Association between Polymorphisms in Vitamin D Pathway-Related Genes, Vitamin D Status, Muscle Mass and Function: A Systematic Review
Abstract
1. Introduction
2. Materials and Methods
2.1. Search Strategy
- (i)
- each of the 14 vitamin D-pathway-related genes (including their aliases variation (MeSH terms])
- (ii)
- vitamin D status (vitamin D[MeSH] OR 25(OH)D OR 25(OH)D2 OR 25(OH) D3 OR Vitamin D Deficiency[MeSH] OR “vitamin D status” OR “vitamin D level” OR “vitamin D inadequacy” OR “hypovitaminosis D” OR “avitaminosis D”) NOT Review[Publication Type], Filters: Humans; English; German
- (iii)
- muscle strength and function (Muscle, Skeletal[MeSH] OR Muscle Strength[MeSH] OR Physical Fitness[MeSH] OR Walking Speed[MeSH] OR “muscle mass” OR “lean body mass” OR “muscle quality” OR “physical performance” OR SPPB OR “short physical performance battery” OR “handgrip strength” OR “chair stand” OR “arm curl”) NOT Review[Publication Type], Filters: Humans; English; German
2.2. Inclusion and Exclusion Criteria
- (i)
- were conducted among humans aged >18 years from both sexes;
- (ii)
- comprised candidate or genome-wide association studies being cross-sectional, cohort, case control, or intervention studies;
- (iii)
- investigated healthy subjects or contained at least a separate control group with normal health status;
- (iv)
- measured at least one genotype in a vitamin D pathway-related gene;
- (v)
- reported vitamin D status (circulating plasma/serum levels, 25(OH)D, 25(OH)D2, or 25(OH)D3 metabolites) and/or reported results for muscle mass or function, muscle strength parameters or scores for the Short Physical Performance Battery (SPPB);
- (vi)
- were published in English or German.
- (i)
- reported only vitamin D intake or vitamin D metabolites from urine;
- (ii)
- investigated children (<18 years old);
- (iii)
- investigated participants with severe chronic or acute illnesses with a known impact on either vitamin D status and/or muscle mass and strength (myopathies, hypocalciuric hypercalcemia), or pregnant or lactating women;
- (iv)
- were published as case reports, systematic reviews, or meta-analyses.
2.3. Study Selection and Data Extraction
2.4. Risk of Bias (Quality) Assessment
3. Results
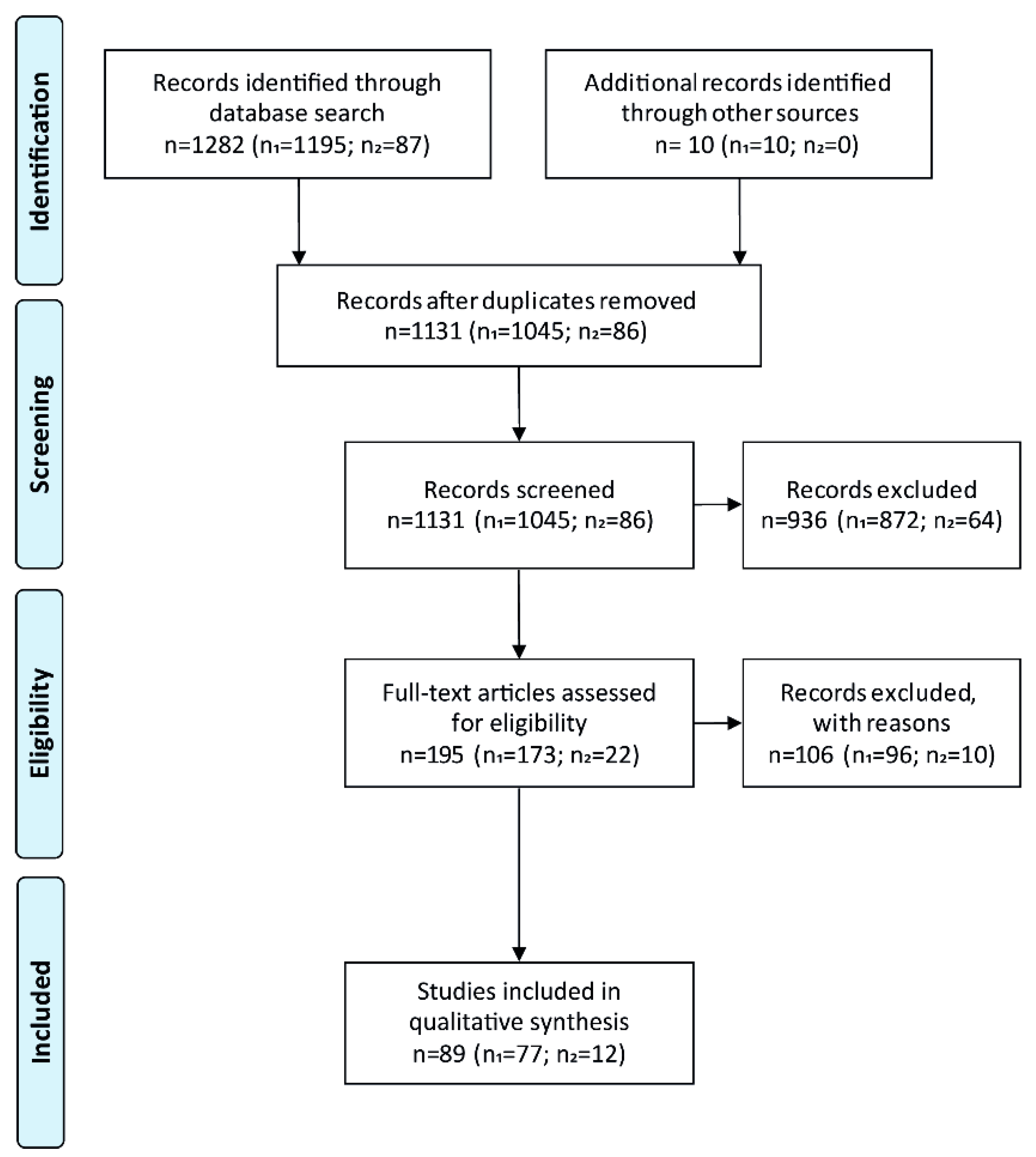
3.1. Study Selection and Characteristics
3.2. Polymorphisms in Vitamin D Pathway-Related Genes and Vitamin D Status
3.3. Polymorphisms in Vitamin D Pathway-Related Genes and Muscle Mass and Function
3.4. Quality of Included Studies
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lips, P.; van Schoor, N.M. The effect of vitamin D on bone and osteoporosis. Best Pract. Res. Clin. Endocrinol. Metab. 2011, 25, 585–591. [Google Scholar] [CrossRef]
- O’Neill, C.M.; Kazantzidis, A.; Ryan, M.J.; Barber, N.; Sempos, C.T.; Durazo-Arvizu, R.A.; Jorde, R.; Grimnes, G.; Eiriksdottir, G.; Gudnason, V.; et al. Seasonal Changes in Vitamin D-Effective UVB Availability in Europe and Associations with Population Serum 25-Hydroxyvitamin D. Nutrients 2016, 8, 533. [Google Scholar] [CrossRef] [PubMed]
- Wacker, M.; Holick, M.F. Sunlight and Vitamin D: A global perspective for health. Dermatoendocrinology 2013, 5, 51–108. [Google Scholar] [CrossRef]
- EFSA Panel on Dietetic Products, Nutrition and Allergies (NDA). Dietary reference values for vitamin D. EFSA J. 2016, 15, e04780. [Google Scholar] [CrossRef]
- Garcia, M.; Seelaender, M.; Sotiropoulos, A.; Coletti, D.; Lancha, A.H., Jr. Vitamin D, muscle recovery, sarcopenia, cachexia, and muscle atrophy. Nutrition 2019, 60, 66–69. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhu, J.; DeLuca, H.F. Where is the vitamin D receptor? Arch. Biochem. Biophys. 2012, 523, 123–133. [Google Scholar] [CrossRef]
- Holick, M.F. Vitamin D status: Measurement, interpretation, and clinical application. Ann. Epidemiol. 2009, 19, 73–78. [Google Scholar] [CrossRef]
- Cashman, K.D.; Dowling, K.G.; Skrabakova, Z.; Gonzalez-Gross, M.; Valtuena, J.; De Henauw, S.; Moreno, L.; Damsgaard, C.T.; Michaelsen, K.F.; Molgaard, C.; et al. Vitamin D deficiency in Europe: Pandemic? Am. J. Clin. Nutr. 2016, 103, 1033–1044. [Google Scholar] [CrossRef]
- Liu, X.; Baylin, A.; Levy, P.D. Vitamin D deficiency and insufficiency among US adults: Prevalence, predictors and clinical implications. Br. J. Nutr. 2018, 119, 928–936. [Google Scholar] [CrossRef]
- van Schoor, N.; Lips, P. Global Overview of Vitamin D Status. Endocrinol. Metab. Clin. N. Am. 2017, 46, 845–870. [Google Scholar] [CrossRef]
- Wicherts, I.S.; van Schoor, N.M.; Boeke, A.J.; Visser, M.; Deeg, D.J.; Smit, J.; Knol, D.L.; Lips, P. Vitamin D status predicts physical performance and its decline in older persons. J. Clin. Endocrinol. Metab. 2007, 92, 2058–2065. [Google Scholar] [CrossRef]
- Remelli, F.; Vitali, A.; Zurlo, A.; Volpato, S. Vitamin D Deficiency and Sarcopenia in Older Persons. Nutrients 2019, 11, 2861. [Google Scholar] [CrossRef] [PubMed]
- Cruz-Jentoft, A.J.; Bahat, G.; Bauer, J.; Boirie, Y.; Bruyere, O.; Cederholm, T.; Cooper, C.; Landi, F.; Rolland, Y.; Sayer, A.A.; et al. Sarcopenia: Revised European consensus on definition and diagnosis. Age Ageing 2019, 48, 16–31. [Google Scholar] [CrossRef] [PubMed]
- Rosendahl-Riise, H.; Spielau, U.; Ranhoff, A.H.; Gudbrandsen, O.A.; Dierkes, J. Vitamin D supplementation and its influence on muscle strength and mobility in community-dwelling older persons: A systematic review and meta-analysis. J. Hum. Nutr. Diet. 2017, 30, 3–15. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Quan, M.; Cao, Z.B. Effect of vitamin D supplementation on upper and lower limb muscle strength and muscle power in athletes: A meta-analysis. PLoS ONE 2019, 14, e0215826. [Google Scholar] [CrossRef] [PubMed]
- Antoniak, A.E.; Greig, C.A. The effect of combined resistance exercise training and vitamin D3 supplementation on musculoskeletal health and function in older adults: A systematic review and meta-analysis. BMJ Open 2017, 7, e014619. [Google Scholar] [CrossRef] [PubMed]
- Dawson-Hughes, B. Vitamin D and muscle function. J. Steroid Biochem. Mol. Biol. 2017, 173, 313–316. [Google Scholar] [CrossRef]
- Ceglia, L. Vitamin D and its role in skeletal muscle. Curr. Opin. Clin. Nutr. Metab. Care 2009, 12, 628–633. [Google Scholar] [CrossRef]
- Arem, H.; Yu, K.; Xiong, X.; Moy, K.; Freedman, N.D.; Mayne, S.T.; Albanes, D.; Arslan, A.A.; Austin, M.; Bamlet, W.R.; et al. Vitamin D metabolic pathway genes and pancreatic cancer risk. PLoS ONE 2015, 10, e0117574. [Google Scholar] [CrossRef]
- Jolliffe, D.A.; Walton, R.T.; Griffiths, C.J.; Martineau, A.R. Single nucleotide polymorphisms in the vitamin D pathway associating with circulating concentrations of vitamin D metabolites and non-skeletal health outcomes: Review of genetic association studies. J. Steroid Biochem. Mol. Biol. 2016, 164, 18–29. [Google Scholar] [CrossRef]
- Puthucheary, Z.; Skipworth, J.R.; Rawal, J.; Loosemore, M.; Van Someren, K.; Montgomery, H.E. Genetic influences in sport and physical performance. Sports Med. 2011, 41, 845–859. [Google Scholar] [CrossRef] [PubMed]
- Wessner, B.; Krasniqi, E.; Boshnjaku, A.; Wagner, K.-H. Association between Polymorphisms in Vitamin D Pathway-Related Genes, Vitamin D Status, Muscle Mass and Function: A Systematic Review; PROSPERO: York, UK, 2018; CRD42018117989. [Google Scholar]
- Moher, D.; Liberati, A.; Tetzlaff, J.; Altman, D.G.; Group, P. Preferred reporting items for systematic reviews and meta-analyses: The PRISMA statement. Int. J. Surg. 2010, 8, 336–341. [Google Scholar] [CrossRef]
- Belinky, F.; Nativ, N.; Stelzer, G.; Zimmerman, S.; Iny Stein, T.; Safran, M.; Lancet, D. PathCards: Multi-source consolidation of human biological pathways. Database 2015, 2015. [Google Scholar] [CrossRef]
