DNA Damage, n-3 Long-Chain PUFA Levels and Proteomic Profile in Brazilian Children and Adolescents
Abstract
1. Introduction
2. Materials and Methods
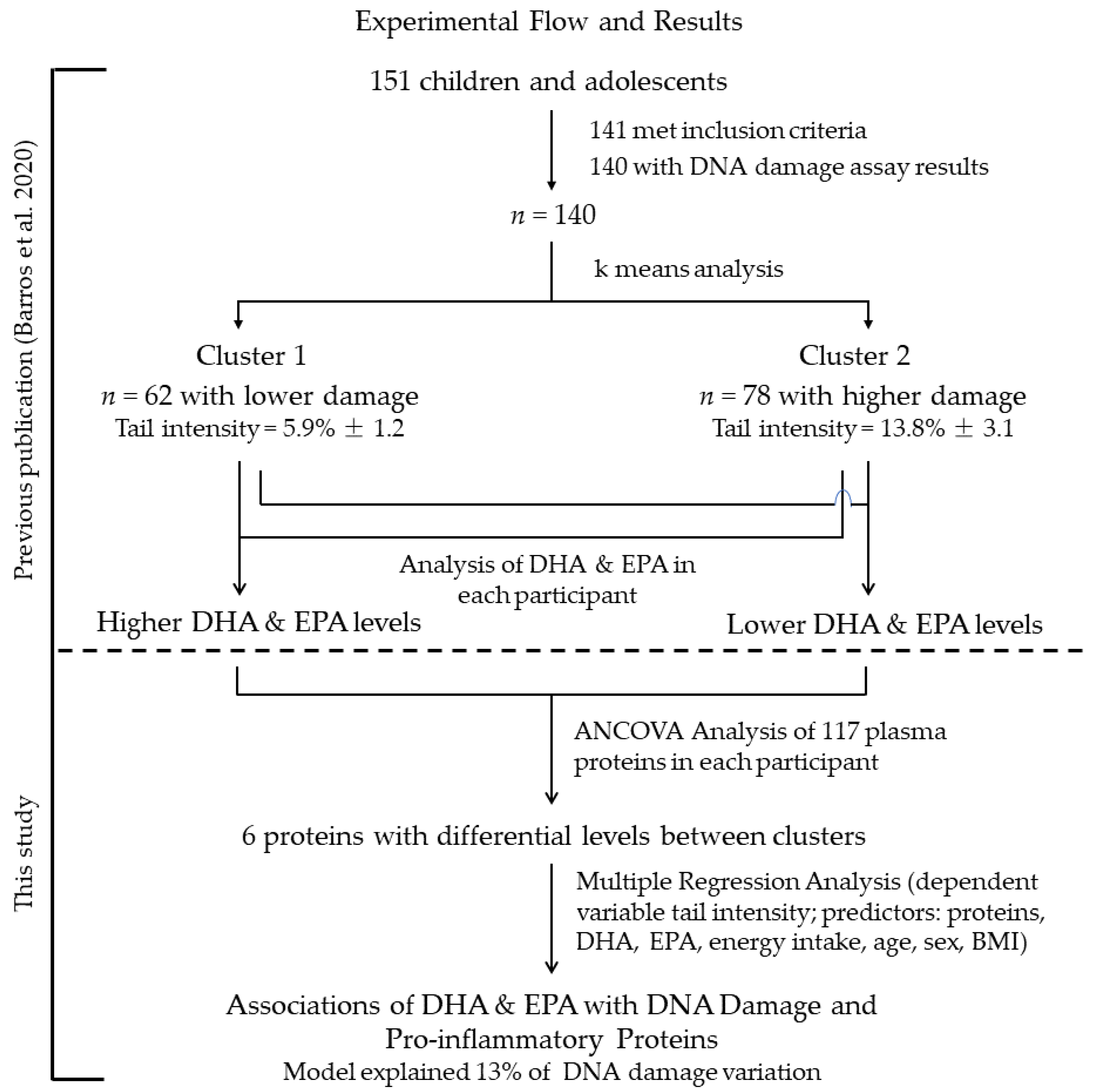
2.1. Study Design and Population
2.2. Fatty Acids Assessment
2.3. DNA Damage Assessment
2.4. Proteomic Analysis
2.5. Statistical Analyses
3. Results
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ramírez De Peña, D.; Martín, A.Á. Genómica nutricional como control de la enfermedad cardiovascular en el futuro próximo. Acta Bioquim. Clin. Lat. 2014, 48, 375–381. [Google Scholar]
- Pisabarro, R. Nutrigenética y nutrigenómica la revolución sanitaria del nuevo milenio. Implicancias clínicas en síndrome metabólico y diabetes tipo 2. Rev. Médica Urug. 2006, 22, 100–107. [Google Scholar]
- Corella, D.; Ordovás, J.M. Papel de las ómicas en la nutrición de precisión: Fortalezas y debilidades. Nutr. Hosp. 2018, 35, 10–18. [Google Scholar] [CrossRef]
- Kussmann, M.; Panchaud, A.; Affolter, M. Proteomics in nutrition: Status quo and outlook for biomarkers and bioactives. J. Proteome Res. 2010, 9, 4876–4887. [Google Scholar] [CrossRef]
- Slyskova, J.; Lorenzo, Y.; Karlsen, A.; Carlsen, M.H.; Novosadova, V.; Blomhoff, R.; Vodicka, P.; Collins, A.R. Both genetic and dietary factors underlie individual differences in DNA damage levels and DNA repair capacity. DNA Repair 2014, 16, 66–73. [Google Scholar] [CrossRef]
- Kryston, T.B.; Georgiev, A.B.; Pissis, P.; Georgakilas, A.G. Role of oxidative stress and DNA damage in human carcinogenesis. Mutat. Res. 2011, 711, 193–201. [Google Scholar] [CrossRef]
- Evans, M.D.; Dizdaroglu, M.; Cooke, M.S. Oxidative DNA damage and disease: Induction, repair and significance. Mutat. Res. 2004, 567, 1–61. [Google Scholar] [CrossRef]
- Demirbag, R.; Yilmaz, R.; Kocyigit, A. Relationship between DNA damage, total antioxidant capacity and coronary artery disease. Mutat. Res. 2005, 570, 197–203. [Google Scholar] [CrossRef]
- Fenech, M. Vitamins Associated with Brain Aging, Mild Cognitive Impairment, and Alzheimer Disease: Biomarkers, Epidemiological and Experimental Evidence, Plausible Mechanisms, and Knowledge Gaps. Adv. Nutr. 2017, 8, 958–970. [Google Scholar] [CrossRef]
- Huang, T.T.; Lampert, E.J.; Coots, C.; Lee, J.M. Targeting the PI3K pathway and DNA damage response as a therapeutic strategy in ovarian cancer. Cancer Treat. Rev. 2020, 86, 102021. [Google Scholar] [CrossRef]
- Rai, R.; Peng, G.; Li, K.; Lin, S.Y. DNA damage response: The players, the network and the role in tumor suppression. Cancer Genom. Proteom. 2007, 4, 99–106. [Google Scholar]
