1. Introduction
Changes in the gut microbiota [
1] and its derived metabolites are closely associated with insulin sensitivity [
2,
3], incretin secretion [
4,
5], and energy homeostasis [
6]. Thus, the gut microbiota has been attracting much attention in metabolic diseases such as obesity and type 2 diabetes mellitus (T2DM).
Previously, we reported the presence of gut dysbiosis and bacterial translocation in patients with T2DM [
7]. In a subsequent randomized controlled trial, we found that probiotic administration reduced the translocation of gut bacteria to blood in patients with T2DM [
8]. Plasma lipopolysaccharide (LPS) from gram-negative bacteria, like translocation of gut bacteria, is an inflammatory mediator that contributes to insulin resistance [
9,
10]. Translocated LPS in the systemic circulation binds to LPS-binding protein (LBP), which is a marker for metabolic syndrome [
11,
12], and our previous study showed clear positive associations between plasma levels of interleukin-6 and LBP in patients with T2DM [
13]. Furthermore, we found higher plasma LBP levels in T2DM patients with obesity and poor glycemic control [
13]. Therefore, a possible therapeutic approach in obese patients with T2DM is to control low-grade chronic inflammation by reducing not only translocation of gut bacteria but also the levels of endotoxins such as LPS. According to the previous review [
14], it has been reported that probiotics administration in patients with metabolic syndrome resulted in improvements in body mass index, lipid, and glucose metabolism, and probiotics also positively affected inflammatory markers such as interleukine-6.
While probiotics may have various beneficial effects on metabolic disease as mentioned above, the combination of one or more probiotics and prebiotics, a mixture generally referred to as a synbiotic, may confer further significant benefits in the human gut environment. Indeed, perioperative synbiotic treatment consisting of two probiotic strains,
