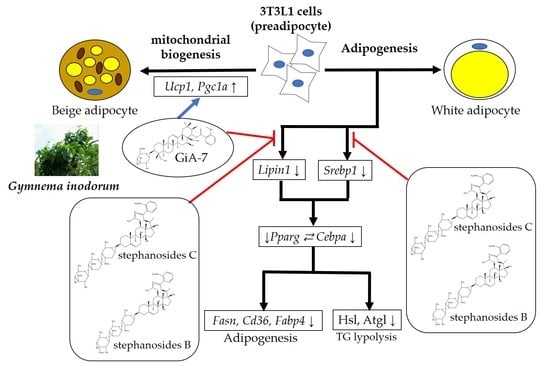
Purified Gymnemic Acids from Gymnema inodorum Tea Inhibit 3T3-L1 Cell Differentiation into Adipocytes
Abstract
1. Introduction
2. Materials and Methods
2.1. Extraction, Isolation and Purification
2.2. Mass Spectrometry
2.3. Nuclear Magnetic Resonance (NMR) Spectroscopy
2.4. Cell Culture
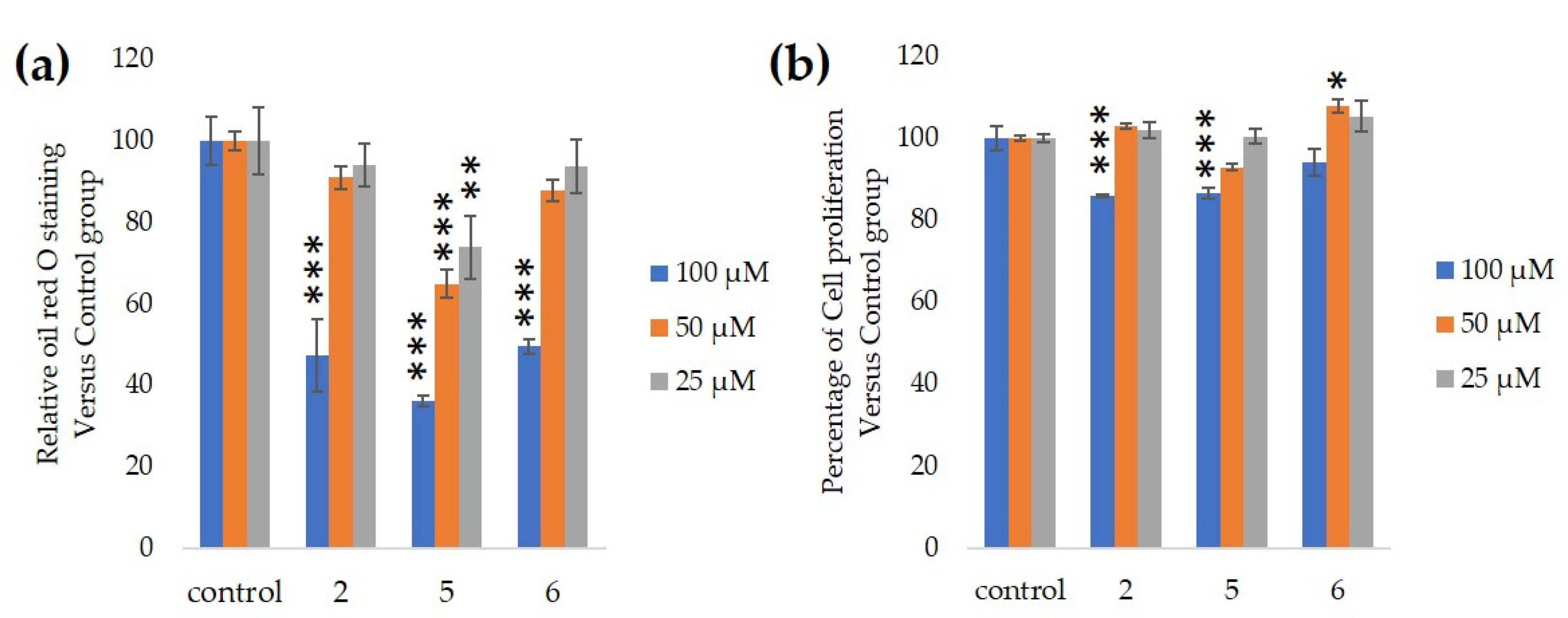
2.5. The Antiadipocyte Differentiation Activity
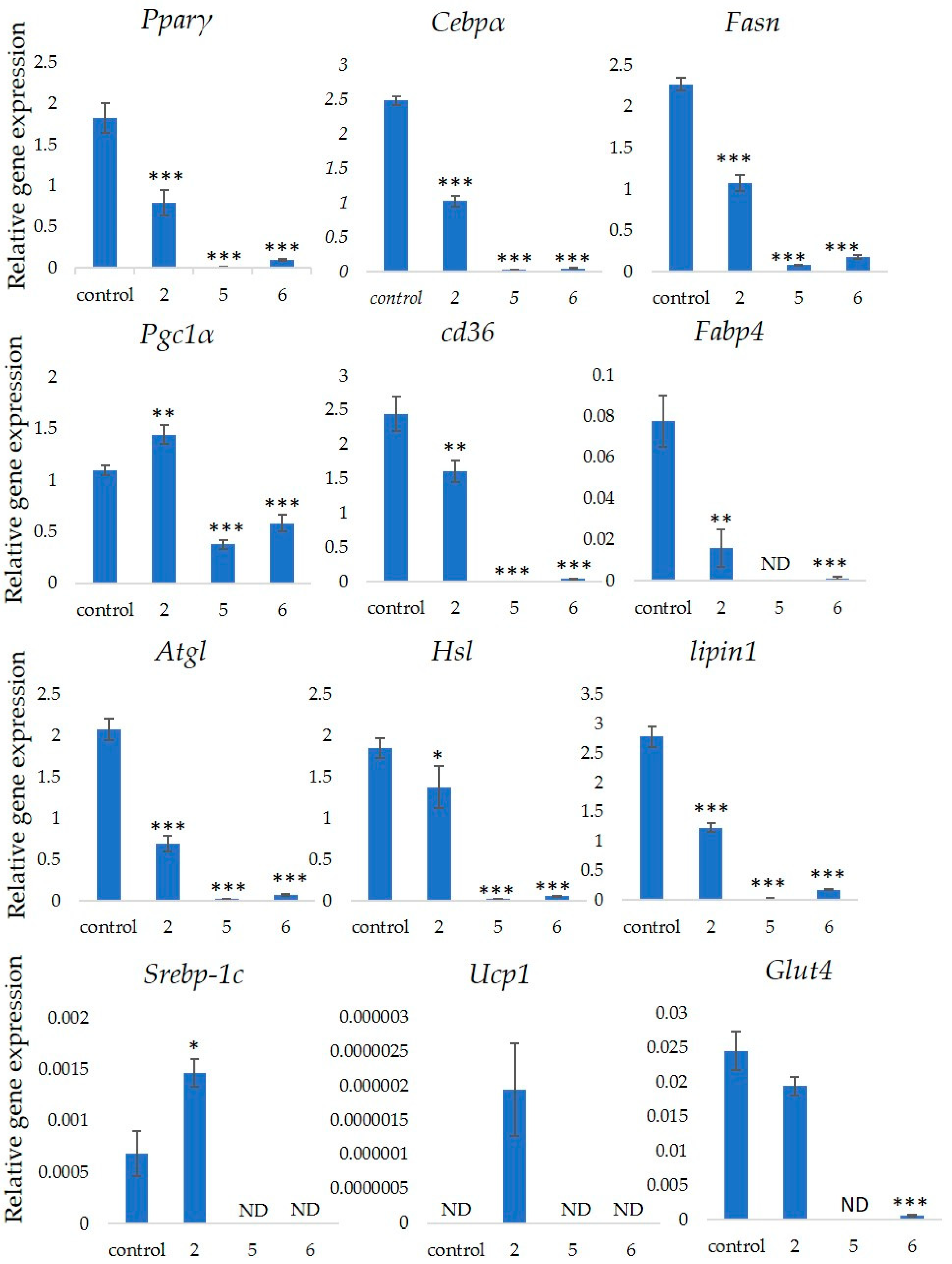
2.6. Quantitative Real-Time PCR
2.7. Statistical Analysis
3. Results and Discussion
4. Conclusions
5. Patents
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Shanmugasundaram, E.; Rajeswari, G.; Baskaran, K.; Kumar, B.R.; Shanmugasundaram, K.R.; Ahmath, B.K. Use of Gymnema sylvestre leaf extract in the control of blood glucose in insulin-dependent diabetes mellitus. J. Ethnopharmacol. 1990, 30, 281–294. [Google Scholar] [CrossRef]
- Shimizu, K.; Iino, A.; Nakajima, J.; Tanaka, K.; Nakajyo, S.; Urakawa, N.; Atsuchi, M.; Wada, T.; Yamashita, C. Suppression of glucose absorption by some fractions extracted from Gymnema sylvestre leaves. J. Vet. Med. Sci. 1997, 59, 245–251. [Google Scholar] [CrossRef] [PubMed]
- Al-Romaiyan, A.; King, A.; Persaud, S.; Jones, P. A novel extract of Gymnema sylvestre improves glucose tolerance in vivo and stimulates insulin secretion and synthesis in vitro. Phytother. Res. 2013, 27, 1006–1011. [Google Scholar] [CrossRef] [PubMed]
- Paliwal, R.; Kathori, S.; Upadhyay, B. Effect of Gurmar (Gymnema sylvestre) powder intervention on the blood glucose levels among diabetics. Stud. Ethno-Med. 2009, 3, 133–135. [Google Scholar] [CrossRef]
- Kanetkar, P.; Singhal, R.; Kamat, M. Recent advances in indian herbal drug research guest editor: Thomas Paul Asir Devasagayam Gymnema sylvestre: A memoir. J. Clin. Biochem. Nutr. 2007, 41, 77–81. [Google Scholar] [CrossRef]
- Shimizu, K.; Ozeki, M.; Iino, A.; Nakajyo, S.; Urakawa, N.; Atsuchi, M. Structure-activity relationships of triterpenoid derivatives extracted from Gymnema inodorum leaves on glucose absorption. Jpn. J. Pharmacol. 2001, 86, 223–229. [Google Scholar] [CrossRef]