- Chou, J.; Quigley, D.A.; Robinson, T.M.; Feng, F.Y.; Ashworth, A. Transcription-associated cyclin-dependent kinases as targets and biomarkers for cancer therapy. Cancer Discov. 2020, 10, 351–370. [Google Scholar] [CrossRef]
- von Stechow, L.; Olsen, J.V. Proteomics insights into DNA damage response and translating this knowledge to clinical strategies. Proteomics 2017, 17, 1600018. [Google Scholar] [CrossRef]
- Ziegler, K.; Kunert, A.T.; Reinmuth-Selzle, K.; Leifke, A.L.; Widera, D.; Weller, M.G.; Schuppan, D.; Fröhlich-Nowoisky, J.; Lucas, K.; Pöschl, U. Chemical modification of pro-inflammatory proteins by peroxynitrite increases activation of TLR4 and NF-κB: Implications for the health effects of air pollution and oxidative stress. Redox Biol. 2020, 37, 101581. [Google Scholar] [CrossRef]
- Elia, A.E.H.; Boardman, A.P.; Wang, D.C.; Huttlin, E.L.; Everley, R.A.; Dephoure, N.; Zhou, C.; Koren, I.; Gygi, S.P.; Elledge, S.J. Quantitative Proteomic Atlas of Ubiquitination and Acetylation in the DNA Damage Response. Mol. Cell 2016, 59, 867–881. [Google Scholar] [CrossRef]
- Barros, T.T.; Venâncio, V.P.; Hernandes, L.C.; Antunes, L.M.G.; Hillesheim, E.; Salomão, R.G.; Mathias, M.G.; Coelho-Landell, C.A.; Toffano, R.B.D.; Almada, M.O.R.V.; et al. DNA damage is inversely associated to blood levels of DHA and EPA fatty acids in Brazilian children and adolescents. Food Funct. 2020, 11, 5115–5121. [Google Scholar] [CrossRef]
- Barden, A.E.; Mas, E.; Mori, T.A. n-3 Fatty acid supplementation and proresolving mediators of inflammation. Curr. Opin. Lipidol. 2016, 27, 26–32. [Google Scholar] [CrossRef]
- Murata, M. Inflammation and Cancer. Environ. Health Prev. Med. 2018, 23, 50. [Google Scholar] [CrossRef]
- Kawanishi, S.; Ohnishi, S.; Ma, N.; Hiraku, Y.; Murata, M. Crosstalk between DNA damage and inflammation in the multiple steps of carcinogenesis. Int. J. Mol. Sci. 2017, 18, 1808. [Google Scholar] [CrossRef]
- Marion-Letellier, R.; Savoye, G.; Ghosh, S. Polyunsaturated fatty acids and inflammation. IUBMB Life 2015, 67, 659–667. [Google Scholar] [CrossRef]
- Newell, M.; Baker, K.; Postovit, L.M.; Field, C.J. A critical review on the effect of docosahexaenoic acid (Dha) on cancer cell cycle progression. Int. J. Mol. Sci. 2017, 18, 1784. [Google Scholar] [CrossRef]
