Abstract
Aflatoxins (AFs) are toxic secondary metabolites produced mostly by Aspergillus species. AF contamination entering the feed and food chain has been a crucial long-term issue for veterinarians, medicals, agroindustry experts, and researchers working in this field. Although different (physical, chemical, and biological) technologies have been developed, tested, and employed to mitigate the detrimental effects of mycotoxins, including AFs, universal methods are still not available to reduce AF levels in feed and food in the last decades. Possible biological control by bacteria, yeasts, and fungi, their excretes, the role of the ruminal degradation, pre-harvest biocontrol by competitive exclusion or biofungicides, and post-harvest technologies and practices based on biological agents currently used to alleviate the toxic effects of AFs are collected in this review. Pre-harvest biocontrol technologies can give us the greatest opportunity to reduce AF production on the spot. Together with post-harvest applications of bacteria or fungal cultures, these technologies can help us strictly reduce AF contamination without synthetic chemicals.
Key Contribution:
Elimination of aflatoxins from commodities is possible only in field applications or under unique microbial processes. The new and promising technologies need to be made available by the authorities to fight against contamination.
1. Introduction
Aflatoxins (AFs) are furanocoumarin derivative mycotoxins biosynthesized by Aspergillus species, among which Aspergillus flavus, A. parasiticus, A. nomiae, and A. pseudotamarii are regarded as primary producers [1]. AFs can contaminate various products (e.g., cereals, oilseeds, nuts, spices, fruits, dried fruits, and milk) [2]. Regarding the background of the AFs production, the regulation of the AF gene clusters is still studied intensively [3]. The ecology of the AF-producer fungi is remarkably complex, and most likely, interactions of the producer fungal species with host plants and the soil micro- and macrobiota should be considered in detail to reach a deeper understanding of the reasons for AF production [4].
AFs are among the most dangerous compounds affecting the physiological processes of animals and humans disadvantageously. AFs are mutagenic, teratogenic, genotoxic, and carcinogenic under long-term exposure [5,6,7]. AFs may enter the feed and food chain at any time point from pre-harvest to human consumption [6,8,9]. These toxins are typically absorbed from the gut in both animals and humans and transferred into different body parts, where they can be chemically bonded or modified. Pathological dysfunction of the liver and kidneys, the gastrointestinal tract, and the immune and reproductive systems have been reported in both humans and livestock under AF exposition [5,6]. AF derivatives like aflatoxin M1 (AFM1) are eventually excreted in animals and cause contamination of milk and milk products [10,11,12,13]. At the same time, AFM1 can be produced in fungal contamination [14]. It may disturb the early development of embryos after getting through the placenta [6,15]. In breastfeeding, AFM1 is threatening even human newborns [12,13]. The direct contamination of fermented milk products like cheese by AF-producer molds and their metabolites has also been described in many cases [16].
Researchers work actively to prevent mycotoxin formation and carry-over because the dangerous effects of AFs on livestock and human health cannot be underestimated. Prevention methods and protocols set into operation pre- and postharvest, for example, good agricultural and manufacturing practices (like deep plowing and grain sorting) and appropriate storage conditions (cold, dry environment), are regarded as the best choices to reduce the AF contamination in feed and food. However, these are not always possible [7].
AFs do not decompose quickly because of their remarkable stability [17,18]. Therefore, preventing the growth of AF-producer fungi and detoxification of contaminated feeds and foods is critical. Several chemical (e.g., fungicides), physical (e.g., radiation), and biological detoxification methods were investigated and applied [19]. The chemical and physical methods may lead to nutrient reduction and sensory property changes, and mount food safety problems [20,21]. The biological prevention methods take advantage of the adverse effects of selected microorganisms, including bacteria, yeasts, and nontoxigenic molds, on the growth and AF production of toxigenic fungal strains. The adverse effects of these techniques are based on space and nutrient competitions (competitive exclusion), or biological interactions like antibiosis. The microorganism-based biological control technologies can be reasonable solutions for controlling and reducing pre- and post-harvest AF contamination in crops and food products. Notably, such biological control technologies have been commercialized and are available on the market [22,23,24].
In this review, special attention is paid to reducing AFs by biological methods in feed and food pre- and post-harvest and emphasizing promising novel and innovative approaches and technologies.
2. Document Analysis
The aim was to gain information on AF and Aspergillus mitigations work with any microbial or biological agents. Therefore, different online libraries (PubMed, Google Scholar, Science Direct, and Mendeley) were searched for the following terms and phrases: “aflatoxin degradation”, “aflatoxin binding”, “aflatoxin AND ruminal degradation”, “non-aflatoxigenic Aspergillus”, “atoxigenic Aspergillus”, “aflatoxin AND biocontrol”, “Aspergillus AND biofungicide”, and “competitive exclusion AND flavus”. The findings were ordered as ruminal, pre-, and post-harvest processes to see all natural and biotechnological possibilities and attempts. Investigations using viable cells in milk or phosphate-buffered saline as a medium were only considered for a clearer comparison of the effects on AF binding or degradation.
3. Ruminal Detoxification of Aflatoxins
It was common to feed ruminants with the fodder of AFs contamination because it was known that ruminal microorganisms could detoxify mycotoxins. However, the scientific literature is still diverse about AFs’ fate in ruminal animals. In 1978, Engel and Hagemeister [25] reported 42% degradation of aflatoxin B1 (AFB1) in vitro applying rumen fluid. The effectiveness of the degradation depended on the animal species and the fodder [26]. The resistance of AFs to ruminal degradation may be caused by the strong inhibitory effect of AFs on rumen microorganisms. It was shown that at high concentration ranges (5 and 10 µg mL−1), AFB1 completely inhibited many ruminal bacteria [27] and harmed ruminal fermentation parameters in vitro [28,29]. Moreover, the fodder composition also impacts the degradation success since the high-starch diet increased the available AFB1 and ochratoxin [30]. Some authors disputed that rumen fluid affected AFB1 in contrast to deoxynivalenol, T-2, or zearalenone mycotoxins [31,32].
4. Pre-Harvest Biocontrol
4.1. Pre-Harvest Biocontrol by Competitive Exclusion
A biocontrol technology relying on endemic non-aflatoxigenic Aspergillus flavus strains as competitors of AF-producer fungi is becoming widespread, as it is remarkably successful, cost-effective, and environmentally friendly.
Carefully selected endemic non-aflatoxigenic genotypes can efficiently reduce AF contamination when applied before plant flowering. Hruska et al. [33] demonstrated via competitive exclusion that the decreased AF level was positively correlated to the low population of the toxigenic A. flavus under treatment, where the biocontrol A. flavus strain showed increased propagation and colonization. Recent studies demonstrated that biocontrol non-aflatoxigenic strains reduced AF concentrations in treated crops by more than 80% under both field and storage conditions [34,35,36]. In Argentina, a native atoxigenic A. flavus strain showed a remarkable biocontrol potential on peanuts, and the pre-harvest application of the biocontrol agent had a carry-over effect and protected peanuts under storage conditions [37]. The main criterion in the strain selection is the high colonization ability of the atoxigenic strain [38]. Researchers showed there was no competitive advantage of the selected atoxigenic fungal strains over the toxigenic strains and could exclude the AF role in peanut infections of A. flavus and A. parasiticus [39]. Moreover, these strains were equally applicable to peanut and maize host plants [40]. Multi-strain biocontrol products of non-aflatoxigenic A. flavus showed both immediate and long-term beneficial effects under different conditions compared with single-strain products [34,38,41,42]. However, the employed biocontrol product’s efficacy depends on several factors, including inoculum rate, formulation, application of herbicide, the soil’s temperature, and the availability of water and substrate [7,43]. Extrolites (e.g., volatile organic carbons and secondary metabolites) secreted by the biocontrol strains may also increase the efficacy of the control, and future biocontrol strategies may take advantage of these not-yet-characterized compounds [44]. Aspergilli have an outstanding secondary metabolite production potential, and it includes aflatrems, aflavarins, aflavinines (only in sclerotium producers), cyclopiazonic acids, kojic acid, and other potentially toxic compounds besides AFs [1]. The possible overproduction of any health hazard metabolites, like cyclopiazonic acid, should be carefully checked in the chosen strains. The potential non-aflatoxigenic biocontrol strains should be real atoxigenic ones [45] without any production of, at least, the known toxic molecules under field conditions.
Formulation and application strategies of the biocontrol agent are of paramount importance to reach the required efficacies. Solid-state fermented rice, encapsulated A. flavus fungal conidia in an extrusion product (Pesta), pregelatinized corn flour granules [46], rice, cracked barley, intact canola seed [39], and sterile sorghum grain [34] were treated with a spore suspension of nontoxigenic A. flavus, A. parasiticus, or both. Application of all formulations significantly decreased the AF contamination of peanuts, and the strains were found to be long-term viable on these matrixes. A more significant or consistent AF reduction was recorded with fluid and granular delivery of biocontrol strains of A. flavus, and both methods were efficient and cost-effective [47]. However, granular delivery has not gained ground in field applications because of the difficulties with applying a granular product through the canopy in the reproductive stage of the maize development. Water-dispersible granule formulations of biocontrol strains can also be useful. Weaver et al. [48] demonstrated that their new formulation with higher wettability and rapid dispersion resulted in more than 49% decrease in AF contaminants in all treatments with the Alfa-Guard biocontrol strain (A. flavus NRRL 21882). In another study, Accinelli et al. [49] evaluated an application method for the biocontrol strains on leaf. The preparation proved to be adherent, the biocontrol strain showed good leaf surface colonization, and it reduced AF level of the kernels by up to 80%–90%. In comparison, the number of AF-producing A. flavus in the soil was not changed significantly [49].
An exciting novel approach was also published by Accinelli et al. [50], who used a starch-based bioplastic formulation to coat corn kernels, which contained two conventional pesticides and spores of the non-aflatoxigenic A. flavus NRRL 30797 strain. Significantly, the additives did not affect the kernel germination adversely or the seedlings’ growth, while the AF contamination was reduced.
The competitive exclusion method was first set in the USA, and similar technologies working with endemic strains have been developed in several African countries [34,41,42,51,52] (Table 1). A non-aflatoxigenic strain (A. flavus NRRL 21882) under the commercial name Afla-Guard (Syngenta, Basel, Switzerland), which is marketed in the USA, has been used successfully on maize, groundnuts, pistachios, and cottonseed for many years [36,40]. A mixture of four endemic non-aflatoxigenic A. flavus strains called Aflasafe (Ibadan, Nigeria) also has been used on maize and groundnuts in various African countries, with an AF contamination reduction rate of 80–99% [35,41,51,53,54,55,56]. A commercial product AF-X1 (A. flavus MUCL54911, Pioneer Hi-Bred, Italy) is applied in Italy to prevent AF contamination [57].

Table 1.
