Abstract
Clostridium perfringens poses a serious threat to small ruminants by causing moderate to severe enterotoxaemia. Due to its ability to produce a wide arsenal of toxins, it is ranked among the most prevalent and important pathogens in livestock. This study focused on the molecular characterization of different Clostridium perfringens types along with their antimicrobial resistance profile. An overall higher prevalence of C. perfringens (46.1%) was detected based on mPCR among sheep and goats (healthy and diseased) in the Punjab province, Pakistan. The majority of the isolates were characterized as type A (82%), followed by type D (18%). Among the isolates from diseased sheep and goats, 27% were positive for cpa, 49% for cpa and cpb2, 9% for cpa and etx, 15% for cpa, cpb2 and etx. In the case of isolates from healthy sheep and goats, 59% were positive for cpa, 34% for cpb2 and cpa, 4% for cpa and etx, and 3% for cpa, cpb2 and etx. The prevalence of the beta2 toxin gene in the diseased sheep and goat population was 64% as compared to 37% in healthy animals. All 184 isolates (100%) were sensitive to rifampin and ceftiofur; the majority (57%) was sensitive to teicoplanin, chloramphenicol, amoxicillin, linezolid and enrofloxacin. A lower proportion of isolates (43%) were sensitive to ciprofloxacin and only 14% were susceptible to erythromycin. The findings of this study highlight the higher prevalence of C. perfringens in small ruminants and indicate that detailed pathogenesis studies are necessary to understand the explicit role of various toxins in causing enteric infections in sheep and goats including how they might be exploited to develop vaccines against these diseases.
Key Contribution:
This is the comprehensive study on phenotypic and genotypic characterization of C. perfringens strains in the relatively novel geographic location. Prevalence was described for the cpb2 gene (significantly enriched in isolates originating from diseased animals) and a few type D isolates from healthy sheep and goats. Besides this, there is some useful information regarding antibiotics resistance profile of these isolates, notably revealing higher intrinsic resistance against tetracycline.
1. Introduction
The livestock industry is one of the fastest growing and financially important sub-sectors of the economy in developing countries like Pakistan, accounting for a 60.54% value addition to the Pakistani agricultural sector and contributing 11.22% to GDP. Furthermore, it makes a significant contribution to rural economic growth, where 40% of the population is dependent on livestock [1]. Small ruminants, especially sheep and goats, are required for the subsistence of poor livestock owners and are a main source of meat, milk, manure and skin [2]. Unfortunately, over the last decade, the caprine and ovine industry has significantly declined due to infectious diseases [3]. Clostridium perfringens is one of the main pathogens responsible for intestinal infections as well as histotoxic diseases in animals, having a low incidence rate (2–8%) but a very high case fatality rate (100%) [4,5]. It is a gram positive, anaerobic, spore-forming microorganism that produces more than 20 extracellular toxins and hydrolytic enzymes, of which six are currently used for typing [6]. The toxins form diverse combinations and can be divided into types A–G. All types produce α-toxin; in addition, type B produces β- and ε-toxin, type C produces β-toxin, type D produces ε-toxin, type E produces ι-toxin, type F produces enterotoxin and type G produces netB toxin [7,8]. All these potent toxins can act locally or are absorbed in the intestine causing enteric infections, which are generically named as enterotoxemia in sheep and goats. However, the severity of these infections is dependent on the type and combination of various toxins, of which epsilon toxin (type D) is the most frequent cause of enterotoxemia in sheep and goats [9]. C. perfringens type A is involved in acute enterotoxemia in the goat kids [10]. Type D is associated with “pulpy kidney disease” and dysentery in sheep [11]. Epsilon toxin, present in C. perfringens type B and D, is mostly involved in disease pathogenesis [7]. Epsilon toxin in goats causes enterocolitis, but in sheep it can also cause abnormal effects on the brain and lungs. The difference in pathogenesis between the two species is due to the different mechanisms by which epsilon toxin modifies water and ion transport in the intestines of sheep and goats [12,13,14]. In developed countries, C. perfringens type F food poisoning is a commonly reported food-borne illness [15]. Strains carrying the CPE gene (cpe), isolated from both humans and animals, produce spores in the intestinal tract. These spores produce enterotoxin (CPE), which is responsible for symptoms such as diarrhea [16,17].
Although C. perfringens can exist as part of the normal intestinal microflora, under certain conditions where the physiological balance of the intestine is altered, abnormal proliferation of this microorganism can result in the production of a wide variety of toxins [18]. These toxins either act locally or are absorbed into the general circulation, resulting in severe effects on the host [19]. Accurate typing of the toxin is very important in disease diagnosis [20]. Traditionally, typing of C. perfringens strains has been performed using a toxin neutralization test with the appropriate antisera in laboratory animals such as guinea pigs [21]. An ELISA (enzyme-linked immunosorbent assay) based method can also be used for typing C. perfringens isolates. However, these methods are often time-consuming and are not always able to identify a range of different toxins, thereby limiting subtyping options [22]. In addition, special culture methods are required to induce sporulation, which aids in the identification of enterotoxin producing strains [23]. Genotyping of C. perfringens isolates, however, can overcome this problem and has become the standard for toxinotyping of C. perfringens. Polymerase chain reaction (PCR) protocols, especially multiplex PCR assay, have been established for simultaneous detection of these toxin genes (cpa, cpb, etx, iap, cpe and cpb2) [24,25,26,27].
Although vaccines and different antimicrobial agents are used to control enterotoxaemia [28], the extensive use of these agents to increase the growth rate of livestock animals and treat gastrointestinal infections likely serves as a way of transmitting antimicrobial-resistant genes or microbes into the human food chain [29,30]. The emerging problem of antimicrobial resistance between pathogenic and commensal bacteria is also of concern [31]. There are very few reports on the antibiogram of C. perfringens isolates recovered from sheep and goats in Pakistan. More comprehensive epidemiological information pertaining to C. perfringens type A and D in these animals is also lacking. As such, the aim of the present study was the isolation, phenotypic, genotypic and molecular typing of C. perfringens type A and D isolates from diseased and healthy sheep and goats in the Punjab province of Pakistan, as well as the analysis of their antimicrobial susceptibility patterns.
