Vitamin D in the Context of Evolution
Abstract
1. Introduction
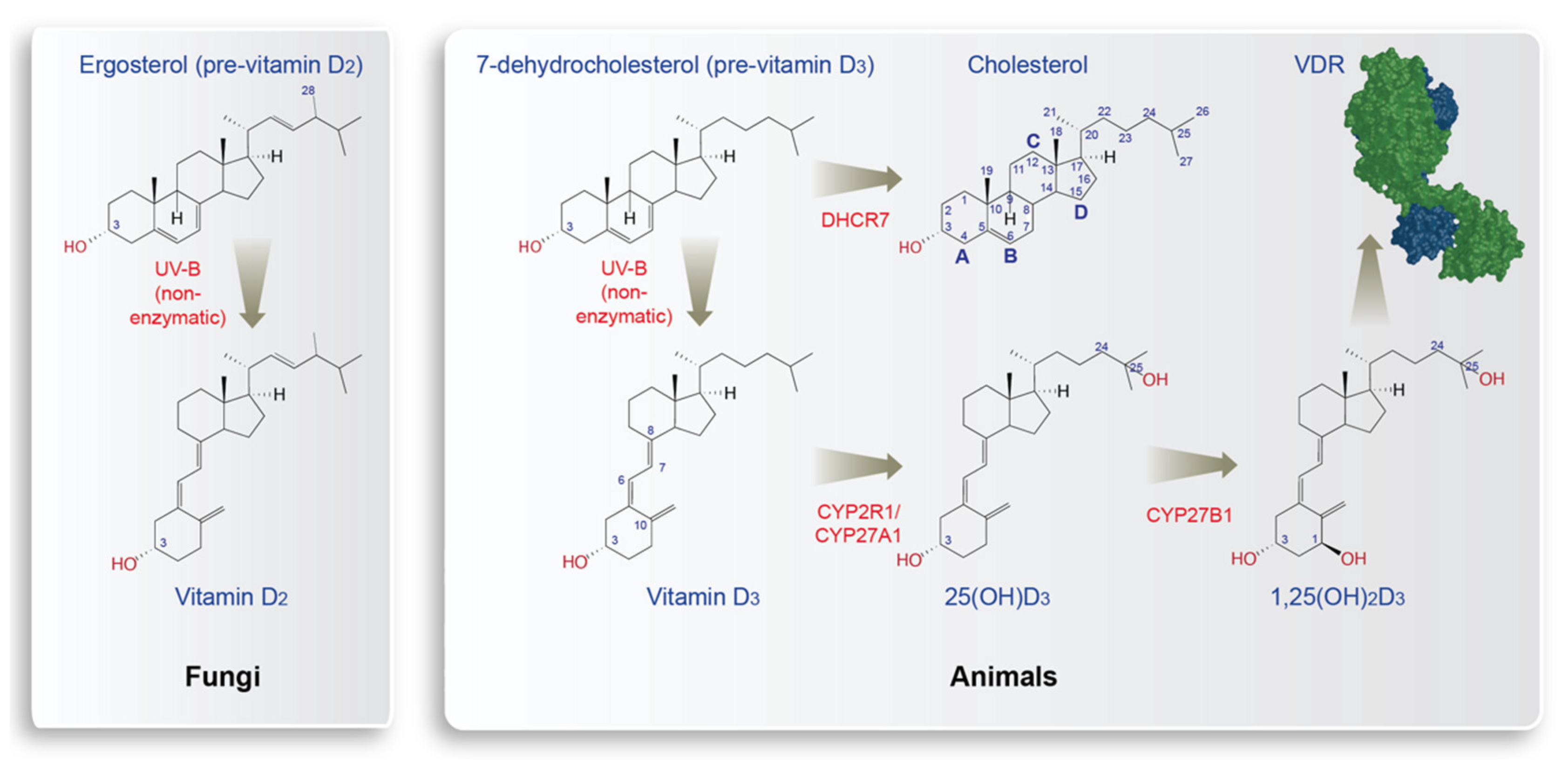
2. Sterols and Vitamin D Synthesis
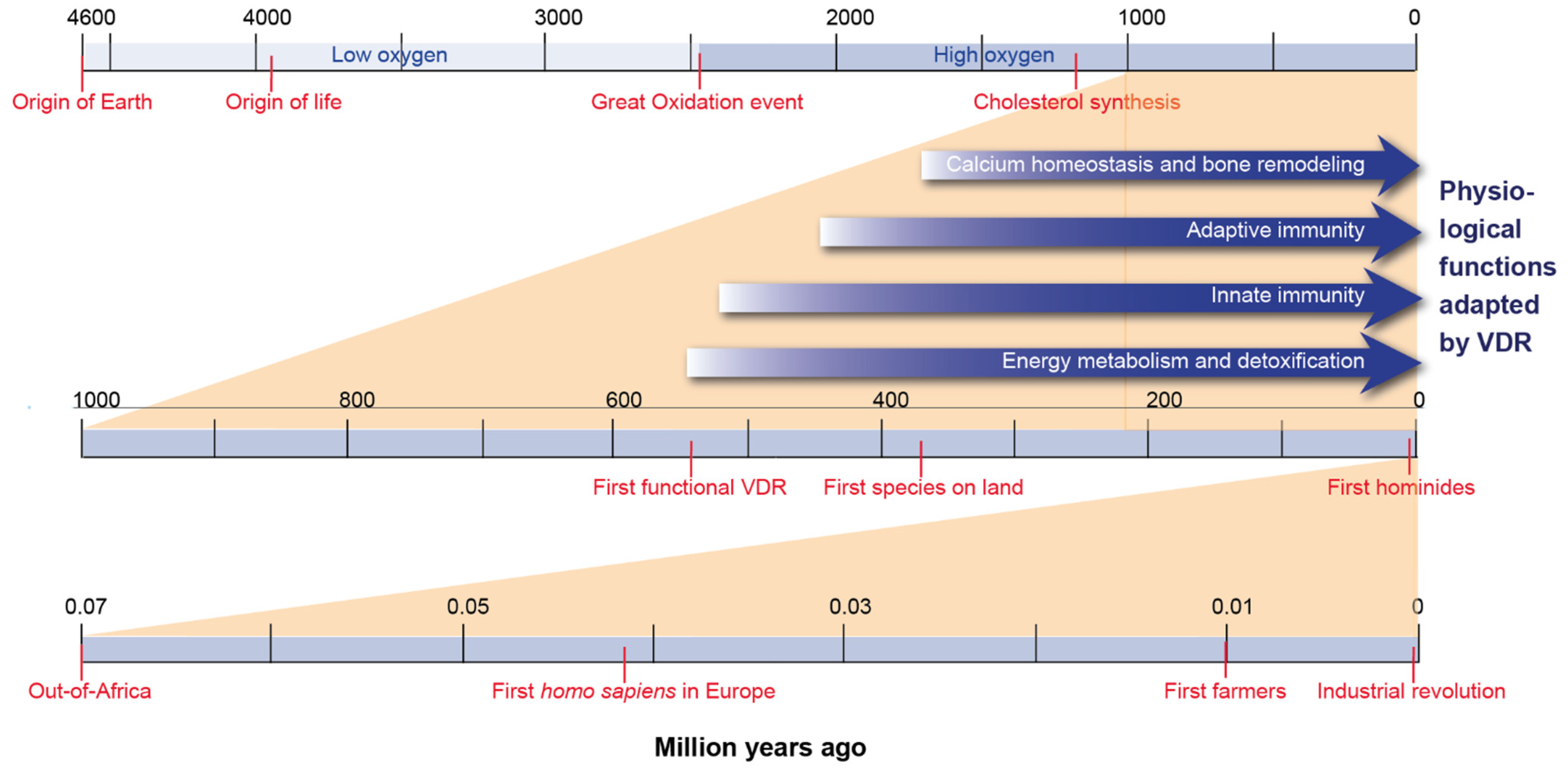
3. Evolution of Vitamin D Endocrinology
4. Evolution of the Physiological Functions of Vitamin D
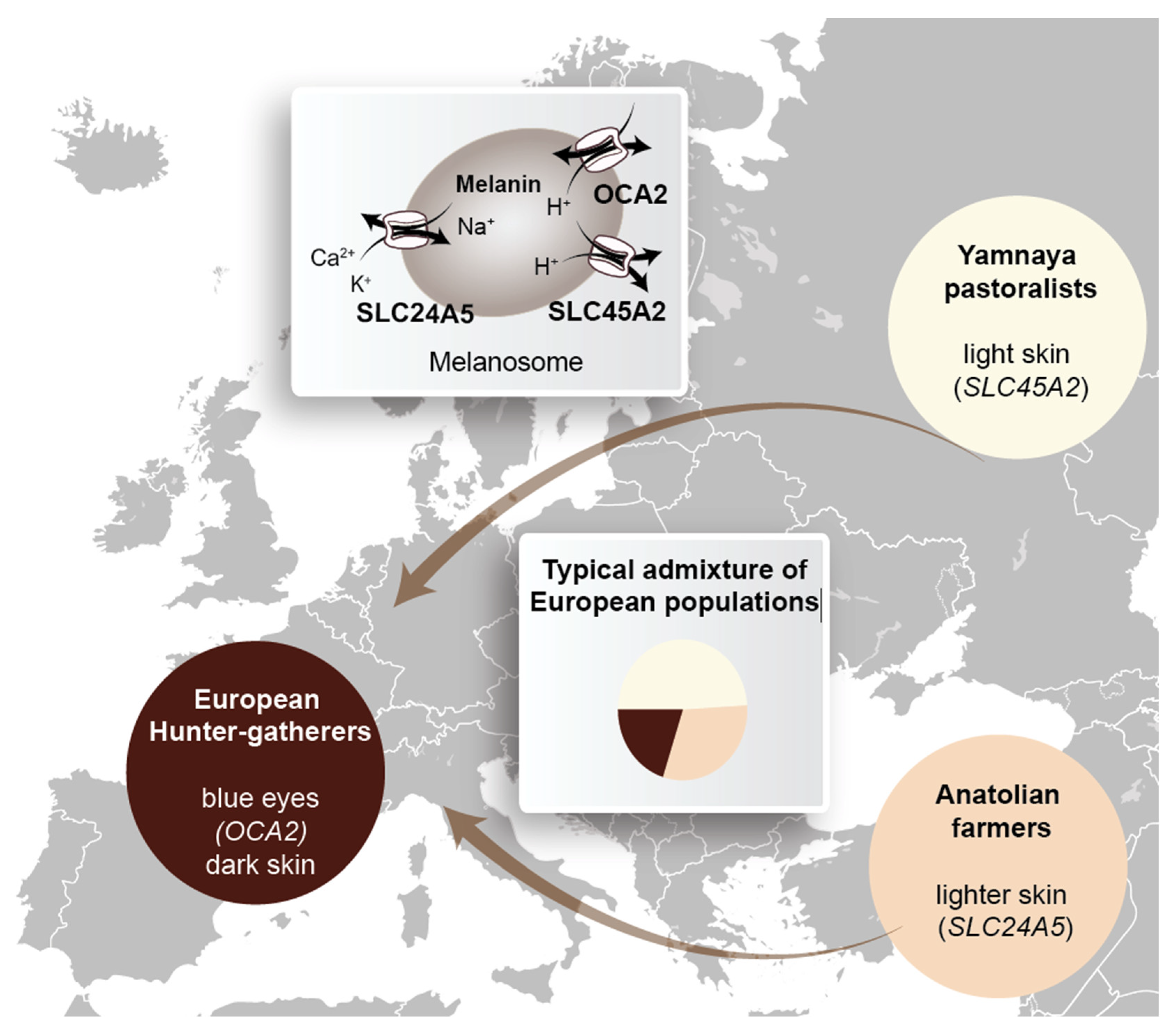
5. How Does the Evolution of Homo sapiens Relate to Vitamin D?
6. How Vitamin D Became a Vitamin?
7. Conclusions
Funding
Conflicts of Interest
References
- Dobzhansky, T. Nothing in biology makes sense except in the light of evolution. Am. Biol. Teach. 1973, 35, 125–129. [Google Scholar] [CrossRef]
- McMollum, E.V.; Simmonds, N.; Becker, J.E.; Shipley, P.G. Studies on experimental rickets: An experimental demonstration of the existence of a vitamin which promotes calcium deposition. J. Biol. Chem. 1922, 52, 293–298. [Google Scholar] [CrossRef]
- Levine, M.A. Diagnosis and management of vitamin D dependent rickets. Front. Pediatr. 2020, 8, 315. [Google Scholar] [CrossRef] [PubMed]
- Bouillon, R.; Carmeliet, G.; Lieben, L.; Watanabe, M.; Perino, A.; Auwerx, J.; Schoonjans, K.; Verstuyf, A. Vitamin D and energy homeostasis-of mice and men. Nat. Rev. Endocrinol. 2014, 10, 79–87. [Google Scholar] [CrossRef] [PubMed]
- Hanel, A.; Carlberg, C. Skin color and vitamin D: An update. Exp. Dermatol. 2020, 29, 864–875. [Google Scholar] [CrossRef] [PubMed]
- Summons, R.E.; Bradley, A.S.; Jahnke, L.L.; Waldbauer, J.R. Steroids, triterpenoids and molecular oxygen. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2006, 361, 951–968. [Google Scholar] [CrossRef]
- Sessions, A.L.; Doughty, D.M.; Welander, P.V.; Summons, R.E.; Newman, D.K. The continuing puzzle of the Great Oxidation Event. Curr. Biol. 2009, 19, R567–R574. [Google Scholar] [CrossRef]
- Raymond, J.; Segre, D. The effect of oxygen on biochemical networks and the evolution of complex life. Science 2006, 311, 1764–1767. [Google Scholar] [CrossRef]
- Desmond, E.; Gribaldo, S. Phylogenomics of sterol synthesis: Insights into the origin, evolution, and diversity of a key eukaryotic feature. Genome Biol. Evol. 2009, 1, 364–381. [Google Scholar] [CrossRef]
- DeBose-Boyd, R.A. Feedback regulation of cholesterol synthesis: Sterol-accelerated ubiquitination and degradation of HMG CoA reductase. Cell Res. 2008, 18, 609–621. [Google Scholar] [CrossRef]
- Japelt, R.B.; Jakobsen, J. Vitamin D in plants: A review of occurrence, analysis, and biosynthesis. Front. Plant Sci. 2013, 4, 136. [Google Scholar] [CrossRef] [PubMed]
- Martin-Creuzburg, D.; Merkel, P. Sterols of freshwater microalgae: Potential implications for zooplankton nutrition. J. Plankton Res. 2016, 38, 865–877. [Google Scholar] [CrossRef]