Lacticaseibacillus paracasei (the previous taxonomic nomenclature was
Lactobacillus casei) strain Shirota (LcS) and
Bifidobacterium breve strain Yakult (BbrY), along with galacto-oligosaccharides (GOS), significantly prevented postoperative infectious complications due to reduction of bacterial translocation [
15]. Thus, our hypothesis is that synbiotics might be expected to effectively inhibit bacterial translocation in metabolic diseases, and subsequently might reduce chronic inflammation. Here, we performed a 24-week, interventional, randomized controlled study to investigate the effects of daily intake of a synbiotic comprising LcS, BbrY, and GOS on chronic inflammation, gut microbiota, fecal organic acids, and bacterial translocation in obese patients with T2DM.
2. Materials and Methods
2.1. Participants
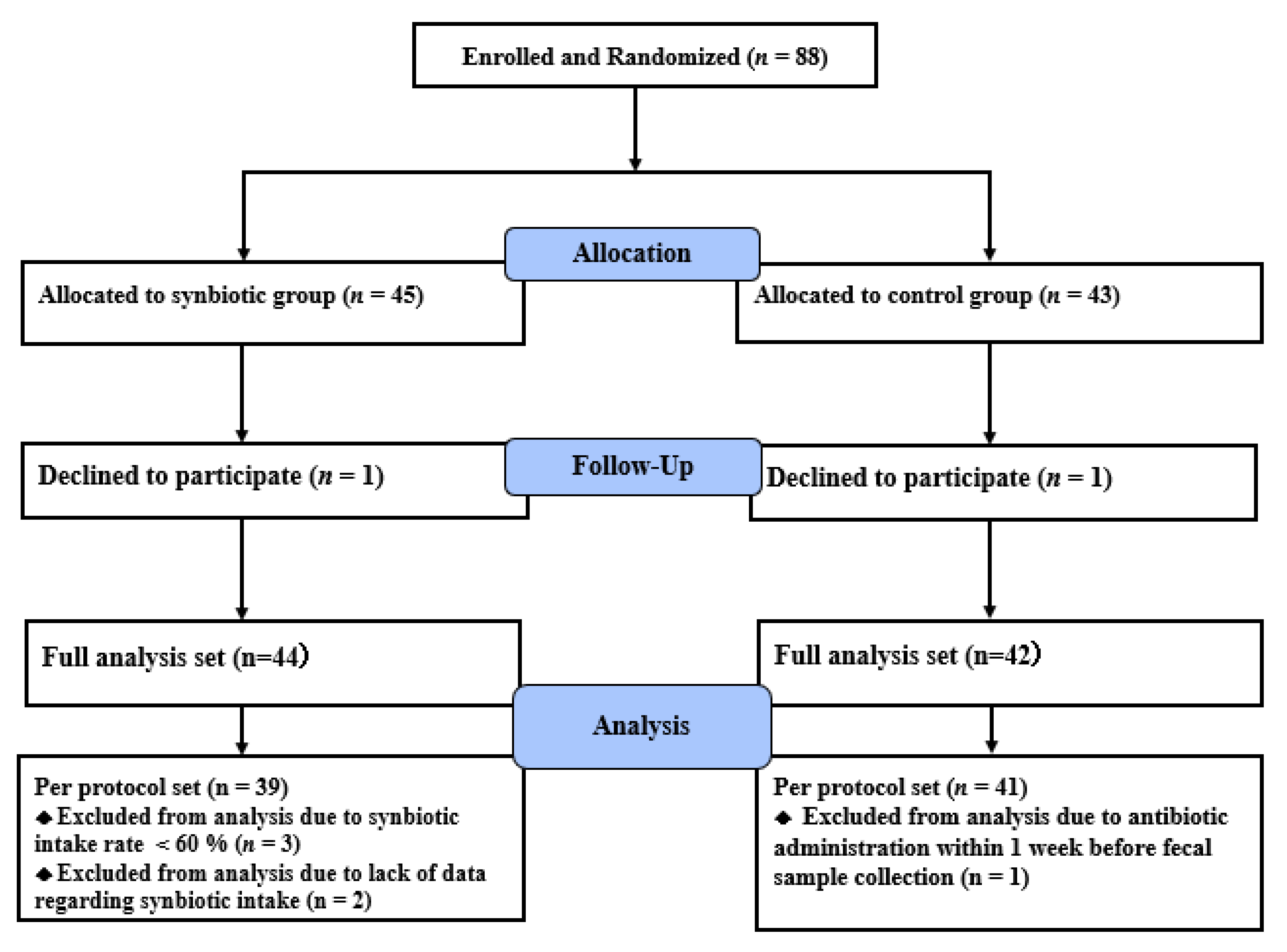
A total of 88 obese patients with T2DM were recruited from the outpatient clinic of Juntendo University Hospital (n = 72) and International Good Will Hospital (n = 16) between July 2018 and April 2019. The following inclusion criteria were applied at study registration: (1) age ≥ 30 but <80 years, (2) HbA1c (NGSP) ≥ 6.0 but <9.0%, (3) body mass index (BMI) ≥ 25.0 kg/m2, and (4) treatment with only diet and exercise or medicines. The selected patients were excluded from the study if any of the following conditions were diagnosed at registration: (1) serious kidney disease (serum creatinine level ≥ 1.5 mg/dL and/or hemodialysis), (2) serious liver disease excluding fatty liver, (3) inflammatory bowel disease, (4) type 1 diabetes mellitus, (5) past history of digestive surgery, (6) allergy to milk, (7) treatment with an α-glucosidase inhibitor, and (8) unsuitability for the study (i.e., irregular visits). This study was registered in the University Hospital Medical Information Network Clinical Trials Registry, a non-profit organization in Japan, and it meets the requirements of the International Committee of Medical Journal Editors (UMIN000032057, registration date: 2 April 2018).
