Effect of Compound Probiotics and Mycotoxin Degradation Enzymes on Alleviating Cytotoxicity of Swine Jejunal Epithelial Cells Induced by Aflatoxin B1 and Zearalenone
Abstract
1. Introduction
2. Results
2.1. The Preliminary Reaction Time, Concentrations, and Relationship of AFB1 and ZEA Determined by the Exposed IPEC-J2 Cell Viability
2.2. Effects of CFSCP + MDE on Alleviating Cell Necrosis and Apoptosis Induced by AFB1 and ZEA
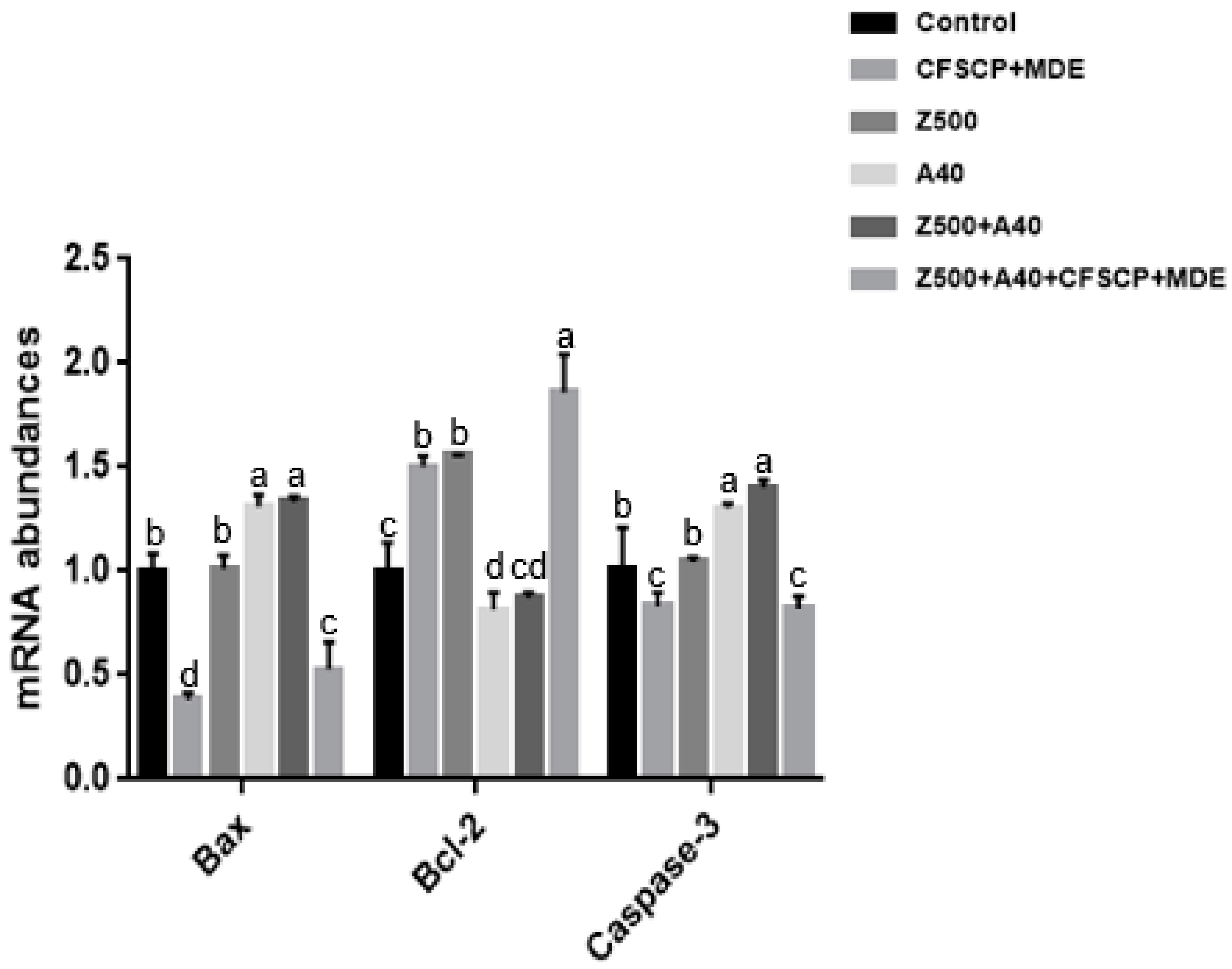
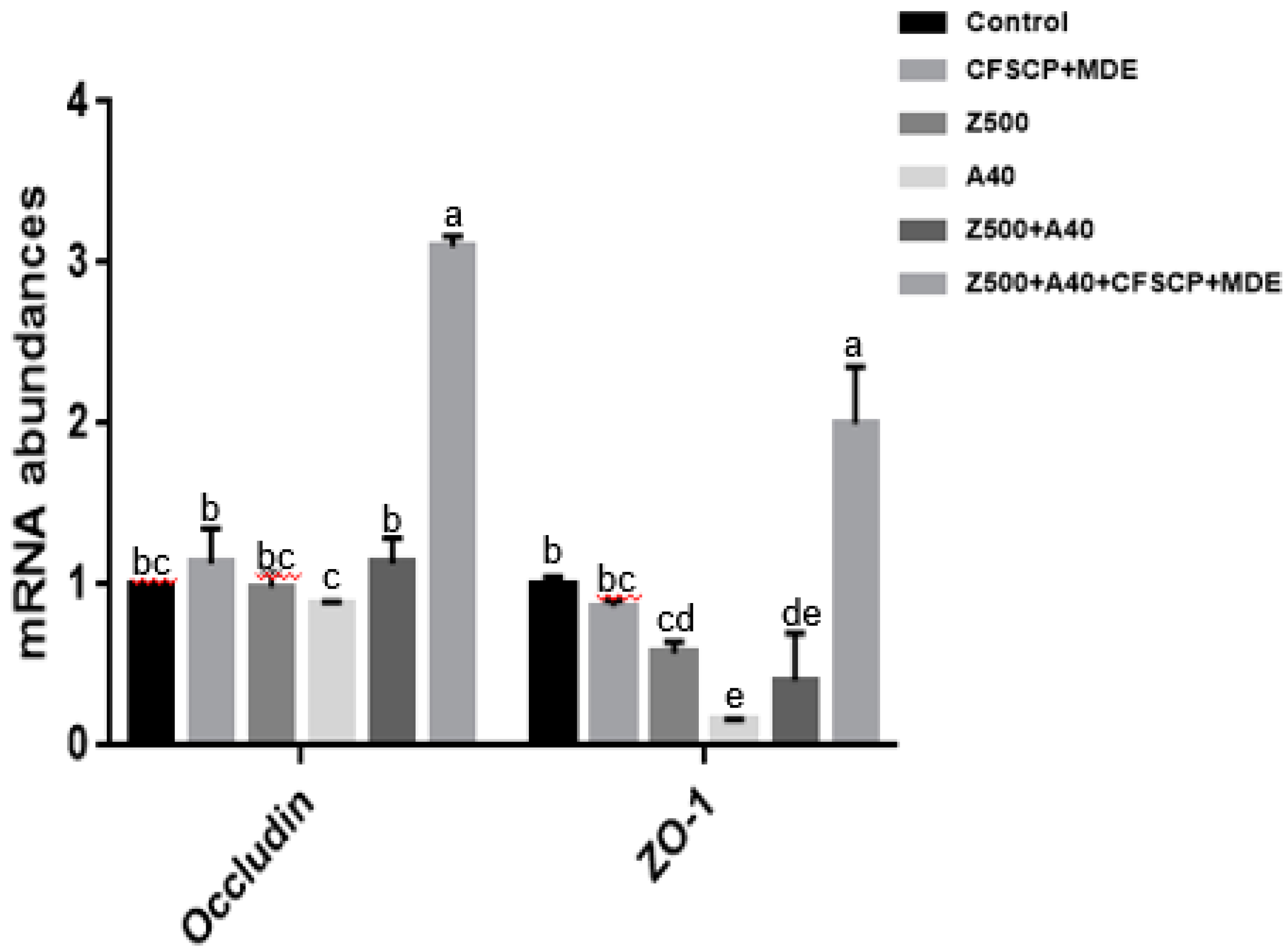
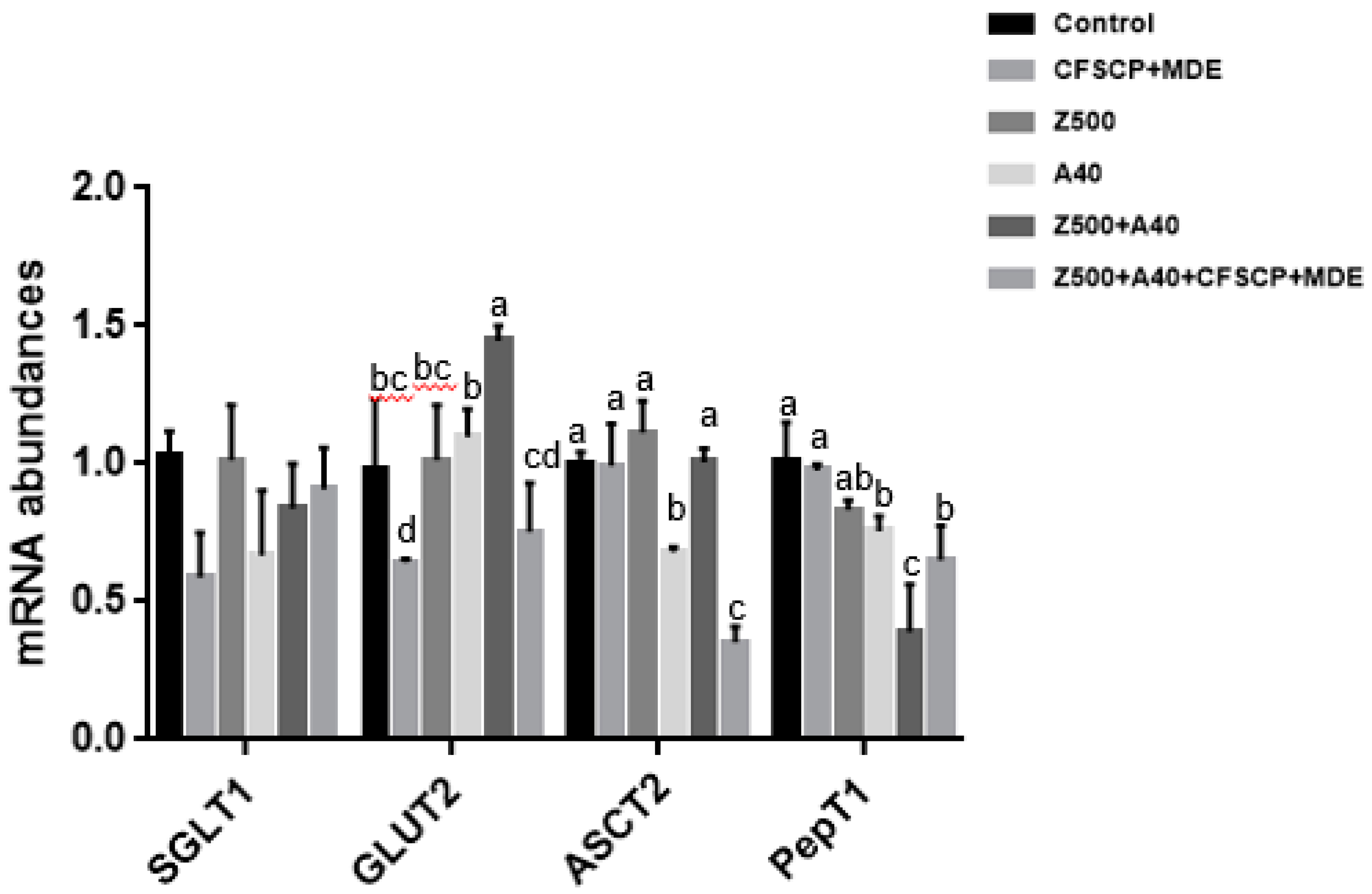
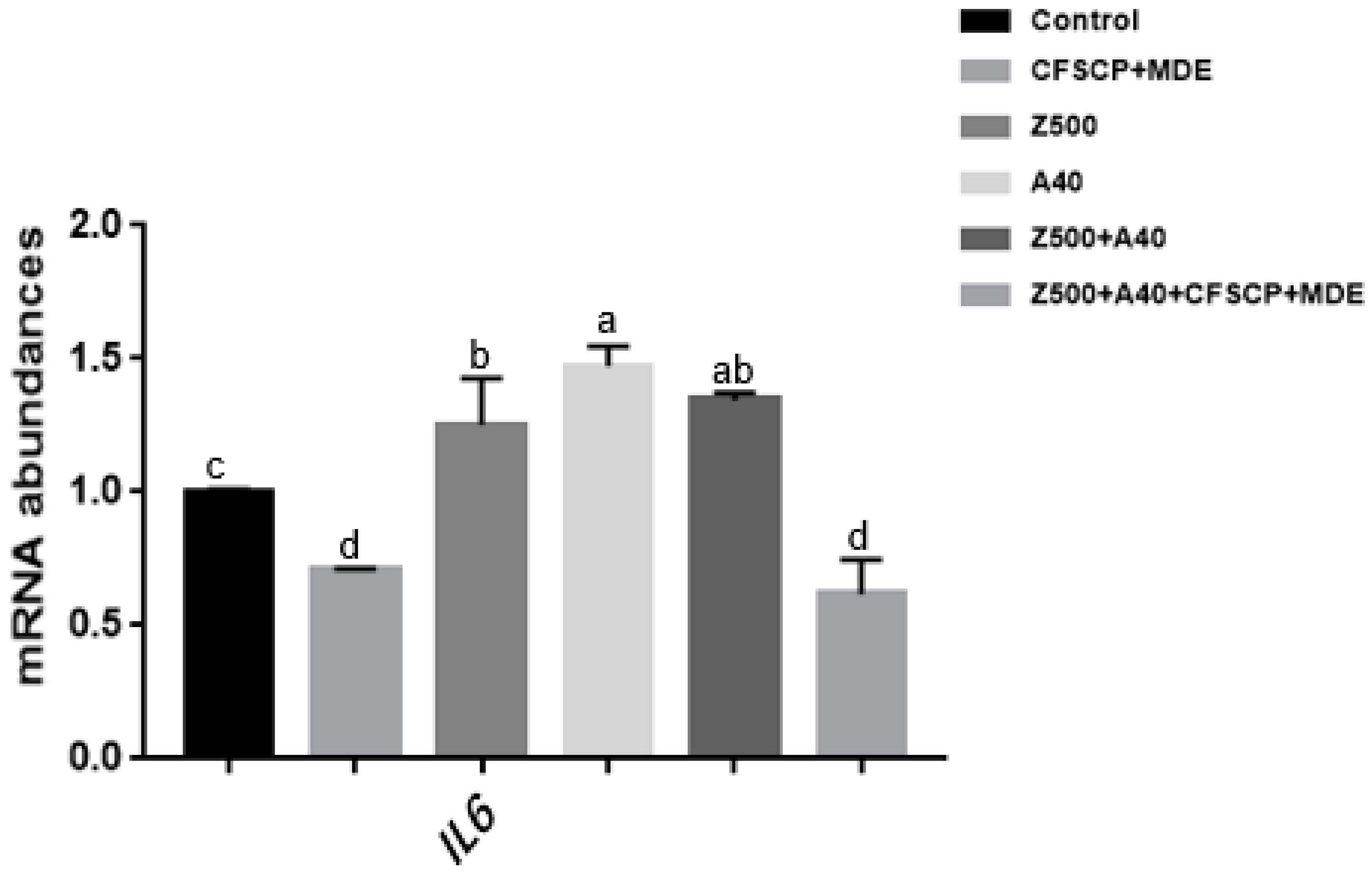
2.3. Effects of CFSCP + MDE on Inflammation, Apoptosis, Tight Junction (TJ) Proteins, and Nutrient Transport Gene mRNA Abundances of IPEC-J2 Cells Induced By AFB1 And ZEA
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Chemicals and Cell Culture
5.2. Microbial Preparation
5.3. Experimental Design
- (1)
- Control group;
- (2)
- CFSCP + MDE;
- (3)
- Toxin group added with 500 or 1000 µg/L ZEA (Z500 or Z1000);
- (4)
- Toxin group added with 40 or 80 µg/L AFB1 (A40 or A80);
- (5)
- Toxin group added with 500 µg/L ZEA and 40 µg/L AFB1 or 1000 µg/L ZEA and 80 µg/L AFB1 (Z500 + A40 or Z1000 + A80);
- (6)
- Toxin group added with 500 µg/L ZEA and 40 µg/L AFB1 plus CFSCP + MDE (Z500 + A40 + CFSCP + MDE).
5.4. Cell Viability Determination
5.5. Determination of Cell Status
5.6. qRT-PCR Analysis of Cytokine Genes
5.7. Data Analyses
Author Contributions
Funding
Conflicts of Interest
References
- Paterson, R.R.M.; Lima, N. How will change affect mycotoxins in food. Food Res. Int. 2010, 42, 1902–1914. [Google Scholar] [CrossRef]
- Wild, C.P.; Miller, J.D.; Groopman, J.D. Mycotoxin Control in Low- and Middle-Income Countries; IARC Working Group Reports; International Agency for Research on Cancer: Lyon, France, 2015. [Google Scholar]
- Rawal, S.; Kim, J.E.; Coulombe, J.R. Aflatoxin B1 in poultry: Toxicology, metabolism and prevention. Res. Vet. Sci. 2010, 89, 325–331. [Google Scholar] [CrossRef] [PubMed]
- Kuiper-Goodman, T.; Scott, P.M.; Watanabe, H. Risk assessment of the mycotoxin zearalenone. Regul. Toxicol. Pharmacol. 1987, 7, 253–306. [Google Scholar] [CrossRef]