- Chiabchalard, A.; Tencomnao, T.; Santiyanont, R. Effect of Gymnema inodorum on postprandial peak plasma glucose levels in healthy human. Afr. J. Biotechnol. 2010, 9, 1079–1085. [Google Scholar]
- Shimizu, K.; Ozeki, M.; Tanaka, K.; Itoh, K.; Nakajyo, S.; Urakawa, N.; Atsuchi, M. Suppression of glucose absorption by extracts from the leaves of Gymnema inodorum. J. Vet. Med. Sci. 1997, 59, 753–757. [Google Scholar] [CrossRef] [PubMed]
- Klungsupya, P.; Muangman, T.; Theangtrong, N.; Khayungarnnawee, A.; Phatvej, W.; Thisayakorn, K.; Rerk-Am, U.; Sematong, T.; Trangvacharakul, S.; Arunpairojana, V. Antioxidant and antihyperglycemic activities of Gymnema inodorum Dence. In Proceedings of the 8th NRCT-JSPS Joint Seminar Innovative Research in Natural Products for Sustainable Development, Bangkok, Thailand, 3–5 February 2009; pp. 207–209. [Google Scholar]
- Rohilla, A.; Ali, S. Alloxan induced diabetes: Mechanisms and effects. Int. J. Res. Pharm. Biomed. Sci. 2012, 3, 819–823. [Google Scholar]
- Risérus, U.; Willett, W.C.; Hu, F.B. Dietary fats and prevention of type 2 diabetes. Prog. Lipid Res. 2009, 48, 44–51. [Google Scholar] [CrossRef]
- Zeyda, M.; Stulnig, T.M. Obesity, inflammation, and insulin resistance—A mini-review. Gerontology 2009, 55, 379–386. [Google Scholar] [CrossRef] [PubMed]
- Hotamisligil, G.S. Inflammation and metabolic disorders. Nature 2006, 444, 860–867. [Google Scholar] [CrossRef] [PubMed]
- Farmer, S.R. Molecular determinants of brown adipocyte formation and function. Genes Dev. 2008, 22, 1269–1275. [Google Scholar] [CrossRef] [PubMed]
- Rosen, E.D.; MacDougald, O.A. Adipocyte differentiation from the inside out. Nat. Rev. Mol. Cell Biol. 2006, 7, 885–896. [Google Scholar] [CrossRef] [PubMed]
- Green, H.; Meuth, M. An established pre-adipose cell line and its differentiation in culture. Cell 1974, 3, 127–133. [Google Scholar] [CrossRef]
- Rubin, C.S.; Hirsch, A.; Fung, C.; Rosen, O.M. Development of hormone receptors and hormonal responsiveness in vitro. Insulin receptors and insulin sensitivity in the preadipocyte and adipocyte forms of 3T3-L1 cells. J. Biol. Chem. 1978, 253, 7570–7578. [Google Scholar]