- Head, B.P.; Patel, H.H.; Insel, P.A. Interaction of membrane/lipid rafts with the cytoskeleton: Impact on signaling and function: Membrane/lipid rafts, mediators of cytoskeletal arrangement and cell signaling. Biochim. Biophys. Acta 2014, 1838, 532–545. [Google Scholar] [CrossRef]
- Capell-Hattam, I.M.; Brown, A.J. Sterol evolution: Cholesterol synthesis in animals is less a required trait than an acquired taste. Curr. Biol. 2020, 30, R886–R888. [Google Scholar] [CrossRef]
- Holick, M.F. The cutaneous photosynthesis of previtamin D3: A unique photoendocrine system. J. Investig. Dermatol. 1981, 77, 51–58. [Google Scholar] [CrossRef] [PubMed]
- Holick, M.F. Vitamin D: A millenium perspective. J. Cell Biochem. 2003, 88, 296–307. [Google Scholar] [CrossRef] [PubMed]
- Holick, M.F.; MacLaughlin, J.A.; Doppelt, S.H. Regulation of cutaneous previtamin D3 photosynthesis in man: Skin pigment is not an essential regulator. Science 1981, 211, 590–593. [Google Scholar] [CrossRef]
- Wacker, M.; Holick, M.F. Sunlight and vitamin D: A global perspective for health. Dermatoendocrinol 2013, 5, 51–108. [Google Scholar] [CrossRef]
- Boyce, D.G.; Lewis, M.R.; Worm, B. Global phytoplankton decline over the past century. Nature 2010, 466, 591–596. [Google Scholar] [CrossRef]
- Byrdwell, W.C.; Horst, R.L.; Phillips, K.M.; Holden, J.M.; Patterson, K.Y.; Harnly, J.M.; Exler, J. Vitamin D levels in fish and shellfish determined by liquid chromatography with ultraviolet detection and mass spectrometry. J. Food Compos. Anal. 2013, 30, 109–119. [Google Scholar] [CrossRef]
- Rajakumar, K. Vitamin D, cod-liver oil, sunlight, and rickets: A historical perspective. Pediatrics 2003, 112, e132–e135. [Google Scholar] [CrossRef] [PubMed]
- Whitfield, G.K.; Dang, H.T.; Schluter, S.F.; Bernstein, R.M.; Bunag, T.; Manzon, L.A.; Hsieh, G.; Dominguez, C.E.; Youson, J.H.; Haussler, M.R.; et al. Cloning of a functional vitamin D receptor from the lamprey (Petromyzon marinus), an ancient vertebrate lacking a calcified skeleton and teeth. Endocrinology 2003, 144, 2704–2716. [Google Scholar] [CrossRef] [PubMed]
- Hanel, A.; Bendik, I.; Carlberg, C. Transcriptome-wide profile of 25-hydroxyvitamin D3 in pimary immune cells from human peripheral blood. Nutrients 2021, 13, 4100. [Google Scholar] [CrossRef] [PubMed]
- Jones, G.; Prosser, D.E.; Kaufmann, M. Cytochrome P450-mediated metabolism of vitamin D. J. Lipid. Res. 2014, 55, 13–31. [Google Scholar] [CrossRef]
- Bouillon, R.; Suda, T. Vitamin D: Calcium and bone homeostasis during evolution. BoneKEy Rep. 2014, 3, 480. [Google Scholar] [CrossRef]
- Escriva, H.; Bertrand, S.; Laudet, V. The evolution of the nuclear receptor superfamily. Essays Biochem. 2004, 40, 11–26. [Google Scholar] [CrossRef]
- Krasowski, M.D.; Ni, A.; Hagey, L.R.; Ekins, S. Evolution of promiscuous nuclear hormone receptors: LXR, FXR, VDR, PXR, and CAR. Mol. Cell Endocrinol. 2011, 334, 39–48. [Google Scholar] [CrossRef]
