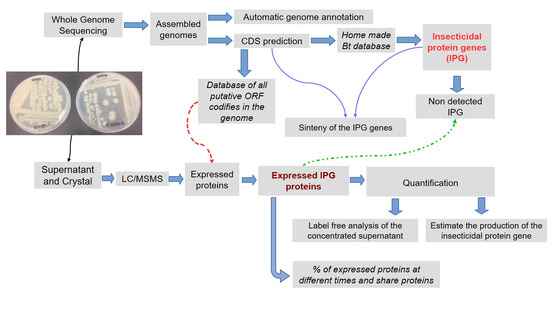
A Genomic and Proteomic Approach to Identify and Quantify the Expressed Bacillus thuringiensis Proteins in the Supernatant and Parasporal Crystal
Abstract
- Genomic and proteomic analysis can identify insecticidal proteins genes and quantify their expressed products.
- The Bt isolates E-SE10.2 and O-V84.2 produce five new Vip-like and Sip-like proteins and eight new Cry-like proteins
1. Introduction
2. Results
2.1. Identification of Vip1-, Vip2-, and Vip4-Type Genes
2.2. Genome Sequencing of the Bt Isolates E-SE10.2 and O-V84.2, Contig Assembly and Gene Annotation
2.3. Global Analysis of the Proteins Identified by in Gel Digestion LC/MSMS Analysis of the Bt Isolates E-SE10.2 and O-V84.2
2.4. Protein Identification of the Expressed Predicted Putative Insecticidal Protein Genes
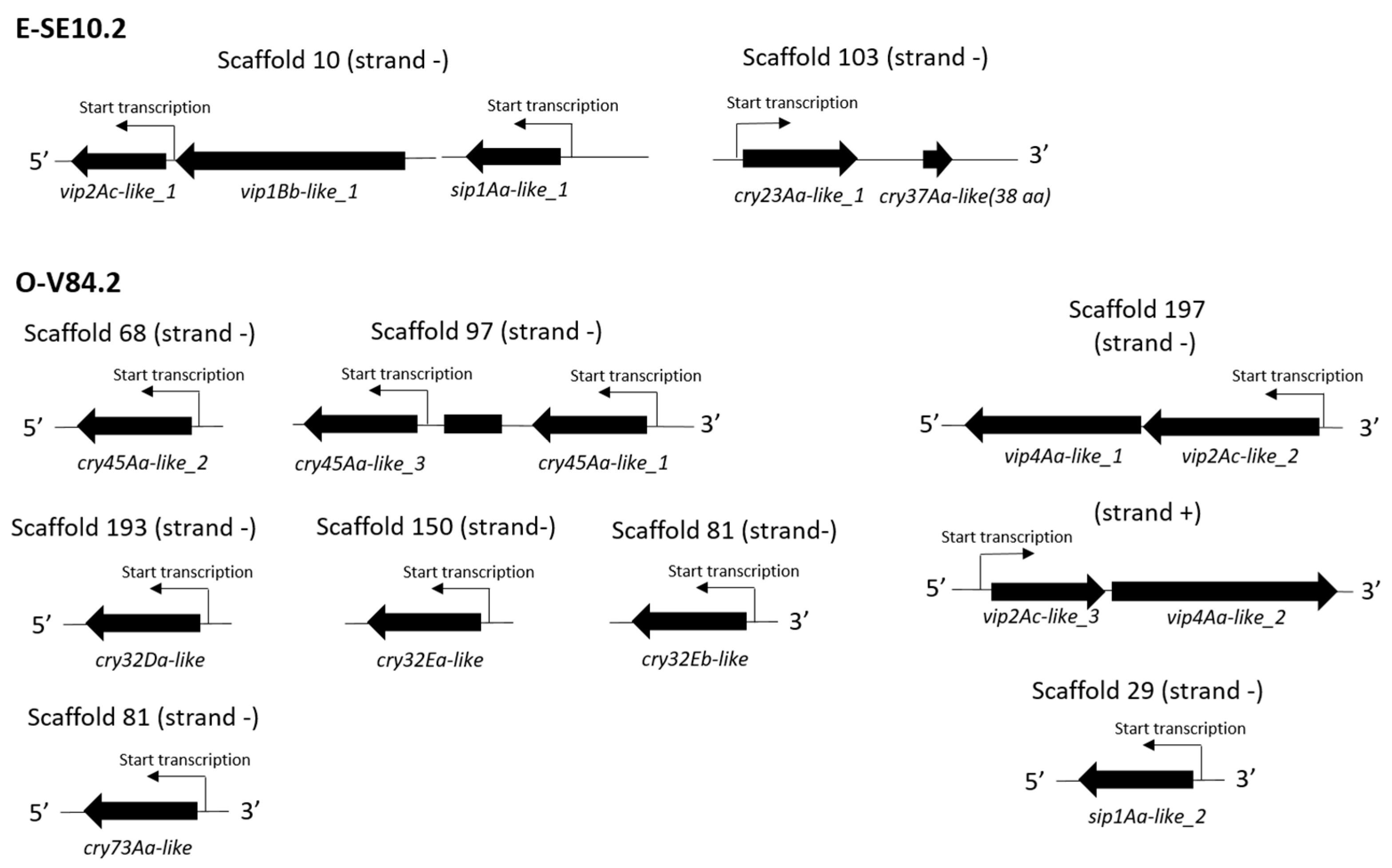
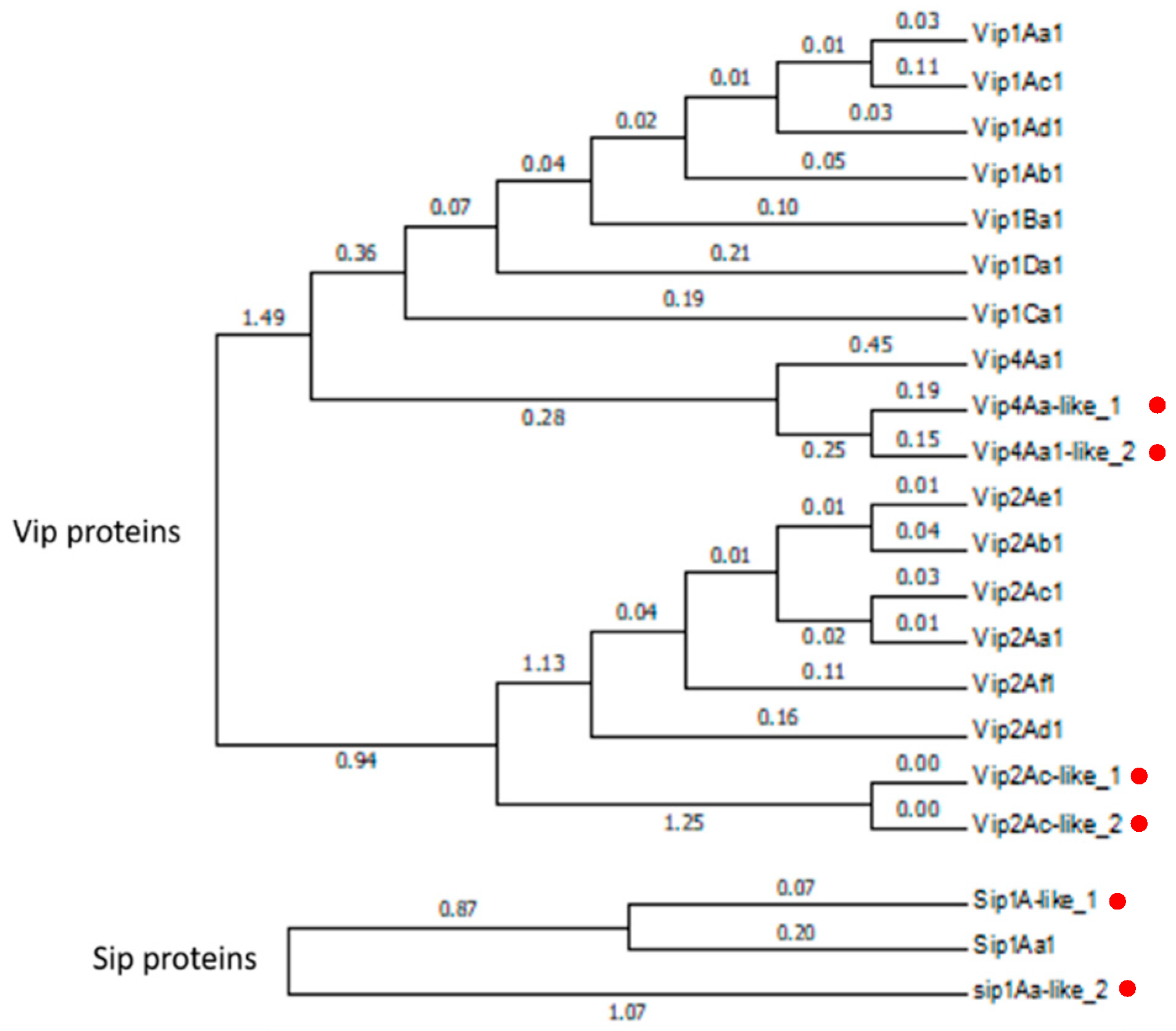
2.5. Gene synteny, Conserved Domains and Phylogenetic Analysis of the Expressed Putative Insecticidal Protein Genes
2.6. Relative Abundance of the Putative Insecticidal Proteins in the Supernatant and in the Crystal of the Bt Isolates E-SE10.2 and O-V84.2
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Bacterial Strains and Growth Conditions for DNA Analysis
5.2. Genomic DNA Preparation
5.3. Identification of Vip1- and Vip2-Type Genes
5.4. Genome Sequencing, Assembly and Annotation Analysis
5.5. Sample Preparation for in Gel Digestion LC/MSMS Analysis and Insecticidal Activity of Bt Isolates
5.6. In Gel Digestion LC/MSMS Analysis
5.7. Protein Identification of the in Gel Digestion LC/MSMS Analysis with Paragon Algorithm and Mascot
5.8. Label Free Analysis of the Concentrated Supernatant 24 h vs. 48 h in Both Bt Isolates
Supplementary Materials
Availability of Data and Material
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Palma, L.; Muñoz, D.; Berry, C.; Murillo, J.; Caballero, P. Bacillus thuringiensis toxins: An overview of their biocidal activity. Toxins 2014, 6, 3296–3325. [Google Scholar] [CrossRef] [PubMed]