- Diekman, M.A.; Green, M.L. Mycotoxins and reproduction in domestic livestock. J. Animal Sci. 1992, 70, 1615–1627. [Google Scholar] [CrossRef]
- D’Mello, J.P.F.; Mac Donald, A.M.C. Mycotoxins. Anim. Feed Sci. Tech. 1997, 69, 155–166. [Google Scholar] [CrossRef]
- Yang, Y.X.; Yu, S.; Tan, Y.; Liu, N.; Wu, A. Individual and combined cytotoxic effects of co-cccurring deoxynivalenol family mycotoxins on human gastric epithelial cells. Toxins 2017, 9, 96. [Google Scholar] [CrossRef]
- Rodriguez-Carrasco, Y.; Jose Ruiz, M.; Font, G.; Berrada, H. Exposure estimates to Fusarium mycotoxins through cereals intake. Chemosphere 2013, 93, 2297–2303. [Google Scholar] [CrossRef]
- FAO. Manual on the Application of the HACCP System in Mycotoxin Prevention and Control; Joint FAO/WHO Food Standards Programme FAO: Rome, Italy, 2002. [Google Scholar]
- San’Ana, A.S.; Rosenthal, A.; De Massaguer, P.R. The fate of patulin in apple juice processing: A review. Food Res. Int. 2008, 41, 441–453. [Google Scholar] [CrossRef]
- Walter, P.P.; Tunde, P.; Istvan, P. Mycotoxins-prevention and decontamination by yeast. J. Basic Microbiol. 2015, 55, 805–818. [Google Scholar]
- Teniola, O.D.; Addo, P.A.; Brost, I.M.; Färber, P.; Jany, K.D.; Alberts, J.F.; Holzapfel, W.H. Degradation of aflatoxin B1 by cell-free extracts of Rhodococcus erythropolis and Mycobacterium fluoranthenivorans sp. nov. DSM44556T. Int. J. Food Microbiol. 2005, 105, 111–117. [Google Scholar] [CrossRef]
- Cserháti, M.; Kriszt, B.; Krifaton, C.; Szoboszlay, S.; Háhn, J.; Toth, S.; Nagy, I.; Kukolya, J. Mycotoxin-degradation profile of Rhodococcus strains. Int. J. Food Microbiol. 2013, 166, 176–185. [Google Scholar] [CrossRef]
- Krifaton, C.; Kriszt, B.; Szoboszlay, S.; Cserháti, M.; Szǔcs, Á.; Kukolya, J. Analysis of aflatoxin B1-degrading microbes by use of a combined toxicity-profiling method. Mutat. Res./Genet. Toxicol. Environ. Mutagen. 2011, 726, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.L.; Yao, D.S.; Liang, L.; Ma, L.; Cheng, W.Q.; Gu, L.Q. Detoxification of Aflatoxin B1 by enzymes isolated from Armillariel latabescans. Food Chem. Toxicol. 1998, 36, 563–574. [Google Scholar] [CrossRef]