- Taira, J.; Ogi, T. Induction of Antioxidant Protein HO-1 Through Nrf2-ARE Signaling Due to Pteryxin in Peucedanum Japonicum Thunb in RAW264. 7 Macrophage Cells. Antioxidants 2019, 8, 621. [Google Scholar] [CrossRef]
- Oishi, K.; Ohyama, S.; Higo-Yamamoto, S. Chronic sleep disorder induced by psychophysiological stress induces glucose intolerance without adipose inflammation in mice. Biochem. Biophys. Res. Commun. 2018, 495, 2616–2621. [Google Scholar] [CrossRef]
- Kanda, Y. Investigation of the freely available easy-to-use software “EZR” for medical statistics. Bone Marrow Transpl. 2013, 48, 452–458. [Google Scholar] [CrossRef] [PubMed]
- Yoshikawa, K.; Okada, N.; Kann, Y.; Arihara, S. Steroidal glycosides from the fresh stem of Stephanotis lutchuensis var. japonica (Asclepiadaceae). Chemical structures of stephanosides AJ. Chem. Pharm. Bull. 1996, 44, 1790–1796. [Google Scholar] [CrossRef][Green Version]
- Drolet, R.; Richard, C.; Sniderman, A.D.; Mailloux, J.; Fortier, M.; Huot, C.; Rheaume, C.; Tchernof, A. Hypertrophy and hyperplasia of abdominal adipose tissues in women. Int. J. Obes. (Lond.) 2008, 32, 283–291. [Google Scholar] [CrossRef]
- Xie, M.; Roy, R. AMP-Activated Kinase Regulates Lipid Droplet Localization and Stability of Adipose Triglyceride Lipase in C. elegans Dauer Larvae. PLoS ONE 2015, 10, e0130480. [Google Scholar] [CrossRef] [PubMed]
- Jiang, D.; Wang, D.; Zhuang, X.; Wang, Z.; Ni, Y.; Chen, S.; Sun, F. Berberine increases adipose triglyceride lipase in 3T3-L1 adipocytes through the AMPK pathway. Lipids Health Dis. 2016, 15, 214. [Google Scholar] [CrossRef]
- Deng, T.; Shan, S.; Li, P.P.; Shen, Z.F.; Lu, X.P.; Cheng, J.; Ning, Z.Q. Peroxisome proliferator-activated receptor-gamma transcriptionally up-regulates hormone-sensitive lipase via the involvement of specificity protein-1. Endocrinology 2006, 147, 875–884. [Google Scholar] [CrossRef] [PubMed]
- Kershaw, E.E.; Schupp, M.; Guan, H.-P.; Gardner, N.P.; Lazar, M.A.; Flier, J.S. PPARγ regulates adipose triglyceride lipase in adipocytes in vitro and in vivo. Am. J. Physiol. Endocrinol. Metab. 2007, 293, E1736–E1745. [Google Scholar] [CrossRef]