2.2. Study Design
The primary endpoint was the change in the level of interleukin-6 (IL-6) from baseline to the end of the study. The secondary endpoints were changes from baseline to the end of the study in high-sensitivity C-reactive protein (hs-CRP), LBP, bacterial counts in feces and blood, fecal organic acid concentrations, diversity of the gut microbiota in feces, and the levels of fasting blood glucose, HbA1c, and lipids. Our previous study showed that the IL-6 level was 1.8 ± 0.9 pg/mL before probiotic administration and 0.20 ± 1.00 pg/mL after the intervention [
8]. In another study, the plasma IL-6 level was reduced by around 40% after synbiotic intervention [
15]. Based on these previous reports, we assumed that the effect size could be 0.65 ± 1.00 pg/mL between control and synbiotic groups. With a two-sided α level of 5% and power (1-β) of 80%, at least 78 patients (39 patients in each group) would permit the detection of IL-6 changes. Therefore, after considering the possibility of a 10% dropout rate, we recruited 88 patients who were then randomly assigned to either a control group or synbiotic group for 24 weeks. Randomization was performed using a dynamic allocation method based on HbA1c and BMI at baseline (Soiken, Inc., Osaka, Japan). During the study period, physicians in charge were permitted to change patients’ diabetes medications as needed. Samples for biochemical assays and for analysis of the gut microbiota in blood and feces and of fecal organic acids were obtained after overnight fasts at each hospital visit (0, 12, and 24 weeks). At 24 weeks, patients with synbiotic intake rate <60% and those who took antibiotics within 1 week before the collection of the fecal samples were excluded from the per protocol set analysis.
2.3. Synbiotic Supplementation
As synbiotic intervention, the following agents were administered orally: 3.0 g dry powder containing at least 3 × 108 living Lacticaseibacillus paracasei YIT 9029 (strain Shirota: LcS) organisms, 3 × 108 living Bifidobacterium breve YIT 12272 (BbrY) organisms, and 7.5 g GOS per day (product name: Yakult Super Synbiotics LBG-P, Yakult Honsha Co., Ltd., Tokyo, Japan). Patients were instructed to take the synbiotic twice a day (2.0 g dry powder and 5.0 g GOS at breakfast and 1.0 g dry powder and 2.5 g GOS at dinner). The nutritional composition of 3.0 g dry powder and of 7.5 g GOS was as follows: energy, 12.0 kcal and 54.9 kcal; protein, 0.03 g and 0.00 g; lipids, 0.0 g and 0.0 g; and carbohydrates, 3.0 g and 15.3 g, respectively. The participants in the synbiotic group consumed the aforementioned dose every day for 24 weeks; this was verified by up to three telephone calls to each patient, as necessary, just before their hospital visits (0, 12, and 24 weeks). In addition, each patient in the synbiotic group was instructed to keep a diary of synbiotic intake, and the control group was told not to take any synbiotic. During the study period, all participants were prohibited from consuming any other probiotics or prebiotics. In addition, the participants in the synbiotic group were instructed to reduce their calorie intake by about 60 kcal/day considering the additional calories from the synbiotic agent.
2.4. Determination of Bacterial Count by rRNA-Targeted Reverse Transcription-Quantitative PCR (RT-qPCR) and qPCR
We examined the gut microbiota composition and plasma levels of the gut bacteria using Yakult Intestinal Flora-SCAN (YIF-SCAN
®), a bacterial rRNA-targeted RT-qPCR system [
16,
17,
18]. The threshold cycle values in the linear range of the assay were applied to the standard curve to obtain the corresponding bacterial cell count in each nucleic acid sample. These data were then used to determine the number of bacteria per sample. The specificity of the RT-qPCR assay using group-, genus- or species-specific primers was determined as previously described [
16,
17,
18,
19]. For the enumeration of LcS and BbrY in feces, qPCR analysis was performed using previously described methods [
20,
21]. The sequences of the primers are listed in
Table S1.