- Liang, Z.H.; Li, J.X.; He, Y.L.; Guan, S.; Wang, N.; Ji, C.; Niu, T.G. AFB1 biodegradation by a new strain-Stenotrophomonas sp. Agric. Sci. China 2008, 7, 1433–1437. [Google Scholar] [CrossRef]
- Altalhi, A.D. Plasmid-inediated detoxification of mycotoxin zearalenone in Pseudomonas sp. ZEA-1. Am. J. Biotechnol. Biochem. 2007, 3, 150–158. [Google Scholar] [CrossRef]
- Kriszt, R.; Krifaton, C.; Szoboszlay, S.; Cserháti, M.; Kriszt, B.; Kukolya, J.; Czéh, Á.; Fehér-Tóth, S.Z.; Török, L.; Szöke, Z.S.; et al. A new zearalenone biodegradation atrategy using non-pathogenic. Rhodococcus pyridinivorans K408 strain. PLoS ONE 2012, 9, e43608. [Google Scholar]
- Zhang, H.Y.; Dong, M.J.; Yang, Q.Y.; Apaliya, M.T.; Li, J.; Zhang, X.Y. Biodegradation of zearalenone by Saccharomyces cerevisiae: Possible involvement of ZEN responsive proteins of the yeast. J. Proteom. 2016, 143, 416–423. [Google Scholar] [CrossRef]
- Zuo, R.Y.; Chang, J.; Yin, Q.Q.; Wang, P.; Yang, Y.R.; Wang, X.; Wang, G.Q.; Zheng, Q.H. Effect of the combined probiotics with aflatoxin B1-degrading enzyme on aflatoxin detoxification, broiler production performance and hepatic enzyme gene expression. Food Chem. Toxicol. 2013, 59, 470–475. [Google Scholar] [CrossRef]
- Huang, W.W.; Chang, J.; Wang, P.; Liu, C.Q.; Yin, Q.Q.; Zhu, Q.; Lu, F.S.; Gao, T.Z. Effect of the combined compound probiotics with mycotoxin-degradation enzyme on detoxifying aflatoxin B1 and zearalenone. J. Toxicol. Sci. 2018, 43, 377–385. [Google Scholar] [CrossRef]
- Jebali, R.; Ben Salah-Abbes, J.; Abbes, S.; Hassan, A.M.; Abdel-Aziem, S.H.; El-Nekeety, A.A.; Oueslati, R.; Abdel-Wahhab, M.A. Lactobacillus plantarum alleviate aflatoxins (B1 and M1) induced disturbances in the intestinal genes expression and DNA fragmentation in mice. Toxicon 2018, 146, 13–23. [Google Scholar] [CrossRef]
- El-Nezami, H.; Haskard, C.; Salminen, E.; Mykkanen, A.; Ahokas, J.; Salminen, S. Lactic acid bacteria and bifidobacteria can reduce dietary exposure to aflatoxins. Br. J. Nutr. 2002, 88, 115–116. [Google Scholar]
