The Development of High-Quality Multispecies Probiotic Formulations: From Bench to Market
Abstract
1. Introduction: Probiotics and the Role of Microorganisms in Human Health
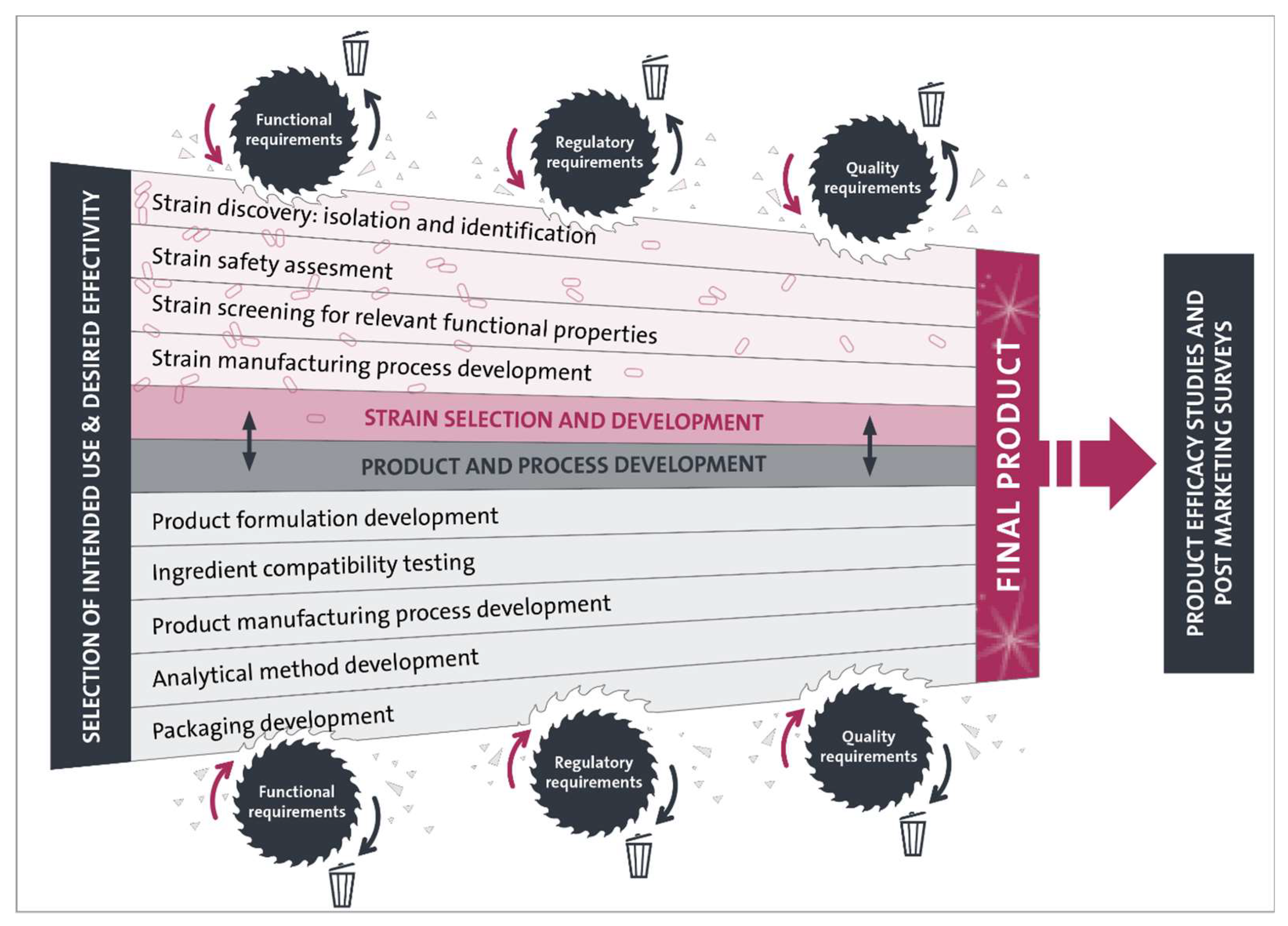
2. Probiotic Strain Selection and Development
2.1. The Concept of Evidence-Based, Indication-Specific and Multispecies Probiotics
2.2. Characterization and Functional Screening of (Candidate) Probiotic Strains
2.3. Identification and Safety Assessment of (Candidate) Probiotic Strains
2.4. Strain Manufacturing Process Development
3. Important Aspects to Consider during Product Formulation and Manufacturing Process Development
3.1. Product Formulation and Manufacturing Process Development
3.2. Quality Properties of Probiotic Formulations
3.2.1. Viability Throughout Shelf Life (Stability)
3.2.2. Gastrointestinal Survival
3.2.3. Activity
3.3. Extensive Quality Monitoring Throughout the Manufacturing Process
3.3.1. Viability and Composition Validation
3.3.2. Safety and Quality Monitoring
4. Clinical Efficacy of a Probiotic Product
4.1. Antibiotic-Associated Diarrhea
4.2. The Infantile Microbiome and Its Effect on the Maturation of the Immune System
4.3. Metabolic Diseases
4.4. Probiotic Impact on the Gut–Brain Axis
4.5. Liver Diseases
4.6. Conclusions and Outlook
Author Contributions
Funding
Conflicts of Interest
References
- Lynch, S.V.; Pedersen, O. The human intestinal microbiome in health and disease. N. Engl. J. Med. 2016, 375, 2369–2379. [Google Scholar] [CrossRef] [PubMed]
- Hill, C.; Guarner, F.; Reid, G.; Gibson, G.R.; Merenstein, D.J.; Pot, B.; Morelli, L.; Canani, R.B.; Flint, H.J.; Salminen, S.; et al. Expert consensus document: The international scientific association for probiotics and prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat. Rev. Gastroenterol. Hepatol. 2014, 11, 506–514. [Google Scholar] [CrossRef] [PubMed]
- Alonso, V.R.; Guarner, F. Linking the gut microbiota to human health. Br. J. Nutr. 2013, 109. [Google Scholar] [CrossRef]
- Gerritsen, J.; Smidt, H.; Rijkers, G.T.; de Vos, W.M. Intestinal microbiota in human health and disease: The impact of probiotics. Genes Nutr. 2011, 6, 209–240. [Google Scholar] [CrossRef]
- De Vos, W.M.; de Vos, E.A.J. Role of the intestinal microbiome in health and disease: From correlation to causation. Nutr. Rev. 2012, 70. [Google Scholar] [CrossRef]
- Plaza-Diaz, J.; Ruiz-Ojeda, F.J.; Gil-Campos, M.; Gil, A. Mechanisms of Action of Probiotics. Adv. Nutr. 2019. [Google Scholar] [CrossRef]
- Gareau, M.G.; Sherman, P.M.; Walker, W.A. Probiotics and the gut microbiota in intestinal health and disease. Nat. Rev. Gastroenterol. Hepatol. 2010, 7, 503–514. [Google Scholar] [CrossRef]