- Mathias, M.G.; Coelho-Landell, C.A.; Scott-Boyer, M.P.; Lacroix, S.; Morine, M.J.; Salomão, R.G.; Toffano, R.B.D.; Almada, M.O.R.V.; Camarneiro, J.M.; Hillesheim, E.; et al. Clinical and Vitamin Response to a Short-Term Multi-Micronutrient Intervention in Brazilian Children and Teens: From Population Data to Interindividual Responses. Mol. Nutr. Food Res. 2018, 62, 1700613. [Google Scholar] [CrossRef]
- Tanner, J.M. Growth at Adolescence, 2nd ed.; Blackwell: Oxford, UK, 1962. [Google Scholar]
- De Onis, M.; Onyango, A.W.; Borghi, E.; Siyam, A.; Nishida, C.; Siekmann, J. Development of a WHO growth reference for school-aged children and adolescents. Bull. World Health Organ. 2007, 85, 660–667. [Google Scholar] [CrossRef]
- Masood, A.; Stark, K.D.; Salem, N. A simplified and efficient method for the analysis of fatty acid methyl esters suitable for large clinical studies. J. Lipid Res. 2005, 46, 2299–2305. [Google Scholar] [CrossRef]
- Ued, F.V.; Mathias, M.G.; Toffano, R.B.D.; Barros, T.T.; Almada, M.O.R.V.; Salomão, R.G.; Coelho-Landell, C.A.; Hillesheim, E.; Camarneiro, J.M.; Camelo-Junior, J.S.; et al. Vitamin B2 and folate concentrations are associated with ARA, EPA and DHA fatty acids in red blood cells of Brazilian children and adolescents. Nutrients 2019, 11, 2918. [Google Scholar] [CrossRef]
- Singh, N.P.; McCoy, M.T.; Tice, R.R.; Schneider, E.L. A simple technique for quantitation of low levels of DNA damage in individual cells. Exp. Cell Res. 1988, 175, 184–191. [Google Scholar] [CrossRef]
- Tice, R.R.; Agurell, E.; Anderson, D.; Burlinson, B.; Hartmann, A.; Kobayashi, H.; Miyamae, Y.; Rojas, E.; Ryu, J.C.; Sasaki, Y.F. Single cell gel/comet assay: Guidelines for in vitro and in vivo genetic toxicology testing. Environ. Mol. Mutagen. 2000, 35, 206–221. [Google Scholar] [CrossRef]
- Kumaravel, T.S.; Vilhar, B.; Faux, S.P.; Jha, A.N. Comet Assay measurements: A perspective. Cell Biol. Toxicol. 2009, 25, 53–64. [Google Scholar] [CrossRef]
- Brody, E.N.; Gold, L.; Lawn, R.M.; Walker, J.J.; Zichi, D. High-content affinity-based proteomics: Unlocking protein biomarker discovery. Expert Rev. Mol. Diagn. 2010, 10, 1013–1022. [Google Scholar] [CrossRef]
- Gold, L.; Ayers, D.; Bertino, J.; Bock, C.; Bock, A.; Brody, E.N.; Carter, J.; Dalby, A.B.; Eaton, B.E.; Fitzwater, T.; et al. Aptamer-based multiplexed proteomic technology for biomarker discovery. PLoS ONE 2010, 5. [Google Scholar] [CrossRef]
- Fant, C.B.; Taatjes, D.J. Regulatory functions of the Mediator kinases CDK8 and CDK19. Transcription 2019, 10, 76–90. [Google Scholar] [CrossRef]
- Galbraith, M.D.; Donner, A.J.; Espinosa, J.M. CDK8: A positive regulator of transcription. Transcription 2010, 1, 4–12. [Google Scholar] [CrossRef]