Aflatoxin elimination by non-aflatoxigenic Aspergillus strains.
4.2. Pre-Harvest Biocontrol by Microbial Biofungicides
Besides atoxigenic A. flavus biocontrol strains, some other promising biocontrol agents are emerging against AF-producer molds. For example, a Trichoderma harzianum strain was applied to restrict A. flavus contamination, with 57% and 65% reduction on AF levels in groundnut [58] and in sweet corn [59], but there were no commercialized products found against A. flavus [60]. However, Lagogianni and Tsitsigiannis [24] evaluated six biofungicides/stimulants (Botector® (Aureobasidium pullulans as anti-Botrytis agent; bio-ferm GmbH, Getzersdorf, Austria), Mycostop® (Streptomyces griseoviridis as anti-Fusarium, Phytophthora, Alternaria, and Pythium agent; Verdera Oy, Espoo, Finland); Serenade Max® (biofungicide, bactericide with Bacillus subtilis QST 713; Bayer, Auckland, New Zealand), Trianum® (Trichoderma harzianum TT-22 as biofungicide against Pythium, Rhizoctonia, Fusarium, and Sclerotinia; Koppert Biological Systems, Berkel en Rodenrijs, The Netherlands); Vacciplant® (biofungicide containing laminarine; Helena Agri-Enterprises, LLC, Collierville, TN, USA) and zeolite inorganic adsorbent) and found most of them useful in reducing A. flavus conidiospores and AFB1 production in vitro. Mycostop® and Botector® treatments decreased (43%) the AFB1 content of maize kernels.
5. Post-Harvest Management of Aflatoxin Contamination
The effects of bacteria and yeasts have also been studied extensively to reduce already manifested AF contamination [4,5,7]. Biological detoxification by microorganisms relies on the binding and transformation of AFs into less toxic metabolites by microbial biomass [4,5,21]. These post-harvest methods are needed as, despite the encouraging results, pre-harvest biocontrol methods have their drawbacks. Interactions in the microbiota are in a flux state, even at the strain level, and the biocontrol effect differs under various environmental conditions [61].
5.1. Bacteria
Lactic acid bacteria (LABs), for example, Lactobacillus acidophilus, Lactococcus lactis subsp. lactis, Lactobacillus selangorensis, Pediococcus acidilactici, Streptococcus thermophilus, Weissella confusa, Enterococcus avium, and Bifidobacterium animalis subsp. lactis, inhibit AF production or remove AFs from feed and food products (Table A1). The sufficient binding of AFs by LAB strains is dependent on the inherent features of the strain, temperature, pH, the matrix itself, and incubation time [20,62]. Asurmendi et al. [63] successfully demonstrated that all LAB strains tested inhibited the growths of aflatoxigenic A. flavus strains and their AFB1 production in brewer’s grains, which is used for feeding pigs. More recently, Saladino et al. [64] tested the beneficial effects of LAB strains on the AF content of bread and found considerable 84%–100% decreases in 4 days. Assaf et al. [65] suggested a method for reducing AFM1 in milk by biofilm-forming probiotic LAB strains. They recorded a 61% reduction of AFM1 by a Lacticaseibacillus rhamnosus (formerly Lactobacillus rhamnosus) GG biofilm. Such capable bacterial biofilms could be formed on glass, metal, plastic surfaces in a test tube, 96-well plate, or flow cell formats [66]. Wacoo et al. [67] found that the allochthonous LAB species (L. brevis, L. casei, L. fermentum, and L. plantarum) isolated from the gut microbiota bound AFs effectively.
The antifungal compounds biosynthesized by LAB can support the reduction of mycotoxin production [20,68]. These compounds usually are organic acids (e.g., acetic, lactic, and propionic acids), carboxylic acids, phenolic compounds, including phenolic acids (benzoic acids, hydroxyphenyl lactic acid, phenyl lactic acid, gallic acid, and tannins), fatty acids (3-hydroxydecanoic acid, coriolic acid, caproic acid, decanoic acid, and ricinoleic acid), volatile organic compounds (e.g., acetoin and diacetyl), cyclopeptides (e.g., cyclo(L-Leu-L-Pro), cyclo(Phe-Pro), cyclo(L-Met-L-Pro), and cyclo(L-Tyr-L-Pro)), ethanol, hydrogen peroxide, proteinaceous compounds, and reuterin [69,70,71,72]. Thus far, the process of the antifungal action of proteinaceous compounds and hydroxy fatty acids has not been elucidated [69]. However, some of them can increase cytoplasmic permeability, which can finally lead to fungal cell death [69]. H2O2 is well known for its oxidizing potential directly on the lipid components and the cellular membranes’ integrant proteins [69].
Several non-lactic acid bacteria, such as Bacillus spp., Brachybacterium spp., Brevundimonas spp., Cellulosimicrobium spp., Enterobacter spp., Escherichia spp., Klebsiella spp., Mycolicibacterium spp., Myxococcus spp., Nocardia spp., Pseudomonas spp., Rhodococcus spp., Streptomyces spp., and Stenotrophomonas spp., can also inhibit the growth and AF production of molds (Table A2). For example, probiotic Enterococcus faecium M74 and EF031 strains reduced the AFB1 content of aqueous solution by 19–38% [73]. A Bacillus subtilis strain also reduced the AFB1 content of contaminated feed and food by 60–95% [74,75]. Moreover, metabolites from Bacillus subtilis (bacillomycin D, fengycins A and B, iturin A, mycosubtilin, and plipastatins A and B), Achromobacter xylosoxidans (cyclo (L-leucyl-L-propyl)), and Streptomyces spp. (blasticidin A, aflastatin A, dioctatin A) are effective inhibitors of AF biosynthesis in vitro and in vivo in crop model systems and field [76]. While the plant-growth-promoting (PGPR) Pseudomonas aeruginosa inhibited A. flavus growth with only 15% in soil [77]. Cellulosimicrobium funkei strains showed outstandingly high (97%) AFB1 biodegradation ability, suggesting that it could be used as a feed additive [78]. Bacillus and Pseudomonas strains isolated from peanut, pistachio, and maize fields also could be promising new biocontrol agents to reduce the growth of aflatoxigenic fungi and the AF contamination of arable crops [79]. According to Wang et al. [80], the culture supernatant of Escherichia coli CG1061 (isolated from healthy chicken cecum) degraded AFB1 by 61.8%, and the strain could colonize the animal gut; therefore, it may also be a suitable candidate for AFB1 detoxification.
Application of active immobilized enzymes of bacterial origin can also be a useful tool to degrade AFs in feeds and foods. In Mycolicibacterium smegmatis (Mycobacterium smegmatis), two families of F420H2-dependent reductases were identified that catalyze AF degradation [81,82]. Meanwhile, an AF degradation enzyme (MADE) was also identified from Myxococcus fulvus [83].
Bacterial volatile organic compounds are also able to hinder or kill fungal cells. Alcaligenes faecalis N1-4 produced several antifungal volatiles and inhibited the growth of A. flavus through in vitro testing. GC-MS/MS analysis detected dimethyl disulfide and methyl isovalerate as the two primary compounds in the strain’s volatile organic carbon spectrum [84]. Dimethyl disulfide hindered the germination of conidia and the growth of A. flavus. These volatile organic compounds repressed the gene expression of 12 genes in the AF biosynthesis pathway, and eight genes were significantly downregulated [84]. In groundnut, rice, maize, and soybean with high water activity, A. flavus infection and AFs contamination were entirely inhibited by Enterobacter asburiae Vt-7 volatile organic compounds (phenyl ethyl alcohol and 1-pentanol) [85]. In Vitro, volatile organic carbons from Streptomyces yanglinensis 3-10 inhibited growth, conidial germination, asexual sporulation, and expression of AFB1 biosynthesis cluster genes in A. flavus and A. parasiticus, and, in vivo, reduced the disease symptoms on peanut kernels [86]. The volatile organic carbons suppressed the mycelial growth of more than 15 plant pathogenic fungi and an oomycete organism. Chemicals, including 2-phenyl ethanol, methyl 2-methyl butyrate, and β-caryophyllene, showed activity against A. flavus and A. parasiticus. Therefore, S. yanglinensis 3-10 may become a promising biofumigant in the control A. flavus and A. parasiticus [86].
Microbial volatile organic carbons are also investigated as plant growth inducers, whose characteristics belong to various groups of chemicals, including alcohols, sulfur compounds, terpenes, ketones, and furans. Microbial volatiles can stimulate growth by modulating hormonal balance, essential nutrients, metabolism, and sugar concentrations. The alterations are coupled mostly to cellular structure and stress response genes [87,88,89].
5.2. Yeasts
Several publications have demonstrated that yeasts, for example, Candida, Debaryomyces, Pichia, Saccharomyces, Saccharomycopsis, Saccharomycodes, Schizosaccharomyces, Trichosporon, and Zygosaccharomyces species, inhibited AF production significantly in aflatoxigenic molds (Table A3). It is considered that yeast supplementation (e.g., Kluyveromyces marxianus and Pichia kudriavzevii) improved AFB1 detoxification in the rumen, reduced the AFM1 content of milk, and improved the performances of dairy cattle [90]. Viable yeast supplement in feed exerts a positive effect on the ruminal environment, total and cellulolytic bacteria, and protozoa [91,92]. Mycotoxin binding of the feed additives, such as bentonite, modified yeast cell-wall extract, or esterified glucomannan, has been shown to reduce the toxic effects of AFB1 in different livestock species by nonspecific binding of the AFs so that they cannot be absorbed in the gastrointestinal tract [93,94,95,96,97]. However, research on the interactions between detoxifying additives and mycotoxins is rare [94,97].
Yeast volatile organic carbons also take part in the hindrance of A. flavus growth and AF production [98]. Additionally, yeasts can bind AFs reversibly and rapidly [4,7]. Consequently, the GRAS baker’s yeast Saccharomyces cerevisiae can be used safely as a feed additive to mitigate aflatoxicosis in livestock, including both broilers and ruminants [21,99,100,101,102,103,104]. Moreover, S. cerevisiae can also be used directly for AF decontamination in food. For example, Shetty et al. [105] reported on the high AF binding capability of S. cerevisiae in indigenous fermented foods from Ghana. Furthermore, S. cerevisiae and S. pastorianus converted the AFB1 content of the raw materials used for wine and beer into a less toxic substance during alcoholic fermentation [106]. Foroughi et al. [107] proposed a unique AF detoxification method for AFM1-contaminated milk. The process relied on baker’s yeast, which had been immobilized on perlite beads, and was suitable to reduce the AFM1 content of all tested milk samples without affecting the milk composition [107]. Such microbial cell immobilization-based methods can be of outstanding practical value and importance when AF decontamination of milk and other drinks is considered.
