Peroxisome Proliferator-Activated Receptors as Molecular Links between Caloric Restriction and Circadian Rhythm
Abstract
1. Introduction
2. Molecular Background of the Circadian Clock
3. Caloric Restriction
4. The Impact of CR on Circadian Rhythm
5. Molecular Pathways Affected by CR and Their Implication in the Circadian Rhythm
5.1. Insulin Signaling in Metabolism and Circadian Rhythm Regulation
5.2. mTOR Signaling and the Circadian Rhythm
5.3. AMPK as an Energy Sensor and Executor of Circadian Metabolic Activities
5.4. SIRT Energy Sensors in the Context of CR and Daily Rhythmicity
6. PPARs
6.1. PPARα
6.1.1. PPARα and Molecular Effectors of CR
6.1.2. PPARα in Circadian Rhythm
6.2. PPARβ/δ
6.2.1. PPARβ/δ and Molecular Effectors of CR
6.2.2. PPARβ/δ in Circadian Rhythm
6.3. PPARγ
6.3.1. PPARγ and Molecular Effectors of CR
6.3.2. PPARγ in Circadian Rhythm
6.4. Systemic Impacts of CR in the Context of Circadian Rhythm and PPAR Activities
6.4.1. Modulation of the Oxidative Status
6.4.2. Reduction of Inflammation
6.4.3. Prevention of Aging
7. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ACC | Acetyl–CoA carboxylase |
| AceCS | Acetyl coenzyme A synthetase |
| AICAR | 5–Aminoimidazole–4–carboxamide ribonucleoside |
| AMP | Adenosine monophosphate () |
| AMPK | Adenosine monophosphate activated protein kinase |
| AP–1 | Activating protein–1 |
| BAT | Brown adipose tissue |
| BHLGB2 | Basic helix–loop–helix domain containing class B 2 |
| BHLH | Basic helix–hoop–helix |
| BMAL1 | Brain and muscle ARNT–like protein 1 |
| CAP | Cbl–associated protein |
| CAT | Catalase |
| CKIε | Casein kinase ε |
| CLC | Cardiotrophin–like cytokine |
| CLOCK | Circadian locomotor output cycles kaput |
| CoA | Acetyl coenzyme A |
| COX–2 | Cyclooxygenase–2 |
| cPLA2 | Cytosolic phospholipase A2 |
| CPT | Carnitine palmitoyltransferase |
| CR | Caloric restriction |
| CREB | cAMP response element–binding protein |
| CRTC2 | cAMP response element–binding protein–regulated transcriptional coactivator–2 |
| CRY | Cryptochrome |
| CYP4A12 | Cytochrome P450 4A12 |
| DEC1 | Differentially expressed in chondrocytes 1 |
| EHHADH | 3-Hydroxyacyl CoA dehydrogenase |
| eIF2α | Eukaryotic initiation factor 2α |
| eNOS | Endothelial NO synthase |
| epSC | Epidermal stem cell |
| ERR | Estrogen–related receptor |
| FA | Fatty acid |
| FAS | Fatty acid synthase |
| FGF21 | Fibroblast growth factor 21 |
| F–L–Leu | FMOC–L–leucine |
| FMO3 | Flavin–containing monooxygenase 3 |
| FOXO | Forkhead |
| G6Pase | Glucose–6–phosphatase |
| GCN2 | General control nonderepressible 2 |
| GHR | Growth hormone receptor |
| GLUT4 | Glucose transporter 4 |
| GP | Glycogen phosphorylase |
| GPx | Glutathione peroxidase |
| GR | Glutathione reductase |
| GRB | Granzyme B |
| GSH | Glutathione |
| GSSG | Oxidized glutathione |
| GST | Glutathione S–transferase |
| GYS | Glycogen synthase |
| HAT | Histone acetyltransferase |
| HDAC5 | Histone deacetylase 5 |
| HFD | High–fat diet |
| HMG–CoAR | 3–Hydroxy 3–methylglutaryl–CoA reductase |
| HMGCS2 | 3–Hydroxy–3–methylglutaryl–CoA synthase 2 |
| ICAM–1 | Intercellular–adhesion–molecule–1 |
| IFNγ | Interferon γ |
| IGF–1 | Insulin–like growth factor 1 |
| iNOS | Inducible nitrogen monoxide synthase |
| IRS | Insulin receptor substrate |
| LCAD | Long–chain acyl–coenzyme A dehydrogenase |
| LKB1 | Liver kinase B1 |
| LPS | Lipopolysaccharide |
| MAPK | Mitogen–activated protein kinase |
| MCAD | Medium–chain acyl–coa dehydrogenase |
| MUP4 | Major urinary protein 4 |
| mTOR | Mammalian target of rapamycin |
| muSC | Skeletal muscle stem cell |
| NAMPT | Nicotinamide phosphoribosyltransferase |
| NAPE-PLD | N-acylphosphatidylethanolamines hydrolyzing phospholipase D |
| NF–κB | Nuclear factor kappa–light–chain enhancer of activated b cells |
| NMN | Nicotinamide mononucleotide |
| NO | Nitrogen monoxide |
| NRF2 | Nuclear factor erythroid 2–related factor 2 |
| NSCLC | Non–small cell lung carcinoma |
| OEA | Oleoylethanolamide |
| PAS | Per–Arnt–Sim |
| PCK1 | Phosphoenolpyruvate carboxykinase 1 |
| PEPCK | Phosphoenolpyruvate–carboxykinase |
| PER | Period |
| PGC–1α | Peroxisome proliferator–activated receptor γ coactivator 1 α |
| PI3 | Phosphatidylinositol 3 |
| PI3K | Phosphoinositide 3–kinase |
| PK2 | Prokineticin 2 |
| PPAR | Peroxisome proliferator–activated receptor |
| PPRE | Peroxisome proliferator–activated receptor response element |
| PRF | Perforin |
| PRX | Peroxidoxin |
| PTEN | Phosphatase and tensin homolog deleted on chromosome 10 |
| S6K1 | Ribosomal S6 kinase 1 |
| SCD–1 | Stearoyl–coa desaturase–1 |
| SCN | Suprachiasmatic nucleus |
| SERPINA12 | Serpin family A member 12 |
| SIRT | Sirtuin |
| SOD–2 | Superoxide dismutase 2 |
| SREBP–1 | Sterol regulatory element–binding protein 1 |
| STAT1 | Signal transducer and activator of transcription 1 |
| RER | Respiratory exchange ratio |
| RHT | Retinohypothalamic tract |
| ROR | Retinoic acid–related orphan receptor |
| RORE | Retinoic acid–related orphan receptor response element |
| ROS | Reactive oxygen species |
| RXR | Retinoid X receptor |
| TAG | Triacylglycerol |
| TGFα | Transforming growth factor α |
| TIM | Timeless |
| TNFα | Tumor necrosis factor α |
| TOR | Target of rapamycin |
| TRF | Time–restricted feeding |
| TSC | Tuberous sclerosis protein |
| TZD | Thiazolidinediones |
| UCP2 | Uncoupling protein 2 |
| WAT | White adipose tissue |
| WT | Wild–type |
| VIP | Vasoactive intestinal polypeptide |
| VLCAD | Very long–chain acyl–coa dehydrogenase |
References
- Panda, S.; Hogenesch, J.B.; Kay, S.A. Circadian rhythms from flies to human. Nature 2002, 417, 329–335. [Google Scholar] [CrossRef] [PubMed]
- Reppert, S.M.; Weaver, D.R. Coordination of circadian timing in mammals. Nature 2002, 418, 935–941. [Google Scholar] [CrossRef] [PubMed]
- Refinetti, R.; Menaker, M. The circadian rhythm of body temperature. Physiol. Behav. 1992, 51, 613–637. [Google Scholar] [CrossRef]
- Bell-Pedersen, D.; Cassone, V.M.; Earnest, D.J.; Golden, S.S.; Hardin, P.E.; Thomas, T.L.; Zoran, M.J. Circadian rhythms from multiple oscillators: Lessons from diverse organisms. Nat. Rev. Genet. 2005, 6, 544–556. [Google Scholar] [CrossRef]
- Schibler, U.; Ripperger, J.; Brown, S.A. Peripheral circadian oscillators in mammals: Time and food. J. Biol. Rhythms 2003, 18, 250–260. [Google Scholar] [CrossRef]
- Panda, S.; Antoch, M.P.; Miller, B.H.; Su, A.I.; Schook, A.B.; Straume, M.; Schultz, P.G.; Kay, S.A.; Takahashi, J.S.; Hogenesch, J.B. Coordinated transcription of key pathways in the mouse by the circadian clock. Cell 2002, 109, 307–320. [Google Scholar] [CrossRef]
- Akhtar, R.A.; Reddy, A.B.; Maywood, E.S.; Clayton, J.D.; King, V.M.; Smith, A.G.; Gant, T.W.; Hastings, M.H.; Kyriacou, C.P. Circadian cycling of the mouse liver transcriptome, as revealed by cDNA microarray, is driven by the suprachiasmatic nucleus. Curr. Biol. 2002, 12, 540–550. [Google Scholar] [CrossRef]
- Kita, Y.; Shiozawa, M.; Jin, W.; Majewski, R.R.; Besharse, J.C.; Greene, A.S.; Jacob, H.J. Implications of circadian gene expression in kidney, liver and the effects of fasting on pharmacogenomic studies. Pharmacogenetics 2002, 12, 55–65. [Google Scholar] [CrossRef]
- Reddy, A.B.; Karp, N.A.; Maywood, E.S.; Sage, E.A.; Deery, M.; O’Neill, J.S.; Wong, G.K.; Chesham, J.; Odell, M.; Lilley, K.S.; et al. Circadian orchestration of the hepatic proteome. Curr. Biol. 2006, 16, 1107–1115. [Google Scholar] [CrossRef]
- Zhang, R.; Lahens, N.F.; Ballance, H.I.; Hughes, M.E.; Hogenesch, J.B. A circadian gene expression atlas in mammals: Implications for biology and medicine. Proc. Natl. Acad. Sci. USA 2014, 111, 16219–16224. [Google Scholar] [CrossRef]
- Dubrovsky, Y.V.; Samsa, W.E.; Kondratov, R.V. Deficiency of circadian protein CLOCK reduces lifespan and increases age-related cataract development in mice. Aging 2010, 2, 936–944. [Google Scholar] [CrossRef] [PubMed]
- Fu, L.; Pelicano, H.; Liu, J.; Huang, P.; Lee, C. The circadian gene Period2 plays an important role in tumor suppression and DNA damage response in vivo. Cell 2002, 111, 41–50. [Google Scholar] [CrossRef]
- Kondratov, R.V.; Kondratova, A.A.; Gorbacheva, V.Y.; Vykhovanets, O.V.; Antoch, M.P. Early aging and age-related pathologies in mice deficient in BMAL1, the core componentof the circadian clock. Genes Dev. 2006, 20, 1868–1873. [Google Scholar] [CrossRef] [PubMed]
- Papagiannakopoulos, T.; Bauer, M.R.; Davidson, S.M.; Heimann, M.; Subbaraj, L.; Bhutkar, A.; Bartlebaugh, J.; Vander Heiden, M.G.; Jacks, T. Circadian rhythm disruption promotes lung tumorigenesis. Cell Metab. 2016, 24, 324–331. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, J.S.; Hong, H.K.; Ko, C.H.; McDearmon, E.L. The genetics of mammalian circadian order and disorder: Implications for physiology and disease. Nat. Rev. Genet. 2008, 9, 764–775. [Google Scholar] [CrossRef]
- Davis, S.; Mirick, D.K. Circadian disruption, shift work and the risk of cancer: A summary of the evidence and studies in Seattle. Cancer Causes Control. 2006, 17, 539–545. [Google Scholar] [CrossRef]
- Anea, C.B.; Zhang, M.; Stepp, D.W.; Simkins, G.B.; Reed, G.; Fulton, D.J.; Rudic, R.D. Vascular disease in mice with a dysfunctional circadian clock. Circulation 2009, 119, 1510–1517. [Google Scholar] [CrossRef]
- Fonken, L.K.; Workman, J.L.; Walton, J.C.; Weil, Z.M.; Morris, J.S.; Haim, A.; Nelson, R.J. Light at night increases body mass by shifting the time of food intake. Proc. Natl. Acad. Sci. USA 2010, 107, 18664–18669. [Google Scholar] [CrossRef]
- Salgado-Delgado, R.; Angeles-Castellanos, M.; Buijs, M.R.; Escobar, C. Internal desynchronization in a model of night-work by forced activity in rats. Neuroscience 2008, 154, 922–931. [Google Scholar] [CrossRef]
- McHill, A.W.; Melanson, E.L.; Higgins, J.; Connick, E.; Moehlman, T.M.; Stothard, E.R.; Wright, K.P., Jr. Impact of circadian misalignment on energy metabolism during simulated nightshift work. Proc. Natl. Acad. Sci. USA 2014, 111, 17302–17307. [Google Scholar] [CrossRef]
- Scheer, F.A.; Hilton, M.F.; Mantzoros, C.S.; Shea, S.A. Adverse metabolic and cardiovascular consequences of circadian misalignment. Proc. Natl. Acad. Sci. USA 2009, 106, 4453–4458. [Google Scholar] [CrossRef] [PubMed]
- Gill, S.; Le, H.D.; Melkani, G.C.; Panda, S. Time-restricted feeding attenuates age-related cardiac decline in Drosophila. Science 2015, 347, 1265–1269. [Google Scholar] [CrossRef] [PubMed]
- Patel, S.A.; Velingkaar, N.; Makwana, K.; Chaudhari, A.; Kondratov, R. Calorie restriction regulates circadian clock gene expression through BMAL1 dependent and independent mechanisms. Sci. Rep. 2016, 6, 25970. [Google Scholar] [CrossRef] [PubMed]
- Stokkan, K.A.; Yamazaki, S.; Tei, H.; Sakaki, Y.; Menaker, M. Entrainment of the circadian clock in the liver by feeding. Science 2001, 291, 490–493. [Google Scholar] [CrossRef] [PubMed]
- Vollmers, C.; Gill, S.; DiTacchio, L.; Pulivarthy, S.R.; Le, H.D.; Panda, S. Time of feeding and the intrinsic circadian clock drive rhythms in hepatic gene expression. Proc. Natl. Acad. Sci. USA 2009, 106, 21453–21458. [Google Scholar] [CrossRef] [PubMed]
- Oike, H.; Nagai, K.; Fukushima, T.; Ishida, N.; Kobori, M. Feeding cues and injected nutrients induce acute expression of multiple clock genes in the mouse liver. PLoS ONE 2011, 6, e23709. [Google Scholar] [CrossRef] [PubMed]
- Tahara, Y.; Otsuka, M.; Fuse, Y.; Hirao, A.; Shibata, S. Refeeding after fasting elicits insulin-dependent regulation of Per2 and Rev-erbalpha with shifts in the liver clock. J. Biol. Rhythms 2011, 26, 230–240. [Google Scholar] [CrossRef]
- Hatori, M.; Vollmers, C.; Zarrinpar, A.; DiTacchio, L.; Bushong, E.A.; Gill, S.; Leblanc, M.; Chaix, A.; Joens, M.; Fitzpatrick, J.A.; et al. Time-restricted feeding without reducing caloric intake prevents metabolic diseases in mice fed a high-fat diet. Cell Metab. 2012, 15, 848–860. [Google Scholar] [CrossRef]
- Sherman, H.; Genzer, Y.; Cohen, R.; Chapnik, N.; Madar, Z.; Froy, O. Timed high-fat diet resets circadian metabolism and prevents obesity. FASEB J. 2012, 26, 3493–3502. [Google Scholar] [CrossRef]
- Kohsaka, A.; Laposky, A.D.; Ramsey, K.M.; Estrada, C.; Joshu, C.; Kobayashi, Y.; Turek, F.W.; Bass, J. High-fat diet disrupts behavioral and molecular circadian rhythms in mice. Cell Metab. 2007, 6, 414–421. [Google Scholar] [CrossRef]
- Salgado-Delgado, R.; Angeles-Castellanos, M.; Saderi, N.; Buijs, R.M.; Escobar, C. Food intake during the normal activity phase prevents obesity and circadian desynchrony in a rat model of night work. Endocrinology 2010, 151, 1019–1029. [Google Scholar] [CrossRef] [PubMed]
- Arble, D.M.; Bass, J.; Laposky, A.D.; Vitaterna, M.H.; Turek, F.W. Circadian timing of food intake contributes to weight gain. Obesity 2009, 17, 2100–2102. [Google Scholar] [CrossRef] [PubMed]
- Gill, S.; Panda, S. A smartphone app reveals erratic diurnal eating patterns in humans that can be modulated for health benefits. Cell Metab. 2015, 22, 789–798. [Google Scholar] [CrossRef] [PubMed]
- Maywood, E.S.; Reddy, A.B.; Wong, G.K.; O’Neill, J.S.; O’Brien, J.A.; McMahon, D.G.; Harmar, A.J.; Okamura, H.; Hastings, M.H. Synchronization and maintenance of timekeeping in suprachiasmatic circadian clock cells by neuropeptidergic signaling. Curr. Biol. 2006, 16, 599–605. [Google Scholar] [CrossRef] [PubMed]
- Lucas, R.J.; Freedman, M.S.; Lupi, D.; Munoz, M.; David-Gray, Z.K.; Foster, R.G. Identifying the photoreceptive inputs to the mammalian circadian system using transgenic and retinally degenerate mice. Behav. Brain Res. 2001, 125, 97–102. [Google Scholar] [CrossRef]
- Kramer, A.; Yang, F.C.; Snodgrass, P.; Li, X.; Scammell, T.E.; Davis, F.C.; Weitz, C.J. Regulation of daily locomotor activity and sleep by hypothalamic EGF receptor signaling. Science 2001, 294, 2511–2515. [Google Scholar] [CrossRef] [PubMed]
- Le Minh, N.; Damiola, F.; Tronche, F.; Schutz, G.; Schibler, U. Glucocorticoid hormones inhibit food-induced phase-shifting of peripheral circadian oscillators. EMBO J. 2001, 20, 7128–7136. [Google Scholar] [CrossRef] [PubMed]
- Cheng, M.Y.; Bullock, C.M.; Li, C.; Lee, A.G.; Bermak, J.C.; Belluzzi, J.; Weaver, D.R.; Leslie, F.M.; Zhou, Q.Y. Prokineticin 2 transmits the behavioural circadian rhythm of the suprachiasmatic nucleus. Nature 2002, 417, 405–410. [Google Scholar] [CrossRef] [PubMed]
- Kraves, S.; Weitz, C.J. A role for cardiotrophin-like cytokine in the circadian control of mammalian locomotor activity. Nat. Neurosci. 2006, 9, 212–219. [Google Scholar] [CrossRef] [PubMed]
- Longo, V.D.; Panda, S. Fasting, circadian rhythms, and time-restricted feeding in healthy lifespan. Cell Metab. 2016, 23, 1048–1059. [Google Scholar] [CrossRef] [PubMed]
- Vitaterna, M.H.; King, D.P.; Chang, A.M.; Kornhauser, J.M.; Lowrey, P.L.; McDonald, J.D.; Dove, W.F.; Pinto, L.H.; Turek, F.W.; Takahashi, J.S. Mutagenesis and mapping of a mouse gene, Clock, essential for circadian behavior. Science 1994, 264, 719–725. [Google Scholar] [CrossRef] [PubMed]
- Gekakis, N.; Staknis, D.; Nguyen, H.B.; Davis, F.C.; Wilsbacher, L.D.; King, D.P.; Takahashi, J.S.; Weitz, C.J. Role of the CLOCK protein in the mammalian circadian mechanism. Science 1998, 280, 1564–1569. [Google Scholar] [CrossRef] [PubMed]
- Kume, K.; Zylka, M.J.; Sriram, S.; Shearman, L.P.; Weaver, D.R.; Jin, X.; Maywood, E.S.; Hastings, M.H.; Reppert, S.M. mCRY1 and mCRY2 are essential components of the negative limb of the circadian clock feedback loop. Cell 1999, 98, 193–205. [Google Scholar] [CrossRef]
- Preitner, N.; Damiola, F.; Lopez-Molina, L.; Zakany, J.; Duboule, D.; Albrecht, U.; Schibler, U. The orphan nuclear receptor REV-ERBalpha controls circadian transcription within the positive limb of the mammalian circadian oscillator. Cell 2002, 110, 251–260. [Google Scholar] [CrossRef]
- Sato, T.K.; Panda, S.; Miraglia, L.J.; Reyes, T.M.; Rudic, R.D.; McNamara, P.; Naik, K.A.; FitzGerald, G.A.; Kay, S.A.; Hogenesch, J.B. A functional genomics strategy reveals Rora as a component of the mammalian circadian clock. Neuron 2004, 43, 527–537. [Google Scholar] [CrossRef]
- Yin, L.; Lazar, M.A. The orphan nuclear receptor Rev-erbalpha recruits the N-CoR/histone deacetylase 3 corepressor to regulate the circadian Bmal1 gene. Mol. Endocrinol. 2005, 19, 1452–1459. [Google Scholar] [CrossRef]
- Sato, T.K.; Yamada, R.G.; Ukai, H.; Baggs, J.E.; Miraglia, L.J.; Kobayashi, T.J.; Welsh, D.K.; Kay, S.A.; Ueda, H.R.; Hogenesch, J.B. Feedback repression is required for mammalian circadian clock function. Nat. Genet. 2006, 38, 312–319. [Google Scholar] [CrossRef]
- Cardone, L.; Hirayama, J.; Giordano, F.; Tamaru, T.; Palvimo, J.J.; Sassone-Corsi, P. Circadian clock control by SUMOylation of BMAL1. Science 2005, 309, 1390–1394. [Google Scholar] [CrossRef]
- Nagel, R.; Clijsters, L.; Agami, R. The miRNA-192/194 cluster regulates the Period gene family and the circadian clock. FEBS J. 2009, 276, 5447–5455. [Google Scholar] [CrossRef]
- Stojkovic, K.; Wing, S.S.; Cermakian, N. A central role for ubiquitination within a circadian clock protein modification code. Front. Mol. Neurosci. 2014, 7, 69. [Google Scholar] [CrossRef]
- Bellet, M.M.; Sassone-Corsi, P. Mammalian circadian clock and metabolism—The epigenetic link. J. Cell Sci. 2010, 123, 3837–3848. [Google Scholar] [CrossRef] [PubMed]
- Reischl, S.; Kramer, A. Kinases and phosphatases in the mammalian circadian clock. FEBS Lett. 2011, 585, 1393–1399. [Google Scholar] [CrossRef] [PubMed]
- Speakman, J.R.; Mitchell, S.E. Caloric restriction. Mol. Aspects Med. 2011, 32, 159–221. [Google Scholar] [CrossRef] [PubMed]
- Masoro, E.J. Overview of caloric restriction and ageing. Mech. Ageing Dev. 2005, 126, 913–922. [Google Scholar] [CrossRef] [PubMed]
- Weindruch, R.; Walford, R.L.; Fligiel, S.; Guthrie, D. The retardation of aging in mice by dietary restriction: Longevity, cancer, immunity and lifetime energy intake. J. Nutr. 1986, 116, 641–654. [Google Scholar] [CrossRef] [PubMed]
- Colman, R.J.; Anderson, R.M.; Johnson, S.C.; Kastman, E.K.; Kosmatka, K.J.; Beasley, T.M.; Allison, D.B.; Cruzen, C.; Simmons, H.A.; Kemnitz, J.W.; et al. Caloric restriction delays disease onset and mortality in rhesus monkeys. Science 2009, 325, 201–204. [Google Scholar] [CrossRef]
- Anderson, R.M.; Shanmuganayagam, D.; Weindruch, R. Caloric restriction and aging: Studies in mice and monkeys. Toxicol. Pathol. 2009, 37, 47–51. [Google Scholar] [CrossRef]
- Abalan, F.; Mayo, W.; Simon, H.; Le Moal, M. Paradoxical effect of severe dietary restriction on Long-Evans rat life span. Int. J. Vitam. Nutr. Res. 2010, 80, 386–393. [Google Scholar] [CrossRef]
- Chung, H.Y.; Kim, H.J.; Kim, J.W.; Yu, B.P. The inflammation hypothesis of aging: Molecular modulation by calorie restriction. Ann. N. Y. Acad. Sci. 2001, 928, 327–335. [Google Scholar] [CrossRef]
- Swindell, W.R. Genes regulated by caloric restriction have unique roles within transcriptional networks. Mech. Ageing Dev. 2008, 129, 580–592. [Google Scholar] [CrossRef]
- Swindell, W.R. Comparative analysis of microarray data identifies common responses to caloric restriction among mouse tissues. Mech. Ageing Dev. 2008, 129, 138–153. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.; Silver, R. Neuroendocrine underpinnings of sex differences in circadian timing systems. J. Steroid. Biochem. Mol. Biol. 2016, 160, 118–126. [Google Scholar] [CrossRef] [PubMed]