2. Results
2.1. Isolation and Identification of C. perfringens
A total of 201 samples (50.31%) from both sheep and goats were found positive and exhibited typical black colonies on tryptose sulphite cycloserine agar plates. Most of isolates showed the presence of Gram-positive rods and possessed subterminal spores. On blood agar, characteristic beta hemolytic colonies with double zone of hemolysis were observed. The microorganism produced an opaque halo around colonies on egg yolk agar, confirming it as C. perfringens whereas most other Clostridia are reported to be lecithinase negative. All culture positive isolates were further resolved through biochemical differential tests and sugar fermentation reactions. Almost all isolates fermented glucose, maltose, sucrose, lactose, mannose and trehalose, while arabinose, cellobiose, mannitol, melezitose, salicin, xylose, sorbitol and rhamnose were not fermented by C. perfringens isolates (Table 1). Two sugars behaved differently in different isolates. Glycerol was fermented by one group of isolates while the other group did not ferment it. Variations were also seen in the case of raffinose sugar, i.e., most of the isolates did not ferment this sugar while only a few fermented it. There were also slight variations in the biochemical properties of different isolates. All isolates hydrolyzed gelatin and were negative for urease production. In addition, very few (2%) isolates hydrolyzed esculin. The indole test was negative in all of the isolates; however, there was one isolate that behaved differently (i.e., indole was positive). Biochemical characterization, therefore, differentiated the isolates into four major groups; these are given in Table 2. It was found that 195 (48.9%) culture isolates belong to C. perfringens on the basis of biochemical characterization.

Table 1.
Carbohydrate fermentation reactions of C. perfringens field isolates. These isolates were placed in groups on the basis of variation in the fermentation reactions of sugars by different isolates.

Table 2.
Biochemical characteristics of C. Perfringens isolates.
2.2. Toxinotyping of Recovered C. perfringens Isolates
All 195 biochemically characterized isolates of C. perfringens were examined for alpha, beta, epsilon, iota, beta2 and enterotoxin gene by using the multiplex PCR technique. The alpha toxin gene (cpa) was found to be present in 184 out of 399 (46.1%) isolates by simple PCR. These 184 isolates were further genotyped using mPCR to further identify them as either type A or type D; and categorized on the basis of presence of major toxin genes. Type A isolates had an alpha gene while type D isolates had alpha and epsilon genes. Neither beta nor iota genes were present in any of the genotyped isolates. In addition to alpha and epsilon genes, beta2 was also found in both type A and type D isolates. Eighty-two percent of the isolates from sheep and goats belonged to C. perfringens Type A while 18% of the isolates were C. perfringens type D (Figure 1).
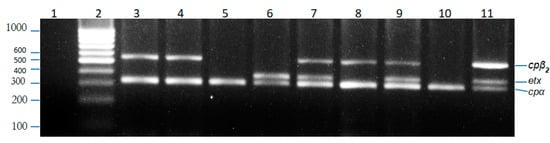
Figure 1.
Agarose gel electrophoresis of PCR products. Lanes: 1, negative control (no template DNA); 2, 1000 bp ladder; 3,4,8, C. perfringens isolate (genotype A, β2); 5,10, C. perfringens isolate (genotype A); 6, C. perfringens isolate (genotype D); 7,9, C. perfringens isolate (genotype D, β2); 11, Positive control.
Among the isolates from healthy sheep and goats, 59% were positive for cpa, 34% for cpb2 and cpa, 4% for cpa and etx, and 3% for cpa, cpb2 and etx. In the case of isolates from diseased sheep and goats, 27% were positive for cpa, 49% for cpa and cpb2, 9% for cpa and etx, and 15% for cpa, cpb2 and etx. Therefore, the percentage of isolates that were positive for cpb2 was significantly higher in the diseased population (64%) of sheep and goats as compared to the healthy population (37%). As far as the presence of etx gene was concerned, 24% of isolates from the diseased population had the etx gene, compared to 7% from the healthy population. The percentage of cpa and cpb2-positive isolates from diseased animals (49%) was significantly higher than that of healthy animals (34%). No isolates from both populations were found to carry cpb, iap and cpe (Table 3 and Table 4).

Table 3.
Multiplex PCR results for the C. perfringens isolates (sheep and goat).

Table 4.
Multiplex PCR results for the C. perfringens isolates (healthy and diseased).
2.3. Antimicrobial Susceptibility Patterns
A total of 184 C. perfringens type A and D isolates confirmed by mPCR were used for in vitro antimicrobial susceptibility testing (Figure 2). All the C. perfringens isolates showed 100% susceptibility to rifampin and ceftiofur, while teicoplanin and enrofloxacin showed “significant susceptibility” against 57% of the C. perfringens isolates. Teicoplanin showed “good susceptibility” against 29% of the isolates while enrofloxacin showed good susceptibility against 43% of the isolates of C. perfringens. Chloramphenicol, amoxicillin and linezolid showed significant susceptibility against 57% of C. perfringens isolates. Erythromycin showed significant susceptibility to 14% isolates while good susceptibility was observed against 72% of the isolates. Norfloxacin showed “moderate susceptibility” against 100% of the isolates. Ciprofloxacin showed significant susceptibility against 43% of the isolates while 29% and 28% of the isolates showed good and moderate susceptibility, respectively. All C. perfringens isolates showed 100% resistance against neomycin and 72% resistance to tetracycline as shown in Table 5.