- Bregman, D.B.; Pestell, R.G.; Kidd, V.J. Cell cycle regulation and RNA polymerase II. Front. Biosci. 2000, 5, D244–D257. [Google Scholar] [CrossRef] [PubMed]
- Liang, J.; Chen, M.; Hughes, D.; Chumanevich, A.A.; Altilia, S.; Kaza, V.; Lim, C.-U.; Kiaris, H.; Mythreye, K.; Pena, M.M.; et al. CDK8 selectively promotes the growth of colon cancer metastases in the liver by regulating gene expression of TIMP3 and matrix metalloproteinases. Cancer Res. 2018, 78, 6594–6606. [Google Scholar] [CrossRef] [PubMed]
- Broude, E.V.; Győrffy, B.; Chumanevich, A.A.; Chen, M.; McDermott, M.S.J.; Shtutman, M.; Catroppo, J.F.; Roninson, I.B. Expression of CDK8 and CDK8-interacting Genes as Potential Biomarkers in Breast Cancer. Curr. Cancer Drug Targets 2015, 15, 739–749. [Google Scholar] [CrossRef] [PubMed]
- Dannappel, M.V.; Sooraj, D.; Loh, J.J.; Firestein, R. Molecular and in vivo Functions of the CDK8 and CDK19 Kinase Modules. Front. Cell Dev. Biol. 2019, 6, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Stieg, D.C.; Chang, K.T.; Cooper, K.F.; Strich, R. Cyclin C regulated oxidative stress responsive transcriptome in mus musculus embryonic fibroblasts. G3 Genes Genomes Genet. 2019, 9, 1901–1908. [Google Scholar] [CrossRef]
- Harden, J.L.; Lewis, S.M.; Lish, S.R.; Suárez-Fariñas, M.; Gareau, D.; Lentini, T.; Johnson-Huang, L.M.; Krueger, J.G.; Lowes, M.A. The Tryptophan Metabolism Enzyme, L-Kynureninase, is a Novel Inflammatory Factor in Psoriasis and other Inflammatory Diseases. J. Allergy Clin. Immunol. 2016, 137, 1830–1840. [Google Scholar] [CrossRef]
- Ci, C.; Wu, C.; Lyu, D.; Chang, X.; He, C.; Liu, W.; Chen, L.; Ding, W. Downregulation of kynureninase restrains cutaneous squamous cell carcinoma proliferation and represses the PI3K/AKT pathway. Clin. Exp. Dermatol. 2020, 45, 194–201. [Google Scholar] [CrossRef]
- Fruman, D.A.; Chiu, H.; Hopkins, B.D.; Bagrodia, S.; Cantley, L.C.; Abraham, R.T. The PI3K Pathway in Human Disease. Cell 2017, 170, 605–635. [Google Scholar] [CrossRef]
- Kawakami, T.; Kawakami, Y.; Kitaura, J. Protein kinase C beta (PKC beta): Normal functions and diseases. J. Biochem. 2002, 132, 677–682. [Google Scholar] [CrossRef]
- Patergnani, S.; Marchi, S.; Rimessi, A.; Bonora, M.; Giorgi, C.; Mehta, K.D.; Pinton, P. PRKCB/protein kinase C, beta and the mitochondrial axis as key regulators of autophagy. Autophagy 2013, 9, 1367–1385. [Google Scholar] [CrossRef] [PubMed]
- National Center for Biotechnology Information (NCBI). Available online: https://www.ncbi.nlm.nih.gov/ (accessed on 11 June 2021).