- Hadden, H.; Soldin, S.J.; Massaro, D. Circadian disruption alters mouse lung clock gene expression and lung mechanics. J. Appl. Physiol. 2012, 113, 385–392. [Google Scholar] [CrossRef] [PubMed]
- Astafev, A.A.; Patel, S.A.; Kondratov, R.V. Calorie restriction effects on circadian rhythms in gene expression are sex dependent. Sci. Rep. 2017, 7, 9716. [Google Scholar] [CrossRef] [PubMed]
- Mendoza, J.; Graff, C.; Dardente, H.; Pevet, P.; Challet, E. Feeding cues alter clock gene oscillations and photic responses in the suprachiasmatic nuclei of mice exposed to a light/dark cycle. J. Neurosci. 2005, 25, 1514–1522. [Google Scholar] [CrossRef] [PubMed]
- Challet, E.; Caldelas, I.; Graff, C.; Pevet, P. Synchronization of the molecular clockwork by light- and food-related cues in mammals. Biol. Chem. 2003, 384, 711–719. [Google Scholar] [CrossRef]
- Challet, E.; Solberg, L.C.; Turek, F.W. Entrainment in calorie-restricted mice: Conflicting zeitgebers and free-running conditions. Am. J. Physiol. 1998, 274, R1751–R1761. [Google Scholar] [CrossRef]
- Mendoza, J.; Drevet, K.; Pevet, P.; Challet, E. Daily meal timing is not necessary for resetting the main circadian clock by calorie restriction. J. Neuroendocrinol. 2008, 20, 251–260. [Google Scholar] [CrossRef]
- Makwana, K.; Gosai, N.; Poe, A.; Kondratov, R.V. Calorie restriction reprograms diurnal rhythms in protein translation to regulate metabolism. FASEB J. 2019, 33, 4473–4489. [Google Scholar] [CrossRef]
- Sato, S.; Solanas, G.; Peixoto, F.O.; Bee, L.; Symeonidi, A.; Schmidt, M.S.; Brenner, C.; Masri, S.; Benitah, S.A.; Sassone-Corsi, P. Circadian reprogramming in the liver identifies metabolic pathways of aging. Cell 2017, 170, 664–677.e611. [Google Scholar] [CrossRef]
- Patel, S.A.; Chaudhari, A.; Gupta, R.; Velingkaar, N.; Kondratov, R.V. Circadian clocks govern calorie restriction-mediated life span extension through BMAL1- and IGF-1-dependent mechanisms. FASEB J. 2016, 30, 1634–1642. [Google Scholar] [CrossRef] [PubMed]
- Zwighaft, Z.; Aviram, R.; Shalev, M.; Rousso-Noori, L.; Kraut-Cohen, J.; Golik, M.; Brandis, A.; Reinke, H.; Aharoni, A.; Kahana, C.; et al. Circadian clock control by polyamine levels through a mechanism that declines with age. Cell Metab. 2015, 22, 874–885. [Google Scholar] [CrossRef] [PubMed]
- Duszka, K.; Gregor, A.; Guillou, H.; Konig, J.; Wahli, W. Peroxisome proliferator-activated receptors and caloric restriction-common pathways affecting metabolism, health, and longevity. Cells 2020, 9, 1708. [Google Scholar] [CrossRef] [PubMed]
- Bluher, M.; Kahn, B.B.; Kahn, C.R. Extended longevity in mice lacking the insulin receptor in adipose tissue. Science 2003, 299, 572–574. [Google Scholar] [CrossRef] [PubMed]
- Tatar, M.; Bartke, A.; Antebi, A. The endocrine regulation of aging by insulin-like signals. Science 2003, 299, 1346–1351. [Google Scholar] [CrossRef]
- Kenyon, C.; Chang, J.; Gensch, E.; Rudner, A.; Tabtiang, R. A C. elegans mutant that lives twice as long as wild type. Nature 1993, 366, 461–464. [Google Scholar] [CrossRef]
- Taguchi, A.; Wartschow, L.M.; White, M.F. Brain IRS2 signaling coordinates life span and nutrient homeostasis. Science 2007, 317, 369–372. [Google Scholar] [CrossRef]
- Selman, C.; Lingard, S.; Choudhury, A.I.; Batterham, R.L.; Claret, M.; Clements, M.; Ramadani, F.; Okkenhaug, K.; Schuster, E.; Blanc, E.; et al. Evidence for lifespan extension and delayed age-related biomarkers in insulin receptor substrate 1 null mice. FASEB J. 2008, 22, 807–818. [Google Scholar] [CrossRef]
- Weindruch, R. The retardation of aging by caloric restriction: Studies in rodents and primates. Toxicol. Pathol. 1996, 24, 742–745. [Google Scholar] [CrossRef]
- Kim, D.H.; Park, M.H.; Lee, E.K.; Choi, Y.J.; Chung, K.W.; Moon, K.M.; Kim, M.J.; An, H.J.; Park, J.W.; Kim, N.D.; et al. The roles of FoxOs in modulation of aging by calorie restriction. Biogerontology 2015, 16, 1–14. [Google Scholar] [CrossRef]
- Oh, W.J.; Jacinto, E. mTOR complex 2 signaling and functions. Cell Cycle 2011, 10, 2305–2316. [Google Scholar] [CrossRef] [PubMed]
- Rowland, A.F.; Fazakerley, D.J.; James, D.E. Mapping insulin/GLUT4 circuitry. Traffic 2011, 12, 672–681. [Google Scholar] [CrossRef] [PubMed]
- Siddle, K. Signalling by insulin and IGF receptors: Supporting acts and new players. J. Mol. Endocrinol. 2011, 47, R1–R10. [Google Scholar] [CrossRef] [PubMed]
- Piscitello, D.; Varshney, D.; Lilla, S.; Vizioli, M.G.; Reid, C.; Gorbunova, V.; Seluanov, A.; Gillespie, D.A.; Adams, P.D. AKT overactivation can suppress DNA repair via p70S6 kinase-dependent downregulation of MRE11. Oncogene 2018, 37, 427–438. [Google Scholar] [CrossRef] [PubMed]
- Jia, Y.; Song, W.; Zhang, F.; Yan, J.; Yang, Q. Akt1 inhibits homologous recombination in Brca1-deficient cells by blocking the Chk1-Rad51 pathway. Oncogene 2013, 32, 1943–1949. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Gan, W.; Guo, C.; Xie, A.; Gao, D.; Guo, J.; Zhang, J.; Willis, N.; Su, A.; Asara, J.M.; et al. Akt-mediated phosphorylation of XLF impairs non-homologous end-joining DNA repair. Mol. Cell 2015, 57, 648–661. [Google Scholar] [CrossRef] [PubMed]
- Zhang, E.E.; Liu, A.C.; Hirota, T.; Miraglia, L.J.; Welch, G.; Pongsawakul, P.Y.; Liu, X.; Atwood, A.; Huss, J.W., 3rd; Janes, J.; et al. A genome-wide RNAi screen for modifiers of the circadian clock in human cells. Cell 2009, 139, 199–210. [Google Scholar] [CrossRef] [PubMed]
- Yamajuku, D.; Inagaki, T.; Haruma, T.; Okubo, S.; Kataoka, Y.; Kobayashi, S.; Ikegami, K.; Laurent, T.; Kojima, T.; Noutomi, K.; et al. Real-time monitoring in three-dimensional hepatocytes reveals that insulin acts as a synchronizer for liver clock. Sci. Rep. 2012, 2, 439. [Google Scholar] [CrossRef]
- Lamia, K.A.; Sachdeva, U.M.; DiTacchio, L.; Williams, E.C.; Alvarez, J.G.; Egan, D.F.; Vasquez, D.S.; Juguilon, H.; Panda, S.; Shaw, R.J.; et al. AMPK regulates the circadian clock by cryptochrome phosphorylation and degradation. Science 2009, 326, 437–440. [Google Scholar] [CrossRef]
- Iwanaga, H.; Yano, M.; Miki, H.; Okada, K.; Azama, T.; Takiguchi, S.; Fujiwara, Y.; Yasuda, T.; Nakayama, M.; Kobayashi, M.; et al. Per2 gene expressions in the suprachiasmatic nucleus and liver differentially respond to nutrition factors in rats. JPEN J. Parenter. Enteral. Nutr. 2005, 29, 157–161. [Google Scholar] [CrossRef]
- Challet, E.; Losee-Olson, S.; Turek, F.W. Reduced glucose availability attenuates circadian responses to light in mice. Am. J. Physiol. 1999, 276, R1063–R1070. [Google Scholar] [CrossRef] [PubMed]
- Hirota, T.; Okano, T.; Kokame, K.; Shirotani-Ikejima, H.; Miyata, T.; Fukada, Y. Glucose down-regulates Per1 and Per2 mRNA levels and induces circadian gene expression in cultured Rat-1 fibroblasts. J. Biol. Chem. 2002, 277, 44244–44251. [Google Scholar] [CrossRef] [PubMed]
- Marcheva, B.; Ramsey, K.M.; Buhr, E.D.; Kobayashi, Y.; Su, H.; Ko, C.H.; Ivanova, G.; Omura, C.; Mo, S.; Vitaterna, M.H.; et al. Disruption of the clock components CLOCK and BMAL1 leads to hypoinsulinaemia and diabetes. Nature 2010, 466, 627–631. [Google Scholar] [CrossRef] [PubMed]
- Vieira, E.; Marroqui, L.; Figueroa, A.L.; Merino, B.; Fernandez-Ruiz, R.; Nadal, A.; Burris, T.P.; Gomis, R.; Quesada, I. Involvement of the clock gene Rev-erb alpha in the regulation of glucagon secretion in pancreatic alpha-cells. PLoS ONE 2013, 8, e69939. [Google Scholar] [CrossRef] [PubMed]
- Sadacca, L.A.; Lamia, K.A.; deLemos, A.S.; Blum, B.; Weitz, C.J. An intrinsic circadian clock of the pancreas is required for normal insulin release and glucose homeostasis in mice. Diabetologia 2011, 54, 120–124. [Google Scholar] [CrossRef] [PubMed]
- Vieira, E.; Merino, B.; Quesada, I. Role of the clock gene Rev-erbalpha in metabolism and in the endocrine pancreas. Diabetes Obes. Metab. 2015, 17 (Suppl. S1), 106–114. [Google Scholar] [CrossRef] [PubMed]
- Muhlbauer, E.; Wolgast, S.; Finckh, U.; Peschke, D.; Peschke, E. Indication of circadian oscillations in the rat pancreas. FEBS Lett. 2004, 564, 91–96. [Google Scholar] [CrossRef]
- Rudic, R.D.; McNamara, P.; Curtis, A.M.; Boston, R.C.; Panda, S.; Hogenesch, J.B.; Fitzgerald, G.A. BMAL1 and CLOCK, two essential components of the circadian clock, are involved in glucose homeostasis. PLoS Biol. 2004, 2, e377. [Google Scholar] [CrossRef]
- Barclay, J.L.; Shostak, A.; Leliavski, A.; Tsang, A.H.; Johren, O.; Muller-Fielitz, H.; Landgraf, D.; Naujokat, N.; van der Horst, G.T.; Oster, H. High-fat diet-induced hyperinsulinemia and tissue-specific insulin resistance in Cry-deficient mice. Am. J. Physiol. Endocrinol. Metab. 2013, 304, E1053–E1063. [Google Scholar] [CrossRef]
- Zhao, Y.; Zhang, Y.; Zhou, M.; Wang, S.; Hua, Z.; Zhang, J. Loss of mPer2 increases plasma insulin levels by enhanced glucose-stimulated insulin secretion and impaired insulin clearance in mice. FEBS Lett. 2012, 586, 1306–1311. [Google Scholar] [CrossRef]
- Delezie, J.; Dumont, S.; Dardente, H.; Oudart, H.; Grechez-Cassiau, A.; Klosen, P.; Teboul, M.; Delaunay, F.; Pevet, P.; Challet, E. The nuclear receptor REV-ERBalpha is required for the daily balance of carbohydrate and lipid metabolism. FASEB J. 2012, 26, 3321–3335. [Google Scholar] [CrossRef] [PubMed]
- Kalsbeek, A.; la Fleur, S.; Fliers, E. Circadian control of glucose metabolism. Mol. Metab. 2014, 3, 372–383. [Google Scholar] [CrossRef] [PubMed]
- Doi, R.; Oishi, K.; Ishida, N. CLOCK regulates circadian rhythms of hepatic glycogen synthesis through transcriptional activation of Gys2. J. Biol. Chem. 2010, 285, 22114–22121. [Google Scholar] [CrossRef] [PubMed]
- Matsuoka, H.; Shima, A.; Kuramoto, D.; Kikumoto, D.; Matsui, T.; Michihara, A. Phosphoenolpyruvate carboxykinase, a key enzyme that controls blood glucose, is a target of retinoic acid receptor-related orphan receptor alpha. PLoS ONE 2015, 10, e0137955. [Google Scholar] [CrossRef] [PubMed]
- Kadiri, S.; Monnier, C.; Ganbold, M.; Ledent, T.; Capeau, J.; Antoine, B. The nuclear retinoid-related orphan receptor-alpha regulates adipose tissue glyceroneogenesis in addition to hepatic gluconeogenesis. Am. J. Physiol Endocrinol. Metab. 2015, 309, E105–E114. [Google Scholar] [CrossRef]
- Kojetin, D.; Wang, Y.; Kamenecka, T.M.; Burris, T.P. Identification of SR8278, a synthetic antagonist of the nuclear heme receptor REV-ERB. ACS Chem. Biol. 2011, 6, 131–134. [Google Scholar] [CrossRef] [PubMed]
- Yin, L.; Wu, N.; Curtin, J.C.; Qatanani, M.; Szwergold, N.R.; Reid, R.A.; Waitt, G.M.; Parks, D.J.; Pearce, K.H.; Wisely, G.B.; et al. Rev-erbalpha, a heme sensor that coordinates metabolic and circadian pathways. Science 2007, 318, 1786–1789. [Google Scholar] [CrossRef]
- Zhang, E.E.; Liu, Y.; Dentin, R.; Pongsawakul, P.Y.; Liu, A.C.; Hirota, T.; Nusinow, D.A.; Sun, X.; Landais, S.; Kodama, Y.; et al. Cryptochrome mediates circadian regulation of cAMP signaling and hepatic gluconeogenesis. Nat. Med. 2010, 16, 1152–1156. [Google Scholar] [CrossRef]
- Lamia, K.A.; Papp, S.J.; Yu, R.T.; Barish, G.D.; Uhlenhaut, N.H.; Jonker, J.W.; Downes, M.; Evans, R.M. Cryptochromes mediate rhythmic repression of the glucocorticoid receptor. Nature 2011, 480, 552–556. [Google Scholar] [CrossRef]
- Zani, F.; Breasson, L.; Becattini, B.; Vukolic, A.; Montani, J.P.; Albrecht, U.; Provenzani, A.; Ripperger, J.A.; Solinas, G. PER2 promotes glucose storage to liver glycogen during feeding and acute fasting by inducing Gys2 PTG and G L expression. Mol. Metab. 2013, 2, 292–305. [Google Scholar] [CrossRef]
- Bhaskar, P.T.; Hay, N. The two TORCs and Akt. Dev. Cell 2007, 12, 487–502. [Google Scholar] [CrossRef] [PubMed]
- Wullschleger, S.; Loewith, R.; Hall, M.N. TOR signaling in growth and metabolism. Cell 2006, 124, 471–484. [Google Scholar] [CrossRef] [PubMed]
- Loewith, R.; Jacinto, E.; Wullschleger, S.; Lorberg, A.; Crespo, J.L.; Bonenfant, D.; Oppliger, W.; Jenoe, P.; Hall, M.N. Two TOR complexes, only one of which is rapamycin sensitive, have distinct roles in cell growth control. Mol. Cell 2002, 10, 457–468. [Google Scholar] [CrossRef]
- Jacinto, E.; Loewith, R.; Schmidt, A.; Lin, S.; Ruegg, M.A.; Hall, A.; Hall, M.N. Mammalian TOR complex 2 controls the actin cytoskeleton and is rapamycin insensitive. Nat. Cell Biol. 2004, 6, 1122–1128. [Google Scholar] [CrossRef] [PubMed]
- Corradetti, M.N.; Guan, K.L. Upstream of the mammalian target of rapamycin: Do all roads pass through mTOR? Oncogene 2006, 25, 6347–6360. [Google Scholar] [CrossRef]
- Hay, N.; Sonenberg, N. Upstream and downstream of mTOR. Genes Dev. 2004, 18, 1926–1945. [Google Scholar] [CrossRef]
- Arsham, A.M.; Neufeld, T.P. Thinking globally and acting locally with TOR. Curr. Opin. Cell Biol. 2006, 18, 589–597. [Google Scholar] [CrossRef]
- Duvel, K.; Yecies, J.L.; Menon, S.; Raman, P.; Lipovsky, A.I.; Souza, A.L.; Triantafellow, E.; Ma, Q.; Gorski, R.; Cleaver, S.; et al. Activation of a metabolic gene regulatory network downstream of mTOR complex 1. Mol. Cell 2010, 39, 171–183. [Google Scholar] [CrossRef]
- Hudson, C.C.; Liu, M.; Chiang, G.G.; Otterness, D.M.; Loomis, D.C.; Kaper, F.; Giaccia, A.J.; Abraham, R.T. Regulation of hypoxia-inducible factor 1alpha expression and function by the mammalian target of rapamycin. Mol. Cell Biol. 2002, 22, 7004–7014. [Google Scholar] [CrossRef]
- Kim, J.E.; Chen, J. regulation of peroxisome proliferator-activated receptor-gamma activity by mammalian target of rapamycin and amino acids in adipogenesis. Diabetes 2004, 53, 2748–2756. [Google Scholar] [CrossRef]
- Zhang, H.H.; Huang, J.; Duvel, K.; Boback, B.; Wu, S.; Squillace, R.M.; Wu, C.L.; Manning, B.D. Insulin stimulates adipogenesis through the Akt-TSC2-mTORC1 pathway. PLoS ONE 2009, 4, e6189. [Google Scholar] [CrossRef] [PubMed]
- Porstmann, T.; Santos, C.R.; Griffiths, B.; Cully, M.; Wu, M.; Leevers, S.; Griffiths, J.R.; Chung, Y.L.; Schulze, A. SREBP activity is regulated by mTORC1 and contributes to Akt-dependent cell growth. Cell Metab. 2008, 8, 224–236. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.T.; Ducker, G.S.; Barczak, A.J.; Barbeau, R.; Erle, D.J.; Shokat, K.M. The mammalian target of rapamycin regulates cholesterol biosynthetic gene expression and exhibits a rapamycin-resistant transcriptional profile. Proc. Natl. Acad. Sci. USA 2011, 108, 15201–15206. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Guan, K.L. mTOR as a central hub of nutrient signalling and cell growth. Nat. Cell Biol. 2019, 21, 63–71. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Zhai, B.; Gygi, S.P.; Goldberg, A.L. mTOR inhibition activates overall protein degradation by the ubiquitin proteasome system as well as by autophagy. Proc. Natl. Acad. Sci. USA 2015, 112, 15790–15797. [Google Scholar] [CrossRef] [PubMed]
- Cao, R.; Lee, B.; Cho, H.Y.; Saklayen, S.; Obrietan, K. Photic regulation of the mTOR signaling pathway in the suprachiasmatic circadian clock. Mol. Cell Neurosci. 2008, 38, 312–324. [Google Scholar] [CrossRef] [PubMed]
- Cao, R.; Anderson, F.E.; Jung, Y.J.; Dziema, H.; Obrietan, K. Circadian regulation of mammalian target of rapamycin signaling in the mouse suprachiasmatic nucleus. Neuroscience 2011, 181, 79–88. [Google Scholar] [CrossRef]
- Cornu, M.; Oppliger, W.; Albert, V.; Robitaille, A.M.; Trapani, F.; Quagliata, L.; Fuhrer, T.; Sauer, U.; Terracciano, L.; Hall, M.N. Hepatic mTORC1 controls locomotor activity, body temperature, and lipid metabolism through FGF21. Proc. Natl. Acad. Sci. USA 2014, 111, 11592–11599. [Google Scholar] [CrossRef]
- Jouffe, C.; Cretenet, G.; Symul, L.; Martin, E.; Atger, F.; Naef, F.; Gachon, F. The circadian clock coordinates ribosome biogenesis. PLoS Biol. 2013, 11, e1001455. [Google Scholar] [CrossRef]
- Khapre, R.V.; Kondratova, A.A.; Patel, S.; Dubrovsky, Y.; Wrobel, M.; Antoch, M.P.; Kondratov, R.V. BMAL1-dependent regulation of the mTOR signaling pathway delays aging. Aging 2014, 6, 48–57. [Google Scholar] [CrossRef]
- Lipton, J.O.; Yuan, E.D.; Boyle, L.M.; Ebrahimi-Fakhari, D.; Kwiatkowski, E.; Nathan, A.; Guttler, T.; Davis, F.; Asara, J.M.; Sahin, M. The circadian protein BMAL1 regulates translation in response to S6K1-mediated phosphorylation. Cell 2015, 161, 1138–1151. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.C.; Ko, M.L.; Ko, G.Y. A new functional role for mechanistic/mammalian target of rapamycin complex 1 (mTORC1) in the circadian regulation of L-type voltage-gated calcium channels in avian cone photoreceptors. PLoS ONE 2013, 8, e73315. [Google Scholar] [CrossRef] [PubMed]
- Dragert, K.; Bhattacharya, I.; Hall, M.N.; Humar, R.; Battegay, E.; Haas, E. Basal mTORC2 activity and expression of its components display diurnal variation in mouse perivascular adipose tissue. Biochem. Biophys. Res. Commun. 2016, 473, 317–322. [Google Scholar] [CrossRef] [PubMed]
- Chang, S.W.; Yoshihara, T.; Machida, S.; Naito, H. Circadian rhythm of intracellular protein synthesis signaling in rat cardiac and skeletal muscles. Biochem. Biophys. Rep. 2017, 9, 153–158. [Google Scholar] [CrossRef] [PubMed]
- Saraf, A.; Luo, J.; Morris, D.R.; Storm, D.R. Phosphorylation of eukaryotic translation initiation factor 4E and eukaryotic translation initiation factor 4E-binding protein (4EBP) and their upstream signaling components undergo diurnal oscillation in the mouse hippocampus: Implications for memory persistence. J. Biol. Chem. 2014, 289, 20129–20138. [Google Scholar] [CrossRef] [PubMed]
- Albert, V.; Cornu, M.; Hall, M.N. mTORC1 signaling in Agrp neurons mediates circadian expression of Agrp and NPY but is dispensable for regulation of feeding behavior. Biochem. Biophys. Res. Commun. 2015, 464, 480–486. [Google Scholar] [CrossRef] [PubMed]
- Khapre, R.V.; Patel, S.A.; Kondratova, A.A.; Chaudhary, A.; Velingkaar, N.; Antoch, M.P.; Kondratov, R.V. Metabolic clock generates nutrient anticipation rhythms in mTOR signaling. Aging 2014, 6, 675–689. [Google Scholar] [CrossRef]
- Zheng, X.; Sehgal, A. AKT and TOR signaling set the pace of the circadian pacemaker. Curr. Biol. 2010, 20, 1203–1208. [Google Scholar] [CrossRef]
- Kijak, E.; Pyza, E. TOR signaling pathway and autophagy are involved in the regulation of circadian rhythms in behavior and plasticity of L2 interneurons in the brain of Drosophila melanogaster. PLoS ONE 2017, 12, e0171848. [Google Scholar] [CrossRef]
- Cao, R.; Robinson, B.; Xu, H.; Gkogkas, C.; Khoutorsky, A.; Alain, T.; Yanagiya, A.; Nevarko, T.; Liu, A.C.; Amir, S.; et al. Translational control of entrainment and synchrony of the suprachiasmatic circadian clock by mTOR/4E-BP1 signaling. Neuron 2013, 79, 712–724. [Google Scholar] [CrossRef]
- Cao, R.; Li, A.; Cho, H.Y.; Lee, B.; Obrietan, K. Mammalian target of rapamycin signaling modulates photic entrainment of the suprachiasmatic circadian clock. J. Neurosci. 2010, 30, 6302–6314. [Google Scholar] [CrossRef] [PubMed]
- Ramanathan, C.; Kathale, N.D.; Liu, D.; Lee, C.; Freeman, D.A.; Hogenesch, J.B.; Cao, R.; Liu, A.C. mTOR signaling regulates central and peripheral circadian clock function. PLoS Genet. 2018, 14, e1007369. [Google Scholar] [CrossRef] [PubMed]
- Feeney, K.A.; Hansen, L.L.; Putker, M.; Olivares-Yanez, C.; Day, J.; Eades, L.J.; Larrondo, L.F.; Hoyle, N.P.; O’Neill, J.S.; van Ooijen, G. Daily magnesium fluxes regulate cellular timekeeping and energy balance. Nature 2016, 532, 375–379. [Google Scholar] [CrossRef] [PubMed]
