AFM1 Detection in Milk by Fab’ Functionalized Si3N4 Asymmetric Mach–Zehnder Interferometric Biosensors
Abstract
1. Introduction
2. Results
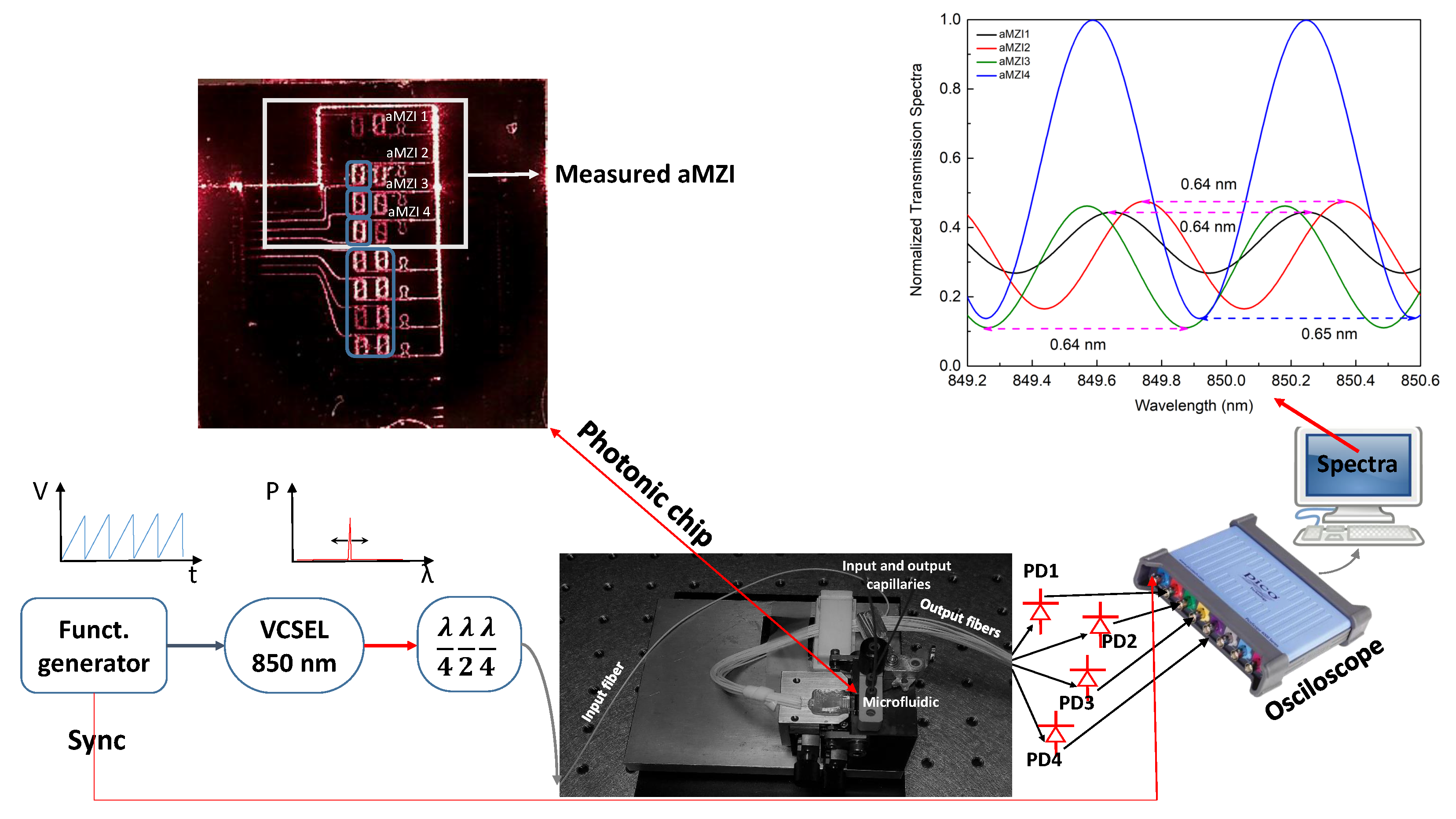
2.1. Experimental Setup
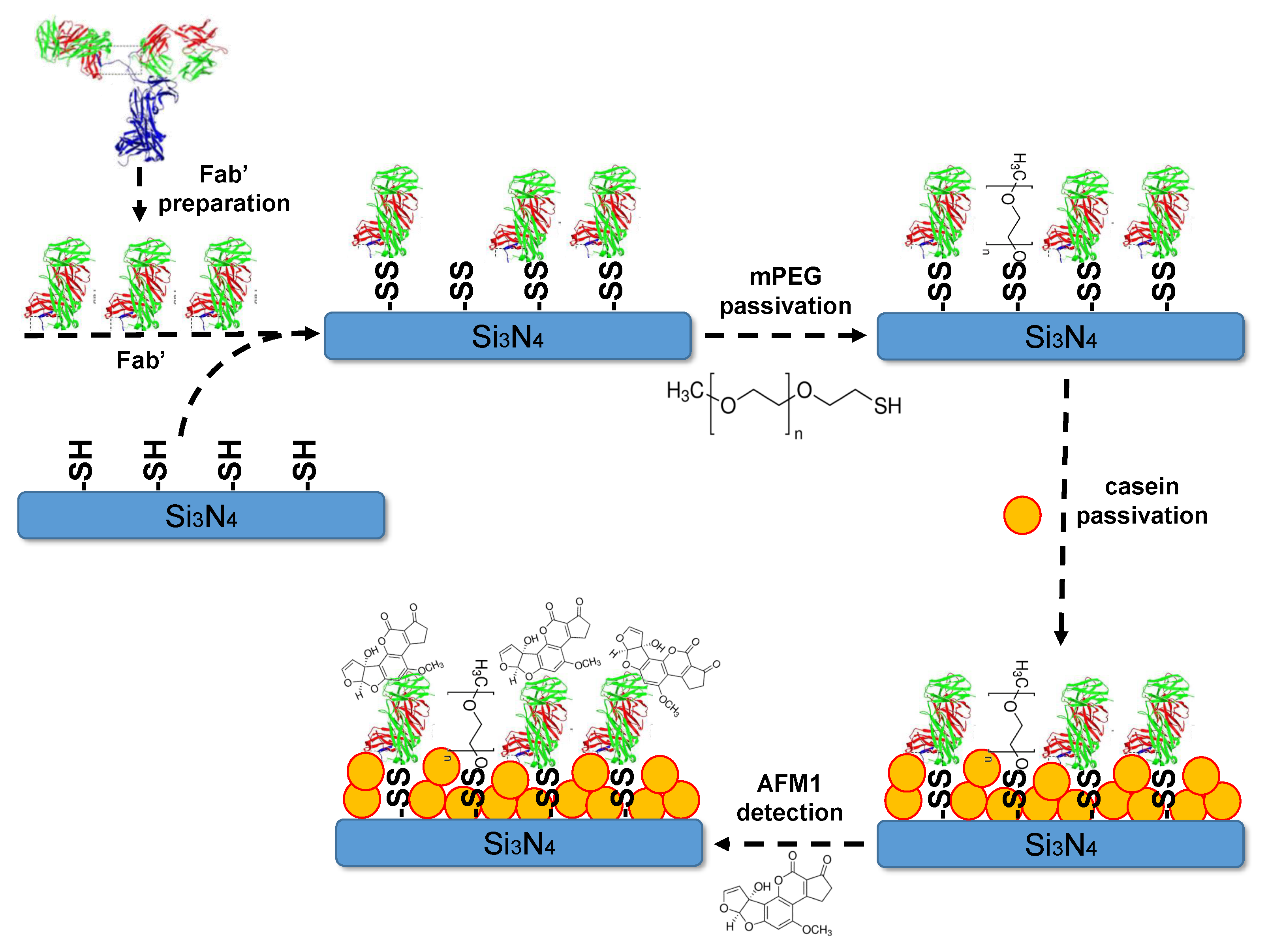
2.2. Surface Preparation
2.3. Theoretical Model
3. Discussion
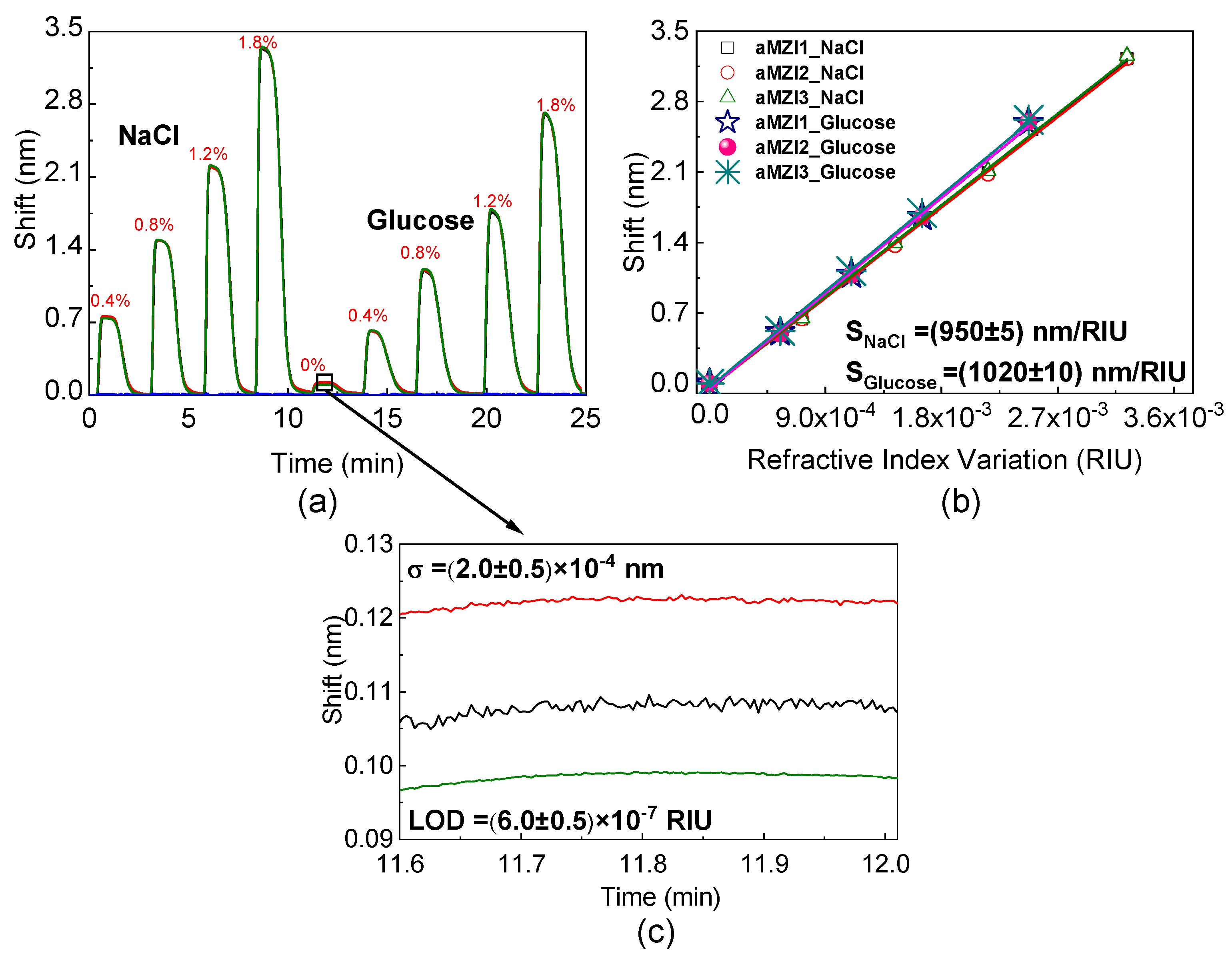
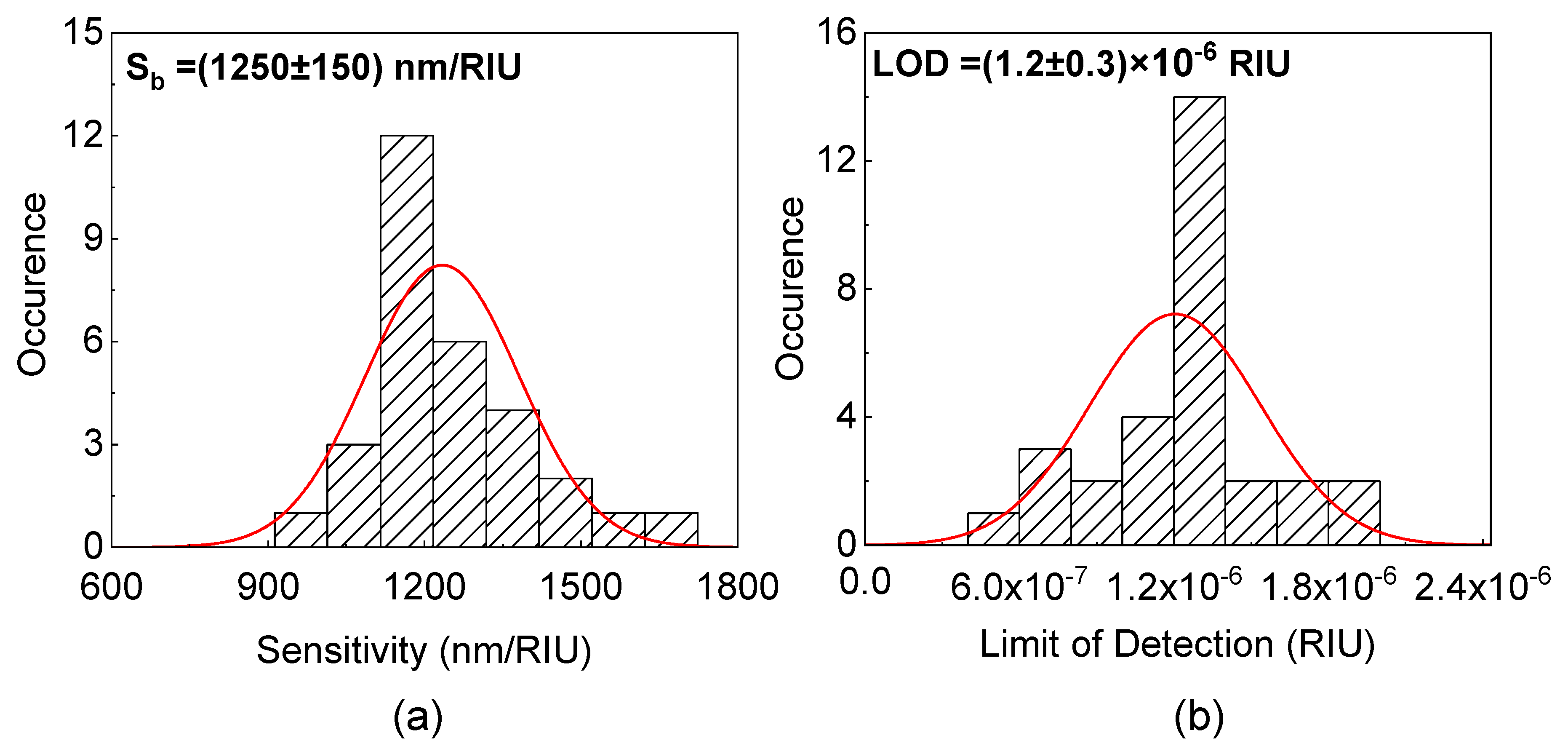
3.1. Bulk Sensitivity and Limit of Detection
3.2. AFM1 Detection
4. Conclusions
5. Materials and Methods
Milk Sample Preparation
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| AFM1 | Aflatoxin M1 |
| AFB1 | Aflatoxin B1 |
| aMZI | Asymmetric Mach–Zehnder interferometer |
| CMOS | Complementary metal oxide semiconductor |
| ELISA | Enzyme-linked immunosorbent assay |
| FSR | Free spectral range |
| HPLC | High performance liquid chromatography |
| hCG | Human chorionic Gonadotropin |
| HRP | Horseradish Peroxidase |
| LOD | Limit Of detection |
| MES | 2-(N-Morpholino)EthaneSulfonic acid |
| MZI | Mach–Zehnder interferometer |
| MRR | Micro ring resonator |
| OTA | Ochratoxin A |
| PDMS | Polydimethylsiloxane |
| RIU | Refractive Index Unit |
| SPR | Surface plasmon resonance |
| TLC | Thin layer chromatography |
| VCSEL | Vertical-cavity surface-emitting laser |
| WGM | Whispering gallery mode |
References
- CAST. Mycotoxins: Economic and Health Risks; Council for Agricultural Science and Technology: Ames, IA, USA, 1989. [Google Scholar]
- Bhat, R. Mycotoxins in Food and Feed: Present Status and Future Concerns. Compr. Rev. Food Sci. Food Saf. 2010, 9, 57–81. [Google Scholar] [CrossRef]