- Kankainen, M.; Paulin, L.; Tynkkynen, S.; von Ossowski, I.; Reunanen, J.; Partanen, P.; Satokari, R.; Vesterlund, S.; Hendrickx, A.P.A.; Lebeer, S.; et al. Comparative genomic analysis of Lactobacillus rhamnosus GG reveals pili containing a human-mucus binding protein. Proc. Natl. Acad. Sci. USA 2009, 106, 17193–17198. [Google Scholar] [CrossRef]
- Papadimitriou, K.; Zoumpopoulou, G.; Foligné, B.; Alexandraki, V.; Kazou, M.; Pot, B.; Tsakalidou, E. Discovering probiotic microorganisms: In vitro, in vivo, genetic and omics approaches. Front. Microbiol. 2015, 6. [Google Scholar] [CrossRef]
- Pearce, S.C.; Coia, H.G.; Karl, J.P.; Pantoja-Feliciano, I.G.; Zachos, N.C.; Racicot, K. Intestinal in vitro and ex vivo models to study host-microbiome interactions and acute stressors. Front. Physiol. 2018, 9. [Google Scholar] [CrossRef]
- Younes, J.A.; Lievens, E.; Hummelen, R.; van der Westen, R.; Reid, G.; Petrova, M.I. Women and Their Microbes: The Unexpected Friendship. Trends Microbiol. 2018, 26, 16–32. [Google Scholar] [CrossRef] [PubMed]
- Lebeer, S.; Vanderleyden, J.; de Keersmaecker, S.C.J. Genes and Molecules of Lactobacilli Supporting Probiotic Action. Microbiol. Mol. Biol. Rev. 2008, 72, 728–764. [Google Scholar] [CrossRef] [PubMed]
- Coman, M.M.; Verdenelli, M.C.; Cecchini, C.; Silvi, S.; Orpianesi, C.; Boylo, N.; Cresci, A. In vitro evaluation of antimicrobial activity of Lactobacillus rhamnosus IMC 501®, Lactobacillus paracasei IMC 502® and SYNBIO® against pathogens. J. Appl. Microbiol. 2014, 117, 518–527. [Google Scholar] [CrossRef] [PubMed]
- Tomás, M.S.J.; Ocaña, V.S.; Wiese, B.; Nader-Macías, M.E. Growth and lactic acid production by vaginal Lactobacillus acidophilus CRL 1259, and inhibition of uropathogenic Escherichia coli. J. Med. Microbiol. 2003, 52, 1117–1124. [Google Scholar] [CrossRef] [PubMed]
- Moens, F.; Weckx, S.; de Vuyst, L. Bifidobacterial inulin-type fructan degradation capacity determines cross-feeding interactions between bifidobacteria and Faecalibacterium prausnitzii. Int. J. Food Microbiol. 2016. [Google Scholar] [CrossRef] [PubMed]
- Van Den Abbeele, P.; Marzorati, M.; Derde, M.; De Weirdt, R.; Vermeiren, J.; Possemiers, S.; Van de Wiele, T. Arabinoxylans, inulin and Lactobacillus reuteri 1063 repress the adherent-invasive Escherichia coli from mucus in a mucosa-comprising gut model. npj Biofilms Microbiomes 2016, 2. [Google Scholar] [CrossRef]
- Ouwehand, A.C.; Salminen, S. In vitro Adhesion Assays for Probiotics and their in vivo Relevance: A Review. Microb. Ecol. Health Dis. 2003, 15, 175–184. [Google Scholar] [CrossRef]
- Atassi, F.; Brassart, D.; Grob, P.; Graf, F.; Servin, A.L. Vaginal Lactobacillus isolates inhibit uropathogenic Escherichia coli. FEMS Microbiol. Lett. 2006, 257, 132–138. [Google Scholar] [CrossRef]