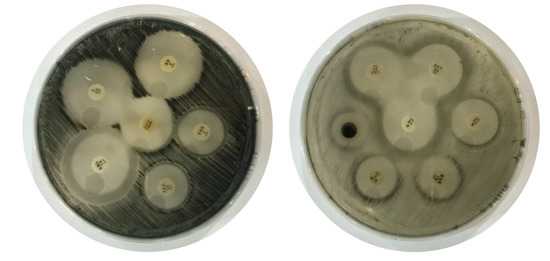
Figure 2.
Antibiotic susceptibility testing of Clostridium perfringens against different drugs.

Table 5.
Antibiotic susceptibility testing of Clostridium perfringens against different antibiotics.
3. Discussion
Clostridium perfringens ranks among the most important pathogens in livestock and humans, and it causes both histotoxic disease and infection originating in the intestines. Toxins produced by C. perfringens types are responsible for enteric diseases in sheep and goats and it has been suggested that economic losses due to C. perfringens infections may be a result of all seven types of the bacterium [32,33]. There may be geographical differences in the prevalent types of the bacterium; also types may vary depending on the animal species in the area [34]. In the present study, the biochemical results were similar to previously reported findings, with few exceptions [35,36]. Very few isolates hydrolyzed esculin and glycerol sugar was weakly fermented in some isolates; in some cases there was no fermentation. Raffinose was only fermented in some isolates [37]. A variation was noted during indole production reaction, although only a single isolate was found as indole positive. This variation is significant as it has not been previously reported in the scientific literature. A study conducted in 1942, however, supports this finding [38].
In this study, C. perfringens type A was present in 82% of samples collected from both healthy and diseased sheep and goats. Different studies have reported C. perfringens type A in 51–87% of samples taken from soil and healthy ruminants [39,40,41]. C. perfringens type A has also been reported as the most prevalent type in North America [42]. Another study conducted in Saudi Arabia indicated that C. perfringens type A was the most prevalent type in sheep, calves and chicken [43]. Unfortunately, there are very few authentic reports citing the prevalence of C. perfringens in Pakistani sheep and goat populations. C. perfringens type D causes diseases in numerous animal species; however, it is more prevalent in sheep and goats. The results of our study indicated that C. perfringens type D is prevalent in both healthy and diseased animals. About 18% of isolates from both healthy and diseased sheep and goats belong to C. perfringens type D. In ruminants, this type is normally present in the intestine even though different studies have isolated this type in less than 20% of the animals and on some farms, no type D microorganism was found at all [41]. In this study, 5 type D isolates out of 68 (7%) samples were found in healthy animals. Of these 5 samples, 2 were isolated from goat and the other 3 isolates belonged to sheep. Type E infections in domestic animals were considered as a rare occurrence. Likewise, no isolates in our study from healthy or diseased animals were found to be C. perfringens type E. In numerous epidemiological studies, C. perfringens types possessing the cpb2 gene have been isolated from cattle, horses, pigs, sheep, goats, poultry, fish, carnivores, domestic wildlife species and humans [44,45]. In this study, the prevalence of beta2 toxin gene was found to be significantly higher in diseased sheep and goats. In diseased animals, the cpb2 gene was found in 64% of the isolates, compared to 37% in healthy animals. The high prevalence of the cpb2 gene in isolates from animals with enteric problems is also consistent with previous studies from different parts of the world [46].
The investigations carried out in control group (healthy sheep and goats) revealed that the most prevalent type in these animals was type A (93%). This is because they have cpa only, while the other 7% of isolates have epsilon toxin gene in addition to the alpha gene, thus, they belong to type D. In one study conducted in Turkey, 61 C. perfringens isolates from healthy sheep were examined by multiplex PCR; of these, 95% belonged to type A and the remaining 5% were type D [47]. These results are similar to those found in previous studies [48,49,50]. A study investigating the predominant flora of ostrich intestine reported type A as the most prevalent type, while in the investigation of fecal samples of diseased (Enteritis, Enterotoxemia) sheep and goats, C. perfringens type A was found in 76% of isolates. Similarly, a study conducted in 2012 in Saudi Arabia reported type A in 73.5% of the genotyped isolates [51]. Type A has also been reported as a predominant type in sheep with enterotoxemia [47]. Based on our findings, besides type A, C. perfringens type D was found in 24% of isolates from diseased sheep and goats. Reports from various studies around the world have found that the prevalence of type D enterotoxemia ranges from 24.13–100% [32]. In Turkey, the prevalence of type D enterotoxemia in diseased animals was reported to be from 38.63–50% [49].
The present study ascertained the distribution of C. perfringens toxin types in sheep and goat populations in relation to health status. The predominance of the alpha toxin gene (C. perfringens type A) is in accordance with the high distribution of this toxin type in neighboring countries like China, India and Bangladesh [52,53]. The high prevalence of alpha and beta2 genes also signifies the need to incorporate these strains into the vaccine that is currently used against C. perfringens infections, which contains only type D strain, and also suggests the use of an oil based/montanide adjuvanted vaccine, which was proven successful against other clinically important diseases of livestock [54]. In addition, this study is the first of its kind to report the presence of epsilon toxin gene (etx) in the feces of healthy sheep and goats raised in Pakistan.
In Pakistan, there are very few reports on the antibiotic resistance patterns of C. perfringens type A and D isolated from an animal source. The present study revealed that all isolates were found to be susceptible to penicillin, as mentioned in different studies reported from other countries [55,56]. In addition to penicillin, all isolates were also found to be susceptible to rifampin and ceftiofur. These results were in agreement to those reported in Brazil and Thailand [57,58]. Amoxicillin and enrofloxacin showed high in vitro activity against 57% of isolates and 60–70% inhibition against the remaining 43% of isolates. Another study showed that all isolates from commercial turkeys were highly susceptible to amoxicillin and enrofloxacin [55]. The teicoplanin activity was also found to be in accordance with the results described against intestinal anaerobic bacteria [59].