5.3. Fungal Biomass, Enzymes, and Antifungal Proteins
Glucomannan from fungal cell wall or peptidoglycan and other cell wall polysaccharides can be effective adsorbents for mycotoxins because of their structural complexity. Saki et al. [108] tested the effect of Mycosorb (patented glucomannan-containing yeast product derived from yeast cell wall; Alltech) on broiler performance, organ weight, protein digestibility, plasma characteristics, and metabolizable energy of the diets. They found that Mycosorb was effective in mitigating the harmful effects of AFs in broiler chickens. Haidukowski et al. [109] also demonstrated that nonviable Pleurotus eryngii mycelia could be used as a practical and economically feasible feed additive for AFB1 detoxification.
Mycotoxin enzymatic degradation is a simple method for usage in food decontamination. However, for AFs degradation, only some fungal enzyme families are known. For example, the spent mushroom substrate crude extract (SMSE) is a rich source of AF-degrading enzymes (e.g., laccase and Mn-peroxidase), and thereby it is a good candidate for the detoxification of AF-contaminated commodities in the future [110]. An extracellular enzyme from the edible mushroom Pleurotus ostreatus showed remarkable AF-degradation activity via cutting the lactone ring of AFB1 [111]. Manganese peroxidase from Phanerochaete sordida YK-624 catalyzed the detoxification by the oxidation of AFB1 to AFB1-8,9-epoxide, and the subsequent hydrolysis to AFB1-8,9-dihydrodiol [112]. Another well-studied AF oxidase, the former AF-detoxifizyme from Armillaria tabescens (Armillariella tabescens) E-20, also attacks the 8,9-unsaturated carbon-carbon bond in AFB1 [113]. Besides enzymes, small-molecular-mass antifungal proteins from filamentous fungi are also characterized by their initiated apoptotic cell death in sensitive fungal pathogens [114,115,116] and regarded as promising future biocontrol agents against many plant-pathogenic and food-deteriorating fungi, including A. flavus.
6. Conclusions and Future Trends
Natural methods reducing the use of synthetic chemicals represent a promising future trend in AF eliminations. Combinations of physical and biological (natural) methods are expected to improve AF decontamination efficiency, both pre- and post-harvest (Figure 1).
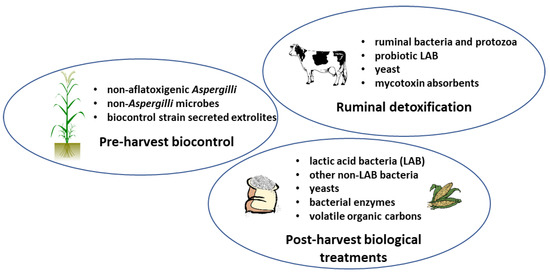
Figure 1.
Three typical areas of mitigation of aflatoxins (AF) contamination. Pre-harvest biocontrol methods with non-aflatoxigenic strains of A. flavus, other non-Aspergilli, and their extrolites are available. In animals fed with contaminated fodder, enteral or ruminal bacteria can degrade or transform AFs to lesser toxic molecules. Besides their beneficial effects on animal health, supplemented probiotic organisms (yeasts and bacteria) can also bind or degrade AFs. In stored food and feed, depending on the water content of the commodity, bacteria, yeasts, or their volatile carbons and enzymes can be used for the AF decontamination in biofilm, immobilized, or encapsulated form.
The most essential requirement for the emerging novel decontamination technologies is that these should not change the physical–chemical properties of the treated feed or food products significantly and no toxic residues of the mycotoxins should be left behind in the decontaminated products. The non-aflatoxigenic, even atoxigenic biocontrol strains are tested mostly for maize, peanuts, groundnuts, pistachios, or cottonseed, while their application in other agricultural sectors like vineyards is also a possibility. When AF contamination occurs in commodities with high water content, like milk, wine, or beer, the application of other technologies like microbial cell immobilization-based methods and enzymatic degradation can have an outstanding practical value and importance. Under the storage of the commodities or in packaging methods, the promising alternatives to synthetic chemicals are the microbial (fungal or bacterial) volatile organic carbons.
Since mammals lack strong natural ruminal or cellular AF degradation, the usual and promising agricultural technology is to help animals with potent probiotic yeasts or bacteria or only their polysaccharides to mitigate the toxic effects. Moreover, pro- or prebiotics are also applied as food supplements. The probiotic supplements have more benefits than the inorganic mycotoxin binders in toxin mitigation, as the microbes have positive physiological effects besides AF binding. Nevertheless, the AF mitigation efficiency is greatly influenced by the nature of the products and the AF contamination level. However, some authors debated the safety of elongated application of LAB or other microbes in food if these cells cannot degrade AFs [117]. Therefore, there is a need to employ starter and probiotic cultures with AF degradation abilities.
It can be stated that there is no general all-purpose decontamination method that could be broadly employed and, hence, one of the main future challenges in this field is to develop new procedures that would support comparable detoxification in a broad spectrum of feed and food matrices. Future research should focus on elaborating these novel technologies and their extensive testing in as versatile feed and food matrices as possible.
Author Contributions
Conceptualization, I.P. and Z.G.; writing—original draft preparation, F.P., P.S., and S.K.; writing—review and editing, T.P.; supervision, I.P.; funding acquisition, I.P. All authors have read and agreed to the published version of the manuscript.
Funding
Project no. 2018-1.2.1-NKP-2018-00002 has been implemented with the support provided from the National Research, Development, and Innovation Fund of Hungary, financed under the 2018-1.2.1-NKP funding scheme. Project no. TKP2020-IKA-04 has been implemented with the support provided from the National Research, Development, and Innovation Fund of Hungary, financed under the 2020-4.1.1-TKP2020 funding scheme (T.P.).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Conflicts of Interest
The authors declare no conflict of interest.
Appendix A

Table A1.
Aflatoxin elimination by lactic acid bacteria. We intended to collect data on the viable cells’ bounding properties, usually in PBS or milk at 4 °C.
Table A1.
Aflatoxin elimination by lactic acid bacteria. We intended to collect data on the viable cells’ bounding properties, usually in PBS or milk at 4 °C.
| Aflatoxin | Bacteria | Strain | Toxin Elimination (%) | References |
|---|---|---|---|---|
| B1 | Bifidobacterium animalis subsp. animalis (formerly Bifidobacterium animalis) | CSCC 1941 | 45.7 | [118] |
| B1 | Bifidobacterium animalis subsp. lactis (formerly Bifidobacterium lactis) | E-94508 | 18 | |
| CSCC 5094 | 34.7 | |||
| CSCC 1906 | 48.7 | |||
| B1 | Bifidobacterium longum | CSCC 5304 | 37.5 | |
| B1 | Enterococcus faecium | M74 | 19.3–30.5 | [73] |
| EF031 | 23.4–37.5 | |||
| B1 | Lactobacillus acidophilus | ATCC 4356 | 48.3 | [119] |
| E-94507 | 18.2 | [118] | ||
| CSCC 5361 | 20.7 | |||
| Chr. Hansen, Denmark | 56.6 | [120] | ||
| CH-2 | 36 | [121] | ||
| B1 | Lactobacillus amylovorus | CSCC 5160 | 59.7 | [118] |
| CSCC 5197 | 57.8 | |||
| B1 | Lactobacillus bulgaricus | Chr. Hansen, Denmark | 16.3 | [120] |
| B1 | Lacticaseibacillus casei (formerly Lactobacillus casei) Shirota | YIT 901, commercial | 21.8 | [119] |
| Yakult | 98 | [122] | ||
| B1 | Lacticaseibacillus casei (formerly Lactobacillus casei) | Chr. Hansen, Denmark | 22.4 | [120] |
| Iranian dairy products | 43 | [123] | ||
| B1 | Lactobacillus delbrueckii | MK9 | 17.3 | [118] |
| B1 | Lactobacillus delbrueckii subsp. bulgaricus | Australian Starter Culture Research Centre Collection, Werribee, Australia | 15.6 | [119] |
| B1 | Limosilactobacillus fermentum (formerly Lactobacillus fermentum) | CSCC 5362 | 22.6 | [118] |
| N.A. | ≥60 | [93] | ||
| Iranian sourdough | 61 | [123] | ||
| PTCC 1744 | 99.9 | [124] | ||
| B1 | Lactobacillus helveticus | Australian Starter Culture Research Centre Collection, Werribee, Australia | 17.5 | [119] |
| Aki4 | 30.1 | [118] | ||
| Chr. Hansen, Denmark | 17.8 | [120] | ||
| B1 | Lactobacillus johnsonii | CSCC 5142 | 30.1 | [118] |
| B1 | Lactiplantibacillus plantarum (formerly Lactobacillus plantarum) | ATCC 8014 | 29.9 | [119] |
| E-79098 | 28.4 | [118] | ||
| N.A. | ≥40 | [93] | ||
| Iranian dairy products | 56 | [123] | ||
| EMCC-1039 | 39.2 | [121] | ||
| B1 | Lacticaseibacillus rhamnosus (formerly Lactobacillus rhamnosus) | DSM 7061 | 76.5 | [119] |
| ATCC 53103 | 54 | [125] | ||
| 78.9 | [119] | |||
| E-97800 | 22.7 | [118] | ||
| Lc 1/3 | 54.6 | |||
| NRRL B-445 | 17.2 | [121] | ||
| B1 | Lactococcus lactis subsp. lactis | Australian Starter Culture Research Centre Collection, Werribee, Australia | 59 | [119] |
| E-90414 | 31.6 | [118] | ||
| Dairy products | 54.3 | [126] | ||
| B1 | Lactococcus lactis subsp. cremoris | Australian Starter Culture Research Centre Collection, Werribee, Australia | 26.9 | [119] |
| MK4 | 5.6 | [118] | ||
| ARH74 (Valio Ltd, Finland) | 41.1 | |||
| B1 | Lactobacillus selangorensis (formerly Paralactobacillus selangorensis) | N.A. | <39 | [93] |
| B1 | Pediococcus acidilactici | N.A. | 45–59 | [93] |
| B1 | Propionibacterium freudenreichii subsp. shermanii JS | Valio Ltd. Finland | 22–82 | [125] |
| [119] | ||||
| B1 | Streptococcus thermophilus | Australian Starter Culture Research Centre Collection, Werribee, Australia | 32.7 | [119] |
| Dairy products | 81 | [126] | ||
| CH-1 | 26.9 | [121] | ||
| B1 | Weissella confuse | N.A. | 15–39 | [93] |
| M1 | Bifidobacterium bifidum | Bb13 | 23.48 | [127] |
| NCC 381 | 18.11 | |||
| PTCC 1644 | 99.9 | [124] | ||
| M1 | Bifidobacterium lactis | FLORA-FIT BI07 (Danisco Ltd.) | 16.89 | [128] |
| M1 | Enterococcus avium | CTC 469 (Tecnolat, Brazil) | 7.36 | [128] |
| M1 | Lactobacillus acidophilus | NCC 12 | 16.05 | [127] |
| NCC 36 | 22.23 | [127] | ||
| NCC 68 | 14.22 | [127] | ||
| LA-5 (Chr. Hansen, Denmark) | 95 | [62] | ||
| M1 | Lactobacillus delbrueckii subsp. bulgaricus | LB340 (Danisco Ltd.) | 30.22 | [128] |
| CH-2 (Chr. Hansen, Denmark) | 18.7 | [129] | ||
| M1 | Lactobacillus gasseri | ATCC 33323 | 21.37 | [128] |
| M1 | Lactiplantibacillus plantarum (formerly Lactobacillus plantarum) | CTC368 | 5.6 | [128] |
| M1 | Lacticaseibacillus rhamnosus (formerly Lactobacillus rhamnosus) | Ezal, France | 20.21 | [127] |
| HOWARU (Danisco Ltd.) | 17.13 | [128] | ||
| M1 | Pediococcus pentosaceus | TR570 (Tecnolat, Brazil) | 8.68 | [128] |
| M1 | Streptococcus thermophilus | ST-36 (Chr. Hansen, Denmark) | 29.42 | [129] |

Table A2.