- Chakroun, M.; Banyuls, N.; Bel, Y.; Escriche, B.; Ferré, J. Bacterial Vegetative Insecticidal Proteins (Vip) from Entomopathogenic Bacteria. Microbiol. Mol. Biol Rev. 2016, 80, 329–350. [Google Scholar] [CrossRef] [PubMed]
- Ferré, J.; Van Rie, J. Biochemistry and genetics of insect resistance to Bacillus thuringiensis. Ann. Rev. Entomol. 2002, 47, 501–533. [Google Scholar] [CrossRef] [PubMed]
- Ferré, J.; Van Rie, J.; MacIntosh, S.C. Insecticidal genetically modified crops and insect resistance management (IRM). In Integration of Insect-Resistanct Genetically Modified Crops within IPM Programs; Romeis, J., Shelton, A.M., Kennedy, G.G., Eds.; Springer: Dordrecht, The Netherlands, 2008. [Google Scholar]
- Tabashnik, B.E.; Van Rensburg, J.B.J.; Carrière, Y. Field-evolved insect resistance to Bt crops: Definition, theory and data. J. Econ. Entomol. 2009, 102, 2011–2025. [Google Scholar] [CrossRef] [PubMed]
- Tabashnik, B.E. ABCs of Insect Resistance to Bt. PLoS Genet. 2015, 11, e1005646. [Google Scholar] [CrossRef] [PubMed]
- Kalman, S.; Kiehne, K.L.; Libs, J.L.; Yamamoto, T. Cloning of a novel cryIC-type gene from a strain of Bacillus thuringiensis subsp. galleriae. Appl. Environ. Microbiol. 1993, 59, 1131–1137. [Google Scholar] [PubMed]
- Kuo, W.S.; Chak, K.F. Identification of novel cry-type genes from Bacillus thuringiensis strains on the basis of restriction fragment length polymorphism of the PCR-amplified DNA. Appl. Environ. Microbiol. 1996, 62, 1369–1377. [Google Scholar] [PubMed]
- Juarez-Perez, V.M.; Ferrandis, M.D.; Frutos, R. PCR-based approach for detection of novel Bacillus thuringiensis cry genes. Appl. Environ. Microbiol. 1997, 63, 2997–3002. [Google Scholar] [PubMed]
- Lin, Y.; Fang, G.; Peng, K. Characterization of the highly variable cry gene regions of Bacillus thuringiensis strain ly4a3 by PCR-SSCP profiling and sequencing. Biotechnol. Lett. 2007, 29, 247–251. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Rodríguez, C.S.; Boets, A.; Van Rie, J.; Ferré, J. Screening and identification of vip genes in Bacillus thuringiensis. J. Appl. Microbiol. 2009, 107, 219–225. [Google Scholar] [CrossRef] [PubMed]
