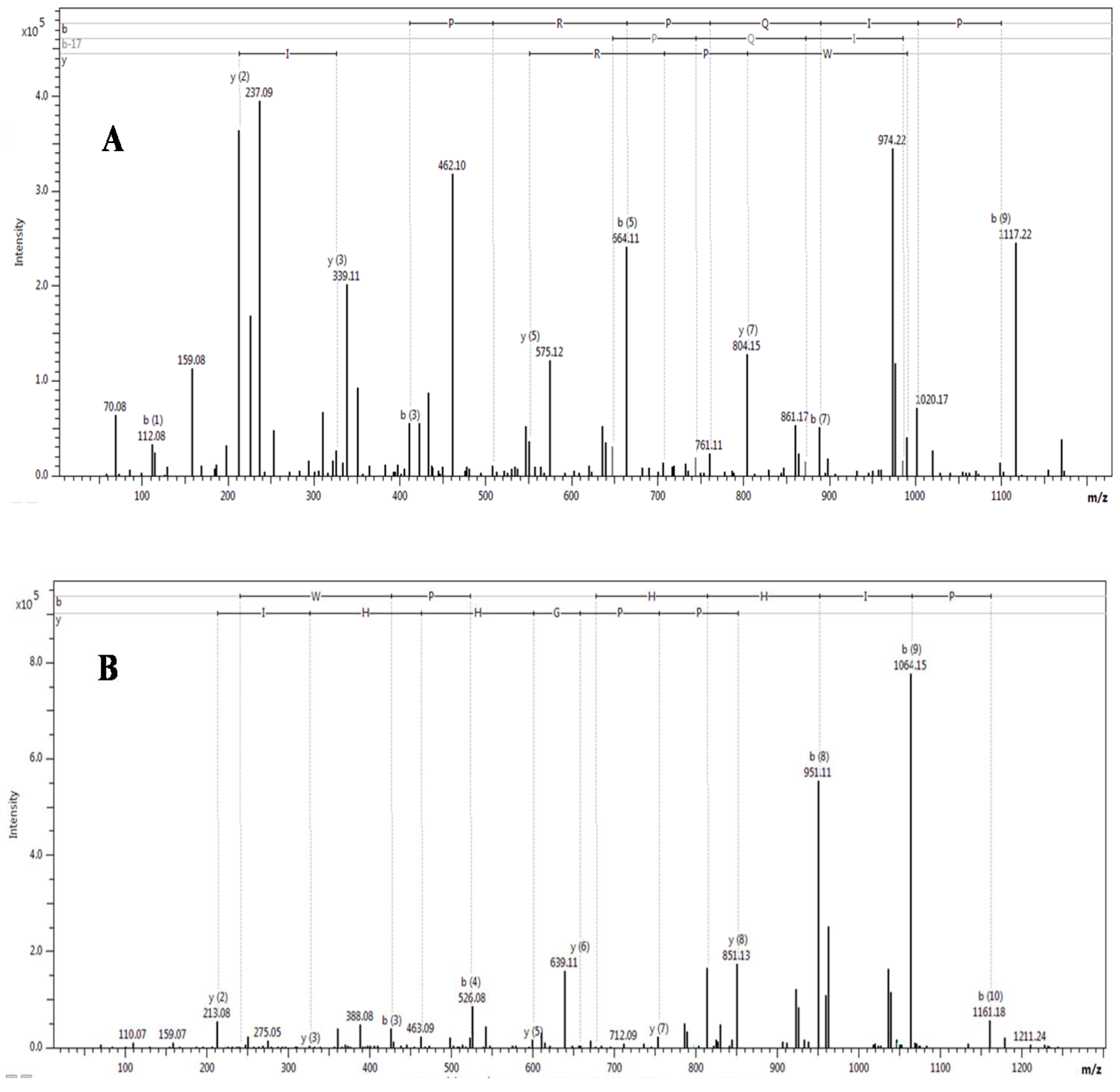
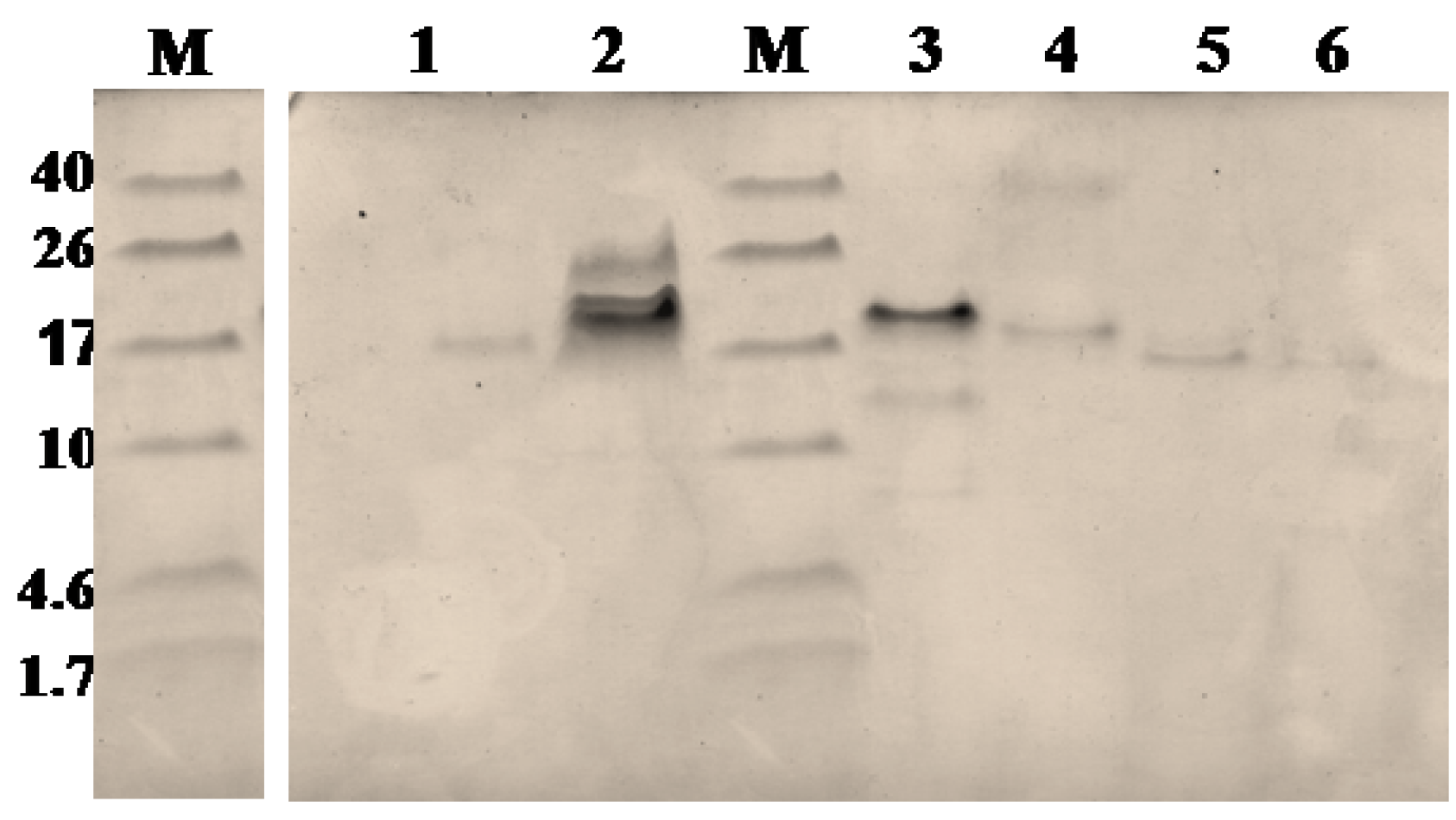
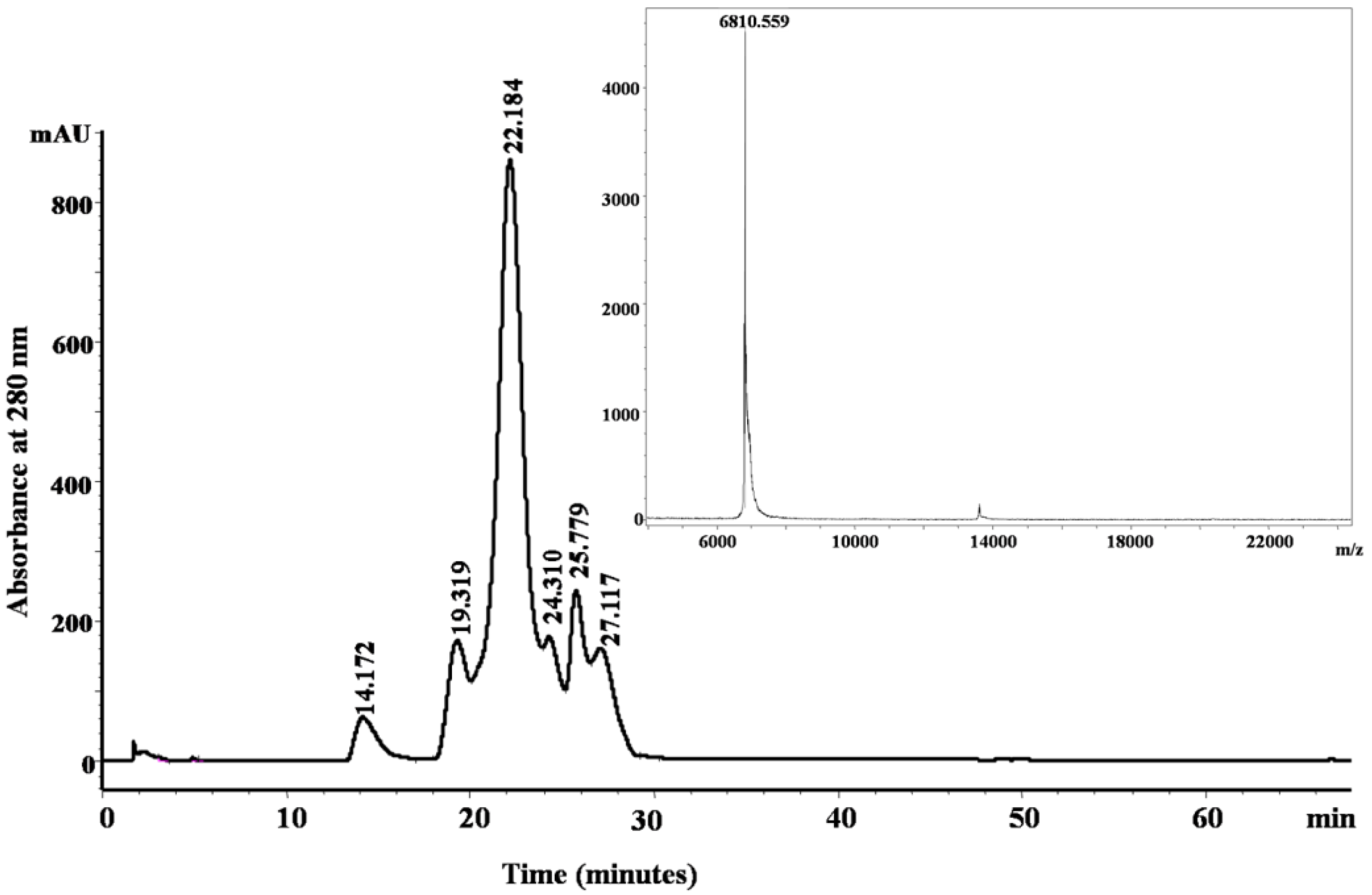
| Observed m/z (M + H)+ | Score/sequence coverage | Sequence determined | Inhibitory activity | Homology with peptide from | Peptide family |
|---|---|---|---|---|---|
| 6647.921* | 185/44% | MTCCNQQSSQPKTTTTCAESSCYKKTWRDHRGTITERGCGCPNVKPGVQINCCKTDECNN | A8HDK0: Notechis scutatus scutatus | Snake venom three-finger toxin | |
| 6687.729 | 121/27% | MTCCNQQSSQPKTTTTCAESSCYKKTWRDHRGTIIERGCGCPNVKPGIQLVCCETNECNN | A8S6A4: Austrelaps superbus | Snake venom three-finger toxin | |
| 6810.559 | 1569/65% | KDRPHFCHLPHDTGPCNRNTQAFYYNPVYHTCLKFIYGGCQGNSNNFKTIDECKRTCAA | Trypsin | B5KL32: Notechis scutatus scutatus | Kunitz/BPTI |
| 6694.233 | 231/45% | KDHPEFCELPADSGPCRGILHAFYYHPVHRTCLEFIYGGCYGNANNFKTIDECKRTCAA | Trypsin, plasma kallikrein, plasmin | Q6ITB3: Notechis scutatus scutatus | Kunitz/BPTI |
| 3347.688 | 54/30% | SGSEVAKIGDGCFGLPLDRIGSASGMGCRSVPKP | ACE | Q3SAE8: Notechis scutatus scutatus | NP |
| 1154.556 | QNPPKPXSGES | Q3SAX8: Oxyuranus scutellatus scutellatus | NP | ||
| 764.201 (M + H)2+ | Glycopeptide | ||||
| 681.251 | Glycopeptide |
3. Discussion
4. Experimental Section
4.1. Material
4.2. Fractionation of the Crude Venoms and Purification of Peptides
4.3. Enzyme Inhibition Assays
4.4. Mass Spectrometric Analyses
5. Conclusions
Acknowledgments
Abbreviations
| ACE | angiotensin-converting enzyme; |
| CTX | cardiotoxin; |
| BPP | bradykinin-potentiating peptide; |
| NP | natriuretic peptide; |
| 3-F | three-finger peptide |
Conflicts of Interest
References
- Fry, B.G.; Wuster, W.; Kini, R.M.; Brusic, V.; Khan, A.; Venkataraman, D.; Rooney, A.P. Molecular evolution and phylogeny of elapid snake venom three-finger toxins. J. Mol. Evol. 2003, 57, 110–129. [Google Scholar] [CrossRef]
- Earl, S.T.; Masci, P.P.; de Jersey, J.; Lavin, M.F.; Dixon, J. Drug development from Australian elapid snake venoms and the Venomics pipeline of candidates for haemostasis: Textilinin-1 (Q8008), Haempatch (Q8009) and CoVase (V0801). Toxicon 2012, 59, 456–463. [Google Scholar] [CrossRef]
