Direct-to-Consumer Nutrigenetics Testing: An Overview
Abstract
1. Introduction
2. Methods
2.1. Identification of Companies Offering Nutrigenetics DTC-GT
2.2. Analysis of Genes and Variants
3. Results
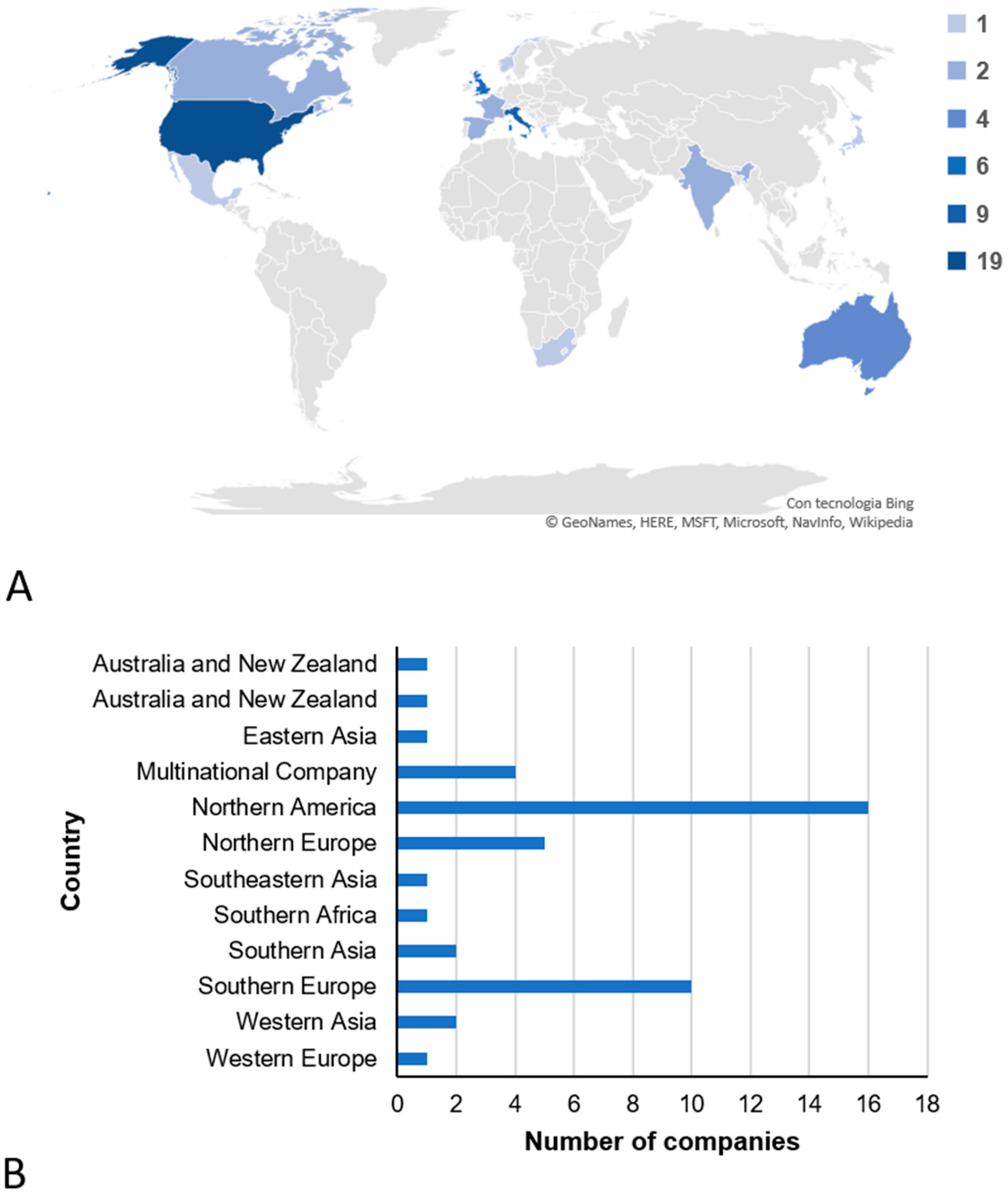
3.1. Companies That Offer Direct-to-Consumer Genetic Testing in the Field of Nutrigenetics
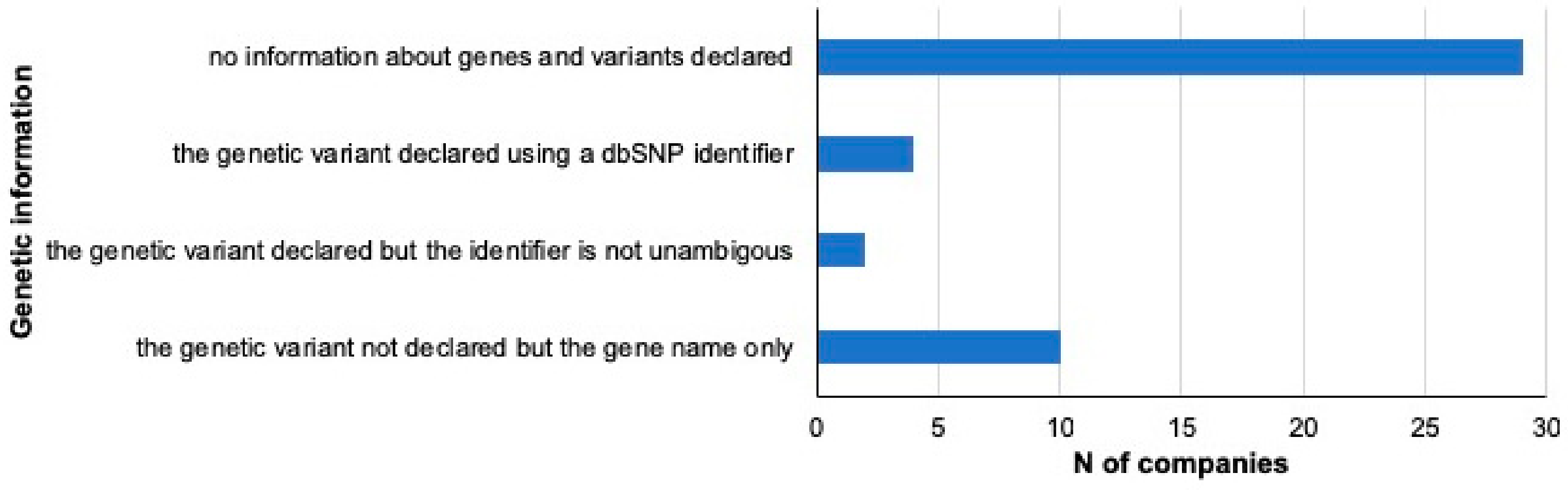
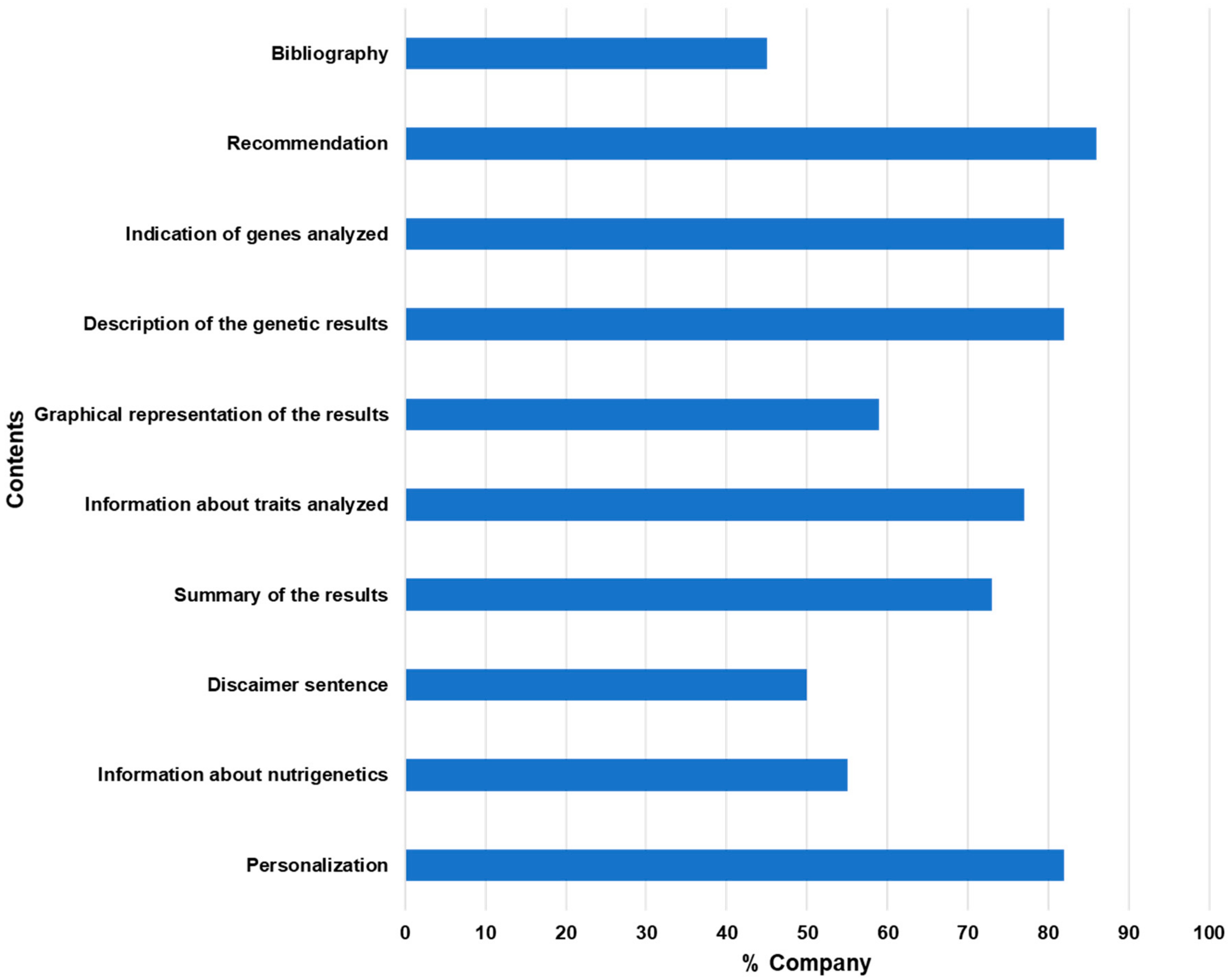
3.2. Traits and Genetic Information
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Crow, D. A New Wave of Genomics for All. Cell 2019, 177, 5–7. [Google Scholar] [CrossRef]
