Diet–Microbiota Interactions in Inflammatory Bowel Disease
Abstract
1. Introduction
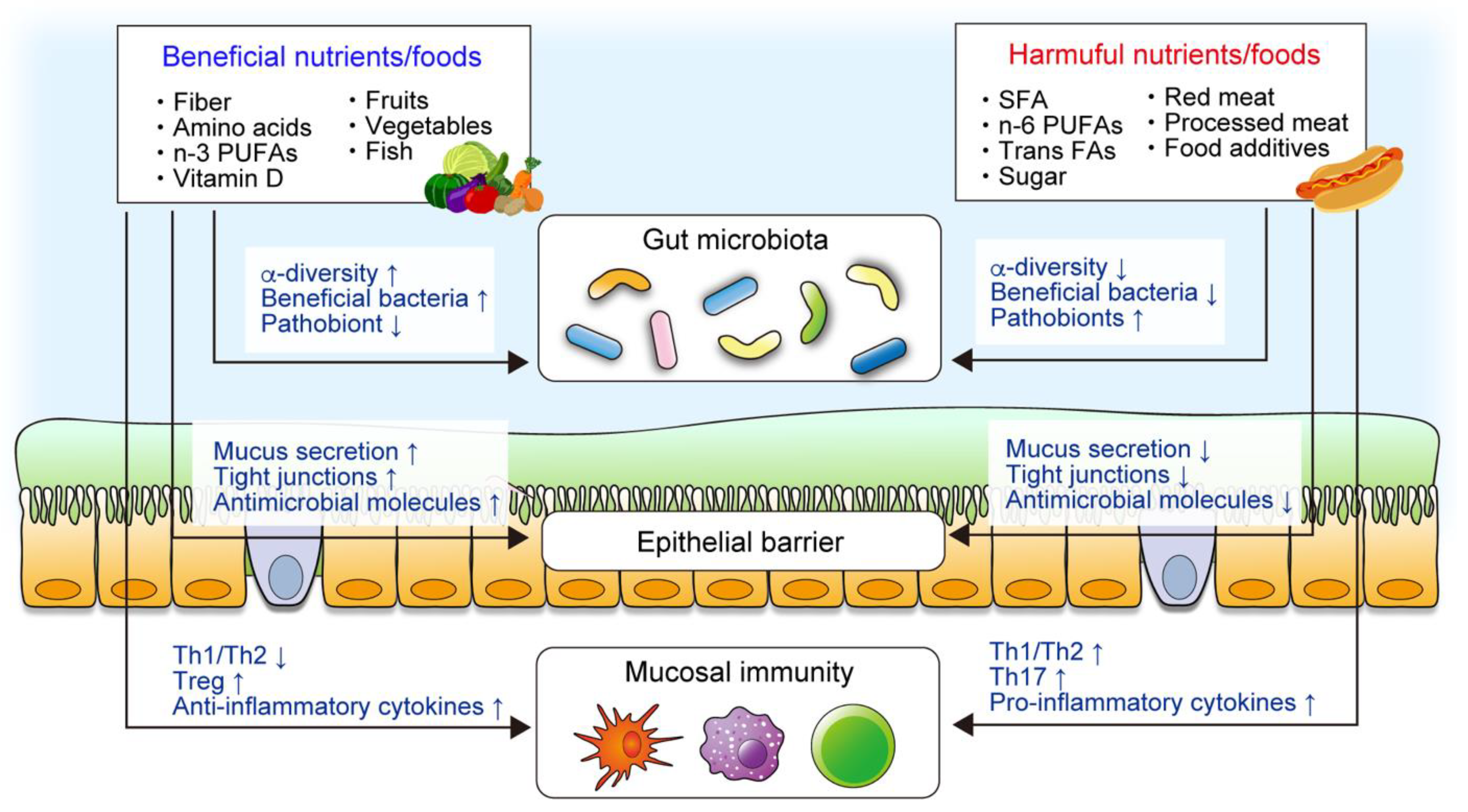
2. The Role of Diet in the Pathogenesis of IBD
2.1. Epidemiological Study
2.2. Animal Study
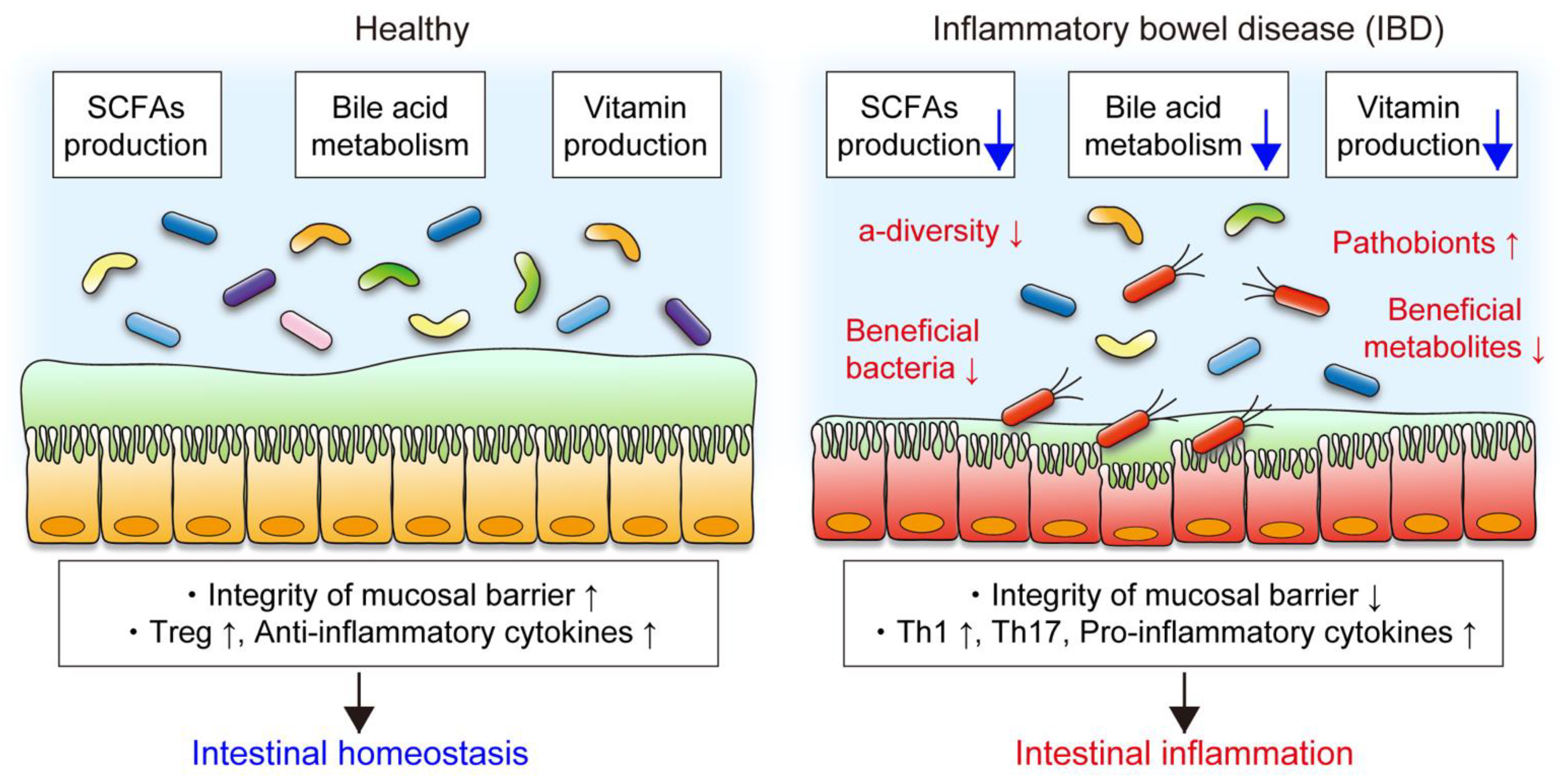
3. Gut Microbiota and IBD
3.1. The Role of the Gut Microbiota in IBD
3.2. Alteration of the Microbial Composition in IBD
3.3. Functional Changes in the Gut Microbiota in IBD
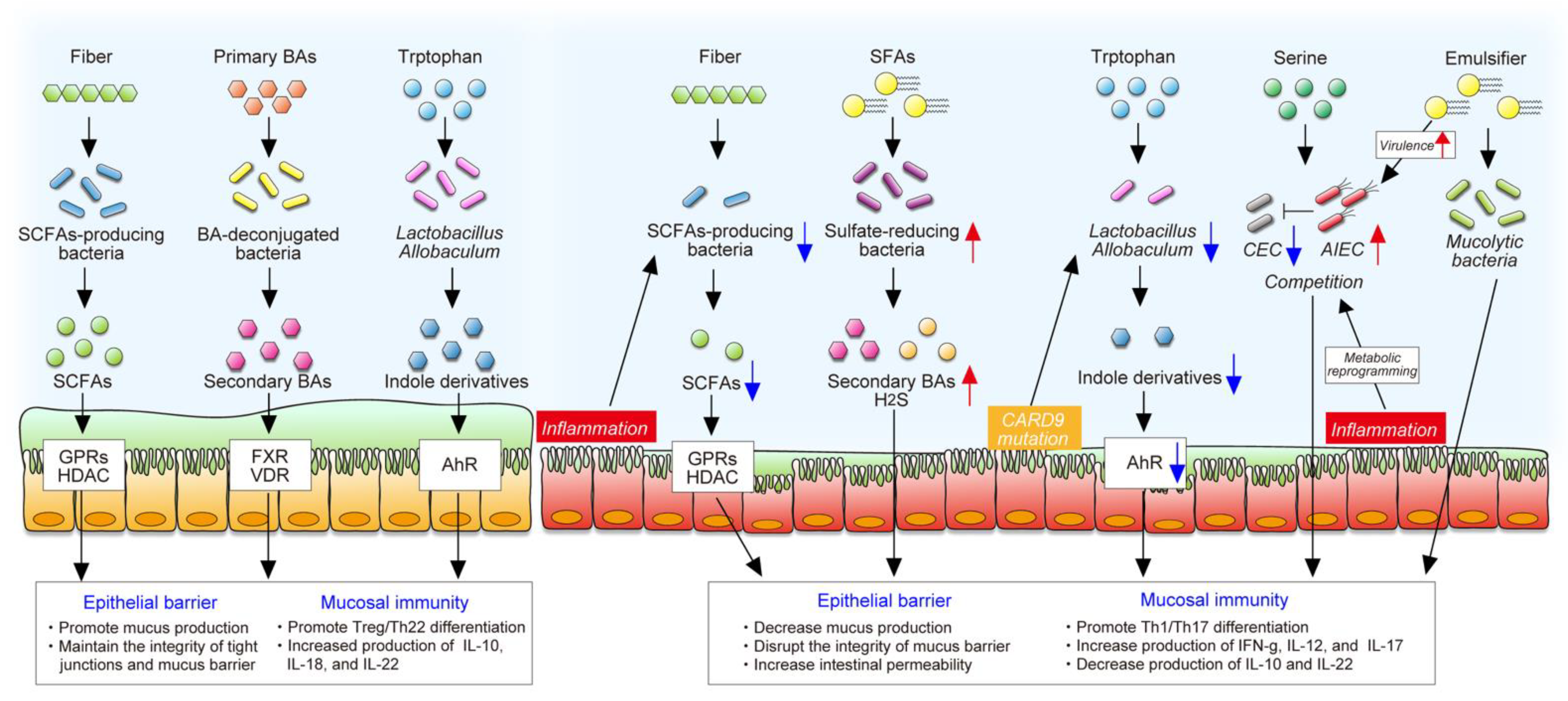
4. Diet–Microbe Interaction in IBD
4.1. Dietary Fibers and SCFAs
4.2. Dietary Fat and Bile Acids
4.3. Dietary Tryptophan and Indole Derivatives
4.4. Dietary L-Serine
4.5. Dietary Emulsifiers
5. The Impact of Nutritional Intervention on the Gut Microbiota in IBD
5.1. Enteral Nutrition
5.2. Dietary Intervention
6. Conclusions and Future Direction
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Ananthakrishnan, A.N. Epidemiology and risk factors for IBD. Nat. Rev. Gastroenterol. Hepatol. 2015, 12, 205–217. [Google Scholar] [CrossRef] [PubMed]
- Kaplan, G.G. The global burden of IBD: From 2015 to 2025. Nat. Rev. Gastroenterol. Hepatol. 2015, 12, 720–727. [Google Scholar] [CrossRef] [PubMed]
- Dahlhamer, J.M.; Zammitti, E.P.; Ward, B.W.; Wheaton, A.G.; Croft, J.B. Prevalence of Inflammatory Bowel Disease Among Adults Aged ≥ 18 Years—United States, 2015. MMWR Morb. Mortal. Wkly. Rep. 2016, 65, 1166–1169. [Google Scholar] [CrossRef] [PubMed]
- Ananthakrishnan, A.N.; Bernstein, C.N.; Iliopoulos, D.; MacPherson, A.; Neurath, M.F.; Ali, R.A.R.; Vavricka, S.R.; Fiocchi, C. Environmental triggers in IBD: A review of progress and evidence. Nat. Rev. Gastroenterol. Hepatol. 2018, 15, 39–49. [Google Scholar] [CrossRef] [PubMed]
- Okabayashi, S.; Kobayashi, T.; Hibi, T. Inflammatory Bowel Disease in Japan-Is It Similar to or Different from Westerns? J. Anus Rectum Colon 2020, 4, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Kanai, T.; Matsuoka, K.; Naganuma, M.; Hayashi, A.; Hisamatsu, T. Diet, microbiota, and inflammatory bowel disease: Lessons from Japanese foods. Korean J. Intern. Med. 2014, 29, 409–415. [Google Scholar] [CrossRef]
- Levine, A.; Boneh, R.S.; Wine, E. Evolving role of diet in the pathogenesis and treatment of inflammatory bowel diseases. Gut 2018, 67, 1726–1738. [Google Scholar] [CrossRef] [PubMed]
- Caruso, R.; Lo, B.C.; Núñez, G. Host–microbiota interactions in inflammatory bowel disease. Nat. Rev. Immunol. 2020, 20, 411–426. [Google Scholar] [CrossRef]
- Statovci, D.; Aguilera, M.; MacSharry, J.; Melgar, S. The Impact of Western Diet and Nutrients on the Microbiota and Immune Response at Mucosal Interfaces. Front. Immunol. 2017, 8, 838. [Google Scholar] [CrossRef]
- Kostic, A.D.; Xavier, R.J.; Gevers, D. The Microbiome in Inflammatory Bowel Disease: Current Status and the Future Ahead. Gastroenterology 2014, 146, 1489–1499. [Google Scholar] [CrossRef]
- Hou, J.K.; Abraham, B.; El-Serag, H. Dietary Intake and Risk of Developing Inflammatory Bowel Disease: A Systematic Review of the Literature. Am. J. Gastroenterol. 2011, 106, 563–573. [Google Scholar] [CrossRef] [PubMed]
- Chan, S.S.M.; Luben, R.; Olsen, A.; Tjonneland, A.; Kaaks, R.; Lindgren, S.; Grip, O.; Bergmann, M.M.; Boeing, H.; Hallmans, G.; et al. Association between high dietary intake of then−3 polyunsaturated fatty acid docosahexaenoic acid and reduced risk of Crohn’s disease. Aliment. Pharmacol. Ther. 2014, 39, 834–842. [Google Scholar] [CrossRef]
- Racine, A.; Carbonnel, F.; Chan, S.S.M.; Hart, A.R.; Bueno-De-Mesquita, H.B.; Oldenburg, B.; Van Schaik, F.D.M.; Tjønneland, A.; Olsen, A.; Dahm, C.C.; et al. Dietary Patterns and Risk of Inflammatory Bowel Disease in Europe. Inflamm. Bowel Dis. 2016, 22, 345–354. [Google Scholar] [CrossRef] [PubMed]
- Ananthakrishnan, A.N.; Khalili, H.; Konijeti, G.G.; Higuchi, L.M.; de Silva, P.; Korzenik, J.R.; Fuchs, C.S.; Willett, W.C.; Richter, J.M.; Chan, A.T. A Prospective Study of Long-term Intake of Dietary Fiber and Risk of Crohn’s Disease and Ulcerative Colitis. Gastroenterology 2013, 145, 970–977. [Google Scholar] [CrossRef] [PubMed]
- Ananthakrishnan, A.N.; Khalili, H.; Song, M.; Higuchi, L.M.; Richter, J.M.; Chan, A.T. Zinc intake and risk of Crohn’s disease and ulcerative colitis: A prospective cohort study. Int. J. Epidemiol. 2015, 44, 1995–2005. [Google Scholar] [CrossRef] [PubMed]