- Groopman, J.D.; Rai, R.V.; Karim, A.A. Aflatoxin Exposure in Human Populations: Measurements and Relationship to Cancer. CRC Crit. Rev. Toxicol. 1988, 19, 113–145. [Google Scholar] [CrossRef] [PubMed]
- Wild, C.; Yun, Y. Mycotoxins and human disease: A largely ignored global health issue. Carcinogenesis 2010, 31, 71–82. [Google Scholar] [CrossRef] [PubMed]
- IARC. List of Classifications; International Agency for Research on Cancer: World Health Organization, Lyon, France, 2012; Volume 56. [Google Scholar]
- Henry, S.H.; Bosch, F.X.; Bowers, J.C. Aflatoxin, Hepatitis and Worldwide Liver Cancer Risks. In Mycotoxins and Food Safety. Advances in Experimental Medicine and Biolog; Springer: Boston, MA, USA, 2002; Volume 504, pp. 229–233. [Google Scholar]
- Polzhofer, K.P. Thermal stability of aflatoxin M1. Z. Lebensm. Unters. Forsch 1977, 164, 80–81. [Google Scholar] [CrossRef] [PubMed]
- Bijl, J.; Peteghem, C.V. Rapid extraction and sample clean-up for the fluorescence densitometric determination of aflatoxin M1 in milk and milk powder. Anal. Chim. Acta 1985, 170, 149–152. [Google Scholar] [CrossRef]
- Behfar, A.; Khorasgani, Z.N.; Alemzadeh, Z.; Goudarzi, M.; Ebrahimi, R.; Tarhani, N. Determination of Aflatoxin M1 Levels in Produced Pasteurized Milk in Ahvaz City by Using HPLC. Jundishapur J. Nat. Pharm. Prod. 2012, 7, 80–84. [Google Scholar] [CrossRef]
- Markaki, P.; Melissari, E. Occurrence of aflatoxin M1 in commercial pasteurized milk determined with ELISA and HPLC. Food Addit. Contam. 1997, 14, 451–456. [Google Scholar] [CrossRef] [PubMed]