Aflatoxin elimination by non-lactic acid bacteria. We intended to collect data on the viable cells’ bounding properties, usually in PBS or milk at 4 °C.
Table A2.
Aflatoxin elimination by non-lactic acid bacteria. We intended to collect data on the viable cells’ bounding properties, usually in PBS or milk at 4 °C.
| Toxin | Bacteria | Strain | Toxin Elimination (%) | References |
|---|---|---|---|---|
| B1 | Bacillus licheniformis | Thai fermented soybean | 74 | [130] |
| B1 | Bacillus stearothermophilus | N.A. | 87 | [131] |
| B1 | Bacillus subtilis | UTBSP1 | 85.66 | [74] |
| B1, G1, M1 | ANSB060 | 81.5, 81, 60 | [132] | |
| B1 | Brachybacterium spp. | Rabbit feces | 74.83 | [133] |
| B1 | Brevundimonas spp. | Yellow cheek feces | 76.86 | [133] |
| B1 | Cellulosimicrobium funkei | Soil around coal factories | 97 | [78] |
| B1 | Enterobacter spp. | Hog deer feces | 76 | [133] |
| B1 | Escherichia coli | Strain CG1061 | 18–62 | [80] |
| B1 | Klebsiella spp. | Rabbit feces | 78 | [133] |
| B1 | Mycolicibacterium fluoranthenivorans (formerly Mycobacterium fluoranthenivorans) | DSM 44556 | >90 | [134] |
| B1 | Mycolicibacterium smegmatis (formerly Mycobacterium smegmatis) | N.A. | >90 | [82] |
| B1, G1, M1 | Myxococcus fulvus | ANSM068, Deer feces | 72, 68, 64 | [83] |
| B1 | Nocardia corynebacterioides (formerly Flavobacterium aurantiacum) | DSM 12,676 | 74.5 | [135] |
| NRRL B-184 | 85 | [136] | ||
| B1, B2M1 | Pseudomonas aeruginosa | Grain kernels and soils | 83, 47, 32 | [137] |
| B1 | Pseudomonas stutzeri | F4 strain, Budorcas taxicolor feces | 90 | [138] |
| B1 | Rhodococcus erythropolis | Strain 4.1491 | 96 | [139] |
| ATCC 4277 | [140] | |||
| 18 strains | 70–100 | [141] | ||
| DSM 14303 | 83 | [134] | ||
| B1 | Streptomyces aureofaciens | ATCC 10762 | 88 | [140] |
| B1 | Streptomyces lividans | TK 24 | 86 | [140] |
| B1 | Stenotrophomonas maltophilia | South American tapir feces | 85 | [133] |

Table A3.
Aflatoxin elimination by some yeast cultures.
Table A3.
Aflatoxin elimination by some yeast cultures.
| Toxin | Fungi | Strain | Toxin Elimination (%) | References |
|---|---|---|---|---|
| B1 | Aureobasidium pullulans | H1 | 68.61 | [142] |
| B1 | Candida albicans | AA17 | 50.34 | [142] |
| AA19 | 64.61 | |||
| B1 | Diutina catenulata (formerly Candida catenulata) | N.A. | <15 | [93] |
| B1 | Candida krusei | N.A. | 15–60 | [93] |
| AUMC 8161 | 100 | [143] | ||
| B1, B2, G1, G2 | Candida parapsilosis | N.A. | 15–72 | [93] |
| IP1698 | 99.94 | [144] | ||
| B1 | Wickerhamiella sorbophila (formerly Candida sorbophila) | ECF16 | 52.59 | [142] |
| ECF85 | 51.49 | |||
| B1 | Debaryomyces hansenii | N.A. | 15–39 | [93] |
| M1 | Kluyveromyces lactis | N.A. | 60–69 | [145] |
| B1 | Komagataella pastoris | EW1 | 71.5 | [142] |
| EW3 | 51.5 | |||
| EW6 | 50.5 | |||
| B1 | Wickerhamomyces anomalus (formerly Pichia anomala) | N.A. | 15 | [93] |
| AUMC 2674 | 100 | [143] | ||
| AFs | Meyerozyma guilliermondii (formerly Pichia guilliermondii) | AUMC 2663 | 80 | [143] |
| B1 | Pichia membranifaciens | N.A. | 40–59 | [93] |
| B1 | Rhodotorula mucilaginosa | various strains | 52.77–70.2 | [142] |
| B1 | Saccharomyces cerevisiae | N.A. | 10–60 | [93] |
| CECT 1891 | - * | [146] | ||
| A18 | 53 | [105] | ||
| 26.1.11 | 48.8 | [105] | ||
| RC 016 | - | [101] | ||
| EB34 | 52.25 | [142] | ||
| EB57 | 51.12 | |||
| M1 | Saccharomyces cerevisiae | SAFLAGER W37/70 | 90.3 | [147] |
| N.A. | 81.3 | [107] | ||
| ATCC 9763 | 75 | |||
| B1 | Saccharomycopsis fibuligera | N.A. | <15 | [93] |
| B1 | Saccharomycodes ludwigii | N.A. | 40–59 | |
| B1 | Schizosaccharomyces pombe | N.A. | 40–59 | |
| B1 | Cutaneotrichosporon mucoides (formerly Trichosporon mucoides) | N.A. | <15 | |
| B1 | Zygosaccharomyces bailii | N.A. | 15–39 |
* no data.
References
- Frisvad, J.C.; Hubka, V.; Ezekiel, C.N.; Hong, S.B.; Nováková, A.; Chen, A.J.; Arzanlou, M.; Larsen, T.O.; Sklenář, F.; Mahakarnchanakul, W.; et al. Taxonomy of Aspergillus section Flavi and their production of aflatoxins, ochratoxins and other mycotoxins. Stud. Mycol. 2019, 93, 1–63. [Google Scholar] [CrossRef]
- Lizárraga-Paulín, E.G.; Miranda-Castro, S.P.; Moreno-Martínez, E.; Torres-Pacheco, I.; Lara-Sagahón, A.V. Novel methods for preventing and controlling aflatoxins in food: A worldwide daily challenge. In Aflatoxins-Recent Advances and Future Prospects; Razzaghi-Abyaneh, M., Ed.; InTech: Rijeka, Croatia, 2013; pp. 93–128. [Google Scholar]
- Caceres, I.; Al Khoury, A.; El Khoury, R.; Lorber, S.; Oswald, I.P.; El Khoury, A.; Atoui, A.; Puel, O.; Bailly, J.-D. Aflatoxin biosynthesis and genetic regulation: A Review. Toxins 2020, 12, 150. [Google Scholar] [CrossRef]
- Pfliegler, V.; Pócsi, I.; Győri, Z.; Pusztahelyi, T. The Aspergilli and their mycotoxins: Metabolic interactions with plants and the soil biota. Front. Microbiol. 2020, 10, 1–45. [Google Scholar] [CrossRef]
- Peles, F.; Sipos, P.; Győri, Z.; Pfliegler, W.P.; Giacometti, F.; Serraino, A.; Pagliuca, G.; Gazzotti, T.; Pócsi, I. Adverse effects, transformation, and channeling of aflatoxins into food raw materials in livestock. Front. Microbiol. 2019, 10, 2861. [Google Scholar] [CrossRef] [PubMed]
- Ráduly, Z.; Szabó, L.; Madar, A.; Pócsi, I.; Csernoch, L. Toxicological and medical aspects of Aspergillus-derived mycotoxins entering the feed and food chain. Front. Microbiol. 2020, 10, 2908. [Google Scholar] [CrossRef]
- Jalili, M. A review on aflatoxins reduction in food. Iran. J. Health Saf. Environ. 2015, 3, 445–459. [Google Scholar]
- Torres, A.M.; Barros, G.G.; Palacios, S.A.; Chulze, S.N.; Battilani, P. Review on pre- and post-harvest management of peanuts to minimize aflatoxin contamination. Food Res. Int. 2014, 62, 11–19. [Google Scholar] [CrossRef]
- Norlia, M.; Jinap, S.; Nor-Khaizura, M.; Radu, S.; Samsudin, N.; Azri, F.A. Aspergillus section Flavi and aflatoxins: Occurrence, detection, and identification in raw peanuts and peanut-based products along the supply chain. Front. Microbiol. 2019, 10, 2602. [Google Scholar] [CrossRef]
- Ketney, O.; Santini, A.; Oancea, S. Recent aflatoxin survey data in milk and milk products: A review. Int. J. Dairy Technol. 2017, 70, 320–331. [Google Scholar] [CrossRef]
- Serraino, A.; Bonilauri, P.; Kerekes, K.; Farkas, Z.; Giacometti, F.; Canever, A.; Zambrini, A.V.; Ambrus, Á. Occurrence of aflatoxin M1 in raw milk marketed in Italy: Exposure assessment and risk characterization. Front. Microbiol. 2019, 10, 2516. [Google Scholar] [CrossRef] [PubMed]
- Maleki, F.; Abdi, S.; Davodian, E.; Haghani, K.; Bakhtiyari, S. Exposure of infants to aflatoxin M1 from mother’s breast milk in Ilam, Western Iran. Osong Public Health Res. Perspec. 2015, 6, 283–287. [Google Scholar] [CrossRef] [PubMed]
- Warth, B.; Braun, D.; Ezekiel, C.N.; Turner, P.C.; Degen, G.H.; Marko, D. Biomonitoring of mycotoxins in human breast milk: Current state and future perspectives. Chem. Res. Toxicol. 2016, 29, 1087–1097. [Google Scholar] [CrossRef] [PubMed]
- Uka, V.; Moore, G.G.; Arroyo-Manzanares, N.; Nebija, D.; De Saeger, S.; Di Mavungu, D.J. Secondary metabolite dereplication and phylogenetic analysis identify various emerging mycotoxins and reveal the high intra-species diversity in Aspergillus flavus. Front. Microbiol. 2019, 10, 667. [Google Scholar] [CrossRef] [PubMed]
- Fisher, W.J.; Schilter, B.; Tritsher, A.M.; Stadler, R.H. Environmental contaminants. Contaminants of milk and dairy products. In Encyclopedia of Dairy Science, 2nd ed.; Fuquay, J.W., Fox, P.F., McSweeney, P.L.H., Eds.; Academic Press: London, UK, 2011; pp. 898–905. [Google Scholar]
- Costamagna, D.; Gaggiotti, M.; Chiericatti, C.A.; Costabel, L.; Audero, G.M.L.; Taverna, M.; Signorini, M.L. Quantification of aflatoxin M1 carry-over rate from feed to soft cheese. Toxicol. Rep. 2019, 6, 782–787. [Google Scholar] [CrossRef]
- Jasutiene, I.; Garmiene, G.; Kulikauskiene, M. Pasteurisation and fermentation effects on Aflatoxin M1 stability. Milchwissenschaft 2006, 61, 75–79. [Google Scholar]