- Webborn, N.; Williams, A.; McNamee, M.; Bouchard, C.; Pitsiladis, Y.; Ahmetov, I.; Ashley, E.; Byrne, N.; Camporesi, S.; Collins, M.; et al. Direct-to-consumer genetic testing for predicting sports performance and talent identification: Consensus statement. Br. J. Sports Med. 2015, 49, 1486–1491. [Google Scholar] [CrossRef]
- Zhang, G.; Nebert, D.W. Personalized medicine: Genetic risk prediction of drug response. Pharmacol. Ther. 2017, 175, 75–90. [Google Scholar] [CrossRef] [PubMed]
- Guasch-Ferré, M.; Dashti, H.S.; Merino, J. Nutritional genomics and direct-to-consumer genetic testing: An overview. Adv. Nutr. 2018, 9, 128–135. [Google Scholar] [CrossRef] [PubMed]
- International Human Genome Sequencing Consortium. Finishing the euchromatic sequence of the human genome. Nature 2004, 431, 931–945. [Google Scholar] [CrossRef]
- dbSNP. Available online: https://www.ncbi.nlm.nih.gov/snp/ (accessed on 15 January 2020).
- Sherry, S.T.; Ward, M.H.; Kholodov, M.; Baker, J.; Phan, L.; Smigielski, E.M.; Sirotkin, K. dbSNP: The NCBI database of genetic variation. Horm. Res. 2001, 29, 308–311. [Google Scholar] [CrossRef] [PubMed]
- Landrum, M.J.; Lee, J.M.; Riley, G.R.; Jang, W.; Rubinstein, W.S.; Church, D.M.; Maglott, D.R. ClinVar: Public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 2014, 42, 980–985. [Google Scholar] [CrossRef]
- RefSNP Report. Available online: https://www.ncbi.nlm.nih.gov/snp/rs1801280 (accessed on 15 January 2020).