- Osset, J.; Bartolomé, R.M.; García, E.; Andreu, A. Assessment of the capacity of Lactobacillus to inhibit the growth of uropathogens and block their adhesion to vaginal epithelial cells. J. Infect. Dis. 2001, 183, 485–491. [Google Scholar] [CrossRef]
- Roselli, M.; Finamore, A.; Britti, M.S.; Mengheri, E. Probiotic bacteria Bifidobacterium animalis MB5 and Lactobacillus rhamnosus GG protect intestinal Caco-2 cells from the inflammation-associated response induced by enterotoxigenic Escherichia coli K88. Br. J. Nutr. 2006, 95, 1177–1184. [Google Scholar] [CrossRef]
- Ussing, H.H.; Zerahn, K. Active Transport of Sodium as the Source of Electric Current in the Short-circuited Isolated Frog Skin. Acta Physiol. Scand. 1951, 23, 110–127. [Google Scholar] [CrossRef] [PubMed]
- Thomson, A.; Smart, K.; Somerville, M.S.; Lauder, S.N.; Appanna, G.; Horwood, G.; Raj, L.S.; Srivastava, B.; Durai, D.; Scurret, M.J.; et al. The Ussing chamber system for measuring intestinal permeability in health and disease. BMC Gastroenterol. 2019, 19, 98. [Google Scholar] [CrossRef] [PubMed]
- Shah, P.; Fritz, J.V.; Glaab, E.; Desai, M.S.; Greenhalgh, K.; Frachet, A.; Niegowska, M.; Estes, M.; Jäger, C.; Seguin-Devaux, C.; et al. A microfluidics-based in vitro model of the gastrointestinal human-microbe interface. Nat. Commun. 2016, 7, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.J.; Li, H.; Collins, J.J.; Ingber, D.E. Contributions of microbiome and mechanical deformation to intestinal bacterial overgrowth and inflammation in a human gut-on-a-chip. Proc. Natl. Acad. Sci. USA 2016, 113, E7–E15. [Google Scholar] [CrossRef] [PubMed]
- Douglas, A.E. Which experimental systems should we use for human microbiome science? PLoS Biol. 2018, 16, e2005245. [Google Scholar] [CrossRef]
- Kostic, A.D.; Howitt, M.R.; Garrett, W.S. Exploring host-microbiota interactions in animal models and humans. Genes Dev. 2013, 27, 701–718. [Google Scholar] [CrossRef]
- Kenny, D.J.; Plichta, D.R.; Vlamakis, H.; Balskus, E.P.; Correspondence, R.J.X. Cholesterol Metabolism by Uncultured Human Gut Bacteria Influences Host Cholesterol Level. Cell Host Microbe 2020. [Google Scholar] [CrossRef]
- Nguyen, M.; Long, S.W.; McDermott, P.F.; Olsen, R.J.; Olson, R.; Stevens, R.L.; Tyson, G.H.; Zhao, S.; Davis, J.J. Using machine learning to predict antimicrobial MICs and associated genomic features for nontyphoidal Salmonella. J. Clin. Microbiol. 2019, 57. [Google Scholar] [CrossRef]
- Armour, C.R.; Nayfach, S.; Pollard, K.S.; Sharpton, T.J. A Metagenomic Meta-analysis Reveals Functional Signatures of Health and Disease in the Human Gut Microbiome. mSystems 2019, 4. [Google Scholar] [CrossRef]
- Rosenberg, E.; De Long, E.F.; Thompson, F.; Lory, S.; Stackebrandt, E. The Prokaryotes: Prokaryotic Biology and Symbiotic Associations; Springer: Berlin/Heidelberg, Germany, 2013. [Google Scholar]