- Winder, W.W.; Hardie, D.G. AMP-activated protein kinase, a metabolic master switch: Possible roles in type 2 diabetes. Am. J. Physiol. 1999, 277, E1–E10. [Google Scholar] [CrossRef] [PubMed]
- Koo, S.H.; Flechner, L.; Qi, L.; Zhang, X.; Screaton, R.A.; Jeffries, S.; Hedrick, S.; Xu, W.; Boussouar, F.; Brindle, P.; et al. The CREB coactivator TORC2 is a key regulator of fasting glucose metabolism. Nature 2005, 437, 1109–1111. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Murphy, J.; Toth, R.; Campbell, D.G.; Morrice, N.A.; Mackintosh, C. Complementary regulation of TBC1D1 and AS160 by growth factors, insulin and AMPK activators. Biochem. J. 2008, 409, 449–459. [Google Scholar] [CrossRef] [PubMed]
- Geraghty, K.M.; Chen, S.; Harthill, J.E.; Ibrahim, A.F.; Toth, R.; Morrice, N.A.; Vandermoere, F.; Moorhead, G.B.; Hardie, D.G.; MacKintosh, C. Regulation of multisite phosphorylation and 14-3-3 binding of AS160 in response to IGF-1, EGF, PMA and AICAR. Biochem. J. 2007, 407, 231–241. [Google Scholar] [CrossRef]
- Jager, S.; Handschin, C.; St-Pierre, J.; Spiegelman, B.M. AMP-activated protein kinase (AMPK) action in skeletal muscle via direct phosphorylation of PGC-1alpha. Proc. Natl. Acad. Sci. USA 2007, 104, 12017–12022. [Google Scholar] [CrossRef]
- McGee, S.L.; van Denderen, B.J.; Howlett, K.F.; Mollica, J.; Schertzer, J.D.; Kemp, B.E.; Hargreaves, M. AMP-activated protein kinase regulates GLUT4 transcription by phosphorylating histone deacetylase 5. Diabetes 2008, 57, 860–867. [Google Scholar] [CrossRef]
- Davies, S.P.; Sim, A.T.; Hardie, D.G. Location and function of three sites phosphorylated on rat acetyl-CoA carboxylase by the AMP-activated protein kinase. Eur. J. Biochem. 1990, 187, 183–190. [Google Scholar] [CrossRef]
- Davies, S.P.; Carling, D.; Munday, M.R.; Hardie, D.G. Diurnal rhythm of phosphorylation of rat liver acetyl-CoA carboxylase by the AMP-activated protein kinase, demonstrated using freeze-clamping. Effects of high fat diets. Eur. J. Biochem. 1992, 203, 615–623. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Xu, S.; Mihaylova, M.M.; Zheng, B.; Hou, X.; Jiang, B.; Park, O.; Luo, Z.; Lefai, E.; Shyy, J.Y.; et al. AMPK phosphorylates and inhibits SREBP activity to attenuate hepatic steatosis and atherosclerosis in diet-induced insulin-resistant mice. Cell Metab. 2011, 13, 376–388. [Google Scholar] [CrossRef] [PubMed]
- Muoio, D.M.; Seefeld, K.; Witters, L.A.; Coleman, R.A. AMP-activated kinase reciprocally regulates triacylglycerol synthesis and fatty acid oxidation in liver and muscle: Evidence that sn-glycerol-3-phosphate acyltransferase is a novel target. Biochem. J. 1999, 338, 783–791. [Google Scholar] [CrossRef] [PubMed]
- Clarke, P.R.; Hardie, D.G. Regulation of HMG-CoA reductase: Identification of the site phosphorylated by the AMP-activated protein kinase in vitro and in intact rat liver. EMBO J. 1990, 9, 2439–2446. [Google Scholar] [CrossRef] [PubMed]
- Hoppe, S.; Bierhoff, H.; Cado, I.; Weber, A.; Tiebe, M.; Grummt, I.; Voit, R. AMP-activated protein kinase adapts rRNA synthesis to cellular energy supply. Proc. Natl. Acad. Sci. USA 2009, 106, 17781–17786. [Google Scholar] [CrossRef]
- Marsin, A.S.; Bouzin, C.; Bertrand, L.; Hue, L. The stimulation of glycolysis by hypoxia in activated monocytes is mediated by AMP-activated protein kinase and inducible 6-phosphofructo-2-kinase. J. Biol. Chem. 2002, 277, 30778–30783. [Google Scholar] [CrossRef]
- Jorgensen, S.B.; Nielsen, J.N.; Birk, J.B.; Olsen, G.S.; Viollet, B.; Andreelli, F.; Schjerling, P.; Vaulont, S.; Hardie, D.G.; Hansen, B.F.; et al. The alpha2-5’AMP-activated protein kinase is a site 2 glycogen synthase kinase in skeletal muscle and is responsive to glucose loading. Diabetes 2004, 53, 3074–3081. [Google Scholar] [CrossRef]
- Hardie, D.G.; Ross, F.A.; Hawley, S.A. AMPK: A nutrient and energy sensor that maintains energy homeostasis. Nat. Rev. Mol. Cell Biol. 2012, 13, 251–262. [Google Scholar] [CrossRef]
- Leclerc, I.; Lenzner, C.; Gourdon, L.; Vaulont, S.; Kahn, A.; Viollet, B. Hepatocyte nuclear factor-4alpha involved in type 1 maturity-onset diabetes of the young is a novel target of AMP-activated protein kinase. Diabetes 2001, 50, 1515–1521. [Google Scholar] [CrossRef]
- Lee, J.M.; Seo, W.Y.; Song, K.H.; Chanda, D.; Kim, Y.D.; Kim, D.K.; Lee, M.W.; Ryu, D.; Kim, Y.H.; Noh, J.R.; et al. AMPK-dependent repression of hepatic gluconeogenesis via disruption of CREB.CRTC2 complex by orphan nuclear receptor small heterodimer partner. J. Biol. Chem. 2010, 285, 32182–32191. [Google Scholar] [CrossRef]
- Dufer, M.; Noack, K.; Krippeit-Drews, P.; Drews, G. Activation of the AMP-activated protein kinase enhances glucose-stimulated insulin secretion in mouse beta-cells. Islets 2010, 2, 156–163. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Chavez, J.A.; Roach, W.G.; Keller, S.R.; Lane, W.S.; Lienhard, G.E. Inhibition of GLUT4 translocation by Tbc1d1, a Rab GTPase-activating protein abundant in skeletal muscle, is partially relieved by AMP-activated protein kinase activation. J. Biol. Chem. 2008, 283, 9187–9195. [Google Scholar] [CrossRef] [PubMed]
- Steinberg, G.R.; Kemp, B.E. AMPK in health and disease. Physiol. Rev. 2009, 89, 1025–1078. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, A.; Okamoto, S.; Lee, S.; Saito, K.; Shiuchi, T.; Minokoshi, Y. Leptin stimulates fatty acid oxidation and peroxisome proliferator-activated receptor alpha gene expression in mouse C2C12 myoblasts by changing the subcellular localization of the alpha2 form of AMP-activated protein kinase. Mol. Cell Biol. 2007, 27, 4317–4327. [Google Scholar] [CrossRef] [PubMed]
- Um, J.H.; Pendergast, J.S.; Springer, D.A.; Foretz, M.; Viollet, B.; Brown, A.; Kim, M.K.; Yamazaki, S.; Chung, J.H. AMPK regulates circadian rhythms in a tissue- and isoform-specific manner. PLoS ONE 2011, 6, e18450. [Google Scholar] [CrossRef] [PubMed]
- Vieira, E.; Nilsson, E.C.; Nerstedt, A.; Ormestad, M.; Long, Y.C.; Garcia-Roves, P.M.; Zierath, J.R.; Mahlapuu, M. Relationship between AMPK and the transcriptional balance of clock-related genes in skeletal muscle. Am. J. Physiol. Endocrinol. Metab. 2008, 295, E1032–E1037. [Google Scholar] [CrossRef]
- Um, J.H.; Yang, S.; Yamazaki, S.; Kang, H.; Viollet, B.; Foretz, M.; Chung, J.H. Activation of 5’-AMP-activated kinase with diabetes drug metformin induces casein kinase Iepsilon (CKIepsilon)-dependent degradation of clock protein mPer2. J. Biol. Chem. 2007, 282, 20794–20798. [Google Scholar] [CrossRef]
- Grimaldi, B.; Sassone-Corsi, P. Circadian rhythms: Metabolic clockwork. Nature 2007, 447, 386–387. [Google Scholar] [CrossRef]
- Liu, C.; Li, S.; Liu, T.; Borjigin, J.; Lin, J.D. Transcriptional coactivator PGC-1alpha integrates the mammalian clock and energy metabolism. Nature 2007, 447, 477–481. [Google Scholar] [CrossRef]
- Minokoshi, Y.; Alquier, T.; Furukawa, N.; Kim, Y.B.; Lee, A.; Xue, B.; Mu, J.; Foufelle, F.; Ferre, P.; Birnbaum, M.J.; et al. AMP-kinase regulates food intake by responding to hormonal and nutrient signals in the hypothalamus. Nature 2004, 428, 569–574. [Google Scholar] [CrossRef]
- Andersson, U.; Filipsson, K.; Abbott, C.R.; Woods, A.; Smith, K.; Bloom, S.R.; Carling, D.; Small, C.J. AMP-activated protein kinase plays a role in the control of food intake. J. Biol. Chem. 2004, 279, 12005–12008. [Google Scholar] [CrossRef] [PubMed]
- Liou, G.G.; Tanny, J.C.; Kruger, R.G.; Walz, T.; Moazed, D. Assembly of the SIR complex and its regulation by O-acetyl-ADP-ribose, a product of NAD-dependent histone deacetylation. Cell 2005, 121, 515–527. [Google Scholar] [CrossRef] [PubMed]
- Vaquero, A.; Scher, M.; Lee, D.; Erdjument-Bromage, H.; Tempst, P.; Reinberg, D. Human SirT1 interacts with histone H1 and promotes formation of facultative heterochromatin. Mol. Cell 2004, 16, 93–105. [Google Scholar] [CrossRef] [PubMed]
- Dang, W. The controversial world of sirtuins. Drug. Discov. Today Technol. 2014, 12, e9–e17. [Google Scholar] [CrossRef] [PubMed]
- Houtkooper, R.H.; Pirinen, E.; Auwerx, J. Sirtuins as regulators of metabolism and healthspan. Nat. Rev. Mol. Cell Biol. 2012, 13, 225–238. [Google Scholar] [CrossRef] [PubMed]
- Shimazu, T.; Hirschey, M.D.; Hua, L.; Dittenhafer-Reed, K.E.; Schwer, B.; Lombard, D.B.; Li, Y.; Bunkenborg, J.; Alt, F.W.; Denu, J.M.; et al. SIRT3 deacetylates mitochondrial 3-hydroxy-3-methylglutaryl CoA synthase 2 and regulates ketone body production. Cell Metab. 2010, 12, 654–661. [Google Scholar] [CrossRef]
- Hirschey, M.D.; Shimazu, T.; Goetzman, E.; Jing, E.; Schwer, B.; Lombard, D.B.; Grueter, C.A.; Harris, C.; Biddinger, S.; Ilkayeva, O.R.; et al. SIRT3 regulates mitochondrial fatty-acid oxidation by reversible enzyme deacetylation. Nature 2010, 464, 121–125. [Google Scholar] [CrossRef]
- Qiu, X.; Brown, K.; Hirschey, M.D.; Verdin, E.; Chen, D. Calorie restriction reduces oxidative stress by SIRT3-mediated SOD2 activation. Cell Metab. 2010, 12, 662–667. [Google Scholar] [CrossRef]
- Preyat, N.; Leo, O. Sirtuin deacylases: A molecular link between metabolism and immunity. J. Leukoc. Biol. 2013, 93, 669–680. [Google Scholar] [CrossRef]
- Satoh, A.; Brace, C.S.; Ben-Josef, G.; West, T.; Wozniak, D.F.; Holtzman, D.M.; Herzog, E.D.; Imai, S. SIRT1 promotes the central adaptive response to diet restriction through activation of the dorsomedial and lateral nuclei of the hypothalamus. J. Neurosci. 2010, 30, 10220–10232. [Google Scholar] [CrossRef]
- Kaeberlein, M.; McVey, M.; Guarente, L. The SIR2/3/4 complex and SIR2 alone promote longevity in Saccharomyces cerevisiae by two different mechanisms. Genes Dev. 1999, 13, 2570–2580. [Google Scholar] [CrossRef] [PubMed]
- Whitaker, R.; Faulkner, S.; Miyokawa, R.; Burhenn, L.; Henriksen, M.; Wood, J.G.; Helfand, S.L. Increased expression of Drosophila Sir2 extends life span in a dose-dependent manner. Aging 2013, 5, 682–691. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.J.; Defossez, P.A.; Guarente, L. Requirement of NAD and SIR2 for life-span extension by calorie restriction in Saccharomyces cerevisiae. Science 2000, 289, 2126–2128. [Google Scholar] [CrossRef] [PubMed]
- Rogina, B.; Helfand, S.L. Sir2 mediates longevity in the fly through a pathway related to calorie restriction. Proc. Natl. Acad. Sci. USA 2004, 101, 15998–16003. [Google Scholar] [CrossRef] [PubMed]
- Tissenbaum, H.A.; Guarente, L. Increased dosage of a sir-2 gene extends lifespan in Caenorhabditis elegans. Nature 2001, 410, 227–230. [Google Scholar] [CrossRef] [PubMed]
- Oberdoerffer, P.; Michan, S.; McVay, M.; Mostoslavsky, R.; Vann, J.; Park, S.K.; Hartlerode, A.; Stegmuller, J.; Hafner, A.; Loerch, P.; et al. SIRT1 redistribution on chromatin promotes genomic stability but alters gene expression during aging. Cell 2008, 135, 907–918. [Google Scholar] [CrossRef] [PubMed]
- Rodgers, J.T.; Lerin, C.; Haas, W.; Gygi, S.P.; Spiegelman, B.M.; Puigserver, P. Nutrient control of glucose homeostasis through a complex of PGC-1alpha and SIRT1. Nature 2005, 434, 113–118. [Google Scholar] [CrossRef]
- Nemoto, S.; Fergusson, M.M.; Finkel, T. SIRT1 functionally interacts with the metabolic regulator and transcriptional coactivator PGC-1{alpha}. J. Biol. Chem. 2005, 280, 16456–16460. [Google Scholar] [CrossRef]
- Lagouge, M.; Argmann, C.; Gerhart-Hines, Z.; Meziane, H.; Lerin, C.; Daussin, F.; Messadeq, N.; Milne, J.; Lambert, P.; Elliott, P.; et al. Resveratrol improves mitochondrial function and protects against metabolic disease by activating SIRT1 and PGC-1alpha. Cell 2006, 127, 1109–1122. [Google Scholar] [CrossRef]
- Daitoku, H.; Hatta, M.; Matsuzaki, H.; Aratani, S.; Ohshima, T.; Miyagishi, M.; Nakajima, T.; Fukamizu, A. Silent information regulator 2 potentiates Foxo1-mediated transcription through its deacetylase activity. Proc. Natl. Acad. Sci. USA 2004, 101, 10042–10047. [Google Scholar] [CrossRef]
- van der Horst, A.; Tertoolen, L.G.; de Vries-Smits, L.M.; Frye, R.A.; Medema, R.H.; Burgering, B.M. FOXO4 is acetylated upon peroxide stress and deacetylated by the longevity protein hSir2(SIRT1). J. Biol. Chem. 2004, 279, 28873–28879. [Google Scholar] [CrossRef] [PubMed]
- Brunet, A.; Sweeney, L.B.; Sturgill, J.F.; Chua, K.F.; Greer, P.L.; Lin, Y.; Tran, H.; Ross, S.E.; Mostoslavsky, R.; Cohen, H.Y.; et al. Stress-dependent regulation of FOXO transcription factors by the SIRT1 deacetylase. Science 2004, 303, 2011–2015. [Google Scholar] [CrossRef] [PubMed]
- Frescas, D.; Valenti, L.; Accili, D. Nuclear trapping of the forkhead transcription factor FoxO1 via Sirt-dependent deacetylation promotes expression of glucogenetic genes. J. Biol. Chem. 2005, 280, 20589–20595. [Google Scholar] [CrossRef] [PubMed]
- Accili, D.; Arden, K.C. FoxOs at the crossroads of cellular metabolism, differentiation, and transformation. Cell 2004, 117, 421–426. [Google Scholar] [CrossRef]
- Martins, R.; Lithgow, G.J.; Link, W. Long live FOXO: Unraveling the role of FOXO proteins in aging and longevity. Aging Cell 2016, 15, 196–207. [Google Scholar] [CrossRef]
- Gross, D.N.; van den Heuvel, A.P.; Birnbaum, M.J. The role of FoxO in the regulation of metabolism. Oncogene 2008, 27, 2320–2336. [Google Scholar] [CrossRef]
- Canto, C.; Gerhart-Hines, Z.; Feige, J.N.; Lagouge, M.; Noriega, L.; Milne, J.C.; Elliott, P.J.; Puigserver, P.; Auwerx, J. AMPK regulates energy expenditure by modulating NAD+ metabolism and SIRT1 activity. Nature 2009, 458, 1056–1060. [Google Scholar] [CrossRef]
- Fulco, M.; Cen, Y.; Zhao, P.; Hoffman, E.P.; McBurney, M.W.; Sauve, A.A.; Sartorelli, V. Glucose restriction inhibits skeletal myoblast differentiation by activating SIRT1 through AMPK-mediated regulation of Nampt. Dev. Cell 2008, 14, 661–673. [Google Scholar] [CrossRef]
- Hou, X.; Xu, S.; Maitland-Toolan, K.A.; Sato, K.; Jiang, B.; Ido, Y.; Lan, F.; Walsh, K.; Wierzbicki, M.; Verbeuren, T.J.; et al. SIRT1 regulates hepatocyte lipid metabolism through activating AMP-activated protein kinase. J. Biol. Chem. 2008, 283, 20015–20026. [Google Scholar] [CrossRef]
- Lan, F.; Cacicedo, J.M.; Ruderman, N.; Ido, Y. SIRT1 modulation of the acetylation status, cytosolic localization, and activity of LKB1. Possible role in AMP-activated protein kinase activation. J. Biol. Chem. 2008, 283, 27628–27635. [Google Scholar] [CrossRef]
- Asher, G.; Gatfield, D.; Stratmann, M.; Reinke, H.; Dibner, C.; Kreppel, F.; Mostoslavsky, R.; Alt, F.W.; Schibler, U. SIRT1 regulates circadian clock gene expression through PER2 deacetylation. Cell 2008, 134, 317–328. [Google Scholar] [CrossRef] [PubMed]
- Rutter, J.; Reick, M.; Wu, L.C.; McKnight, S.L. Regulation of clock and NPAS2 DNA binding by the redox state of NAD cofactors. Science 2001, 293, 510–514. [Google Scholar] [CrossRef] [PubMed]
- Oike, H.; Kobori, M. Resveratrol regulates circadian clock genes in Rat-1 fibroblast cells. Biosci. Biotechnol. Biochem. 2008, 72, 3038–3040. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Wang, Y.; Song, Y.; Cheng, X.R.; Xia, S.; Rahman, M.R.; Shi, Y.; Le, G. Resveratrol restores the circadian rhythmic disorder of lipid metabolism induced by high-fat diet in mice. Biochem. Biophys. Res. Commun. 2015, 458, 86–91. [Google Scholar] [CrossRef]
- Miranda, J.; Portillo, M.P.; Madrid, J.A.; Arias, N.; Macarulla, M.T.; Garaulet, M. Effects of resveratrol on changes induced by high-fat feeding on clock genes in rats. Br. J. Nutr. 2013, 110, 1421–1428. [Google Scholar] [CrossRef]
- Nakahata, Y.; Kaluzova, M.; Grimaldi, B.; Sahar, S.; Hirayama, J.; Chen, D.; Guarente, L.P.; Sassone-Corsi, P. The NAD+-dependent deacetylase SIRT1 modulates CLOCK-mediated chromatin remodeling and circadian control. Cell 2008, 134, 329–340. [Google Scholar] [CrossRef]
- Ramsey, K.M.; Yoshino, J.; Brace, C.S.; Abrassart, D.; Kobayashi, Y.; Marcheva, B.; Hong, H.K.; Chong, J.L.; Buhr, E.D.; Lee, C.; et al. Circadian clock feedback cycle through NAMPT-mediated NAD+ biosynthesis. Science 2009, 324, 651–654. [Google Scholar] [CrossRef]
- Nakahata, Y.; Sahar, S.; Astarita, G.; Kaluzova, M.; Sassone-Corsi, P. Circadian control of the NAD+ salvage pathway by CLOCK-SIRT1. Science 2009, 324, 654–657. [Google Scholar] [CrossRef]
- Peek, C.B.; Affinati, A.H.; Ramsey, K.M.; Kuo, H.Y.; Yu, W.; Sena, L.A.; Ilkayeva, O.; Marcheva, B.; Kobayashi, Y.; Omura, C.; et al. Circadian clock NAD+ cycle drives mitochondrial oxidative metabolism in mice. Science 2013, 342, 1243417. [Google Scholar] [CrossRef]
- Sahar, S.; Masubuchi, S.; Eckel-Mahan, K.; Vollmer, S.; Galla, L.; Ceglia, N.; Masri, S.; Barth, T.K.; Grimaldi, B.; Oluyemi, O.; et al. Circadian control of fatty acid elongation by SIRT1 protein-mediated deacetylation of acetyl-coenzyme A synthetase 1. J. Biol. Chem. 2014, 289, 6091–6097. [Google Scholar] [CrossRef]
- Hallows, W.C.; Lee, S.; Denu, J.M. Sirtuins deacetylate and activate mammalian acetyl-CoA synthetases. Proc. Natl. Acad. Sci. USA 2006, 103, 10230–10235. [Google Scholar] [CrossRef] [PubMed]
- Nuclear Receptors Nomenclature Committee. A unified nomenclature system for the nuclear receptor superfamily. Cell 1999, 97, 161–163. [Google Scholar] [CrossRef]
- Feige, J.N.; Gelman, L.; Michalik, L.; Desvergne, B.; Wahli, W. From molecular action to physiological outputs: Peroxisome proliferator-activated receptors are nuclear receptors at the crossroads of key cellular functions. Prog. Lipid. Res. 2006, 45, 120–159. [Google Scholar] [CrossRef] [PubMed]
- Michalik, L.; Auwerx, J.; Berger, J.P.; Chatterjee, V.K.; Glass, C.K.; Gonzalez, F.J.; Grimaldi, P.A.; Kadowaki, T.; Lazar, M.A.; O’Rahilly, S.; et al. International Union of Pharmacology. LXI. Peroxisome proliferator-activated receptors. Pharmacol. Rev. 2006, 58, 726–741. [Google Scholar] [CrossRef]
- Moller, D.E.; Berger, J.P. Role of PPARs in the regulation of obesity-related insulin sensitivity and inflammation. Int. J. Obes. Relat. Metab. Disord. 2003, 27 (Suppl. S3), S17–S21. [Google Scholar] [CrossRef]
- Tan, C.K.; Zhuang, Y.; Wahli, W. Synthetic and natural Peroxisome Proliferator-Activated Receptor (PPAR) agonists as candidates for the therapy of the metabolic syndrome. Expert. Opin. Ther. Targets 2017, 21, 333–348. [Google Scholar] [CrossRef]
- Cheng, H.S.; Tan, W.R.; Low, Z.S.; Marvalim, C.; Lee, J.Y.H.; Tan, N.S. Exploration and development of PPAR modulators in health and disease: An update of clinical evidence. Int. J. Mol. Sci. 2019, 20, 5055. [Google Scholar] [CrossRef]
- Desvergne, B.; Wahli, W. Peroxisome proliferator-activated receptors: Nuclear control of metabolism. Endocr. Rev. 1999, 20, 649–688. [Google Scholar] [CrossRef]
- Green, S.; Wahli, W. Peroxisome proliferator-activated receptors: Finding the orphan a home. Mol. Cell Endocrinol. 1994, 100, 149–153. [Google Scholar] [CrossRef]
- Masternak, M.M.; Bartke, A. PPARs in calorie restricted and genetically long-lived mice. PPAR Res. 2007, 2007, 28436. [Google Scholar] [CrossRef]
- Yang, Y.B.; Wu, X.L.; Ke, B.; Huang, Y.J.; Chen, S.Q.; Su, Y.Q.; Qin, J. Effects of caloric restriction on peroxisome proliferator-activated receptors and positive transcription elongation factor b expression in obese rats. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 4369–4378. [Google Scholar] [PubMed]