Norfloxacin, neomycin and tetracycline were least active against C. perfringens isolates from sheep and goats. The poor efficacy of these antibiotics has also been previously reported in piglets [57]. Reduced susceptibility to tetracycline detected in C. perfringens isolates across all host species is consistent with the findings of several studies from various countries [60,61]. In this study, 72% of isolates were found to be resistant to tetracycline, which is more than previous findings [62]. This higher incidence of tetracycline resistance in C. perfringens isolates is the result of the excessive use of this antibiotic in the sampling areas, often following poor or incorrect veterinary advice. In the present study, linezolid and chloramphenicol showed significant susceptibility against 57% of C. perfringens isolates while ciprofloxacin and erythromycin showed intermediate susceptibility. Previous studies have indicated that almost all isolates were susceptible to linezolid, chloramphenicol while resistance against erythromycin antibiotic was reported for fewer isolates [56,62,63,64].
Our findings showed that tetracycline is a poor treatment choice for C. perfringens-associated infections due to the decreased susceptibility among C. perfringens isolates. Therefore, the use of tetracycline, particularly at low doses, could select for tetracycline resistant strains and result in the transmission of tetracycline resistance.
4. Conclusions
In Pakistan, enteric diseases in small ruminants are causing annual losses of billions of rupees to the sheep and goat industry. The high distribution of toxins in strains from clinically infected sheep and goats also highlights their role in producing diseased conditions. Consequently, these initial findings could be further used for epidemiological investigations, prophylaxis plans and control strategies by formulating appropriate vaccines.
5. Materials and Methods
5.1. Sample Site and Isolation Source
A total n = 399 samples were collected aseptically in sterile plastic bags from sheep and goats in selected districts (Faisalabad, Okara, DG Khan, Muzaffargarh, Bhakkar, Layyah) in the Punjab province, Pakistan over a one year period using a convenient random sampling technique. Fecal samples were collected from healthy and diseased animals following the inclusion and exclusion criteria. Healthy sheep and goats included animals that were assessed as healthy with no history of diarrhea and with normal fecal consistency. In addition, these animals had not received any other medical treatment in the last month. Diseased animals included sheep and goats suffering from enteric disease (having diarrhea or history of diarrhea) which may include fever, abdominal discomfort, lack of appetite and a history of weight loss. Vaccinated animals (alum precipitated enterotoxemia vaccine) were also included according to their health status either in the healthy or diseased group. Animals undergoing any medical treatment in the last month were excluded from the study. Collected samples were transported immediately to the laboratory under refrigerated conditions.
5.2. Isolation of C. perfringens
The fecal samples were diluted in phosphate buffered saline (PBS) 1:10 and placed in a water bath for 10 min at 80 °C to kill the non-spore forming microorganisms. The processed samples were sub-cultured in reinforced clostridial media (RCM) and then inoculated on tryptose sulfite cycloserine agar (TSC) and 5% blood agar plates. The inoculated plates were kept in anaerobic jars at 37 °C for 24 h. Anaerobiosis was created by using an anaerobe sachet. After initial identification based on characteristic colony morphology and Gram staining, the colonies presenting C. perfringens characteristics were streaked on egg yolk agar in order to confirm the lecithinase and lipase activity. Further biochemical characterization was carried out using API 20A kits (bioMérieux, Marcy l’Étoile, France).
5.3. Molecular Typing of C. perfringens Through Multiplex PCR (mPCR)
Following biochemical characterization, isolates found positive for C. perfringens were subjected to molecular typing for identification of the prevalent toxin types. Molecular typing of C. perfringens was performed by multiplex PCR technique. Accordingly, previously developed primers cpa, etx, cpb, iap, cpe and cpb2 (Table 6) were used simultaneously for their ability to identify C. perfringens types [65,66,67].

Table 6.
Oligonucleotide primers for multiplex PCR detection of C. perfringens toxin genes.
Genomic DNA was extracted from all C. perfringens isolates using a manual method [34]. Colonies from 12–18 h pure culture of C. perfringens were suspended in 1 mL distilled water (about 106 cells per mL). Centrifugation was done at 6000× g for 5 min. The supernatant was discarded and cells were resuspended in 200 μL cold Tris EDTA (TE) Buffer. DNA was extracted using EZ-10 Spin Column DNA Gel Extraction Kit (Bio Basic Inc., Toronto, Canada). Multiplex PCR was performed in a thermal cycler with final volume of 25 µL for each reaction mixture. The reaction mixture contained 1.25 U Taq DNA polymerase, 1X PCR Buffer, 4 mM MgCl2, 250 µM dNTPs, 0.12 µM forward and reverse primers of alpha, epsilon, beta, iota and entero gene, 0.16 µM forward and reverse primer of beta2 gene, 2 µL of sample DNA (150–200 ng/µL). The PCR conditions consisted of a pre-denaturation phase at 95 °C for 10 min and 40 cycles of 94 °C for 45 s, 55 °C for 90 s, 72 °C for 90 s followed by 72 °C for 10 min. The PCR product was electrophoresed on 1.5% agarose gel. DNA bands were observed under UV transillumination and photographed using Alpha Imager Mini Imaging System (San Jose, CA, USA).
5.4. Antimicrobial Susceptibility Testing
Antibiotic susceptibility assays were performed according to the disk diffusion method described in [68]. Thirteen different antibiotics were selected on the basis of clinical relevance and veterinary farm practices, belonging to different antimicrobial groups. The antibiotic discs used (Oxoid, Basingstoke, UK) were as follows: amoxicillin (10 µg), ceftiofur (30 µg), chloramphenicol (30 µg), ciprofloxacin (5 µg), enrofloxacin (5 µg), erythromycin (15 µg), linezolid (30 µg), neomycin (10 µg), norfloxacin (10 µg), penicillin G (10 µg), rifampin (5 µg), teicoplanin (30 µg) and tetracycline (30 µg). Perfringens agar medium (Oxoid, UK) was used as a growth medium for C. perfringens. A single colony from a 24 h fresh culture was dispensed in sterilized saline solution by using a sterilized inoculating loop. After thorough mixing, the turbidity of the cultures was compared with standard McFarland solution. If the broth was more turbid, they were adjusted through the addition of normal saline. Bacterial lawn was made on the petri plates by dipping sterile cotton swab in bacterial inoculum. Antibiotic discs were placed on the inoculated plate at an appropriate distance from each other. Penicillin was taken as a positive control. The clear zones of inhibition appeared around the discs of antibiotics after 24 h of incubation at 37 °C.