- Schleifer, K.H. Classification of Bacteria and Archaea: Past, present and future. Syst. Appl. Microbiol. 2009, 32, 533–542. [Google Scholar] [CrossRef]
- Tindall, B.J.; Rosselló-Móra, R.; Busse, H.J.; Ludwig, W.; Kämpfer, P. Notes on the characterization of prokaryote strains for taxonomic purposes. Int. J. Syst. Evol. Microbiol. 2010, 60, 249–266. [Google Scholar] [CrossRef] [PubMed]
- Sanders, M.E.; Merenstein, D.; Merrifield, C.A.; Hutkins, R. Probiotics for human use. Nutr. Bull. 2018, 43, 212–225. [Google Scholar] [CrossRef]
- Zheng, J.; Wittouck, S.; Salvetti, E.; Franz, C.M.A.P.; Harris, H.M.B.; Mattarelli, P.; O’Toole, P.W.; Pot, B.; Vandamme, P.; Walter, J.; et al. A taxonomic note on the genus Lactobacillus: Description of 23 novel genera, emended description of the genus Lactobacillus Beijerinck 1901, and union of Lactobacillaceae and Leuconostocaceae. Int. J. Syst. Evol. Microbiol. 2020, 70, 2782–2858. [Google Scholar] [CrossRef] [PubMed]
- Doron, S.; Snydman, D.R. Risk and Safety of Probiotics. Clin. Infect. Dis. 2015, 60, 129–134. [Google Scholar] [CrossRef]
- Kumar, H.; Salminen, S.; Verhagen, H.; Rowland, I.; Heimbach, J.; Bañares, S.; Young, T.; Nomoto, K.; Lalonde, M. Novel probiotics and prebiotics: Road to the market. Curr. Opin. Biotechnol. 2015, 32, 99–103. [Google Scholar] [CrossRef]
- Marteau, P. Safety aspects of probiotic products. Scand. J. Nutr. Naringsforsk. 2001, 45, 22–24. [Google Scholar] [CrossRef][Green Version]
- Miquel, S.; Beaumont, M.; Martín, R.; Langella, P.; Braesco, V.; Thomas, M. A proposed framework for an appropriate evaluation scheme for microorganisms as novel foods with a health claim in Europe. Microb. Cell Factories 2015, 14, 48. [Google Scholar] [CrossRef]
- Pariza, M.W.; Gillies, K.O.; Kraak-Ripple, S.F.; Leyer, G.; Smith, A.B. Determining the safety of microbial cultures for consumption by humans and animals. Regul. Toxicol. Pharmacol. 2015, 73, 164–171. [Google Scholar] [CrossRef]
- Salminen, S.; von Wright, A.; Morelli, L.; Marteau, P.; Brassart, D.; de Vos, W.M.; Fondén, R.; Saxelin, M.; Collins, K.; Mogensen, G.; et al. Demonstration of safety of probiotics—A review. Int. J. Food Microbiol. 1998, 44, 93–106. [Google Scholar] [CrossRef]
- Sanders, M.E.; Akkermans, L.M.A.; Haller, D.; Hammerman, C.; Heimbach, J.; Hörmannsperger, G.; Huys, G.; Levy, D.D.; Lutgendorff, F.; Mack, D.; et al. Safety assessment of probiotics for human use. Gut Microbes 2010, 1, 1–22. [Google Scholar] [CrossRef]
- Bringing a Probiotic-Containing Functional Food to the Market: Microbiological, Product, Regulatory and Labeling Issues—PubMed. Available online: https://pubmed.ncbi.nlm.nih.gov/10532385/ (accessed on 29 June 2020).