- Ananthakrishnan, A.N.; Khalili, H.; Song, M.; Higuchi, L.M.; Lochhead, P.; Richter, J.M.; Chan, A.T. Genetic Polymorphisms in Fatty Acid Metabolism Modify the Association Between Dietary n3. Inflamm. Bowel Dis. 2017, 23, 1898–1904. [Google Scholar] [CrossRef]
- Wirtz, S.; Popp, V.; Kindermann, M.; Gerlach, K.; Weigmann, B.; Fichtner-Feigl, S.; Neurath, M.F. Chemically induced mouse models of acute and chronic intestinal inflammation. Nat. Protoc. 2017, 12, 1295–1309. [Google Scholar] [CrossRef]
- Keubler, L.M.; Buettner, M.; Häger, C.; Bleich, A. A Multihit Model. Inflamm. Bowel Dis. 2015, 21, 1967–1975. [Google Scholar] [CrossRef]
- Ma, X.; Torbenson, M.; Hamad, A.R.A.; Soloski, M.J.; Li, Z. High-fat diet modulates non-CD1d-restricted natural killer T cells and regulatory T cells in mouse colon and exacerbates experimental colitis. Clin. Exp. Immunol. 2007, 151, 130–138. [Google Scholar] [CrossRef]
- Kim, K.-A.; Gu, W.; Lee, I.-A.; Joh, E.-H.; Kim, D.-H. High Fat Diet-Induced Gut Microbiota Exacerbates Inflammation and Obesity in Mice via the TLR4 Signaling Pathway. PLoS ONE 2012, 7, e47713. [Google Scholar] [CrossRef]
- Cheng, L.; Jin, H.; Qiang, Y.; Wu, S.; Yan, C.; Han, M.; Xiao, T.; Yan, N.; An, H.; Zhou, X.; et al. High fat diet exacerbates dextran sulfate sodium induced colitis through disturbing mucosal dendritic cell homeostasis. Int. Immunopharmacol. 2016, 40, 1–10. [Google Scholar] [CrossRef]
- Islam, J.; Sato, S.; Watanabe, K.; Watanabe, T.; Ardiansyah; Hirahara, K.; Aoyama, Y.; Tomita, S.; Aso, H.; Komai, M.; et al. Dietary tryptophan alleviates dextran sodium sulfate-induced colitis through aryl hydrocarbon receptor in mice. J. Nutr. Biochem. 2017, 42, 43–50. [Google Scholar] [CrossRef] [PubMed]
- Coburn, L.A.; Gong, X.; Singh, K.; Asim, M.; Scull, B.P.; Allaman, M.M.; Williams, C.S.; Rosen, M.J.; Washington, M.K.; Barry, D.P.; et al. L-arginine Supplementation Improves Responses to Injury and Inflammation in Dextran Sulfate Sodium Colitis. PLoS ONE 2012, 7, e33546. [Google Scholar] [CrossRef]
- Shiomi, Y.; Nishiumi, S.; Ooi, M.; Hatano, N.; Shinohara, M.; Yoshie, T.; Kondo, Y.; Furumatsu, K.; Shiomi, H.; Kutsumi, H.; et al. GCMS-based metabolomic study in mice with colitis induced by dextran sulfate sodium. Inflamm. Bowel Dis. 2011, 17, 2261–2274. [Google Scholar] [CrossRef]
- Tsune, I.; Ikejima, K.; Hirose, M.; Yoshikawa, M.; Enomoto, N.; Takei, Y.; Sato, N. Dietary glycine prevents chemical-induced experimental colitis in the rat. Gastroenterology 2003, 125, 775–785. [Google Scholar] [CrossRef]
- Andou, A.; Hisamatsu, T.; Okamoto, S.; Chinen, H.; Kamada, N.; Kobayashi, T.; Hashimoto, M.; Okutsu, T.; Shimbo, K.; Takeda, T.; et al. Dietary Histidine Ameliorates Murine Colitis by Inhibition of Proinflammatory Cytokine Production from Macrophages. Gastroenterology 2009, 136, 564–574.e2. [Google Scholar] [CrossRef] [PubMed]
- Khan, S.; Waliullah, S.; Godfrey, V.; Khan, A.W.; Ramachandran, R.A.; Cantarel, B.L.; Behrendt, C.; Peng, L.; Hooper, L.V.; Zaki, H. Dietary simple sugars alter microbial ecology in the gut and promote colitis in mice. Sci. Transl. Med. 2020, 12, eaay6218. [Google Scholar] [CrossRef]
- Levine, A.; Wine, E.; Assa, A.; Boneh, R.S.; Shaoul, R.; Kori, M.; Cohen, S.; Peleg, S.; Shamaly, H.; On, A.; et al. Crohn’s Disease Exclusion Diet Plus Partial Enteral Nutrition Induces Sustained Remission in a Randomized Controlled Trial. Gastroenterology 2019, 157, 440–450.e8. [Google Scholar] [CrossRef]
- Kanauchi, O.; Suga, T.; Tochihara, M.; Hibi, T.; Naganuma, M.; Homma, T.; Asakura, H.; Nakano, H.; Takahama, K.; Fujiyama, Y.; et al. Treatment of ulcerative colitis by feeding with germinated barley foodstuff: First report of a multicenter open control trial. J. Gastroenterol. 2002, 37, 67–72. [Google Scholar] [CrossRef]
- Casellas, F.; Borruel, N.; Torrejón, A.; Varela, E.; Antolin, M.; Guarner, F.; Malagelada, J.-R. Oral oligofructose-enriched inulin supplementation in acute ulcerative colitis is well tolerated and associated with lowered faecal calprotectin. Aliment. Pharmacol. Ther. 2007, 25, 1061–1067. [Google Scholar] [CrossRef]
- Benjamin, J.L.; Hedin, C.R.H.; Koutsoumpas, A.; Ng, S.C.; McCarthy, N.E.; Hart, A.L.; Kamm, M.A.; Sanderson, J.D.; Knight, S.C.; Forbes, A.; et al. Randomised, double-blind, placebo-controlled trial of fructo-oligosaccharides in active Crohn’s disease. Gut 2011, 60, 923–929. [Google Scholar] [CrossRef] [PubMed]