- Schnepf, H.E.; Whiteley, H.R. Cloning and expression of the Bacillus thuringiensis crystal protein gene in Escherichia coli. Proc. Natl. Acad. Sci. USA 1981, 78, 2893–2897. [Google Scholar] [CrossRef] [PubMed]
- Balasubramanian, P.; Jayakumar, R.; Shambharkar, P.; Unnamalai, N.; Pandian, S.K.; Kumaraswami, N.S.; Ilangovan, R.; Sekar, V. Cloning and characterization of the crystal protein-encoding gene of Bacillus thuringiensis subsp. yunnanensis. Appl. Environ. Microbiol. 2002, 68, 408–411. [Google Scholar] [CrossRef] [PubMed]
- Kongsuwan, K.; Gough, J.; Kemp, D.; McDevitt, A.; Akhurst, R. Characterization of a new Bacillus thuringiensis endotoxin, Cry47Aa, from strains that are toxic to the Australian sheep blowfly, Lucilia cuprina. FEMS Microbiol. Lett. 2005, 252, 127–136. [Google Scholar] [CrossRef] [PubMed]
- Fang, J.; Xu, X.; Wang, P.; Zhao, J.Z.; Shelton, A.M.; Cheng, J.; Feng, M.G.; Shen, Z. Characterization of chimeric Bacillus thuringiensis Vip3 toxins. Appl. Environ. Microbiol. 2003, 73, 956–961. [Google Scholar] [CrossRef] [PubMed]
- Metzker, M.L. Sequencing technologies–The next generation. Nat. Rev. Genet. 2010, 11, 31–46. [Google Scholar] [CrossRef] [PubMed]
- Shendure, J.; Ji, H. Next-generation DNA sequencing. Nat. Biotechnol. 2008, 26, 1135–1145. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.Y.; Ding, X.Z.; Sun, Y.J.; Yang, Q.; Xiao, X.Q.; Cao, Z.P.; Xia, L. Proteomic analysis of Bacillus thuringiensis at different growth phases by using an automated online two-dimensional liquid chromatography tandem mass spectrometry strategy. Appl. Environ. Microb. 2012, 78, 5270–5279. [Google Scholar] [CrossRef] [PubMed]