- Reschly, E.J.; Bainy, A.C.; Mattos, J.J.; Hagey, L.R.; Bahary, N.; Mada, S.R.; Ou, J.; Venkataramanan, R.; Krasowski, M.D. Functional evolution of the vitamin D and pregnane X receptors. BMC Evol. Biol. 2007, 7, 222. [Google Scholar] [CrossRef]
- Hanel, A.; Carlberg, C. Vitamin D and evolution: Pharmacologic implications. Biochem. Pharmacol. 2020, 173, 113595. [Google Scholar] [CrossRef]
- Makishima, M.; Lu, T.T.; Xie, W.; Whitfield, G.K.; Domoto, H.; Evans, R.M.; Haussler, M.R.; Mangelsdorf, D.J. Vitamin D receptor as an intestinal bile acid sensor. Science 2002, 296, 1313–1316. [Google Scholar] [CrossRef]
- Makishima, M.; Okamoto, A.Y.; Repa, J.J.; Tu, H.; Learned, R.M.; Luk, A.; Hull, M.V.; Lustig, K.D.; Mangelsdorf, D.J.; Shan, B. Identification of a nuclear receptor for bile acids. Science 1999, 284, 1362–1365. [Google Scholar] [CrossRef] [PubMed]
- Staudinger, J.L.; Goodwin, B.; Jones, S.A.; Hawkins-Brown, D.; MacKenzie, K.I.; LaTour, A.; Liu, Y.; Klaassen, C.D.; Brown, K.K.; Reinhard, J.; et al. The nuclear receptor PXR is a lithocholic acid sensor that protects against liver toxicity. Proc. Natl. Acad. Sci. USA 2001, 98, 3369–3374. [Google Scholar] [CrossRef] [PubMed]
- Guo, G.L.; Lambert, G.; Negishi, M.; Ward, J.M.; Brewer, H.B., Jr.; Kliewer, S.A.; Gonzalez, F.J.; Sinal, C.J. Complementary roles of farnesoid X receptor, pregnane X receptor, and constitutive androstane receptor in protection against bile acid toxicity. J. Biol. Chem. 2003, 278, 45062–45071. [Google Scholar] [CrossRef] [PubMed]
- Reschly, E.J.; Krasowski, M.D. Evolution and function of the NR1I nuclear hormone receptor subfamily (VDR, PXR, and CAR) with respect to metabolism of xenobiotics and endogenous compounds. Curr. Drug Metab. 2006, 7, 349–365. [Google Scholar] [CrossRef] [PubMed]
- Carlberg, C. Nutrigenomics of vitamin D. Nutrients 2019, 11, 676. [Google Scholar] [CrossRef]
- Müller, M.; Kersten, S. Nutrigenomics: Goals and strategies. Nat. Rev. Genet. 2003, 4, 315–322. [Google Scholar] [CrossRef]
- Muller, M.; Mentel, M.; van Hellemond, J.J.; Henze, K.; Woehle, C.; Gould, S.B.; Yu, R.Y.; van der Giezen, M.; Tielens, A.G.; Martin, W.F. Biochemistry and evolution of anaerobic energy metabolism in eukaryotes. Microbiol. Mol. Biol. Rev. 2012, 76, 444–495. [Google Scholar] [CrossRef]
- Seuter, S.; Neme, A.; Carlberg, C. Epigenome-wide effects of vitamin D and their impact on the transcriptome of human monocytes involve CTCF. Nucleic Acids Res. 2016, 44, 4090–4104. [Google Scholar] [CrossRef]
- Pasing, Y.; Fenton, C.G.; Jorde, R.; Paulssen, R.H. Changes in the human transcriptome upon vitamin D supplementation. J. Steroid Biochem. Mol. Biol. 2017, 173, 93–99. [Google Scholar] [CrossRef]