- Barbarino, J.M.; Whirl-Carrillo, M.; Altman, R.B.; Klein, T.E. PharmGKB: A worldwide resource for pharmacogenomic information. Wiley Interdiscip. Rev. Syst. Biol. Med. 2018, 10, 1–13. [Google Scholar] [CrossRef]
- Nyaga, D.M.; Vickers, M.H.; Perry, J.K.; O’Sullivan, J.M. The genetic architecture of type 1 diabetes mellitus. Mol. Cell. Endocrinol. 2018, 477, 70–80. [Google Scholar] [CrossRef]
- Cuyvers, E.; Sleegers, K. Genetic variations underlying Alzheimer’s disease: Evidence from genome-wide association studies and beyond. Lancet Neurol. 2016, 15, 857–868. [Google Scholar] [CrossRef]
- Marchini, J.; Howie, B. Genotype imputation for genome-wide association studies. Nat. Rev. Genet. 2010, 11, 499–511. [Google Scholar] [CrossRef] [PubMed]
- Tam, V.; Patel, N.; Turcotte, M.; Bossé, Y.; Paré, G.; Meyre, D. Benefits and limitations of genome-wide association studies. Nat. Rev. Genet. 2019, 20, 467–484. [Google Scholar] [CrossRef] [PubMed]
- Buniello, A.; Macarthur, J.A.L.; Cerezo, M.; Harris, L.W.; Hayhurst, J.; Malangone, C.; McMahon, A.; Morales, J.; Mountjoy, E.; Sollis, E.; et al. The NHGRI-EBI GWAS Catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res. 2019, 47, D1005–D1012. [Google Scholar] [CrossRef] [PubMed]
- GWAS Catalog. Available online: https://www.ebi.ac.uk/gwas/ (accessed on 22 September 2019).
- Zenin, A.; Tsepilov, Y.; Sharapov, S.; Getmantsev, E.; Menshikov, L.I.; Fedichev, P.O.; Aulchenko, Y. Identification of 12 genetic loci associated with human healthspan. Commun. Biol. 2019, 2, 41. [Google Scholar] [CrossRef] [PubMed]
- Costa, V.; Casamassimi, A.; Ciccodicola, A. Nutritional genomics era: Opportunities toward a genome-tailored nutritional regimen. J. Nutr. Biochem. 2010, 21, 457–467. [Google Scholar] [CrossRef] [PubMed]
- Loos, R.J.F. From nutrigenomics to personalizing diets: Are we ready for precision medicine? Am. J. Clin. Nutr. 2019, 109. [Google Scholar] [CrossRef]
- Wray, N.R.; Yang, J.; Hayes, B.J.; Price, A.L.; Goddard, M.E.; Visscher, P.M. A commentary on Pitfalls of predicting complex traits from SNPs. Nat. Rev. Genet. 2013, 14, 507–515. [Google Scholar] [CrossRef]
- Tenesa, A.; Haley, C.S. The heritability of human disease: Estimation, uses and abuses. Nat. Rev. Genet. 2013, 14, 139–149. [Google Scholar] [CrossRef]
- Moore, D.S.; Shenk, D. The heritability fallacy. Wiley Interdiscip. Rev. Cogn. Sci. 2016, 8. [Google Scholar] [CrossRef]
- Schrempft, S.; van Jaarsveld, C.H.M.; Fisher, A.; Herle, M.; Smith, A.D.; Fildes, A.; Llewellyn, C.H. Variation in the Heritability of Child Body Mass Index by Obesogenic Home Environment. JAMA Pediatr. 2018, 172, 1153–1160. [Google Scholar] [CrossRef]
- Duncan, L.; Shen, H.; Gelaye, B.; Meijsen, J.; Ressler, K.; Feldman, M.; Peterson, R.; Domingue, B. Analysis of polygenic risk score usage and performance in diverse human populations. Nat. Commun. 2019, 10, 3328. [Google Scholar] [CrossRef] [PubMed]
- Jordan, B. Chroniques génomiques Balayage du génome des personnes à risque. Médecine/Sciences. 2018, 34, 1116–1119. [Google Scholar] [CrossRef] [PubMed]
- Regalado, A. More than 26 Million People Have Taken an At-Home Ancestry Test. Mit. Technol. Rev. 2019. Available online: www.technologyreview.com/s/612880/more-than-26-million-people-have-taken-an-at-home-ancestry-test/ (accessed on 22 September 2019).
- Friend, L.; Rivilin, A.; O’Neil, J.; Browne, R. Direct-to-Consumer Genetic Testing: Opportunities and Risks in a Rapidly Evolving Market. KPMG International: 2018. Available online: https://assets.kpmg/content/dam/kpmg/xx/pdf/2018/08/direct-to-consumer-genetic-testing.pdf (accessed on 22 September 2019).