- Sas, K.; Szabó, E.; Vécsei, L. Mitochondria, oxidative stress and the kynurenine system, with a focus on ageing and neuroprotection. Molecules 2018, 23, 191. [Google Scholar] [CrossRef]
- Donner, A.J.; Ebmeier, C.C.; Taatjes, D.J.; Espinosa, J.M. CDK8 is a positive regulator of transcriptional elongation within the serum response network. Nat. Struct. Mol. Biol. 2010, 17, 194–201. [Google Scholar] [CrossRef]
- Espinosa, J.M.; Verdun, R.E.; Emerson, B.M. p53 Functions through Stress- and Promoter- Specific Recruitment of Transcription Initiation Components before and after DNA Damage. Mol. Cell 2003, 12, 1015–1027. [Google Scholar] [CrossRef]
- Cavalcanti Júnior, G.B.; Klumb2, C.E.; Maia, R.C. p53 in hematological malignancies. Rev. Bras. Cancerol. 2002, 48, 419–427. [Google Scholar] [CrossRef]
- Poss, Z.C.; Ebmeier, C.C.; Odell, A.T.; Tangpeerachaikul, A.; Lee, T.; Pelish, H.E.; Shair, M.D.; Dowell, R.D.; Old, W.M.; Taatjes, D.J. Identification of Mediator kinase substrates in human cells using cortistatin A and quantitative phosphoproteomics. Cell Rep. 2016, 15, 436–450. [Google Scholar] [CrossRef] [PubMed]
- Porter, D.C.; Farmaki, E.; Altilia, S.; Schools, G.P.; West, D.K.; Chen, M.; Chang, B.D.; Puzyrev, A.T.; Lim, C.U.; Rokow-Kittell, R.; et al. Cyclin-dependent kinase 8 mediates chemotherapy-induced tumor-promoting paracrine activities. PNAS 2012, 109, 13799–13804. [Google Scholar] [CrossRef]
- Stieg, D.C.; Cooper, K.F.; Strich, R. The extent of cyclin C promoter occupancy directs changes in stress-dependent transcription. J. Biol. Chem. 2020, 295, 16280–16291. [Google Scholar] [CrossRef]
- Zhao, X.; Feng, D.; Wang, Q.; Abdulla, A.; Xie, X.; Zhou, J.; Sun, Y.; Yang, E.S.; Liu, L.; Vaitheesvaran, B.; et al. Regulation of lipogenesis by cyclin-dependent kinase 8-mediated control of SREBP-1. J. Clin. Invest. 2012, 122, 2417–2427. [Google Scholar] [CrossRef]
- Yang, J.; Nie, J.; Ma, X.; Wei, Y.; Peng, Y.; Wei, X. Targeting PI3K in cancer: Mechanisms and advances in clinical trials. Mol. Cancer 2019, 18, 26. [Google Scholar] [CrossRef]
- Donato, J.; Frazão, R.; Elias, C.F. The PI3K signaling pathway mediates the biological effects of leptin. Arq. Bras. Endocrinol. Metabol. 2010, 54, 591–602. [Google Scholar] [CrossRef]
- Kma, L.; Baruah, T.J. The interplay of ROS and the PI3K/Akt pathway in autophagy regulation. Biotechnol. Appl. Biochem. 2021, 1–17. [Google Scholar] [CrossRef]
- Koundouros, N.; Poulogiannis, G. Phosphoinositide 3-Kinase/Akt signaling and redox metabolism in cancer. Front. Oncol. 2018, 8, 2–10. [Google Scholar] [CrossRef]
- Mathew, R.; Karp, C.M.; Beaudoin, B.; Vuong, N.; Chen, G.; Chen, H.Y.; Bray, K.; Reddy, A.; Bhanot, G.; Gelinas, C.; et al. Autophagy Suppresses Tumorigenesis through Elimination of p62. Cell 2009, 137, 1062–1075. [Google Scholar] [CrossRef]