- Round, J.L.; Mazmanian, S.K. The gut microbiota shapes intestinal immune responses during health and disease. Nat. Rev. Immunol. 2009, 9, 313–323. [Google Scholar] [CrossRef]
- Johansson, M.E.; Jakobsson, H.E.; Holmén-Larsson, J.; Schütte, A.; Ermund, A.; Rodríguez-Piñeiro, A.M.; Arike, L.; Wising, C.; Svensson, F.; Bäckhed, F.; et al. Normalization of Host Intestinal Mucus Layers Requires Long-Term Microbial Colonization. Cell Host Microbe 2015, 18, 582–592. [Google Scholar] [CrossRef]
- Sellon, R.K.; Tonkonogy, S.; Schultz, M.; Dieleman, L.A.; Grenther, W.; Balish, E.; Rennick, D.M.; Sartor, R.B. Resident Enteric Bacteria Are Necessary for Development of Spontaneous Colitis and Immune System Activation in Interleukin-10-Deficient Mice. Infect. Immun. 1998, 66, 5224–5231. [Google Scholar] [CrossRef] [PubMed]
- Stepankova, R.; Powrie, F.; Kofronova, O.; Kozakova, H.; Hudcovic, T.; Hrncir, T.; Uhlig, H.; Read, S.; Rehakova, Z.; Benada, O.; et al. Segmented filamentous bacteria in a defined bacterial cocktail induce intestinal inflammation in SCID mice reconstituted with CD45RBhigh CD4+ T cells. Inflamm. Bowel Dis. 2007, 13, 1202–1211. [Google Scholar] [CrossRef]
- Britton, G.J.; Contijoch, E.J.; Mogno, I.; Vennaro, O.H.; Llewellyn, S.R.; Ng, R.; Li, Z.; Mortha, A.; Merad, M.; Das, A.; et al. Microbiotas from Humans with Inflammatory Bowel Disease Alter the Balance of Gut Th17 and RORγt+ Regulatory T Cells and Exacerbate Colitis in Mice. Immunity 2019, 50, 212–224.e4. [Google Scholar] [CrossRef]
- Nagao-Kitamoto, H.; Shreiner, A.B.; Gillilland, M.G.; Kitamoto, S.; Ishii, C.; Hirayama, A.; Kuffa, P.; El-Zaatari, M.; Grasberger, H.; Seekatz, A.M.; et al. Functional Characterization of Inflammatory Bowel Disease–Associated Gut Dysbiosis in Gnotobiotic Mice. Cell. Mol. Gastroenterol. Hepatol. 2016, 2, 468–481. [Google Scholar] [CrossRef] [PubMed]
- Imdad, A.; Nicholson, M.R.; Tanner-Smith, E.E.; Zackular, J.P.; Gomez-Duarte, O.G.; Beaulieu, D.B.; Acra, S. Fecal transplantation for treatment of inflammatory bowel disease. Cochrane Database Syst. Rev. 2018, 11, CD012774. [Google Scholar] [CrossRef]
- Su, G.L.; Ko, C.W.; Bercik, P.; Falck-Ytter, Y.; Sultan, S.; Weizman, A.V.; Morgan, R.L. AGA Clinical Practice Guidelines on the Role of Probiotics in the Management of Gastrointestinal Disorders. Gastroenterology 2020, 159, 697–705. [Google Scholar] [CrossRef]
- Cho, I.; Blaser, M.J. The human microbiome: At the interface of health and disease. Nat. Rev. Genet. 2012, 13, 260–270. [Google Scholar] [CrossRef]
- Rigottier-Gois, L. Dysbiosis in inflammatory bowel diseases: The oxygen hypothesis. ISME J. 2013, 7, 1256–1261. [Google Scholar] [CrossRef] [PubMed]
- Palmela, C.; Chevarin, C.; Xu, Z.; Torres, J.; Sevrin, G.; Hirten, R.; Barnich, N.; Ng, S.C.; Colombel, J.-F. Adherent-invasive Escherichia coli in inflammatory bowel disease. Gut 2018, 67, 574–587. [Google Scholar] [CrossRef] [PubMed]
- Jellbauer, S.; Raffatellu, M. An intestinal arsonist: Pathobiont ignites IBD and flees the scene. Gut 2013, 63, 1034–1035. [Google Scholar] [CrossRef]
- Carvalho, F.A.; Barnich, N.; Sauvanet, P.; Darcha, C.; Gelot, A.; Darfeuille-Michaud, A. Crohn’s disease-associated Escherichia coli LF82 aggravates colitis in injured mouse colon via signaling by flagellin. Inflamm. Bowel Dis. 2008, 14, 1051–1060. [Google Scholar] [CrossRef]
- Carvalho, F.A.; Barnich, N.; Sivignon, A.; Darcha, C.; Chan, C.H.; Stanners, C.P.; Darfeuille-Michaud, A. Crohn’s disease adherent-invasive Escherichia coli colonize and induce strong gut inflammation in transgenic mice expressing human CEACAM. J. Exp. Med. 2009, 206, 2179–2189. [Google Scholar] [CrossRef] [PubMed]
- Imai, J.; Kitamoto, S.; Sugihara, K.; Nagao-Kitamoto, H.; Hayashi, A.; Morhardt, T.L.; Kuffa, P.; Higgins, P.D.R.; Barnich, N.; Kamada, N. Flagellin-mediated activation of IL-33-ST2 signaling by a pathobiont promotes intestinal fibrosis. Mucosal Immunol. 2019, 12, 632–643. [Google Scholar] [CrossRef]
- Kitamoto, S.; Nagao-Kitamoto, H.; Hein, R.; Schmidt, T.; Kamada, N. The Bacterial Connection between the Oral Cavity and the Gut Diseases. J. Dent. Res. 2020, 99, 1021–1029. [Google Scholar] [CrossRef]
- Hajishengallis, G.; Chavakis, T. Local and systemic mechanisms linking periodontal disease and inflammatory comorbidities. Nat. Rev. Immunol. 2021, 28, 1–15. [Google Scholar] [CrossRef]