- Wood, L. Global Direct-to-Consumer Genetic Testing (DTC-GT) Market: Focus on Direct-to-Consumer Genetic Testing Market by Product Type, Distribution Channel, 15 Countries Mapping, and Competitive Landscape—Analysis and Forecast, 2019–2028. Research and Markets: 2019. Available online: https://www.globenewswire.com/newsrelease/2019/05/28/1851338/0/en/Global-6-3-Bn-Direct-to-Consumer-Genetic-Testing-DTC-GT-Market-to-2028.html (accessed on 22 September 2019).
- University College Dublin, National University of Ireland Dublin. Final Report Summary—FOOD4ME (Personalised Nutrition: An Integrated Analysis of Opportunities and Challenges). CORDIS EU Research Results: 2019. Available online: http://www.food4me.org/images/Food4MeWB-PRINT-14-05-15.pdf (accessed on 22 September 2019).
- Locke, A.E.; Kahali, B.; Berndt, S.I.; Justice, A.E.; Pers, T.H.; Day, F.R.; Powell, C.; Vedantam, S.; Buchkovich, M.L.; Yang, J.; et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 2015, 518, 197–206. [Google Scholar] [CrossRef] [PubMed]
- Loos, R.J.F.; Yeo, G.S.H. The bigger picture of FTO—The first GWAS-identified obesity gene. Nat. Rev. Endocrinol. 2014, 10, 51–61. [Google Scholar] [CrossRef]
- Yengo, L.; Sidorenko, J.; Kemper, K.E.; Zheng, Z.; Wood, A.R.; Weedon, M.N.; Frayling, T.M.; Hirschhorn, J.; Yang, J.; Visscher, P.M.; et al. Meta-analysis of genome-wide association studies for height and body mass index in ~700 000 individuals of European ancestry. Hum. Mol. Genet. 2018, 27, 3641–3649. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Ojeda, F.J.; Anguita-Ruiz, A.; Leis, R.; Aguilera, C.M. Genetic factors and molecular mechanisms of Vitamin D and obesity relationship. Ann. Nutr. Metab. 2018, 73, 89–99. [Google Scholar] [CrossRef] [PubMed]
- Basoli, V.; Santaniello, S.; Cruciani, S.; Ginesu, G.C.; Cossu, M.L.; Delitala, A.P.; Serra, P.A.; Ventura, C.; Maioli, M. Melatonin and vitamin D interfere with the adipogenic fate of adipose-derived stem cells. Int. J. Mol. Sci. 2017, 18, 981. [Google Scholar] [CrossRef] [PubMed]
- Cruciani, S.; Santaniello, S.; Garroni, G.; Fadda, A.; Balzano, F.; Bellu, E.; Sarais, G.; Fais, G.; Mulas, M.; Maioli, M. Myrtus polyphenols, from antioxidants to anti-inflammatory molecules: Exploring a network involving cytochromes P450 and Vitamin D. Molecules 2019, 24, 1515. [Google Scholar] [CrossRef] [PubMed]
- Santaniello, S.; Cruciani, S.; Basoli, V.; Balzano, F.; Bellu, E.; Garroni, G.; Ginesu, G.C.; Cossu, M.L.; Facchin, F.; Delitala, A.P.; et al. Melatonin and vitamin D orchestrate adipose derived stem cell fate by modulating epigenetic regulatory genes. Int. J. Med. Sci. 2018, 15, 1631–1639. [Google Scholar] [CrossRef]
- Jiang, X.; O’Reilly, P.F.; Aschard, H.; Hsu, Y.H.; Richards, J.B.; Dupuis, J.; Ingelsson, E.; Karasik, D.; Pilz, S.; Berry, D.; et al. Genome-wide association study in 79,366 European-ancestry individuals informs the genetic architecture of 25-hydroxyvitamin D levels. Nat. Commun. 2018, 9, 1–12. [Google Scholar] [CrossRef]
- Dudbridge, F. Power and Predictive Accuracy of Polygenic Risk Scores. PLoS Genet. 2013, 9, e003348. [Google Scholar] [CrossRef]
| Classification | N° Companies | |
|---|---|---|
| DTC-GT COMPANY | 45 | |
| NOT DTC-GT COMPANY | 18 | |
| DNA test services for hospital or specialized personnel | 8 | |
| Specialist web sites | 3 | |