- Niderkorn, V.; Morgavi, D.P.; Pujos, E.; Tissandier, A.; Boudra, H. Screening of fermentative bacteria for their ability to bind and biotransform deoxynivalenol, zearalenone and fumonisins in an in vitro simulated corn silage model. Food Addit. Contam. 2007, 24, 406–415. [Google Scholar] [CrossRef] [PubMed]
- Bouhet, S.; Oswald, I.P. The effects of mycotoxins, fungal food contaminants, on the intestinal epithelial cell derived innate immune response. Vet. Immunol. Immunopathol. 2005, 108, 199–209. [Google Scholar] [CrossRef] [PubMed]
- Bouhet, S.; Oswald, I.P. The intestine as a possible target for fumonisin toxicity. Mol. Nutr. Food Res. 2007, 51, 925–931. [Google Scholar] [CrossRef] [PubMed]
- Oswald, I.P. Role of intestinal epithelial cells in the innate immune defence of pig intestine. Vet Res. 2006, 37, 359–368. [Google Scholar] [CrossRef] [PubMed]
- Pinton, P.; Oswald, I.P. Effect of deoxynivalenol and other type B trichothecenes on the intestine: A review. Toxins 2014, 6, 1615–1643. [Google Scholar] [CrossRef] [PubMed]
- Robert, H.; Payros, D.; Pinton, P.; Théodorou, V.; Mercier-Bonin, M.; Oswald, I.P. Impact of mycotoxins on the intestine: Are mucus and microbiota new targets? J. Toxicol. Environ. Health Part B Crit. Rev. 2017, 280, 1–27. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Li, G.; Wen, K.; Bui, T.; Cao, D.; Zhang, Y. Porcine small intestinal epithelial cell line (IPEC-J2) of rotavirus infection as a new model for the study of innate immune responses to rotaviruses and probiotics. Viral Immunol. 2010, 23, 135–149. [Google Scholar] [CrossRef]
- Arce, C.; Ramírez-Boo, M.; Lucena, C.; Garrido, J.J. Innate immune activation of swine intestinal epithelial cell lines (IPEC-J2 and IPI-2I) in response to LPS from Salmonella typhimurium. Comp. Immunol. Microb. 2010, 33, 161–174. [Google Scholar] [CrossRef]
- Brosnahan, A.J.; Brown, D.R. Porcine IPEC-J2 intestinal epithelial cells in microbiological investigations. Vet. Microbiol. 2012, 156, 229–237. [Google Scholar] [CrossRef]