- Little, J.; Higgins, J.P.; Ioannidis, J.P.; Moher, D.; Gagnon, F.; von Elm, E.; Khoury, M.J.; Cohen, B.; Davey-Smith, G.; Grimshaw, J.; et al. STrengthening the REporting of Genetic Association Studies (STREGA): An extension of the STROBE statement. PLoS Med. 2009, 6, e22. [Google Scholar] [CrossRef]
- Ahn, J.; Albanes, D.; Berndt, S.I.; Peters, U.; Chatterjee, N.; Freedman, N.D.; Abnet, C.C.; Huang, W.Y.; Kibel, A.S.; Crawford, E.D.; et al. Vitamin D-related genes, serum vitamin D concentrations and prostate cancer risk. Carcinogenesis 2009, 30, 769–776. [Google Scholar] [CrossRef]
- Barry, E.L.; Rees, J.R.; Peacock, J.L.; Mott, L.A.; Amos, C.I.; Bostick, R.M.; Figueiredo, J.C.; Ahnen, D.J.; Bresalier, R.S.; Burke, C.A.; et al. Genetic variants in CYP2R1, CYP24A1, and VDR modify the efficacy of vitamin D3 supplementation for increasing serum 25-hydroxyvitamin D levels in a randomized controlled trial. J. Clin. Endocrinol. Metab. 2014, 99, E2133–E2137. [Google Scholar] [CrossRef] [PubMed]
- Batai, K.; Murphy, A.B.; Shah, E.; Ruden, M.; Newsome, J.; Agate, S.; Dixon, M.A.; Chen, H.Y.; Deane, L.A.; Hollowell, C.M.; et al. Common vitamin D pathway gene variants reveal contrasting effects on serum vitamin D levels in African Americans and European Americans. Hum. Genet. 2014, 133, 1395–1405. [Google Scholar] [CrossRef]
- Brouwer-Brolsma, E.M.; Vaes, A.M.M.; van der Zwaluw, N.L.; van Wijngaarden, J.P.; Swart, K.M.A.; Ham, A.C.; van Dijk, S.C.; Enneman, A.W.; Sohl, E.; van Schoor, N.M.; et al. Relative importance of summer sun exposure, vitamin D intake, and genes to vitamin D status in Dutch older adults: The B-PROOF study. J. Steroid Biochem. Mol. Biol. 2016, 164, 168–176. [Google Scholar] [CrossRef]
- Bu, F.X.; Armas, L.; Lappe, J.; Zhou, Y.; Gao, G.; Wang, H.W.; Recker, R.; Zhao, L.J. Comprehensive association analysis of nine candidate genes with serum 25-hydroxy vitamin D levels among healthy Caucasian subjects. Hum. Genet. 2010, 128, 549–556. [Google Scholar] [CrossRef]
- Cheung, C.L.; Lau, K.S.; Sham, P.C.; Tan, K.C.; Kung, A.W. Genetic variant in vitamin D binding protein is associated with serum 25-hydroxyvitamin D and vitamin D insufficiency in southern Chinese. J. Hum. Genet. 2013, 58, 749–751. [Google Scholar] [CrossRef]
- Clendenen, T.V.; Ge, W.; Koenig, K.L.; Axelsson, T.; Liu, M.; Afanasyeva, Y.; Andersson, A.; Arslan, A.A.; Chen, Y.; Hallmans, G.; et al. Genetic Polymorphisms in Vitamin D Metabolism and Signaling Genes and Risk of Breast Cancer: A Nested Case-Control Study. PLoS ONE 2015, 10, e0140478. [Google Scholar] [CrossRef]
- Cooper, J.D.; Smyth, D.J.; Walker, N.M.; Stevens, H.; Burren, O.S.; Wallace, C.; Greissl, C.; Ramos-Lopez, E.; Hypponen, E.; Dunger, D.B.; et al. Inherited variation in vitamin D genes is associated with predisposition to autoimmune disease type 1 diabetes. Diabetes 2011, 60, 1624–1631. [Google Scholar] [CrossRef] [PubMed]
- Davies, J.R.; Chang, Y.M.; Snowden, H.; Chan, M.; Leake, S.; Karpavicius, B.; Haynes, S.; Kukalizch, K.; Randerson-Moor, J.; Elliott, F.; et al. The determinants of serum vitamin D levels in participants in a melanoma case-control study living in a temperate climate. Cancer Causes Control. 2011, 22, 1471–1482. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Didriksen, A.; Grimnes, G.; Hutchinson, M.S.; Kjaergaard, M.; Svartberg, J.; Joakimsen, R.M.; Jorde, R. The serum 25-hydroxyvitamin D response to vitamin D supplementation is related to genetic factors, BMI, and baseline levels. Eur. J. Endocrinol. 2013, 169, 559–567. [Google Scholar] [CrossRef] [PubMed]
- Elkum, N.; Alkayal, F.; Noronha, F.; Ali, M.M.; Melhem, M.; Al-Arouj, M.; Bennakhi, A.; Behbehani, K.; Alsmadi, O.; Abubaker, J. Vitamin D insufficiency in Arabs and South Asians positively associates with polymorphisms in GC and CYP2R1 genes. PLoS ONE 2014, 9, e113102. [Google Scholar] [CrossRef]
- Engelman, C.D.; Fingerlin, T.E.; Langefeld, C.D.; Hicks, P.J.; Rich, S.S.; Wagenknecht, L.E.; Bowden, D.W.; Norris, J.M. Genetic and environmental determinants of 25-hydroxyvitamin D and 1,25-dihydroxyvitamin D levels in Hispanic and African Americans. J. Clin. Endocrinol. Metab. 2008, 93, 3381–3388. [Google Scholar] [CrossRef] [PubMed]
- Engelman, C.D.; Meyers, K.J.; Iyengar, S.K.; Liu, Z.; Karki, C.K.; Igo, R.P., Jr.; Truitt, B.; Robinson, J.; Sarto, G.E.; Wallace, R.; et al. Vitamin D intake and season modify the effects of the GC and CYP2R1 genes on 25-hydroxyvitamin D concentrations. J. Nutr. 2013, 143, 17–26. [Google Scholar] [CrossRef]
- Fohner, A.E.; Wang, Z.; Yracheta, J.; O’Brien, D.M.; Hopkins, S.E.; Black, J.; Philip, J.; Wiener, H.W.; Tiwari, H.K.; Stapleton, P.L.; et al. Genetics, Diet, and Season Are Associated with Serum 25-Hydroxycholecalciferol Concentration in a Yup’ik Study Population from Southwestern Alaska. J. Nutr. 2016, 146, 318–325. [Google Scholar] [CrossRef]
- Fu, L.; Yun, F.; Oczak, M.; Wong, B.Y.; Vieth, R.; Cole, D.E. Common genetic variants of the vitamin D binding protein (DBP) predict differences in response of serum 25-hydroxyvitamin D [25(OH)D] to vitamin D supplementation. Clin. Biochem. 2009, 42, 1174–1177. [Google Scholar] [CrossRef]
- Gaffney-Stomberg, E.; Lutz, L.J.; Shcherbina, A.; Ricke, D.O.; Petrovick, M.; Cropper, T.L.; Cable, S.J.; McClung, J.P. Association Between Single Gene Polymorphisms and Bone Biomarkers and Response to Calcium and Vitamin D Supplementation in Young Adults Undergoing Military Training. J. Bone Miner. Res. 2017, 32, 498–507. [Google Scholar] [CrossRef]
- Gilbert, R.; Bonilla, C.; Metcalfe, C.; Lewis, S.; Evans, D.M.; Fraser, W.D.; Kemp, J.P.; Donovan, J.L.; Hamdy, F.C.; Neal, D.E.; et al. Associations of vitamin D pathway genes with circulating 25-hydroxyvitamin-D, 1,25-dihydroxyvitamin-D, and prostate cancer: A nested case-control study. Cancer Causes Control. 2015, 26, 205–218. [Google Scholar] [CrossRef] [PubMed]
- Gozdzik, A.; Zhu, J.; Wong, B.Y.; Fu, L.; Cole, D.E.; Parra, E.J. Association of vitamin D binding protein (VDBP) polymorphisms and serum 25(OH)D concentrations in a sample of young Canadian adults of different ancestry. J. Steroid Biochem. Mol. Biol. 2011, 127, 405–412. [Google Scholar] [CrossRef]
- Hansen, J.G.; Gao, W.; Dupuis, J.; O’Connor, G.T.; Tang, W.; Kowgier, M.; Sood, A.; Gharib, S.A.; Palmer, L.J.; Fornage, M.; et al. Association of 25-Hydroxyvitamin D status and genetic variation in the vitamin D metabolic pathway with FEV1 in the Framingham Heart Study. Respir. Res. 2015, 16, 81. [Google Scholar] [CrossRef] [PubMed]
- Janssens, W.; Bouillon, R.; Claes, B.; Carremans, C.; Lehouck, A.; Buysschaert, I.; Coolen, J.; Mathieu, C.; Decramer, M.; Lambrechts, D. Vitamin D deficiency is highly prevalent in COPD and correlates with variants in the vitamin D-binding gene. Thorax 2010, 65, 215–220. [Google Scholar] [CrossRef]
- Jolliffe, D.A.; Hanifa, Y.; Witt, K.D.; Venton, T.R.; Rowe, M.; Timms, P.M.; Hypponen, E.; Walton, R.T.; Griffiths, C.J.; Martineau, A.R. Environmental and genetic determinants of vitamin D status among older adults in London, UK. J. Steroid Biochem. Mol. Biol. 2016, 164, 30–35. [Google Scholar] [CrossRef]
- Junaid, K.; Rehman, A.; Jolliffe, D.A.; Wood, K.; Martineau, A.R. High prevalence of vitamin D deficiency among women of child-bearing age in Lahore Pakistan, associating with lack of sun exposure and illiteracy. BMC Womens Health 2015, 15, 83. [Google Scholar] [CrossRef]
- Lafi, Z.M.; Irshaid, Y.M.; El-Khateeb, M.; Ajlouni, K.M.; Hyassat, D. Association of rs7041 and rs4588 Polymorphisms of the Vitamin D Binding Protein and the rs10741657 Polymorphism of CYP2R1 with Vitamin D Status Among Jordanian Patients. Genet. Test. Mol. Biomark. 2015, 19, 629–636. [Google Scholar] [CrossRef]