- Raters, M.; Matissek, R. Thermal stability of aflatoxin B1 and ochratoxin A. Mycotoxin Res. 2008, 24, 130–134. [Google Scholar] [CrossRef]
- Assaf, J.C.; Nahle, S.; Chokr, A.; Louka, N.; Atoui, A.; El Khoury, A. Assorted methods for decontamination of aflatoxin M1 in milk using microbial adsorbents. Toxins 2019, 11, 304. [Google Scholar] [CrossRef]
- Ahlberg, S.H.; Joutsjoki, V.; Korhonen, H.J. Potential of lactic acid bacteria in aflatoxin risk mitigation. Int. J. Food Microbiol. 2015, 207, 87–102. [Google Scholar] [CrossRef]
- Tian, F.; Chun, H.S. Natural products for preventing and controlling aflatoxin contamination of food. In Aflatoxin-Control, Analysis, Detection and Health Risks; Abdulra’uf, L., Ed.; IntechOpen: London, UK, 2017; pp. 13–44. [Google Scholar]
- Medeiros, F.H.V.; de Martins, S.J.; Zucchi, T.D.; Melo, I.S.; de Batista, L.R.; Machado, J.D.C. Biological control of mycotoxin-producing molds. Ciênc. Agrotec. 2012, 36, 483–497. [Google Scholar] [CrossRef]
- Tsitsigiannis, D.I.; Dimakopoulou, M.; Antoniou, P.P.; Tjamos, E.C. Biological control strategies of mycotoxigenic fungi and associated mycotoxins in Mediterranean basin crops. Phytopathol. Mediterr. 2012, 51, 158–174. [Google Scholar]
- Lagogianni, C.S.; Tsitsigiannis, D.I. Effective biopesticides and biostimulants to reduce aflatoxins in maize fields. Front. Microbiol. 2019, 10, 2645. [Google Scholar] [CrossRef] [PubMed]
- Engel, V.G.; Hagemeister, H. 1978 Untersuchungenueber den verblieb von aflatoxin B1 im Verdaaundtarkt von Kuehen. In Biological Detoxification of Fungal Toxin and its use in Plant Breeding, Feed and food production. Nat. Toxins 1999, 7, 1–23. [Google Scholar]
- Upadhaya, S.D.; Sung, H.G.; Lee, C.H.; Lee, S.Y.; Kim, S.W.; Jo, K.J.; Ha, J.K. Comparative study on the aflatoxin B1 degradation ability of rumen fluid from Holstein steers and Korean native goats. J. Vet. Sci. 2009, 10, 29–34. [Google Scholar] [CrossRef] [PubMed]
- Jouany, J.-P.; Yiannikouris, A.; Bertin, G. Risk assessment of mycotoxins in ruminants and ruminant products. Opt. Mediterr. A 2009, 85, 205–224. [Google Scholar]
- Khodabandehloo, M.; Malecky, M.; Aliarabi, H.; Saki, A.A.; Alipour, D. In Vitro evaluation of aflatoxin B1 effect on gas production and ruminal fermentation parameters. Iran. J. Vet. Res. 2019, 20, 263–269. [Google Scholar]
- Singh, R.; Park, S.; Koo, J.; Balasubramanian, B. Influence of various concentrations of aflatoxin B1 on in vitro rumen fermentation of a buffalo diet. Korean J. Agric. Sci. 2020, 47, 131–138. [Google Scholar]
- Pantaya, D.; Morgavi, D.P.; Silberberg, M.; Chaucheyras-Durand, F.; Martin, C.; Wiryawan, K.G.; Boudra, H. Bioavailability of aflatoxin B1 and ochratoxin A, but not fumonisin B1 or deoxynivalenol, is increased in starch-induced low ruminal pH in nonlactating dairy cows. J. Dairy Sci. 2016, 99, 9759–9767. [Google Scholar] [CrossRef]
- Kiessling, K.H.; Pettersson, H.; Sandholm, K.; Olsen, M. Metabolism of aflatoxin, ochratoxin, zearalenone, and three trichothecenes by intact rumen fluid, rumen protozoa, and rumen bacteria. Appl. Environ. Microbiol. 1984, 47, 1070–1073. [Google Scholar] [CrossRef]
- Westlake, K.; Mackie, R.I.; Dutton, M.F. In Vitro metabolism of mycotoxins by bacterial, protozoal, and ovine ruminal fluid preparations. Anim. Feed Sci. Technol. 1989, 25, 169–178. [Google Scholar] [CrossRef]
- Hruska, Z.; Rajasekaran, K.; Yao, H.; Kincaid, R.; Darlington, D.; Brown, R.L.; Bhatnagar, D.; Cleveland, T.E. Co-inoculation of aflatoxigenic and non-aflatoxigenic strains of Aspergillus flavus to study fungal invasion, colonization, and competition in maize kernels. Front. Microbiol. 2014, 5, 122. [Google Scholar] [CrossRef]
- Atehnkeng, J.; Ojiambo, P.S.; Cotty, P.J.; Bandyopadhyay, R. Field efficacy of a mixture of atoxigenic Aspergillus flavus Link:Fr vegetative compatibility groups in preventing aflatoxin contamination in maize (Zea mays L.). Biol. Control. 2014, 72, 62–70. [Google Scholar] [CrossRef]
- Bandyopadhyay, R.; Ortega-Beltran, A.; Akande, A.; Mutegi, C.; Atehnkeng, J.; Kaptoge, L.; Senghor, A.L.; Adhikari, B.N.; Cotty, P.J. Biological control of aflatoxins in Africa: Current status and potential challenges in the face of climate change. World Mycotoxin J. 2016, 9, 771–789. [Google Scholar] [CrossRef]
- Lewis, M.H.; Carbone, I.; Luis, J.M.; Payne, G.A.; Bowen, K.L.; Hagan, A.K.; Kemerait, R.; Heiniger, R.; Ojiambo, P.S. Biocontrol strains differentially shift the genetic structure of indigenous soil populations of Aspergillus flavus. Front. Microbiol. 2019, 10, 1738. [Google Scholar] [CrossRef]
- Chulze, S.N.; Palazzini, J.M.; Torres, A.M.; Barros, G.; Ponsone, M.L.; Geisen, R.; Schmidt-Heydt, M.; Köhl, J. Biological control as a strategy to reduce the impact of mycotoxins in peanuts, grapes and cereals in Argentina. Food Addit. Contam. Part A Chem. Anal. Control Expo. Risk Assess. 2015, 32, 471–479. [Google Scholar] [CrossRef] [PubMed]
- Agbetiameh, D.; Ortega-Beltran, A.; Awuah, R.T.; Atehnkeng, J.; Islam, M.-S.; Callicott, K.A.; Cotty, P.J.; Bandyopadhyay, R. Potential of atoxigenic Aspergillus flavus vegetative compatibility groups associated with maize and groundnut in Ghana as biocontrol agents for aflatoxin management. Front. Microbiol. 2019, 10, 2069. [Google Scholar] [CrossRef]
- Pitt, J.I.; Hocking, A.D. Mycotoxins in Australia: Biocontrol of aflatoxin in peanuts. Mycopathologia 2006, 162, 233–243. [Google Scholar] [CrossRef]
- Dorner, J.W.; Cole, R.J.; Wicklow, D.T. Aflatoxin reduction in corn through field application of competitive fungi. J. Food Prot. 1999, 62, 650–656. [Google Scholar] [CrossRef]
- Probst, C.; Bandyopadhyay, R.; Price, L.E.; Cotty, P.J. Identification of atoxigenic Aspergillus flavus isolates to reduce aflatoxin contamination of maize in Kenya. Plant. Dis. 2011, 95, 212–218. [Google Scholar] [CrossRef]
- Unnevehr, L.; Grace, D. (Eds.) Tackling Aflatoxins: An Overview of Challenges and Solutions in Aflatoxins. In Finding Solutions for Improved Food Safety; International Food Policy Research Institute: Washington, DC, USA, 2013. [Google Scholar]
- Mohale, S.; Medina, A.; Magan, N. Effect of environmental factors on in vitro and in situ interactions between atoxigenic and toxigenic A. flavus strains and control of aflatoxin contamination of maize. Biocontrol Sci. Technol. 2013, 23, 776–793. [Google Scholar] [CrossRef]
- Moore, G.G.; Lebar, M.D.; Carter-Wientjes, C.H. The role of extrolites secreted by non-aflatoxigenic Aspergillus flavus in biocontrol efficacy. J. Appl. Microbiol. 2018, 126, 1257–1264. [Google Scholar] [CrossRef]
- Abbas, H.K.; Zablotowicz, R.M.; Horn, B.W.; Phillips, N.A.; Johnson, B.J.; Jin, X.; Abel, C.A. Comparison of major biocontrol strains of non-aflatoxigenic Aspergillus flavus for the reduction of aflatoxins and cyclopiazonic acid in maize. Food Addit. Contam. Part A Chem. Anal. Control Expo. Risk Assess. 2011, 28, 198–208. [Google Scholar] [CrossRef] [PubMed]
- Dorner, J.W.; Cole, R.J.; Connick, W.J.; Daigle, D.J.; McGuire, M.R.; Shasha, B.S. Evaluation of biological control formulations to reduce aflatoxin contamination in peanuts. Biol. Control 2003, 26, 318–324. [Google Scholar] [CrossRef]
- Accinelli, C.; Abbas, H.K.; Vicaria, A.; Shier, W.T. Evaluation of recycled bioplastic pellets and a sprayable formulation for application of an Aspergillus flavus biocontrol strain. Crop. Prot. 2015, 72, 9–15. [Google Scholar] [CrossRef]
- Weaver, M.A.; Abbas, H.K.; Jin, X.; Elliott, B. Efficacy of water-dispersible formulations of biological control strains of Aspergillus flavus for aflatoxin management in corn. Food Addit. Contam. Part A Chem. Anal. Control Expo. Risk Assess. 2016, 33, 346–351. [Google Scholar] [PubMed]
- Accinelli, C.; Abbas, H.K.; Vicaria, A.; Shier, W.T. Leaf application of a sprayable bioplastic-based formulation of biocontrol Aspergillus flavus strains for reduction of aflatoxins in corn. Pest Manag. Sci. 2016, 72, 1521–1528. [Google Scholar] [CrossRef] [PubMed]