- Sembongi, H.; Miranda, M.; Han, G.S.; Fakas, S.; Grimsey, N.; Vendrell, J.; Carman, G.M.; Siniossoglou, S. Distinct roles of the phosphatidate phosphatases lipin 1 and 2 during adipogenesis and lipid droplet biogenesis in 3T3-L1 cells. J. Biol. Chem. 2013, 288, 34502–34513. [Google Scholar] [CrossRef]
- Kim, H.E.; Bae, E.; Jeong, D.-Y.; Kim, M.-J.; Jin, W.-J.; Park, S.-W.; Han, G.-S.; Carman, G.M.; Koh, E.; Kim, K.-S. Lipin1 regulates PPARγ transcriptional activity. Biochem. J. 2013, 453, 49–60. [Google Scholar] [CrossRef]
- Zhang, P.; Takeuchi, K.; Csaki, L.S.; Reue, K. Lipin-1 phosphatidic phosphatase activity modulates phosphatidate levels to promote peroxisome proliferator-activated receptor gamma (PPARgamma) gene expression during adipogenesis. J. Biol. Chem. 2012, 287, 3485–3494. [Google Scholar] [CrossRef] [PubMed]
- Phan, J.; Peterfy, M.; Reue, K. Lipin expression preceding peroxisome proliferator-activated receptor-gamma is critical for adipogenesis in vivo and in vitro. J. Biol. Chem. 2004, 279, 29558–29564. [Google Scholar] [CrossRef]
- Puigserver, P.; Spiegelman, B.M. Peroxisome proliferator-activated receptor-γ coactivator 1α (PGC-1α): Transcriptional coactivator and metabolic regulator. Endocr. Rev. 2003, 24, 78–90. [Google Scholar] [CrossRef]
- Kobayashi, M.; Fujii, N.; Narita, T.; Higami, Y. SREBP-1c-Dependent Metabolic Remodeling of White Adipose Tissue by Caloric Restriction. Int. J. Mol. Sci. 2018, 19, 3335. [Google Scholar] [CrossRef] [PubMed]
- Sharp, L.Z.; Shinoda, K.; Ohno, H.; Scheel, D.W.; Tomoda, E.; Ruiz, L.; Hu, H.; Wang, L.; Pavlova, Z.; Gilsanz, V. Human BAT possesses molecular signatures that resemble beige/brite cells. PLoS ONE 2012, 7, e49452. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Bostrom, P.; Sparks, L.M.; Ye, L.; Choi, J.H.; Giang, A.H.; Khandekar, M.; Virtanen, K.A.; Nuutila, P.; Schaart, G.; et al. Beige adipocytes are a distinct type of thermogenic fat cell in mouse and human. Cell 2012, 150, 366–376. [Google Scholar] [CrossRef] [PubMed]
- Cannon, B.; Nedergaard, J. Neither brown nor white. Nature 2012, 488, 286–287. [Google Scholar] [CrossRef] [PubMed]