- Chandra Pal, H.; Athar, M.; Elmets, C.A.; Afaq, F. Fisetin inhibits UVB-induced cutaneous inflammation and activation of PI3K/AKT/NFκB signaling pathways in SKH-1 hairless mice. Photochem. Photobiol. 2015, 91, 225–234. [Google Scholar] [CrossRef]
- Nguyen, L.N.; Nguyen, L.N.T.; Zhao, J.; Schank, M.; Dang, X.; Cao, D.; Khanal, S.; Thakuri, B.K.C.; Zhang, J.; Lu, Z.; et al. Immune activation induces telomeric DNA damage and promotes short-lived effector T cell differentiation in chronic HCV infection. Hepatology 2021. [Google Scholar] [CrossRef]
- Denys, A.; Hichami, A.; Khan, N.A. n-3 PUFAs modulate T-cell activation via protein kinase C-α and -ε and the NF-κB signaling pathway. J. Lipid Res. 2005, 46, 752–758. [Google Scholar] [CrossRef]
- Simopoulos, A.P. Omega-3 fatty acids in inflammation and autoimmune diseases. J. Am. Coll. Nutr. 2002, 21, 495–505. [Google Scholar] [CrossRef]
- Kim, N.; Jeong, S.; Jing, K.; Shin, S.; Kim, S.; Heo, J.Y.; Kweon, G.R.; Park, S.K.; Wu, T.; Park, J., II; et al. Docosahexaenoic acid induces cell death in human non-small cell lung cancer cells by repressing mTOR via AMPK activation and PI3K/Akt inhibition. Biomed Res. Int. 2015, 2015, 239764. [Google Scholar] [CrossRef] [PubMed]
- Mándi, Y.; Vécsei, L. The kynurenine system and immunoregulation. J. Neural. Transm. 2012, 119, 197–209. [Google Scholar] [CrossRef]
- Kolodziej, L.R.; Paleolog, E.M.; Williams, R.O. Kynurenine metabolism in health and disease. Amino Acids 2011, 41, 1173–1183. [Google Scholar] [CrossRef] [PubMed]
- Braidy, N.; Grant, R. Kynurenine pathway metabolism and neuroinflammatory disease. Neural Regen. Res. 2017, 12, 39–42. [Google Scholar] [CrossRef]
- Braidy, N.; Berg, J.; Clement, J.; Khorshidi, F.; Poljak, A.; Jayasena, T.; Grant, R.; Sachdev, P. Role of Nicotinamide Adenine Dinucleotide and Related Precursors as Therapeutic Targets for Age-Related Degenerative Diseases: Rationale, Biochemistry, Pharmacokinetics, and Outcomes. Antioxid. Redox Signal. 2019, 30, 251–294. [Google Scholar] [CrossRef] [PubMed]
- Gomes, M.G.; Souza, L.C.; Goes, A.R.; Del Fabbro, L.; Filho, C.B.; Donato, F.; Prigol, M.; Luchese, C.; Roman, S.S.; Puntel, R.L.; et al. Fish oil ameliorates sickness behavior induced by lipopolysaccharide in aged mice through the modulation of kynurenine pathway. J. Nutr. Biochem. 2018, 58, 37–48. [Google Scholar] [CrossRef] [PubMed]
- Sheak, J.R.; Yan, S.; Weise-Cross, L.; Ahmadian, R.; Walker, B.R.; Jernigan, N.L.; Resta, T.C. PKCβ and reactive oxygen species mediate enhanced pulmonary vasoconstrictor reactivity following chronic hypoxia in neonatal rats. Am. J. Physiol. Heart Circ. Physiol. 2020, 318, H470–H483. [Google Scholar] [CrossRef]
- Sakata, K.; Kondo, T.; Mizuno, N.; Shoji, M.; Yasui, H.; Yamamori, T.; Inanami, O.; Yokoo, H.; Yoshimura, N.; Hattori, Y. Roles of ROS and PKC-βII in ionizing radiation-induced eNOS activation in human vascular endothelial cells. Vascul. Pharmacol. 2015, 70, 55–65. [Google Scholar] [CrossRef]