- Millers, E.K.; Johnson, L.A.; Birrell, G.W.; Masci, P.P.; Lavin, M.F.; de Jersey, J.; Guddat, L.W. The structure of human microplasmin in complex withTextilinin-1, an Aprotinin-like inhibitor from the Australian brown snake. PLoS ONE 2013, 8, e54104. [Google Scholar]
- Cushman, D.W.; Ondetti, M.A. Design of angiotensin converting enzyme inhibitors. Nat. Med. 1999, 5, 1110–1113. [Google Scholar] [CrossRef]
- Akif, M.; Georgiadis, D.; Mahajan, A.; Dive, V.; Sturrock, E.D.; Isaac, R.E.; Acharya, K.R. High-resolution crystal structures of Drosophila melanogaster angiotensin-converting enzyme in complex with novel inhibitors and antihypertensive drugs. J. Mol. Biol. 2010, 400, 502–517. [Google Scholar] [CrossRef]
- King, G.F. Venoms as a platform for human drugs: Translating toxins into therapeutics. Expert. Opin. Biol. Th. 2011, 11, 1469–1484. [Google Scholar]
- Steckelings, U.M.; Artuc, M.; Wollschlager, T.; Wiehstutz, S.; Henz, B.M. Angiotensin-converting enzyme inhibitors as inducers of adverse cutaneous reactions. Acta Derm. Venereol. 2001, 81, 321–325. [Google Scholar] [CrossRef]
- Masuyer, G.; Schwager, S.L.; Sturrock, E.D.; Isaac, R.E.; Acharya, K.R. Molecular recognition and regulation of human angiotensin-I converting enzyme (ACE) activity by natural inhibitory peptides. Sci. Rep. 2012, 2, ep00717. [Google Scholar]
- Fry, B.G. Structure-function properties of venom components from Australian elapids. Toxicon 1999, 37, 11–32. [Google Scholar] [CrossRef]
- St Pierre, L.; Earl, S.T.; Filippovich, I.; Sorokina, N.; Masci, P.P.; De Jersey, J.; Lavin, M.F. Common evolution of waprin and kunitz-like toxin families in Australian venomous snakes. Cell. Mol. Life Sci. 2008, 65, 4039–4054. [Google Scholar] [CrossRef]
- Dubovskii, P.V.; Lesovoy, D.M.; Dubinnyi, M.A.; Utkin, Y.N.; Arseniev, A.S. Interaction of the P-type cardiotoxin with phospholipid membranes. Eur. J. Biochem. 2003, 270, 2038–2046. [Google Scholar] [CrossRef]