- Gupta, N.; Benhamida, J.; Bhargava, V.; Goodman, D.; Kain, E.; Kerman, I.; Nguyen, N.; Ollikainen, N.; Rodriguez, J.; Wang, J.; et al. Comparative proteogenomics: Combining massspectrometry and comparative genomics to analyse multiple genomes. Genome Res. 2008, 18, 1133–1142. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.K.; Yang, Y.H.; Chen, Z.; Zhang, J.; Lin, Y.; Wang, Y.; Xiong, Q.; Li, T.; Ge, F.; Bryant, D.A.; et al. Proteogenomic analysis and global discovery of posttranslational Modifications in prokaryotes. Proc. Natl. Acad. Sci. USA 2014, 111, E5633–E5642. [Google Scholar] [CrossRef] [PubMed]
- Quan, M.; Xie, J.; Liu, X.; Li, Y.; Rang, J.; Zhang, T.; Zhou, F.; Xia, L.; Hu, S.; Sun, Y.; et al. Comparative analysis of genomics and proteomics in the new isolates X022 reveals the metabolic regulation mechanism of Carbon flux following Cu2+ Treatment. Front. Microbiol. 2016, 7, 792. [Google Scholar] [CrossRef] [PubMed]
- Rang, J.; He, H.; Wang, T.; Ding, X.; Zuo, M.; Quan, M.; Sun, Y.; Yu, Z.; Hu, S.; Xia, L. Comparative analysis of genomics and proteomic in Bacillus thuringiensis 4.0718. PLoS ONE 2015, 10, e0119065. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Liu, T.; Liang, X.; Tang, C.; Zhu, J.; Wang, S.; Li, S.; Deng, Q.; Wang, L.; Zheng, A.; et al. Rapid detection of vip1-type genes from Bacillus cereus and characterization of a novel vip binary toxin gene. FEMS Microbiol. Lett. 2011, 325, 30. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Luo, R.; Liu, B.; Xie, Y.; Li, Z.; Huang, W.; Yuan, J.; He, G.; Chen, Y.; Pan, Q.; Liu, Y.; et al. SOAPdenovo2: An empirically improved memory-efficient short-read de novo assembler. GigaScience 2012, 1, 18. [Google Scholar] [CrossRef] [PubMed]
- Aziz, R.K.; Bartels, D.; Best, A.A.; DeJongh, M.; Disz, T.; Edwards, R.A.; Formsma, K.; Gerdes, S.; Glass, E.M.; Kubal, M.; et al. The RAST Server: Rapid annotations using subsystems technology. BMC Genom. 2008, 9, 75. [Google Scholar] [CrossRef] [PubMed]
- Marchler-Bauer, A.; Derbyshire, M.K.; Gonzales, N.R.; Lu, S.; Chitsaz, F.; Geer, L.Y.; Geer, R.C.; He, J.; Gwadz, M.; Hurwitz, D.I.; et al. CDD: NCBI’s conserved domain database. Nucleic Acids Res. 2015, 43, 222–226. [Google Scholar] [CrossRef] [PubMed]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Shilov, I.V.; Seymour, S.L.; Patel, A.A.; Loboda, A.; Tang, W.H.; Keating, S.P.; Hunter, C.L.; Nuwaysir, L.M.; Schaeffer, D.A. The Paragon Algorithm, a next generation search engine that uses sequence temperature values and feature probabilities to identify peptides from tandem mass spectra. Mol. Cell. Proteom. 