The diameter of the zone around each antibiotic was measured in millimeters (mm) and calculated as the mean of three replicates. The percentage of inhibition of antibiotics was calculated using the formula described earlier by comparing it with the standard antibiotic (Penicillin) [68,69,70].
5.5. Criteria for Bacterial Inhibition
The bacterial inhibition was carried out for C. perfringens isolates using the criteria mentioned (Table 7).

Table 7.
Inhibition Criteria for C. perfringens isolates.
5.6. Statistical Analysis
The C. perfringens isolates identified from fecal samples of the control group and diseased group of animals were screened by multiplex PCR (mPCR) and the results were presented in a tabularized form. The health status-wise prevalence of C. perfringens types in different groups was compared by Pearson’s chi-square test (χ2) using SPSS software, 22.0 version (IBM, New York, NY, USA).
Author Contributions
Conceptualization, M.M. and Z.I.; methodology, M.M.; validation, M.M. and Z.I.; formal analysis, M.M.; A.S. and A.M.D.; investigation, M.M. and Z.I.; resources, Z.I. and M.S.; data curation, M.M.; writing—original draft preparation, M.M.; A.S.; M.K.F.S.; N.Q. and A.M.D.; writing—review and editing, M.M.; Z.I.; S.L.; M.K.F.S.; A.S.; A.M.D. and M.S.; supervision, Z.I.; project administration, Z.I.; funding acquisition, Z.I. and M.S. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by “Scientific Innovation strategy-construction of high level Academy of Agricultural sciences (BZ201908)”, “Rural Revitalization special strategic project of Guangdong (201817SY0003)”, “Pakistan Science Foundation PSF/NSLP/C-IU(249)”, “Guangdong Provincial special fund for modern Agriculture Industry Technology Innovation teams (2019KJ119)”, “The key Research and Development Program of Guangdong province (2019B020218004)”, “Discipline Team Building Projects of Guangdong Academy of Agricultural Sciences in the 13th Five-Year Period (201623TD)”, “Science and Technology Program of Guangzhou (201906040005)”.
Acknowledgments
The authors acknowledge Charlotte Thomas and Omar Alfituri from the University of Edinburgh for English style editing and proof reading.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Ministry of Finance and Economic Affairs, Government of Pakistan. Pakistan Economic Survey 2018-19. Available online: http://www.finance.gov.pk/survey_1819.html (accessed on 12 January 2020).
- Devendra, C. Small ruminants in Asia; Contribution to food security, poverty alleviation and opportunities for productivity enhancement. In Proceedings of the International Workshop on Small Ruminant Production and Development in South East Asia, Ho Chi Minh City, Vietnam, 2–4 March 2005; pp. 19–32. [Google Scholar]
- Uzal, F.A.; Giannitti, F.; Finnie, J.W.; García, J.P. Diseases produced by Clostridium perfringens type D. In Clostridial Diseases of Animals; Wiley Blackwell: Ames, IA, USA, 2016; pp. 157–172. [Google Scholar]
- Radostits, O.M.; Gay, C.C.; Hinchcliff, K.W.; Constable, P.D. Veterinary Medicine E-Book: A Textbook of the Diseases of Cattle, Horses, Sheep, Pigs and Goats; Elsevier Health Sciences: Nothingam, UK, 2006. [Google Scholar]
- Stevens, D.L.; Aldape, M.J.; Bryant, A.E. Life-threatening clostridial infections. Anaerobe 2012, 18, 254–259. [Google Scholar] [CrossRef] [PubMed]
- Hill, K.K.; Smith, T.J. Genetic diversity within Clostridium botulinum serotypes, botulinum neurotoxin gene clusters and toxin subtypes. In Botulinum Neurotoxins; Springer: Los Alamos, NM, USA, 2012; pp. 1–20. [Google Scholar]
- Garcia, J.; Adams, V.; Beingesser, J.; Hughes, M.L.; Poon, R.; Lyras, D.; Hill, A.; McClane, B.A.; Rood, J.I.; Uzal, F.A. Epsilon toxin is essential for the virulence of Clostridium perfringens type D infection in sheep, goats, and mice. Infect. Immun. 2013, 81, 2405–2414. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Adams, V.; Bannam, T.L.; Miyamoto, K.; Garcia, J.P.; Uzal, F.A.; Rood, J.I.; McClane, B.A. Toxin plasmids of Clostridium perfringens. Microbiol. Mol. Biol. Rev. 2013, 77, 208–233. [Google Scholar] [CrossRef] [PubMed]
- Uzal, F.A.; Songer, J.G. Diagnosis of Clostridium perfringens intestinal infections in sheep and goats. J. Vet. Diagn. Investig. 2008, 20, 253–265. [Google Scholar] [CrossRef] [PubMed]
- Dray, T. Clostridium perfringens type A and type A and β2 toxin associated with enterotoxemia in a 5-week-old goat. Can. Vet. J. 2004, 45, 251. [Google Scholar] [PubMed]
- Lewis, C.J. Control of important clostridial diseases of sheep. Vet. Clin. Food Anim. Pract. 2011, 27, 121–126. [Google Scholar] [CrossRef]
- Alves, G.G.; de Ávila, R.A.M.; Chávez-Olórtegui, C.D.; Lobato, F.C.F. Clostridium perfringens epsilon toxin: The third most potent bacterial toxin known. Anaerobe 2014, 30, 102–107. [Google Scholar] [CrossRef]
- Bokori-Brown, M.; Savva, C.G.; da Costa, S.P.F.; Naylor, C.E.; Basak, A.K.; Titball, R.W. Molecular basis of toxicity of Clostridium perfringens epsilon toxin. FEBS J. 2011, 278, 4589–4601. [Google Scholar] [CrossRef]
- Uzal, F.A.; McClane, B.A.; Cheung, J.K.; Theoret, J.; Garcia, J.P.; Moore, R.J.; Rood, J.I. Animal models to study the pathogenesis of human and animal Clostridium perfringens infections. Vet. Microbiol. 2015, 179, 23–33. [Google Scholar] [CrossRef]