- Haberman, Y.; Tickle, T.L.; Dexheimer, P.J.; Kim, M.-O.; Tang, D.; Karns, R.; Baldassano, R.N.; Noe, J.D.; Rosh, J.; Markowitz, J.; et al. Pediatric Crohn disease patients exhibit specific ileal transcriptome and microbiome signature. J. Clin. Investig. 2014, 124, 3617–3633. [Google Scholar] [CrossRef]
- Gevers, D.; Kugathasan, S.; Denson, L.A.; Vázquez-Baeza, Y.; Van Treuren, W.; Ren, B.; Schwager, E.; Knights, D.; Song, S.J.; Yassour, M.; et al. The Treatment-Naive Microbiome in New-Onset Crohn’s Disease. Cell Host Microbe 2014, 15, 382–392. [Google Scholar] [CrossRef]
- Atarashi, K.; Suda, W.; Luo, C.; Kawaguchi, T.; Motoo, I.; Narushima, S.; Kiguchi, Y.; Yasuma, K.; Watanabe, E.; Tanoue, T.; et al. Ectopic colonization of oral bacteria in the intestine drives TH1 cell induction and inflammation. Science 2017, 358, 359–365. [Google Scholar] [CrossRef] [PubMed]
- Papageorgiou, S.N.; Hagner, M.; Nogueira, A.V.B.; Franke, A.; Jäger, A.; Deschner, J. Inflammatory bowel disease and oral health: Systematic review and a meta-analysis. J. Clin. Periodontol. 2017, 44, 382–393. [Google Scholar] [CrossRef] [PubMed]
- Kitamoto, S.; Nagao-Kitamoto, H.; Jiao, Y.; Gillilland, M.G.; Hayashi, A.; Imai, J.; Sugihara, K.; Miyoshi, M.; Brazil, J.C.; Kuffa, P.; et al. The Intermucosal Connection between the Mouth and Gut in Commensal Pathobiont-Driven Colitis. Cell 2020, 182, 447–462.e14. [Google Scholar] [CrossRef]
- The Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 2012, 486, 207–214. [Google Scholar] [CrossRef]
- Franzosa, E.A.; Sirota-Madi, A.; Avila-Pacheco, J.; Fornelos, N.; Haiser, H.J.; Reinker, S.; Vatanen, T.; Hall, A.B.; Mallick, H.; McIver, L.J.; et al. Gut microbiome structure and metabolic activity in inflammatory bowel disease. Nat. Microbiol. 2019, 4, 293–305. [Google Scholar] [CrossRef]
- Morgan, X.C.; Tickle, T.L.; Sokol, H.; Gevers, D.; Devaney, K.L.; Ward, D.V.; Reyes, J.A.; Shah, S.A.; Leleiko, N.; Snapper, S.B.; et al. Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome Biol. 2012, 13, R79. [Google Scholar] [CrossRef]
- Becattini, S.; Sorbara, M.T.; Kim, S.G.; Littmann, E.L.; Dong, Q.; Walsh, G.; Wright, R.; Amoretti, L.; Fontana, E.; Hohl, T.M.; et al. Rapid transcriptional and metabolic adaptation of intestinal microbes to host immune activation. Cell Host Microbe 2021, 29, 378–393.e5. [Google Scholar] [CrossRef]
- Kitamoto, S.; Alteri, C.J.; Rodrigues, M.; Nagao-Kitamoto, H.; Sugihara, K.; Himpsl, S.D.; Bazzi, M.; Miyoshi, M.; Nishioka, T.; Hayashi, A.; et al. Dietary l-serine confers a competitive fitness advantage to Enterobacteriaceae in the inflamed gut. Nat. Microbiol. 2020, 5, 116–125. [Google Scholar] [CrossRef]
- Franzosa, E.A.; Morgan, X.C.; Segata, N.; Waldron, L.; Reyes, J.; Earl, A.M.; Giannoukos, G.; Boylan, M.R.; Ciulla, D.; Gevers, D.; et al. Relating the metatranscriptome and metagenome of the human gut. Proc. Natl. Acad. Sci. USA 2014, 111, E2329–E2338. [Google Scholar] [CrossRef] [PubMed]
- Schirmer, M.; Franzosa, E.A.; Lloyd-Price, J.; McIver, L.J.; Schwager, R.; Poon, T.W.; Ananthakrishnan, A.N.; Andrews, E.; Barron, G.; Lake, K.; et al. Dynamics of metatranscription in the inflammatory bowel disease gut microbiome. Nat. Microbiol. 2018, 3, 337–346. [Google Scholar] [CrossRef]
- Lloyd-Price, J.; Ibdmdb, I.; Arze, C.; Ananthakrishnan, A.N.; Schirmer, M.; Avila-Pacheco, J.; Poon, T.W.; Andrews, E.; Ajami, N.J.; Bonham, K.S.; et al. Multi-omics of the gut microbial ecosystem in inflammatory bowel diseases. Nat. Cell Biol. 2019, 569, 655–662. [Google Scholar] [CrossRef] [PubMed]
- Aden, K.; Rehman, A.; Waschina, S.; Pan, W.-H.; Walker, A.; Lucio, M.; Nunez, A.M.; Bharti, R.; Zimmerman, J.; Bethge, J.; et al. Metabolic Functions of Gut Microbes Associate with Efficacy of Tumor Necrosis Factor Antagonists in Patients with Inflammatory Bowel Diseases. Gastroenterology 2019, 157, 1279–1292.e11. [Google Scholar] [CrossRef] [PubMed]
- Makki, K.; Deehan, E.C.; Walter, J.; Bäckhed, F. The Impact of Dietary Fiber on Gut Microbiota in Host Health and Disease. Cell Host Microbe 2018, 23, 705–715. [Google Scholar] [CrossRef]
- Davie, J.R. Inhibition of Histone Deacetylase Activity by Butyrate. J. Nutr. 2003, 133, 2485S–2493S. [Google Scholar] [CrossRef] [PubMed]