2007, 6, 1638–1655. [Google Scholar] [CrossRef] [PubMed]
- Ishihama, Y.; Oda, Y.; Tabata, T.; Sato, T.; Nagasu, T.; Rappsilber, J.; Mann, M. Exponentially Modified Protein Abundance Index (emPAI) for Estimation of Absolute Protein Amount in Proteomics by the Number of Sequenced Peptides per Protein. Mol. Cell. Proteom. 2005, 4, 1265–1272. [Google Scholar] [CrossRef] [PubMed]
- Conesa, A.; Götz, S.; García-Gómez, J.M.; Terol, J.; Talón, M.; Robles, M. Blast2GO: A universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 2005, 21, 3674–3676. [Google Scholar] [CrossRef] [PubMed]
| Name of Isolate | Identified with vip1 Primers | Identified with vip2 Primers | ||||
|---|---|---|---|---|---|---|
| Similarity (%) ‡ | Coverage (%) * | Closest Homolog | Similarity (%) ‡ | Coverage (%) * | Closest Homolog | |
| V-J20.2 | 100 | 44 | vip1Bb1 | 97 | 70 | vip2Bb1 |
| V-LE1.1 | 100 | 40 | vip1Bb1 | 99 | 72 | vip2Bb1 |
| V-V54.26 | 99 | 49 | vip1Ca1 | 99 | 71 | vip2Ac1 |
| V-V54.31 | 100 | 49 | vip1Ca1 | 98 | 73 | vip2Ac1 |
| E-SE10.2 | 62 | 30 | vip1Bb3 | No DNA amplification | ||
| E-TE7.43 | 100 | 49 | vip1Ca1 | 99 | 64 | vip2Ac1 |
| E-TE16.5 | 98 | 43 | vip1Ca1 | 98 | 73 | vip2Ac1 |
| E-TE18.40 | 100 | 30 | vip1Ca1 | 100 | 45 | vip2Ac1 |
| O-V84.2 | 40 | 40 | vip4Aa1 | No DNA amplification | ||
| Features | E-SE10.2 | O-V84.2 | ||
|---|---|---|---|---|
| Gene Content | Length (Mb) | Gene Content | Length (Mb) | |
| Genome Content * | 6216 | 6.1 | 6525 | 6.3 |
| Coding sequences ‡ | 6156 (99%) | 4.8 | 6457 (98.9%) | 5 |
| Annotated genes | 4398 (70%) | 4.05 | 4615 (71.4%) | 4.21 |
| Hypothetical genes | 1758 (28.2%) | 0.75 | 1842 (28.2%) | 0.79 |
| Predicted insecticidal genes § | 6 | 0.002 | 18 | 0.4 |
| tRNAs | 60 | 0.004 | 68 | 0.005 |
| Sample | Gene Identity ‡ | Closest Homolog * | Similarity (%) | Coverage (%) |