- Masternak, M.M.; Al-Regaiey, K.A.; Del Rosario Lim, M.M.; Jimenez-Ortega, V.; Panici, J.A.; Bonkowski, M.S.; Kopchick, J.J.; Wang, Z.; Bartke, A. Caloric restriction and growth hormone receptor knockout: Effects on expression of genes involved in insulin action in the heart. Exp. Gerontol. 2006, 41, 417–429. [Google Scholar] [CrossRef] [PubMed]
- Masternak, M.M.; Al-Regaiey, K.A.; Del Rosario Lim, M.M.; Jimenez-Ortega, V.; Panici, J.A.; Bonkowski, M.S.; Kopchick, J.J.; Bartke, A. Effects of caloric restriction and growth hormone resistance on the expression level of peroxisome proliferator-activated receptors superfamily in liver of normal and long-lived growth hormone receptor/binding protein knockout mice. J. Gerontol. A Biol. Sci. Med. Sci. 2005, 60, 1394–1398. [Google Scholar] [CrossRef] [PubMed]
- Masternak, M.M.; Al-Regaiey, K.; Bonkowski, M.S.; Panici, J.; Sun, L.; Wang, J.; Przybylski, G.K.; Bartke, A. Divergent effects of caloric restriction on gene expression in normal and long-lived mice. J. Gerontol. A Biol. Sci. Med. Sci. 2004, 59, 784–788. [Google Scholar] [CrossRef] [PubMed]
- Masternak, M.M.; Al-Regaiey, K.A.; Del Rosario Lim, M.M.; Bonkowski, M.S.; Panici, J.A.; Przybylski, G.K.; Bartke, A. Caloric restriction results in decreased expression of peroxisome proliferator-activated receptor superfamily in muscle of normal and long-lived growth hormone receptor/binding protein knockout mice. J. Gerontol. A Biol. Sci. Med. Sci. 2005, 60, 1238–1245. [Google Scholar] [CrossRef]
- Poynter, M.E.; Daynes, R.A. Peroxisome proliferator-activated receptor alpha activation modulates cellular redox status, represses nuclear factor-kappaB signaling, and reduces inflammatory cytokine production in aging. J. Biol. Chem. 1998, 273, 32833–32841. [Google Scholar] [CrossRef]
- Duszka, K.; Ellero-Simatos, S.; Ow, G.S.; Defernez, M.; Paramalingam, E.; Tett, A.; Ying, S.; Konig, J.; Narbad, A.; Kuznetsov, V.A.; et al. Complementary intestinal mucosa and microbiota responses to caloric restriction. Sci. Rep. 2018, 8, 11338. [Google Scholar] [CrossRef]
- Dhahbi, J.M.; Tsuchiya, T.; Kim, H.J.; Mote, P.L.; Spindler, S.R. Gene expression and physiologic responses of the heart to the initiation and withdrawal of caloric restriction. J. Gerontol. A Biol. Sci. Med. Sci. 2006, 61, 218–231. [Google Scholar] [CrossRef]
- Sung, B.; Park, S.; Yu, B.P.; Chung, H.Y. Modulation of PPAR in aging, inflammation, and calorie restriction. J. Gerontol. A Biol. Sci. Med. Sci. 2004, 59, 997–1006. [Google Scholar] [CrossRef]
- Nakamura, K.; Inoue, I.; Takahashi, S.; Komoda, T.; Katayama, S. Cryptochrome and period proteins are regulated by the CLOCK/BMAL1 Gene: Crosstalk between the PPARs/RXR alpha-regulated and CLOCK/BMAL1-regulated systems. PPAR Res. 2008, 2008, 348610. [Google Scholar] [CrossRef]
- Braissant, O.; Foufelle, F.; Scotto, C.; Dauca, M.; Wahli, W. Differential expression of peroxisome proliferator-activated receptors (PPARs): Tissue distribution of PPAR-alpha, -beta, and -gamma in the adult rat. Endocrinology 1996, 137, 354–366. [Google Scholar] [CrossRef] [PubMed]
- Corton, J.C.; Apte, U.; Anderson, S.P.; Limaye, P.; Yoon, L.; Latendresse, J.; Dunn, C.; Everitt, J.I.; Voss, K.A.; Swanson, C.; et al. Mimetics of caloric restriction include agonists of lipid-activated nuclear receptors. J. Biol. Chem. 2004, 279, 46204–46212. [Google Scholar] [CrossRef] [PubMed]
- Peeters, A.; Baes, M. Role of PPAR in hepatic carbohydrate metabolism. PPAR Res. 2010, 2010, 572405. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Xiao, G.; Trujillo, C.; Chang, V.; Blanco, L.; Joseph, S.B.; Bassilian, S.; Saad, M.F.; Tontonoz, P.; Lee, W.N.; et al. Peroxisome proliferator-activated receptor alpha (PPARalpha) influences substrate utilization for hepatic glucose production. J. Biol. Chem. 2002, 277, 50237–50244. [Google Scholar] [CrossRef] [PubMed]
- Mandard, S.; Stienstra, R.; Escher, P.; Tan, N.S.; Kim, I.; Gonzalez, F.J.; Wahli, W.; Desvergne, B.; Muller, M.; Kersten, S. Glycogen synthase 2 is a novel target gene of peroxisome proliferator-activated receptors. Cell Mol. Life Sci. 2007, 64, 1145–1157. [Google Scholar] [CrossRef] [PubMed]
- Finck, B.N.; Lehman, J.J.; Leone, T.C.; Welch, M.J.; Bennett, M.J.; Kovacs, A.; Han, X.; Gross, R.W.; Kozak, R.; Lopaschuk, G.D.; et al. The cardiac phenotype induced by PPARalpha overexpression mimics that caused by diabetes mellitus. J. Clin. Investig. 2002, 109, 121–130. [Google Scholar] [CrossRef] [PubMed]
- Gremlich, S.; Nolan, C.; Roduit, R.; Burcelin, R.; Peyot, M.L.; Delghingaro-Augusto, V.; Desvergne, B.; Michalik, L.; Prentki, M.; Wahli, W. Pancreatic islet adaptation to fasting is dependent on peroxisome proliferator-activated receptor alpha transcriptional up-regulation of fatty acid oxidation. Endocrinology 2005, 146, 375–382. [Google Scholar] [CrossRef]
- Ravnskjaer, K.; Boergesen, M.; Rubi, B.; Larsen, J.K.; Nielsen, T.; Fridriksson, J.; Maechler, P.; Mandrup, S. Peroxisome proliferator-activated receptor alpha (PPARalpha) potentiates, whereas PPARgamma attenuates, glucose-stimulated insulin secretion in pancreatic beta-cells. Endocrinology 2005, 146, 3266–3276. [Google Scholar] [CrossRef]
- Maida, A.; Lamont, B.J.; Cao, X.; Drucker, D.J. Metformin regulates the incretin receptor axis via a pathway dependent on peroxisome proliferator-activated receptor-alpha in mice. Diabetologia 2011, 54, 339–349. [Google Scholar] [CrossRef]
- Sugden, M.C.; Greenwood, G.K.; Smith, N.D.; Holness, M.J. Peroxisome proliferator-activated receptor-alpha activation during pregnancy attenuates glucose-stimulated insulin hypersecretion in vivo by increasing insulin sensitivity, without impairing pregnancy-induced increases in beta-cell glucose sensing and responsiveness. Endocrinology 2003, 144, 146–153. [Google Scholar] [CrossRef][Green Version]
- Murakami, K.; Nambu, S.; Koh, H.; Kobayashi, M.; Shigeta, Y. Clofibrate enhances the affinity of insulin receptors in non-insulin dependent diabetes mellitus. Br. J. Clin. Pharmacol. 1984, 17, 89–91. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, M.; Shigeta, Y.; Hirata, Y.; Omori, Y.; Sakamoto, N.; Nambu, S.; Baba, S. Improvement of glucose tolerance in NIDDM by clofibrate. Randomized double-blind study. Diabetes Care 1988, 11, 495–499. [Google Scholar] [CrossRef]
- Guerre-Millo, M.; Gervois, P.; Raspe, E.; Madsen, L.; Poulain, P.; Derudas, B.; Herbert, J.M.; Winegar, D.A.; Willson, T.M.; Fruchart, J.C.; et al. Peroxisome proliferator-activated receptor alpha activators improve insulin sensitivity and reduce adiposity. J. Biol. Chem. 2000, 275, 16638–16642. [Google Scholar] [CrossRef] [PubMed]
- Aasum, E.; Belke, D.D.; Severson, D.L.; Riemersma, R.A.; Cooper, M.; Andreassen, M.; Larsen, T.S. Cardiac function and metabolism in Type 2 diabetic mice after treatment with BM 17.0744, a novel PPAR-alpha activator. Am. J. Physiol. Heart Circ. Physiol. 2002, 283, H949–H957. [Google Scholar] [CrossRef] [PubMed]
- Patsouris, D.; Mandard, S.; Voshol, P.J.; Escher, P.; Tan, N.S.; Havekes, L.M.; Koenig, W.; Marz, W.; Tafuri, S.; Wahli, W.; et al. PPARalpha governs glycerol metabolism. J. Clin. Investig. 2004, 114, 94–103. [Google Scholar] [CrossRef] [PubMed]
- Lalloyer, F.; Vandewalle, B.; Percevault, F.; Torpier, G.; Kerr-Conte, J.; Oosterveer, M.; Paumelle, R.; Fruchart, J.C.; Kuipers, F.; Pattou, F.; et al. Peroxisome proliferator-activated receptor alpha improves pancreatic adaptation to insulin resistance in obese mice and reduces lipotoxicity in human islets. Diabetes 2006, 55, 1605–1613. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sugden, M.C.; Bulmer, K.; Gibbons, G.F.; Knight, B.L.; Holness, M.J. Peroxisome-proliferator-activated receptor-alpha (PPARalpha) deficiency leads to dysregulation of hepatic lipid and carbohydrate metabolism by fatty acids and insulin. Biochem. J. 2002, 364, 361–368. [Google Scholar] [CrossRef] [PubMed]
- Kersten, S.; Seydoux, J.; Peters, J.M.; Gonzalez, F.J.; Desvergne, B.; Wahli, W. Peroxisome proliferator-activated receptor alpha mediates the adaptive response to fasting. J. Clin. Investig. 1999, 103, 1489–1498. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, T.; Cook, W.S.; Qi, C.; Yeldandi, A.V.; Reddy, J.K.; Rao, M.S. Defect in peroxisome proliferator-activated receptor alpha-inducible fatty acid oxidation determines the severity of hepatic steatosis in response to fasting. J. Biol. Chem. 2000, 275, 28918–28928. [Google Scholar] [CrossRef] [PubMed]
- Leone, T.C.; Weinheimer, C.J.; Kelly, D.P. A critical role for the peroxisome proliferator-activated receptor alpha (PPARalpha) in the cellular fasting response: The PPARalpha-null mouse as a model of fatty acid oxidation disorders. Proc. Natl. Acad. Sci. USA 1999, 96, 7473–7478. [Google Scholar] [CrossRef] [PubMed]
- Fu, J.; Gaetani, S.; Oveisi, F.; Verme, J.L.; Serrano, A.; De Fonseca, F.R.; Rosengarth, A.; Luecke, H.; Di Giacomo, B.; Tarzia, G.; et al. Oleylethanolamide regulates feeding and body weight through activation of the nuclear receptor PPAR-α. Nat. Cell Biol. 2003, 425, 90–93. [Google Scholar] [CrossRef] [PubMed]
- Fu, J.; DiPatrizio, N.V.; Guijarro, A.; Schwartz, G.J.; Li, X.; Gaetani, S.; Astarita, G.; Piomelli, D. Sympathetic activity controls fat-induced oleoylethanolamide signaling in small intestine. J. Neurosci. 2011, 31, 5730–5736. [Google Scholar] [CrossRef] [PubMed]
- Magotti, P.; Bauer, I.; Igarashi, M.; Babagoli, M.; Marotta, R.; Piomelli, D.; Garau, G. Structure of Human N -Acylphosphatidylethanolamine-Hydrolyzing Phospholipase D: Regulation of Fatty Acid Ethanolamide Biosynthesis by Bile Acids. Structure 2015, 23, 598–604. [Google Scholar] [CrossRef] [PubMed]
- Gaetani, S.; Oveisi, F.; Piomelli, D. Modulation of Meal Pattern in the Rat by the Anorexic Lipid Mediator Oleoylethanolamide. Neuropsychopharmacology 2003, 28, 1311–1316. [Google Scholar] [CrossRef] [PubMed]
- Fu, J.; Oveisi, F.; Gaetani, S.; Lin, E.; Piomelli, D. Oleoylethanolamide, an endogenous PPAR-α agonist, lowers body weight and hyperlipidemia in obese rats. Neuropharmacology 2005, 48, 1147–1153. [Google Scholar] [CrossRef] [PubMed]
- Caillon, A.; Duszka, K.; Wahli, W.; Rohner-Jeanrenaud, F.; Altirriba, J. The OEA effect on food intake is independent from the presence of PPARα in the intestine and the nodose ganglion, while the impact of OEA on energy expenditure requires the presence of PPARα in mice. Metabolism 2018, 87, 13–17. [Google Scholar] [CrossRef] [PubMed]
- De Fonseca, F.R.; Navarro, M.; Gómez, R.; Escuredo, L.; Nava, F.; Fu, J.; Murillo-Rodríguez, E.; Giuffrida, A.; LoVerme, J.; Gaetani, S.; et al. An anorexic lipid mediator regulated by feeding. Nat. Cell Biol. 2001, 414, 209–212. [Google Scholar] [CrossRef]
- Gaetani, S.; Fu, J.; Cassano, T.; DiPasquale, P.; Romano, A.; Righetti, L.; Cianci, S.; Laconca, L.; Giannini, E.; Scaccianoce, S.; et al. The fat-induced satiety factor oleoylethanolamide suppresses feeding through central release of oxytocin. J. Neurosci. 2010, 30, 8096–8101. [Google Scholar] [CrossRef]
- Hankir, M.K.; Seyfried, F.; Hintschich, C.A.; Diep, T.-A.; Kleberg, K.; Kranz, M.; Deuther-Conrad, W.; Tellez, L.A.; Rullmann, M.; Patt, M.; et al. Gastric Bypass Surgery Recruits a Gut PPAR-α-Striatal D1R Pathway to Reduce Fat Appetite in Obese Rats. Cell Metab. 2017, 25, 335–344. [Google Scholar] [CrossRef]
- Pan, W.; Liu, C.; Zhang, J.; Gao, X.; Yu, S.; Tan, H.; Yu, J.; Qian, D.; Li, J.; Bian, S.; et al. Association Between Single Nucleotide Polymorphisms in PPARA and EPAS1 Genes and High-Altitude Appetite Loss in Chinese Young Men. Front. Physiol. 2019, 10. [Google Scholar] [CrossRef]
- Chakravarthy, M.V.; Zhu, Y.; López, M.; Yin, L.; Wozniak, D.F.; Coleman, T.; Hu, Z.; Wolfgang, M.; Vidal-Puig, A.; Lane, M.D.; et al. Brain fatty acid synthase activates PPARα to maintain energy homeostasis. J. Clin. Investig. 2007, 117, 2539–2552. [Google Scholar] [CrossRef] [PubMed]
- Iroz, A.; Montagner, A.; Benhamed, F.; Levavasseur, F.; Polizzi, A.; Anthony, E.; Régnier, M.; Fouché, E.; Lukowicz, C.; Cauzac, M.; et al. A Specific ChREBP and PPARα Cross-Talk Is Required for the Glucose-Mediated FGF21 Response. Cell Rep. 2017, 21, 403–416. [Google Scholar] [CrossRef] [PubMed]
- Laeger, T.; Henagan, T.M.; Albarado, D.C.; Redman, L.M.; Bray, G.A.; Noland, R.C.; Münzberg, H.; Hutson, S.M.; Gettys, T.W.; Schwartz, M.W.; et al. FGF21 is an endocrine signal of protein restriction. J. Clin. Investig. 2014, 124, 3913–3922. [Google Scholar] [CrossRef] [PubMed]
- Sengupta, S.; Peterson, T.R.; Laplante, M.; Oh, S.; Sabatini, D.M. mTORC1 controls fasting-induced ketogenesis and its modulation by ageing. Nature 2010, 468, 1100–1104. [Google Scholar] [CrossRef] [PubMed]
- Pawlak, M.; Lefebvre, P.; Staels, B. Molecular mechanism of PPARalpha action and its impact on lipid metabolism, inflammation and fibrosis in non-alcoholic fatty liver disease. J. Hepatol. 2015, 62, 720–733. [Google Scholar] [CrossRef] [PubMed]
- Pentinmikko, N.; Iqbal, S.; Mana, M.; Andersson, S.; Cognetta, A.B., 3rd; Suciu, R.M.; Roper, J.; Luopajarvi, K.; Markelin, E.; Gopalakrishnan, S.; et al. Notum produced by Paneth cells attenuates regeneration of aged intestinal epithelium. Nature 2019, 571, 398–402. [Google Scholar] [CrossRef] [PubMed]
- Okuda, Y.; Kawai, K.; Yamashita, K. Age-related change in ketone body metabolism: Diminished glucagon effect on ketogenesis in adult rats. Endocrinology 1987, 120, 2152–2157. [Google Scholar] [CrossRef]
- Sanguino, E.; Ramon, M.; Michalik, L.; Wahli, W.; Alegret, M.; Sanchez, R.M.; Vazquez-Carrera, M.; Laguna, J.C. Lack of hypotriglyceridemic effect of gemfibrozil as a consequence of age-related changes in rat liver PPARalpha. Biochem. Pharmacol. 2004, 67, 157–166. [Google Scholar] [CrossRef]
- Sastre, J.; Pallardo, F.V.; Pla, R.; Pellin, A.; Juan, G.; O’Connor, J.E.; Estrela, J.M.; Miquel, J.; Vina, J. Aging of the liver: Age-associated mitochondrial damage in intact hepatocytes. Hepatology 1996, 24, 1199–1205. [Google Scholar] [CrossRef]
- Jiao, M.; Ren, F.; Zhou, L.; Zhang, X.; Zhang, L.; Wen, T.; Wei, L.; Wang, X.; Shi, H.; Bai, L.; et al. Peroxisome proliferator-activated receptor alpha activation attenuates the inflammatory response to protect the liver from acute failure by promoting the autophagy pathway. Cell Death Dis. 2014, 5, e1397. [Google Scholar] [CrossRef]
- Kim, Y.S.; Lee, H.M.; Kim, J.K.; Yang, C.S.; Kim, T.S.; Jung, M.; Jin, H.S.; Kim, S.; Jang, J.; Oh, G.T.; et al. PPAR-alpha Activation Mediates Innate Host Defense through Induction of TFEB and Lipid Catabolism. J. Immunol. 2017, 198, 3283–3295. [Google Scholar] [CrossRef] [PubMed]
- Hardie, D.G.; Carling, D.; Carlson, M. The AMP-activated/SNF1 protein kinase subfamily: Metabolic sensors of the eukaryotic cell? Annu. Rev. Biochem. 1998, 67, 821–855. [Google Scholar] [CrossRef] [PubMed]
- Viollet, B.; Andreelli, F. AMP-activated protein kinase and metabolic control. Handb. Exp. Pharmacol. 2011, 13, 303–330. [Google Scholar] [CrossRef]
- Lee, W.J.; Kim, M.; Park, H.S.; Kim, H.S.; Jeon, M.J.; Oh, K.S.; Koh, E.H.; Won, J.C.; Kim, M.S.; Oh, G.T.; et al. AMPK activation increases fatty acid oxidation in skeletal muscle by activating PPARalpha and PGC-1. Biochem. Biophys. Res. Commun. 2006, 340, 291–295. [Google Scholar] [CrossRef] [PubMed]
- Yoon, M.J.; Lee, G.Y.; Chung, J.J.; Ahn, Y.H.; Hong, S.H.; Kim, J.B. Adiponectin increases fatty acid oxidation in skeletal muscle cells by sequential activation of AMP-activated protein kinase, p38 mitogen-activated protein kinase, and peroxisome proliferator-activated receptor alpha. Diabetes 2006, 55, 2562–2570. [Google Scholar] [CrossRef] [PubMed]
- Bai, F.; Liu, Y.; Tu, T.; Li, B.; Xiao, Y.; Ma, Y.; Qin, F.; Xie, J.; Zhou, S.; Liu, Q. Metformin regulates lipid metabolism in a canine model of atrial fibrillation through AMPK/PPAR-alpha/VLCAD pathway. Lipids Health Dis. 2019, 18, 109. [Google Scholar] [CrossRef]
- Joly, E.; Roduit, R.; Peyot, M.L.; Habinowski, S.A.; Ruderman, N.B.; Witters, L.A.; Prentki, M. Glucose represses PPARalpha gene expression via AMP-activated protein kinase but not via p38 mitogen-activated protein kinase in the pancreatic beta-cell. J. Diabetes 2009, 1, 263–272. [Google Scholar] [CrossRef]
- Ravnskjaer, K.; Boergesen, M.; Dalgaard, L.T.; Mandrup, S. Glucose-induced repression of PPARalpha gene expression in pancreatic beta-cells involves PP2A activation and AMPK inactivation. J. Mol. Endocrinol. 2006, 36, 289–299. [Google Scholar] [CrossRef]
- Bronner, M.; Hertz, R.; Bar-Tana, J. Kinase-independent transcriptional co-activation of peroxisome proliferator-activated receptor alpha by AMP-activated protein kinase. Biochem. J. 2004, 384, 295–305. [Google Scholar] [CrossRef]
- Manickam, R.; Duszka, K.; Wahli, W. PPARs and Microbiota in Skeletal Muscle Health and Wasting. Int. J. Mol. Sci. 2020, 21, 8056. [Google Scholar] [CrossRef]
- Chanda, D.; Lee, C.H.; Kim, Y.H.; Noh, J.R.; Kim, D.K.; Park, J.H.; Hwang, J.H.; Lee, M.R.; Jeong, K.H.; Lee, I.K.; et al. Fenofibrate differentially regulates plasminogen activator inhibitor-1 gene expression via adenosine monophosphate-activated protein kinase-dependent induction of orphan nuclear receptor small heterodimer partner. Hepatology 2009, 50, 880–892. [Google Scholar] [CrossRef] [PubMed]
- Liangpunsakul, S.; Wou, S.E.; Wineinger, K.D.; Zeng, Y.; Cyganek, I.; Jayaram, H.N.; Crabb, D.W. Effects of WY-14,643 on the phosphorylation and activation of AMP-dependent protein kinase. Arch. Biochem. Biophys. 2009, 485, 10–15. [Google Scholar] [CrossRef] [PubMed]
- Ren, G.; Rimando, A.M.; Mathews, S.T. AMPK activation by pterostilbene contributes to suppression of hepatic gluconeogenic gene expression and glucose production in H4IIE cells. Biochem. Biophys. Res. Commun. 2018, 498, 640–645. [Google Scholar] [CrossRef] [PubMed]
- Purushotham, A.; Schug, T.T.; Xu, Q.; Surapureddi, S.; Guo, X.; Li, X. Hepatocyte-specific deletion of SIRT1 alters fatty acid metabolism and results in hepatic steatosis and inflammation. Cell Metab. 2009, 9, 327–338. [Google Scholar] [CrossRef] [PubMed]
- Oka, S.; Alcendor, R.; Zhai, P.; Park, J.Y.; Shao, D.; Cho, J.; Yamamoto, T.; Tian, B.; Sadoshima, J. PPARalpha-Sirt1 complex mediates cardiac hypertrophy and failure through suppression of the ERR transcriptional pathway. Cell Metab. 2011, 14, 598–611. [Google Scholar] [CrossRef]
- Oka, S.; Zhai, P.; Alcendor, R.; Park, J.Y.; Tian, B.; Sadoshima, J. Suppression of ERR targets by a PPARalpha/Sirt1 complex in the failing heart. Cell Cycle 2012, 11, 856–864. [Google Scholar] [CrossRef][Green Version]
- Planavila, A.; Iglesias, R.; Giralt, M.; Villarroya, F. Sirt1 acts in association with PPARalpha to protect the heart from hypertrophy, metabolic dysregulation, and inflammation. Cardiovasc. Res. 2011, 90, 276–284. [Google Scholar] [CrossRef]
- Yang, X.; Downes, M.; Yu, R.T.; Bookout, A.L.; He, W.; Straume, M.; Mangelsdorf, D.J.; Evans, R.M. Nuclear receptor expression links the circadian clock to metabolism. Cell 2006, 126, 801–810. [Google Scholar] [CrossRef]
- Lemberger, T.; Saladin, R.; Vazquez, M.; Assimacopoulos, F.; Staels, B.; Desvergne, B.; Wahli, W.; Auwerx, J. Expression of the peroxisome proliferator-activated receptor alpha gene is stimulated by stress and follows a diurnal rhythm. J. Biol. Chem. 1996, 271, 1764–1769. [Google Scholar] [CrossRef]