2.5. 16S rRNA Gene Sequencing for Microbiota Analysis
Bacterial DNA in feces was extracted as previously described [
22]. The V1-2 regions of the 16S rRNA gene in each sample were amplified using the forward 27Fmod2 and reverse 338R primers [
23]. Amplification and sequencing were performed using an ABI PRISM
® 7500 Real-Time PCR System (Applied Biosystems, Framingham, MA, USA) and a MiSeq sequencer with MiSeq Reagent Kits v2 (Illumina, SanDiego, CA, USA) as previously described [
24]. The sequences generated from the MiSeq platform were analyzed using the open-source software package Quantitative Insights Into Microbial Ecology 2 (QIIME2) (2020.2) [
25], and the SILVA 138 database (
https://www.arb-silva.de/) was used to annotate taxonomic information. α-diversities represented as the number of observed operational taxonomic units (OTUs), the Shannon index, and phylogenetic diversity (PD) were estimated for 5000 randomly selected sequences to account for differences in sampling effort between the samples.
2.6. Measurement of Organic Acids and pH in Fecal Samples
The pH of stool specimens was analyzed using a handheld pH meter (model IQ150; IQ Scientific Instruments, San Diego, CA, USA). The concentration of organic acids in each sample was measured using a high-performance liquid chromatography system equipped with 432 electroconductivity detectors (Waters), as previously described [
19]. In addition, all assays were performed blindly, including those involving the gut microbiota and organic acids.
2.7. Biochemical Assays
Serum lipids (total cholesterol, high-density lipoprotein cholesterol, and triglycerides), fasting blood glucose, and HbA1c were measured with standard techniques. The plasma levels of hs-CRP and IL-6 were measured by latex nephelometry, chemiluminescent enzyme immunoassay, and enzyme-linked immunosorbent assay, respectively, at a private laboratory (SRL Laboratory, Tokyo, Japan). The plasma level of LBP was measured using a Human LBP ELISA Kit (RayBiotech, GA, USA).
2.8. Statistical Analyses
All statistical analyses were performed by a private company (Soiken, Inc., Osaka, Japan) with SAS software version 9.4 (SAS Institute, Cary, NC, USA). Normally distributed data were expressed as mean ± standard deviation and were analyzed by the t-test. Data with skewed distribution were expressed as median (interquartile range) and were analyzed by the Wilcoxon rank sum test. The detection rates of fecal and blood bacteria and fecal organic acids in both groups were analyzed by Fisher’s exact probability test.