- Davidson, L.A.; Brown, R.E.; Chang, W.-C.L.; Morris, J.S.; Wang, N.; Carroll, R.J.; Turner, N.D.; Lupton, J.R.; Chapkin, R.S. Morphodensitometric analysis of protein kinase C β II expression in rat colon: Modulation by diet and relation to in situ cell proliferation and apoptosis. Carcinogenesis 2000, 21, 1513–1519. [Google Scholar] [CrossRef][Green Version]
- Kliemann, M.; Prá, D.; Müller, L.L.; Hermes, L.; Horta, J.A.; Reckziegel, M.B.; Burgos, M.S.; Maluf, S.W.; Franke, S.I.R.; da Silva, J. DNA damage in children and adolescents with cardiovascular disease risk factors. An. Acad. Bras. Cienc. 2012, 84, 833–840. [Google Scholar] [CrossRef]

| Characteristics | Cluster 1 (n = 62) | Cluster 2 (n = 78) | p-Value |
|---|---|---|---|
| Age (years old) | 11.4 ± 1.1 | 11.4 ± 1.1 | 0.989 3 |
| Sex (n; %) | 0.728 4 | ||
| Male | 26 (41.9) | 35 (44.9) | |
| Female | 36 (58.1) | 43 (55.1) | |
| Sexual Maturity Rate (n; %) 1 | 0.497 4 | ||
| 1 | 4 (6.4) | 10 (12.8) | |
| 2 | 29 (46.8) | 32 (41.0) | |
| 3 | 21 (33.9) | 27 (34.6) | |
| 4 | 5 (8.1) | 8 (10.3) | |
| 5 | 3 (4.8) | 1 (1.3) | |
| Nutritional status (n; %) 2 | 0.332 4 | ||
| Eutrophic | 29 (46.8) | 43 (55.1) | |
| Overweight | 19 (30.6) | 14 (17.9) | |
| Obese | 14 (22.6) | 21 (26.9) | |
| Energy Intake (kcal/day) | 2061.6 ± 664.5 | 2065.2 ± 848.6 | 0.978 3 |
| Tail intensity (%) | 5.9 ± 1.2 | 13.8 ± 3.1 | <0.001 |
| DHA (mg/dL) | 6.2 ± 1.6 | 5.4 ± 1.3 | 0.003 3 |
| EPA (mg/dL) | 0.6 ± 0.2 | 0.5 ± 0.1 | <0.001 3 |
| Protein (pM) | Cluster 1 (n = 62) | Cluster 2 (n = 78) | p-Value 1 |
|---|---|---|---|
| CDK8–CCNC | 2265.5 (1989.0; 2468.1) | 2399.5 (2093.7; 2708.4) | 0.008 |
| KYNU | 985.4 (837.7; 1150.2) | 1061.8 (906.0; 1207.8) | 0.028 |
| PIK3CA–PIK3R1 | 1332.7 (1132.4; 1523.0) | 1424.0 (1228.6; 1614.8) | 0.044 |
| PRKCB | 24,843.2 (20,265.4; 27,851.2) | 25,460.1 (21,616.7; 29,323.1) | 0.024 |
| Gene ID | Full Name | Function | Action |
|---|---|---|---|
| CDK8 1024 | Cyclindependent kinase 8 | This kinase and its regulatory subunit, cyclin C, are components of the mediator transcriptional regulatory complex involved in both transcriptional activation and repression. This kinase phosphorylates the carboxy-terminal domain of RNA polymerase II [32,33,34] | Pro-inflammatory [35,36] |
| CCNC 892 | Cyclin C | Cyclin C interacts with cyclin-dependent kinase 8 and induces the phosphorylation of RNA polymerase II. The level of mRNAs for the gene that encodes this protein reaches its peak in the G1 phase of the cell cycle [32,37] | Anti- and pro-inflammatory [38] |
| KYNU 8942 | Kynureninase | KYNU is a pyridoxal-5′-phosphate-dependent enzyme that catalyzes the cleavage of L-kynurenine and L-3-hydroxykynurenine. KYNU is involved in biosynthesis of NAD cofactors from tryptophan through kynurenine pathway [39]. | Pro-inflammatory [39,40] |