- Biancardi, A. Determination of aflatoxin M1 residues in milk. A comparative assessment of ELISA and IAC-HPLC methods. Ind. Aliment. 1997, 36, 870–876. [Google Scholar]
- Abouzied, M.; Bentley, P.; Klein, F.; Rice, J. A Sensitive ELISA for the Detection and Quantitation of Aflatoxin M1 (AFM1) in Milk and Dairy Products; Neogen Corporation: Lansing, MI, USA, 2015. [Google Scholar]
- Wang, Y.; Dostalek, J.; Knoll, W. Long range surface plasmon enhanced fluorescence spectroscopy for the detection of Aflatoxin M1 in milk. Biosens. Bioelectron. 2009, 24, 2264–2267. [Google Scholar] [CrossRef]
- Xu, F.; Zhen, G.; Yu, F.; Kuennemann, E.; Textor, M.; Knoll, W. Combined Affinity and Catalytic Biosensor: In Situ Enzymatic Activity Monitoring of Surface-Bound Enzymes. J. Am. Chem. Soc. 2005, 127, 1308–13085. [Google Scholar] [CrossRef]
- Salter, R.S.; Douglas, D.; Tess, M.; Markovsky, R.J.; Saul, S.J. Collaborative Study of the Charm ROSA Safe Level Aflatoxin M1 Quantitative (SLAFMQ) Lateral Flow Test for Raw Bovine Milk; Charm Sciences, Inc.: Lawrence, MA, USA, 2018. [Google Scholar]
- Tothill, I.E. Biosensors and nanomaterials and their application for mycotoxin determination. World Mycotoxin J. 2011, 4, 361–374. [Google Scholar] [CrossRef]
- Parker, C.O.; Lanyon, H.; Manning, M.; Arrigan, D.W.M.; Tothill, I.E. Electrochemical Immunochip Sensor for Aflatoxin M1 Detection. Anal. Chem. 2009, 81, 5291–5298. [Google Scholar] [CrossRef] [PubMed]