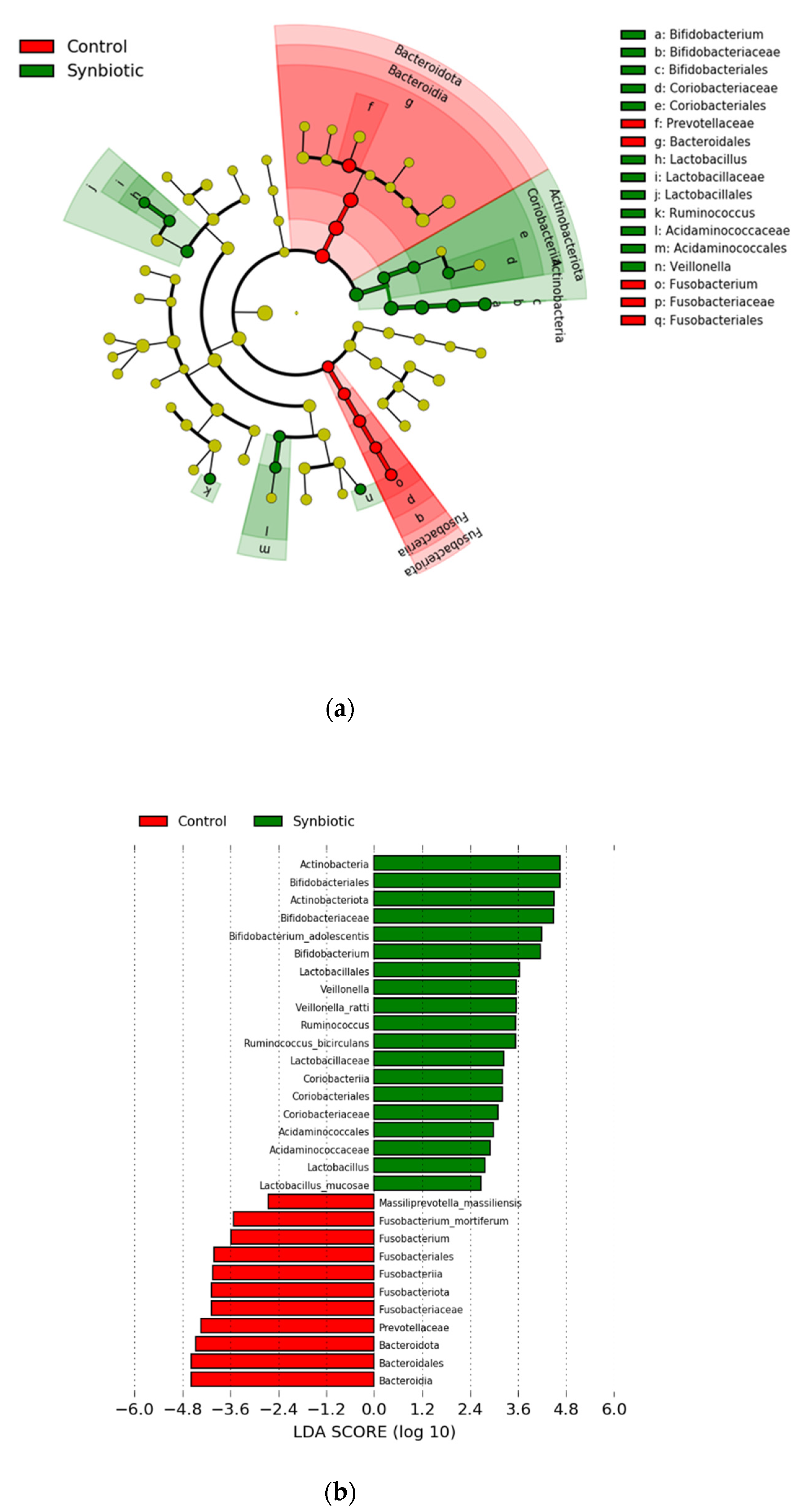
P < 0.05 was considered statistically significant. For microbiota analysis, RT-qPCR-negative samples were analyzed using half values of the lower limit (logarithm) that each corresponding primer sets could detect. Then, for enumeration of LcS and BbrY in feces, qPCR-negative samples were excluded from the statistical analysis. Differences in the relative abundance of microbial features were determined by linear discriminant analysis (LDA) effect size (LEfSe) analysis using the Galaxy web application (
http://huttenhower.sph.harvard.edu/galaxy/) [
26]. Bacterial abundance profiles were calculated at taxonomic levels from phylum to species in terms of percent abundance, and a logarithmic LDA score ≥ 2.0 was used as a threshold.
4. Discussion
The present study assessed the effect of synbiotic administration on several inflammatory markers, including IL-6, LBP, and hs-CRP, in obese patients with T2DM. The results showed no differences in the levels of inflammatory markers between the synbiotic and control groups. So far, several studies have investigated the effects of probiotics/synbiotics on inflammatory markers in T2DM. While some found that IL-6 levels were reduced [
27,
28], others did not [
29,
30]. Thus, this issue remains controversial. The reasons for these inconsistencies remain unknown, but they may be due to study differences in probiotics/synbiotics as well as patient age, ethnicity, and eating habits.
In the present study, the effects of a synbiotic on the gut microbiota was investigated quantitatively and qualitatively using RT-qPCR and 16S rRNA amplicon analysis. The relative abundances of Bifidobacteriaceae was significantly increased in feces after synbiotic administration. Of the increased Bifidobacteriaceae, the two species of
Bifidobacterium adolescentis and
B. pseudocatenulatum were increased after synbiotic administration. Interestingly, metformin was reported to directly cause the growth of
B. adolescentis and also
A. muciniphila [
31]. Furthermore,
B. adolescentis is positively associated with GLP-1 secretion [
32] and exhibits inhibitory activity against dipeptidyl peptidase-4 [
33]. Therefore, this bacterium might exert incretin-mediated and/or unknown antidiabetic effects via metformin. In addition, some strains of
B. pseudocatenulatum have beneficial effects on inflammation [
34] and metabolism [
35]. Taken together, the bacteria that showed an increase in relative abundance in response to synbiotic administration might play important roles in glucose metabolism.