- Canaple, L.; Rambaud, J.; Dkhissi-Benyahya, O.; Rayet, B.; Tan, N.S.; Michalik, L.; Delaunay, F.; Wahli, W.; Laudet, V. Reciprocal regulation of brain and muscle Arnt-like protein 1 and peroxisome proliferator-activated receptor alpha defines a novel positive feedback loop in the rodent liver circadian clock. Mol. Endocrinol. 2006, 20, 1715–1727. [Google Scholar] [CrossRef]
- Oishi, K.; Shirai, H.; Ishida, N. CLOCK is involved in the circadian transactivation of peroxisome-proliferator-activated receptor alpha (PPARalpha) in mice. Biochem. J. 2005, 386, 575–581. [Google Scholar] [CrossRef] [PubMed]
- Gervois, P.; Chopin-Delannoy, S.; Fadel, A.; Dubois, G.; Kosykh, V.; Fruchart, J.C.; Najib, J.; Laudet, V.; Staels, B. Fibrates increase human REV-ERBalpha expression in liver via a novel peroxisome proliferator-activated receptor response element. Mol. Endocrinol. 1999, 13, 400–409. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Schmutz, I.; Ripperger, J.A.; Baeriswyl-Aebischer, S.; Albrecht, U. The mammalian clock component PERIOD2 coordinates circadian output by interaction with nuclear receptors. Genes Dev. 2010, 24, 345–357. [Google Scholar] [CrossRef] [PubMed]
- Shirai, H.; Oishi, K.; Kudo, T.; Shibata, S.; Ishida, N. PPARalpha is a potential therapeutic target of drugs to treat circadian rhythm sleep disorders. Biochem. Biophys. Res. Commun. 2007, 357, 679–682. [Google Scholar] [CrossRef] [PubMed]
- Oishi, K.; Shirai, H.; Ishida, N. PPARalpha is involved in photoentrainment of the circadian clock. Neuroreport 2008, 19, 487–489. [Google Scholar] [CrossRef]
- Raspe, E.; Duez, H.; Mansen, A.; Fontaine, C.; Fievet, C.; Fruchart, J.C.; Vennstrom, B.; Staels, B. Identification of Rev-erbalpha as a physiological repressor of apoC-III gene transcription. J. Lipid Res. 2002, 43, 2172–2179. [Google Scholar] [CrossRef]
- Turek, F.W.; Joshu, C.; Kohsaka, A.; Lin, E.; Ivanova, G.; McDearmon, E.; Laposky, A.; Losee-Olson, S.; Easton, A.; Jensen, D.R.; et al. Obesity and metabolic syndrome in circadian Clock mutant mice. Science 2005, 308, 1043–1045. [Google Scholar] [CrossRef]
- Grimaldi, B.; Bellet, M.M.; Katada, S.; Astarita, G.; Hirayama, J.; Amin, R.H.; Granneman, J.G.; Piomelli, D.; Leff, T.; Sassone-Corsi, P. PER2 controls lipid metabolism by direct regulation of PPARgamma. Cell Metab. 2010, 12, 509–520. [Google Scholar] [CrossRef]
- Dallmann, R.; Viola, A.U.; Tarokh, L.; Cajochen, C.; Brown, S.A. The human circadian metabolome. Proc. Natl. Acad. Sci. USA 2012, 109, 2625–2629. [Google Scholar] [CrossRef]
- Chua, E.C.; Shui, G.; Lee, I.T.; Lau, P.; Tan, L.C.; Yeo, S.C.; Lam, B.D.; Bulchand, S.; Summers, S.A.; Puvanendran, K.; et al. Extensive diversity in circadian regulation of plasma lipids and evidence for different circadian metabolic phenotypes in humans. Proc. Natl. Acad. Sci. USA 2013, 110, 14468–14473. [Google Scholar] [CrossRef]
- Pan, X.; Hussain, M.M. Diurnal regulation of microsomal triglyceride transfer protein and plasma lipid levels. J. Biol. Chem. 2007, 282, 24707–24719. [Google Scholar] [CrossRef] [PubMed]
- Adamovich, Y.; Rousso-Noori, L.; Zwighaft, Z.; Neufeld-Cohen, A.; Golik, M.; Kraut-Cohen, J.; Wang, M.; Han, X.; Asher, G. Circadian clocks and feeding time regulate the oscillations and levels of hepatic triglycerides. Cell Metab. 2014, 19, 319–330. [Google Scholar] [CrossRef] [PubMed]
- Zvonic, S.; Ptitsyn, A.A.; Conrad, S.A.; Scott, L.K.; Floyd, Z.E.; Kilroy, G.; Wu, X.; Goh, B.C.; Mynatt, R.L.; Gimble, J.M. Characterization of peripheral circadian clocks in adipose tissues. Diabetes 2006, 55, 962–970. [Google Scholar] [CrossRef] [PubMed]
- Barger, P.M.; Kelly, D.P. PPAR signaling in the control of cardiac energy metabolism. Trends Cardiovasc. Med. 2000, 10, 238–245. [Google Scholar] [CrossRef]
- Wu, X.; Liu, Z.; Shi, G.; Xing, L.; Wang, X.; Gu, X.; Qu, Z.; Dong, Z.; Xiong, J.; Gao, X.; et al. The circadian clock influences heart performance. J. Biol. Rhythms 2011, 26, 402–411. [Google Scholar] [CrossRef] [PubMed]
- Oishi, K.; Uchida, D.; Ishida, N. Circadian expression of FGF21 is induced by PPARalpha activation in the mouse liver. FEBS Lett. 2008, 582, 3639–3642. [Google Scholar] [CrossRef]
- Patel, D.D.; Knight, B.L.; Wiggins, D.; Humphreys, S.M.; Gibbons, G.F. Disturbances in the normal regulation of SREBP-sensitive genes in PPAR alpha-deficient mice. J. Lipid Res. 2001, 42, 328–337. [Google Scholar]
- Girroir, E.E.; Hollingshead, H.E.; He, P.; Zhu, B.; Perdew, G.H.; Peters, J.M. Quantitative expression patterns of peroxisome proliferator-activated receptor-beta/delta (PPARbeta/delta) protein in mice. Biochem. Biophys. Res. Commun 2008, 371, 456–461. [Google Scholar] [CrossRef]
- Braissant, O.; Wahli, W. Differential expression of peroxisome proliferator-activated receptor-alpha, -beta, and -gamma during rat embryonic development. Endocrinology 1998, 139, 2748–2754. [Google Scholar] [CrossRef]
- Tan, N.S.; Michalik, L.; Desvergne, B.; Wahli, W. Peroxisome proliferator-activated receptor (PPAR)-beta as a target for wound healing drugs: What is possible? Am. J. Clin. Dermatol. 2003, 4, 523–530. [Google Scholar] [CrossRef]
- Di-Poi, N.; Tan, N.S.; Michalik, L.; Wahli, W.; Desvergne, B. Antiapoptotic role of PPARbeta in keratinocytes via transcriptional control of the Akt1 signaling pathway. Mol. Cell 2002, 10, 721–733. [Google Scholar] [CrossRef]
- Letavernier, E.; Perez, J.; Joye, E.; Bellocq, A.; Fouqueray, B.; Haymann, J.P.; Heudes, D.; Wahli, W.; Desvergne, B.; Baud, L. Peroxisome proliferator-activated receptor beta/delta exerts a strong protection from ischemic acute renal failure. J. Am. Soc. Nephrol. 2005, 16, 2395–2402. [Google Scholar] [CrossRef] [PubMed]
- Michalik, L.; Wahli, W. Involvement of PPAR nuclear receptors in tissue injury and wound repair. J. Clin. Investig. 2006, 116, 598–606. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.X.; Lee, C.H.; Tiep, S.; Yu, R.T.; Ham, J.; Kang, H.; Evans, R.M. Peroxisome-proliferator-activated receptor delta activates fat metabolism to prevent obesity. Cell 2003, 113, 159–170. [Google Scholar] [CrossRef]
- Nadra, K.; Anghel, S.I.; Joye, E.; Tan, N.S.; Basu-Modak, S.; Trono, D.; Wahli, W.; Desvergne, B. Differentiation of trophoblast giant cells and their metabolic functions are dependent on peroxisome proliferator-activated receptor beta/delta. Mol. Cell Biol. 2006, 26, 3266–3281. [Google Scholar] [CrossRef]
- Varnat, F.; Heggeler, B.B.; Grisel, P.; Boucard, N.; Corthesy-Theulaz, I.; Wahli, W.; Desvergne, B. PPARbeta/delta regulates paneth cell differentiation via controlling the hedgehog signaling pathway. Gastroenterology 2006, 131, 538–553. [Google Scholar] [CrossRef] [PubMed]
- Doktorova, M.; Zwarts, I.; Zutphen, T.V.; Dijk, T.H.; Bloks, V.W.; Harkema, L.; Bruin, A.; Downes, M.; Evans, R.M.; Verkade, H.J.; et al. Intestinal PPARdelta protects against diet-induced obesity, insulin resistance and dyslipidemia. Sci. Rep. 2017, 7, 846. [Google Scholar] [CrossRef] [PubMed]
- Dreyer, C.; Krey, G.; Keller, H.; Givel, F.; Helftenbein, G.; Wahli, W. Control of the peroxisomal beta-oxidation pathway by a novel family of nuclear hormone receptors. Cell 1992, 68, 879–887. [Google Scholar] [CrossRef]
- Daoudi, M.; Hennuyer, N.; Borland, M.G.; Touche, V.; Duhem, C.; Gross, B.; Caiazzo, R.; Kerr-Conte, J.; Pattou, F.; Peters, J.M.; et al. PPARbeta/delta activation induces enteroendocrine L cell GLP-1 production. Gastroenterology 2011, 140, 1564–1574. [Google Scholar] [CrossRef]
- Schuler, M.; Ali, F.; Chambon, C.; Duteil, D.; Bornert, J.M.; Tardivel, A.; Desvergne, B.; Wahli, W.; Chambon, P.; Metzger, D. PGC1alpha expression is controlled in skeletal muscles by PPARbeta, whose ablation results in fiber-type switching, obesity, and type 2 diabetes. Cell Metab. 2006, 4, 407–414. [Google Scholar] [CrossRef]
- Liu, S.; Brown, J.D.; Stanya, K.J.; Homan, E.; Leidl, M.; Inouye, K.; Bhargava, P.; Gangl, M.R.; Dai, L.; Hatano, B.; et al. A diurnal serum lipid integrates hepatic lipogenesis and peripheral fatty acid use. Nature 2013, 502, 550–554. [Google Scholar] [CrossRef] [PubMed]
- Philp, A.; MacKenzie, M.G.; Belew, M.Y.; Towler, M.C.; Corstorphine, A.; Papalamprou, A.; Hardie, D.G.; Baar, K. Glycogen content regulates peroxisome proliferator activated receptor- partial differential (PPAR- partial differential) activity in rat skeletal muscle. PLoS ONE 2013, 8, e77200. [Google Scholar] [CrossRef] [PubMed]
- Manickam, R.; Wahli, W. Roles of Peroxisome Proliferator-Activated Receptor beta/delta in skeletal muscle physiology. Biochimie 2017, 136, 42–48. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.X.; Zhang, C.L.; Yu, R.T.; Cho, H.K.; Nelson, M.C.; Bayuga-Ocampo, C.R.; Ham, J.; Kang, H.; Evans, R.M. Regulation of muscle fiber type and running endurance by PPAR delta. PLoS Biol. 2004, 2, e294. [Google Scholar] [CrossRef] [PubMed]
- Kocalis, H.E.; Turney, M.K.; Printz, R.L.; Laryea, G.N.; Muglia, L.J.; Davies, S.S.; Stanwood, G.D.; McGuinness, O.P.; Niswender, K.D. Neuron-Specific Deletion of Peroxisome Proliferator-Activated Receptor Delta (PPARδ) in Mice Leads to Increased Susceptibility to Diet-Induced Obesity. PLoS ONE 2012, 7, e42981. [Google Scholar] [CrossRef] [PubMed]
- Foster, D.A. Phosphatidic acid and lipid-sensing by mTOR. Trends Endocrinol. Metab. 2013, 24, 272–278. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Ritzenthaler, J.D.; Zhong, X.; Zheng, Y.; Roman, J.; Han, S. Nicotine stimulates PPARbeta/delta expression in human lung carcinoma cells through activation of PI3K/mTOR and suppression of AP-2 alpha. Cancer Res. 2009, 69, 6445–6453. [Google Scholar] [CrossRef] [PubMed]
- Han, S.; Ritzenthaler, J.D.; Zheng, Y.; Roman, J. PPARbeta/delta agonist stimulates human lung carcinoma cell growth through inhibition of PTEN expression: The involvement of PI3K and NF-kappa B signals. Am. J. Physiol. Lung Cell Mol. Physiol. 2008, 294, L1238–L1249. [Google Scholar] [CrossRef] [PubMed]
- Oliver, W.R., Jr.; Shenk, J.L.; Snaith, M.R.; Russell, C.S.; Plunket, K.D.; Bodkin, N.L.; Lewis, M.C.; Winegar, D.A.; Sznaidman, M.L.; Lambert, M.H.; et al. A selective peroxisome proliferator-activated receptor delta agonist promotes reverse cholesterol transport. Proc. Natl. Acad. Sci. USA 2001, 98, 5306–5311. [Google Scholar] [CrossRef]
- Tanaka, T.; Yamamoto, J.; Iwasaki, S.; Asaba, H.; Hamura, H.; Ikeda, Y.; Watanabe, M.; Magoori, K.; Ioka, R.X.; Tachibana, K.; et al. Activation of peroxisome proliferator-activated receptor delta induces fatty acid beta-oxidation in skeletal muscle and attenuates metabolic syndrome. Proc. Natl. Acad. Sci. USA 2003, 100, 15924–15929. [Google Scholar] [CrossRef]
- Riserus, U.; Sprecher, D.; Johnson, T.; Olson, E.; Hirschberg, S.; Liu, A.; Fang, Z.; Hegde, P.; Richards, D.; Sarov-Blat, L.; et al. Activation of peroxisome proliferator-activated receptor (PPAR)delta promotes reversal of multiple metabolic abnormalities, reduces oxidative stress, and increases fatty acid oxidation in moderately obese men. Diabetes 2008, 57, 332–339. [Google Scholar] [CrossRef] [PubMed]
- Cohen, G.; Riahi, Y.; Shamni, O.; Guichardant, M.; Chatgilialoglu, C.; Ferreri, C.; Kaiser, N.; Sasson, S. Role of lipid peroxidation and PPAR-delta in amplifying glucose-stimulated insulin secretion. Diabetes 2011, 60, 2830–2842. [Google Scholar] [CrossRef] [PubMed]
- Iglesias, J.; Barg, S.; Vallois, D.; Lahiri, S.; Roger, C.; Yessoufou, A.; Pradevand, S.; McDonald, A.; Bonal, C.; Reimann, F.; et al. PPARbeta/delta affects pancreatic beta cell mass and insulin secretion in mice. J. Clin. Investig. 2012, 122, 4105–4117. [Google Scholar] [CrossRef] [PubMed]
- Serrano-Marco, L.; Rodriguez-Calvo, R.; El Kochairi, I.; Palomer, X.; Michalik, L.; Wahli, W.; Vazquez-Carrera, M. Activation of peroxisome proliferator-activated receptor-beta/-delta (PPAR-beta/-delta) ameliorates insulin signaling and reduces SOCS3 levels by inhibiting STAT3 in interleukin-6-stimulated adipocytes. Diabetes 2011, 60, 1990–1999. [Google Scholar] [CrossRef] [PubMed]
- Serrano-Marco, L.; Barroso, E.; El Kochairi, I.; Palomer, X.; Michalik, L.; Wahli, W.; Vazquez-Carrera, M. The peroxisome proliferator-activated receptor (PPAR) beta/delta agonist GW501516 inhibits IL-6-induced signal transducer and activator of transcription 3 (STAT3) activation and insulin resistance in human liver cells. Diabetologia 2012, 55, 743–751. [Google Scholar] [CrossRef]
- Salvado, L.; Barroso, E.; Gomez-Foix, A.M.; Palomer, X.; Michalik, L.; Wahli, W.; Vazquez-Carrera, M. PPARbeta/delta prevents endoplasmic reticulum stress-associated inflammation and insulin resistance in skeletal muscle cells through an AMPK-dependent mechanism. Diabetologia 2014, 57, 2126–2135. [Google Scholar] [CrossRef]
- Lee, C.H.; Olson, P.; Hevener, A.; Mehl, I.; Chong, L.W.; Olefsky, J.M.; Gonzalez, F.J.; Ham, J.; Kang, H.; Peters, J.M.; et al. PPARdelta regulates glucose metabolism and insulin sensitivity. Proc. Natl. Acad. Sci. USA 2006, 103, 3444–3449. [Google Scholar] [CrossRef]
- Gan, Z.; Burkart-Hartman, E.M.; Han, D.H.; Finck, B.; Leone, T.C.; Smith, E.Y.; Ayala, J.E.; Holloszy, J.; Kelly, D.P. The nuclear receptor PPARbeta/delta programs muscle glucose metabolism in cooperation with AMPK and MEF2. Genes Dev. 2011, 25, 2619–2630. [Google Scholar] [CrossRef] [PubMed]
- Manio, M.C.; Inoue, K.; Fujitani, M.; Matsumura, S.; Fushiki, T. Combined pharmacological activation of AMPK and PPARdelta potentiates the effects of exercise in trained mice. Physiol Rep. 2016, 4. [Google Scholar] [CrossRef]
- Narkar, V.A.; Downes, M.; Yu, R.T.; Embler, E.; Wang, Y.X.; Banayo, E.; Mihaylova, M.M.; Nelson, M.C.; Zou, Y.; Juguilon, H.; et al. AMPK and PPARdelta agonists are exercise mimetics. Cell 2008, 134, 405–415. [Google Scholar] [CrossRef]
- Okazaki, M.; Iwasaki, Y.; Nishiyama, M.; Taguchi, T.; Tsugita, M.; Nakayama, S.; Kambayashi, M.; Hashimoto, K.; Terada, Y. PPARbeta/delta regulates the human SIRT1 gene transcription via Sp1. Endocr. J. 2010, 57, 403–413. [Google Scholar] [CrossRef] [PubMed]
- Barroso, E.; Eyre, E.; Palomer, X.; Vazquez-Carrera, M. The peroxisome proliferator-activated receptor beta/delta (PPARbeta/delta) agonist GW501516 prevents TNF-alpha-induced NF-kappaB activation in human HaCaT cells by reducing p65 acetylation through AMPK and SIRT1. Biochem. Pharmacol. 2011, 81, 534–543. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.Y.; Kang, E.S.; Ham, S.A.; Hwang, J.S.; Yoo, T.S.; Lee, H.; Paek, K.S.; Park, C.; Lee, H.T.; Kim, J.H.; et al. The PPARdelta-mediated inhibition of angiotensin II-induced premature senescence in human endothelial cells is SIRT1-dependent. Biochem. Pharmacol. 2012, 84, 1627–1634. [Google Scholar] [CrossRef] [PubMed]
- Gong, K.; Qu, B.; Wang, C.; Zhou, J.; Liao, D.; Zheng, W.; Pan, X. Peroxisome proliferator-activated receptor alpha facilitates osteogenic differentiation in MC3T3-E1 cells via the sirtuin 1-dependent signaling pathway. Mol. Cells 2017, 40, 393–400. [Google Scholar] [CrossRef] [PubMed]
- Gong, K.; Qu, B.; Liao, D.; Liu, D.; Wang, C.; Zhou, J.; Pan, X. MiR-132 regulates osteogenic differentiation via downregulating Sirtuin1 in a peroxisome proliferator-activated receptor beta/delta-dependent manner. Biochem. Biophys. Res. Commun. 2016, 478, 260–267. [Google Scholar] [CrossRef] [PubMed]
- Qu, B.; Ma, Y.; Yan, M.; Gong, K.; Liang, F.; Deng, S.; Jiang, K.; Ma, Z.; Pan, X. Sirtuin1 promotes osteogenic differentiation through downregulation of peroxisome proliferator-activated receptor gamma in MC3T3-E1 cells. Biochem. Biophys. Res. Commun. 2016, 478, 439–445. [Google Scholar] [CrossRef] [PubMed]
- Gatfield, D.; Le Martelot, G.; Vejnar, C.E.; Gerlach, D.; Schaad, O.; Fleury-Olela, F.; Ruskeepaa, A.L.; Oresic, M.; Esau, C.C.; Zdobnov, E.M.; et al. Integration of microRNA miR-122 in hepatic circadian gene expression. Genes Dev. 2009, 23, 1313–1326. [Google Scholar] [CrossRef]
- Challet, E.; Denis, I.; Rochet, V.; Aioun, J.; Gourmelen, S.; Lacroix, H.; Goustard-Langelier, B.; Papillon, C.; Alessandri, J.M.; Lavialle, M. The role of PPARbeta/delta in the regulation of glutamatergic signaling in the hamster suprachiasmatic nucleus. Cell Mol. Life Sci. 2013, 70, 2003–2014. [Google Scholar] [CrossRef]
- Anghel, S.I.; Wahli, W. Fat poetry: A kingdom for PPAR gamma. Cell Res. 2007, 17, 486–511. [Google Scholar] [CrossRef]
- Lehmann, J.M.; Moore, L.B.; Smith-Oliver, T.A.; Wilkison, W.O.; Willson, T.M.; Kliewer, S.A. An antidiabetic thiazolidinedione is a high affinity ligand for peroxisome proliferator-activated receptor gamma (PPAR gamma). J. Biol. Chem. 1995, 270, 12953–12956. [Google Scholar] [CrossRef]
- Imai, T.; Takakuwa, R.; Marchand, S.; Dentz, E.; Bornert, J.M.; Messaddeq, N.; Wendling, O.; Mark, M.; Desvergne, B.; Wahli, W.; et al. Peroxisome proliferator-activated receptor gamma is required in mature white and brown adipocytes for their survival in the mouse. Proc. Natl. Acad. Sci. USA 2004, 101, 4543–4547. [Google Scholar] [CrossRef] [PubMed]
- Cerbone, A.; Toaldo, C.; Laurora, S.; Briatore, F.; Pizzimenti, S.; Dianzani, M.U.; Ferretti, C.; Barrera, G. 4-Hydroxynonenal and PPARgamma ligands affect proliferation, differentiation, and apoptosis in colon cancer cells. Free Radic. Biol. Med. 2007, 42, 1661–1670. [Google Scholar] [CrossRef] [PubMed]
- Martinasso, G.; Oraldi, M.; Trombetta, A.; Maggiora, M.; Bertetto, O.; Canuto, R.A.; Muzio, G. Involvement of PPARs in cell proliferation and apoptosis in human colon cancer specimens and in normal and cancer cell lines. PPAR Res. 2007, 2007, 93416. [Google Scholar] [CrossRef] [PubMed]
- Theocharis, S.; Margeli, A.; Vielh, P.; Kouraklis, G. Peroxisome proliferator-activated receptor-gamma ligands as cell-cycle modulators. Cancer Treat. Rev. 2004, 30, 545–554. [Google Scholar] [CrossRef] [PubMed]
- Xu, W.P.; Zhang, X.; Xie, W.F. Differentiation therapy for solid tumors. J. Dig. Dis. 2014, 15, 159–165. [Google Scholar] [CrossRef]
- Chen, G.G.; Lee, J.F.; Wang, S.H.; Chan, U.P.; Ip, P.C.; Lau, W.Y. Apoptosis induced by activation of peroxisome-proliferator activated receptor-gamma is associated with Bcl-2 and NF-kappaB in human colon cancer. Life Sci. 2002, 70, 2631–2646. [Google Scholar] [CrossRef]
- Sharma, C.; Pradeep, A.; Wong, L.; Rana, A.; Rana, B. Peroxisome proliferator-activated receptor gamma activation can regulate beta-catenin levels via a proteasome-mediated and adenomatous polyposis coli-independent pathway. J. Biol. Chem. 2004, 279, 35583–35594. [Google Scholar] [CrossRef]
- Leonardini, A.; Laviola, L.; Perrini, S.; Natalicchio, A.; Giorgino, F. Cross-Talk between PPARgamma and Insulin Signaling and Modulation of Insulin Sensitivity. PPAR Res. 2009, 2009, 818945. [Google Scholar] [CrossRef]
- Tontonoz, P.; Spiegelman, B.M. Fat and beyond: The diverse biology of PPARgamma. Annu. Rev. Biochem. 2008, 77, 289–312. [Google Scholar] [CrossRef]
- Duszka, K.; Oresic, M.; Le May, C.; Konig, J.; Wahli, W. PPAR gamma modulates long chain fatty acid processing in the intestinal epithelium. Int. J. Mol. Sci. 2017, 18, 2559. [Google Scholar] [CrossRef]
- Duszka, K.; Picard, A.; Ellero-Simatos, S.; Chen, J.; Defernez, M.; Paramalingam, E.; Pigram, A.; Vanoaica, L.; Canlet, C.; Parini, P.; et al. Intestinal PPARgamma signalling is required for sympathetic nervous system activation in response to caloric restriction. Sci. Rep. 2016, 6, 36937. [Google Scholar] [CrossRef] [PubMed]