- Furusawa, Y.; Obata, Y.; Fukuda, S.; Endo, T.A.; Nakato, G.; Takahashi, D.; Nakanishi, Y.; Uetake, C.; Kato, K.; Kato, T.; et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 2013, 504, 446–450. [Google Scholar] [CrossRef]
- Singh, N.; Gurav, A.; Sivaprakasam, S.; Brady, E.; Padia, R.; Shi, H.; Thangaraju, M.; Prasad, P.D.; Manicassamy, S.; Munn, D.H.; et al. Activation of Gpr109a, receptor for niacin and the commensal metabolite butyrate, suppresses colonic inflammation and carcinogenesis. Immunity 2014, 40, 128–139. [Google Scholar] [CrossRef] [PubMed]
- Fukuda, S.; Toh, H.; Hase, K.; Oshima, K.; Nakanishi, Y.; Yoshimura, K.; Tobe, T.; Clarke, J.M.; Topping, D.L.; Suzuki, T.; et al. Bifidobacteria can protect from enteropathogenic infection through production of acetate. Nature 2011, 469, 543–547. [Google Scholar] [CrossRef]
- Kelly, C.J.; Zheng, L.; Campbell, E.L.; Saeedi, B.; Scholz, C.C.; Bayless, A.J.; Wilson, K.E.; Glover, L.E.; Kominsky, D.J.; Magnuson, A.; et al. Crosstalk between Microbiota-Derived Short-Chain Fatty Acids and Intestinal Epithelial HIF Augments Tissue Barrier Function. Cell Host Microbe 2015, 17, 662–671. [Google Scholar] [CrossRef]
- Paassen, N.B.-V.; Vincent, A.; Puiman, P.J.; Van Der Sluis, M.; Bouma, J.; Boehm, G.; Van Goudoever, J.B.; Van Seuningen, I.; Renes, I.B. The regulation of intestinal mucin MUC2 expression by short-chain fatty acids: Implications for epithelial protection. Biochem. J. 2009, 420, 211–219. [Google Scholar] [CrossRef][Green Version]
- Desai, M.S.; Seekatz, A.M.; Koropatkin, N.M.; Kamada, N.; Hickey, C.A.; Wolter, M.; Pudlo, N.A.; Kitamoto, S.; Terrapon, N.; Muller, A.; et al. A Dietary Fiber-Deprived Gut Microbiota Degrades the Colonic Mucus Barrier and Enhances Pathogen Susceptibility. Cell 2016, 167, 1339–1353.e21. [Google Scholar] [CrossRef] [PubMed]
- Schroeder, B.O.; Birchenough, G.M.; Ståhlman, M.; Arike, L.; Johansson, M.E.; Hansson, G.C.; Bäckhed, F. Bifidobacteria or Fiber Protects against Diet-Induced Microbiota-Mediated Colonic Mucus Deterioration. Cell Host Microbe 2018, 23, 27–40.e7. [Google Scholar] [CrossRef] [PubMed]
- Thibault, R.; De Coppet, P.; Daly, K.; Bourreille, A.; Cuff, M.; Bonnet, C.; Mosnier, J.; Galmiche, J.; Shirazi–Beechey, S.; Segain, J. Down-Regulation of the Monocarboxylate Transporter 1 Is Involved in Butyrate Deficiency During Intestinal Inflammation. Gastroenterology 2007, 133, 1916–1927. [Google Scholar] [CrossRef] [PubMed]
- Chapman, M.A.; Grahn, M.F.; Boyle, M.A.; Hutton, M.; Rogers, J.; Williams, N.S. Butyrate oxidation is impaired in the colonic mucosa of sufferers of quiescent ulcerative colitis. Gut 1994, 35, 73–76. [Google Scholar] [CrossRef] [PubMed]
- De Preter, V.; Arijs, I.; Windey, K.; Vanhove, W.; Vermeire, S.; Schuit, F.; Rutgeerts, P.; Verbeke, K. Impaired butyrate oxidation in ulcerative colitis is due to decreased butyrate uptake and a defect in the oxidation pathway. Inflamm. Bowel Dis. 2012, 18, 1127–1136. [Google Scholar] [CrossRef] [PubMed]
- De Preter, V.; Rutgeerts, P.; Verbeke, K.; Schuit, F.; Arijs, I. Impaired Expression of Genes Involved in the Butyrate Oxidation Pathway in Crohn’s Disease Patients. Inflamm. Bowel Dis. 2013, 19, E43–E44. [Google Scholar] [CrossRef]
- Silveira, A.L.M.; Ferreira, A.V.M.; De Oliveira, M.C.; Rachid, M.A.; Sousa, L.F.D.C.; Martins, F.D.S.; Gomes-Santos, A.C.; Vieira, A.T.; Teixeira, M.M. Preventive rather than therapeutic treatment with high fiber diet attenuates clinical and inflammatory markers of acute and chronic DSS-induced colitis in mice. Eur. J. Nutr. 2015, 56, 179–191. [Google Scholar] [CrossRef]
- Ishisono, K.; Mano, T.; Yabe, T.; Kitaguchi, K. Dietary Fiber Pectin Ameliorates Experimental Colitis in a Neutral Sugar Side Chain-Dependent Manner. Front. Immunol. 2019, 10, 2979. [Google Scholar] [CrossRef]
- Scheppach, W.; Sommer, H.; Kirchner, T.; Paganelli, G.-M.; Bartram, P.; Christl, S.; Richter, F.; Dusel, G.; Kasper, H. Effect of butyrate enemas on the colonic mucosa in distal ulcerative colitis. Gastroenterology 1992, 103, 51–56. [Google Scholar] [CrossRef]
- Breuer, R.I.; Soergel, K.H.; Lashner, B.A.; Christ, M.L.; Hanauer, S.B.; Vanagunas, A.; Harig, J.M.; Keshavarzian, A.; Robinson, M.; Sellin, J.H.; et al. Short chain fatty acid rectal irrigation for left-sided ulcerative colitis: A randomised, placebo controlled trial. Gut 1997, 40, 485–491. [Google Scholar] [CrossRef]
