Hesperidin Is a Potential Inhibitor against SARS-CoV-2 Infection
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Lines and Cell Culture
2.2. Measurement of Luciferase Intensity of SARS-CoV-2 Spike Protein-Pseudotyped Lentiviral Particles
2.3. Measurement of 3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium Bromide (MTT)
2.4. Docking-Pose Prediction
2.5. FRET-Based Enzyme Activity Assay
2.6. Western Blotting
2.7. Statistical Analysis
3. Results
3.1. Molecular Docking Reveals HT and HD as Potential Multiple-Target Inhibitors against COVID-19
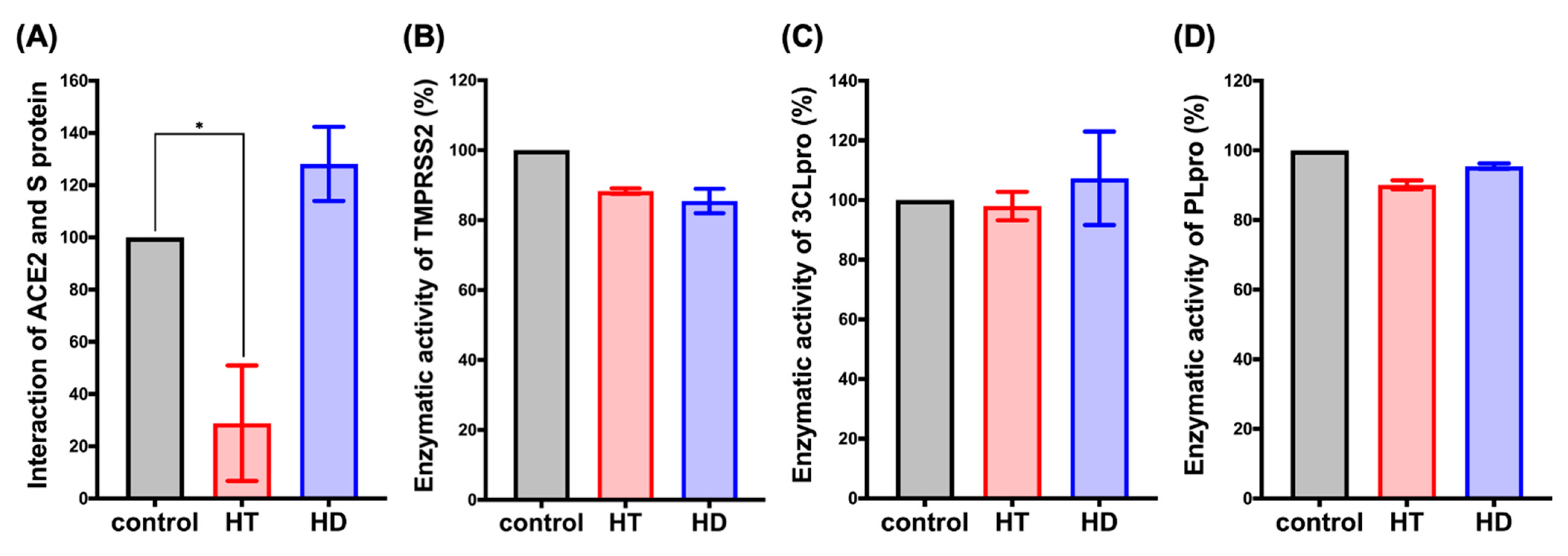
3.2. HT Suppresses the Interaction between ACE2 and the Spike Protein of SARS-CoV-2 In Vitro
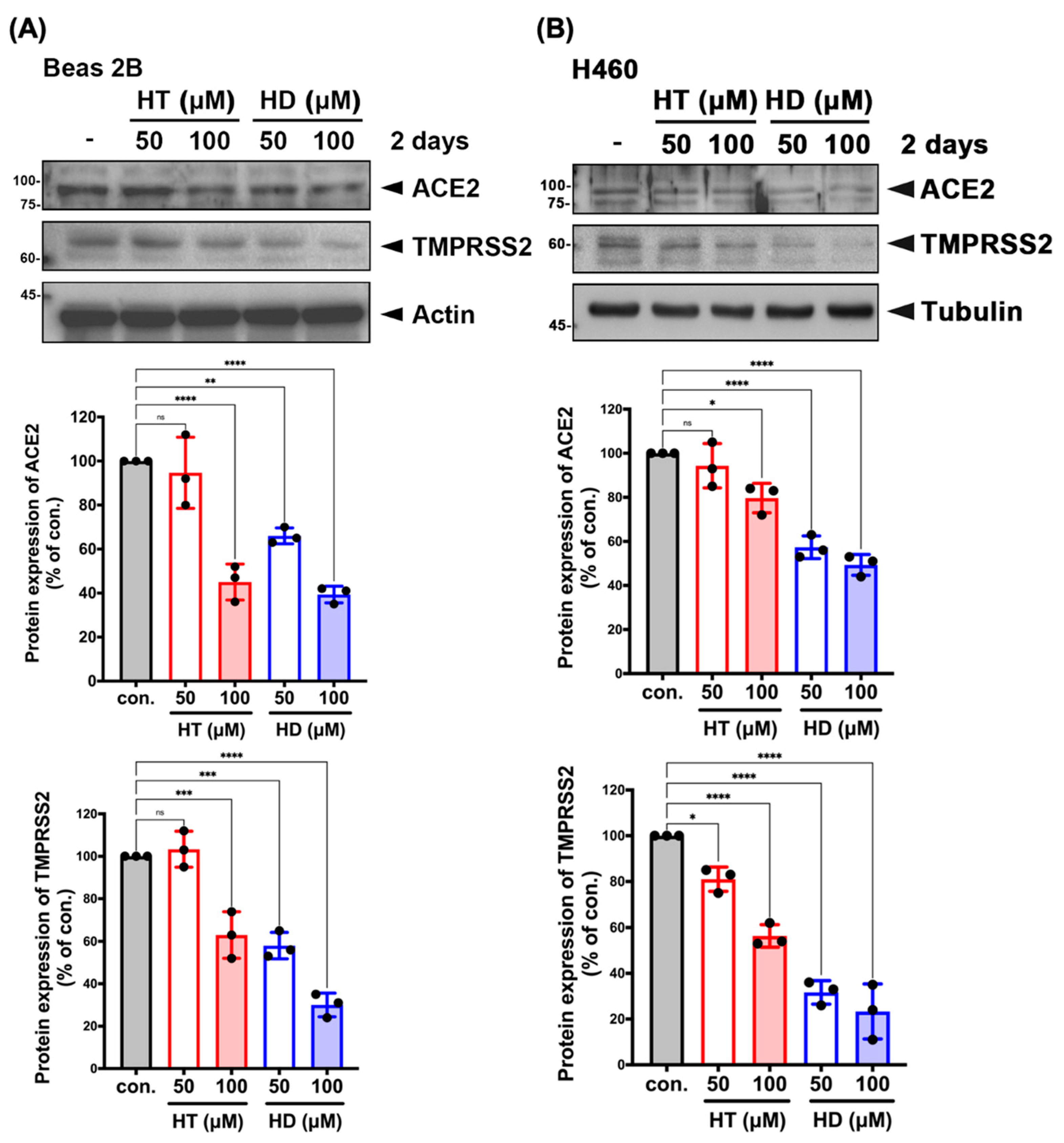
3.3. HT and HD Downregulated the Protein Expression of ACE2 and TMPRSS2 in Normal and Malignant Lung Cells
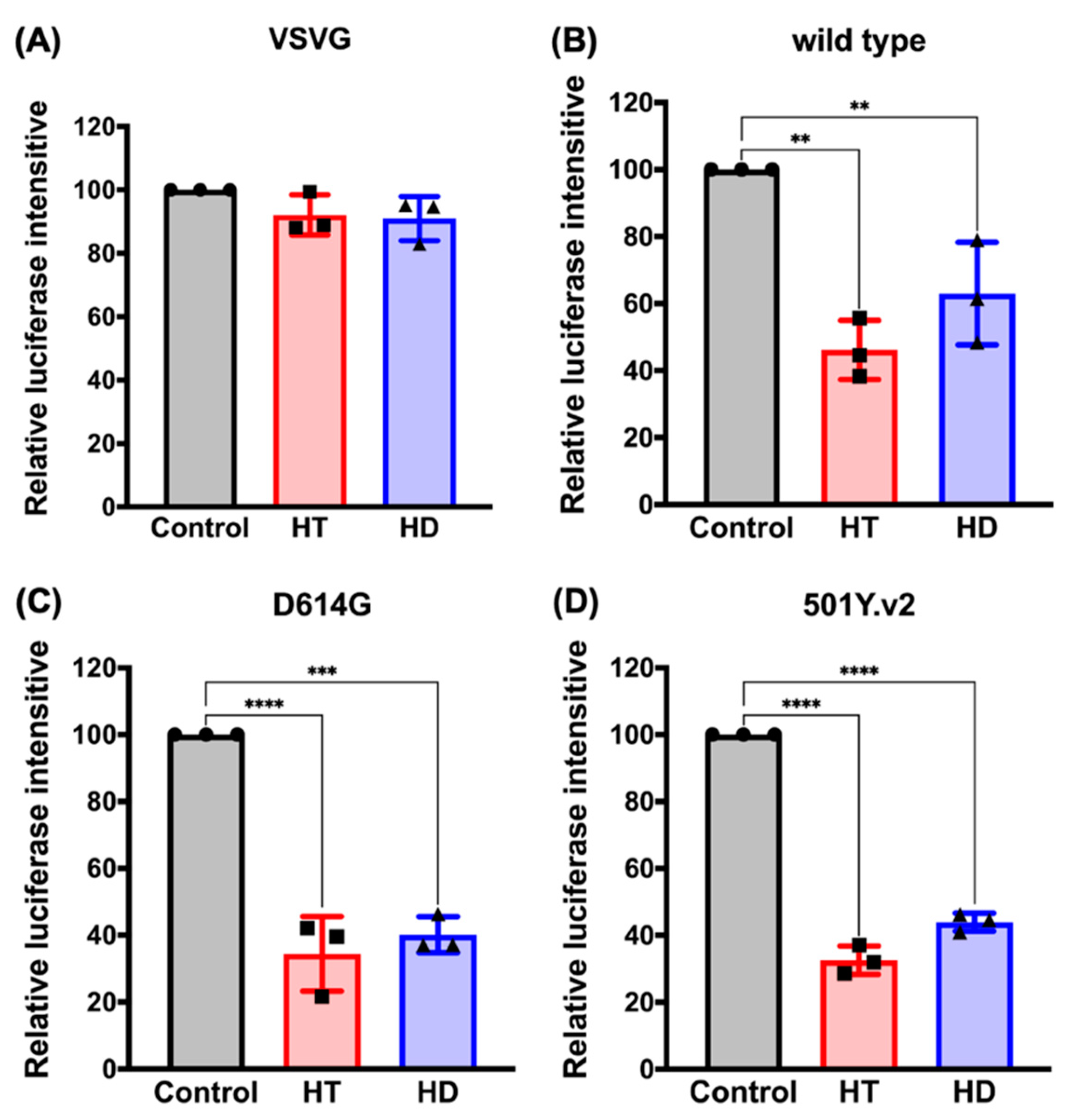
3.4. HT and HD Block the Cellular Entry of Vpp of SARS-CoV-2 Spike Protein (SARS-CoV-2-S)
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Acknowledgments
Conflicts of Interest
References
- Del Rio, C.; Malani, P.N. COVID-19-New Insights on a Rapidly Changing Epidemic. JAMA 2020, 323, 1339–1340. [Google Scholar] [CrossRef]
- Gao, Y.; Yan, L.; Huang, Y.; Liu, F.; Zhao, Y.; Cao, L.; Wang, T.; Sun, Q.; Ming, Z.; Zhang, L.; et al. Structure of the RNA-dependent RNA polymerase from COVID-19 virus. Science 2020, 368, 779–782. [Google Scholar] [CrossRef]
- Chan, J.F.; Yuan, S.; Kok, K.H.; To, K.K.; Chu, H.; Yang, J.; Xing, F.; Liu, J.; Yip, C.C.; Poon, R.W.; et al. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: A study of a family cluster. Lancet 2020, 395, 514–523. [Google Scholar] [CrossRef]