- Larcombe, L.; Mookherjee, N.; Slater, J.; Slivinski, C.; Singer, M.; Whaley, C.; Denechezhe, L.; Matyas, S.; Turner-Brannen, E.; Nickerson, P.; et al. Vitamin D in a northern Canadian first nation population: Dietary intake, serum concentrations and functional gene polymorphisms. PLoS ONE 2012, 7, e49872. [Google Scholar] [CrossRef]
- Li, L.H.; Yin, X.Y.; Wu, X.H.; Zhang, L.; Pan, S.Y.; Zheng, Z.J.; Wang, J.G. Serum 25(OH)D and vitamin D status in relation to VDR, GC and CYP2R1 variants in Chinese. Endocr. J. 2014, 61, 133–141. [Google Scholar] [CrossRef]
- Lu, L.; Sheng, H.; Li, H.; Gan, W.; Liu, C.; Zhu, J.; Loos, R.J.; Lin, X. Associations between common variants in GC and DHCR7/NADSYN1 and vitamin D concentration in Chinese Hans. Hum. Genet. 2012, 131, 505–512. [Google Scholar] [CrossRef]
- McKay, G.J.; Young, I.S.; McGinty, A.; Bentham, G.C.; Chakravarthy, U.; Rahu, M.; Seland, J.; Soubrane, G.; Tomazzoli, L.; Topouzis, F.; et al. Associations between Serum Vitamin D and Genetic Variants in Vitamin D Pathways and Age-Related Macular Degeneration in the European Eye Study. Ophthalmology 2017, 124, 90–96. [Google Scholar] [CrossRef]
- Medlej-Hashim, M.; Jounblat, R.; Hamade, A.; Ibrahim, J.N.; Rizk, F.; Azzi, G.; Abdallah, M.; Nakib, L.; Lahoud, M.; Nabout, R. Hypovitaminosis D in a Young Lebanese Population: Effect of GC Gene Polymorphisms on Vitamin D and Vitamin D Binding Protein Levels. Ann. Hum. Genet. 2015, 79, 394–401. [Google Scholar] [CrossRef]
- Nimitphong, H.; Saetung, S.; Chanprasertyotin, S.; Chailurkit, L.O.; Ongphiphadhanakul, B. Changes in circulating 25-hydroxyvitamin D according to vitamin D binding protein genotypes after vitamin D(3) or D(2)supplementation. Nutr. J. 2013, 12, 39. [Google Scholar] [CrossRef]
- Nissen, J.; Rasmussen, L.B.; Ravn-Haren, G.; Andersen, E.W.; Hansen, B.; Andersen, R.; Mejborn, H.; Madsen, K.H.; Vogel, U. Common variants in CYP2R1 and GC genes predict vitamin D concentrations in healthy Danish children and adults. PLoS ONE 2014, 9, e89907. [Google Scholar] [CrossRef]
- Nissen, J.; Vogel, U.; Ravn-Haren, G.; Andersen, E.W.; Madsen, K.H.; Nexo, B.A.; Andersen, R.; Mejborn, H.; Bjerrum, P.J.; Rasmussen, L.B.; et al. Common variants in CYP2R1 and GC genes are both determinants of serum 25-hydroxyvitamin D concentrations after UVB irradiation and after consumption of vitamin D(3)-fortified bread and milk during winter in Denmark. Am. J. Clin. Nutr. 2015, 101, 218–227. [Google Scholar] [CrossRef]
- Ordonez-Mena, J.M.; Maalmi, H.; Schottker, B.; Saum, K.U.; Holleczek, B.; Wang, T.J.; Burwinkel, B.; Brenner, H. Genetic Variants in the Vitamin D Pathway, 25(OH)D Levels, and Mortality in a Large Population-Based Cohort Study. J. Clin. Endocrinol. Metab. 2017, 102, 470–477. [Google Scholar] [CrossRef]
- Perna, L.; Felix, J.F.; Breitling, L.P.; Haug, U.; Raum, E.; Burwinkel, B.; Schottker, B.; Brenner, H. Genetic variations in the vitamin D binding protein and season-specific levels of vitamin D among older adults. Epidemiology 2013, 24, 104–109. [Google Scholar] [CrossRef]
- Powe, C.E.; Evans, M.K.; Wenger, J.; Zonderman, A.B.; Berg, A.H.; Nalls, M.; Tamez, H.; Zhang, D.; Bhan, I.; Karumanchi, S.A.; et al. Vitamin D-binding protein and vitamin D status of black Americans and white Americans. N. Engl. J. Med. 2013, 369, 1991–2000. [Google Scholar] [CrossRef]
- Rees, J.R.; Mott, L.A.; Barry, E.L.; Baron, J.A.; Bostick, R.M.; Figueiredo, J.C.; Bresalier, R.S.; Robertson, D.J.; Peacock, J.L. Lifestyle and Other Factors Explain One-Half of the Variability in the Serum 25-Hydroxyvitamin D Response to Cholecalciferol Supplementation in Healthy Adults. J. Nutr. 2016, 146, 2312–2324. [Google Scholar] [CrossRef]
- Reimers, L.L.; Crew, K.D.; Bradshaw, P.T.; Santella, R.M.; Steck, S.E.; Sirosh, I.; Terry, M.B.; Hershman, D.L.; Shane, E.; Cremers, S.; et al. Vitamin D-related gene polymorphisms, plasma 25-hydroxyvitamin D, and breast cancer risk. Cancer Causes Control. 2015, 26, 187–203. [Google Scholar] [CrossRef]
- Robien, K.; Butler, L.M.; Wang, R.; Beckman, K.B.; Walek, D.; Koh, W.P.; Yuan, J.M. Genetic and environmental predictors of serum 25-hydroxyvitamin D concentrations among middle-aged and elderly Chinese in Singapore. Br. J. Nutr. 2013, 109, 493–502. [Google Scholar] [CrossRef]
- Sadat-Ali, M.; Al-Turki, H.A.; Azam, M.Q.; Al-Elq, A.H. Genetic influence on circulating vitamin D among Saudi Arabians. Saudi Med. J. 2016, 37, 996–1001. [Google Scholar] [CrossRef]
- Sedky, N.K.; Hassanein, S.I.; Gad, M.Z. Independent assortment of GC gene polymorphism (rs2282679) and 25-hydroxyvitamin D levels in coronary artery disease. Can. J. Physiol. Pharm. 2018, 96, 345–351. [Google Scholar] [CrossRef]
- Signorello, L.B.; Shi, J.; Cai, Q.; Zheng, W.; Williams, S.M.; Long, J.; Cohen, S.S.; Li, G.; Hollis, B.W.; Smith, J.R.; et al. Common variation in vitamin D pathway genes predicts circulating 25-hydroxyvitamin D Levels among African Americans. PLoS ONE 2011, 6, e28623. [Google Scholar] [CrossRef]
- Sinotte, M.; Diorio, C.; Berube, S.; Pollak, M.; Brisson, J. Genetic polymorphisms of the vitamin D binding protein and plasma concentrations of 25-hydroxyvitamin D in premenopausal women. Am. J. Clin. Nutr. 2009, 89, 634–640. [Google Scholar] [CrossRef]
- Slater, N.A.; Rager, M.L.; Havrda, D.E.; Harralson, A.F. Genetic Variation in CYP2R1 and GC Genes Associated With Vitamin D Deficiency Status. J. Pharm. Pract. 2017, 30, 31–36. [Google Scholar] [CrossRef]
- Szili, B.; Szabo, B.; Horvath, P.; Bakos, B.; Kirschner, G.; Kosa, J.P.; Toldy, E.; Putz, Z.; Lakatos, P.; Tabak, A.; et al. Impact of genetic influence on serum total- and free 25-hydroxyvitamin-D in humans. J. Steroid Biochem. Mol. Biol. 2018, 183, 62–67. [Google Scholar] [CrossRef]
- Theodoratou, E.; Palmer, T.; Zgaga, L.; Farrington, S.M.; McKeigue, P.; Din, F.V.; Tenesa, A.; Davey-Smith, G.; Dunlop, M.G.; Campbell, H. Instrumental variable estimation of the causal effect of plasma 25-hydroxy-vitamin D on colorectal cancer risk: A mendelian randomization analysis. PLoS ONE 2012, 7, e37662. [Google Scholar] [CrossRef]
- Thongthai, P.; Chailurkit, L.O.; Chanprasertyothin, S.; Nimitphong, H.; Sritara, P.; Aekplakorn, W.; Ongphiphadhanakul, B. Vitamin D binding protein gene polymorphism as a risk factor for vitamin D deficiency in Thais. Endocr. Pract. 2015, 21, 221–225. [Google Scholar] [CrossRef]
- Touvier, M.; Deschasaux, M.; Montourcy, M.; Sutton, A.; Charnaux, N.; Kesse-Guyot, E.; Assmann, K.E.; Fezeu, L.; Latino-Martel, P.; Druesne-Pecollo, N.; et al. Determinants of vitamin D status in Caucasian adults: Influence of sun exposure, dietary intake, sociodemographic, lifestyle, anthropometric, and genetic factors. J. Investig. Dermatol. 2015, 135, 378–388. [Google Scholar] [CrossRef]
- Voipio, A.J.; Pahkala, K.A.; Viikari, J.S.; Mikkila, V.; Magnussen, C.G.; Hutri-Kahonen, N.; Kahonen, M.; Lehtimaki, T.; Mannisto, S.; Loo, B.M.; et al. Determinants of serum 25(OH)D concentration in young and middle-aged adults. The Cardiovascular Risk in Young Finns Study. Ann. Med. 2015, 47, 253–262. [Google Scholar] [CrossRef]
- Wang, W.; Ingles, S.A.; Torres-Mejia, G.; Stern, M.C.; Stanczyk, F.Z.; Schwartz, G.G.; Nelson, D.O.; Fejerman, L.; Wolff, R.K.; Slattery, M.L.; et al. Genetic variants and non-genetic factors predict circulating vitamin D levels in Hispanic and non-Hispanic White women: The Breast Cancer Health Disparities Study. Int. J. Mol. Epidemiol. Genet. 2014, 5, 31–46. [Google Scholar]