- Accinelli, C.; Abbas, H.K.; Little, N.; Kotowicz, J.K.; Shier, W.T. Biological control of aflatoxin production in corn using non-aflatoxigenic Aspergillus flavus administered as a bioplastic-based seed coating. Crop. Prot. 2018, 107, 87–92. [Google Scholar] [CrossRef]
- Senghor, L.A.; Ortega-Beltran, A.; Atehnkeng, J.; Callicott, K.A.; Cotty, P.J.; Bandyopadhyay, R. The atoxigenic biocontrol product Aflasafe SN01 is a valuable tool to mitigate aflatoxin contamination of both maize and groundnut cultivated in Senegal. Plant Dis. 2020, 104, 510–520. [Google Scholar] [CrossRef]
- Atehnkeng, J.; Ojiambo, P.S.; Donner, M.; Ikotun, T.; Sikora, R.A.; Cotty, P.J.; Bandyopadhyay, R. Distribution and toxigenicity of Aspergillus species isolated from maize kernels from three agro-ecological zones in Nigeria. Int. J. Food Microbiol. 2008, 122, 74–84. [Google Scholar] [CrossRef]
- Hell, K.; Mutegi, C. Aflatoxin control and prevention strategies in key crops of Sub-Saharan Africa. Afr. J. Microbiol. Res. 2011, 5, 459–466. [Google Scholar]
- Grace, D.; Mahuku, G.; Hoffmann, V.; Atherstone, C.; Upadhyaya, H.D.; Bandyopadhyay, R. International agricultural research to reduce food risks: Case studies on aflatoxins. Food Secur. 2015, 7, 569–582. [Google Scholar] [CrossRef]
- Udomkun, P.; Wiredu, A.N.; Nagle, M.; Müller, J.; Vanlauwe, B.; Bandyopadhyay, R. Innovative technologies to manage aflatoxins in foods and feeds and the profitability of application—A review. Food Control 2017, 76, 127–138. [Google Scholar] [CrossRef] [PubMed]
- Shenge, K.C.; Adhikari, B.N.; Akande, A.; Callicott, K.A.; Atehnkeng, J.; Ortega-Beltran, A.; Kumar, P.L.; Bandyopadhyay, R.; Cotty, P.J. Monitoring Aspergillus flavus genotypes in a multi-genotype aflatoxin biocontrol product with quantitative pyrosequencing. Front. Microbiol. 2019, 10, 2529. [Google Scholar] [CrossRef] [PubMed]
- Mauro, A.; Garcia-Cela, E.; Pietri, A.; Cotty, P.J.; Battilani, P. Biological control products for aflatoxin prevention in Italy: Commercial field evaluation of atoxigenic Aspergillus flavus active ingredients. Toxins 2018, 10, 30. [Google Scholar] [CrossRef] [PubMed]
- Kifle, M.H.; Yobo, K.S.; Laing, M.D. Biocontrol of Aspergillus flavus in groundnut using Trichoderma harzianum strain kd. J. Plant Dis. Prot. 2017, 124, 51–56. [Google Scholar] [CrossRef]
- Sivparsad, B.J.; Laing, M.D. Pre-harvest silk treatment with Trichoderma harzianum reduces aflatoxin contamination in sweetcorn. J. Plant Dis. Prot. 2016, 123, 285–293. [Google Scholar] [CrossRef]
- Ren, X.; Zhang, Q.; Zhang, W.; Mao, J.; Li, P. Control of aflatoxigenic molds by antagonistic microorganisms: Inhibitory behaviors, bioactive compounds, related mechanisms, and influencing factors. Toxins 2020, 12, 24. [Google Scholar] [CrossRef]
- Magan, N.; Aldred, D. Post-harvest control strategies: Minimizing mycotoxins in the food chain. Int. J. Food Microbiol. 2007, 119, 131–139. [Google Scholar] [CrossRef] [PubMed]
- Adibpour, N.; Soleimanian-Zad, S.; Sarabi-Jamab, M.; Tajalli, F. Effect of storage time and concentration of aflatoxin M1 on toxin binding capacity of L. acidophilus in fermented milk product. J. Agric. Sci. Technol. 2016, 18, 1209–1220. [Google Scholar]
- Asurmendi, P.; Pascual, L.; Dalcero, A.; Barberis, L. Incidence of lactic acid bacteria and Aspergillus flavus in brewer’s grains and evaluation of potential antifungal activity of these bacteria. J. Stored Prod. Res. 2014, 56, 33–37. [Google Scholar] [CrossRef]
- Saladino, F.; Luz, C.; Manyes, L.; Fernandez-Franzon, M.; Meca, G. In Vitro antifungal activity of lactic acid bacteria against mycotoxigenic fungi and their application in loaf bread shelf-life improvement. Food Control 2016, 67, 273–277. [Google Scholar] [CrossRef]
- Assaf, J.C.; Khoury, A.; Chokr, A.; Louka, N.; Atoui, A. A novel method for elimination of aflatoxin M1 in milk using Lactobacillus rhamnosus G.G. biofilm. Int. J. Dairy Technol. 2019, 72, 248–256. [Google Scholar] [CrossRef]
- Gu, H.; Ren, D. Materials, and surface engineering to control bacterial adhesion and biofilm formation: A review of recent advances. Front. Chem. Sci. Eng. 2014, 8, 20–33. [Google Scholar] [CrossRef]
- Wacoo, A.P.; Atukunda, P.; Muhoozi, G.; Braster, M.; Wagner, M.; Broek, T.J.V.D.; Sybesma, W.; Westerberg, A.C.; Iversen, P.O.; Kort, R. Aflatoxins: Occurrence, exposure, and binding to Lactobacillus species from the gut microbiota of rural Ugandan children. Microorganisms 2020, 8, 347. [Google Scholar] [CrossRef] [PubMed]
- Sadiq, F.A.; Yan, B.; Tian, F.; Zhao, J.; Zhang, H.; Wei, C. Lactic acid bacteria as antifungal and anti-mycotoxigenic agents: A comprehensive review. Compr. Rev. Food Sci. Food Saf. 2019, 18, 1403–1436. [Google Scholar] [CrossRef] [PubMed]
- Dalié, D.; Deschamps, A.; Richard-Forget, F. Lactic acid bacteria–Potential for control of mold growth and mycotoxins: A review. Food Control 2010, 21, 370–380. [Google Scholar] [CrossRef]
- Crowley, S.; Mahony, J.; van Sinderen, D. Current perspectives on antifungal lactic acid bacteria as natural bio-preservatives. Trends Food Sci. Technol. 2013, 33, 93–109. [Google Scholar] [CrossRef]
- Le Lay, C.; Coton, E.; Le Blay, G.; Chobert, J.M.; Haertlé, T.; Choiset, Y.; Longa, N.G.V.; Meslet-Cladièrea, L.; Mounie, J. Identification and quantification of antifungal compounds produced by lactic acid bacteria and propionibacteria. Int. J. Food Microbiol. 2016, 239, 79–85. [Google Scholar] [CrossRef]
- Leyva Salas, M.; Mounier, J.; Valence, F.; Coton, M.; Thierry, A.; Coton, E. Antifungal microbial agents for food biopreservation—A review. Microorganisms 2017, 5, 37. [Google Scholar] [CrossRef]
- Topcu, A.; Bulat, T.; Wishah, R.; Boyaci, I.H. Detoxification of aflatoxin B1 and patulin by Enterococcus faecium strains. Int. J. Food Microbiol. 2010, 139, 202–205. [Google Scholar] [CrossRef]
- Farzaneh, M.; Shi, Z.Q.; Ghassempour, A.; Sedaghat, N.; Ahmadzadeh, M.; Mirabolfathy, M.; Javan-Nikkhah, M. Aflatoxin B1 degradation by Bacillus subtilis UTBSP1 isolated from pistachio nuts of Iran. Food Control 2012, 23, 100–106. [Google Scholar] [CrossRef]
- Fan, Y.; Zhao, L.; Ma, Q.; Li, X.; Shi, H.; Zhou, T.; Zhang, J.; Ji, C. Effects of Bacillus subtilis ANSB060 on growth performance, meat quality and aflatoxin residues in broilers fed moldy peanut meal naturally contaminated with aflatoxins. Food Chem. Toxicol. 2013, 59, 748–753. [Google Scholar] [CrossRef]
- Razzaghi-Abyaneh, M.; Shams-Ghahfarokhi, M.; Chang, P.K. Aflatoxins: Mechanisms of inhibition by antagonistic plants and microorganisms. In Aflatoxins: Biochemistry and Molecular Biology; Guevara-Gonzalez, R.G., Ed.; IntechOpen: London, UK, 2011; pp. 285–304. [Google Scholar]
- Chandra, H.; Kumari, P.; Bisht, R.; Prasad, R.; Yadav, S. Plant growth promoting Pseudomonas aeruginosa from Valeriana wallichii displays antagonistic potential against three phytopathogenic fungi. Mol. Biol. Rep. 2020, 47, 6015–6026. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.H.; Zhang, N.; Sun, R.R.; Gao, X.; Gu, C.; Krumm, C.S.; Qi, D.S. A novel strain of Cellulosimicrobium funkei can biologically detoxify aflatoxin B1 in ducklings. Microb. Biotechnol. 2015, 8, 490–498. [Google Scholar] [CrossRef] [PubMed]
- Shams-Ghahfarokhi, M.; Kalantari, S.; Razzaghi-Abyaneh, M. Terrestrial bacteria from agricultural soils: Versatile weapons against aflatoxigenic fungi. In Aflatoxins–Recent Advances and Future Prospects; Razzaghi-Abyaneh, M., Ed.; InTech: Rijeka, Croatia, 2013; pp. 23–39. [Google Scholar]
- Wang, L.; Wu, J.; Liu, Z.; Shi, Y.; Liu, J.; Xu, X.; Hao, S.; Mu, P.; Deng, F.; Deng, Y. Aflatoxin B1 degradation and detoxification by Escherichia coli CG1061 isolated from chicken cecum. Front. Pharmacol. 2019, 9, 1548. [Google Scholar] [CrossRef] [PubMed]
- Taylor, M.C.; Jackson, C.J.; Tattersall, D.B.; French, N.; Peat, T.S.; Newman, J. Identification, and characterization of two families of F420H2-dependent reductases from Mycobacteria that catalyse aflatoxin degradation. Mol. Microbiol. 2010, 78, 561–575. [Google Scholar] [CrossRef] [PubMed]