- Walls, A.C.; Park, Y.J.; Tortorici, M.A.; Wall, A.; McGuire, A.T.; Veesler, D. Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell 2020, 181, 281–292.e286. [Google Scholar] [CrossRef]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280.e278. [Google Scholar] [CrossRef]
- Dömling, A.; Gao, L. Chemistry and Biology of SARS-CoV-2. Chem 2020, 6, 1283–1295. [Google Scholar] [CrossRef]
- Báez-Santos, Y.M.; St John, S.E.; Mesecar, A.D. The SARS-coronavirus papain-like protease: Structure, function and inhibition by designed antiviral compounds. Antivir. Res. 2015, 115, 21–38. [Google Scholar] [CrossRef]
- Bouhaddou, M.; Memon, D.; Meyer, B.; White, K.M.; Rezelj, V.V.; Correa Marrero, M.; Polacco, B.J.; Melnyk, J.E.; Ulferts, S.; Kaake, R.M.; et al. The Global Phosphorylation Landscape of SARS-CoV-2 Infection. Cell 2020, 182, 685–712. [Google Scholar] [CrossRef]
- Jan, J.T.; Cheng, T.R.; Juang, Y.P.; Ma, H.H.; Wu, Y.T.; Yang, W.B.; Cheng, C.W.; Chen, X.; Chou, T.H.; Shie, J.J.; et al. Identification of existing pharmaceuticals and herbal medicines as inhibitors of SARS-CoV-2 infection. Proc. Natl. Acad. Sci. USA 2021, 118, e2021579118. [Google Scholar] [CrossRef]
- Jackson, L.A.; Anderson, E.J.; Rouphael, N.G.; Roberts, P.C.; Makhene, M.; Coler, R.N.; McCullough, M.P.; Chappell, J.D.; Denison, M.R.; Stevens, L.J.; et al. An mRNA Vaccine against SARS-CoV-2-Preliminary Report. N. Engl. J. Med. 2020, 383, 1920–1931. [Google Scholar] [CrossRef]
- Iddir, M.; Brito, A.; Dingeo, G.; Fernandez Del Campo, S.S.; Samouda, H.; La Frano, M.R.; Bohn, T. Strengthening the Immune System and Reducing Inflammation and Oxidative Stress through Diet and Nutrition: Considerations during the COVID-19 Crisis. Nutrients 2020, 12, 1562. [Google Scholar] [CrossRef]