- Waterhouse, M.; Tran, B.; Armstrong, B.K.; Baxter, C.; Ebeling, P.R.; English, D.R.; Gebski, V.; Hill, C.; Kimlin, M.G.; Lucas, R.M.; et al. Environmental, personal, and genetic determinants of response to vitamin D supplementation in older adults. J. Clin. Endocrinol. Metab. 2014, 99, E1332–E1340. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Cheng, J.; Yang, K. Vitamin D-Related Gene Polymorphisms, Plasma 25-Hydroxy-Vitamin D, Cigarette Smoke and Non-Small Cell Lung Cancer (NSCLC) Risk. Int. J. Mol. Sci. 2016, 17, 1597. [Google Scholar] [CrossRef]
- Xu, W.; Sun, J.; Wang, W.; Wang, X.; Jiang, Y.; Huang, W.; Zheng, X.; Wang, Q.; Ning, Z.; Pei, Y.; et al. Association of genetic variants of vit D binding protein (DBP/GC) and of the enzyme catalyzing its 25-hydroxylation (DCYP2R1) and serum vit D in postmenopausal women. Hormones 2014, 13, 345–352. [Google Scholar] [CrossRef]
- Xu, X.; Mao, J.; Zhang, M.; Liu, H.; Li, H.; Lei, H.; Han, L.; Gao, M. Vitamin D Deficiency in Uygurs and Kazaks Is Associated with Polymorphisms in CYP2R1 and DHCR7/NADSYN1 Genes. Med. Sci. Monit. 2015, 21, 1960–1968. [Google Scholar] [CrossRef][Green Version]
- Yao, P.; Sun, L.; Lu, L.; Ding, H.; Chen, X.; Tang, L.; Xu, X.; Liu, G.; Hu, Y.; Ma, Y.; et al. Effects of Genetic and Nongenetic Factors on Total and Bioavailable 25(OH)D Responses to Vitamin D Supplementation. J. Clin. Endocrinol. Metab. 2017, 102, 100–110. [Google Scholar] [CrossRef]
- Yao, S.; Hong, C.C.; Bandera, E.V.; Zhu, Q.; Liu, S.; Cheng, T.D.; Zirpoli, G.; Haddad, S.A.; Lunetta, K.L.; Ruiz-Narvaez, E.A.; et al. Demographic, lifestyle, and genetic determinants of circulating concentrations of 25-hydroxyvitamin D and vitamin D-binding protein in African American and European American women. Am. J. Clin. Nutr. 2017, 105, 1362–1371. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Zhao, L.J.; Zhou, Y.; Badr, R.; Watson, P.; Ye, A.; Zhou, B.; Zhang, J.; Deng, H.W.; Recker, R.R.; et al. SNP rs11185644 of RXRA gene is identified for dose-response variability to vitamin D3 supplementation: A randomized clinical trial. Sci. Rep. 2017, 7, 40593. [Google Scholar] [CrossRef]
- Zhang, Z.; He, J.W.; Fu, W.Z.; Zhang, C.Q.; Zhang, Z.L. An analysis of the association between the vitamin D pathway and serum 25-hydroxyvitamin D levels in a healthy Chinese population. J. Bone Miner. Res. 2013, 28, 1784–1792. [Google Scholar] [CrossRef]
- Arabi, A.; Khoueiry-Zgheib, N.; Awada, Z.; Mahfouz, R.; Al-Shaar, L.; Hoteit, M.; Rahme, M.; Baddoura, R.; Halabi, G.; Singh, R.; et al. CYP2R1 polymorphisms are important modulators of circulating 25-hydroxyvitamin D levels in elderly females with vitamin insufficiency, but not of the response to vitamin D supplementation. Osteoporos. Int. 2017, 28, 279–290. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Yu, F.; Yu, S.; Zhang, D.; Wang, J.; Han, H.; Sun, H.; Xue, Y.; Ba, Y.; Wang, C.; et al. Triangular relationship between CYP2R1 gene polymorphism, serum 25(OH)D3 levels and T2DM in a Chinese rural population. Gene 2018, 678, 172–176. [Google Scholar] [CrossRef] [PubMed]
- Abbas, S.; Nieters, A.; Linseisen, J.; Slanger, T.; Kropp, S.; Mutschelknauss, E.J.; Flesch-Janys, D.; Chang-Claude, J. Vitamin D receptor gene polymorphisms and haplotypes and postmenopausal breast cancer risk. Breast Cancer Res. 2008, 10, R31. [Google Scholar] [CrossRef]
- Arabi, A.; Mahfoud, Z.; Zahed, L.; El-Onsi, L.; El-Hajj Fuleihan, G. Effect of age, gender and calciotropic hormones on the relationship between vitamin D receptor gene polymorphisms and bone mineral density. Eur. J. Clin. Nutr. 2010, 64, 383–391. [Google Scholar] [CrossRef]
- Carvalho, A.Y.; Bishop, K.S.; Han, D.Y.; Ellett, S.; Jesuthasan, A.; Lam, W.J.; Ferguson, L.R. The role of Vitamin D level and related single nucleotide polymorphisms in Crohn’s disease. Nutrients 2013, 5, 3898–3909. [Google Scholar] [CrossRef] [PubMed]
- Ghorbanihaghjo, A.; Argani, H.; Samadi, N.; Valizadeh, S.; Halajzadeh, J.; Yousefi, B.; Rashtchizadeh, N. Relationship between vitamin D receptor gene FokI and ApaI polymorphisms and serum levels of fetuin-A, vitamin D, and parathyroid hormone in patients on hemodialysis. Iran. J. Kidney Dis. 2014, 8, 394–400. [Google Scholar]
- Issa, C.T.; Silva, A.S.; Toscano, L.T.; Medeiros, M.S.; Persuhn, D.C.; da Silva Diniz, A.; de Carvalho Costa, M.J.; Rodrigues Goncalves Mda, C. Relationship between cardiometabolic profile, vitamin D status and BsmI polymorphism of the VDR gene in non-institutionalized elderly subjects: Cardiometabolic profile, vitamin D status and BsmI polymorphism of the VDR gene in non-institutionalized elderly subjects. Exp. Gerontol. 2016, 81, 56–64. [Google Scholar] [CrossRef]
- Kahraman, H.; Duman, B.S.; Alagol, F.; Tanakol, R.; Yilmazer, S. Lack of association between vitamin D receptor gene polymorphism (BsmI) and osteomalacia. J. Bone Miner. Metab. 2004, 22, 39–43. [Google Scholar] [CrossRef]
- Laaksonen, M.; Karkkainen, M.; Outila, T.; Vanninen, T.; Ray, C.; Lamberg-Allardt, C. Vitamin D receptor gene BsmI-polymorphism in Finnish premenopausal and postmenopausal women: Its association with bone mineral density, markers of bone turnover, and intestinal calcium absorption, with adjustment for lifestyle factors. J. Bone Miner. Metab. 2002, 20, 383–390. [Google Scholar] [CrossRef]
- Macdonald, H.M.; McGuigan, F.E.; Stewart, A.; Black, A.J.; Fraser, W.D.; Ralston, S.; Reid, D.M. Large-scale population-based study shows no evidence of association between common polymorphism of the VDR gene and BMD in British women. J. Bone Miner. Res. 2006, 21, 151–162. [Google Scholar] [CrossRef]
- Mayer, O.; Seidlerova, J.; Cerna, V.; Kucerova, A.; Karnosova, P.; Hronova, M.; Wohlfahrt, P.; Fuchsova, R.; Filipovsky, J.; Cifkova, R.; et al. Serum Vitamin D Status, Vitamin D Receptor Polymorphism, and Glucose Homeostasis in Healthy Subjects. Horm. Metab. Res. 2018, 50, 56–64. [Google Scholar] [CrossRef]
- Santos, B.R.; Lecke, S.B.; Spritzer, P.M. Apa-I polymorphism in VDR gene is related to metabolic syndrome in polycystic ovary syndrome: A cross-sectional study. Reprod. Biol. Endocrinol. 2018, 16, 38. [Google Scholar] [CrossRef] [PubMed]
- Schuch, N.J.; Garcia, V.C.; Vivolo, S.R.; Martini, L.A. Relationship between Vitamin D Receptor gene polymorphisms and the components of metabolic syndrome. Nutr. J. 2013, 12, 96. [Google Scholar] [CrossRef] [PubMed]
- Tanabe, R.; Kawamura, Y.; Tsugawa, N.; Haraikawa, M.; Sogabe, N.; Okano, T.; Hosoi, T.; Goseki-Sone, M. Effects of Fok-I polymorphism in vitamin D receptor gene on serum 25-hydroxyvitamin D, bone-specific alkaline phosphatase and calcaneal quantitative ultrasound parameters in young adults. Asia Pac. J. Clin. Nutr. 2015, 24, 329–335. [Google Scholar] [CrossRef] [PubMed]
- Vupputuri, M.R.; Goswami, R.; Gupta, N.; Ray, D.; Tandon, N.; Kumar, N. Prevalence and functional significance of 25-hydroxyvitamin D deficiency and vitamin D receptor gene polymorphisms in Asian Indians. Am. J. Clin. Nutr. 2006, 83, 1411–1419. [Google Scholar] [CrossRef]
- Yao, S.; Zirpoli, G.; Bovbjerg, D.H.; Jandorf, L.; Hong, C.C.; Zhao, H.; Sucheston, L.E.; Tang, L.; Roberts, M.; Ciupak, G.; et al. Variants in the vitamin D pathway, serum levels of vitamin D, and estrogen receptor negative breast cancer among African-American women: A case-control study. Breast Cancer Res. 2012, 14, R58. [Google Scholar] [CrossRef]
- Zhou, J.C.; Zhu, Y.M.; Chen, Z.; Mo, J.L.; Xie, F.Z.; Wen, Y.H.; Guo, P.; Peng, J.; Xu, J.; Wang, J.; et al. Oral vitamin D supplementation has a lower bioavailability and reduces hypersecretion of parathyroid hormone and insulin resistance in obese Chinese males. Public Health Nutr. 2015, 18, 2211–2219. [Google Scholar] [CrossRef]
- Abu El Maaty, M.A.; Hassanein, S.I.; Sleem, H.M.; Gad, M.Z. Effect of polymorphisms in the NADSYN1/DHCR7 locus (rs12785878 and rs1790349) on plasma 25-hydroxyvitamin D levels and coronary artery disease incidence. J. Nutr. Nutr. 2013, 6, 327–335. [Google Scholar] [CrossRef]