|---|---|---|---|---|
| E-SE10.2 | vip1Ad-like_1 | AGC08395.1 | 55 | 24 |
| vip1Bb-like_1 | AAR40282.1 | 61 | 99 | |
| vip2Aa-like | 1QS1_A | 41 | 23 | |
| vip2Ac-like_1 | AAO86513.1 | 30 | 30 | |
| sip1Aa-like_1 | ABC71340.1 | 75 | 98 | |
| cry23Aa-like | AAF76375.1 | 75 | 98 | |
| O-V84.2 | vip1Ad-like_2 | AGC08395.1 | 26 | 34 |
| vip1Ba-like | AAR40886.1 | 28 | 30 | |
| vip1Da-like | CAI40767.1 | 37 | 12 | |
| vip2Ac-like_2 | AAO86513.1 | 33 | 47 | |
| vip2Ac-like_3 | AAO86513.1 | 37 | 41 | |
| vip2Bb-like | AKI69695.1 | 30 | 43 | |
| vip4Aa1-like_1 | AEB52299.1 | 40 | 80 | |
| vip4Aa1-like_2 | AEB52299.1 | 40 | 83 | |
| vip4Aa-like_3 | AEB52299.1 | 49 | 94 | |
| vip4Aa-like_4 | AEB52299.1 | 52 | 97 | |
| sip1Aa-like_3 | ABC71340.1 | 32 | 33 | |
| cry45Aa-like_1 | BAD22577.1 | 61 | 100 | |
| cry45Aa-like_2 | BAD22577.1 | 69 | 99 | |
| cry45Aa-like_3 | BAD22577.1 | 68 | 85 | |
| cry32Ea-like | ADK66923.1 | 47 | 98 | |
| cry32Eb-like | AGU13828.1 | 51 | 41 | |
| cry32Da-like | BAB78603.1 | 40 | 98 | |
| cry73Aa-like | AEH76822.1 | 88 | 80 |
| Sample | Protein Identity | Mass Protein (kDa) | SignalIP Server 4.1 | Supernatant (LB) | Spore/Crystal Mixture (CCY) | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 24 h | 48 h | 72 h | ||||||||||
| Rep. 1 | Rep. 2 | Rep. 3 | Rep. 1 | Rep. 2 | Rep. 3 | Rep. 1 | Rep. 2 | Rep. 3 | ||||
| E-SE10.2 | Vip2Ac-like_1 | 51.6 | Yes | +/+ | +/+ | +/+ | +/+ | +/+ | +/+ | −/− | −/− | −/− |
| Sip1Aa-like_1 | 40.7 | Yes | +/+ | +/+ | +/+ | +/+ | +/+ | +/+ | −/− | −/− | −/− | |
| Cry23Aa-like | 29.3 | No | +/+ | +/+ | +/+ | +/+ | +/+ | +/+ | +/+ | +/+ | +/+ | |
| O-V84.2 | Vip4Aa-like_1 | 97.5 | Yes | +/+ | +/+ | +/+ | +/+ | +/+ | +/+ | −/− | −/− | −/− |
| Vip2Ac-like_2 | 80.79 | Yes | +/+ | +/+ | +/+ | −/+ | +/+ | +/+ | −/− | −/− | −/− | |
| Vip4Aa-like_2 | 87.5 | Yes | +/+ | +/+ | −/+ | −/− | −/− | −/− | −/− | −/− | −/− | |
| Vip2Ac-like_3 | 23.2 | Yes | −/− | −/+ | −/− | −/− | −/− | −/− | −/− | −/− | −/− | |
| Sip1Aa-like_2 | 38.7 | Yes | −/− | −/+ | +/+ | −/− | −/− | −/− | −/− | −/− | −/− | |