- Manthey, J.A.; Grohmann, K. Flavonoids of the orange subfamily Aurantioideae. Adv. Exp. Med. Biol. 1998, 439, 85–101. [Google Scholar] [CrossRef]
- González-Barrio, R.; Trindade, L.M.; Manzanares, P.; de Graaff, L.H.; Tomás-Barberán, F.A.; Espín, J.C. Production of bioavailable flavonoid glucosides in fruit juices and green tea by use of fungal alpha-L-rhamnosidases. J. Agric. Food Chem. 2004, 52, 6136–6142. [Google Scholar] [CrossRef]
- Jin, M.J.; Kim, U.; Kim, I.S.; Kim, Y.; Kim, D.H.; Han, S.B.; Kim, D.H.; Kwon, O.S.; Yoo, H.H. Effects of gut microflora on pharmacokinetics of hesperidin: A study on non-antibiotic and pseudo-germ-free rats. J. Toxicol. Environ. Health A 2010, 73, 1441–1450. [Google Scholar] [CrossRef]
- Li, C.; Schluesener, H. Health-promoting effects of the citrus flavanone hesperidin. Crit. Rev. Food Sci. Nutr. 2017, 57, 613–631. [Google Scholar] [CrossRef]
- Ding, Z.; Sun, G.; Zhu, Z. Hesperidin attenuates influenza A virus (H1N1) induced lung injury in rats through its anti-inflammatory effect. Antivir. Ther. 2018, 23, 611–615. [Google Scholar] [CrossRef]
- Kandeil, A.; Mostafa, A.; Kutkat, O.; Moatasim, Y.; Al-Karmalawy, A.A.; Rashad, A.A.; Kayed, A.E.; Kayed, A.E.; El-Shesheny, R.; Kayali, G.; et al. Bioactive Polyphenolic Compounds Showing Strong Antiviral Activities against Severe Acute Respiratory Syndrome Coronavirus 2. Pathogens 2021, 10, 758. [Google Scholar] [CrossRef]
- Bellavite, P.; Donzelli, A. Hesperidin and SARS-CoV-2: New Light on the Healthy Function of Citrus Fruits. Antioxidants 2020, 9, 742. [Google Scholar] [CrossRef]
- Haggag, Y.A.; El-Ashmawy, N.E.; Okasha, K.M. Is hesperidin essential for prophylaxis and treatment of COVID-19 Infection? Med. Hypotheses 2020, 144, 109957. [Google Scholar] [CrossRef]