- Paniel, N.; Radoi, A.; Marty, J.L. Development of an electrochemical biosensor for the detection of Aflatoxin M1 in milk. Sensors 2010, 10, 9439–9448. [Google Scholar] [CrossRef] [PubMed]
- Sibanda, L.; Saeger, S.; Peteghem, C.V. Development of a portable field immunoassay for the detection of Aflatoxin M in milk. Int. J. Food Microbiol. 1999, 48, 203–209. [Google Scholar] [CrossRef]
- Istamboulie, G.; Paniel, N.; Zara, L.; Granados, L.R.; Barthelmebs, L.; Noguer, T. Development of an impedimetric aptasensor for the determination of aflatoxin M1 in milk. Talanta 2016, 146, 464–469. [Google Scholar] [CrossRef] [PubMed]
- Karczmarczyk, A.; Dubiak-Szepietowska, M.; Vorobii, M.; Rodriguez-Emmeneggerd, C.; Dostálek, J.; Feller, K.H. Sensitive and rapid detection of aflatoxin M1 in milk utilizing enhanced SPR and p(HEMA) brushes. Biosens. Bioelectron. 2016, 81, 159–165. [Google Scholar] [CrossRef] [PubMed]
- Nirschl, M.; Reuter, F.; Voros, J. Review of Transducer Principles for Label-Free Biomolecular Interaction Analysis. Biosensors 2011, 1, 70–92. [Google Scholar] [CrossRef]
- Lafleur, J.P.; Jonsson, A.; Senkbeil, S.; Kutter, J.P. Recent advances in lab-on-a-chip for biosensing applications. Biosens. Bioelectron. 2016, 76, 213–233. [Google Scholar] [CrossRef]
- Lockwood, D.; Pavesi, L. Silicon Photonics II: Components and Integration Applications, 2nd ed.; Springer: Berlin, Germany, 2011. [Google Scholar]
- Zhu, H.; White, I.M.; Suter, J.D.; Dale, P.S.; Fan, X. Analysis of biomolecule detection with optofluidic ring resonator sensors. Opt. Express 2007, 15, 9139–9146. [Google Scholar] [CrossRef]