- St Pierre, L.; Fischer, H.; Adams, D.J.; Schenning, M.; Lavidis, N.; de Jersey, J.; Masci, P.P.; Lavin, M.F. Distinct activities of novel neurotoxins from Australian venomous snakes for nicotinic acetylcholine receptors. Cell. Mol. Life Sci. 2007, 64, 2829–2840. [Google Scholar] [CrossRef]
- Syka, J.E.; Coon, J.J.; Schroeder, M.J.; Shabanowitz, J.; Hunt, D.F. Peptide and protein sequence analysis by electron transfer dissociation mass spectrometry. Proc. Natl. Acad. Sci. USA 2004, 101, 9528–9533. [Google Scholar]
- Quinton, L.; Gilles, N.; Smargiasso, N.; Kiehne, A.; de Pauw, E. An unusual family of glycosylated peptides isolated from Dendroaspis angusticeps venom and characterized by combination of collision induced and electron transfer dissociation. J. Am. Soc. Mass Spectrom. 2011, 22, 1891–1897. [Google Scholar] [CrossRef]
- Calvete, J.J.; Ghezellou, P.; Paiva, O.; Matainaho, T.; Ghassempour, A.; Goudarzi, H.; Kraus, F.; Sanz, L.; Williams, D.J. Snake venomics of two poorly known Hydrophiinae: Comparative proteomics of the venoms of terrestrial Toxicocalamus longissimus and marine Hydrophis cyanocinctus. J. Proteomics 2012, 75, 4091–4101. [Google Scholar] [CrossRef]
- Gregoire, J.; Rochat, H. Amino acid sequences of neurotoxins I and III of the elapid snake Naja mossambica mossambica. Eur. J. Biochem. 1977, 80, 283–293. [Google Scholar] [CrossRef]
- Georgieva, D.; Ohler, M.; Seifert, J.; von Bergen, M.; Arni, R.K.; Genov, N.; Betzel, C. Snake venomic of Crotalus durissus terrificus--correlation with pharmacological activities. J. Proteome Res. 2010, 9, 2302–2316. [Google Scholar] [CrossRef]
- Petras, D.; Sanz, L.; Segura, A.; Herrera, M.; Villalta, M.; Solano, D.; Vargas, M.; Leon, G.; Warrell, D.A.; Theakston, R.D.; et al. Snake venomics of African spitting cobras: Toxin composition and assessment of congeneric cross-reactivity of the pan-African EchiTAb-Plus-ICP antivenom by antivenomics and neutralization approaches. J. Proteome Res. 2011, 10, 1266–1280. [Google Scholar] [CrossRef]