- Wang, Y.C.; Yang, W.H.; Yang, C.S.; Hou, M.H.; Tsai, C.L.; Chou, Y.Z.; Hung, M.C.; Chen, Y. Structural basis of SARS-CoV-2 main protease inhibition by a broad-spectrum anti-coronaviral drug. Am. J. Cancer Res. 2020, 10, 2535–2545. [Google Scholar]
- Alreshidi, F.S.; Ginawi, I.A.; Hussain, M.A.; Arif, J.M. Piperaquine- and Aspirin- Mediated Protective Role of HSP70 and HSP90 as Modes to Strengthen the Natural Immunity against Potent SARS-CoV-2. Biointerface Res. Appl. Chem. 2021, 11, 12364–12379. [Google Scholar] [CrossRef]
- Yumnam, S.; Venkatarame Gowda Saralamma, V.; Raha, S.; Lee, H.J.; Lee, W.S.; Kim, E.K.; Lee, S.J.; Heo, J.D.; Kim, G.S. Proteomic profiling of human HepG2 cells treated with hesperidin using antibody array. Mol. Med. Rep. 2017, 16, 5386–5392. [Google Scholar] [CrossRef]
- Zhang, L.; Jackson, C.B.; Mou, H.; Ojha, A.; Peng, H.; Quinlan, B.D.; Rangarajan, E.S.; Pan, A.; Vanderheiden, A.; Suthar, M.S.; et al. SARS-CoV-2 spike-protein D614G mutation increases virion spike density and infectivity. Nat. Commun. 2020, 11, 6013. [Google Scholar] [CrossRef]
- Tegally, H.; Wilkinson, E.; Giovanetti, M.; Iranzadeh, A.; Fonseca, V.; Giandhari, J.; Doolabh, D.; Pillay, S.; San, E.J.; Msomi, N.; et al. Emergence of a SARS-CoV-2 variant of concern with mutations in spike glycoprotein. Nature 2021, 592, 438–443. [Google Scholar] [CrossRef]
- Mascola, J.R.; Graham, B.S.; Fauci, A.S. SARS-CoV-2 Viral Variants-Tackling a Moving Target. JAMA 2021, 325, 1261–1262. [Google Scholar] [CrossRef] [PubMed]
- Baric, R.S. Emergence of a Highly Fit SARS-CoV-2 Variant. N. Engl. J. Med. 2020, 383, 2684–2686. [Google Scholar] [CrossRef]