- Selvaraj, P.; Prabhu Anand, S.; Harishankar, M.; Alagarasu, K. Plasma 1,25 dihydroxy vitamin D3 level and expression of vitamin d receptor and cathelicidin in pulmonary tuberculosis. J. Clin. Immunol. 2009, 29, 470–478. [Google Scholar] [CrossRef]
- Majid, H.; Khan, A.H.; Moatter, T. R990G polymorphism of calcium sensing receptor gene is associated with high parathyroid hormone levels in subjects with vitamin D deficiency: A cross-sectional study. Biomed. Res. Int. 2015, 2015, 407159. [Google Scholar] [CrossRef]
- Al-Daghri, N.M.; Al-Attas, O.S.; Krishnaswamy, S.; Yakout, S.M.; Mohammed, A.K.; Alenad, A.M.; Chrousos, G.P.; Alokail, M.S. Association between promoter region genetic variants of PTH SNPs and serum 25(OH)-vitamin D level. Int. J. Clin. Exp. Pathol. 2015, 8, 8463–8471. [Google Scholar]
- Hopkinson, N.S.; Li, K.W.; Kehoe, A.; Humphries, S.E.; Roughton, M.; Moxham, J.; Montgomery, H.; Polkey, M.I. Vitamin D receptor genotypes influence quadriceps strength in chronic obstructive pulmonary disease. Am. J. Clin. Nutr. 2008, 87, 385–390. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Ma, L.H.; Wang, H.Y.; Zhang, W.; Tian, Q.; Cao, D.N.; Zheng, G.X.; Sun, Y.L. Association between polymorphisms of vitamin D receptor gene ApaI, BsmI and TaqI and muscular strength in young Chinese women. Int. J. Sports Med. 2006, 27, 182–186. [Google Scholar] [CrossRef] [PubMed]
- Iki, M.; Saito, Y.; Dohi, Y.; Kajita, E.; Nishino, H.; Yonemasu, K.; Kusaka, Y. Greater trunk muscle torque reduces postmenopausal bone loss at the spine independently of age, body size, and vitamin D receptor genotype in Japanese women. Calcif. Tissue Int. 2002, 71, 300–307. [Google Scholar] [CrossRef]
- Gussago, C.; Arosio, B.; Guerini, F.R.; Ferri, E.; Costa, A.S.; Casati, M.; Bollini, E.M.; Ronchetti, F.; Colombo, E.; Bernardelli, G.; et al. Impact of vitamin D receptor polymorphisms in centenarians. Endocrine 2016, 53, 558–564. [Google Scholar] [CrossRef] [PubMed]
- Onder, G.; Capoluongo, E.; Danese, P.; Settanni, S.; Russo, A.; Concolino, P.; Bernabei, R.; Landi, F. Vitamin D receptor polymorphisms and falls among older adults living in the community: Results from the ilSIRENTE study. J. Bone Miner. Res. 2008, 23, 1031–1036. [Google Scholar] [CrossRef]
- Walsh, S.; Ludlow, A.T.; Metter, E.J.; Ferrucci, L.; Roth, S.M. Replication study of the vitamin D receptor (VDR) genotype association with skeletal muscle traits and sarcopenia. Aging Clin. Exp. Res. 2016, 28, 435–442. [Google Scholar] [CrossRef]
- Bahat, G.; Saka, B.; Erten, N.; Ozbek, U.; Coskunpinar, E.; Yildiz, S.; Sahinkaya, T.; Karan, M.A. BsmI polymorphism in the vitamin D receptor gene is associated with leg extensor muscle strength in elderly men. Aging Clin. Exp. Res. 2010, 22, 198–205. [Google Scholar] [CrossRef]
- Barr, R.; Macdonald, H.; Stewart, A.; McGuigan, F.; Rogers, A.; Eastell, R.; Felsenberg, D.; Gluer, C.; Roux, C.; Reid, D.M. Association between vitamin D receptor gene polymorphisms, falls, balance and muscle power: Results from two independent studies (APOSS and OPUS). Osteoporos. Int. 2010, 21, 457–466. [Google Scholar] [CrossRef]
- Windelinckx, A.; De Mars, G.; Beunen, G.; Aerssens, J.; Delecluse, C.; Lefevre, J.; Thomis, M.A. Polymorphisms in the vitamin D receptor gene are associated with muscle strength in men and women. Osteoporos. Int. 2007, 18, 1235–1242. [Google Scholar] [CrossRef]
- Grundberg, E.; Brandstrom, H.; Ribom, E.L.; Ljunggren, O.; Mallmin, H.; Kindmark, A. Genetic variation in the human vitamin D receptor is associated with muscle strength, fat mass and body weight in Swedish women. Eur. J. Endocrinol. 2004, 150, 323–328. [Google Scholar] [CrossRef] [PubMed]
- Roth, S.M.; Zmuda, J.M.; Cauley, J.A.; Shea, P.R.; Ferrell, R.E. Vitamin D receptor genotype is associated with fat-free mass and sarcopenia in elderly men. J. Gerontol. A Biol. Sci. Med. Sci. 2004, 59, 10–15. [Google Scholar] [CrossRef] [PubMed]
- Geusens, P.; Vandevyver, C.; Vanhoof, J.; Cassiman, J.J.; Boonen, S.; Raus, J. Quadriceps and grip strength are related to vitamin D receptor genotype in elderly nonobese women. J. Bone Miner. Res. 1997, 12, 2082–2088. [Google Scholar] [CrossRef]
- Cheng, J.B.; Levine, M.A.; Bell, N.H.; Mangelsdorf, D.J.; Russell, D.W. Genetic evidence that the human CYP2R1 enzyme is a key vitamin D 25-hydroxylase. Proc. Natl. Acad. Sci. USA 2004, 101, 7711–7715. [Google Scholar] [CrossRef]
- Duan, L.; Xue, Z.; Ji, H.; Zhang, D.; Wang, Y. Effects of CYP2R1 gene variants on vitamin D levels and status: A systematic review and meta-analysis. Gene 2018, 678, 361–369. [Google Scholar] [CrossRef]
- Ramos-Lopez, E.; Bruck, P.; Jansen, T.; Herwig, J.; Badenhoop, K. CYP2R1 (vitamin D 25-hydroxylase) gene is associated with susceptibility to type 1 diabetes and vitamin D levels in Germans. Diabetes Metab. Res. Rev. 2007, 23, 631–636. [Google Scholar] [CrossRef]
- Bikle, D.D.; Schwartz, J. Vitamin D Binding Protein, Total and Free Vitamin D Levels in Different Physiological and Pathophysiological Conditions. Front. Endocrinol. 2019, 10, 317. [Google Scholar] [CrossRef]
- Wang, T.J.; Zhang, F.; Richards, J.B.; Kestenbaum, B.; van Meurs, J.B.; Berry, D.; Kiel, D.P.; Streeten, E.A.; Ohlsson, C.; Koller, D.L.; et al. Common genetic determinants of vitamin D insufficiency: A genome-wide association study. Lancet 2010, 376, 180–188. [Google Scholar] [CrossRef]
- Abboud, M.; Puglisi, D.A.; Davies, B.N.; Rybchyn, M.; Whitehead, N.P.; Brock, K.E.; Cole, L.; Gordon-Thomson, C.; Fraser, D.R.; Mason, R.S. Evidence for a specific uptake and retention mechanism for 25-hydroxyvitamin D (25OHD) in skeletal muscle cells. Endocrinology 2013, 154, 3022–3030. [Google Scholar] [CrossRef]
- Fedirko, V.; Mandle, H.B.; Zhu, W.; Hughes, D.J.; Siddiq, A.; Ferrari, P.; Romieu, I.; Riboli, E.; Bueno-de-Mesquita, B.; van Duijnhoven, F.J.B.; et al. Vitamin D-Related Genes, Blood Vitamin D Levels and Colorectal Cancer Risk in Western European Populations. Nutrients 2019, 11, 1954. [Google Scholar] [CrossRef]
- Jorde, R.; Wilsgaard, T.; Grimnes, G. Polymorphisms in the vitamin D system and mortality—The Tromso study. J. Steroid Biochem. Mol. Biol. 2019, 195, 105481. [Google Scholar] [CrossRef]
- Malik, S.; Fu, L.; Juras, D.J.; Karmali, M.; Wong, B.Y.; Gozdzik, A.; Cole, D.E. Common variants of the vitamin D binding protein gene and adverse health outcomes. Crit. Rev. Clin. Lab. Sci. 2013, 50, 1–22. [Google Scholar] [CrossRef]
- Kato, S.; Takeyama, K.; Kitanaka, S.; Murayama, A.; Sekine, K.; Yoshizawa, T. In vivo function of VDR in gene expression-VDR knock-out mice. J. Steroid Biochem. Mol. Biol. 1999, 69, 247–251. [Google Scholar] [CrossRef]
- Girgis, C.M.; Clifton-Bligh, R.J.; Hamrick, M.W.; Holick, M.F.; Gunton, J.E. The roles of vitamin D in skeletal muscle: Form, function, and metabolism. Endocr. Rev. 2013, 34, 33–83. [Google Scholar] [CrossRef]
- Bouillon, R.; Carmeliet, G.; Verlinden, L.; van Etten, E.; Verstuyf, A.; Luderer, H.F.; Lieben, L.; Mathieu, C.; Demay, M. Vitamin D and human health: Lessons from vitamin D receptor null mice. Endocr. Rev. 2008, 29, 726–776. [Google Scholar] [CrossRef]
- Holick, M.F. Vitamin D deficiency. N. Engl. J. Med. 2007, 357, 266–281. [Google Scholar] [CrossRef] [PubMed]
- Laczmanski, L.; Jakubik, M.; Bednarek-Tupikowska, G.; Rymaszewska, J.; Sloka, N.; Lwow, F. Vitamin D receptor gene polymorphisms in Alzheimer’s disease patients. Exp. Gerontol. 2015, 69, 142–147. [Google Scholar] [CrossRef]
- Uitterlinden, A.G.; Fang, Y.; Van Meurs, J.B.; Pols, H.A.; Van Leeuwen, J.P. Genetics and biology of vitamin D receptor polymorphisms. Gene 2004, 338, 143–156. [Google Scholar] [CrossRef]