- Zlatanovic, S.; Mirkarimi, L.W.; Sigalas, M.M.; Bynum, M.A.; Chow, E.; Robotti, K.M.; Burr, G.W.; Esener, S.; Grot, A. Photonic crystal microcavity sensor for ultracompact monitoring of reaction kinetics and protein concentration. Sens. Actuators B Chem. 2009, 141, 13–19. [Google Scholar] [CrossRef]
- Baaske, M.D.; Foreman, M.R.; Vollmer, F. Single-molecule nucleic acid interactions monitored on a label-free microcavity biosensor platform. Nat. Nanotechnol. 2014, 9, 933–939. [Google Scholar] [CrossRef] [PubMed]
- Heideman, R.G.; Kooyman, R.P.H.; Greve, J. Performance of a highly sensitive optical waveguide Mach–Zehnder interferometer immunosensor. Sens. Actuators B Chem. 1993, 10, 209–217. [Google Scholar] [CrossRef]
- Heideman, R.G.; Lambeck, P.V. Remote opto-chemical sensing with extreme sensitivity: Design, fabrication and performance of a pigtailed integrated optical phase-modulated Mach–Zehnder interferometer system. Sens. Actuators B Chem. 1999, 61, 100–127. [Google Scholar] [CrossRef]
- Liu, Q.; Tu, X.; Kim, K.W.; Kee, J.S.; Shin, Y.; Han, K.; Yoon, Y.J.; Lo, G.Q.; Park, M.K. Highly sensitive Mach–Zehnder interferometer biosensor based on silicon nitride slot waveguide. Sens. Actuators B Chem. 2013, 188, 681–688. [Google Scholar] [CrossRef]
- Duval, D.; Gonzalez-Guerrero, A.B.; Dante, S.; Lechuga, L.M. Interferometric waveguide biosensor based on Si-technology for point-of-care diagnostic. In Proceedings of the Silicon Photonics and Photonic Integrated Circuits III, Brussels, Belgium, 16–19 April 2012; Volume 8431, p. 84310P. [Google Scholar]