- Fiorucci, S.; Carino, A.; Baldoni, M.; Santucci, L.; Costanzi, E.; Graziosi, L.; Distrutti, E.; Biagioli, M. Bile Acid Signaling in Inflammatory Bowel Diseases. Dig. Dis. Sci. 2021, 66, 674–693. [Google Scholar] [CrossRef] [PubMed]
- Hang, S.; Paik, D.; Yao, L.; Kim, E.; Trinath, J.; Lu, J.; Ha, S.; Nelson, B.N.; Kelly, S.P.; Wu, L.; et al. Bile acid metabolites control TH17 and Treg cell differentiation. Nat. Cell Biol. 2019, 576, 143–148. [Google Scholar] [CrossRef]
- Campbell, C.; McKenney, P.T.; Konstantinovsky, D.; Isaeva, O.I.; Schizas, M.; Verter, J.; Mai, C.; Jin, W.-B.; Guo, C.-J.; Violante, S.; et al. Bacterial metabolism of bile acids promotes generation of peripheral regulatory T cells. Nat. Cell Biol. 2020, 581, 475–479. [Google Scholar] [CrossRef]
- Song, X.; Sun, X.; Oh, S.F.; Wu, M.; Zhang, Y.; Zheng, W.; Geva-Zatorsky, N.; Jupp, R.; Mathis, D.; Benoist, C.; et al. Microbial bile acid metabolites modulate gut RORγ+ regulatory T cell homeostasis. Nat. Cell Biol. 2020, 577, 410–415. [Google Scholar] [CrossRef]
- Devkota, S.; Wang, Y.; Musch, M.W.; Leone, V.; Fehlner-Peach, H.; Nadimpalli, A.; Antonopoulos, D.A.; Jabri, B.; Chang, E.B. Dietary-fat-induced taurocholic acid promotes pathobiont expansion and colitis in Il10−/− mice. Nat. Cell Biol. 2012, 487, 104–108. [Google Scholar] [CrossRef]
- Narula, N.; Dhillon, A.; Zhang, D.; Sherlock, M.E.; Tondeur, M.; Zachos, M. Enteral nutritional therapy for induction of remission in Crohn’s disease. Cochrane Database Syst. Rev. 2018, 4, CD000542. [Google Scholar] [CrossRef] [PubMed]
- Fritsch, J.; Garces, L.; Quintero, M.A.; Pignac-Kobinger, J.; Santander, A.M.; Fernández, I.; Ban, Y.J.; Kwon, D.; Phillips, M.C.; Knight, K.; et al. Low-Fat, High-Fiber Diet Reduces Markers of Inflammation and Dysbiosis and Improves Quality of Life in Patients with Ulcerative Colitis. Clin. Gastroenterol. Hepatol. 2020. [Google Scholar] [CrossRef]
- Gao, J.; Xu, K.; Liu, H.; Liu, G.; Bai, M.; Peng, C.; Li, T.; Yin, Y. Impact of the Gut Microbiota on Intestinal Immunity Mediated by Tryptophan Metabolism. Front. Cell. Infect. Microbiol. 2018, 8, 13. [Google Scholar] [CrossRef] [PubMed]
- Zelante, T.; Iannitti, R.G.; Cunha, C.; De Luca, A.; Giovannini, G.; Pieraccini, G.; Zecchi, R.; D’Angelo, C.; Massi-Benedetti, C.; Fallarino, F.; et al. Tryptophan catabolites from microbiota engage aryl hydrocarbon receptor and balance mucosal reactivity via interleukin-22. Immunity 2013, 39, 372–385. [Google Scholar] [CrossRef] [PubMed]
- Peterson, L.W.; Artis, D. Intestinal epithelial cells: Regulators of barrier function and immune homeostasis. Nat. Rev. Immunol. 2014, 14, 141–153. [Google Scholar] [CrossRef]
- Rivas, M.A.; National Institute of Diabetes and Digestive Kidney Diseases Inflammatory Bowel Disease Genetics Consortium (NIDDK IBDGC); Beaudoin, M.; Gardet, A.; Stevens, C.; Sharma, Y.; Zhang, C.K.; Boucher, G.; Ripke, S.; Ellinghaus, D.; et al. Deep resequencing of GWAS loci identifies independent rare variants associated with inflammatory bowel disease. Nat. Genet. 2011, 43, 1066–1073. [Google Scholar] [CrossRef]
- Lamas, B.; Richard, M.L.; Leducq, V.; Pham, H.-P.; Michel, M.-L.; Da Costa, G.; Bridonneau, C.; Jegou, S.; Hoffmann, T.W.; Natividad, J.M.; et al. CARD9 impacts colitis by altering gut microbiota metabolism of tryptophan into aryl hydrocarbon receptor ligands. Nat. Med. 2016, 22, 598–605. [Google Scholar] [CrossRef]
- Newman, A.C.; Maddocks, O.D. Serine and Functional Metabolites in Cancer. Trends Cell Biol. 2017, 27, 645–657. [Google Scholar] [CrossRef]
- Maddocks, O.D.K.; Athineos, D.; Cheung, E.C.; Lee, P.; Zhang, T.; Broek, N.J.F.V.D.; Mackay, G.M.; Labuschagne, C.F.; Gay, D.; Kruiswijk, F.; et al. Modulating the therapeutic response of tumours to dietary serine and glycine starvation. Nature 2017, 544, 372–376. [Google Scholar] [CrossRef] [PubMed]
- Ma, E.H.; Bantug, G.; Griss, T.; Condotta, S.; Johnson, R.M.; Samborska, B.; Mainolfi, N.; Suri, V.; Guak, H.; Balmer, M.L.; et al. Serine Is an Essential Metabolite for Effector T Cell Expansion. Cell Metab. 2017, 25, 345–357. [Google Scholar] [CrossRef]
- Rodriguez, A.E.; Ducker, G.S.; Billingham, L.K.; Martinez, C.A.; Mainolfi, N.; Suri, V.; Friedman, A.; Manfredi, M.G.; Weinberg, S.E.; Rabinowitz, J.D.; et al. Serine Metabolism Supports Macrophage IL-1β Production. Cell Metab. 2019, 29, 1003–1011.e4. [Google Scholar] [CrossRef]