- Lindström, M.; Heikinheimo, A.; Lahti, P.; Korkeala, H. Novel insights into the epidemiology of Clostridium perfringens type A food poisoning. Food Microbiol. 2011, 28, 192–198. [Google Scholar] [CrossRef]
- Yadav, J.P.; Das, S.C.; Dhaka, P.; Vijay, D.; Kumar, M.; Mukhopadhyay, A.K.; Chowdhury, G.; Chauhan, P.; Singh, R.; Dhama, K. Molecular characterization and antimicrobial resistance profile of Clostridium perfringens type A isolates from humans, animals, fish and their environment. Anaerobe 2017, 47, 120–124. [Google Scholar] [CrossRef] [PubMed]
- Sarker, M.R.; Carman, R.J.; McClane, B.A. Inactivation of the gene (cpe) encoding Clostridium perfringens enterotoxin eliminates the ability of two cpe-positive C. perfringens type A human gastrointestinal disease isolates to affect rabbit ileal loops. Mol. Microbiol. 1999, 33, 946–958. [Google Scholar] [CrossRef] [PubMed]
- Bourlioux, P.; Koletzko, B.; Guarner, F.; Braesco, V. The intestine and its microflora are partners for the protection of the host: Report on the Danone Symposium “The Intelligent Intestine”, held in Paris, June 14, 2002. Am. J. Clin. Nutr. 2003, 78, 675–683. [Google Scholar] [CrossRef] [PubMed]
- Popoff, M.R.; Stiles, B.G. 17 Clostridial Toxins vs. Other Bacterial Toxins. In Handbook on Clostridia; Duerre, P., Ed.; Taylor&Francis Group: Boca Raton, FL, USA, 2005; p. 323. [Google Scholar]
- Kumar, N.V.; Sreenivasulu, D.; Reddy, Y. Prevalence of Clostridium perfringens toxin genotypes in enterotoxemia suspected sheep flocks of Andhra Pradesh. Vet. World 2014, 7, 1132–1136. [Google Scholar] [CrossRef]
- Labbé, R.G.; Juneja, V. Clostridium perfringens gastroenteritis. In Foodborne Infections and Intoxications; Elsevier: Amsterdam, The Netherlands, 2013; pp. 99–112. [Google Scholar]
- Hadimli, H.H.; Erganiş, O.; Sayin, Z.; Aras, Z. Toxinotyping of Clostridium perfringens isolates by ELISA and PCR from lambs suspected of enterotoxemia. Turk. J. Vet. Anim. Sci. 2012, 36, 409–415. [Google Scholar]
- Gajdács, M.; Spengler, G.; Urbán, E. Identification and antimicrobial susceptibility testing of anaerobic bacteria: Rubik’s cube of clinical microbiology? Antibiotics 2017, 6, 25. [Google Scholar] [CrossRef]
- Baums, C.G.; Schotte, U.; Amtsberg, G.; Goethe, R. Diagnostic multiplex PCR for toxin genotyping of Clostridium perfringens isolates. Vet. Microbiol. 2004, 100, 11–16. [Google Scholar] [CrossRef]
- Chukwu, E.E.; Nwaokorie, F.O.; Coker, A.O.; Avila-Campos, M.J.; Solis, R.L.; Llanco, L.A.; Ogunsola, F.T. Detection of toxigenic Clostridium perfringens and Clostridium botulinum from food sold in Lagos, Nigeria. Anaerobe 2016, 42, 176–181. [Google Scholar] [CrossRef]
- Crespo, R.; Fisher, D.J.; Shivaprasad, H.; Fernández-Miyakawa, M.E.; Uzal, F.A. Toxinotypes of Clostridium perfringens isolated from sick and healthy avian species. J. Vet. Diagn. Investig. 2007, 19, 329–333. [Google Scholar] [CrossRef]
- Lebrun, M.; Mainil, J.; Linden, A. Cattle enterotoxaemia and Clostridium perfringens: Description, diagnosis and prophylaxis. Vet. Rec. J. Br. Vet. Assoc. 2010, 167, 13–22. [Google Scholar] [CrossRef]
- Chandran, D.; Naidu, S.S.; Sugumar, P.; Rani, G.S.; Vijayan, S.P.; Mathur, D.; Garg, L.C.; Srinivasan, V.A. Development of a recombinant epsilon toxoid vaccine against enterotoxemia and its use as a combination vaccine with live attenuated sheep pox virus against enterotoxemia and sheep pox. Clin. Vaccine Immunol. 2010, 17, 1013–1016. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.A.; Bahadar, S.; Ullah, N.; Ullah, S.; Shakeeb, U.; Khan, A.Z.; Khan, I.U.; Kalhoro, N.H.; Shah, M.B.; Malik, M.I.U. Distribution and antimicrobial resistance patterns of Clostridium Perfringens isolated from vaccinated and unvaccinated goats. Small Rumin. Res. 2019, 173, 70–73. [Google Scholar] [CrossRef]
- Marshall, B.M.; Levy, S.B. Food animals and antimicrobials: Impacts on human health. Clin. Microbiol. Rev. 2011, 24, 718–733. [Google Scholar] [CrossRef] [PubMed]
- Oliver, S.P.; Murinda, S.E.; Jayarao, B.M. Impact of antibiotic use in adult dairy cows on antimicrobial resistance of veterinary and human pathogens: A comprehensive review. Foodborne Pathog. Dis. 2011, 8, 337–355. [Google Scholar] [CrossRef] [PubMed]
- Greco, G.; Madio, A.; Buonavoglia, D.; Totaro, M.; Corrente, M.; Martella, V.; Buonavoglia, C. Clostridium perfringens toxin-types in lambs and kids affected with gastroenteric pathologies in Italy. Vet. J. 2005, 170, 346–350. [Google Scholar] [CrossRef] [PubMed]
- Kamber, U.; Gokce, H.; Elmali, M. Clostridium perfringens and its toxins in minced meat from Kars, Turkey. Food Addit. Contam. 2007, 24, 673–678. [Google Scholar] [CrossRef]
- Yoo, H.S.; Lee, S.U.; Park, K.Y.; Park, Y.H. Molecular typing and epidemiological survey of prevalence of Clostridium perfringens types by multiplex PCR. J. Clin. Microbiol. 1997, 35, 228–232. [Google Scholar] [CrossRef]
- Ahsani, M.; Mohammadabadi, M.; Shamsaddini, M. Clostridium perfringens isolate typing by multiplex PCR. J. Venom. Anim. Tox. Incl. Trop. Dis. 2010, 16, 573–578. [Google Scholar] [CrossRef]
- Costin, S. Regnum Prokaryotae. Available online: https://www.tgw1916.net/citation.html (accessed on 2 October 2012).