- Sugihara, Y.; Nojima, H.; Matsuda, H.; Murakami, T.; Yoshikawa, M.; Kimura, I. Antihyperglycemic effects of gymnemic acid IV, a compound derived from Gymnema sylvestre leaves in streptozotocin-diabetic mice. J. Asian Nat. Prod. Res. 2000, 2, 321–327. [Google Scholar] [CrossRef]
- Wang, L.; Luo, H.; Miyoshi, M.; Imoto, T.; Hiji, Y.; Sasaki, T. Inhibitory effect of gymnemic acid on intestinal absorption of oleic acid in rats. Can. J. Physiol. Pharmacol. 1998, 76, 1017–1023. [Google Scholar] [CrossRef]
- Liu, H.-M.; Kiuchi, F.; Tsuda, Y. Isolation and structure elucidation of gymnemic acids, antisweet principles of Gymnema sylvestre. Chem. Pharm. Bull. 1992, 40, 1366–1375. [Google Scholar] [CrossRef]
- Asare-Anane, H.; Huang, G.; Amiel, S.; Jones, P.; Persaud, S. Stimulation of insulin secretion by an aqueous extract of Gymnema sylvestre: Role of intracellular calcium. In Proceedings of the 196th Meeting of the Society for Endocrinology and Society for Endocrinology Joint Endocrinology and Diabetes Day, London, UK, 7–9 November 2005. [Google Scholar]
- Libman, A.; Zhang, H.; Ma, C.; Southavong, B.; Sydara, K.; Bouamanivong, S.; Tan, G.T.; Fong, H.H.; Soejarto, D.D. A first new antimalarial pregnane glycoside from Gongronema napalense. Asian J. Tradit. Med. 2008, 3, 203. [Google Scholar]
| Target Gene | Direction | Primer Sequence (5′–3′) |
|---|---|---|
| Pparγ | Forward | AACTCTGGGAGATTCTCCTGTTGA |
| Reverse | TGGTAATTTCTTGTGAAGTGCTCATA | |
| Fasn | Forward | GGAGGTGGTGATAGCCGGTAT |
| Reverse | TGGGTAATCCATAGAGCCCAG | |
| Cebpα | Forward | AAGAAGTCGGTGGACAAGAACAG |
| Reverse | GTTGCGTTGTTTGGCTTTATCTC | |
| Pgc1α | Forward | GTAGGCCCAGGTACGACAGC |
| Reverse | GCTCTTTGCGGTATTCATCCC | |
| Lipin-1 | Forward | CCATAGAGATGAGCTCGGAT |
| Reverse | AACTGGGATACGATGCTGACT | |
| Atgl | Forward | CTTGAGCAGCTAGAACAATG |
| Reverse | GGACACCTCAATAATGTTGGC | |
| Hsl | Forward | GCTGGAGGAGTGTTTTTTTGC |
| Reverse | AGTTGAACCAAGCAGGTCACA | |
| Srebp-1c | Forward | ATCGGCGCGGAAGCTGTCGGGGTAGCGTC |
| Reverse | ACTGTCTTGGTTGTTGATGAGCTGGAGCAT | |
| Glut4 | Forward | CTGTCGCTGGTTTCTCCAAC |
| Reverse | CAGGAGGACGGCAAATAGAA | |
| Ucp1 | Forward | GGCAACAAGAGCTGACAGTAAAT |