- Soccio, R.E.; Chen, E.R.; Lazar, M.A. Thiazolidinediones and the promise of insulin sensitization in type 2 diabetes. Cell Metab. 2014, 20, 573–591. [Google Scholar] [CrossRef] [PubMed]
- Henke, B.R. Peroxisome proliferator-activated receptor alpha/gamma dual agonists for the treatment of type 2 diabetes. J. Med. Chem. 2004, 47, 4118–4127. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.H.; Hong, S.H.; Park, Y.J.; Sung, J.H.; Suh, W.; Lee, K.W.; Jung, K.; Lim, C.; Kim, J.H.; Kim, H.; et al. MD001, a novel peroxisome proliferator-activated receptor alpha/gamma agonist, improves glucose and lipid metabolism. Sci. Rep. 2019, 9, 1656. [Google Scholar] [CrossRef] [PubMed]
- Jain, M.R.; Giri, S.R.; Bhoi, B.; Trivedi, C.; Rath, A.; Rathod, R.; Ranvir, R.; Kadam, S.; Patel, H.; Swain, P.; et al. Dual PPARalpha/gamma agonist saroglitazar improves liver histopathology and biochemistry in experimental NASH models. Liver Int. 2018, 38, 1084–1094. [Google Scholar] [CrossRef] [PubMed]
- Marion-Letellier, R.; Dechelotte, P.; Iacucci, M.; Ghosh, S. Dietary modulation of peroxisome proliferator-activated receptor gamma. Gut 2009, 58, 586–593. [Google Scholar] [CrossRef] [PubMed]
- Schwab, M.; Reynders, V.; Loitsch, S.; Steinhilber, D.; Stein, J.; Schroder, O. Involvement of different nuclear hormone receptors in butyrate-mediated inhibition of inducible NF kappa B signalling. Mol. Immunol. 2007, 44, 3625–3632. [Google Scholar] [CrossRef]
- Nepelska, M.; de Wouters, T.; Jacouton, E.; Beguet-Crespel, F.; Lapaque, N.; Dore, J.; Arulampalam, V.; Blottiere, H.M. Commensal gut bacteria modulate phosphorylation-dependent PPARgamma transcriptional activity in human intestinal epithelial cells. Sci. Rep. 2017, 7, 43199. [Google Scholar] [CrossRef]
- Voltan, S.; Martines, D.; Elli, M.; Brun, P.; Longo, S.; Porzionato, A.; Macchi, V.; D’Inca, R.; Scarpa, M.; Palu, G.; et al. Lactobacillus crispatus M247-derived H2O2 acts as a signal transducing molecule activating peroxisome proliferator activated receptor-gamma in the intestinal mucosa. Gastroenterology 2008, 135, 1216–1227. [Google Scholar] [CrossRef]
- Are, A.; Aronsson, L.; Wang, S.; Greicius, G.; Lee, Y.K.; Gustafsson, J.A.; Pettersson, S.; Arulampalam, V. Enterococcus faecalis from newborn babies regulate endogenous PPARgamma activity and IL-10 levels in colonic epithelial cells. Proc. Natl. Acad. Sci. USA 2008, 105, 1943–1948. [Google Scholar] [CrossRef]
- Couvigny, B.; de Wouters, T.; Kaci, G.; Jacouton, E.; Delorme, C.; Dore, J.; Renault, P.; Blottiere, H.M.; Guedon, E.; Lapaque, N. Commensal streptococcus salivarius modulates PPAR gamma transcriptional activity in human intestinal epithelial cells. PLoS ONE 2015, 10, e0125371. [Google Scholar] [CrossRef] [PubMed]
- Lu, M.; Sarruf, D.A.; Talukdar, S.; Sharma, S.; Li, P.; Bandyopadhyay, G.; Nalbandian, S.; Fan, W.; Gayen, J.R.; Mahata, S.K.; et al. Brain PPAR-γ promotes obesity and is required for the insulin–sensitizing effect of thiazolidinediones. Nat. Med. 2011, 17, 618–622. [Google Scholar] [CrossRef] [PubMed]
- Ryan, K.K.; Li, B.; Grayson, B.E.; Matter, E.K.; Woods, S.C.; Seeley, R.J. A role for central nervous system PPAR-γ in the regulation of energy balance. Nat. Med. 2011, 17, 623–626. [Google Scholar] [CrossRef] [PubMed]
- Koleva, D.I.; Orbetzova, M.M.; Atanassova, P.K. Adipose Tissue Hormones and Appetite and Body Weight Regulators an Insulin Resistance/Гормоны Жировой Ткани И Регуляторы Аппетита И Массы Тела При Инсулиновой Резистентности. Folia Med. 2013, 55, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Adamo, K.B.; Dent, R.; Langefeld, C.D.; Cox, M.; Williams, K.; Carrick, K.M.; Stuart, J.S.; Sundseth, S.S.; Harper, M.E.; McPherson, R.; et al. Peroxisome proliferator-activated receptor gamma 2 and acyl-CoA synthetase 5 polymorphisms influence diet response. Obesity 2007, 15, 1068–1075. [Google Scholar] [CrossRef] [PubMed]
- Matsuo, T.; Nakata, Y.; Katayama, Y.; Iemitsu, M.; Maeda, S.; Okura, T.; Kim, M.K.; Ohkubo, H.; Hotta, K.; Tanaka, K. PPARG genotype accounts for part of individual variation in body weight reduction in response to calorie restriction. Obesity 2009, 17, 1924–1931. [Google Scholar] [CrossRef]
- Deeb, S.S.; Fajas, L.; Nemoto, M.; Pihlajamaki, J.; Mykkanen, L.; Kuusisto, J.; Laakso, M.; Fujimoto, W.; Auwerx, J. A Pro12Ala substitution in PPARgamma2 associated with decreased receptor activity, lower body mass index and improved insulin sensitivity. Nat. Genet. 1998, 20, 284–287. [Google Scholar] [CrossRef]
- Masugi, J.; Tamori, Y.; Mori, H.; Koike, T.; Kasuga, M. Inhibitory effect of a proline-to-alanine substitution at codon 12 of peroxisome proliferator-activated receptor-gamma 2 on thiazolidinedione-induced adipogenesis. Biochem. Biophys. Res. Commun. 2000, 268, 178–182. [Google Scholar] [CrossRef]
- Nicklas, B.J.; van Rossum, E.F.; Berman, D.M.; Ryan, A.S.; Dennis, K.E.; Shuldiner, A.R. Genetic variation in the peroxisome proliferator-activated receptor-gamma2 gene (Pro12Ala) affects metabolic responses to weight loss and subsequent weight regain. Diabetes 2001, 50, 2172–2176. [Google Scholar] [CrossRef]
- Memisoglu, A.; Hu, F.B.; Hankinson, S.E.; Manson, J.E.; De Vivo, I.; Willett, W.C.; Hunter, D.J. Interaction between a peroxisome proliferator-activated receptor gamma gene polymorphism and dietary fat intake in relation to body mass. Hum. Mol. Genet. 2003, 12, 2923–2929. [Google Scholar] [CrossRef]
- Robitaille, J.; Despres, J.P.; Perusse, L.; Vohl, M.C. The PPAR-gamma P12A polymorphism modulates the relationship between dietary fat intake and components of the metabolic syndrome: Results from the Quebec Family Study. Clin. Genet. 2003, 63, 109–116. [Google Scholar] [CrossRef] [PubMed]
- Luan, J.; Browne, P.O.; Harding, A.H.; Halsall, D.J.; O’Rahilly, S.; Chatterjee, V.K.; Wareham, N.J. Evidence for gene-nutrient interaction at the PPARgamma locus. Diabetes 2001, 50, 686–689. [Google Scholar] [CrossRef] [PubMed]
- Lindi, V.; Schwab, U.; Louheranta, A.; Laakso, M.; Vessby, B.; Hermansen, K.; Storlien, L.; Riccardi, G.; Rivellese, A.A.; Matti, I.J. Impact of the Pro12Ala polymorphism of the PPAR-gamma2 gene on serum triacylglycerol response to n-3 fatty acid supplementation. Mol. Genet. Metab. 2003, 79, 52–60. [Google Scholar] [CrossRef]
- Nolan, J.J.; Ludvik, B.; Beerdsen, P.; Joyce, M.; Olefsky, J. Improvement in glucose tolerance and insulin resistance in obese subjects treated with troglitazone. N. Engl. J. Med. 1994, 331, 1188–1193. [Google Scholar] [CrossRef] [PubMed]
- Buchanan, T.A.; Xiang, A.H.; Peters, R.K.; Kjos, S.L.; Marroquin, A.; Goico, J.; Ochoa, C.; Tan, S.; Berkowitz, K.; Hodis, H.N.; et al. Preservation of pancreatic beta-cell function and prevention of type 2 diabetes by pharmacological treatment of insulin resistance in high-risk hispanic women. Diabetes 2002, 51, 2796–2803. [Google Scholar] [CrossRef] [PubMed]
- Knowler, W.C.; Hamman, R.F.; Edelstein, S.L.; Barrett-Connor, E.; Ehrmann, D.A.; Walker, E.A.; Fowler, S.E.; Nathan, D.M.; Kahn, S.E.; Diabetes Prevention Program Research. Prevention of type 2 diabetes with troglitazone in the Diabetes Prevention Program. Diabetes 2005, 54, 1150–1156. [Google Scholar] [CrossRef]
- Rocchi, S.; Picard, F.; Vamecq, J.; Gelman, L.; Potier, N.; Zeyer, D.; Dubuquoy, L.; Bac, P.; Champy, M.F.; Plunket, K.D.; et al. A unique PPAR gamma ligand with potent insulin-sensitizing yet weak adipogenic activity. Mol. Cell 2001, 8, 737–747. [Google Scholar] [CrossRef]
- Dunn, F.L.; Higgins, L.S.; Fredrickson, J.; DePaoli, A.M.; INT131-004 Study Group. Selective modulation of PPARgamma activity can lower plasma glucose without typical thiazolidinedione side-effects in patients with Type 2 diabetes. J. Diabetes Complicat. 2011, 25, 151–158. [Google Scholar] [CrossRef]
- Beekmann, K.; Rubio, L.; de Haan, L.H.; Actis-Goretta, L.; van der Burg, B.; van Bladeren, P.J.; Rietjens, I.M. The effect of quercetin and kaempferol aglycones and glucuronides on peroxisome proliferator-activated receptor-gamma (PPAR-gamma). Food Funct. 2015, 6, 1098–1107. [Google Scholar] [CrossRef]
- Fang, X.K.; Gao, J.; Zhu, D.N. Kaempferol and quercetin isolated from Euonymus alatus improve glucose uptake of 3T3-L1 cells without adipogenesis activity. Life Sci. 2008, 82, 615–622. [Google Scholar] [CrossRef]
- Moitra, J.; Mason, M.M.; Olive, M.; Krylov, D.; Gavrilova, O.; Marcus-Samuels, B.; Feigenbaum, L.; Lee, E.; Aoyama, T.; Eckhaus, M.; et al. Life without white fat: A transgenic mouse. Genes Dev. 1998, 12, 3168–3181. [Google Scholar] [CrossRef] [PubMed]
- Reitman, M.L.; Arioglu, E.; Gavrilova, O.; Taylor, S.I. Lipoatrophy revisited. Trends Endocrinol. Metab. 2000, 11, 410–416. [Google Scholar] [CrossRef]
- Larsen, T.M.; Toubro, S.; Astrup, A. PPARgamma agonists in the treatment of type II diabetes: Is increased fatness commensurate with long-term efficacy? Int. J. Obes. Relat. Metab. Disord. 2003, 27, 147–161. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.M.; Doyle, P.J.; Iglesias, M.A.; Watson, D.G.; Cooney, G.J.; Kraegen, E.W. Peroxisome proliferator-activated receptor (PPAR)-alpha activation lowers muscle lipids and improves insulin sensitivity in high fat-fed rats: Comparison with PPAR-gamma activation. Diabetes 2001, 50, 411–417. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, A.K.; Garg, A. A novel heterozygous mutation in peroxisome proliferator-activated receptor-gamma gene in a patient with familial partial lipodystrophy. J. Clin. Endocrinol. Metab. 2002, 87, 408–411. [Google Scholar] [CrossRef] [PubMed]
- Hegele, R.A.; Cao, H.; Frankowski, C.; Mathews, S.T.; Leff, T. PPARG F388L, a transactivation-deficient mutant, in familial partial lipodystrophy. Diabetes 2002, 51, 3586–3590. [Google Scholar] [CrossRef] [PubMed]
- Demir, T.; Onay, H.; Savage, D.B.; Temeloglu, E.; Uzum, A.K.; Kadioglu, P.; Altay, C.; Ozen, S.; Demir, L.; Cavdar, U.; et al. Familial partial lipodystrophy linked to a novel peroxisome proliferator activator receptor -gamma (PPARG) mutation, H449L: A comparison of people with this mutation and those with classic codon 482 Lamin A/C (LMNA) mutations. Diabet Med. 2016, 33, 1445–1450. [Google Scholar] [CrossRef]
- Ludtke, A.; Buettner, J.; Schmidt, H.H.; Worman, H.J. New PPARG mutation leads to lipodystrophy and loss of protein function that is partially restored by a synthetic ligand. J. Med. Genet. 2007, 44, e88. [Google Scholar] [CrossRef]
- Ludtke, A.; Buettner, J.; Wu, W.; Muchir, A.; Schroeter, A.; Zinn-Justin, S.; Spuler, S.; Schmidt, H.H.; Worman, H.J. Peroxisome proliferator-activated receptor-gamma C190S mutation causes partial lipodystrophy. J. Clin. Endocrinol. Metab. 2007, 92, 2248–2255. [Google Scholar] [CrossRef]
- Barroso, I.; Gurnell, M.; Crowley, V.E.; Agostini, M.; Schwabe, J.W.; Soos, M.A.; Maslen, G.L.; Williams, T.D.; Lewis, H.; Schafer, A.J.; et al. Dominant negative mutations in human PPARgamma associated with severe insulin resistance, diabetes mellitus and hypertension. Nature 1999, 402, 880–883. [Google Scholar] [CrossRef]
- Savage, D.B.; Tan, G.D.; Acerini, C.L.; Jebb, S.A.; Agostini, M.; Gurnell, M.; Williams, R.L.; Umpleby, A.M.; Thomas, E.L.; Bell, J.D.; et al. Human metabolic syndrome resulting from dominant-negative mutations in the nuclear receptor peroxisome proliferator-activated receptor-gamma. Diabetes 2003, 52, 910–917. [Google Scholar] [CrossRef] [PubMed]
- Collino, M.; Aragno, M.; Castiglia, S.; Miglio, G.; Tomasinelli, C.; Boccuzzi, G.; Thiemermann, C.; Fantozzi, R. Pioglitazone improves lipid and insulin levels in overweight rats on a high cholesterol and fructose diet by decreasing hepatic inflammation. Br. J. Pharmacol. 2010, 160, 1892–1902. [Google Scholar] [CrossRef] [PubMed]
- Iwata, M.; Haruta, T.; Usui, I.; Takata, Y.; Takano, A.; Uno, T.; Kawahara, J.; Ueno, E.; Sasaoka, T.; Ishibashi, O.; et al. Pioglitazone ameliorates tumor necrosis factor-alpha-induced insulin resistance by a mechanism independent of adipogenic activity of peroxisome proliferator–activated receptor-gamma. Diabetes 2001, 50, 1083–1092. [Google Scholar] [CrossRef] [PubMed]
- Smith, U.; Gogg, S.; Johansson, A.; Olausson, T.; Rotter, V.; Svalstedt, B. Thiazolidinediones (PPARgamma agonists) but not PPARalpha agonists increase IRS-2 gene expression in 3T3-L1 and human adipocytes. FASEB J. 2001, 15, 215–220. [Google Scholar] [CrossRef] [PubMed]
- Rieusset, J.; Auwerx, J.; Vidal, H. Regulation of gene expression by activation of the peroxisome proliferator-activated receptor gamma with rosiglitazone (BRL 49653) in human adipocytes. Biochem. Biophys. Res. Commun. 1999, 265, 265–271. [Google Scholar] [CrossRef] [PubMed]
- Baumann, C.A.; Chokshi, N.; Saltiel, A.R.; Ribon, V. Cloning and characterization of a functional peroxisome proliferator activator receptor-gamma-responsive element in the promoter of the CAP gene. J. Biol. Chem. 2000, 275, 9131–9135. [Google Scholar] [CrossRef]
- Ribon, V.; Johnson, J.H.; Camp, H.S.; Saltiel, A.R. Thiazolidinediones and insulin resistance: Peroxisome proliferatoractivated receptor gamma activation stimulates expression of the CAP gene. Proc. Natl. Acad. Sci. USA 1998, 95, 14751–14756. [Google Scholar] [CrossRef]
- Kramer, D.; Shapiro, R.; Adler, A.; Bush, E.; Rondinone, C.M. Insulin-sensitizing effect of rosiglitazone (BRL-49653) by regulation of glucose transporters in muscle and fat of Zucker rats. Metabolism 2001, 50, 1294–1300. [Google Scholar] [CrossRef]
- Standaert, M.L.; Kanoh, Y.; Sajan, M.P.; Bandyopadhyay, G.; Farese, R.V. Cbl, IRS-1, and IRS-2 mediate effects of rosiglitazone on PI3K, PKC-lambda, and glucose transport in 3T3/L1 adipocytes. Endocrinology 2002, 143, 1705–1716. [Google Scholar] [CrossRef]
- Kim, H.I.; Kim, J.W.; Kim, S.H.; Cha, J.Y.; Kim, K.S.; Ahn, Y.H. Identification and functional characterization of the peroxisomal proliferator response element in rat GLUT2 promoter. Diabetes 2000, 49, 1517–1524. [Google Scholar] [CrossRef]
- Samaha, F.F.; Szapary, P.O.; Iqbal, N.; Williams, M.M.; Bloedon, L.T.; Kochar, A.; Wolfe, M.L.; Rader, D.J. Effects of rosiglitazone on lipids, adipokines, and inflammatory markers in nondiabetic patients with low high-density lipoprotein cholesterol and metabolic syndrome. Arterioscler. Thromb. Vasc. Biol. 2006, 26, 624–630. [Google Scholar] [CrossRef] [PubMed]
- Szalkowski, D.; White-Carrington, S.; Berger, J.; Zhang, B. Antidiabetic thiazolidinediones block the inhibitory effect of tumor necrosis factor-alpha on differentiation, insulin-stimulated glucose uptake, and gene expression in 3T3-L1 cells. Endocrinology 1995, 136, 1474–1481. [Google Scholar] [CrossRef] [PubMed]
- Pajvani, U.B.; Hawkins, M.; Combs, T.P.; Rajala, M.W.; Doebber, T.; Berger, J.P.; Wagner, J.A.; Wu, M.; Knopps, A.; Xiang, A.H.; et al. Complex distribution, not absolute amount of adiponectin, correlates with thiazolidinedione-mediated improvement in insulin sensitivity. J. Biol. Chem. 2004, 279, 12152–12162. [Google Scholar] [CrossRef] [PubMed]
- Yamauchi, T.; Kamon, J.; Waki, H.; Terauchi, Y.; Kubota, N.; Hara, K.; Mori, Y.; Ide, T.; Murakami, K.; Tsuboyama-Kasaoka, N.; et al. The fat-derived hormone adiponectin reverses insulin resistance associated with both lipoatrophy and obesity. Nat. Med. 2001, 7, 941–946. [Google Scholar] [CrossRef] [PubMed]
- Panasyuk, G.; Espeillac, C.; Chauvin, C.; Pradelli, L.A.; Horie, Y.; Suzuki, A.; Annicotte, J.S.; Fajas, L.; Foretz, M.; Verdeguer, F.; et al. PPARgamma contributes to PKM2 and HK2 expression in fatty liver. Nat. Commun. 2012, 3, 672. [Google Scholar] [CrossRef]
- Armoni, M.; Harel, C.; Karni, S.; Chen, H.; Bar-Yoseph, F.; Ver, M.R.; Quon, M.J.; Karnieli, E. FOXO1 represses peroxisome proliferator-activated receptor-gamma1 and -gamma2 gene promoters in primary adipocytes. A novel paradigm to increase insulin sensitivity. J. Biol. Chem. 2006, 281, 19881–19891. [Google Scholar] [CrossRef]
- Fan, W.; Imamura, T.; Sonoda, N.; Sears, D.D.; Patsouris, D.; Kim, J.J.; Olefsky, J.M. FOXO1 transrepresses peroxisome proliferator-activated receptor gamma transactivation, coordinating an insulin-induced feed-forward response in adipocytes. J. Biol. Chem. 2009, 284, 12188–12197. [Google Scholar] [CrossRef]
- Dowell, P.; Otto, T.C.; Adi, S.; Lane, M.D. Convergence of peroxisome proliferator-activated receptor gamma and Foxo1 signaling pathways. J. Biol. Chem. 2003, 278, 45485–45491. [Google Scholar] [CrossRef]
- Zhang, Y.; Gu, M.; Cai, W.; Yu, L.; Feng, L.; Zhang, L.; Zang, Q.; Wang, Y.; Wang, D.; Chen, H.; et al. Dietary component isorhamnetin is a PPARgamma antagonist and ameliorates metabolic disorders induced by diet or leptin deficiency. Sci. Rep. 2016, 6, 19288. [Google Scholar] [CrossRef]
- Yamauchi, T.; Waki, H.; Kamon, J.; Murakami, K.; Motojima, K.; Komeda, K.; Miki, H.; Kubota, N.; Terauchi, Y.; Tsuchida, A.; et al. Inhibition of RXR and PPARgamma ameliorates diet-induced obesity and type 2 diabetes. J. Clin. Investig. 2001, 108, 1001–1013. [Google Scholar] [CrossRef]
- Kubota, N.; Terauchi, Y.; Miki, H.; Tamemoto, H.; Yamauchi, T.; Komeda, K.; Satoh, S.; Nakano, R.; Ishii, C.; Sugiyama, T.; et al. PPAR gamma mediates high-fat diet-induced adipocyte hypertrophy and insulin resistance. Mol. Cell 1999, 4, 597–609. [Google Scholar] [CrossRef]
- Miles, P.D.; Barak, Y.; He, W.; Evans, R.M.; Olefsky, J.M. Improved insulin-sensitivity in mice heterozygous for PPAR-gamma deficiency. J. Clin. Investig. 2000, 105, 287–292. [Google Scholar] [CrossRef] [PubMed]
- Heikkinen, S.; Argmann, C.; Feige, J.N.; Koutnikova, H.; Champy, M.F.; Dali-Youcef, N.; Schadt, E.E.; Laakso, M.; Auwerx, J. The Pro12Ala PPARgamma2 variant determines metabolism at the gene-environment interface. Cell Metab. 2009, 9, 88–98. [Google Scholar] [CrossRef] [PubMed]
- Altshuler, D.; Hirschhorn, J.N.; Klannemark, M.; Lindgren, C.M.; Vohl, M.C.; Nemesh, J.; Lane, C.R.; Schaffner, S.F.; Bolk, S.; Brewer, C.; et al. The common PPARgamma Pro12Ala polymorphism is associated with decreased risk of type 2 diabetes. Nat. Genet. 2000, 26, 76–80. [Google Scholar] [CrossRef] [PubMed]
- Picard, F.; Auwerx, J. PPAR(gamma) and glucose homeostasis. Annu. Rev. Nutr. 2002, 22, 167–197. [Google Scholar] [CrossRef] [PubMed]
- Cho, H.J.; Park, J.; Lee, H.W.; Lee, Y.S.; Kim, J.B. Regulation of adipocyte differentiation and insulin action with rapamycin. Biochem. Biophys. Res. Commun. 2004, 321, 942–948. [Google Scholar] [CrossRef] [PubMed]
- Gagnon, A.; Lau, S.; Sorisky, A. Rapamycin-sensitive phase of 3T3-L1 preadipocyte differentiation after clonal expansion. J. Cell Physiol. 2001, 189, 14–22. [Google Scholar] [CrossRef]
- Polak, P.; Cybulski, N.; Feige, J.N.; Auwerx, J.; Ruegg, M.A.; Hall, M.N. Adipose-specific knockout of raptor results in lean mice with enhanced mitochondrial respiration. Cell Metab. 2008, 8, 399–410. [Google Scholar] [CrossRef]
- Bell, A.; Grunder, L.; Sorisky, A. Rapamycin inhibits human adipocyte differentiation in primary culture. Obes. Res. 2000, 8, 249–254. [Google Scholar] [CrossRef]
- Carnevalli, L.S.; Masuda, K.; Frigerio, F.; Le Bacquer, O.; Um, S.H.; Gandin, V.; Topisirovic, I.; Sonenberg, N.; Thomas, G.; Kozma, S.C. S6K1 plays a critical role in early adipocyte differentiation. Dev. Cell 2010, 18, 763–774. [Google Scholar] [CrossRef]