- Korber, B.; Fischer, W.M.; Gnanakaran, S.; Yoon, H.; Theiler, J.; Abfalterer, W.; Hengartner, N.; Giorgi, E.E.; Bhattacharya, T.; Foley, B.; et al. Tracking Changes in SARS-CoV-2 Spike: Evidence that D614G Increases Infectivity of the COVID-19 Virus. Cell 2020, 182, 812–827.e819. [Google Scholar] [CrossRef]
- Abdool Karim, S.S.; de Oliveira, T. New SARS-CoV-2 Variants-Clinical, Public Health, and Vaccine Implications. N. Engl. J. Med. 2021, 384, 1866–1868. [Google Scholar] [CrossRef]
- Dong, W.; Wei, X.; Zhang, F.; Hao, J.; Huang, F.; Zhang, C.; Liang, W. A dual character of flavonoids in influenza A virus replication and spread through modulating cell-autonomous immunity by MAPK signaling pathways. Sci. Rep. 2014, 4, 7237. [Google Scholar] [CrossRef]
- Pappalardo, F.; Fichera, E.; Paparone, N.; Lombardo, A.; Pennisi, M.; Russo, G.; Leotta, M.; Pappalardo, F.; Pedretti, A.; De Fiore, F.; et al. A computational model to predict the immune system activation by citrus-derived vaccine adjuvants. Bioinformatics 2016, 32, 2672–2680. [Google Scholar] [CrossRef][Green Version]
- Hu, B.; Guo, H.; Zhou, P.; Shi, Z.L. Characteristics of SARS-CoV-2 and COVID-19. Nat. Rev. Microbiol. 2021, 19, 141–154. [Google Scholar] [CrossRef] [PubMed]
- Geleris, J.; Sun, Y.; Platt, J.; Zucker, J.; Baldwin, M.; Hripcsak, G.; Labella, A.; Manson, D.K.; Kubin, C.; Barr, R.G.; et al. Observational Study of Hydroxychloroquine in Hospitalized Patients with Covid-19. N. Engl. J. Med. 2020, 382, 2411–2418. [Google Scholar] [CrossRef] [PubMed]
- Hifumi, T.; Isokawa, S.; Otani, N.; Ishimatsu, S. Adverse events associated with nafamostat mesylate and favipiravir treatment in COVID-19 patients. Crit. Care 2020, 24, 497. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Kandhare, A.D.; Mukherjee, A.A.; Bodhankar, S.L. Acute and sub-chronic oral toxicity studies of hesperidin isolated from orange peel extract in Sprague Dawley rats. Regul. Toxicol. Pharmacol. 2019, 105, 77–85. [Google Scholar] [CrossRef]