- Mohiuddin, M. Molecular Epidemiology of Clostridium Perfringens Isolated from Small Ruminants. Ph.D. Thesis, Isra University, Islamabad, Pakistan, 2016. [Google Scholar]
- Reed, R. Nitrate, nitrite and indole reactions of gas gangrene anaerobes. J. Bacteriol. 1942, 44, 425. [Google Scholar] [CrossRef]
- Itodo, A.; Adesiyun, A.; Adekeye, J.; Umoh, J. Toxin-types of Clostridium perfringens strains isolated from sheep, cattle and paddock soils in Nigeria. Vet. Microbiol. 1986, 12, 93–96. [Google Scholar] [CrossRef]
- Sipos, W.; Fischer, L.; Schindler, M.; Schmoll, F. Genotyping of Clostridium perfringens isolated from domestic and exotic ruminants and swine. J. Vet. Med. Ser. B 2003, 50, 360–362. [Google Scholar] [CrossRef] [PubMed]
- Uzal, F.; Marcellino, R. Clostridium perfringens in clinically healthy sheep of Patagonia, Argentina. In Proceedings of the 6th Biennial Congress of the Anaerobe Society of the Americas, Park City, UT, USA, 29 June–2 July 2002. [Google Scholar]
- Sobel, J.; Mixter, C.G.; Kolhe, P.; Gupta, A.; Guarner, J.; Zaki, S.; Hoffman, N.A.; Songer, J.G.; Fremont-Smith, M.; Fischer, M. Necrotizing enterocolitis associated with Clostridium perfringens type A in previously healthy North American adults. J. Am. Coll. Surg. 2005, 201, 48–56. [Google Scholar] [CrossRef] [PubMed]
- Omer, S.A.; Al-Olayan, E.M.; Babiker, S.E.H.; Aljulaifi, M.Z.; Alagaili, A.N.; Mohammed, O.B. Genotyping of Clostridium perfringens Isolates from Domestic Livestock in Saudi Arabia. BioMed Res. Int. 2020, 2020, 9035341. [Google Scholar] [CrossRef] [PubMed]
- van Asten, A.J.; Nikolaou, G.N.; Gröne, A. The occurrence of cpb2-toxigenic Clostridium perfringens and the possible role of the β2-toxin in enteric disease of domestic animals, wild animals and humans. Vet. J. 2010, 183, 135–140. [Google Scholar] [CrossRef] [PubMed]
- Mohiuddin, M.; Iqbal, Z.; Rahman, S.U. Prevalence of Clostridium perfringens β2-toxin in sheep and goat population in Punjab, Pakistan. Thai J. Vet. Med. 2016, 46, 491–496. [Google Scholar]
- Jabbari, A.; Tekyei, F.; Esmaelizad, M.; Pilehchian, L.R. Occurrence of Beta2 toxigenic Clostridium perfringens isolates with different toxin types in Iran. Arch. Razi Inst. 2012, 67, 133–137. [Google Scholar]
- Kalender, H.; Ertas, H.; Cetinkaya, B.; Muz, A.; Arslan, N.; Kilic, A. Typing of isolates of Clostridium perfringens from healthy and diseased sheep by multiplex PCR. Vet. Med. Praha 2005, 50, 439. [Google Scholar] [CrossRef]
- Grant, K.A.; Kenyon, S.; Nwafor, I.; Plowman, J.; Ohai, C.; Halford-Maw, R.; Peck, M.W.; McLauchlin, J. The identification and characterization of Clostridium perfringens by real-time PCR, location of enterotoxin gene, and heat resistance. Foodborne Pathog. Dis. 2008, 5, 629–639. [Google Scholar] [CrossRef]
- Özcan, C.; Gürçay, M. Enterotoxaemia incidence in small ruminants in Elazığ and the surrounding provinces in 1994–1998. Turk. J. Vet. Anim. Sci. 2000, 24, 283–286. [Google Scholar]
- Wang, G.; Zhou, J.; Zheng, F.; Lin, G.; Cao, X.; Gong, X.; Qiu, C. Detection of different genotypes of Clostridium perfringens in feces of healthy dairy cattle from China using real-time duplex PCR assay. Pak. Vet. J. 2011, 31, 120–124. [Google Scholar]
- Al-Humiany, A.A. Microbiological studies on enteritis caused by Clostridium perfringens type A, in sheep in Saudi Arabia. J. Appl. Sci. Res. 2012, 2012, 836–844. [Google Scholar]
- Chai, T.; Ma, R.; Chang, W.; Zhang, S. Spread and Pathogenic Mechanism of Clostridium perfringens. Chin. J. Prev. Vet. Med. 2001, 1, 70–72. [Google Scholar]
- Milton, A.A.P.; Agarwal, R.K.; Priya, G.B.; Saminathan, M.; Aravind, M.; Reddy, A.; Athira, C.; Ramees, T.; Sharma, A.K.; Kumar, A. Prevalence and molecular typing of Clostridium perfringens in captive wildlife in India. Anaerobe 2017, 44, 55–57. [Google Scholar] [CrossRef] [PubMed]