- Misiakos, K.; Raptis, I.; Salapatas, A.; Makarona, E.; Botsialas, A.; Hoekman, M.; Stoffer, R.; Jobst, G. Broad-band Mach–Zehnder interferometers as high performance refractive index sensors: Theory and monolithic implementation. Opt. Express 2014, 22, 8856–8870. [Google Scholar] [CrossRef] [PubMed]
- SYMPHONY. Available online: http://www.symphony-project.eu (accessed on 12 July 2019).
- Chalyan, T. Aptamer- and Fab’- functionalized microring resonators for Aflatoxin M1 detection. IEEE J. Sel. Top. Quantum Electron. 2017, 23, 1–8. [Google Scholar] [CrossRef]
- Chalyan, T.; Pasquardini, L.; Falke, F.; Zanetti, M.; Guider, R.; Gandolfi, D.; Schreuder, E.; Pederzolli, C.; Heideman, R.G.; Pavesi, L. Biosensors based on Si3N4 Asymmetric Mach–Zehnder Interferometers. In Proceedings of the SPIE Optical Sensing and Detection IV, Brussels, Belgium, 3–7 April 2016; Volume 9899, p. 98991S. [Google Scholar]
- Worhoff, K. TriPleX: A versatile dielectric photonic platform. Adv. Opt. Technol. 2015, 4, 189–207. [Google Scholar] [CrossRef]
- Roy, D.; Park, J.W. Spatially nanoscale-controlled functional surfaces toward efficient bioactive platforms. J. Mater. Chem. B 2015, 3, 5135–5149. [Google Scholar] [CrossRef][Green Version]
- Yoshimoto, K.; Nishio, M.; Sugasawa, H.; Nagasaki, Y. Direct Observation of Adsorption-Induced Inactivation of Antibody Fragments Surrounded by Mixed-PEG Layer on a Gold Surface. J. Am. Chem. Soc. 2010, 132, 7982–7989. [Google Scholar] [CrossRef] [PubMed]