- Ferreira, G.B.; Vanherwegen, A.S.; Eelen, G.; Gutierrez, A.C.; Van Lommel, L.; Marchal, K.; Verlinden, L.; Verstuyf, A.; Nogueira, T.; Georgiadou, M.; et al. Vitamin D3 induces tolerance in human dendritic cells by activation of intracellular metabolic pathways. Cell Rep. 2015, 10, 711–725. [Google Scholar] [CrossRef]
- Hanel, A.; Carlberg, C. Time-resolved gene expression analysis monitors the regulation of inflammatory mediators and attenuation of adaptive immune response by vitamin D. Int. J. Mol. Sci. 2022, 23, 911. [Google Scholar] [CrossRef] [PubMed]
- Mullican, S.E.; Dispirito, J.R.; Lazar, M.A. The orphan nuclear receptors at their 25-year reunion. J. Mol. Endocrinol. 2013, 51, T115–T140. [Google Scholar] [CrossRef]
- Sigueiro, R.; Bianchetti, L.; Peluso-Iltis, C.; Chalhoub, S.; Dejaegere, A.; Osz, J.; Rochel, N. Advances in vitamin D receptor function and evolution based on the 3D structure of the Lamprey ligand-binding domain. J. Med. Chem. 2022, 65, 5821–5829. [Google Scholar] [CrossRef]
- Rochel, N.; Wurtz, J.M.; Mitschler, A.; Klaholz, B.; Moras, D. Crystal structure of the nuclear receptor for vitamin D bound to its natural ligand. Mol. Cell 2000, 5, 173–179. [Google Scholar] [CrossRef]
- Kollitz, E.M.; Zhang, G.; Hawkins, M.B.; Whitfield, G.K.; Reif, D.M.; Kullman, S.W. Molecular cloning, functional characterization, and evolutionary analysis of vitamin D receptors isolated from basal vertebrates. PLoS ONE 2015, 10, e0122853. [Google Scholar] [CrossRef] [PubMed]
- Kollitz, E.M.; Hawkins, M.B.; Whitfield, G.K.; Kullman, S.W. Functional diversification of vitamin D receptor paralogs in teleost fish after a whole genome duplication event. Endocrinology 2014, 155, 4641–4654. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Flajnik, M.F. A cold-blooded view of adaptive immunity. Nat. Rev. Immunol. 2018, 18, 438–453. [Google Scholar] [CrossRef]
- Cooper, M.D.; Alder, M.N. The evolution of adaptive immune systems. Cell 2006, 124, 815–822. [Google Scholar] [CrossRef]
- Ganeshan, K.; Nikkanen, J.; Man, K.; Leong, Y.A.; Sogawa, Y.; Maschek, J.A.; Van Ry, T.; Chagwedera, D.N.; Cox, J.E.; Chawla, A. Energetic trade-offs and hypometabolic states promote disease tolerance. Cell 2019, 177, 399–413.e12. [Google Scholar] [CrossRef]
- Vanherwegen, A.S.; Eelen, G.; Ferreira, G.B.; Ghesquiere, B.; Cook, D.P.; Nikolic, T.; Roep, B.; Carmeliet, P.; Telang, S.; Mathieu, C.; et al. Vitamin D controls the capacity of human dendritic cells to induce functional regulatory T cells by regulation of glucose metabolism. J. Steroid Biochem. Mol. Biol. 2019, 187, 134–145. [Google Scholar] [CrossRef]
- Heikkinen, S.; Väisänen, S.; Pehkonen, P.; Seuter, S.; Benes, V.; Carlberg, C. Nuclear hormone 1α,25-dihydroxyvitamin D3 elicits a genome-wide shift in the locations of VDR chromatin occupancy. Nucleic Acids Res. 2011, 39, 9181–9193. [Google Scholar] [CrossRef] [PubMed]