- Rychen, G.; Aquilina, G.; Azimonti, G.; Bampidis, V.; de Lourdes, B.M.; Bories, G.; Chesson, A.; Cocconcelli, P.S.; Flachowsky, G.; Gropp, J.; et al. Guidance on the characterisation of microorganisms used as feed additives or as production organisms. EFSA J. 2018, 16. [Google Scholar] [CrossRef]
- Public Consultation on the EFSA Statement on the Requirements for. Available online: https://www.efsa.europa.eu/en/consultations/call/public-consultation-efsa-statement-requirements-whole-genome (accessed on 29 June 2020).
- Chen, L.; Yang, J.; Yu, J.; Yao, Z.; Sun, L.; Shen, Y.; Jin, Q. VFDB: A reference database for bacterial virulence factors. Nucleic Acids Res. 2005, 33, D325–D328. [Google Scholar] [CrossRef] [PubMed]
- Guidance on the assessment of bacterial susceptibility to antimicrobials of human and veterinary importance. EFSA J. 2012, 10. [CrossRef]
- McArthur, A.G.; Waglechner, N.; Nizam, F.; Yan, A.; Azad, M.A.; Baylay, A.J.; Bhullar, K.; Canova, M.J.; De Pascale, G.; Ejim, L.; et al. The comprehensive antibiotic resistance database. Antimicrob. Agents Chemother. 2013, 57, 3348–3357. [Google Scholar] [CrossRef]
- Gupta, S.K.; Padmanabhan, B.R.; Diene, S.M.; Lopez-Rojas, R.; Kempf, M.; Landraud, L.; Rolain, J.-M. ARG-annot, a new bioinformatic tool to discover antibiotic resistance genes in bacterial genomes. Antimicrob. Agents Chemother. 2014, 58, 212–220. [Google Scholar] [CrossRef]
- Partridge, S.R.; Kwong, S.M.; Firth, N.; Jensen, S.O. Mobile genetic elements associated with antimicrobial resistance. Clin. Microbiol. Rev. 2018, 31. [Google Scholar] [CrossRef]
- Bourdichon, F.; Laulund, S.; Tenning, P. Inventory of microbial species with a rationale: A comparison of the IDF/EFFCA inventory of microbial food cultures with the EFSA Biohazard Panel qualified presumption of safety. FEMS Microbiol. Lett. 2019, 366. [Google Scholar] [CrossRef]
- Mogensen, G.; Salminen, S.; O’Brien, J.; Ouwehand, A.; Holzapfel, W.; Shortt, C.; Fondén, R.; Miller, G.D.; Donohue, D.; Playne, M.; et al. Food microorganisms—Health benefits, safety evaluation and strains with documented history of use in foods & Inventory of microorganisms with a documented history of use in food. Bull. IDF 2002, 377, 4–19. [Google Scholar]
- Generally Recognized as Safe (GRAS) Notification Program FDA. Available online: https://www.fda.gov/animal-veterinary/animal-food-feeds/generally-recognized-safe-gras-notification-program (accessed on 29 June 2020).