| Cry45Aa-like_1 | 30.6 | No | −/− | −/− | −/− | −/− | −/− | −/− | +/+ | +/+ | +/+ | |
| Cry45Aa-like_2 | 29.3 | No | −/− | −/− | −/− | −/− | −/− | −/− | +/+ | +/+ | +/+ | |
| Cry45Aa-like_3 | 25.6 | No | −/− | −/− | −/− | −/− | −/− | −/− | +/+ | +/+ | +/+ | |
| Cry32Ea-like | 151.2 | No | −/− | −/− | −/− | −/− | −/− | −/− | +/+ | +/+ | +/+ | |
| Cry32Da-like | 153.7 | No | −/− | −/− | −/− | −/− | −/− | −/− | +/+ | +/+ | +/+ | |
| Cry32Eb-like | 76.8 | No | −/− | −/− | −/− | −/− | −/− | −/− | +/+ | +/+ | +/+ | |
| Cry73Aa-like | 72.2 | No | −/− | −/− | −/− | −/− | −/− | −/− | +/+ | +/+ | +/+ | |
| Supernatant | 24 h (% Weight) | 48 h (% Weight) | ||||
|---|---|---|---|---|---|---|
| R1 | R2 | R3 | R1 | R2 | R3 | |
| E-SE10.2 | ||||||
| Non Secretable toxins | 98.56 | 99.06 | 99.36 | 97.87 | 99.26 | 99.63 |
| Flagellin protein FlaA | 55.21 | 73.82 | 87.19 | 39.75 | 36.04 | 43.19 |
| Secretable toxins | 1.44 | 0.94 | 0.64 | 2.13 | 0.74 | 0.37 |
| Vip2Ac-like_1 | 0.09 | 0.06 | 0.03 | 0.09 | 0.10 | 0.06 |
| Sip1A-like_1 | 0.09 | 0.03 | 0.02 | 0.06 | 0.08 | 0.02 |
| Cry23Aa-like * | 1.26 | 0.85 | 0.59 | 1.98 | 0.56 | 0.29 |
| O-V84.2 | ||||||
| Non Secretable toxins | 99.99 | 99.99 | 99.99 | 99.99 | 99.99 | 99.99 |
| Flagellin protein FlaA | 99.20 | 99.37 | 98.35 | 84.14 | 99.15 | 88.35 |
| Secretable toxins | 0.0032 | 0.0012 | 0.0030 | 0.0111 | 0.00007 | 0.0075 |
| Vip4Aa-like_1 | 0.0014 | 0.0008 | 0.0016 | 0.0019 | 0.00004 | 0.0029 |
| Vip4Aa-like_2 | 0.0005 | 0.0001 | 0.0002 | - | - | - |
| Vip2Ac-like_2 | 0.0013 | 0.0002 | 0.0010 | 0.0092 | 0.00003 | 0.0046 |
| Crystal | 72 h (% Weight) | |||||
| R1 | R2 | R3 | ||||
| E-SE10.2 | ||||||
| Non-crystal toxins | 69.52 | 95.26 | 97.51 | |||
| Crystal toxins | 30.48 | 4.74 | 2.49 | |||
| Cry23A-like | 30.48 | 4.74 | 2.49 | |||
| O-V84.2 | ||||||
| Non-crystal toxins | 52.86 | 51.12 | 53.25 | |||
| Crystal toxins | 47.14 | 48.88 | 46.75 | |||
| Cry45Aa-like_1 | 2.82 | 2.46 | 1.41 | |||
| Cry45Aa-like_2 | 3.05 | 2.03 | 3.38 | |||