- Wu, Y.P.; Zhu, C.; Chen, Z.; Chen, Z.J.; Zhang, W.N.; Ma, X.Y.; Wang, L.; Yang, X.F.; Jiang, Z.Y. Protective effects of Lactobacillus plantarum on epithelial barrier disruption caused by enterotoxigenic Escherchia coli in intestinal porcine epithelial cells. Vet. Immunol. Immunop. 2016, 172, 55–63. [Google Scholar] [CrossRef] [PubMed]
- Lei, M.Y.; Zhang, N.Y.; Qi, D.S. In vitro investigation of individual and combined cytotoxic effects of aflatoxin B1 and other selected mycotoxins on the cell line porcine kidney 15. Exp. Toxicol. Pathol. 2013, 65, 1149–1157. [Google Scholar] [CrossRef] [PubMed]
- Tatay, E.; Meca, G.; Font, G.; Ruiz, M.J. Interactive effects of zearalenone and its metabolites on cytotoxicity and metabolization in ovarian CHO-K1 cells. Toxicol. In Vitro 2014, 28, 95–103. [Google Scholar] [CrossRef] [PubMed]
- Mechoud, M.A.; Juarez, G.E.; de Valdez, G.F.; Rodriguez, A.V. Lactobacillus reuteri CRL 1098 and Lactobacillus acidophilus CRL 1014 differently reduce in vitro immunotoxic effect induced by ochratoxin A. Food Chem. Toxicol. 2012, 50, 4310–4315. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Yuan, H.; Guo, C.; Lu, Y.; Deng, S.; Yang, Y. Zearalenone induces apoptosis and necrosis in porcine granulosa cells via a caspase-3- and caspase-9-dependent mitochondrial signaling pathway. J. Cell Physiol. 2012, 227, 1814–1820. [Google Scholar] [CrossRef] [PubMed]
- Zain, M.E. Impact of mycotoxins on humans and animals. J. Saudi Chem. Soc. 2011, 15, 129–144. [Google Scholar] [CrossRef]
- Xiao, Z.; Liu, L.; Tao, W.; Pei, X.; Wang, G.; Wang, M. Clostridium tyrobutyricum protect intestinal barrier function from LPS-induced apoptosis via P38/JNK signaling pathway in IPEC-J2 cells. Cell Physiol. Biochem. 2018, 46, 1779–1792. [Google Scholar] [CrossRef]
- Becker, H.M.; Apladas, A.; Scharl, M.; Fried, M.; Rogler, G. Probiotic Escherichia coli Nissle 1917 and commensal E. coli K12 differentially affect the inflammasome in intestinal epithelial cells. Digestion 2014, 89, 110–118. [Google Scholar] [CrossRef]
- Maresca, M.; Mahfoud, R.; Garmy, N.; Fantini, J. The mycotoxin deoxynivalenol affects nutrient absorption in human intestinal epithelial cells. J. Nutr. 2002, 132, 2723–2731. [Google Scholar] [CrossRef]