| Multinational companies providing kit to specialists or specialized centers | 2 | |
| Biochemical analysis centres for patients | 2 | |
| Other | 3 | |
| Services and Information Provided | N° Companies | |
|---|---|---|
| DNA KIT | Provided | 41 |
| Purchase online | 35 | |
| Order by email or phone | 5 | |
| Not clear purchase modality | 1 | |
| Not Provided | 4 | |
| COST | Indicated | 36 |
| 0-100 $ | 9 | |
| 100-200 $ | 13 | |
| >200 $ | 14 | |
| Not Indicated | 9 | |
| REPORT | Provided | 22 |
| Not provided | 23 | |
| Category | Traits | N° of Companies |
|---|---|---|
| Food intolerance | ||
| Lactose intolerance | 18 | |
| Fructose intolerance | 2 | |
| Food Sensitivity | ||
| Caffeine sensitivity | 15 | |
| Alcohol sensitivity | 8 | |
| Salt Sensitivity | 5 | |
| Sulphites sensitivity | 1 | |
| Nickel sensitivity | 1 | |
| Saturated Fat Sensitivity | 3 | |
| Sensitivity and response to particular foods | 2 | |
| Refined carbohydrate Sensitivity | 1 | |
| Macronutrients | ||
| Lipid metabolism | 13 | |
| Carbohydrate metabolism | 6 | |
| Protein metabolism | 1 | |
| Whole grains metabolism | 1 | |
| Micronutrients | ||
| Vitamin D metabolism | 13 | |
| Vitamin C metabolism | 10 | |
| Vitamin A metabolism | 9 | |
| Vitamina B6 metabolism | 9 | |
| Vitamin B12 metabolism | 9 | |
| Vitamin B9 metabolism | 6 | |
| Vitamina E metabolism | 6 | |
| Iron metabolism | 4 | |
| Vitamina B metabolism | 4 | |
| Calcium metabolism | 2 | |
| Metabolism of vitamins and minerals | 1 | |
| Sodium metabolism | 1 | |
| Vitamina F metabolism | 1 | |
| Vitamin K metabolism | 1 | |
| Eating behaviour | ||
| Weight menagement | 12 | |
| Hunger and appetite control | 4 | |
| Regulation of metabolism and eating behavior | 3 | |
| Eating habits | 1 | |
| Food craving | 1 | |
| Feeling full | 1 | |
| Physiological parameters | ||
| Levels of lipid profile | 5 | |
| Basal metabolism | 1 | |
| BMI | 1 | |
| Nutrient response | 2 | |
| Nutritional needs | 3 | |
| Blood pressure | 1 | |
| Microbiome | 1 | |
| Bitter taste | 2 | |
| Oxidative stress | ||
| Antioxidant and detoxifying capacity | 3 | |
| Oxidative stress | 1 | |
| Antioxidant needs | 1 | |
| Glutathione | 1 | |
| Hormones | ||
| Insulin sensivity | 1 | |
| Insulin resistance | 1 | |
| Risk of developing insulin resistance | 1 | |
| Other | ||
| N.D. items | 15 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Floris, M.; Cano, A.; Porru, L.; Addis, R.; Cambedda, A.; Idda, M.L.; Steri, M.; Ventura, C.; Maioli, M. Direct-to-Consumer Nutrigenetics Testing: An Overview. Nutrients 2020, 12, 566. https://doi.org/10.3390/nu12020566
Floris M, Cano A, Porru L, Addis R, Cambedda A, Idda ML, Steri M, Ventura C, Maioli M. Direct-to-Consumer Nutrigenetics Testing: An Overview. Nutrients. 2020; 12(2):566. https://doi.org/10.3390/nu12020566
Chicago/Turabian StyleFloris, Matteo, Antonella Cano, Laura Porru, Roberta Addis, Antonio Cambedda, Maria Laura Idda, Maristella Steri, Carlo Ventura, and Margherita Maioli. 2020. "Direct-to-Consumer Nutrigenetics Testing: An Overview" Nutrients 12, no. 2: 566. https://doi.org/10.3390/nu12020566
APA StyleFloris, M., Cano, A., Porru, L., Addis, R., Cambedda, A., Idda, M. L., Steri, M., Ventura, C., & Maioli, M. (2020). Direct-to-Consumer Nutrigenetics Testing: An Overview. Nutrients, 12(2), 566. https://doi.org/10.3390/nu12020566