- Mohiuddin, M.; Mudasser, H.; Zahid, I.; Iftikhar, H. Immune response of rabbits to hemorrhagic septicemia vaccine formulations adjuvanted with montanide ISA-206, paraffin oil and alum. Asian J. Agri. Biol. 2012, 2, 161–167. [Google Scholar]
- Gharaibeh, S.; Al Rifai, R.; Al-Majali, A. Molecular typing and antimicrobial susceptibility of Clostridium perfringens from broiler chickens. Anaerobe 2010, 16, 586–589. [Google Scholar] [CrossRef]
- Silva, R.O.S.; Ferreira Junior, F.C.; Marques, M.V.R.; Oliveira Junior, C.A.; Martins, N.R.D.S.; Lobato, F.C.F. Genotyping and antimicrobial susceptibility of Clostridium perfringens isolated from Tinamidae, Cracidae and Ramphastidae species in Brazil. Ciência Rural 2014, 44, 486–491. [Google Scholar] [CrossRef][Green Version]
- Salvarani, F.M.; Silva, R.O.S.; Pires, P.S.; Cruz Júnior, E.C.D.C.; Albefaro, I.S.; Guedes, R.M.D.C.; Lobato, F.C.F. Antimicrobial susceptibility of Clostridium perfringens isolated from piglets with or without diarrhea in Brazil. Braz. J. Microbiol. 2012, 43, 1030–1033. [Google Scholar] [CrossRef]
- Tansuphasiri, U.; Matra, W.; Sangsuk, L. Antimicrobial resistance among Clostridium perfringens isolated from various sources in Thailand. Southeast Asian J. Trop. Med. Public Health 2005, 36, 954–961. [Google Scholar]
- Citron, D.M.; Tyrrell, K.L.; Merriam, C.V.; Goldstein, E.J. Comparative in vitro activities of LFF571 against Clostridium difficile and 630 other intestinal strains of aerobic and anaerobic bacteria. Antimicrob. Agents Chemother. 2012, 56, 2493–2503. [Google Scholar] [CrossRef]
- Gholamiandehkordi, A.; Eeckhaut, V.; Lanckriet, A.; Timbermont, L.; Bjerrum, L.; Ducatelle, R.; Haesebrouck, F.; Van Immerseel, F. Antimicrobial resistance in Clostridium perfringens isolates from broilers in Belgium. Vet. Res. Commun. 2009, 33, 1031–1037. [Google Scholar] [CrossRef]
- Johansson, A.; Greko, C.; Engström, B.; Karlsson, M. Antimicrobial susceptibility of Swedish, Norwegian and Danish isolates of Clostridium perfringens from poultry, and distribution of tetracycline resistance genes. Vet. Microbiol. 2004, 99, 251–257. [Google Scholar] [CrossRef] [PubMed]
- Gad, W.; Hauck, R.; Krüger, M.; Hafez, H. Prevalence of Clostridium perfringens in commercial turkey and layer flocks. Arch. Geflkd 2011, 75, 74–79. [Google Scholar]
- Behra-Miellet, J.; Calvet, L.; Dubreuil, L. Activity of linezolid against anaerobic bacteria. Int. J. Antimicrob. Agents 2003, 22, 28–34. [Google Scholar] [CrossRef]
- Miranda, C.; Rojo, M.D. Clostridium Perfringens: Infecciones de Piel y Tejidos Blandos. 2010. Available online: https://www.seimc.org/Contenidos/Ccs/Revisionestematicas/Bacteriologia/Clostper.Pdf (accessed on 12 October 2020).
- Bailey, M. The Development and Use of Multiplex PCR Protocols for the Detection of Clostridium Perfringens Toxin Encoding Genes Cpa, Cpb, Etx, Ia, Cpe, NetB, and TpeL. Master‘s Thesis, Auburn University, Auburn, AL, USA, 2013. [Google Scholar]
- Ferrarezi, M.C.; Cardoso, T.C.; Dutra, I.S. Genotyping of Clostridium perfringens isolated from calves with neonatal diarrhea. Anaerobe 2008, 14, 328–331. [Google Scholar] [CrossRef] [PubMed]
- Guran, H.S.; Vural, A.; Erkan, M.E. The prevalence and molecular typing of Clostridium perfringens in ground beef and sheep meats. J. Für Verbrauch. Und Lebensm. 2014, 9, 121–128. [Google Scholar] [CrossRef]
- Hudzicki, J. Kirby-Bauer disk diffusion susceptibility test protocol. Am. Soc. Microbiol. 2009, 2009, 1–23. [Google Scholar]
- Debalke, D.; Birhan, M.; Kinubeh, A.; Yayeh, M. Assessments of Antibacterial Effects of Aqueous-Ethanolic Extracts of Sida rhombifolia’s Aerial Part. Sci. World J. 2018, 2018, 8429809. [Google Scholar] [CrossRef]
- Hafidh, R.R.; Abdulamir, A.S.; Vern, L.S.; Bakar, F.A.; Abas, F.; Jahanshiri, F.; Sekawi, Z. Inhibition of Growth of Highly Resistant Bacterial and Fungal Pathogens by a Natural Product. Open Microbiol. J. 2011, 5, 96–106. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).