- Gombart, A.F.; Borregaard, N.; Koeffler, H.P. Human cathelicidin antimicrobial peptide (CAMP) gene is a direct target of the vitamin D receptor and is strongly up-regulated in myeloid cells by 1,25-dihydroxyvitamin D3. FASEB J. 2005, 19, 1067–1077. [Google Scholar] [CrossRef] [PubMed]
- Bscheider, M.; Butcher, E.C. Vitamin D immunoregulation through dendritic cells. Immunology 2016, 148, 227–236. [Google Scholar] [CrossRef] [PubMed]
- Coussens, A.K.; Wilkinson, R.J.; Hanifa, Y.; Nikolayevskyy, V.; Elkington, P.T.; Islam, K.; Timms, P.M.; Venton, T.R.; Bothamley, G.H.; Packe, G.E.; et al. Vitamin D accelerates resolution of inflammatory responses during tuberculosis treatment. Proc. Natl. Acad. Sci. USA 2012, 109, 15449–15454. [Google Scholar] [CrossRef] [PubMed]
- Carlberg, C.; Seuter, S.; Nurmi, T.; Tuomainen, T.P.; Virtanen, J.K.; Neme, A. In vivo response of the human epigenome to vitamin D: A proof-of-principle study. J. Steroid Biochem. Mol. Biol. 2018, 180, 142–148. [Google Scholar] [CrossRef]
- Venkatesh, B.; Lee, A.P.; Ravi, V.; Maurya, A.K.; Lian, M.M.; Swann, J.B.; Ohta, Y.; Flajnik, M.F.; Sutoh, Y.; Kasahara, M.; et al. Elephant shark genome provides unique insights into gnathostome evolution. Nature 2014, 505, 174–179. [Google Scholar] [CrossRef]
- Van de Peppel, J.; van Leeuwen, J.P. Vitamin D and gene networks in human osteoblasts. Front. Physiol. 2014, 5, 137. [Google Scholar] [CrossRef]
- Kim, S.; Yamazaki, M.; Zella, L.A.; Meyer, M.B.; Fretz, J.A.; Shevde, N.K.; Pike, J.W. Multiple enhancer regions located at significant distances upstream of the transcriptional start site mediate RANKL gene expression in response to 1,25-dihydroxyvitamin D3. J. Steroid Biochem. Mol. Biol. 2007, 103, 430–434. [Google Scholar] [CrossRef]
- Karsenty, G.; Khosla, S. The crosstalk between bone remodeling and energy metabolism: A translational perspective. Cell Metab. 2022, 34, 805–817. [Google Scholar] [CrossRef]
- Tsukasaki, M.; Takayanagi, H. Osteoimmunology: Evolving concepts in bone-immune interactions in health and disease. Nat. Rev. Immunol. 2019, 19, 626–642. [Google Scholar] [CrossRef]
- Cortes, M.; Chen, M.J.; Stachura, D.L.; Liu, S.Y.; Kwan, W.; Wright, F.; Vo, L.T.; Theodore, L.N.; Esain, V.; Frost, I.M.; et al. Developmental vitamin D availability impacts hematopoietic stem cell production. Cell Rep. 2016, 17, 458–468. [Google Scholar] [CrossRef] [PubMed]
- Hublin, J.J.; Ben-Ncer, A.; Bailey, S.E.; Freidline, S.E.; Neubauer, S.; Skinner, M.M.; Bergmann, I.; Le Cabec, A.; Benazzi, S.; Harvati, K.; et al. New fossils from Jebel Irhoud, Morocco and the pan-African origin of Homo sapiens. Nature 2017, 546, 289–292. [Google Scholar] [CrossRef] [PubMed]
- Greaves, M. Was skin cancer a selective force for black pigmentation in early hominin evolution? Proc. Biol. Sci. 2014, 281, 20132955. [Google Scholar] [CrossRef]