- Chalyan, T.; Guider, R.; Pasquardini, L.; Zanetti, M.; Falke, F.; Schreuder, E.; Heideman, R.G.; Pederzolli, C.; Pavesi, L. Asymmetric Mach–Zehnder Interferometer Based Biosensors for Aflatoxin M1 Detection. Biosensors 2016, 1, 1. [Google Scholar] [CrossRef]
- Chalyan, T.; Besselink, G.A.J.; Heideman, R.G.; Pavesi, L. Use of microring resonators for biospecific interaction analysis. In Proceedings of the SPIE Optical Sensing, Imaging, and Photon Counting: Nanostructured Devices and Applications 2017, San Diego, CA, USA, 6–10 August 2017; Volume 10353, p. 103530Q. [Google Scholar]
- Stone, D.C.; Ellis, J. Stats Tutorial-Instrumental Analysis and Calibration; Department of Chemistry, University of Toronto: Toronto, ON, Canada, 2011. [Google Scholar]
- Malhotra, S.; Pandey, A.K.; Rajput, Y.S.; Sharma, R. Selection of aptamers for aflatoxin M1 and their characterization. J. Mol. Recognit. 2014, 27, 493–500. [Google Scholar] [CrossRef] [PubMed]
- Anti-Aflatoxin Antibody [1C6] (ab685). Available online: http://www.abcam.com/aflatoxin-antibody-1c6-ab685.html (accessed on 12 July 2019).
- Matabaro, E.; Ishimwe, N.; Uwimbabazi, E.; Lee, B.H. Current Immunoassay Methods for the Rapid detection of Aflatoxin in Milk and Dairy Products. Compr. Rev. Food Sci. Food Saf. 2017, 16, 808–820. [Google Scholar] [CrossRef]
| Method | LOD (ppt) | Detection Time (min) | Reference |
|---|---|---|---|
| TLC | 5 | 4–5 | [8] |
| HPLC | 4.5 | 72 | [9,10] |
| ELISA | 4.3 | 3 | [11,12] |