| Reverse | GGCCCTTGTAAACAACAAAATAC | |
| Fabp4 | Forward | CCGCAGACGACAGGA |
| Reverse | CTCATGCCCTTTCATAAACT | |
| 36b4 | Forward | CTTCATTGTGGGAGCAGACA |
| Reverse | TCTCCAGAGCTGGGTTGTTC |
| C-No. | Carbon Type | GiA-7 | Compound 2 |
|---|---|---|---|
| 1 | —CH2— | 39.7 | 39.7 |
| 2 | —CH2— | 26.2 | 26.4 |
| 3 | >CH—O— | 82.3 | 82.9 |
| 4 | >C< | 43.9 | 44.0 |
| 5 | >CH— | 48.1 | 48.1 |
| 6 | —CH2— | 18.8 | 18.9 |
| 7 | —CH2— | 33.2 | 33.2 |
| 8 | >C< | 41.2 | 41.3 |
| 9 | >CH— | 48.2 | 48.2 |
| 10 | >C< | 37.5 | 37.5 |
| 11 | —CH2— | 24.8 | 24.8 |
| 12 | —CH= | 124.9 | 125.0 |
| 13 | >C= | 142.8 | 142.8 |
| 14 | >C< | 43.9 | 44.0 |
| 15 | —CH2— | 37.0 | 37.0 |
| 16 | >CH—O— | 66.8 | 66.8 |
| 17 | >C< | 46.5 | 46.5 |
| 18 | >CH— | 44.9 | 44.9 |
| 19 | —CH2— | 47.1 | 47.1 |
| 20 | >C< | 33.0 | 33.1 |
| 21 | —CH2— | 39.9 | 39.9 |
| 22 | >CH— | 74.3 | 74.3 |
| 23 | —CH2—O— | 64.8 | 64.7 |
| 24 | —CH3 | 13.4 | 13.4 |
| 25 | —CH3 | 16.7 | 16.7 |
| 26 | —CH3 | 17.5 | 17.5 |
| 27 | —CH3 | 28.0 | 28.0 |
| 28 | —CH2—O— | 61.2 | 61.1 |
| 29 | —CH3 | 33.5 | 33.5 |
| 30 | —CH3 | 25.6 | 25.6 |
| O-NMAt | |||
| N1 | >C=O | 169.6 | 169.7 |
| N2 | >C= | 112.1 | 112.1 |
| N3 | >C= | 153.0 | 153.0 |
| N4 | —CH= | 111.9 | 112.0 |
| N5 | —CH= | 135.6 | 135.6 |
| N6 | —CH= | 115.3 | 115.4 |
| N7 | —CH= | 133.0 | 133.0 |
| N8 | —CH3 | 29.7 | 29.7 |
| β-glu | |||
| 1’ | —O—CH—O— | 105.3 | 105.7 |
| 2’ | >CH—O— | 75.0 | 75.2 |
| 3’ | >CH—O— | 78.0 | 78.0 |
| 4’ | >CH—O— | 73.5 | 73.5 |
| 5’ | >CH—O— | 76.6 | 78.0 |
| 6’ | —COO— |
| C-No. | Carbon Type | Stephanoside C | Compound 5 |
|---|---|---|---|
| 1 | —CH2— | 38.9 | 38.9 |
| 2 | —CH2— | 30.0 | 30.0 |
| 3 | >CH—O— | 77.8 | 77.7 |
| 4 | —CH2— | 39.3 | 39.4 |
| 5 | >C= | 139.3 | 139.4 |
| 6 | —CH= | 119.5 | 119.5 |
| 7 | —CH2— | 35.0 | 35.0 |
| 8 | >C< | 74.4 | 74.4 |
| 9 | >CH— | 44.1 | 44.2 |
| 10 | >C< | 37.3 | 37.4 |
| 11 | —CH2— | 25.7 | 25.7 |
| 12 | >CH— | 74.7 | 74.6 |
| 13 | >C< | 57.0 | 57.0 |
| 14 | >C< | 89.0 | 89.0 |
| 15 | —CH2— | 33.8 | 33.8 |
| 16 | —CH2— | 34.0 | 34.0 |
| 17 | >C< | 87.7 | 87.7 |
| 18 | —CH3 | 11.4 | 11.4 |
| 19 | —CH3 | 18.1 | 18.1 |