| PIK3CA 5290 | Phosphatidylinositol 3-kinase catalytic subunit alpha | PIK3CA is the catalytic subunit of phosphatidylinositol 3-kinase (PI3K). It uses ATP to phosphorylate phosphatidylinositol, phosphatidylinositol-4-phosphate, and phosphatidylinositol 4,5-bisphosphate. The gene that encodes this protein has been found to be oncogenic [41]. | Pro-inflammatory [10,41] |
| PIK3R1 5295 | Phosphatidylinositol 3-kinase regulatory subunit 1 | PIK3R1 is the regulatory subunit of phosphatidylinositol 3-kinase, which phosphorylates the inositol ring of phosphatidylinositol at the 3-prime position. Phosphatidylinositol 3-kinase plays an important role in metabolic actions of insulin [41]. | Pro-inflammatory [10,41] |
| PRKCB 5579 | Protein kinase C beta | PRKCB is a member of protein kinase C (PKC) family. PKC family members phosphorylate a wide variety of protein targets and are involved in diverse cellular signaling pathways. PRKCB has been involved in many different cellular functions, such as B cell activation, apoptosis, endothelial cell proliferation, and intestinal sugar absorption [42]. | Pro-inflammatory [43] |
| Protein (pM) | R | R Square | Adjusted R Square | Durbin–Watson | p-Value 1 |
|---|---|---|---|---|---|
| All proteins | 0.44 | 0.19 | 0.13 | 1.93 | 0.002 |
| CDK8–CCNC | 0.42 | 0.18 | 0.13 | 1.98 | 0.001 |
| KYNU | 0.42 | 0.18 | 0.13 | 1.94 | 0.001 |
| PIK3CA–PIK3R1 | 0.41 | 0.17 | 0.12 | 1.98 | 0.002 |
| PRKCB | 0.42 | 0.18 | 0.13 | 1.97 | 0.001 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Barros, T.T.d.; Venancio, V.d.P.; Hernandes, L.C.; Antunes, L.M.G.; Hillesheim, E.; Salomão, R.G.; Mathias, M.G.; Coelho-Landell, C.A.; Toffano, R.B.D.; Almada, M.O.R.d.V.; et al. DNA Damage, n-3 Long-Chain PUFA Levels and Proteomic Profile in Brazilian Children and Adolescents. Nutrients 2021, 13, 2483. https://doi.org/10.3390/nu13082483
Barros TTd, Venancio VdP, Hernandes LC, Antunes LMG, Hillesheim E, Salomão RG, Mathias MG, Coelho-Landell CA, Toffano RBD, Almada MORdV, et al. DNA Damage, n-3 Long-Chain PUFA Levels and Proteomic Profile in Brazilian Children and Adolescents. Nutrients. 2021; 13(8):2483. https://doi.org/10.3390/nu13082483
Chicago/Turabian StyleBarros, Tamiris Trevisan de, Vinicius de Paula Venancio, Lívia Cristina Hernandes, Lusania Maria Greggi Antunes, Elaine Hillesheim, Roberta Garcia Salomão, Mariana Giaretta Mathias, Carolina Almeida Coelho-Landell, Roseli Borges Donegá Toffano, Maria Olimpia Ribeiro do Vale Almada, and et al. 2021. "DNA Damage, n-3 Long-Chain PUFA Levels and Proteomic Profile in Brazilian Children and Adolescents" Nutrients 13, no. 8: 2483. https://doi.org/10.3390/nu13082483
APA StyleBarros, T. T. d., Venancio, V. d. P., Hernandes, L. C., Antunes, L. M. G., Hillesheim, E., Salomão, R. G., Mathias, M. G., Coelho-Landell, C. A., Toffano, R. B. D., Almada, M. O. R. d. V., Camelo-Junior, J. S., Moco, S., Cominetti, O., Ued, F. d. V., Kaput, J., & Monteiro, J. P. (2021). DNA Damage, n-3 Long-Chain PUFA Levels and Proteomic Profile in Brazilian Children and Adolescents. Nutrients, 13(8), 2483. https://doi.org/10.3390/nu13082483