- Vuolo, L.; Di Somma, C.; Faggiano, A.; Colao, A. Vitamin D and cancer. Front. Endocrinol. 2012, 3, 58. [Google Scholar] [CrossRef] [PubMed]
- Kelishadi, R.; Heidari-Beni, M.; Akbarian, S.A.; Hasan Tajadini, M.; Haghjooy Javanmard, S. Genetic Variation in Cytochrome P450 2R1 and Vitamin D Binding Protein Genes are associated with Vitamin D Deficiency in Adolescents. Int. J. Vitam. Nutr. Res. 2020, 90, 339–345. [Google Scholar] [CrossRef]
- Yuan, C.; Renfro, L.; Ambadwar, P.B.; Ou, F.S.; McLeod, H.L.; Innocenti, F.; Meyerhardt, J.A.; Wolpin, B.M.; Goldberg, R.M.; Grothey, A.; et al. Influence of genetic variation in the vitamin D pathway on plasma 25-hydroxyvitamin D3 levels and survival among patients with metastatic colorectal cancer. Cancer Causes Control. 2019, 30, 757–765. [Google Scholar] [CrossRef]
- Carter, G.D. Accuracy of 25-hydroxyvitamin D assays: Confronting the issues. Curr. Drug Targets 2011, 12, 19–28. [Google Scholar] [CrossRef] [PubMed]
- Giustina, A.; Bouillon, R.; Binkley, N.; Sempos, C.; Adler, R.A.; Bollerslev, J.; Dawson-Hughes, B.; Ebeling, P.R.; Feldman, D.; Heijboer, A.; et al. Controversies in Vitamin D: A Statement From the Third International Conference. JBMR Plus 2020, 4, e10417. [Google Scholar] [CrossRef]
- Beaudart, C.; Buckinx, F.; Rabenda, V.; Gillain, S.; Cavalier, E.; Slomian, J.; Petermans, J.; Reginster, J.Y.; Bruyere, O. The effects of vitamin D on skeletal muscle strength, muscle mass, and muscle power: A systematic review and meta-analysis of randomized controlled trials. J. Clin. Endocrinol. Metab. 2014, 99, 4336–4345. [Google Scholar] [CrossRef]
- Broe, K.E.; Chen, T.C.; Weinberg, J.; Bischoff-Ferrari, H.A.; Holick, M.F.; Kiel, D.P. A higher dose of vitamin d reduces the risk of falls in nursing home residents: A randomized, multiple-dose study. J. Am. Geriatr. Soc. 2007, 55, 234–239. [Google Scholar] [CrossRef]
- Endo, I.; Inoue, D.; Mitsui, T.; Umaki, Y.; Akaike, M.; Yoshizawa, T.; Kato, S.; Matsumoto, T. Deletion of vitamin D receptor gene in mice results in abnormal skeletal muscle development with deregulated expression of myoregulatory transcription factors. Endocrinology 2003, 144, 5138–5144. [Google Scholar] [CrossRef]
- McCarthy, E.K.; Kiely, M. Vitamin D and muscle strength throughout the life course: A review of epidemiological and intervention studies. J. Hum. Nutr. Diet. 2015, 28, 636–645. [Google Scholar] [CrossRef]
- Bjork, A.; Ribom, E.; Johansson, G.; Scragg, R.; Mellstrom, D.; Grundberg, E.; Ohlsson, C.; Karlsson, M.; Ljunggren, O.; Kindmark, A. Variations in the vitamin D receptor gene are not associated with measures of muscle strength, physical performance, or falls in elderly men. Data from MrOS Sweden. J. Steroid Biochem. Mol. Biol. 2019, 187, 160–165. [Google Scholar] [CrossRef]
- Wang, Y.; Han, H.; Wang, J.; Shen, F.; Yu, F.; Wang, L.; Yu, S.C.; Zhang, D.D.; Sun, H.L.; Xue, Y.; et al. Polymorphisms in CYP2R1 Gene Associated with Serum Vitamin D Levels and Status in a Chinese Rural Population. Biomed. Environ. Sci. 2019, 32, 550–553. [Google Scholar] [CrossRef]
- Tuncel, G.; Temel, S.G.; Ergoren, M.C. Strong association between VDR FokI (rs2228570) gene variant and serum vitamin D levels in Turkish Cypriots. Mol. Biol. Rep. 2019, 46, 3349–3355. [Google Scholar] [CrossRef]
- Ragia, G.; Archontogeorgis, K.; Simmaco, M.; Gentile, G.; Borro, M.; Zissimopoulos, A.; Froudarakis, M.; Manolopoulos, V.G.; Steiropoulos, P. Genetics of Obstructive Sleep Apnea: Vitamin D Receptor Gene Variation Affects Both Vitamin D Serum Concentration and Disease Susceptibility. OMICS 2019, 23, 45–53. [Google Scholar] [CrossRef]
- Santos, B.R.; Costa, N.C.; Silva, T.R.; Oppermann, K.; Magalhaes, J.A.; Casanova, G.; Spritzer, P.M. Prevalence of vitamin D deficiency in women from southern Brazil and association with vitamin D-binding protein levels and GC-DBP gene polymorphisms. PLoS ONE 2019, 14, e0226215. [Google Scholar] [CrossRef]
- Khan, A.H.; Jafri, L.; Siddiqui, A.; Naureen, G.; Morris, H.; Moatter, T. Polymorphisms in the GC Gene for Vitamin D Binding Protein and Their Association with Vitamin D and Bone Mass in Young Adults. J. Coll. Physicians Surg. Pak. 2019, 29, 715–719. [Google Scholar] [CrossRef]
- Pooyan, S.; Rahimi, M.H.; Mollahosseini, M.; Khorrami-Nezhad, L.; Maghbooli, Z.; Mirzaei, K. The Association between Vitamin D Deficiency and variants of Vitamin D Binding protein gene among Healthy Iranian Adults. Int. J. Vitam. Nutr. Res. 2020, 90, 249–256. [Google Scholar] [CrossRef]
- Yu, S.; Li, X.; Wang, Y.; Mao, Z.; Xie, Y.; Zhang, L.; Wang, C.; Li, W. Family-based Association between Allele T of rs4646536 in CYP27B1 and vitamin D deficiency. J. Clin. Lab. Anal. 2019, 33, e22898. [Google Scholar] [CrossRef]
- Ye, X.; Jia, J.; Zhang, N.; Ding, H.; Zhan, Y. Associations of genetic polymorphisms of the vitamin D pathway with blood pressure in a Han Chinese population. Clin. Exp. Hypertens. 2019, 41, 460–465. [Google Scholar] [CrossRef]
- Bjork, A.; Mellstrom, D.; Ohlsson, C.; Karlsson, M.; Mallmin, H.; Johansson, G.; Ljunggren, O.; Kindmark, A. Haplotypes in the CYP2R1 gene are associated with levels of 25(OH)D and bone mineral density, but not with other markers of bone metabolism (MrOS Sweden). PLoS ONE 2018, 13, e0209268. [Google Scholar] [CrossRef] [PubMed]
- Manousaki, D.; Mitchell, R.; Dudding, T.; Haworth, S.; Harroud, A.; Forgetta, V.; Shah, R.L.; Luan, J.; Langenberg, C.; Timpson, N.J.; et al. Genome-wide Association Study for Vitamin D Levels Reveals 69 Independent Loci. Am. J. Hum. Genet. 2020, 106, 327–337. [Google Scholar] [CrossRef]
| Name Gene/ID | Description | Aliases | Studies | SNPs Investigated in Association with Circulating Vitamin D Levels | |
|---|---|---|---|---|---|
| Significant Associations | Non-Significant | ||||
| GC ID:2638 | GC, vitamin D binding protein | DBP, DBP-maf, DBP/GC, GRD3, Gc-MAF, GcMAF, HEL-S-51, VDB, VDBG, VDBP | 68 different SNPs reported by 56 studies [26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67,68,69,70,71,72,73,74,75,76,77,78,79,80,81] | rs115316390, rs1155563 *, rs11939173, rs12512631 *, rs16846876 *, rs16847015, rs17467825 *, rs222020 *, rs222040, rs222054, rs2282679 *, rs2298849 *, rs2298850 *, rs3755967 *, rs4588 *, rs7041 *, rs705119, rs705120, rs842999, rs9016 | rs10011000, rs10488854, rs12640179, rs12644050, rs13117483, rs1352841, rs1352843, rs1352844, rs1352845, rs1491709, rs1491710, rs1491711, rs1491718, rs1491719, rs1565572, rs16846912, rs16846943, rs16847019, rs16847024, rs16847028, rs16847039, rs16847050, rs17383291, rs17830803, rs1873590, rs188812, rs2070741, rs222003, rs222010, rs222014, rs222016, rs222017, rs222023, rs222029, rs222035, rs222043, rs222049, rs2276461, rs3733359, rs3737549, rs3775152, rs4694105, rs4752, rs6817912, rs6835052, rs705117, rs705124, rs843006 |
| CYP2R1 ID:120227 | cytochrome P450 family 2 subfamily R member 1 | - | 29 different SNPs reported by 41 studies [27,28,29,30,31,32,33,34,35,36,38,39,42,44,46,47,48,50,51,52,55,56,57,60,62,63,65,67,68,69,72,73,74,76,77,78,79,80,81,82,83] | rs10500804 *, rs10741657 *, rs10766197 *, rs10832306, rs11023374 *, rs11023380 * rs12794714 *, rs1562902 *, rs1993116 *, rs2060793 *, rs7116978, rs7935792 | rs1037379, rs10832312, rs10832313, rs11023371, rs114050796, rs11819875, rs12418214, rs12419657, rs1496167, rs16930609, rs16930625, rs206793, rs7117967, rs7125348, rs7129781, rs7936142, rs952301 |