In our previous research using only LcS as a probiotic [
8], no changes in fecal organic acids were found in T2DM, but in the present study, which instead used a synbiotic, the fecal acetic acid concentration was significantly increased. It was reported that endogenous
Bifidobacterium belonging to the predominant obligate anaerobes in humans can grow using the GOS in synbiotics and efficiently produce acetic acids as well as BbrY [
36,
37]. Therefore, it is considered that GOS might be a key regulator in increasing fecal acetic acids, with BbrY playing a secondary role. It was also demonstrated that short chain fatty acids, especially acetic acid, can improve the function of the gut barrier [
38]. Furthermore, Kimura et al. reported that acetic acid promoted glucose metabolism via the activation of G protein-coupled receptor 43, which suppresses insulin signaling in adipocytes [
39]. Therefore, increasing the production of acetic acid might also play an important role in glycemic control.
The counts of two lactobacilli genera, specifically the
Lactobacillus and
Lacticaseibacillus, also increased after synbiotic administration. This was not surprising for
Lacticaseibacillus, since it was administered as the LcS probiotic, but that was not the case for
Lactobacillus.
L. gasseri is a microorganism that is vaginally transmitted from mother to infant at birth [
40], and is considered one of the primary microbiota to be involved in GOS fermentation [
41]. In this study, therefore,
L. gasseri may have utilized the GOS in the synbiotic, leading to an increase in fecal count of
Lactobacillus. It has been reported that this bacterium has anti-pathogenic activity, for instance via the production of bacteriocin, and that it contributes to the maintenance of gut homeostasis [
42]. In addition, a previous study showed a positive correlation between HbA1c and the bacterial count of
Lactobacillus [
43], suggesting important roles of this subgroup in glycemic control. However, the precise mechanism remains unknown, and further studies to investigate the pathophysiological roles of these bacteria in T2DM are necessary.
B. vulgatus, which in this study demonstrated a decrease in bacterial count in response to synbiotic administration, was recently identified as the main species driving the association between biosynthesis of branched-chain amino acids and insulin resistance in obese patients [
2]. Therefore, a decrease in this bacterium following exposure to a synbiotic might play an important role in insulin resistance in obese patients with T2DM. Although this study suggested that glycemic control might be affected by changes in various gut bacteria, it did not improve after synbiotic administration. One reason may be that changes in glycemic control were difficult to evaluate because the mean HbA1c level at baseline was not very high (synbiotic group, HbA1c 7.4 ± 0.7%).
Interestingly, Bacteroidaceae abundance and
A. muciniphila count were decreased in this study. These two bacteria are known as mucolytic bacteria [
44]. Therefore, it was suggested that endogenous mucolytic Bacteroidaceae and
A. muciniphila were relatively reduced due to the increase of mucolytic
Bifidobacterium by synbiotic administration.
Our study has several limitations. First, since the study did not use a double-blind design, patients in the synbiotic group may have been aware of the effects of the synbiotic on the gut microbiota, which might have biased the results. Second, the study did not directly evaluate gut barrier function or plasma LPS levels. Therefore, it remains unknown whether synbiotic administration definitely reduced plasma LPS levels in T2DM. Third, the detection rate (2.4%) of gut bacteria in blood was very low compared with a previous study (22.0%) [
13]. Therefore, we could not evaluate the effects of the synbiotic on the translocation of live gut bacteria to the blood. Additionally, it is known that probiotic bacteria regulate intestinal permeability by tumor necrosis factor (TNF)-α-dependent mechanisms [
45]. However, in this study plasma levels of TNF-α were not evaluated. Therefore, the evaluation will be necessary for future study.