- Blanchard, P.G.; Festuccia, W.T.; Houde, V.P.; St-Pierre, P.; Brule, S.; Turcotte, V.; Cote, M.; Bellmann, K.; Marette, A.; Deshaies, Y. Major involvement of mTOR in the PPARgamma-induced stimulation of adipose tissue lipid uptake and fat accretion. J. Lipid Res. 2012, 53, 1117–1125. [Google Scholar] [CrossRef] [PubMed]
- Koh, Y.K.; Lee, M.Y.; Kim, J.W.; Kim, M.; Moon, J.S.; Lee, Y.J.; Ahn, Y.H.; Kim, K.S. Lipin1 is a key factor for the maturation and maintenance of adipocytes in the regulatory network with CCAAT/enhancer-binding protein alpha and peroxisome proliferator-activated receptor gamma 2. J. Biol. Chem. 2008, 283, 34896–34906. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.E.; Bae, E.; Jeong, D.Y.; Kim, M.J.; Jin, W.J.; Park, S.W.; Han, G.S.; Carman, G.M.; Koh, E.; Kim, K.S. Lipin1 regulates PPARgamma transcriptional activity. Biochem. J. 2013, 453, 49–60. [Google Scholar] [CrossRef] [PubMed]
- Finck, B.N.; Gropler, M.C.; Chen, Z.; Leone, T.C.; Croce, M.A.; Harris, T.E.; Lawrence, J.C., Jr.; Kelly, D.P. Lipin 1 is an inducible amplifier of the hepatic PGC-1alpha/PPARalpha regulatory pathway. Cell Metab. 2006, 4, 199–210. [Google Scholar] [CrossRef]
- Kim, J.B.; Wright, H.M.; Wright, M.; Spiegelman, B.M. ADD1/SREBP1 activates PPARgamma through the production of endogenous ligand. Proc. Natl. Acad. Sci. USA 1998, 95, 4333–4337. [Google Scholar] [CrossRef]
- Bakan, I.; Laplante, M. Connecting mTORC1 signaling to SREBP-1 activation. Curr. Opin Lipidol. 2012, 23, 226–234. [Google Scholar] [CrossRef]
- Purnell, P.T.; Liu, S.C.; Cardiff, R.; Gregg, J. Induction of PPAR gamma signaling and autophagy as a mechanism mediating acquired rapamycin resistance in breast cancer models. Cancer Res. 2008, 68, 3210. [Google Scholar]
- Assumpcao, J.A.F.; Magalhaes, K.G.; Correa, J.R. The role of ppargamma and autophagy in ros production, lipid droplets biogenesis and its involvement with colorectal cancer cells modulation. Cancer Cell. Int. 2017, 17, 82. [Google Scholar] [CrossRef]
- Kang, J.H.; Chang, Y.C.; Maurizi, M.R. 4-O-carboxymethyl ascochlorin causes ER stress and induced autophagy in human hepatocellular carcinoma cells. J. Biol. Chem. 2012, 287, 15661–15671. [Google Scholar] [CrossRef]
- Pellerito, O.; Notaro, A.; Sabella, S.; De Blasio, A.; Vento, R.; Calvaruso, G.; Giuliano, M. WIN induces apoptotic cell death in human colon cancer cells through a block of autophagic flux dependent on PPARgamma down-regulation. Apoptosis 2014, 19, 1029–1042. [Google Scholar] [CrossRef]
- Weng, J.R.; Bai, L.Y.; Chiu, C.F.; Hu, J.L.; Chiu, S.J.; Wu, C.Y. Cucurbitane triterpenoid from momordica charantia induces apoptosis and autophagy in breast cancer cells, in part, through peroxisome proliferator-activated receptor gamma activation. Evid. Based Complement. Alternat. Med. 2013, 2013, 935675. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Zhang, W.; Liang, B.; Casimiro, M.C.; Whitaker-Menezes, D.; Wang, M.; Lisanti, M.P.; Lanza-Jacoby, S.; Pestell, R.G.; Wang, C. PPARgamma activation induces autophagy in breast cancer cells. Int. J. Biochem. Cell Biol. 2009, 41, 2334–2342. [Google Scholar] [CrossRef] [PubMed]
- Yan, S.; Yang, X.; Chen, T.; Xi, Z.; Jiang, X. The PPARgamma agonist Troglitazone induces autophagy, apoptosis and necroptosis in bladder cancer cells. Cancer Gene Ther. 2014, 21, 188–193. [Google Scholar] [CrossRef] [PubMed]
- Cerquetti, L.; Sampaoli, C.; Amendola, D.; Bucci, B.; Masuelli, L.; Marchese, R.; Misiti, S.; De Venanzi, A.; Poggi, M.; Toscano, V.; et al. Rosiglitazone induces autophagy in H295R and cell cycle deregulation in SW13 adrenocortical cancer cells. Exp. Cell Res. 2011, 317, 1397–1410. [Google Scholar] [CrossRef]
- Xu, F.; Li, J.; Ni, W.; Shen, Y.W.; Zhang, X.P. Peroxisome proliferator-activated receptor-gamma agonist 15d-prostaglandin J2 mediates neuronal autophagy after cerebral ischemia-reperfusion injury. PLoS ONE 2013, 8, e55080. [Google Scholar] [CrossRef]
- Li, H.; Zhang, Q.; Yang, X.; Wang, L. PPAR-gamma agonist rosiglitazone reduces autophagy and promotes functional recovery in experimental traumaticspinal cord injury. Neurosci. Lett. 2017, 650, 89–96. [Google Scholar] [CrossRef] [PubMed]
- Mahmood, D.F.; Jguirim-Souissi, I.; Khadija el, H.; Blondeau, N.; Diderot, V.; Amrani, S.; Slimane, M.N.; Syrovets, T.; Simmet, T.; Rouis, M. Peroxisome proliferator-activated receptor gamma induces apoptosis and inhibits autophagy of human monocyte-derived macrophages via induction of cathepsin L: Potential role in atherosclerosis. J. Biol. Chem. 2011, 286, 28858–28866. [Google Scholar] [CrossRef]
- Lamontagne, J.; Pepin, E.; Peyot, M.L.; Joly, E.; Ruderman, N.B.; Poitout, V.; Madiraju, S.R.; Nolan, C.J.; Prentki, M. Pioglitazone acutely reduces insulin secretion and causes metabolic deceleration of the pancreatic beta-cell at submaximal glucose concentrations. Endocrinology 2009, 150, 3465–3474. [Google Scholar] [CrossRef]
- Boyle, J.G.; Logan, P.J.; Ewart, M.A.; Reihill, J.A.; Ritchie, S.A.; Connell, J.M.; Cleland, S.J.; Salt, I.P. Rosiglitazone stimulates nitric oxide synthesis in human aortic endothelial cells via AMP-activated protein kinase. J. Biol. Chem. 2008, 283, 11210–11217. [Google Scholar] [CrossRef]
- Ceolotto, G.; Gallo, A.; Papparella, I.; Franco, L.; Murphy, E.; Iori, E.; Pagnin, E.; Fadini, G.P.; Albiero, M.; Semplicini, A.; et al. Rosiglitazone reduces glucose-induced oxidative stress mediated by NAD(P)H oxidase via AMPK-dependent mechanism. Arterioscler. Thromb. Vasc. Biol. 2007, 27, 2627–2633. [Google Scholar] [CrossRef]
- Wang, L.; Yin, Y.; Hou, G.; Kang, J.; Wang, Q. Peroxisome Proliferator-Activated Receptor (PPAR gamma) Plays a Protective Role in Cigarette Smoking-Induced Inflammation via AMP-Activated Protein Kinase (AMPK) Signaling. Med. Sci. Monit. 2018, 24, 5168–5177. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Zhu, X.; Lin, W.; Zhou, Y.; Cai, W.; Qiu, L. Interactions of TLR4 and PPAR gamma, dependent on AMPK signalling pathway contribute to anti-inflammatory effects of vaccariae hypaphorine in endothelial cells. Cell Physiol. Biochem. 2017, 42, 1227–1239. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Zhou, L.; Shao, L.; Qian, L.; Fu, X.; Li, G.; Luo, T.; Gu, Y.; Li, F.; Li, J.; et al. Troglitazone acutely activates AMP-activated protein kinase and inhibits insulin secretion from beta cells. Life Sci. 2007, 81, 160–165. [Google Scholar] [CrossRef] [PubMed]
- LeBrasseur, N.K.; Kelly, M.; Tsao, T.S.; Farmer, S.R.; Saha, A.K.; Ruderman, N.B.; Tomas, E. Thiazolidinediones can rapidly activate AMP-activated protein kinase in mammalian tissues. Am. J. Physiol. Endocrinol. Metab. 2006, 291, E175–E181. [Google Scholar] [CrossRef] [PubMed]
- Shen, Z.; Liang, X.; Rogers, C.Q.; Rideout, D.; You, M. Involvement of adiponectin-SIRT1-AMPK signaling in the protective action of rosiglitazone against alcoholic fatty liver in mice. Am. J. Physiol. Gastrointest. Liver Physiol. 2010, 298, G364–G374. [Google Scholar] [CrossRef] [PubMed]
- Saha, A.K.; Avilucea, P.R.; Ye, J.M.; Assifi, M.M.; Kraegen, E.W.; Ruderman, N.B. Pioglitazone treatment activates AMP-activated protein kinase in rat liver and adipose tissue in vivo. Biochem. Biophys. Res. Commun. 2004, 314, 580–585. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.M.; Dzamko, N.; Hoy, A.J.; Iglesias, M.A.; Kemp, B.; Kraegen, E. Rosiglitazone treatment enhances acute AMP-activated protein kinase-mediated muscle and adipose tissue glucose uptake in high-fat-fed rats. Diabetes 2006, 55, 2797–2804. [Google Scholar] [CrossRef]
- Coletta, D.K.; Sriwijitkamol, A.; Wajcberg, E.; Tantiwong, P.; Li, M.; Prentki, M.; Madiraju, M.; Jenkinson, C.P.; Cersosimo, E.; Musi, N.; et al. Pioglitazone stimulates AMP-activated protein kinase signalling and increases the expression of genes involved in adiponectin signalling, mitochondrial function and fat oxidation in human skeletal muscle in vivo: A randomised trial. Diabetologia 2009, 52, 723–732. [Google Scholar] [CrossRef]
- Xiao, X.; Su, G.; Brown, S.N.; Chen, L.; Ren, J.; Zhao, P. Peroxisome proliferator-activated receptors gamma and alpha agonists stimulate cardiac glucose uptake via activation of AMP-activated protein kinase. J. Nutr. Biochem. 2010, 21, 621–626. [Google Scholar] [CrossRef]
- Zhang, F.; Dey, D.; Branstrom, R.; Forsberg, L.; Lu, M.; Zhang, Q.; Sjoholm, A. BLX-1002, a novel thiazolidinedione with no PPAR affinity, stimulates AMP-activated protein kinase activity, raises cytosolic Ca2+, and enhances glucose-stimulated insulin secretion in a PI3K-dependent manner. Am. J. Physiol. Cell Physiol. 2009, 296, C346–C354. [Google Scholar] [CrossRef]
- Chang, T.J.; Chen, W.P.; Yang, C.; Lu, P.H.; Liang, Y.C.; Su, M.J.; Lee, S.C.; Chuang, L.M. Serine-385 phosphorylation of inwardly rectifying K+ channel subunit (Kir6.2) by AMP-dependent protein kinase plays a key role in rosiglitazone-induced closure of the K(ATP) channel and insulin secretion in rats. Diabetologia 2009, 52, 1112–1121. [Google Scholar] [CrossRef] [PubMed]
- Sozio, M.S.; Lu, C.; Zeng, Y.; Liangpunsakul, S.; Crabb, D.W. Activated AMPK inhibits PPAR-{alpha} and PPAR-{gamma} transcriptional activity in hepatoma cells. Am. J. Physiol. Gastrointest. Liver Physiol. 2011, 301, G739–G747. [Google Scholar] [CrossRef] [PubMed]
- Han, L.; Zhou, R.; Niu, J.; McNutt, M.A.; Wang, P.; Tong, T. SIRT1 is regulated by a PPAR {gamma}-SIRT1 negative feedback loop associated with senescence. Nucleic Acids Res. 2010, 38, 7458–7471. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Zhang, Y.; Xiao, F.; Liu, Y.; Wang, J.; Gao, H.; Rong, S.; Yao, Y.; Li, J.; Xu, G. The peroxisome proliferator-activated receptor gamma agonist pioglitazone prevents NF-kappaB activation in cisplatin nephrotoxicity through the reduction of p65 acetylation via the AMPK-SIRT1/p300 pathway. Biochem. Pharmacol. 2016, 101, 100–111. [Google Scholar] [CrossRef] [PubMed]
- Picard, F.; Kurtev, M.; Chung, N.; Topark-Ngarm, A.; Senawong, T.; Machado De Oliveira, R.; Leid, M.; McBurney, M.W.; Guarente, L. Sirt1 promotes fat mobilization in white adipocytes by repressing PPAR-gamma. Nature 2004, 429, 771–776. [Google Scholar] [CrossRef] [PubMed]
- Floyd, Z.E.; Wang, Z.Q.; Kilroy, G.; Cefalu, W.T. Modulation of peroxisome proliferator-activated receptor gamma stability and transcriptional activity in adipocytes by resveratrol. Metabolism 2008, 57, S32–S38. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.; Markan, K.; Temple, K.A.; Deplewski, D.; Brady, M.J.; Cohen, R.N. The nuclear receptor corepressors NCoR and SMRT decrease peroxisome proliferator-activated receptor gamma transcriptional activity and repress 3T3-L1 adipogenesis. J. Biol. Chem. 2005, 280, 13600–13605. [Google Scholar] [CrossRef]
- Wang, N.; Yang, G.; Jia, Z.; Zhang, H.; Aoyagi, T.; Soodvilai, S.; Symons, J.D.; Schnermann, J.B.; Gonzalez, F.J.; Litwin, S.E.; et al. Vascular PPARgamma controls circadian variation in blood pressure and heart rate through Bmal1. Cell Metab. 2008, 8, 482–491. [Google Scholar] [CrossRef]
- Fontaine, C.; Dubois, G.; Duguay, Y.; Helledie, T.; Vu-Dac, N.; Gervois, P.; Soncin, F.; Mandrup, S.; Fruchart, J.C.; Fruchart-Najib, J.; et al. The orphan nuclear receptor Rev-Erbalpha is a peroxisome proliferator-activated receptor (PPAR) gamma target gene and promotes PPARgamma-induced adipocyte differentiation. J. Biol. Chem. 2003, 278, 37672–37680. [Google Scholar] [CrossRef]
- Austin, S.; Medvedev, A.; Yan, Z.H.; Adachi, H.; Hirose, T.; Jetten, A.M. Induction of the nuclear orphan receptor RORgamma during adipocyte differentiation of D1 and 3T3-L1 cells. Cell Growth Differ. 1998, 9, 267–276. [Google Scholar]
- Nakahata, Y.; Akashi, M.; Trcka, D.; Yasuda, A.; Takumi, T. The in vitro real-time oscillation monitoring system identifies potential entrainment factors for circadian clocks. BMC Mol. Biol. 2006, 7, 5. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.; Jia, Z.; Aoyagi, T.; McClain, D.; Mortensen, R.M.; Yang, T. Systemic PPARgamma deletion impairs circadian rhythms of behavior and metabolism. PLoS ONE 2012, 7, e38117. [Google Scholar] [CrossRef]
- Anan, F.; Masaki, T.; Fukunaga, N.; Teshima, Y.; Iwao, T.; Kaneda, K.; Umeno, Y.; Okada, K.; Wakasugi, K.; Yonemochi, H.; et al. Pioglitazone shift circadian rhythm of blood pressure from non-dipper to dipper type in type 2 diabetes mellitus. Eur. J. Clin. Investig. 2007, 37, 709–714. [Google Scholar] [CrossRef] [PubMed]
- Kumar Jha, P.; Challet, E.; Kalsbeek, A. Circadian rhythms in glucose and lipid metabolism in nocturnal and diurnal mammals. Mol. Cell Endocrinol 2015, 418, 74–88. [Google Scholar] [CrossRef]
- Chawla, A.; Lazar, M.A. Induction of Rev-ErbA alpha, an orphan receptor encoded on the opposite strand of the alpha-thyroid hormone receptor gene, during adipocyte differentiation. J. Biol. Chem. 1993, 268, 16265–16269. [Google Scholar]
- Cho, H.; Zhao, X.; Hatori, M.; Yu, R.T.; Barish, G.D.; Lam, M.T.; Chong, L.W.; DiTacchio, L.; Atkins, A.R.; Glass, C.K.; et al. Regulation of circadian behaviour and metabolism by REV-ERB-alpha and REV-ERB-beta. Nature 2012, 485, 123–127. [Google Scholar] [CrossRef]
- Shimba, S.; Ishii, N.; Ohta, Y.; Ohno, T.; Watabe, Y.; Hayashi, M.; Wada, T.; Aoyagi, T.; Tezuka, M. Brain and muscle Arnt-like protein-1 (BMAL1), a component of the molecular clock, regulates adipogenesis. Proc. Natl. Acad. Sci. USA 2005, 102, 12071–12076. [Google Scholar] [CrossRef]
- Guo, B.; Chatterjee, S.; Li, L.; Kim, J.M.; Lee, J.; Yechoor, V.K.; Minze, L.J.; Hsueh, W.; Ma, K. The clock gene, brain and muscle Arnt-like 1, regulates adipogenesis via Wnt signaling pathway. FASEB J. 2012, 26, 3453–3463. [Google Scholar] [CrossRef]
- Douris, N.; Kojima, S.; Pan, X.; Lerch-Gaggl, A.F.; Duong, S.Q.; Hussain, M.M.; Green, C.B. Nocturnin regulates circadian trafficking of dietary lipid in intestinal enterocytes. Curr. Biol. 2011, 21, 1347–1355. [Google Scholar] [CrossRef]
- Katewa, S.D.; Akagi, K.; Bose, N.; Rakshit, K.; Camarella, T.; Zheng, X.; Hall, D.; Davis, S.; Nelson, C.S.; Brem, R.B.; et al. Peripheral circadian clocks mediate dietary restriction-dependent changes in lifespan and fat metabolism in drosophila. Cell Metab. 2016, 23, 143–154. [Google Scholar] [CrossRef]
- Eckel-Mahan, K.L.; Patel, V.R.; de Mateo, S.; Orozco-Solis, R.; Ceglia, N.J.; Sahar, S.; Dilag-Penilla, S.A.; Dyar, K.A.; Baldi, P.; Sassone-Corsi, P. Reprogramming of the circadian clock by nutritional challenge. Cell 2013, 155, 1464–1478. [Google Scholar] [CrossRef] [PubMed]
- Murakami, M.; Tognini, P.; Liu, Y.; Eckel-Mahan, K.L.; Baldi, P.; Sassone-Corsi, P. Gut microbiota directs PPARgamma-driven reprogramming of the liver circadian clock by nutritional challenge. EMBO Rep. 2016, 17, 1292–1303. [Google Scholar] [CrossRef] [PubMed]
- Merry, B.J. Molecular mechanisms linking calorie restriction and longevity. Int. J. Biochem. Cell Biol. 2002, 34, 1340–1354. [Google Scholar] [CrossRef]
- Sanz, A.; Caro, P.; Ibanez, J.; Gomez, J.; Gredilla, R.; Barja, G. Dietary restriction at old age lowers mitochondrial oxygen radical production and leak at complex I and oxidative DNA damage in rat brain. J. Bioenerg. Biomembr. 2005, 37, 83–90. [Google Scholar] [CrossRef] [PubMed]
- Yu, B.P. Aging and oxidative stress: Modulation by dietary restriction. Free Radic. Biol. Med. 1996, 21, 651–668. [Google Scholar] [CrossRef]
- Barja, G. Aging in vertebrates, and the effect of caloric restriction: A mitochondrial free radical production-DNA damage mechanism? Biol. Rev. Camb. Philos. Soc. 2004, 79, 235–251. [Google Scholar] [CrossRef] [PubMed]
- Barja, G. Free radicals and aging. Trends Neurosci. 2004, 27, 595–600. [Google Scholar] [CrossRef]
- Forster, M.J.; Sohal, B.H.; Sohal, R.S. Reversible effects of long-term caloric restriction on protein oxidative damage. J. Gerontol. A Biol. Sci. Med. Sci. 2000, 55, B522–B529. [Google Scholar] [CrossRef]
- Lambert, A.J.; Merry, B.J. Lack of effect of caloric restriction on bioenergetics and reactive oxygen species production in intact rat hepatocytes. J. Gerontol. A Biol. Sci. Med. Sci. 2005, 60, 175–180. [Google Scholar] [CrossRef]
- Lambert, A.J.; Portero-Otin, M.; Pamplona, R.; Merry, B.J. Effect of ageing and caloric restriction on specific markers of protein oxidative damage and membrane peroxidizability in rat liver mitochondria. Mech. Ageing Dev. 2004, 125, 529–538. [Google Scholar] [CrossRef]
- Patel, S.A.; Velingkaar, N.S.; Kondratov, R.V. Transcriptional control of antioxidant defense by the circadian clock. Antioxid. Redox. Signal. 2014, 20, 2997–3006. [Google Scholar] [CrossRef] [PubMed]
- Hardeland, R.; Coto-Montes, A.; Poeggeler, B. Circadian rhythms, oxidative stress, and antioxidative defense mechanisms. Chronobiol. Int. 2003, 20, 921–962. [Google Scholar] [CrossRef] [PubMed]
- Tomas-Zapico, C.; Coto-Montes, A.; Martinez-Fraga, J.; Rodriguez-Colunga, M.J.; Tolivia, D. Effects of continuous light exposure on antioxidant enzymes, porphyric enzymes and cellular damage in the Harderian gland of the Syrian hamster. J. Pineal. Res. 2003, 34, 60–68. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.; Singh, R.K.; Tripathi, A.K.; Cornelissen, G.; Schwartzkopff, O.; Otsuka, K.; Halberg, F. Chronomics of circulating plasma lipid peroxides and anti-oxidant enzymes and other related molecules in cirrhosis of liver. In the memory of late Shri Chetan Singh. Biomed. Pharmacother. 2005, 59 (Suppl. S1), S229–S235. [Google Scholar] [CrossRef]
- Sani, M.; Sebai, H.; Gadacha, W.; Boughattas, N.A.; Reinberg, A.; Mossadok, B.A. Catalase activity and rhythmic patterns in mouse brain, kidney and liver. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 2006, 145, 331–337. [Google Scholar] [CrossRef] [PubMed]
- Fanjul-Moles, M.L.; Prieto-Sagredo, J.; Lopez, D.S.; Bartolo-Orozco, R.; Cruz-Rosas, H. Crayfish Procambarus clarkii retina and nervous system exhibit antioxidant circadian rhythms coupled with metabolic and luminous daily cycles. Photochem. Photobiol. 2009, 85, 78–87. [Google Scholar] [CrossRef] [PubMed]
- Kanabrocki, E.L.; Murray, D.; Hermida, R.C.; Scott, G.S.; Bremner, W.F.; Ryan, M.D.; Ayala, D.E.; Third, J.L.; Shirazi, P.; Nemchausky, B.A.; et al. Circadian variation in oxidative stress markers in healthy and type II diabetic men. Chronobiol. Int. 2002, 19, 423–439. [Google Scholar] [CrossRef] [PubMed]
- Krishnan, N.; Davis, A.J.; Giebultowicz, J.M. Circadian regulation of response to oxidative stress in Drosophila melanogaster. Biochem. Biophys. Res. Commun. 2008, 374, 299–303. [Google Scholar] [CrossRef] [PubMed]
- Lapenna, D.; De Gioia, S.; Mezzetti, A.; Porreca, E.; Ciofani, G.; Marzio, L.; Capani, F.; Di Ilio, C.; Cuccurullo, F. Circadian variations in antioxidant defences and lipid peroxidation in the rat heart. Free Radic. Res. Commun. 1992, 17, 187–194. [Google Scholar] [CrossRef] [PubMed]
- Diaz-Munoz, M.; Hernandez-Munoz, R.; Suarez, J.; Chagoya de Sanchez, V. Day-night cycle of lipid peroxidation in rat cerebral cortex and their relationship to the glutathione cycle and superoxide dismutase activity. Neuroscience 1985, 16, 859–863. [Google Scholar] [CrossRef]
- Martin, V.; Sainz, R.M.; Mayo, J.C.; Antolin, I.; Herrera, F.; Rodriguez, C. Daily rhythm of gene expression in rat superoxide dismutases. Endocr. Res. 2003, 29, 83–95. [Google Scholar] [CrossRef] [PubMed]
- Jang, Y.S.; Lee, M.H.; Lee, S.H.; Bae, K. Cu/Zn superoxide dismutase is differentially regulated in period gene-mutant mice. Biochem. Biophys. Res. Commun. 2011, 409, 22–27. [Google Scholar] [CrossRef] [PubMed]