- Li, C.H.; Li, W.Y.; Hsu, I.N.; Liao, Y.Y.; Yang, C.Y.; Taylor, M.C.; Liu, Y.F.; Huang, W.H.; Chang, H.H.; Huang, H.L.; et al. Recombinant aflatoxin-degrading F420H2-dependent reductase from Mycobacterium smegmatis protects mammalian cells from aflatoxin toxicity. Toxins 2019, 11, 259. [Google Scholar] [CrossRef]
- Zhao, L.H.; Guan, S.; Gao, X.; Ma, Q.G.; Lei, Y.P.; Bai, X.M. Preparation, purification, and characteristics of an aflatoxin degradation enzyme from Myxococcus fulvus ANSM068. J. Appl. Microbiol. 2011, 110, 147–155. [Google Scholar] [CrossRef]
- Gong, A.-D.; Wu, N.-N.; Kong, X.-W.; Zhang, Y.-M.; Hu, M.-J.; Gong, S.-J.; Dong, F.-Y.; Wang, J.-H.; Zhao, Z.-Y.; Liao, Y.-C. Inhibitory effect of volatiles emitted from Alcaligenes faecalis N1-4 on Aspergillus flavus and aflatoxins in storage. Front. Microbiol. 2019, 10, 1–12. [Google Scholar] [CrossRef]
- Gong, A.-D.; Dong, F.-Y.; Hu, M.-J.; Kong, X.-W.; Wei, F.-F.; Gong, S.-J.; Zhang, Y.-M.; Zhang, J.-B.; Wu, A.-B. Antifungal activity of volatile emitted from Enterobacter asburiae Vt-7 against Aspergillus flavus and aflatoxins in peanuts during storage. Food Control 2019, 106, 106718. [Google Scholar] [CrossRef]
- Lyu, A.; Yang, L.; Wu, M.; Zhang, J.; Li, G. High efficacy of the volatile organic compounds of Streptomyces yanglinensis 3-10 in suppression of Aspergillus contamination on peanut kernels. Front. Microbiol. 2020, 11, 142. [Google Scholar] [CrossRef]
- Zhang, H.; Kim, M.S.; Krishnamachari, V.; Payton, P.; Sun, Y.; Grimson, M.; Farag, M.A.; Ryu, C.-M.; Allen, R.; Melo, I.S.; et al. Rhizobacterial volatile emissions regulate auxin homeostasis and cell expansion in Arab. Planta 2007, 226, 839–851. [Google Scholar] [CrossRef] [PubMed]
- Kanchiswamy, C.N.; Malnoy, M.; Maffei, M.E. Bioprospecting bacterial and fungal volatiles for sustainable agriculture. Trends Plant Sci. 2015, 20, 206–211. [Google Scholar] [CrossRef]
- Fincheira, P.; Quiroz, A. Microbial volatiles as plant growth inducers. Microbiol. Res. 2018, 208, 63–75. [Google Scholar] [CrossRef]
- Intanoo, M.; Kongkeitkajorn, M.B.; Suriyasathaporn, W.; Phasuk, Y.; Bernard, J.K.; Pattarajinda, V. Effect of supplemental Kluyveromyces marxianus and Pichia kudriavzevii on aflatoxin M1 excretion in milk of lactating dairy cows. Animals 2020, 10, 709. [Google Scholar] [CrossRef] [PubMed]
- Jouany, J.-P.; Morgavi, D. Use of “natural” products as alternative to antibiotic feed additives in ruminant production. Animal 2007, 1, 1443–1466. [Google Scholar] [CrossRef] [PubMed]
- Chaucheyras-Durand, F.; Walker, N.; Bach, A. Effects of active dry yeasts on the rumen microbial ecosystem: Past, present, and future. Anim. Feed Sci. Tech. 2008, 145, 5–26. [Google Scholar] [CrossRef]
- Shetty, P.H.; Jespersen, L. Saccharomyces cerevisiae and lactic acid bacteria as potential mycotoxin decontaminating agents. Trends Food Sci. Technol. 2006, 17, 48–55. [Google Scholar] [CrossRef]
- Xiong, J.L.; Wang, Y.M.; Nennich, T.D.; Li, Y.; Liu, J.X. Transfer of dietary aflatoxin B1, to milk aflatoxin M1, and effect of inclusion of adsorbent in the diet of dairy cows. J. Dairy Sci. 2015, 98, 2545–2554. [Google Scholar] [CrossRef]
- Diaz, D.E.; Hagler, W.M.; Blackwelder, J.T.; Eve, J.A.; Hopkins, B.A.; Anderson, K.L.; Jones, F.T.; Whitlow, L.W. Aflatoxin binders II: Reduction of aflatoxin M1 in milk by sequestering agents of cows consuming aflatoxin in feed. Mycopathologia 2004, 157, 233–241. [Google Scholar] [CrossRef]
- Queiroz, O.C.M.; Han, J.H.; Staples, C.R.; Adesogan, A.T. Effect of adding a mycotoxin-sequestering agent on milk aflatoxin M1 concentration and the performance and immune response of dairy cattle fed an aflatoxin B1-contaminated diet. J. Dairy Sci. 2012, 95, 5901–5908. [Google Scholar] [CrossRef]
- Anjos, F.R.D.; Ledoux, D.R.; Rottinghaus, G.E.; Chimonyo, M. Efficacy of Mozambican bentonite and diatomaceous earth in reducing the toxic effects of aflatoxins in chicks. World Mycotoxin J. 2016, 9, 63–72. [Google Scholar] [CrossRef]
- Hua, S.S.T.; Beck, J.J.; Sarreal, S.B.L.; Gee, W. A major volatile compound 2-phenylethanol from the biocontrol yeast, Pichia anomala, inhibits growth and expression of aflatoxin biosynthesis genes of Aspergillus flavus. Mycotoxin Res. 2014, 30, 71–78. [Google Scholar] [CrossRef] [PubMed]
- Çelýk, K.; Denlý, M.; Savas, T. Reduction of toxic effects of aflatoxin B1 by using baker yeast (Saccharomyces cerevisiae) in growing broiler chicks’ diets. Revista brasileira de zootecnia 2003, 32, 615–619. [Google Scholar] [CrossRef]
- Santin, E.; Paulillo, A.C.; Maiorka, A.; Nakaghi, L.S.O.; Macari, M.; Silva, A.V.F.; Alessi, A.C. Evaluation of the efficacy of Saccharomyces cerevisiae cell wall to ameliorate the toxic effects of aflatoxin in broilers. Int. J. Poult. Sci. 2003, 2, 341–344. [Google Scholar]
- Armando, M.R.; Dogi, C.A.; Pizzolitto, R.P.; Escobar, F.; Peirano, M.S.; Salvano, M.A.; Sabini, L.I.; Combina, M.; Dalcero, A.M.; Cavaglieri, L.R. Saccharomyces cerevisiae strains from animal environment with in vitro aflatoxin B1 binding ability and anti-pathogenic bacterial influence. World Mycotoxin J. 2011, 4, 59–68. [Google Scholar] [CrossRef]
- Dogi, C.A.; Armando, R.; Luduena, R.; de Moreno de LeBlanc, A.; Rosa, C.A.; Dalcero, A.; Cavaglieri, L. Saccharomyces cerevisiae strains retain their viability and aflatoxin B1 binding ability under gastrointestinal conditions and improve ruminal fermentation. Food Addit. Contam. Part A Chem. Anal. Control Expo. Risk Assess. 2011, 28, 1705–1711. [Google Scholar]
- Pizzolitto, R.P.; Armando, M.R.; Combina, M.; Cavaglieri, L.R.; Dalcero, A.M.; Salvano, M.A. Evaluation of Saccharomyces cerevisiae strains as probiotic agent with aflatoxin B1 adsorption ability for use in poultry feedstuffs. J. Environ. Sci. Health B 2012, 47, 933–941. [Google Scholar] [CrossRef]
- Kim, S.; Lee, H.; Lee, S.; Lee, J.; Ha, J.; Choi, Y.; Yoon, Y.; Choi, K.-Y. Microbe-mediated aflatoxin decontamination of dairy products and feeds. J. Dairy Sci. 2017, 100, 871–880. [Google Scholar] [CrossRef]
- Shetty, P.H.; Hald, B.; Jespersen, L. Surface binding of aflatoxin B1 by Saccharomyces cerevisiae strains with potential decontaminating abilities in indigenous fermented foods. Int. J. Food Microbiol. 2007, 113, 41–46. [Google Scholar] [CrossRef]
- Inoue, T.; Nagatomi, Y.; Uyama, A.; Naoki, M. Degradation of aflatoxin B1 during the fermentation of alcoholic beverages. Toxins 2013, 5, 1219–1229. [Google Scholar] [CrossRef]
- Foroughi, M.; Sarabi Jamab, M.; Keramat, J.; Foroughi, M. Immobilization of Saccharomyces cerevisiae on perlite beads for the decontamination of aflatoxin M1 in milk. J. Food Sci. 2018, 83, 2008–2013. [Google Scholar] [CrossRef] [PubMed]
- Saki, A.; Rahmani, A.; Mahmoudi, H.; Tabatabaei, M.M.; Zamani, P.; Khosravi, A.R. The ameliorative effect of Mycosorb in aflatoxin contaminated diet of broiler chickens. J. Livest. Sci. Technol. 2018, 6, 39–47. [Google Scholar]
- Haidukowski, M.; Casamassima, E.; Cimmarusti, M.T.; Branà, M.T.; Longobardi, F.; Acquafredda, P.; Logrieco, A.; Altomare, C. Aflatoxin B1-adsorbing capability of Pleurotus eryngii mycelium: Efficiency and modeling of the process. Front. Microbiol. 2019, 10, 1386. [Google Scholar] [CrossRef] [PubMed]
- Branà, M.T.; Sergio, L.; Haidukowski, M.; Logrieco, A.F.; Altomare, C. Degradation of aflatoxin B1 by a sustainable enzymatic extract from spent mushroom substrate of Pleurotus eryngii. Toxins 2020, 12, 49. [Google Scholar] [CrossRef]
- Motomura, M.; Toyomasu, T.; Mizuno, K.; Shinozawa, T. Purification and characterization of an aflatoxin degradation enzyme from Pleurotus ostreatus. Microbiol. Res. 2003, 158, 237–242. [Google Scholar] [CrossRef]
- Wang, J.; Ogata, M.; Hirai, H.; Kawagishi, H. Detoxification of aflatoxin B1 by manganese peroxidase from the white-rot fungus Phanerochaete sordida YK-624. FEMS Microbiol. Lett. 2011, 314, 164–169. [Google Scholar] [CrossRef]