- Jang, Y.; Kim, E.K.; Shim, W.S. Phytotherapeutic effects of the fruits of Poncirus trifoliata (L.) Raf. on cancer, inflammation, and digestive dysfunction. Phytother. Res. 2018, 32, 616–624. [Google Scholar] [CrossRef]
- Heo, Y.; Cho, Y.; Ju, K.S.; Cho, H.; Park, K.H.; Choi, H.; Yoon, J.K.; Moon, C.; Kim, Y.B. Antiviral activity of Poncirus trifoliata seed extract against oseltamivir-resistant influenza virus. J. Microbiol. 2018, 56, 586–592. [Google Scholar] [CrossRef] [PubMed]
- Hussain, S.; Curk, F.; Dhuique-Mayer, C.; Urban, L.; Ollitrault, P.; Luro, F.; Morillon, R. Autotetraploid trifoliate orange (Poncirus trifoliata) rootstocks do not impact clementine quality but reduce fruit yields and highly modify rootstock/scion physiology. Sci. Hortic. 2012, 134, 100–107. [Google Scholar] [CrossRef]
- Tundis, R.; Bonesi, M.; Sicari, V.; Pellicanò, T.M.; Tenuta, M.C.; Leporini, M.; Menichini, F.; Loizzo, M.R. Poncirus trifoliata (L.) Raf.: Chemical composition, antioxidant properties and hypoglycaemic activity via the inhibition of α-amylase and α-glucosidase enzymes. J. Funct. Foods 2016, 25, 477–485. [Google Scholar] [CrossRef]
- Chen, Y.C. Beware of docking! Trends Pharmacol. Sci. 2015, 36, 78–95. [Google Scholar] [CrossRef]
- Walsh, E.E.; Frenck, R.W., Jr.; Falsey, A.R.; Kitchin, N.; Absalon, J.; Gurtman, A.; Lockhart, S.; Neuzil, K.; Mulligan, M.J.; Bailey, R.; et al. Safety and Immunogenicity of Two RNA-Based Covid-19 Vaccine Candidates. N. Engl. J. Med. 2020, 383, 2439–2450. [Google Scholar] [CrossRef] [PubMed]