- Rees, B.; Samama, J.P.; Thierry, J.C.; Gilibert, M.; Fischer, J.; Schweitz, H.; Lazdunski, M.; Moras, D. Crystal structure of a snake venom cardiotoxin. Proc. Natl. Acad. Sci. USA 1987, 84, 3132–3136. [Google Scholar] [CrossRef]
- Chien, K.Y.; Chiang, C.M.; Hseu, Y.C.; Vyas, A.A.; Rule, G.S.; Wu, W. Two distinct types of cardiotoxin as revealed by the structure and activity relationship of their interaction with zwitterionic phospholipid dispersions. J. Biol. Chem. 1994, 269, 14473–14483. [Google Scholar]
- Fry, B.G.; Lumsden, N.G.; Wuster, W.; Wickramaratna, J.C.; Hodgson, W.C.; Kini, R.M. Isolation of a neurotoxin (alpha-colubritoxin) from a nonvenomous colubrid: Evidence for early origin of venom in snakes. J. Mol. Evol. 2003, 57, 446–452. [Google Scholar] [CrossRef]
- Kini, R.M. Molecular moulds with multiple missions: Functional sites in three-finger toxins. Clin. Exp. Pharmacol. Physiol. 2002, 29, 815–822. [Google Scholar] [CrossRef]
- Bilwes, A.; Rees, B.; Moras, D.; Ménez, R.; Ménez, A. X-ray structure at 1.55 A of toxin gamma, a cardiotoxin from Naja nigricollis venom. Crystal packing reveals a model for insertion into membranes. J. Mol. Biol. 1994, 239, 122–136. [Google Scholar] [CrossRef]
- O’Connell, J.F.; Bougis, P.E.; Wüthrich, K. Determination of the nuclear-magnetic-resonance solution structure of cardiotoxin CTX IIb from Naja mossambica mossambica. Eur. J. Biochem. 1993, 213, 891–900. [Google Scholar] [CrossRef]
- Nam, S.; Smith, D.M.; Dou, Q.P. Ester bond-containing tea polyphenols potently inhibit proteasome activity in vitro and in vivo. J. Biol. Chem. 2001, 276, 13322–13330. [Google Scholar] [CrossRef]
- Peters, J.M.; Franke, W.W.; Kleinschmidt, J.A. Distinct 19 S and 20 S subcomplexes of the 26 S proteasome and their distribution in the nucleus and the cytoplasm. J. Biol. Chem. 1994, 269, 7709–7718. [Google Scholar]
- Wang, J.; Maldonado, M.A. The ubiquitin-proteasome system and its role in inflammatory and autoimmune diseases. Cell. Mol. Immunol. 2006, 3, 255–261. [Google Scholar]