| VDR ID:7421 | vitamin D receptor | NR1I1, PPP1R163 | 111 different SNPs reported by 41 studies [27,28,29,30,33,37,41,42,46,47,50,52,55,56,57,60,61,62,63,65,67,71,74,75,77,78,84,85,86,87,88,89,90,91,92,93,94,95,96,97,98] | rs10783219 *, rs11568820, rs1544410 *, rs2228570 * (merged rs10735810), rs2239186 *, rs2408876, rs4516035, rs7139166, rs731236 *, rs7968585, rs7975232 * | rs10083198, rs10747524, rs10783215, rs10783218, rs10875693, rs10875694, rs10875695, rs10875702, rs10875712, rs11168264, rs11168266, rs11168268, rs11168275, rs11168277, rs11168287, rs11168288, rs11168292, rs11168302, rs11168314, rs11540149, rs11574024, rs11574026, rs11574027, rs11574038, rs11574042, rs11574044, rs11574065, rs11574077, rs11574110, rs11574113, rs11574138, rs11574141, rs11574143, rs11608702, rs11834903, rs12308082, rs12314197, rs12370156, rs12717991, rs12721364, rs12721365, rs12721370, rs1540339, rs17882106, rs1859281, rs1989969, rs2071358, rs2107301, rs2189480, rs2238135, rs2238136, rs2238138, rs2239179, rs2239180, rs2239181, rs2239182, rs2239184, rs2239185, rs2248098, rs2254210, rs2283342, rs2525044, rs2525045, rs2544027, rs2544038, rs2853559, rs2853560, rs2853564, rs3782905, rs3819545, rs3847987, rs4077869, rs4328262, rs4334089, rs4442605, rs4760648, rs4760655, rs4760658, rs4760674, rs6580642, rs7136534, rs7299460, rs7302038, rs7302235, rs7305032, rs7310552, rs7311030, rs739837, rs757343, rs7962898, rs7963776, rs7965281, rs7967152, rs7971418, rs7975128, rs7976091, rs881383, rs886441, rs9729, rs987849 |
| CYP24A1 ID:1591 | cytochrome P450 family 24 subfamily A member 1 | CP24, CYP24, HCAI, HCINF1, P450-CC24 | 65 different SNPs reported by 31 studies [26,27,28,29,30,31,32,33,35,38,42,46,52,55,56,57,60,61,62,65,68,69,71,72,73,74,75,78,80,81,97] | rs17216707, rs2209314 *, rs2762939 *, rs2762941, rs6013897 *, rs73913757 | rs11907350, rs13038432, rs1555439, rs1570669, rs1570670, rs17219315, rs1870969, rs2021940, rs2181874, rs2244719, rs2245153, rs2248137, rs2248359, rs2248461, rs2274130, rs2296239, rs2296241, rs2426496, rs2426498, rs2585413, rs2585422, rs2585423, rs2585428, rs2585439, rs2762926, rs2762929, rs2762932, rs35051736, rs3787555, rs3787557, rs3886163, rs4809957, rs4809958, rs4809959, rs4809960, rs6013905, rs6022990, rs6022999, rs6023005, rs6023009, rs6023012, rs6068810, rs6068812, rs6068816, rs6068824, rs6097797, rs6097801, rs6097805, rs6097809, rs6127112, rs6127119, rs73913755, rs751090, rs765058, rs765059, rs8124792, rs912505, rs927650, rs927651 |
| DHCR7 ID:1717 | 7-dehydrocholesterol reductase | SLOS | 25 different SNPs reported by 28 studies [27,28,29,31,32,33,35,36,38,39,42,44,46,51,52,55,56,57,60,67,69,72,73,74,77,78,81,99] | rs11603330, rs12785878 *, rs1790349 * | rs11233570, rs11234027, rs11606033, rs12419279, rs12800438, rs1540129, rs1540130, rs1790325, rs1790329, rs1790334, rs1790373, rs1792272, rs1792284, rs3794060, rs3829251, rs4316537, rs4944957, rs4945008, rs7122671, rs7944926, rs7950649, rs949178 |
| CYP27B1 ID:1594 | cytochrome P450 family 27 subfamily B member 1 | CP2B, CYP1, CYP1alpha, CYP27B, P450c1, PDDR, VDD1, VDDR, VDDRI, VDR | 15 different SNPs reported by 22 studies [26,27,28,30,33,37,41,42,46,52,55,56,60,62,65,74,75,77,80,81,97,100] | rs10877012 * | rs1021469, rs1048691, rs10877011, rs10877013, rs12368653, rs2269720, rs3782130, rs4646536, rs4646537, rs4760169, rs703842, rs8176344, rs8176345, −1077 |
| CYP27A1 ID:1593 | cytochrome P450 family 27 subfamily A member 1 | CP27, CTX, CYP27 | 27 different SNPs reported by 7 studies [26,28,30,33,74,80,81] | -- | rs116071925, rs11677711, rs12623740, rs12694443, rs12987009, rs12990447, rs13013510, rs13382651, rs17470271, rs4646535, rs4674338, rs4674344, rs4674345, rs6436084, rs6436094, rs645163, rs647952, rs6709815, rs6716642, rs6723334, rs6740004, rs6751527, rs7566656, rs7568196, rs7594289, rs7603709, rs933994 |
| CASR ID:846 | calcium-sensing receptor | CAR, EIG8, FHH, FIH, GPRC2A, HHC, HHC1, HYPOC1, NSHPT, PCAR1, hCasR | 71 different SNPs reported by 6 studies [27,60,71,74,80,101] | rs17251221, rs1801725, | rs10222633, rs1042636, rs10934578, rs11715859, rs11716910, rs12485716, rs13093602, rs13324814, rs13327652, rs1354162, rs1393198, rs1501892, rs1501898, rs1501900, rs16832787, rs17203502, rs17203516, rs17282008, rs1801726, rs1814740, rs1847029, rs1973490, rs1979869, rs2036399, rs2134223, rs2134224, rs2173961, rs2202127, rs2221266, rs2270916, rs2279802, rs3749203, rs3749207, rs3792288, rs3792291, rs3804592, rs3804593, rs3804595, rs3845918, rs4677900, rs4678013, rs4678029, rs4678031, rs4678035, rs4678173, rs4678174, rs6438705, rs6438706, rs6438712, rs6764205, rs6764544, rs6768109, rs6776158, rs6799828, rs7614486, rs7617603, rs7628990, rs7635128, rs7639847, rs7644981, rs7647405, rs7648041, rs937626, rs9740, rs9820206, rs9826770, rs9866419, rs9875101, rs9875636 |
| PTH ID: 5741 | parathyroid hormone [Homo sapiens (human)] | PTH1 | 12 different SNPs reported by 6 studies [30,74,80,81,96,102] | rs10500783, rs1459015, | rs2593570, rs6254, rs6256, rs6264, rs694, rs10500784, rs177706, rs192802, rs3099597, rs751610 |
| CYP3A4 ID:1576 | cytochrome P450 family 3 subfamily A member 4 | CP33, CP34, CYP3A, CYP3A3, CYPIIIA3, CYPIIIA4, HLP, NF-25, P450C3, P450PCN1 | 9 different SNPs reported by 5 studies [28,30,62,80,81] | rs2242480 | rs12333983, rs2246709, rs2687116, rs2740574, rs35599367, rs3735451, rs4646437, rs6956344 |
| RXRA ID: 6256 | retinoid X receptor alpha | NR2B1 | 48 different SNPs reported by 3 studies [52,71,80] | rs11185644 * | rs1045570, rs10785870, rs10881577, rs10881578, rs10881580, rs10881582, rs10881583, rs11102986, rs11103473, rs11103482, rs11185647, rs11185659, rs12004589, rs12004786, rs1536475, rs1805348, rs1805352, rs12339187, rs3118523, rs3118526, rs3118536, rs3118570, rs3118571, rs3132294, rs3132296, rs3132299, rs3132300, rs34677682, rs35603635, rs3818738, rs3818739, rs3818740, rs4240705, rs4917347, rs4917352, rs4917353, rs4917354, rs6537944, rs7039190, rs7861779, rs7864987, rs7871655, rs842196, rs872298, rs877954, rs881657, rs914853 |
| CUBN ID:8029 | cubilin [Homo sapiens (human)] | IFCR, MGA1, gp280 | 14 different SNPs reported by 1 study [81] | -- | rs10904881, rs11254370, rs1687705, rs1801222, rs1801223, rs1801224, rs1801225, rs1801231, rs1801232, rs1801234, rs1801241, rs2271462, rs3740165, rs703064 |
| RXRB ID:6257 | retinoid X receptor beta [Homo sapiens (human)] | DAUDI6, H-2RIIBP, NR2B2, RCoR-1 | 4 different SNPs reported by 1 study [81] | -- | rs6531, rs2076310, rs3117040, rs9277935 |
| SNP/Tra-ditional Name | Region, Reference | Participants (Number, Gender, Age) | Outcomes for Muscle Mass and Function | Main Findings |
|---|---|---|---|---|
| rs7975232/ ApaI | Tianjin, China [104] | n = 109 f (AA: 19.57 ± 0.53 y, Aa: 20.00 ± 1.20 y, aa: 19.89 ± 1.05 y) | Concentric and eccentric peak torque of knee extensors and flexors at 30°/s, 60°/s, and 180°/s; concentric peak torque of elbow extensors and flexors at 30°/s and 120°/s | aa + aA genotypes→higher knee extension peak torque at 120°s, higher elbow flexion at 120°/s and 30°/s than AA |
| Fuki Prefecture, Japan [105] | n = 180 f; 60.1 ± 6.6 y | Handgrip strength; isokinetic concentric peak torque of knee extensors and flexors; isometric and isokinetic concentric and eccentric peak torque of trunk flexors and extensors | No significant differences in any muscle strength parameter between ApaI genotype groups (AA, Aa, aa) (data not shown in article) | |
| Northern Italy [106] | Centenarians (n = 102, 102.3 ± 0.3 y) versus septuagenarians (n = 163; 73.0 ± 0.6 y) | Handgrip strength | FF→significantly higher handgrip strength than Ff + ff | |
| rs1544410/ BsmI | Sirente area, Italy [107] | n = 259 (87 m + 172 f); 85.0 ± 4.5 y | Handgrip strength, short physical performance battery (SPPB) | No significant differences between genotypes (BB, Bb, bb) in handgrip strength and SPPB score |