- Fajuyigbe, D.; Lwin, S.M.; Diffey, B.L.; Baker, R.; Tobin, D.J.; Sarkany, R.P.E.; Young, A.R. Melanin distribution in human epidermis affords localized protection against DNA photodamage and concurs with skin cancer incidence difference in extreme phototypes. FASEB J. 2018, 32, 3700–3706. [Google Scholar] [CrossRef] [PubMed]
- Luxwolda, M.F.; Kuipers, R.S.; Kema, I.P.; Dijck-Brouwer, D.A.; Muskiet, F.A. Traditionally living populations in East Africa have a mean serum 25-hydroxyvitamin D concentration of 115 nmol/l. Br. J. Nutr. 2012, 108, 1557–1561. [Google Scholar] [CrossRef] [PubMed]
- Pagani, L.; Lawson, D.J.; Jagoda, E.; Morseburg, A.; Eriksson, A.; Mitt, M.; Clemente, F.; Hudjashov, G.; DeGiorgio, M.; Saag, L.; et al. Genomic analyses inform on migration events during the peopling of Eurasia. Nature 2016, 538, 238–242. [Google Scholar] [CrossRef] [PubMed]
- Skoglund, P.; Mathieson, I. Ancient genomics of modern humans: The first decade. Annu. Rev. Genomics Hum. Genet. 2018, 19, 381–404. [Google Scholar] [CrossRef]
- O’Neill, C.M.; Kazantzidis, A.; Ryan, M.J.; Barber, N.; Sempos, C.T.; Durazo-Arvizu, R.A.; Jorde, R.; Grimnes, G.; Eiriksdottir, G.; Gudnason, V.; et al. Seasonal changes in vitamin D-effective UVB availability in Europe and associations with population serum 25-hydroxyvitamin D. Nutrients 2016, 8, 533. [Google Scholar] [CrossRef]
- Jablonski, N.G.; Chaplin, G. The evolution of human skin coloration. J. Hum. Evol. 2000, 39, 57–106. [Google Scholar] [CrossRef]
- Deng, L.; Xu, S. Adaptation of human skin color in various populations. Hereditas 2018, 155, 1–12. [Google Scholar] [CrossRef]
- Cichorek, M.; Wachulska, M.; Stasiewicz, A.; Tyminska, A. Skin melanocytes: Biology and development. Postepy Dermatol. Alergol. 2013, 30, 30–41. [Google Scholar] [CrossRef] [PubMed]
- Pavan, W.J.; Sturm, R.A. The genetics of human skin and hair pigmentation. Annu. Rev. Genomics Hum. Genet. 2019, 20, 41–72. [Google Scholar] [CrossRef] [PubMed]
- Lindgren, J.; Moyer, A.; Schweitzer, M.H.; Sjovall, P.; Uvdal, P.; Nilsson, D.E.; Heimdal, J.; Engdahl, A.; Gren, J.A.; Schultz, B.P.; et al. Interpreting melanin-based coloration through deep time: A critical review. Proc. Biol. Sci. 2015, 282, 20150614. [Google Scholar] [CrossRef] [PubMed]
- Ju, D.; Mathieson, I. The evolution of skin pigmentation associated variation in West Eurasia. Proc Nat Acad Sci USA 2021, 118, e2009227118. [Google Scholar] [CrossRef]
- Gunther, T.; Malmstrom, H.; Svensson, E.M.; Omrak, A.; Sanchez-Quinto, F.; Kilinc, G.M.; Krzewinska, M.; Eriksson, G.; Fraser, M.; Edlund, H.; et al. Population genomics of Mesolithic Scandinavia: Investigating early postglacial migration routes and high-latitude adaptation. PLoS Biol. 2018, 16, e2003703. [Google Scholar] [CrossRef]