| ROSA | 500 | 0.15 | [15] |
| Bilayer Lipid Membranes | 16 | 0.5 | [16] |
| Microelectrodes Arrays | 8 | 2 | [17] |
| Electrochemical | 10 | 0.5 | [18] |
| Field Immunoassay | 50 | 3 | [19] |
| DNA-aptasensor | 20 | 4 | [20] |
| SPR | 0.6 | 1 | [13] |
| SPR with gold nanoparticles | 18 | 1 | [21] |
| aMZI | 16.8 | 1.5 | This work |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chalyan, T.; Potrich, C.; Schreuder, E.; Falke, F.; Pasquardini, L.; Pederzolli, C.; Heideman, R.; Pavesi, L. AFM1 Detection in Milk by Fab’ Functionalized Si3N4 Asymmetric Mach–Zehnder Interferometric Biosensors. Toxins 2019, 11, 409. https://doi.org/10.3390/toxins11070409
Chalyan T, Potrich C, Schreuder E, Falke F, Pasquardini L, Pederzolli C, Heideman R, Pavesi L. AFM1 Detection in Milk by Fab’ Functionalized Si3N4 Asymmetric Mach–Zehnder Interferometric Biosensors. Toxins. 2019; 11(7):409. https://doi.org/10.3390/toxins11070409
Chicago/Turabian StyleChalyan, Tatevik, Cristina Potrich, Erik Schreuder, Floris Falke, Laura Pasquardini, Cecilia Pederzolli, Rene Heideman, and Lorenzo Pavesi. 2019. "AFM1 Detection in Milk by Fab’ Functionalized Si3N4 Asymmetric Mach–Zehnder Interferometric Biosensors" Toxins 11, no. 7: 409. https://doi.org/10.3390/toxins11070409
APA StyleChalyan, T., Potrich, C., Schreuder, E., Falke, F., Pasquardini, L., Pederzolli, C., Heideman, R., & Pavesi, L. (2019). AFM1 Detection in Milk by Fab’ Functionalized Si3N4 Asymmetric Mach–Zehnder Interferometric Biosensors. Toxins, 11(7), 409. https://doi.org/10.3390/toxins11070409