| Baltimore, USA [108] | n = 864 (489 m + 375 f); 22–90 y; | Total and appendicular fat-free mass (DXA); handgrip strength; concentric peak torque of knee extensors at 30°/s and 180°/s; isometric peak torque at 120° and 140° knee ankle | No significant differences in fat-free mass Only modest differences in strength measurements: knee extensor isokinetic peak torque at 30°/s in females (bb > bB > BB) | |
| Northern Italy [106] | Centenarians (n = 102, 102.3 ± 0.3 y) versus septuagenarians (n = 163; 73.0 ± 0.6 y) | Handgrip strength | No significant differences in handgrip strength between genotypes | |
| Istanbul, Turkey [109] | n = 120 m; 69.0 ± 6.9 y | Fat-free mass (BIA); isokinetic peak torque of knee extensors, flexors at 60°/s | BB→higher knee extensor strength as compared to Bb + bb; No differences for flexors, muscle mass, and vitamin D level | |
| UK, Germany, France [110] | APUSS cohort (n = 3234 f; 54.3 ± 2.3 y); OPUS cohort (n = 1970 f; 66.9 ± 7.0 y) | Handgrip strength; chair rise test (difficulty and power with force plate); data only from OPUS cohort | BB + bB genotypes→higher max power, fewer difficulties to stand up from a chair No differences in handgrip strength | |
| London, UK [103] | COPD (n = 107; 75 m + 32 f; 63.5 ± 9.5y); age-matched controls (n= 104; 48 m + 56 f; 61.8 ± 8.5 y) | Handgrip strength; quadriceps strength, calculated as quadriceps maximum voluntary contraction force; fat-free mass (BIA) | No significant differences between genotypes for fat-free mass and strength measures | |
| Leuven, Belgium [111] | 493(253 m (54.9 ± 10.2 y) + 240 f (41.5 ± 13.2 y)) | Fat-free mass (estimated from skinfolds); handgrip strength; Isometric knee extension strength at 150°, 120°, and 90° knee ankle; Isometric knee flexion strength at 120° | No significant differences in any of the measured parameters [data not shown] | |
| Tianjin, China [104] | n = 109f (AA: 19.57 ± 0.53 y, Aa: 20.00 ± 1.20 y, aa: 19.89 ± 1.05 y) | Concentric and eccentric peak torque of knee extensors and flexors at 30°/s, 60°/s, and 180°/s; concentric peak torque of elbow extensors and flexors at 30°/s and 120°/s | BB + Bb genotypes→higher knee flexion peak torque at 180°/s than bb group No other differences | |
| Uppsala, Sweden [112] | n = 175 f; 29.6 ± 5.9 | Handgrip strength; Isokinetic knee-flexion and extension strength at 90°/s; lean body mass (DXA) | BB→higher hamstring strength as compared to bb No differences in lean mass | |
| Monongahela Valley, USA [113] | n = 302 m; 58−93 y | Peak and average isometric quadriceps strength at 125° knee ankle; appendicular and total fat-free mass (DXA) | No differences in any measured parameter | |
| n/A [114] | n = 501 f; >70 y (mean age: 75 y) | isometric muscle strength of the quadriceps; handgrip strength | bb + Bb→higher quadriceps strength than BB in non-obese females, but not in obese females | |
| rs2228570/ FokI (rs10735810) | Sirente area, Italy [107] | n = 259 (87 m + 172f ); 85.0 ± 4.5 y | Handgrip strength, short physical performance battery (SPPB) | No significant differences between genotypes (FF, Ff, ff) in handgrip strength and SPPB score |
| Baltimore, USA [108] | n = 864 (489 m+ 375 f); 22–90 y; | Total and appendicular fat-free mass (DXA); handgrip strength; concentric peak torque of knee extensors at 30°/s and 180°/s; isometric peak torque at 120° and 140° knee ankle | No significant differences in fat-free mass; Males: no significant differences for any strength variable measured; Females: ff→higher isometric quadriceps strength (120°) in comparison to Ff and FF | |
| Northern Italy [106] | Centenarians (n = 102, 102.3 ± 0.3 y) versus septuagenarians (n = 163; 73.0 ± 0.6 y) | Handgrip strength | FF→significantly higher handgrip strength than Ff + ff | |
| Istanbul, Turkey [109] | n = 120 m; 69.0 ± 6.9 y | Fat-free mass (BIA); isokinetic peak torque of knee extensors, flexors at 60°/s | No significant differences between genotypes (FF, Ff, ff) | |
| UK, Germany, France [110] | APUSS cohort (n = 3234 f; 54.3 ± 2.3 y); OPUS cohort (n = 1970 f; 66.9 ± 7.0 y) | Handgrip strength; chair rise test (difficulty and power with force plate); data only from OPUS cohort | No significant differences between genotypes (FF, Ff, ff) | |
| London, UK [103] | cases with stable COPD (n = 107; 63.5 ± 9.5 y) and healthy age-matched controls (n= 104; 61.8 ± 8.5 y) | Handgrip strength; quadriceps strength, calculated as quadriceps maximum voluntary contraction force; fat-free mass (BIA) | FF→significantly lower quadriceps strength than Ff + ff | |
| Leuven, Belgium [111] | 493(253 m (54.9 ± 10.2 y) + 240 f (41.5 ± 13.2 y)) | Fat-free mass (estimated from skinfolds); handgrip strength; Maximal isometric knee extension strength at 150°, 120°, and 90° knee ankle; Maximal isometric knee flexion strength at 120° knee ankle; | Females: Ff→lower isometric knee extension strength at 120° and 90° than both FF and ff Males: no significant differences for any of the strength measurements | |
| Monongahela Valley, USA [113] | n = 302 m; 58–93 y | Peak and average isometric quadriceps strength at 125° knee ankle; appendicular and total fat-free mass (DXA) | FF→significantly lower appendicular and total fat-free mass than Ff and ff; FF→significantly lower peak and average isometric quadriceps strength than ff | |
| rs731236/TaqI | Tianjin, China [104] | n = 109f (AA: 19.57 ± 0.53 y, Aa: 20.00 ± 1.20 y, aa: 19.89 ± 1.05 y) | Concentric and eccentric peak torque of knee extensors and flexors at 30°/s, 60°/s, and 180°/s; concentric peak torque of elbow extensors and flexors at 30°/s and 120°/s | No significant differences between genotypes (TT, Tt, tt) |
| Fuki Prefecture, Japan [105] | n = 180 f; 60.1 ± 6.6 y | Handgrip strength; isokinetic concentric peak torque of knee extensors and flexors; isometric and isokinetic concentric and eccentric peak torque of trunk flexors and extensors | No significant differences between genotypes (TT, Tt, tt) | |
| Baltimore, USA [108] | n = 864 (489 m + 375 f); 22–90 y; | Total and appendicular fat-free mass (DXA); handgrip strength; concentric peak torque of knee extensors at 30°/s and 180°/s; isometric peak torque at 120° and 140° knee ankle | No significant differences in fat-free mass; Males: no significant differences for any strength variable measured; Females: ff→higher isometric quadriceps strength (120°) in comparison to Ff and FF | |
| Northern Italy [106] | Centenarians (n = 102, 102.3 ± 0.3 y) versus septuagenarians (n = 163; 73.0 ± 0.6 y) | Handgrip strength | FF→significantly higher handgrip strength than Ff + ff | |
| Istanbul, Turkey [109] | n = 120 m; 69.0 ± 6.9 y | Fat-free mass (BIA); isokinetic peak torque of knee extensors, flexors at 60°/s | No significant differences between genotypes (TT, Tt, tt) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Krasniqi, E.; Boshnjaku, A.; Wagner, K.-H.; Wessner, B. Association between Polymorphisms in Vitamin D Pathway-Related Genes, Vitamin D Status, Muscle Mass and Function: A Systematic Review. Nutrients 2021, 13, 3109. https://doi.org/10.3390/nu13093109
Krasniqi E, Boshnjaku A, Wagner K-H, Wessner B. Association between Polymorphisms in Vitamin D Pathway-Related Genes, Vitamin D Status, Muscle Mass and Function: A Systematic Review. Nutrients. 2021; 13(9):3109. https://doi.org/10.3390/nu13093109
Chicago/Turabian StyleKrasniqi, Ermira, Arben Boshnjaku, Karl-Heinz Wagner, and Barbara Wessner. 2021. "Association between Polymorphisms in Vitamin D Pathway-Related Genes, Vitamin D Status, Muscle Mass and Function: A Systematic Review" Nutrients 13, no. 9: 3109. https://doi.org/10.3390/nu13093109
APA StyleKrasniqi, E., Boshnjaku, A., Wagner, K.-H., & Wessner, B. (2021). Association between Polymorphisms in Vitamin D Pathway-Related Genes, Vitamin D Status, Muscle Mass and Function: A Systematic Review. Nutrients, 13(9), 3109. https://doi.org/10.3390/nu13093109