- Zhang, H.; Hua, R.; Zhang, B.; Zhang, X.; Yang, H.; Zhou, X. Serine Alleviates Dextran Sulfate Sodium-Induced Colitis and Regulates the Gut Microbiota in Mice. Front. Microbiol. 2018, 9, 3062. [Google Scholar] [CrossRef] [PubMed]
- Laudisi, F.; Stolfi, C.; Monteleone, G. Impact of Food Additives on Gut Homeostasis. Nutrients 2019, 11, 2334. [Google Scholar] [CrossRef] [PubMed]
- Bancil, A.S.; Sandall, A.M.; Rossi, M.; Chassaing, B.; Lindsay, J.O.; Whelan, K. Food Additive Emulsifiers and Their Impact on Gut Microbiome, Permeability, and Inflammation: Mechanistic Insights in Inflammatory Bowel Disease. J. Crohns Coliti 2020, 10, jjaa254. [Google Scholar] [CrossRef] [PubMed]
- Chassaing, B.; Koren, O.; Goodrich, J.K.; Poole, A.C.; Srinivasan, S.; Ley, R.E.; Gewirtz, A.T. Dietary emulsifiers impact the mouse gut microbiota promoting colitis and metabolic syndrome. Nat. Cell Biol. 2015, 519, 92–96. [Google Scholar] [CrossRef] [PubMed]
- Chassaing, B.; Van De Wiele, T.; De Bodt, J.; Marzorati, M.; Gewirtz, A.T. Dietary emulsifiers directly alter human microbiota composition and gene expression ex vivo potentiating intestinal inflammation. Gut 2017, 66, 1414–1427. [Google Scholar] [CrossRef] [PubMed]
- Viennois, E.; Bretin, A.; Dubé, P.E.; Maue, A.C.; Dauriat, C.J.; Barnich, N.; Gewirtz, A.T.; Chassaing, B. Dietary Emulsifiers Directly Impact Adherent-Invasive E. coli Gene Expression to Drive Chronic Intestinal Inflammation. Cell Rep. 2020, 33, 108229. [Google Scholar] [CrossRef]
- Sandall, A.M.; Cox, S.R.; Lindsay, J.O.; Gewirtz, A.T.; Chassaing, B.; Rossi, M.; Whelan, K. Emulsifiers Impact Colonic Length in Mice and Emulsifier Restriction is Feasible in People with Crohn’s Disease. Nutrients 2020, 12, 2827. [Google Scholar] [CrossRef] [PubMed]
- Halmos, E.P.; Gibson, P.R. Dietary management of IBD—insights and advice. Nat. Rev. Gastroenterol. Hepatol. 2015, 12, 133–146. [Google Scholar] [CrossRef]
- Gerasimidis, K.; Bertz, M.; Hanske, L.; Junick, J.; Biskou, O.; Aguilera, M.; Garrick, V.; Russell, R.K.; Blaut, M.; McGrogan, P.; et al. Decline in Presumptively Protective Gut Bacterial Species and Metabolites Are Paradoxically Associated with Disease Improvement in Pediatric Crohn’s Disease During Enteral Nutrition. Inflamm. Bowel Dis. 2014, 20, 861–871. [Google Scholar] [CrossRef] [PubMed]
- Quince, C.; Ijaz, U.Z.; Loman, N.; Eren, M.A.; Saulnier, D.; Russell, J.; Haig, S.J.; Calus, S.T.; Quick, J.; Barclay, A.; et al. Extensive Modulation of the Fecal Metagenome in Children with Crohn’s Disease During Exclusive Enteral Nutrition. Am. J. Gastroenterol. 2015, 110, 1718–1729. [Google Scholar] [CrossRef] [PubMed]
- Lewis, J.D.; Chen, E.Z.; Baldassano, R.N.; Otley, A.R.; Griffiths, A.M.; Lee, D.; Bittinger, K.; Bailey, A.; Friedman, E.S.; Hoffmann, C.; et al. Inflammation, Antibiotics, and Diet as Environmental Stressors of the Gut Microbiome in Pediatric Crohn’s Disease. Cell Host Microbe 2015, 18, 489–500. [Google Scholar] [CrossRef]
- Cox, S.R.; Lindsay, J.O.; Fromentin, S.; Stagg, A.J.; McCarthy, N.E.; Galleron, N.; Ibraim, S.B.; Roume, H.; Levenez, F.; Pons, N.; et al. Effects of Low FODMAP Diet on Symptoms, Fecal Microbiome, and Markers of Inflammation in Patients with Quiescent Inflammatory Bowel Disease in a Randomized Trial. Gastroenterology 2020, 158, 176–188.e7. [Google Scholar] [CrossRef]
- Halmos, E.P.; Christophersen, C.T.; Bird, A.R.; Shepherd, S.J.; Muir, J.G.; Gibson, P.R. Consistent Prebiotic Effect on Gut Microbiota With Altered FODMAP Intake in Patients with Crohn’s Disease: A Randomised, Controlled Cross-Over Trial of Well-Defined Diets. Clin Transl Gastroenterol. 2016, 7, e164. [Google Scholar] [CrossRef]
- Svolos, V.; Hansen, R.; Nichols, B.; Quince, C.; Ijaz, U.Z.; Papadopoulou, R.T.; Edwards, C.A.; Watson, D.; Alghamdi, A.; Brejnrod, A.; et al. Treatment of Active Crohn’s Disease with an Ordinary Food-based Diet That Replicates Exclusive Enteral Nutrition. Gastroenterology 2019, 156, 1354–1367.e6. [Google Scholar] [CrossRef]
- Borrelli, O.; Cordischi, L.; Cirulli, M.; Paganelli, M.; Labalestra, V.; Uccini, S.; Russo, P.M.; Cucchiara, S. Polymeric Diet Alone Versus Corticosteroids in the Treatment of Active Pediatric Crohn’s Disease: A Randomized Controlled Open-Label Trial. Clin. Gastroenterol. Hepatol. 2006, 4, 744–753. [Google Scholar] [CrossRef]