- Fenster, K.; Freeburg, B.; Hollard, C.; Wong, C.; Laursen, R.R.; Ouwehand, A. The Production and Delivery of Probiotics: A Review of a Practical Approach. Microorganisms 2019, 7, 83. [Google Scholar] [CrossRef]
- Jackson, S.A.; Schoeni, J.L.; Vegge, C.; Pane, M.; Stahl, B.; Bradley, M.; Goldman, V.S.; Burguière, P.; Atwater, J.B.; Sanders, M.E.; et al. Improving end-user trust in the quality of commercial probiotic products. Front. Microbiol. 2019, 10. [Google Scholar] [CrossRef]
- Hansen, S.J.Z.; Tang, P.; Kiefer, A.; Galles, K.; Wong, C.; Morovic, W. Droplet Digital PCR Is an Improved Alternative Method for High-Quality Enumeration of Viable Probiotic Strains. Front. Microbiol. 2010, 10. [Google Scholar] [CrossRef] [PubMed]
- Kiefer, A.; Tang, P.; Arndt, S.; Fallico, V.; Wong, C. Optimization of Viability Treatment Essential for Accurate Droplet Digital PCR Enumeration of Probiotics. Front. Microbiol. 2010, 11, 1811. [Google Scholar] [CrossRef]
- Ritchie, M.L.; Romanuk, T.N. A meta-analysis of probiotic efficacy for gastrointestinal diseases. PLoS ONE 2012, 7. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Shen, C.; Ma, L. Treatment efficacy of probiotics on atopic dermatitis, zooming in on infants: A systematic review and meta-analysis. Int. J. Dermatol. 2018, 57, 635–641. [Google Scholar] [CrossRef] [PubMed]
- Huang, R.; Wang, K.; Hu, J. Effect of probiotics on depression: A systematic review and meta-analysis of randomized controlled trials. Nutrients 2016, 8, 483. [Google Scholar] [CrossRef] [PubMed]
- Koning, C.J.M.; Jonkers, D.M.A.E.; Stobberingh, E.E.; Mulder, L.; Rombouts, F.M.; Stockbrügger, R.W. The effect of a multispecies probiotic on the intestinal microbiota and bowel movements in healthy volunteers taking the antibiotic amoxycillin. Am. J. Gastroenterol. 2008, 103, 178–189. [Google Scholar] [CrossRef]
- Hell, M.; Bernhofer, C.; Stalzer, P.; Kern, J.M.; Claassen, E. Probiotics in Clostridium difficile infection: Reviewing the need for a multistrain probiotic. Benef. Microbes 2013, 4, 39–51. [Google Scholar] [CrossRef]
- Van Wietmarschen, H.A.; Busch, M.; van Oostveen, A.; Pot, G.; Jong, M.C. Probiotics use for antibiotic-associated diarrhea: A pragmatic participatory evaluation in nursing homes. BMC Gastroenterol. 2020, 20, 151. [Google Scholar] [CrossRef]
- Niers, L.; Martín, R.; Rijkers, G.; Sengers, F.; Timmerman, H.; van Uden, N.; Smidt, H.; Kimpen, J.; Hoekstra, M. The effects of selected probiotic strains on the development of eczema (the PandA study). Allergy Eur. J. Allergy Clin. Immunol. 2009, 64, 1349–1358. [Google Scholar] [CrossRef]
- Hofmann, H. Probiotikum gegen Verdauungsbeschwerden in der Schwangerschaft und Säuglingskoliken. Gynäkologie Akt. 2015, 8–9. [Google Scholar] [CrossRef]
- Szulińska, M.; Łoniewski, I.; van Hemert, S.; Sobieska, M.; Bogdański, P. Dose-dependent effects of multispecies probiotic supplementation on the lipopolysaccharide (LPS) level and cardiometabolic profile in obese postmenopausal women: A 12-week randomized clinical trial. Nutrients 2018, 10, 773. [Google Scholar] [CrossRef]
- Sabico, S.; Al-Mashharawi, A.; Al-Daghri, N.M.; Yakout, S.; Alnaami, A.M.; Alokail, M.S.; McTernan, P.G. Effects of a multi-strain probiotic supplement for 12 weeks in circulating endotoxin levels and cardiometabolic profiles of medication naïve T2DM patients: A randomized clinical trial. J. Transl. Med. 2017, 15, 249. [Google Scholar] [CrossRef] [PubMed]