- Nakagome, S.; Alkorta-Aranburu, G.; Amato, R.; Howie, B.; Peter, B.M.; Hudson, R.R.; Di Rienzo, A. Estimating the ages of selection signals from different epochs in human history. Mol. Biol. Evol. 2016, 33, 657–669. [Google Scholar] [CrossRef]
- Fu, Q.; Posth, C.; Hajdinjak, M.; Petr, M.; Mallick, S.; Fernandes, D.; Furtwangler, A.; Haak, W.; Meyer, M.; Mittnik, A.; et al. The genetic history of Ice Age Europe. Nature 2016, 534, 200–205. [Google Scholar] [CrossRef]
- Prufer, K.; Racimo, F.; Patterson, N.; Jay, F.; Sankararaman, S.; Sawyer, S.; Heinze, A.; Renaud, G.; Sudmant, P.H.; de Filippo, C.; et al. The complete genome sequence of a Neanderthal from the Altai Mountains. Nature 2014, 505, 43–49. [Google Scholar] [CrossRef]
- Peyregne, S.; Slon, V.; Mafessoni, F.; de Filippo, C.; Hajdinjak, M.; Nagel, S.; Nickel, B.; Essel, E.; Le Cabec, A.; Wehrberger, K.; et al. Nuclear DNA from two early Neandertals reveals 80,000 years of genetic continuity in Europe. Sci. Adv. 2019, 5, eaaw5873. [Google Scholar] [CrossRef]
- Petr, M.; Paabo, S.; Kelso, J.; Vernot, B. Limits of long-term selection against Neandertal introgression. Proc. Natl. Acad. Sci. USA 2019, 116, 1639–1644. [Google Scholar] [CrossRef]
- Villalba-Mouco, V.; van de Loosdrecht, M.S.; Posth, C.; Mora, R.; Martinez-Moreno, J.; Rojo-Guerra, M.; Salazar-Garcia, D.C.; Royo-Guillen, J.I.; Kunst, M.; Rougier, H.; et al. Survival of late Pleistocene hunter-gatherer ancestry in the Iberian peninsula. Curr. Biol. 2019, 29, 1169–1177.e7. [Google Scholar] [CrossRef] [PubMed]
- Mathieson, I.; Lazaridis, I.; Rohland, N.; Mallick, S.; Patterson, N.; Roodenberg, S.A.; Harney, E.; Stewardson, K.; Fernandes, D.; Novak, M.; et al. Genome-wide patterns of selection in 230 ancient Eurasians. Nature 2015, 528, 499–503. [Google Scholar] [CrossRef] [PubMed]
- Mathieson, I.; Alpaslan-Roodenberg, S.; Posth, C.; Szecsenyi-Nagy, A.; Rohland, N.; Mallick, S.; Olalde, I.; Broomandkhoshbacht, N.; Candilio, F.; Cheronet, O.; et al. The genomic history of southeastern Europe. Nature 2018, 555, 197–203. [Google Scholar] [CrossRef] [PubMed]
- Haak, W.; Lazaridis, I.; Patterson, N.; Rohland, N.; Mallick, S.; Llamas, B.; Brandt, G.; Nordenfelt, S.; Harney, E.; Stewardson, K.; et al. Massive migration from the steppe was a source for Indo-European languages in Europe. Nature 2015, 522, 207–211. [Google Scholar] [CrossRef] [PubMed]
- Damgaard, P.B.; Marchi, N.; Rasmussen, S.; Peyrot, M.; Renaud, G.; Korneliussen, T.; Moreno-Mayar, J.V.; Pedersen, M.W.; Goldberg, A.; Usmanova, E.; et al. 137 ancient human genomes from across the Eurasian steppes. Nature 2018, 557, 369–374. [Google Scholar] [CrossRef] [PubMed]