- Hunder, G.; Schumann, K.; Strugala, G.; Gropp, J.; Fichtl, B.; Forth, W. Influence of subchronic exposure to low dietary deoxynivalenol, a trichothecene mycotoxin, on intestinal absorption of nutrients in mice. Food Chem. Toxicol. 1991, 29, 809–814. [Google Scholar] [CrossRef]
- Scalise, M.; Pochini, L.; Console, L.; Losso, M.A.; Indiveri, C. The human SLC1A5 (ASCT2) amino acid transporter: From function to structure and role in cell biology. Front. Cell Dev. Biol. 2018, 6, 96. [Google Scholar] [CrossRef] [PubMed]
- Mertl, M.; Daniel, H.; Kottra, G. Substrate-induced changes in the density of peptide transporter PEPT1 expressed in Xenopus oocytes. Am. J. Physiol. Cell Physiol. 2008, 295, C1332–C1343. [Google Scholar] [CrossRef] [PubMed]
- Ashida, H.; Ogawa, M.; Kim, M.; Mimuro, H.; Sesakawa, C. Bacteria and host interactions in the gut epithelial barrier. Nat. Chem. Biol. 2012, 8, 36–45. [Google Scholar] [CrossRef] [PubMed]
- Bertrand, G.; Todd, J.A. Modulation of intestinal functions following mycotoxin ingestion: Meta-analysis of published experiments in animals. Toxins 2013, 5, 396–430. [Google Scholar]
- Gu, M.J.; Song, S.K.; Park, S.M.; Lee, I.K.; Yun, C.H. Bacillus subtilis protects porcine intestinal barrier from deoxynivalenol via improved zonula occludens-1 expression. Asian Australas. J. Anim. Sci. 2013, 27, 580–586. [Google Scholar] [CrossRef] [PubMed]
- Mennigen, R.; Nolte, K.; Rijcken, E.; Utech, M.; Loeffler, B.; Senninger, N.; Bruewer, M. Probiotic mixture VSL#3 protects the epithelial barrier by maintaining tight junction protein expression and preventing apoptosis in a murine model of colitis. Am. J. Physiol. Gastrointest. Liver Physiol. 2009, 296, G1140–G1149. [Google Scholar]
- Huang, S.; Zheng, N.; Fan, C.; Cheng, M.; Wang, S.; Jabar, A.; Wang, J.; Cheng, J. Effects of aflatoxin B1 combined with ochratoxin A and/or zearalenone on metabolism, immune function, and antioxidant status in lactating dairy goats. Asian-Australas. J. Anim. Sci. 2018, 31, 505–513. [Google Scholar] [CrossRef]