| 20 | >CH—O— | 75.0 | 75.0 |
| 21 | —CH3 | 15.6 | 15.7 |
| 12-O-Acetyl moiety | |||
| A1 | —COO— | 171.5 | 171.4 |
| A2 | —CH3 | 22.1 | 22.1 |
| 20-O-N-Methylanthraniloyl moiety | |||
| N1 | —COO— | 111.0 | 111.1 |
| N2 | >C= | 152.7 | 152.7 |
| N3 | >C= | 111.6 | 111.6 |
| N4 | CH= | 135.1 | 135.2 |
| N5 | —CH= | 114.8 | 114.8 |
| N6 | —CH= | 132.7 | 132.7 |
| N7 | —CH= | 168.3 | 168.3 |
| N8 | —CH3 | 29.7 | 29.6 |
| C-No. | Carbon Type | Stephanoside C | Compound 5 |
|---|---|---|---|
| d-Cymarose | |||
| 1′ | —O—CH—O— | 96.5 | 96.5 |
| 2′ | —CH2— | 37.3 | 37.4 |
| 3′ | >CH—O— | 78.0 | 78.0 |
| 4′ | >CH—O— | 83.5 | 83.5 |
| 5′ | >CH—O— | 69.1 | 69.1 |
| 6′ | —CH3 | 18.7 | 18.7 |
| O-Me | —O—CH3 | 59.0 | 59.0 |
| d-Olenadrose | |||
| 1′′ | —O—CH—O— | 102.2 | 102.1 |
| 2′′ | —CH2— | 37.7 | 37.8 |
| 3′′ | >CH—O— | 79.3 | 79.3 |
| 4′′ | >CH—O— | 83.2 | 83.2 |
| 5′′ | >CH—O— | 72.1 | 72.2 |
| 6′′ | —CH3 | 19.0 | 19.0 |
| O-Me | —O—CH3 | 57.4 | 57.4 |
| d-Thevetose | |||
| 1′′′ | ―O—CH—O— | 104.2 | 104.3 |
| 2′′′ | >CH—O— | 75.3 | 75.4 |
| 3′′′ | >CH—O— | 88.2 | 88.3 |
| 4′′′ | >CH—O— | 76.1 | 76.1 |
| 5′′′ | >CH—O— | 72.9 | 73.0 |
| 6′′′ | —CH3 | 18.8 | 18.8 |
| O-Me | —O—CH3 | 61.1 | 61.1 |
| C-No. | Carbon Type | Stephanoside B | Compound 6 |
|---|---|---|---|
| 1 | —CH2— | 38.8 | 38.8 |
| 2 | —CH2— | 30.0 | 29.9 |
| 3 | >CH—O— | 77.7 | 77.6 |
| 4 | —CH2— | 39.3 | 39.2 |
| 5 | >C= | 139.3 | 139.2 |
| 6 | —CH= | 119.4 | 119.4 |
| 7 | —CH2— | 34.9 | 34.9 |
| 8 | >C< | 74.3 | 74.3 |
| 9 | >CH— | 44.1 | 44.0 |
| 10 | >C< | 37.3 | 37.2 |
| 11 | —CH2— | 25.6 | 25.6 |
| 12 | >CH— | 74.6 | 74.6 |
| 13 | >C< | 56.9 | 56.9 |
| 14 | >C< | 88.9 | 88.9 |
| 15 | —CH2— | 33.8 | 33.7 |
| 16 | —CH2— | 33.9 | 33.9 |
| 17 | >C< | 87.6 | 87.6 |
| 18 | —CH3 | 11.3 | 11.3 |
| 19 | —CH3 | 18.1 | 18.0 |
| 20 | >CH—O— | 74.9 | 74.9 |
| 21 | —CH3 | 15.6 | 15.6 |
| 12-O-Acetyl moiety | |||
| A1 | —COO— | 171.3 | 171.3 |
| A2 | —CH3 | 22.0 | 22.1 |
| 20-O-N-Methylanthraniloyl moiety | |||
| N1 | —COO— | 111.0 | 111.0 |
| N2 | >C= | 152.6 | 152.6 |
| N3 | >C= | 111.5 | 111.5 |
| N4 | —CH= | 135.1 | 135.1 |
| N5 | —CH= | 114.7 | 114.7 |