- Mays, S.; Prowse, T.; George, M.; Brickley, M. Latitude, urbanization, age, and sex as risk factors for vitamin D deficiency disease in the Roman Empire. Am. J. Phys. Anthropol. 2018, 167, 484–496. [Google Scholar] [CrossRef]
- Kuan, V.; Martineau, A.R.; Griffiths, C.J.; Hypponen, E.; Walton, R. DHCR7 mutations linked to higher vitamin D status allowed early human migration to northern latitudes. BMC Evol. Biol. 2013, 13, 144. [Google Scholar] [CrossRef]
- Wang, T.J.; Zhang, F.; Richards, J.B.; Kestenbaum, B.; van Meurs, J.B.; Berry, D.; Kiel, D.P.; Streeten, E.A.; Ohlsson, C.; Koller, D.L.; et al. Common genetic determinants of vitamin D insufficiency: A genome-wide association study. Lancet 2010, 376, 180–188. [Google Scholar] [CrossRef]
- Jiang, X.; O’Reilly, P.F.; Aschard, H.; Hsu, Y.H.; Richards, J.B.; Dupuis, J.; Ingelsson, E.; Karasik, D.; Pilz, S.; Berry, D.; et al. Genome-wide association study in 79,366 European-ancestry individuals informs the genetic architecture of 25-hydroxyvitamin D levels. Nat. Commun. 2018, 9, 260. [Google Scholar] [CrossRef]
- Revez, J.A.; Lin, T.; Qiao, Z.; Xue, A.; Holtz, Y.; Zhu, Z.; Zeng, J.; Wang, H.; Sidorenko, J.; Kemper, K.E.; et al. Genome-wide association study identifies 143 loci associated with 25 hydroxyvitamin D concentration. Nat. Commun. 2020, 11, 1647. [Google Scholar] [CrossRef]
- Hardy, A. Commentary: Bread and alum, syphilis and sunlight: Rickets in the nineteenth century. Int. J. Epidemiol. 2003, 32, 337–340. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Shen, F.; Petryk, A.; Tang, J.; Chen, X.; Sergi, C. “English Disease”: Historical notes on rickets, the bone–lung link and child neglect issues. Nutrients 2016, 8, 722. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.J.; Wang, X.H.; Liu, Z.D.; Cao, W.L.; Han, Y.; Ma, A.G.; Xu, S.F. Vitamin D deficiency and the risk of tuberculosis: A meta-analysis. Drug Des. Devel. Ther. 2017, 11, 91–102. [Google Scholar] [CrossRef] [PubMed]
- Brace, S.; Diekmann, Y.; Booth, T.J.; van Dorp, L.; Faltyskova, Z.; Rohland, N.; Mallick, S.; Olalde, I.; Ferry, M.; Michel, M.; et al. Ancient genomes indicate population replacement in Early Neolithic Britain. Nat. Ecol. Evol. 2019, 3, 765–771. [Google Scholar] [CrossRef]
- Holick, M.F. Vitamin D deficiency. N. Engl. J. Med. 2007, 357, 266–281. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Carlberg, C. Vitamin D in the Context of Evolution. Nutrients 2022, 14, 3018. https://doi.org/10.3390/nu14153018
Carlberg C. Vitamin D in the Context of Evolution. Nutrients. 2022; 14(15):3018. https://doi.org/10.3390/nu14153018
Chicago/Turabian StyleCarlberg, Carsten. 2022. "Vitamin D in the Context of Evolution" Nutrients 14, no. 15: 3018. https://doi.org/10.3390/nu14153018
APA StyleCarlberg, C. (2022). Vitamin D in the Context of Evolution. Nutrients, 14(15), 3018. https://doi.org/10.3390/nu14153018