- Baden, L.R.; El Sahly, H.M.; Essink, B.; Kotloff, K.; Frey, S.; Novak, R.; Diemert, D.; Spector, S.A.; Rouphael, N.; Creech, C.B.; et al. Efficacy and Safety of the mRNA-1273 SARS-CoV-2 Vaccine. N. Engl. J. Med. 2021, 384, 403–416. [Google Scholar] [CrossRef] [PubMed]
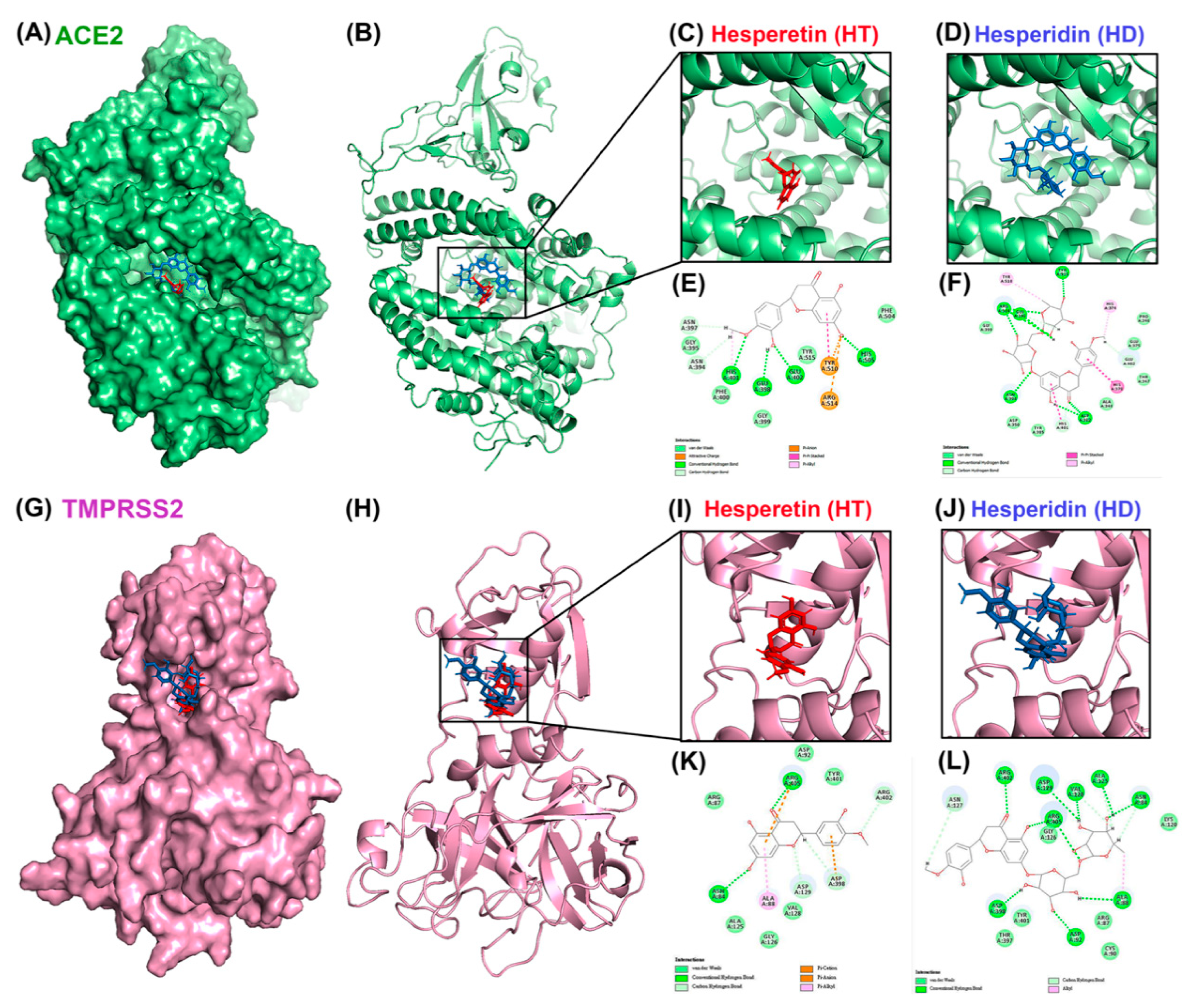
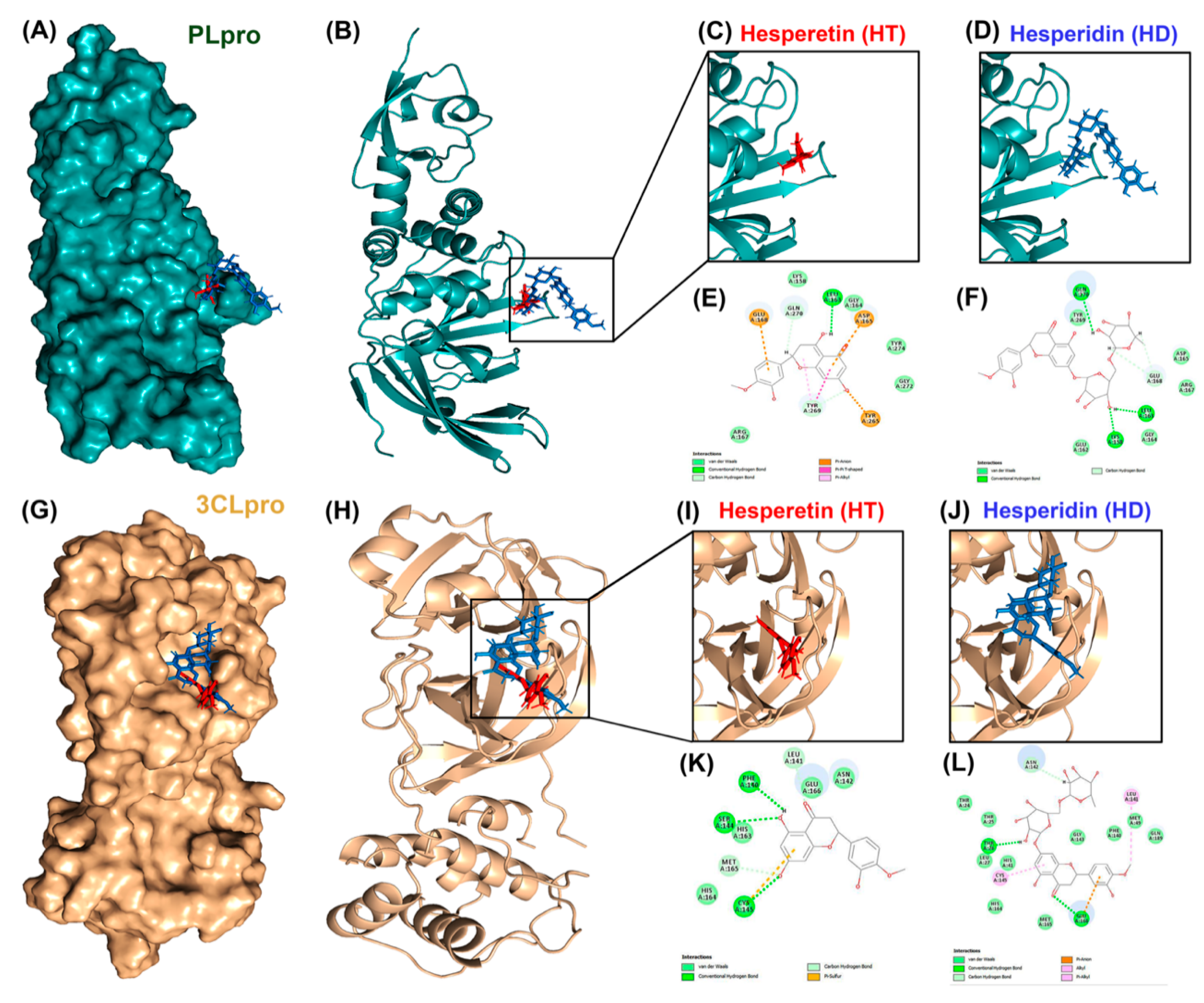
| c-Docker Energy Value (kcal/mol) | Residues | Distance (Å) | Types | ||
|---|---|---|---|---|---|
| ACE2 | HT | −34.81 | Ala348 | 1.9 | H-bond (H34, O) |
| Asp382 | 2.76 | CH-bond (H31, OD1) | |||
| Glu398 | 2.42 | H-bond (H36, O) | |||
| 2.44 | CH-bond (HA, O21) | ||||
| Glu402 | 2.23 | H-bond (HN, O20) | |||
| HD | −1.65 | Asp206 | 2.97/2.97 | CH-bond (H52, OD2)/(H54, OD2) | |
| Thr347 | 2.74 | CH-bond (HA, O14) | |||
| Ala348 | 2.66 | H-bond (H73, O) | |||
| Glu375 | 2.02/2.17 | H-bond (H74, OE1)/(H75, OE1) | |||
| Asp382 | 2.09 | H-bond (HD2, O9) | |||
| 2.59 | CH-bond (H57, OD1) | ||||
| Glu398 | 2.57 | CH-bond (H52, OE1) | |||
| His401 | 2.76 | p-s | |||
| Glu402 | 2.69 | CH-bond (H65, OE1) | |||
| Arg514 | 4.78 | p-cation | |||
| TMPRSS2 | HT | −30.56 | Lys254 | 3.92 | p-cation |
| Gly378 | 2.15 | H-bond (H67, O) | |||
| HD | −7.2 | His203 | 4.81 | p-alkyl | |
| Lys254 | 1.94 | H-bond (HZ1, O15) | |||
| 1.75 | H-bond (HZ3, O14) | ||||
| 4.08 | p-cation | ||||
| 2.65 | CH-bond (HE1, O6) | ||||
| Glu301 | 2.68/2.74 | H-bond | |||
| (H72, O)/(H71, OE2) | |||||
| 2.47 | CH-bond (H56, OE2) | ||||
| PLpro | HT | −13.69 | Asp165 | 2.98 | CH-bond (H31, OD2) |
| Pro248 | 4.97 | p-alkyl | |||
| HD | 17 | Leu163 | 2.28 | H-bond (H77, O) | |
| Gly164 | 5.21 | Amide-π stracked | |||
| Asp165 | 2.96 | p-anion | |||
| Glu168 | 2.94 | H-bond (H71, OE2) | |||
| Pro249 | 2.64 | CH-bond (HD2, O6) | |||
| Tyr265 | 5.47 | p-p T shaped | |||
| Gln270 | 2.44 | CH-bond (H60, OE1) | |||