- Gao, X.; Ma, Q.G.; Zhao, H.L.; Lei, Y.P.; Shan, Y.J.; Ji, C. Isolation of Bacillus subtilis: Screening for aflatoxins B1, M1 and G1 detoxification. Eur. Food Res. Technol. 2011, 232, 957–962. [Google Scholar] [CrossRef]
- Fotakis, G.; Timbrell, J.A. In vitro cytotoxicity assays: Comparison of LDH, neutral red, MTT and protein assay in hepatoma cell lines following exposure to cadmium chloride. Toxicol. Lett. 2006, 160, 171–177. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
| Groups | 6 h | 12 h | 18 h | 24 h | 48 h |
|---|---|---|---|---|---|
| Z500 | 83.48 ± 4.26Cb | 101.79 ± 6.60Aa | 94.18 ± 6.39Bab | 90.15 ± 6.21Bc | 103.72 ± 3.35Aa |
| A40 | 82.83 ± 3.48Cbc | 95.04 ± 3.04Bb | 94.94 ± 3.98Bab | 97.72 ± 2.35Ba | 103.48 ± 3.33Aa |
| Z500 + A40 | 81.02 ± 4.02Ccd | 85.70 ± 5.77BCc | 87.24 ± 6.62BCcd | 88.63 ± 4.27ABbc | 94.56 ± 4.21Abc |
| Z1000 | 79.40 ± 3.67Bcd | 92.02 ± 5.61Ab | 93.50 ± 3.69Aab | 91.45 ± 6.47Ab | 94.91 ± 5.90Abc |
| A80 | 81.58 ± 3.37Cbc | 92.30 ± 3.21ABb | 91.73 ± 3.63Bbc | 95.84 ± 4.66ABa | 97.27 ± 5.90Ab |
| Z1000 + A80 | 75.45 ± 5.81Cd | 85.35 ± 7.28ABc | 83.99 ± 4.44Bd | 81.64 ± 2.72Bd | 90.96 ± 3.78Ac |
| CFSCP + MDE | 98.56 ± 0.45Aa | 99.12 ± 0.67Aa | 98.36 ± 0.06Aa | 98.96 ± 0.43Aa | 98.15 ± 0.55Ab |
| Groups | Q1 | Q2 | Q3 | Q4 |
|---|---|---|---|---|
| Control group | 1.66 ± 0.09d | 0.96 ± 0.24b | 0.75 ± 0.30c | 96.63 ± 0.38a |
| CFSCP + MDE | 2.44 ± 0.24c | 2.44 ± 0.42a | 2.14 ± 0.36b | 92.97 ± 1.00b |
| Z500 | 5.36 ± 0.45a | 0.90 ± 0.39b | 0.89 ± 0.14c | 92.86 ± 0.56b |
| A40 | 4.19 ± 0.30b | 2.33 ± 0.37a | 2.94 ± 0.09a | 90.54 ± 0.57c |
| Z500 + A40 | 5.13 ± 0.84a | 2.85 ± 0.68a | 2.50 ± 0.25b | 89.52 ± 0.09c |
| Z500 + A40 + CFSCP + MDE | 2.94 ± 0.32c | 2.56 ± 0.43a | 2.20 ± 0.25b | 92.30 ± 1.00b |
| Gene | Primer Sequence (5′-3′) | Accession Number | Size (bp) |
|---|---|---|---|
| GAPDH | F: ATGACCACAGTCCATGCCATC | XM-004387206.1 | 271 |
| R: CCTGCTTCACCACCTTCTTG | |||
| Cell apoptosis genes | |||
| Bcl-2 | F: AGAGCCGTTTCGTCCCTTTC | XM-003122573.2 | 270 |
| R: GCACGTTTCCTAGCGAGCAT | |||
| Bax | F: ATGATCGCAGCCGTGGACACG | XM-003355975.1 | 296 |
| R: ACGAAGATGGTCACCGTCTGC | |||
| Caspase-3 | F:TTGGACTGTGGGATTGAGACG | NM-214131.1 | 165 |