- Triantafillidis, J.K.; Vagianos, C.; Papalois, A.E. The Role of Enteral Nutrition in Patients with Inflammatory Bowel Disease: Current Aspects. BioMed Res. Int. 2015, 2015, 197167. [Google Scholar] [CrossRef]
- Sigall-Boneh, R.; Pfeffer-Gik, T.; Segal, I.; Zangen, T.; Boaz, M.; Levine, A. Partial Enteral Nutrition with a Crohn’s Disease Exclusion Diet Is Effective for Induction of Remission in Children and Young Adults with Crohn’s Disease. Inflamm. Bowel Dis. 2014, 20, 1353–1360. [Google Scholar] [CrossRef] [PubMed]
- Boneh, R.S.; Shabat, C.S.; Yanai, H.; Chermesh, I.; Ben Avraham, S.; Boaz, M.; Levine, A. Dietary Therapy with the Crohn’s Disease Exclusion Diet is a Successful Strategy for Induction of Remission in Children and Adults Failing Biological Therapy. J. Crohns Coliti 2017, 11, 1205–1212. [Google Scholar] [CrossRef] [PubMed]
- Cohen, S.A.; Gold, B.D.; Oliva, S.; Lewis, J.; Stallworth, A.; Koch, B.; Eshee, L.; Mason, D. Clinical and Mucosal Improvement with Specific Carbohydrate Diet in Pediatric Crohn Disease. J. Pediatr. Gastroenterol. Nutr. 2014, 59, 516–521. [Google Scholar] [CrossRef] [PubMed]
- Limketkai, B.N.; Iheozor-Ejiofor, Z.; Gjuladin-Hellon, T.; Parian, A.; Matarese, L.E.; Bracewell, K.; Macdonald, J.K.; Gordon, M.; Mullin, G.E. Dietary interventions for induction and maintenance of remission in inflammatory bowel disease. Cochrane Database Syst. Rev. 2019, 2, CD012839. [Google Scholar] [CrossRef] [PubMed]
- Sugihara, K.; Morhardt, T.L.; Kamada, N. The Role of Dietary Nutrients in Inflammatory Bowel Disease. Front. Immunol. 2019, 9, 3183. [Google Scholar] [CrossRef]
- Hurtado-Lorenzo, A.; Honig, G.; Heller, C. Precision Nutrition Initiative: Toward Personalized Diet Recommendations for Patients with Inflammatory Bowel Diseases. Crohn’s Colitis 2020, 2, otaa087. [Google Scholar] [CrossRef]
| Year | Study Design | Subject | Nutritonal Intervention | Results | Ref | |
|---|---|---|---|---|---|---|
| Clinical | Gut Microbiota/Metabolites | |||||
| 2014 | Cohort | CD: 15, HC: 21 | EEN (PD) | no data | Diversity ↓, Faecalibacterium prausnitzii ↓, butyrate ↓, sulfide ↑ | [104] |
| 2015 | Cohort | CD: 23, HC: 21 | EEN (PD) | 62% remission rate | Diversity ↓, Bifidobacterium ↓, Ruminococcus ↓, Faecalibacterium ↓, Akkermansia ↓, Lactococcus ↑ | [105] |
| 2015 | Cohort | CD: 90 | EEN vs. PEN vs. Anti-TNF | 45% clinical response | Haemophilus ↓, Streptococcus ↓, Dialister ↓, Dorea ↓, Gordonibacter ↓, Alistipes ↑ | [106] |
| 2020 | RCT | IBD: 52 | Low-FODMAP diet vs. control diet | Relief of GI symptoms ↑, IBS score ↓, HR-QOL ↑ | Bifidobacterium adolescentis↓, B. longum↓, Faecalibacterium prausnitzii↓, B. dentium↑ | [107] |
| 2016 | Cross-over | CD: 9 | Low-FODMAP diet vs. Australian diet | GI symptoms ↓, Fecal calprotectin → | Clostridium cluster XIVa ↓, Akkermansia muciniphila ↓, Ruminococcus torques ↑, SCFA → | [108] |
| 2019 | RCT | CD: 78 | CDED + PEN vs. EEN (PD) | % remission rate→, sustained remission ↑, tolerability ↑ | Haemophilus ↓, Veillonella ↓, Bifidobacterium ↓, Prevotella ↓, Anaerostipes ↓, Oscillibacter ↑, Roseburia ↑ | [28] |
| 2020 | Cross-over | UC: 17 | Low-fat diet vs. improved standard American diet | QOL ↑, amyloid A ↓ | Actinobacteria ↓, Bacteroidetes ↑, Faecalibacterium prausnitzii ↑, Prevotella ↑, acetate ↑, Trp ↑, lauric acid ↓ | [86] |
| 2019 | RCT/cohort | HC: 25 (microbiota) CD: 5 (clinical) | CD-TREAT vs. EEN (PD) vs. habitual diet | 80% clinical response, fecal calprotectin ↓ | Prevotella ↑, Escherichia Shigella ↑, Eisenbergiella ↑, Lachnoclostridium ↑, Bifidobacterium ↓, Faecalibacterium ↓, Ruminococcus ↓ | [109] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sugihara, K.; Kamada, N. Diet–Microbiota Interactions in Inflammatory Bowel Disease. Nutrients 2021, 13, 1533. https://doi.org/10.3390/nu13051533
Sugihara K, Kamada N. Diet–Microbiota Interactions in Inflammatory Bowel Disease. Nutrients. 2021; 13(5):1533. https://doi.org/10.3390/nu13051533
Chicago/Turabian StyleSugihara, Kohei, and Nobuhiko Kamada. 2021. "Diet–Microbiota Interactions in Inflammatory Bowel Disease" Nutrients 13, no. 5: 1533. https://doi.org/10.3390/nu13051533
APA StyleSugihara, K., & Kamada, N. (2021). Diet–Microbiota Interactions in Inflammatory Bowel Disease. Nutrients, 13(5), 1533. https://doi.org/10.3390/nu13051533