| 2.31 | Unfavorable donor-donor | ||||
| (HE22, H75) | |||||
| 3CLpro | HT | −17.94 | Phe140 | 1.94 | H-bond (H34, O) |
| Asn142 | 2.96 | H-bond (HD21, O18) | |||
| Glu166 | 2.75 | H-bond (H34, OE1) | |||
| 3.07 | p-cation | ||||
| Gln189 | 2.19 | H-bond (HE22, O21) | |||
| HD | 6.21 | His41 | 2.34 | H-bond (HE2, O15) | |
| 4.6 | p-alkyl | ||||
| Asn142 | 2.63 | CH-bond (H46, OD1) | |||
| 2.81/2.85 | H-bond (HN, O2)/(HD21, O3) | ||||
| Gly143 | 2.8 | H-bond (HN, O14) | |||
| Cys145 | 2.27 | H-bond (HG, O15) | |||
| Met165 | 5.17 | Alkyl | |||
| Glu166 | 2.47/2.69 | CH-bond (H61, O)/(H59, O) | |||
| 1.95 | H-bond (H73, O) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cheng, F.-J.; Huynh, T.-K.; Yang, C.-S.; Hu, D.-W.; Shen, Y.-C.; Tu, C.-Y.; Wu, Y.-C.; Tang, C.-H.; Huang, W.-C.; Chen, Y.; et al. Hesperidin Is a Potential Inhibitor against SARS-CoV-2 Infection. Nutrients 2021, 13, 2800. https://doi.org/10.3390/nu13082800
Cheng F-J, Huynh T-K, Yang C-S, Hu D-W, Shen Y-C, Tu C-Y, Wu Y-C, Tang C-H, Huang W-C, Chen Y, et al. Hesperidin Is a Potential Inhibitor against SARS-CoV-2 Infection. Nutrients. 2021; 13(8):2800. https://doi.org/10.3390/nu13082800
Chicago/Turabian StyleCheng, Fang-Ju, Thanh-Kieu Huynh, Chia-Shin Yang, Dai-Wei Hu, Yi-Cheng Shen, Chih-Yen Tu, Yang-Chang Wu, Chih-Hsin Tang, Wei-Chien Huang, Yeh Chen, and et al. 2021. "Hesperidin Is a Potential Inhibitor against SARS-CoV-2 Infection" Nutrients 13, no. 8: 2800. https://doi.org/10.3390/nu13082800
APA StyleCheng, F.-J., Huynh, T.-K., Yang, C.-S., Hu, D.-W., Shen, Y.-C., Tu, C.-Y., Wu, Y.-C., Tang, C.-H., Huang, W.-C., Chen, Y., & Ho, C.-Y. (2021). Hesperidin Is a Potential Inhibitor against SARS-CoV-2 Infection. Nutrients, 13(8), 2800. https://doi.org/10.3390/nu13082800