- Bagga, D.; Reichert, J.L.; Koschutnig, K.; Aigner, C.S.; Holzer, P.; Koskinen, K.; Moissl-Eichinger, C.; Schöpf, V. Probiotics drive gut microbiome triggering emotional brain signatures. Gut Microbes 2018. [Google Scholar] [CrossRef] [PubMed]
- Bagga, D.; Aigner, C.S.; Reichert, J.L.; Cecchetto, C.; Fischmeister, F.P.S.; Holzer, P.; Moissl-Eichinger, C.; Schöpf, V. Influence of 4-week multi-strain probiotic administration on resting-state functional connectivity in healthy volunteers. Eur. J. Nutr. 2018. [Google Scholar] [CrossRef] [PubMed]
- Reininghaus, E.Z.; Wetzlmair, L.-C.; Fellendorf, F.T.; Platzer, M.; Queissner, R.; Birner, A.; Pilz, R.; Hamm, C.; Maget, A.; Koidl, C.; et al. The Impact of Probiotic Supplements on Cognitive Parameters in Euthymic Individuals with Bipolar Disorder: A Pilot Study. Neuropsychobiology 2018, 1–8. [Google Scholar] [CrossRef]
- Leblhuber, F.; Steiner, K.; Schuetz, B.; Fuchs, D.; Gostner, J.M. Probiotic Supplementation in Patients with Alzheimer’s Dementia—An Explorative Intervention Study. Curr. Alzheimer Res. 2018. [Google Scholar] [CrossRef]
- Abildgaard, A.; Elfving, B.; Hokland, M.; Lund, S.; Wegener, G. Probiotic treatment protects against the pro-depressant-like effect of high-fat diet in Flinders Sensitive Line rats. Brain Behav. Immun. 2017, 65, 33–42. [Google Scholar] [CrossRef]
- Abildgaard, A.; Elfving, B.; Hokland, M.; Wegener, G.; Lund, S. Probiotic treatment reduces depressive-like behaviour in rats independently of diet. Psychoneuroendocrinology 2017, 79, 40–48. [Google Scholar] [CrossRef]
- Steenbergen, L.; Sellaro, R.; van Hemert, S.; Bosch, J.A.; Colzato, L.S. A randomized controlled trial to test the effect of multispecies probiotics on cognitive reactivity to sad mood. Brain Behav. Immun. 2015, 48, 258–264. [Google Scholar] [CrossRef]
- Chahwan, B.; Kwan, S.; Isik, A.; van Hemert, S.; Burke, C.; Roberts, L. Gut feelings: A randomised, triple-blind, placebo-controlled trial of probiotics for depressive symptoms. J. Affect. Disord. 2019, 253, 317–326. [Google Scholar] [CrossRef]
- Papalini, S.; Michels, F.; Kohn, N.; Wegman, J.; van Hemert, S.; Roelofs, K.; Arias-Vasquez, A.; Aarts, E. Stress matters: Randomized controlled trial on the effect of probiotics on neurocognition. Neurobiol. Stress 2019, 10, 100141. [Google Scholar] [CrossRef]
- Horvath, A.; Leber, B.; Schmerboeck, B.; Tawdrous, M.; Zettel, G.; Hartl, A.; Madl, T.; Stryeck, S.; Fuchs, D.; Lemesch, S.; et al. Randomised clinical trial: The effects of a multispecies probiotic vs. placebo on innate immune function, bacterial translocation and gut permeability in patients with cirrhosis. Aliment. Pharmacol. Ther. 2016, 44, 926–935. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Grumet, L.; Tromp, Y.; Stiegelbauer, V. The Development of High-Quality Multispecies Probiotic Formulations: From Bench to Market. Nutrients 2020, 12, 2453. https://doi.org/10.3390/nu12082453
Grumet L, Tromp Y, Stiegelbauer V. The Development of High-Quality Multispecies Probiotic Formulations: From Bench to Market. Nutrients. 2020; 12(8):2453. https://doi.org/10.3390/nu12082453
Chicago/Turabian StyleGrumet, Lukas, Yorick Tromp, and Verena Stiegelbauer. 2020. "The Development of High-Quality Multispecies Probiotic Formulations: From Bench to Market" Nutrients 12, no. 8: 2453. https://doi.org/10.3390/nu12082453
APA StyleGrumet, L., Tromp, Y., & Stiegelbauer, V. (2020). The Development of High-Quality Multispecies Probiotic Formulations: From Bench to Market. Nutrients, 12(8), 2453. https://doi.org/10.3390/nu12082453