- Cron, K.R.; Zhu, K.; Kushwaha, D.S.; Hsieh, G.; Merzon, D.; Rameseder, J.; Chen, C.C.; D’Andrea, A.D.; Kozono, D. Proteasome inhibitors block DNA repair and radiosensitize non-small cell lung cancer. PLoS ONE 2013, 8, e73710. [Google Scholar] [CrossRef]
- Almond, J.B.; Cohen, G.M. The proteasome: a novel target for cancer chemotherapy. Leukemia 2002, 16, 433–443. [Google Scholar] [CrossRef]
- Lin, K.L.; Su, J.C.; Chien, C.M.; Chuang, P.W.; Chang, L.S.; Lin, S.R. Down-regulation of the JAK2/PI3K-mediated signaling activation is involved in Taiwan cobra cardiotoxin III-induced apoptosis of human breast MDA-MB-231 cancer cells. Toxicon 2010, 55, 1263–1273. [Google Scholar] [CrossRef]
- Fry, B.G.; Wuster, W. Assembling an arsenal: Origin and evolution of the snake venom proteome inferred from phylogenetic analysis of toxin sequences. Mol. Biol. Evol. 2004, 21, 870–883. [Google Scholar] [CrossRef]
- Munawar, A.; Trusch, M.; Georgieva, D.; Spencer, P.; Frochaux, V.; Harder, S.; Arni, R.K.; Duhalov, D.; Genov, N.; Schluter, H.; et al. Venom peptide analysis of Vipera ammodytes meridionalis (Viperinae) and Bothrops jararacussu (Crotalinae) demonstrates subfamily-specificity of the peptidome in the family Viperidae. Mol. Biosyst. 2011, 7, 3298–3307. [Google Scholar] [CrossRef]
- El-Saadani, M.A.; El-Sayed, M.F. A bradykinin potentiating peptide from Egyptian cobra venom strongly affects rat atrium contractile force and cellular calcium regulation. Compa. Biochem. Physiol. C Toxicol. Pharmacol. 2003, 136, 387–395. [Google Scholar] [CrossRef]
- Vyas, V.K.; Brahmbhatt, K.; Bhatt, H.; Parmar, U.; Patidar, R. Therapeutic potential of snake venom in cancer therapy: Current perspectives. Asian Pac. J. Trop. Biomed. 2013, 3, 156–162. [Google Scholar] [CrossRef]
- St Pierre, L.; Flight, S.; Masci, P.P.; Hanchard, K.J.; Lewis, R.J.; Alewood, P.F.; de Jersey, J.; Lavin, M.F. Cloning and characterisation of natriuretic peptides from the venom glands of Australian elapids. Biochimie 2006, 88, 1923–1931. [Google Scholar] [CrossRef]