| N6 | —CH= | 132.6 | 132.6 |
| N7 | —CH= | 168.2 | 168.2 |
| N8 | —CH2 | 29.6 | 29.5 |
| C-No. | Carbon Type | Stephanoside B | Compound 6 |
|---|---|---|---|
| d-Cymarose | |||
| 1′ | —O—CH—O— | 96.4 | 96.4 |
| 2′ | —CH2— | 37.3 | 37.2 |
| 3′ | >CH—O— | 77.9 | 77.9 |
| 4′ | >CH—O— | 83.5 | 83.5 |
| 5′ | >CH—O— | 69.0 | 68.9 |
| 6′ | —CH3 | 18.7 | 18.7 |
| O—Me | —O—CH3 | 58.9 | 58.8 |
| d-Olenadrose | |||
| 1′′ | —O—CH—O— | 101.9 | 101.9 |
| 2′′ | —CH2— | 37.5 | 37.6 |
| 3′′ | >CH—O— | 79.3 | 79.2 |
| 4′′ | >CH—O— | 82.8 | 82.9 |
| 5′′ | >CH—O— | 72.0 | 72.0 |
| 6′′ | —CH3 | 19.0 | 18.9 |
| O-Me | —O—CH3 | 57.2 | 57.2 |
| d-Allomethylose | |||
| 1‴ | —O—CH—O— | 102.2 | 102.1 |
| 2‴ | >CH—O— | 73.2 | 73.3 |
| 3‴ | >CH—O— | 84.0 | 84.1 |
| 4‴ | >CH—O— | 74.6 | 74.5 |
| 5‴ | >CH—O— | 71.0 | 71.0 |
| 6‴ | —CH3 | 18.7 | 18.7 |
| O-Me | —O—CH3 | 62.1 | 62.1 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Saiki, P.; Kawano, Y.; Ogi, T.; Klungsupya, P.; Muangman, T.; Phantanaprates, W.; Kongchinda, P.; Pinnak, N.; Miyazaki, K. Purified Gymnemic Acids from Gymnema inodorum Tea Inhibit 3T3-L1 Cell Differentiation into Adipocytes. Nutrients 2020, 12, 2851. https://doi.org/10.3390/nu12092851
Saiki P, Kawano Y, Ogi T, Klungsupya P, Muangman T, Phantanaprates W, Kongchinda P, Pinnak N, Miyazaki K. Purified Gymnemic Acids from Gymnema inodorum Tea Inhibit 3T3-L1 Cell Differentiation into Adipocytes. Nutrients. 2020; 12(9):2851. https://doi.org/10.3390/nu12092851
Chicago/Turabian StyleSaiki, Papawee, Yasuhiro Kawano, Takayuki Ogi, Prapaipat Klungsupya, Thanchanok Muangman, Wimonsri Phantanaprates, Papitchaya Kongchinda, Nantaporn Pinnak, and Koyomi Miyazaki. 2020. "Purified Gymnemic Acids from Gymnema inodorum Tea Inhibit 3T3-L1 Cell Differentiation into Adipocytes" Nutrients 12, no. 9: 2851. https://doi.org/10.3390/nu12092851
APA StyleSaiki, P., Kawano, Y., Ogi, T., Klungsupya, P., Muangman, T., Phantanaprates, W., Kongchinda, P., Pinnak, N., & Miyazaki, K. (2020). Purified Gymnemic Acids from Gymnema inodorum Tea Inhibit 3T3-L1 Cell Differentiation into Adipocytes. Nutrients, 12(9), 2851. https://doi.org/10.3390/nu12092851