- Wu, Y.Z.; Lu, F.P.; Jiang, H.L.; Tan, C.P.; Yao, D.S.; Xie, C.F. The furofuran-ring selectivity, hydrogen peroxide-production and low Km value are the three elements for highly effective detoxification of aflatoxin oxidase. Food Chem. Toxicol. 2015, 76, 125–131. [Google Scholar] [CrossRef]
- Delgado, J.; Owens, R.A.; Doyle, S.; Asensio, M.A.; Núñez, F. Impact of the antifungal protein PgAFP from Penicillium chrysogenum on the protein profile in Aspergillus flavus. Appl. Microbiol. Biotechnol. 2015, 99, 8701–8715. [Google Scholar] [CrossRef]
- Delgado, J.; Owens, R.A.; Doyle, S.; Núñez, F.; Asensio, M.A. Quantitative proteomics reveals new insights into calcium-mediated resistance mechanisms in Aspergillus flavus against the antifungal protein PgAFP in cheese. Food Microbiol. 2017, 66, 1–10. [Google Scholar] [CrossRef]
- Leiter, É.; Gáll, T.; Csernoch, L.; Pócsi, I. Biofungicide utilizations of antifungal proteins of filamentous ascomycetes: Current and foreseeable developments. BioControl 2017, 62, 125–138. [Google Scholar] [CrossRef]
- Ahlberg, S.; Randolph, D.; Okoth, S.; Lindahl, J. Aflatoxin binders in foods for human consumption-can this be promoted safely and ethically? Toxins 2019, 11, 410. [Google Scholar] [CrossRef] [PubMed]
- Peltonen, K.; EI-Nezami, H.; Haskard, C.; Ahokas, J.; Salminen, S. Aflatoxin B1 binding by dairy strains of lactic acid bacteria and bifidobacteria. J. Dairy Sci. 2001, 84, 2152–2156. [Google Scholar] [CrossRef]
- Haskard, C.A.; El-Nezami, H.S.; Kankaanpaa, P.E.; Salminen, S.; Ahokas, J.T. Surface binding of aflatoxin B1 by lactic acid bacteria. Appl. Environ. Microbiol. 2001, 67, 3086–3091. [Google Scholar] [CrossRef] [PubMed]
- Azab, R.M.; Tawakkol, W.M.; Hamad, A.R.M.; Abou-Elmagd, M.K.; El-Agrab, H.M.; Refai, M.K. Detection and estimation of aflatoxin B1 in feeds and its biodegradation by bacteria and fungi. Egypt. J. Nat. Toxins 2005, 2, 39–54. [Google Scholar]
- Marrez, D.A.; Shahy, E.M.; El-Sayed, H.S.; Sultan, Y.Y. Detoxification of aflatoxin B1 in milk using lactic acid bacteria. J. Biol. Sci. 2018, 18, 144–151. [Google Scholar] [CrossRef]
- Liew, W.P.P.; Nurul-Adilah, Z.; Than, L.T.L.; Mohd-Redzwan, S. The binding efficiency and interaction of Lactobacillus casei Shirota toward aflatoxin B1. Front. Microbiol. 2018, 9, 1503. [Google Scholar] [CrossRef]
- Fazeli, M.R.; Hajimohammadali, M.; Moshkani, A.; Samadi, N.; Jamalifar, H.; Khoshayand, M.R. Aflatoxin B1 binding capacity of autochthonous strains of lactic acid bacteria. J. Food Protect. 2009, 72, 189–192. [Google Scholar] [CrossRef]
- Ghazvini, R.D.; Kouhsari, E.; Zibafar, E.; Hashemi, S.J.; Amini, A.; Niknejad, F. Antifungal activity and aflatoxin degradation of Bifidobacterium bifidum and Lactobacillus fermentum against toxigenic Aspergillus parasiticus. Open Microbiol. J. 2016, 10, 1–5. [Google Scholar] [CrossRef]
- EI-Nezarni, H.; Mykkiinen, H.; Kankaanpi, H.; Salminen, S.; Ahokas, J. Ability of Lactobacillus and Propionibacterium strains to remove aflatoxin B1 from the chicken duodenum. J. Food Protect. 2000, 63, 549–552. [Google Scholar]
- Shahin, A.A.M. Removal of aflatoxin B1 from contaminated liquid media by dairy lactic acid bacteria. Int. J. Agric. Biol. 2007, 9, 71–75. [Google Scholar]
- Kabak, B.; Var, I. Factors affecting the removal of aflatoxin M1 from food model by Lactobacillus and Bifidobacterium strains. J. Environ. Sci. Health B 2008, 43, 617–624. [Google Scholar] [CrossRef] [PubMed]
- Bovo, F.; Corassin, C.H.; Rosim, R.E.; Oliveira, C.A.F. Efficiency of lactic acid bacteria strains for decontamination of aflatoxin M1 in phosphate buffer saline solution and in skimmed milk. Food Bioproc. Technol. 2013, 6, 2230–2234. [Google Scholar] [CrossRef]
- Sarimehmetoglu, B.; Küplülü, Ö. Binding ability of aflatoxin M1 to yoghurt bacteria. Vet. Fak. Derg. 2004, 51, 195–198. [Google Scholar]
- Petchkongkaew, A.; Taillandier, P.; Gasaluck, P.; Lebrihi, A. Isolation of Bacillus spp. from Thai fermented soybean (Thuanao): Screening for aflatoxin B1 and ochratoxin A detoxification. J. Appl. Microbiol. 2008, 104, 1495–1502. [Google Scholar] [CrossRef]
- Smith, J.E.; Harran, G. Microbial degradation of mycotoxins. Intern. Biodeter. Biodeg. 1993, 32, 205–211. [Google Scholar] [CrossRef]
- Gao, X.; Ma, Q.; Zhao, L.; Lei, Y.; Shan, Y.; Cheng, J. Isolation of Bacillus subtilis: Screening for aflatoxins B1, M1, and G1 detoxication. Eur. Food Res. Technol. 2011, 232, 957–962. [Google Scholar] [CrossRef]
- Guan, S.; Ji, C.; Zhou, T.; Li, J.; Ma, Q.; Niu, T. Aflatoxin B1 degradation by Stenotrophomonas maltophilia and other microbes selected using coumarin medium. Int. J. Mol. Sci. 2008, 9, 1489–1503. [Google Scholar] [CrossRef]
- Teniola, O.D.; Addo, P.A.; Brost, I.M.; Farber, P.; Jany, K.D.; Alberts, J.F.; van Zyl, W.H.; Steyn, P.S.; Holzapfel, W.H. Degradation of aflatoxin B1 by cell-free extracts of Rhodococcus erythropolis and Mycobacterium fluoranthenivorans sp. nov. DSM44556(T). Int. J. Food Microbiol. 2005, 105, 111–117. [Google Scholar] [CrossRef]
- Smiley, R.D.; Draughon, F.A. Preliminary evidence that degradation of aflatoxin B1 by Flavobacterium aurantiacum is enzymatic. J. Food Protect. 2000, 63, 415–418. [Google Scholar] [CrossRef]
- D’Souza, D.H.; Brackett, R.E. Aflatoxin B1 degradation by Flavobacterium aurantiacum in the presence of reducing conditions and seryl and sulfhydryl group inhibitors. J. Food Protect. 2001, 64, 268–271. [Google Scholar] [CrossRef]
- Sangare, L.; Zhao, Y.; Folly, Y.M.; Chang, J.; Li, J.; Selvaraj, J.N.; Xing, F.; Zhou, L.; Wang, Y.; Liu, Y. Aflatoxin B1 degradation by a Pseudomonas strain. Toxins 2014, 6, 3028–3040. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Li, W.; Yang, W.; Li, H.; Liu, X.; Cao, Y. Isolation and characterisation of an aflatoxin B1-degrading bacterium. Wei Sheng Wu Xue Bao 2012, 52, 1129–1136. [Google Scholar] [PubMed]
- Kong, Q.; Zhai, C.; Guan, B.; Li, C.; Shan, S.; Yu, J. Mathematic modeling for optimum conditions on aflatoxin B1 degradation by the aerobic bacterium Rhodococcus erythropolis. Toxins 2012, 4, 1181–1195. [Google Scholar] [CrossRef]
- Eshelli, M.; Harvey, L.; Edrada-Ebel, R.; McNeil, B. Metabolomics of the bio-degradation process of aflatoxin B1 by actinomycetes at an initial pH of 6.0. Toxins 2015, 7, 439–456. [Google Scholar] [CrossRef] [PubMed]
- Cserhati, M.; Kriszt, B.; Krifaton, C.; Szoboszlay, S.; Hahn, J.; Toth, S.; Nagy, I.; Kukolya, J. Mycotoxin-degradation profile of Rhodococcus strains. Int. J. Food Microbiol. 2013, 166, 176–185. [Google Scholar] [CrossRef]
- García-Béjar, B.; Arévalo-Villena, M.; Guisantes-Batan, E.; Rodríguez-Flores, J.; Briones, A. Study of the bioremediatory capacity of wild yeasts. Sci. Rep. 2020, 10, 11265. [Google Scholar] [CrossRef]
- Zohri, A.A.; Abdel-Kareem, M.M. Four strains of yeasts: As effective biocontrol agents against both growth and mycotoxins formation by selected 11 toxigenic fungi. Glob. Adv. Res. J. Microbiol. 2018, 7, 132–135. [Google Scholar]
- Niknejad, F.; Zaini, F.; Faramarzi, M.; Amini, M.; Kordbacheh, P.; Mahmoudi, M.; Safara, M. Candida parapsilosis as a potent biocontrol agent against growth and aflatoxin production by Aspergillus species. Iran. J. Public Health 2012, 41, 72–80. [Google Scholar]
- Ghanbari, R.; Rezaie, S.; Noorbakhsh, F.; Khaniki, G.J.; Soleimani, M.; Aghaee, E.M. Biocontrol effect of Kluyveromyces lactis on aflatoxin expression and production in Aspergillus parasiticus. FEMS Microbiol. Lett. 2019, 366, fnz114. [Google Scholar] [CrossRef]
- Bueno, D.; Casale, C.H.; Pizzolitto, R.P.; Salano, M.A.; Olivier, G. Physical adsorption of aflatoxin B1 by lactic acid bacteria and Saccharomyces cerevisiae: A theoretical model. J. Food Protect. 2007, 70, 2148–2154. [Google Scholar] [CrossRef]
- Corassin, C.H.; Bovo, F.; Rosim, R.E.; Oliveira, C.A.F. Efficiency of Saccharomyces cerevisiae and lactic acid bacteria strains to bind aflatoxin M1 in UHT skim milk. Food Control 2013, 31, 80–83. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).