- Hodgson, W.C.; Isbister, G.K. The application of toxins and venoms to cardiovascular drug discovery. Curr. Opin. Pharmacol. 2009, 9, 173–176. [Google Scholar] [CrossRef]
- Johns, D.G.; Ao, Z.; Heidrich, B.J.; Hunsberger, G.E.; Graham, T.; Payne, L.; Elshourbagy, N.; Lu, Q.; Aiyar, N.; Douglas, S.A. Dendroaspis natriuretic peptide binds to the natriuretic peptide clearance receptor. Biochem. Biophys. Res. Commun. 2007, 358, 145–149. [Google Scholar] [CrossRef]
- Zupunski, V.; Kordis, D.; Gubensek, F. Adaptive evolution in the snake venom Kunitz/BPTI protein family. FEBS Letters 2003, 547, 131–136. [Google Scholar] [CrossRef]
- Helland, R.; Leiros, I.; Berglund, G.I.; Willassen, N.P.; Smalas, A.O. The crystal structure of anionic salmon trypsin in complex with bovine pancreatic trypsin inhibitor. Eur. J. Biochem. 1998, 256, 317–324. [Google Scholar]
- Scheidig, A.J.; Hynes, T.R.; Pelletier, L.A.; Wells, J.A.; Kossiakoff, A.A. Crystal structures of bovine chymotrypsin and trypsin complexed to the inhibitor domain of Alzheimer’s amyloid beta-protein precursor (APPI) and basic pancreatic trypsin inhibitor (BPTI): Engineering of inhibitors with altered specificities. Protein Sci. 1997, 6, 1806–1824. [Google Scholar]
- Millers, E.K.; Trabi, M.; Masci, P.P.; Lavin, M.F.; de Jersey, J.; Guddat, L.W. Crystal structure of textilinin-1, a Kunitz-type serine protease inhibitor from the venom of the Australian common brown snake (Pseudonaja textilis). Febs. J. 2009, 276, 3163–3175. [Google Scholar] [CrossRef]
- Ritonja, A.; Meloun, B.; Gubensek, F. The primary structure of Vipera ammodytes venom trypsin inhibitor I. Biochim. Biophys. Acta 1983, 748, 429–435. [Google Scholar] [CrossRef]
- Tang, J.; Yu, C.L.; Williams, S.R.; Springman, E.; Jeffery, D.; Sprengeler, P.A.; Estevez, A.; Sampang, J.; Shrader, W.; Spencer, J.; et al. Expression, crystallization, and three-dimensional structure of the catalytic domain of human plasma kallikrein. J. Biol. Chem. 2005, 280, 41077–41089. [Google Scholar] [CrossRef]