| R: CGCTGCACAAAGTGACTGGA | |||
| Cytokines gene | |||
| IL-6 | F: GCTCTCTGTGAGGCTGCAGTTC | NM_213867.1 | 107 |
| R: AAGGTGTGGAATGCGTATTTATGC | |||
| Barrier function genes | |||
| ZO-1 | F: CCTGAGTTTGATAGTGGCGTTGA | XM-003353439.2 | 269 |
| R: AAATAGATTTCCTGCCCAATTCC | |||
| Occludin | F: ACCCAGCAACGACATA | NM_001163647.2 | 155 |
| R: TCACGATAACGAGCATA | |||
| Nutrient transporter genes | |||
| SGLT1 | F: TCATCATCGTCCTGGTCGTCTC | M34044.1 | 144 |
| R: CTTCTGGGGCTTCTTGAATGTC | |||
| GLUT2 | F: ATTGTCACAGGCATTCTTGTTAGTCA | NM_001097417.1 | 273 |
| R: TTCACTTGATGCTTCTTCCCTTTC | |||
| PepT1 | F: CAGACTTCGACCACAACGGA | NM_214347.1 | 99 |
| R: TTATCCCGCCAGTACCCAGA | |||
| ASCT2 | F: CTGGTCTCCTGGATCATGTGG | DQ231578.1 | 172 |
| R: CAGGAAGCGGTAGGGGTTTT | |||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, W.; Chang, J.; Wang, P.; Liu, C.; Yin, Q.; Song, A.; Gao, T.; Dang, X.; Lu, F. Effect of Compound Probiotics and Mycotoxin Degradation Enzymes on Alleviating Cytotoxicity of Swine Jejunal Epithelial Cells Induced by Aflatoxin B1 and Zearalenone. Toxins 2019, 11, 12. https://doi.org/10.3390/toxins11010012
Huang W, Chang J, Wang P, Liu C, Yin Q, Song A, Gao T, Dang X, Lu F. Effect of Compound Probiotics and Mycotoxin Degradation Enzymes on Alleviating Cytotoxicity of Swine Jejunal Epithelial Cells Induced by Aflatoxin B1 and Zearalenone. Toxins. 2019; 11(1):12. https://doi.org/10.3390/toxins11010012
Chicago/Turabian StyleHuang, Weiwei, Juan Chang, Ping Wang, Chaoqi Liu, Qingqiang Yin, Andong Song, Tianzeng Gao, Xiaowei Dang, and Fushan Lu. 2019. "Effect of Compound Probiotics and Mycotoxin Degradation Enzymes on Alleviating Cytotoxicity of Swine Jejunal Epithelial Cells Induced by Aflatoxin B1 and Zearalenone" Toxins 11, no. 1: 12. https://doi.org/10.3390/toxins11010012
APA StyleHuang, W., Chang, J., Wang, P., Liu, C., Yin, Q., Song, A., Gao, T., Dang, X., & Lu, F. (2019). Effect of Compound Probiotics and Mycotoxin Degradation Enzymes on Alleviating Cytotoxicity of Swine Jejunal Epithelial Cells Induced by Aflatoxin B1 and Zearalenone. Toxins, 11(1), 12. https://doi.org/10.3390/toxins11010012