| Cry45Aa-like_3 | 1.88 | 5.29 | 2.20 | |||
| Cry32Ea-like | 24.30 | 25.06 | 25.89 | |||
| Cry32Da-like | 6.10 | 5.40 | 4.60 | |||
| Cry32Eb-like | 6.20 | 4.46 | 5.02 | |||
| Cry73Aa-like | 2.79 | 4.18 | 4.25 | |||
| Bt Isolate | Proteins | t-Value † | p-Value ᶲ | Mean Peaks Area | Standard Deviation Peaks Area | Fold Change 24/48 § | Status | ||
|---|---|---|---|---|---|---|---|---|---|
| 24 h | 48 h | 24 h | 48 h | ||||||
| E-SE10.2 | Vip2Ac-like_1 | 0.71 | 0.52 | 307,838 | 266,602 | 54,005 | 84,577 | 1.15 | No differences |
| Sip1Aa-like_1 | 0.89 | 0.42 | 110,172 | 76,938 | 47,032 | 44,040 | 1.43 | No differences | |
| Cry23Aa-like * | 0.32 | 0.77 | 5,796,029 | 4,858,544 | 4,951,202 | 1,257,383 | 1.19 | No differences | |
| O-V84.2 | Vip4Aa-like_1 | 4.07 | 0.04 | 134,357 | 68,825 | 26,801 | 7636 | 1.95 | Increased |
| Vip2Ac-like_2 | 0.56 | 0.61 | 32,544 | 27,573 | 13,419 | 7512 | 1.18 | No differences | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gomis-Cebolla, J.; Scaramal Ricietto, A.P.; Ferré, J. A Genomic and Proteomic Approach to Identify and Quantify the Expressed Bacillus thuringiensis Proteins in the Supernatant and Parasporal Crystal. Toxins 2018, 10, 193. https://doi.org/10.3390/toxins10050193
Gomis-Cebolla J, Scaramal Ricietto AP, Ferré J. A Genomic and Proteomic Approach to Identify and Quantify the Expressed Bacillus thuringiensis Proteins in the Supernatant and Parasporal Crystal. Toxins. 2018; 10(5):193. https://doi.org/10.3390/toxins10050193
Chicago/Turabian StyleGomis-Cebolla, Joaquín, Ana Paula Scaramal Ricietto, and Juan Ferré. 2018. "A Genomic and Proteomic Approach to Identify and Quantify the Expressed Bacillus thuringiensis Proteins in the Supernatant and Parasporal Crystal" Toxins 10, no. 5: 193. https://doi.org/10.3390/toxins10050193
APA StyleGomis-Cebolla, J., Scaramal Ricietto, A. P., & Ferré, J. (2018). A Genomic and Proteomic Approach to Identify and Quantify the Expressed Bacillus thuringiensis Proteins in the Supernatant and Parasporal Crystal. Toxins, 10(5), 193. https://doi.org/10.3390/toxins10050193