- Carmona, A.K.; Schwager, S.L.; Juliano, M.A.; Juliano, L.; Sturrock, E.D. A continuous fluorescence resonance energy transfer angiotensin I-converting enzyme assay. Nat. Protoc. 2006, 1, 1971–1976. [Google Scholar] [CrossRef]
- Perkins, D.N.; Pappin, D.J.; Creasy, D.M.; Cottrell, J.S. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 1999, 20, 3551–3567. [Google Scholar] [CrossRef]
- Boeckmann, B.; Bairoch, A.; Apweiler, R.; Blatter, M.C.; Estreicher, A.; Gasteiger, E.; Martin, M.J.; Michoud, K.; O’Donovan, C.; Phan, I.; et al. The SWISS-PROT protein knowledgebase and its supplement TrEMBL in 2003. Nucleic Acids Res. 2003, 31, 365–370. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Munawar, A.; Trusch, M.; Georgieva, D.; Hildebrand, D.; Kwiatkowski, M.; Behnken, H.; Harder, S.; Arni, R.; Spencer, P.; Schlüter, H.; et al. Elapid Snake Venom Analyses Show the Specificity of the Peptide Composition at the Level of Genera Naja and Notechis. Toxins 2014, 6, 850-868. https://doi.org/10.3390/toxins6030850
Munawar A, Trusch M, Georgieva D, Hildebrand D, Kwiatkowski M, Behnken H, Harder S, Arni R, Spencer P, Schlüter H, et al. Elapid Snake Venom Analyses Show the Specificity of the Peptide Composition at the Level of Genera Naja and Notechis. Toxins. 2014; 6(3):850-868. https://doi.org/10.3390/toxins6030850
Chicago/Turabian StyleMunawar, Aisha, Maria Trusch, Dessislava Georgieva, Diana Hildebrand, Marcel Kwiatkowski, Henning Behnken, Sönke Harder, Raghuvir Arni, Patrick Spencer, Hartmut Schlüter, and et al. 2014. "Elapid Snake Venom Analyses Show the Specificity of the Peptide Composition at the Level of Genera Naja and Notechis" Toxins 6, no. 3: 850-868. https://doi.org/10.3390/toxins6030850
APA StyleMunawar, A., Trusch, M., Georgieva, D., Hildebrand, D., Kwiatkowski, M., Behnken, H., Harder, S., Arni, R., Spencer, P., Schlüter, H., & Betzel, C. (2014). Elapid Snake Venom Analyses Show the Specificity of the Peptide Composition at the Level of Genera Naja and Notechis. Toxins, 6(3), 850-868. https://doi.org/10.3390/toxins6030850