- Baydas, G.; Gursu, M.F.; Yilmaz, S.; Canpolat, S.; Yasar, A.; Cikim, G.; Canatan, H. Daily rhythm of glutathione peroxidase activity, lipid peroxidation and glutathione levels in tissues of pinealectomized rats. Neurosci. Lett. 2002, 323, 195–198. [Google Scholar] [CrossRef]
- Agapito, M.T.; Redondo, I.; Plaza, R.; Lopez-Burillo, S.; Recio, J.M.; Pablos, M.I. Relationships between melatonin, glutathione peroxidase, glutathione reductase, and catalase. Endogenous rhythms on cerebral cortex in Gallus domesticus. Adv. Exp. Med. Biol. 1999, 460, 377–381. [Google Scholar] [PubMed]
- Coto-Montes, A.; Boga, J.A.; Tomas-Zapico, C.; Rodriguez-Colunga, M.J.; Martinez-Fraga, J.; Tolivia-Cadrecha, D.; Menendez, G.; Hardeland, R.; Tolivia, D. Physiological oxidative stress model: Syrian hamster Harderian gland-sex differences in antioxidant enzymes. Free Radic. Biol. Med. 2001, 30, 785–792. [Google Scholar] [CrossRef]
- Davies, M.H.; Bozigian, H.P.; Merrick, B.A.; Birt, D.F.; Schnell, R.C. Circadian variations in glutathione-S-transferase and glutathione peroxidase activities in the mouse. Toxicol. Lett. 1983, 19, 23–27. [Google Scholar] [CrossRef]
- Jaeschke, H.; Wendel, A. Diurnal fluctuation and pharmacological alteration of mouse organ glutathione content. Biochem. Pharmacol. 1985, 34, 1029–1033. [Google Scholar] [CrossRef]
- Maurice, D.V.; Lightsey, S.F.; Hsu, K.T.; Rhoades, J.F. Comparison of glutathione S-transferase activity in the rat and birds: Tissue distribution and rhythmicity in chicken (Gallus domesticus) liver. Comp. Biochem. Physiol. B 1991, 100, 471–474. [Google Scholar] [CrossRef]
- Chakravarty, S.; Rizvi, S.I. Circadian modulation of human erythrocyte plasma membrane redox system by melatonin. Neurosci. Lett. 2012, 518, 32–35. [Google Scholar] [CrossRef]
- Jung-Hynes, B.; Schmit, T.L.; Reagan-Shaw, S.R.; Siddiqui, I.A.; Mukhtar, H.; Ahmad, N. Melatonin, a novel Sirt1 inhibitor, imparts antiproliferative effects against prostate cancer in vitro in culture and in vivo in TRAMP model. J. Pineal. Res. 2011, 50, 140–149. [Google Scholar] [CrossRef]
- Patterson, A.D.; Shah, Y.M.; Matsubara, T.; Krausz, K.W.; Gonzalez, F.J. Peroxisome proliferator-activated receptor alpha induction of uncoupling protein 2 protects against acetaminophen-induced liver toxicity. Hepatology 2012, 56, 281–290. [Google Scholar] [CrossRef] [PubMed]
- Sekulic-Jablanovic, M.; Petkovic, V.; Wright, M.B.; Kucharava, K.; Huerzeler, N.; Levano, S.; Brand, Y.; Leitmeyer, K.; Glutz, A.; Bausch, A.; et al. Effects of peroxisome proliferator activated receptors (PPAR)-gamma and -alpha agonists on cochlear protection from oxidative stress. PLoS ONE 2017, 12, e0188596. [Google Scholar] [CrossRef] [PubMed]
- Chung, S.S.; Kim, M.; Youn, B.S.; Lee, N.S.; Park, J.W.; Lee, I.K.; Lee, Y.S.; Kim, J.B.; Cho, Y.M.; Lee, H.K.; et al. Glutathione peroxidase 3 mediates the antioxidant effect of peroxisome proliferator-activated receptor gamma in human skeletal muscle cells. Mol. Cell Biol. 2009, 29, 20–30. [Google Scholar] [CrossRef] [PubMed]
- Girnun, G.D.; Domann, F.E.; Moore, S.A.; Robbins, M.E. Identification of a functional peroxisome proliferator-activated receptor response element in the rat catalase promoter. Mol. Endocrinol. 2002, 16, 2793–2801. [Google Scholar] [CrossRef]
- Okuno, Y.; Matsuda, M.; Miyata, Y.; Fukuhara, A.; Komuro, R.; Shimabukuro, M.; Shimomura, I. Human catalase gene is regulated by peroxisome proliferator activated receptor-gamma through a response element distinct from that of mouse. Endocr. J. 2010, 57, 303–309. [Google Scholar] [CrossRef]
- Ding, G.; Fu, M.; Qin, Q.; Lewis, W.; Kim, H.W.; Fukai, T.; Bacanamwo, M.; Chen, Y.E.; Schneider, M.D.; Mangelsdorf, D.J.; et al. Cardiac peroxisome proliferator-activated receptor gamma is essential in protecting cardiomyocytes from oxidative damage. Cardiovasc. Res. 2007, 76, 269–279. [Google Scholar] [CrossRef]
- Zhang, F.; Liu, F.; Yan, M.; Ji, H.; Hu, L.; Li, X.; Qian, J.; He, X.; Zhang, L.; Shen, A.; et al. Peroxisome proliferator-activated receptor-gamma agonists suppress iNOS expression induced by LPS in rat primary Schwann cells. J. Neuroimmunol. 2010, 218, 36–47. [Google Scholar] [CrossRef]
- Vandewalle, B.; Moerman, E.; Lefebvre, B.; Defrance, F.; Gmyr, V.; Lukowiak, B.; Kerr Conte, J.; Pattou, F. PPARgamma-dependent and -independent effects of rosiglitazone on lipotoxic human pancreatic islets. Biochem. Biophys. Res. Commun. 2008, 366, 1096–1101. [Google Scholar] [CrossRef]
- Kleinhenz, J.M.; Kleinhenz, D.J.; You, S.; Ritzenthaler, J.D.; Hansen, J.M.; Archer, D.R.; Sutliff, R.L.; Hart, C.M. Disruption of endothelial peroxisome proliferator-activated receptor-gamma reduces vascular nitric oxide production. Am. J. Physiol. Heart Circ. Physiol. 2009, 297, H1647–H1654. [Google Scholar] [CrossRef]
- Ricote, M.; Li, A.C.; Willson, T.M.; Kelly, C.J.; Glass, C.K. The peroxisome proliferator-activated receptor-gamma is a negative regulator of macrophage activation. Nature 1998, 391, 79–82. [Google Scholar] [CrossRef]
- Li, M.; Pascual, G.; Glass, C.K. Peroxisome proliferator-activated receptor gamma-dependent repression of the inducible nitric oxide synthase gene. Mol. Cell Biol. 2000, 20, 4699–4707. [Google Scholar] [CrossRef] [PubMed]
- Chan, S.H.; Wu, K.L.; Kung, P.S.; Chan, J.Y. Oral intake of rosiglitazone promotes a central antihypertensive effect via upregulation of peroxisome proliferator-activated receptor-gamma and alleviation of oxidative stress in rostral ventrolateral medulla of spontaneously hypertensive rats. Hypertension 2010, 55, 1444–1453. [Google Scholar] [CrossRef] [PubMed]
- Paradis, E.; Clavel, S.; Bouillaud, F.; Ricquier, D.; Richard, D. Uncoupling protein 2: A novel player in neuroprotection. Trends Mol. Med. 2003, 9, 522–525. [Google Scholar] [CrossRef] [PubMed]
- Nagy, L.; Tontonoz, P.; Alvarez, J.G.; Chen, H.; Evans, R.M. Oxidized LDL regulates macrophage gene expression through ligand activation of PPAR gamma. Cell 1998, 93, 229–240. [Google Scholar] [CrossRef]
- Ishii, T.; Itoh, K.; Ruiz, E.; Leake, D.S.; Unoki, H.; Yamamoto, M.; Mann, G.E. Role of Nrf2 in the regulation of CD36 and stress protein expression in murine macrophages: Activation by oxidatively modified LDL and 4-hydroxynonenal. Circ. Res. 2004, 94, 609–616. [Google Scholar] [CrossRef] [PubMed]
- Febbraio, M.; Hajjar, D.P.; Silverstein, R.L. CD36: A class B scavenger receptor involved in angiogenesis, atherosclerosis, inflammation, and lipid metabolism. J. Clin. Investig. 2001, 108, 785–791. [Google Scholar] [CrossRef] [PubMed]
- Fuenzalida, K.; Quintanilla, R.; Ramos, P.; Piderit, D.; Fuentealba, R.A.; Martinez, G.; Inestrosa, N.C.; Bronfman, M. Peroxisome proliferator-activated receptor gamma up-regulates the Bcl-2 anti-apoptotic protein in neurons and induces mitochondrial stabilization and protection against oxidative stress and apoptosis. J. Biol. Chem. 2007, 282, 37006–37015. [Google Scholar] [CrossRef] [PubMed]
- Ren, Y.; Sun, C.; Sun, Y.; Tan, H.; Wu, Y.; Cui, B.; Wu, Z. PPAR gamma protects cardiomyocytes against oxidative stress and apoptosis via Bcl-2 upregulation. Vascul. Pharmacol. 2009, 51, 169–174. [Google Scholar] [CrossRef] [PubMed]
- Osburn, W.O.; Kensler, T.W. Nrf2 signaling: An adaptive response pathway for protection against environmental toxic insults. Mutat. Res. 2008, 659, 31–39. [Google Scholar] [CrossRef] [PubMed]
- Cho, H.Y.; Gladwell, W.; Wang, X.; Chorley, B.; Bell, D.; Reddy, S.P.; Kleeberger, S.R. Nrf2-regulated PPAR{gamma} expression is critical to protection against acute lung injury in mice. Am. J. Respir. Crit. Care Med. 2010, 182, 170–182. [Google Scholar] [CrossRef]
- Park, E.Y.; Cho, I.J.; Kim, S.G. Transactivation of the PPAR-responsive enhancer module in chemopreventive glutathione S-transferase gene by the peroxisome proliferator-activated receptor-gamma and retinoid X receptor heterodimer. Cancer Res. 2004, 64, 3701–3713. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Tabbi-Anneni, I.; Gunda, V.; Wang, L. Transcription factor Nrf2 regulates SHP and lipogenic gene expression in hepatic lipid metabolism. Am. J. Physiol. Gastrointest. Liver Physiol. 2010, 299, G1211–G1221. [Google Scholar] [CrossRef] [PubMed]
- Pekovic-Vaughan, V.; Gibbs, J.; Yoshitane, H.; Yang, N.; Pathiranage, D.; Guo, B.; Sagami, A.; Taguchi, K.; Bechtold, D.; Loudon, A.; et al. The circadian clock regulates rhythmic activation of the NRF2/glutathione-mediated antioxidant defense pathway to modulate pulmonary fibrosis. Genes Dev. 2014, 28, 548–560. [Google Scholar] [CrossRef] [PubMed]
- Froy, O.; Chapnik, N. Circadian oscillation of innate immunity components in mouse small intestine. Mol. Immunol. 2007, 44, 1954–1960. [Google Scholar] [CrossRef] [PubMed]
- Arjona, A.; Sarkar, D.K. Circadian oscillations of clock genes, cytolytic factors, and cytokines in rat NK cells. J. Immunol. 2005, 174, 7618–7624. [Google Scholar] [CrossRef] [PubMed]
- Arjona, A.; Sarkar, D.K. Evidence supporting a circadian control of natural killer cell function. Brain Behav. Immun. 2006, 20, 469–476. [Google Scholar] [CrossRef] [PubMed]
- Arjona, A.; Boyadjieva, N.; Sarkar, D.K. Circadian rhythms of granzyme B, perforin, IFN-gamma, and NK cell cytolytic activity in the spleen: Effects of chronic ethanol. J. Immunol. 2004, 172, 2811–2817. [Google Scholar] [CrossRef]
- Druzd, D.; Matveeva, O.; Ince, L.; Harrison, U.; He, W.; Schmal, C.; Herzel, H.; Tsang, A.H.; Kawakami, N.; Leliavski, A.; et al. Lymphocyte circadian clocks control lymph node trafficking and adaptive immune responses. Immunity 2017, 46, 120–132. [Google Scholar] [CrossRef]
- Bellet, M.M.; Deriu, E.; Liu, J.Z.; Grimaldi, B.; Blaschitz, C.; Zeller, M.; Edwards, R.A.; Sahar, S.; Dandekar, S.; Baldi, P.; et al. Circadian clock regulates the host response to Salmonella. Proc. Natl. Acad. Sci. USA 2013, 110, 9897–9902. [Google Scholar] [CrossRef]
- Kundu, P.; Ling, T.W.; Korecka, A.; Li, Y.; D’Arienzo, R.; Bunte, R.M.; Berger, T.; Arulampalam, V.; Chambon, P.; Mak, T.W.; et al. Absence of intestinal PPARgamma aggravates acute infectious colitis in mice through a lipocalin-2-dependent pathway. PLoS Pathog. 2014, 10, e1003887. [Google Scholar] [CrossRef]
- Silver, A.C.; Arjona, A.; Walker, W.E.; Fikrig, E. The circadian clock controls toll-like receptor 9-mediated innate and adaptive immunity. Immunity 2012, 36, 251–261. [Google Scholar] [CrossRef] [PubMed]
- Logan, R.W.; Arjona, A.; Sarkar, D.K. Role of sympathetic nervous system in the entrainment of circadian natural-killer cell function. Brain Behav. Immun. 2011, 25, 101–109. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, Y.; Ishimaru, K.; Tahara, Y.; Shibata, S.; Nakao, A. Disruption of the suprachiasmatic nucleus blunts a time of day-dependent variation in systemic anaphylactic reaction in mice. J. Immunol. Res. 2014, 2014, 474217. [Google Scholar] [CrossRef] [PubMed]
- Guerrero-Vargas, N.N.; Salgado-Delgado, R.; Basualdo Mdel, C.; Garcia, J.; Guzman-Ruiz, M.; Carrero, J.C.; Escobar, C.; Buijs, R.M. Reciprocal interaction between the suprachiasmatic nucleus and the immune system tunes down the inflammatory response to lipopolysaccharide. J. Neuroimmunol. 2014, 273, 22–30. [Google Scholar] [CrossRef] [PubMed]
- Gibbs, J.; Ince, L.; Matthews, L.; Mei, J.; Bell, T.; Yang, N.; Saer, B.; Begley, N.; Poolman, T.; Pariollaud, M.; et al. An epithelial circadian clock controls pulmonary inflammation and glucocorticoid action. Nat. Med. 2014, 20, 919–926. [Google Scholar] [CrossRef] [PubMed]
- Adams, K.L.; Castanon-Cervantes, O.; Evans, J.A.; Davidson, A.J. Environmental circadian disruption elevates the IL-6 response to lipopolysaccharide in blood. J. Biol. Rhythms 2013, 28, 272–277. [Google Scholar] [CrossRef] [PubMed]
- Guerrero-Vargas, N.N.; Guzman-Ruiz, M.; Fuentes, R.; Garcia, J.; Salgado-Delgado, R.; Basualdo Mdel, C.; Escobar, C.; Markus, R.P.; Buijs, R.M. Shift work in rats results in increased inflammatory response after lipopolysaccharide administration: A role for food consumption. J. Biol. Rhythms 2015, 30, 318–330. [Google Scholar] [CrossRef]
- Stapleton, C.M.; Jaradat, M.; Dixon, D.; Kang, H.S.; Kim, S.C.; Liao, G.; Carey, M.A.; Cristiano, J.; Moorman, M.P.; Jetten, A.M. Enhanced susceptibility of staggerer (RORalphasg/sg) mice to lipopolysaccharide-induced lung inflammation. Am. J. Physiol. Lung Cell Mol. Physiol. 2005, 289, L144–L152. [Google Scholar] [CrossRef]
- Arjona, A.; Sarkar, D.K. The circadian gene mPer2 regulates the daily rhythm of IFN-gamma. J. Interferon. Cytokine Res. 2006, 26, 645–649. [Google Scholar] [CrossRef]
- Liu, J.; Malkani, G.; Shi, X.; Meyer, M.; Cunningham-Runddles, S.; Ma, X.; Sun, Z.S. The circadian clock Period 2 gene regulates gamma interferon production of NK cells in host response to lipopolysaccharide-induced endotoxic shock. Infect. Immun. 2006, 74, 4750–4756. [Google Scholar] [CrossRef]
- Gibbs, J.E.; Blaikley, J.; Beesley, S.; Matthews, L.; Simpson, K.D.; Boyce, S.H.; Farrow, S.N.; Else, K.J.; Singh, D.; Ray, D.W.; et al. The nuclear receptor REV-ERBalpha mediates circadian regulation of innate immunity through selective regulation of inflammatory cytokines. Proc. Natl. Acad. Sci. USA 2012, 109, 582–587. [Google Scholar] [CrossRef] [PubMed]
- Sato, S.; Sakurai, T.; Ogasawara, J.; Takahashi, M.; Izawa, T.; Imaizumi, K.; Taniguchi, N.; Ohno, H.; Kizaki, T. A circadian clock gene, Rev-erbalpha, modulates the inflammatory function of macrophages through the negative regulation of Ccl2 expression. J. Immunol. 2014, 192, 407–417. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, K.D.; Fentress, S.J.; Qiu, Y.; Yun, K.; Cox, J.S.; Chawla, A. Circadian gene Bmal1 regulates diurnal oscillations of Ly6C(hi) inflammatory monocytes. Science 2013, 341, 1483–1488. [Google Scholar] [CrossRef] [PubMed]
- Huo, M.; Huang, Y.; Qu, D.; Zhang, H.; Wong, W.T.; Chawla, A.; Huang, Y.; Tian, X.Y. Myeloid Bmal1 deletion increases monocyte recruitment and worsens atherosclerosis. FASEB J. 2017, 31, 1097–1106. [Google Scholar] [CrossRef] [PubMed]
- Dzhagalov, I.; Giguere, V.; He, Y.W. Lymphocyte development and function in the absence of retinoic acid-related orphan receptor alpha. J. Immunol. 2004, 173, 2952–2959. [Google Scholar] [CrossRef]
- Narasimamurthy, R.; Hatori, M.; Nayak, S.K.; Liu, F.; Panda, S.; Verma, I.M. Circadian clock protein cryptochrome regulates the expression of proinflammatory cytokines. Proc. Natl. Acad. Sci. USA 2012, 109, 12662–12667. [Google Scholar] [CrossRef] [PubMed]
- Spengler, M.L.; Kuropatwinski, K.K.; Comas, M.; Gasparian, A.V.; Fedtsova, N.; Gleiberman, A.S.; Gitlin, I.I.; Artemicheva, N.M.; Deluca, K.A.; Gudkov, A.V.; et al. Core circadian protein CLOCK is a positive regulator of NF-kappaB-mediated transcription. Proc. Natl. Acad. Sci. USA 2012, 109, E2457–E2465. [Google Scholar] [CrossRef]
- Morgan, M.J.; Liu, Z.G. Crosstalk of reactive oxygen species and NF-kappaB signaling. Cell Res. 2011, 21, 103–115. [Google Scholar] [CrossRef]
- Goto, M.; Katayama, K.I.; Shirakawa, F.; Tanaka, I. Involvement of NF-kappaB p50/p65 heterodimer in activation of the human pro-interleukin-1beta gene at two subregions of the upstream enhancer element. Cytokine 1999, 11, 16–28. [Google Scholar] [CrossRef]
- Kim, H.J.; Kim, K.W.; Yu, B.P.; Chung, H.Y. The effect of age on cyclooxygenase-2 gene expression: NF-kappaB activation and IkappaBalpha degradation. Free Radic. Biol. Med. 2000, 28, 683–692. [Google Scholar] [CrossRef]
- Libermann, T.A.; Baltimore, D. Activation of interleukin-6 gene expression through the NF-kappa B transcription factor. Mol. Cell Biol. 1990, 10, 2327–2334. [Google Scholar] [CrossRef] [PubMed]
- Yao, J.; Mackman, N.; Edgington, T.S.; Fan, S.T. Lipopolysaccharide induction of the tumor necrosis factor-alpha promoter in human monocytic cells. Regulation by Egr-1, c-Jun, and NF-kappaB transcription factors. J. Biol. Chem. 1997, 272, 17795–17801. [Google Scholar] [CrossRef] [PubMed]
- Taylor, B.S.; de Vera, M.E.; Ganster, R.W.; Wang, Q.; Shapiro, R.A.; Morris, S.M., Jr.; Billiar, T.R.; Geller, D.A. Multiple NF-kappaB enhancer elements regulate cytokine induction of the human inducible nitric oxide synthase gene. J. Biol. Chem. 1998, 273, 15148–15156. [Google Scholar] [CrossRef] [PubMed]
- Ricote, M.; Huang, J.; Fajas, L.; Li, A.; Welch, J.; Najib, J.; Witztum, J.L.; Auwerx, J.; Palinski, W.; Glass, C.K. Expression of the peroxisome proliferator-activated receptor gamma (PPARgamma) in human atherosclerosis and regulation in macrophages by colony stimulating factors and oxidized low density lipoprotein. Proc. Natl. Acad. Sci. USA 1998, 95, 7614–7619. [Google Scholar] [CrossRef] [PubMed]
- Jiang, C.; Ting, A.T.; Seed, B. PPAR-gamma agonists inhibit production of monocyte inflammatory cytokines. Nature 1998, 391, 82–86. [Google Scholar] [CrossRef]
- Okada, M.; Yan, S.F.; Pinsky, D.J. Peroxisome proliferator-activated receptor-gamma (PPAR-gamma) activation suppresses ischemic induction of Egr-1 and its inflammatory gene targets. FASEB J. 2002, 16, 1861–1868. [Google Scholar] [CrossRef]
- Barbier, O.; Torra, I.P.; Duguay, Y.; Blanquart, C.; Fruchart, J.C.; Glineur, C.; Staels, B. Pleiotropic actions of peroxisome proliferator-activated receptors in lipid metabolism and atherosclerosis. Arterioscler. Thromb. Vasc. Biol. 2002, 22, 717–726. [Google Scholar] [CrossRef]
- Narala, V.R.; Adapala, R.K.; Suresh, M.V.; Brock, T.G.; Peters-Golden, M.; Reddy, R.C. Leukotriene B4 is a physiologically relevant endogenous peroxisome proliferator-activated receptor-alpha agonist. J. Biol. Chem. 2010, 285, 22067–22074. [Google Scholar] [CrossRef]
- Wahli, W.; Michalik, L. PPARs at the crossroads of lipid signaling and inflammation. Trends Endocrinol. Metab. 2012, 23, 351–363. [Google Scholar] [CrossRef]
- Delerive, P.; De Bosscher, K.; Besnard, S.; Vanden Berghe, W.; Peters, J.M.; Gonzalez, F.J.; Fruchart, J.C.; Tedgui, A.; Haegeman, G.; Staels, B. Peroxisome proliferator-activated receptor alpha negatively regulates the vascular inflammatory gene response by negative cross-talk with transcription factors NF-kappa B and AP-1. J. Biol. Chem. 1999, 274, 32048–32054. [Google Scholar] [CrossRef]
- Crisafulli, C.; Cuzzocrea, S. The role of endogenous and exogenous ligands for the peroxisome proliferator-activated receptor alpha (PPAR-alpha) in the regulation of inflammation in macrophages. Shock 2009, 32, 62–73. [Google Scholar] [CrossRef] [PubMed]
- Benetti, E.; Mastrocola, R.; Rogazzo, M.; Chiazza, F.; Aragno, M.; Fantozzi, R.; Collino, M.; Minetto, M.A. High sugar intake and development of skeletal muscle insulin resistance and inflammation in mice: A protective role for PPAR- delta agonism. Mediat. Inflamm. 2013, 2013, 509502. [Google Scholar] [CrossRef] [PubMed]
- Ding, G.; Cheng, L.; Qin, Q.; Frontin, S.; Yang, Q. PPARdelta modulates lipopolysaccharide-induced TNFalpha inflammation signaling in cultured cardiomyocytes. J. Mol. Cell Cardiol. 2006, 40, 821–828. [Google Scholar] [CrossRef] [PubMed]
- Peters, J.M.; Hollingshead, H.E.; Gonzalez, F.J. Role of peroxisome-proliferator-activated receptor beta/delta (PPARbeta/delta) in gastrointestinal tract function and disease. Clin. Sci. 2008, 115, 107–127. [Google Scholar] [CrossRef] [PubMed]
- Lewis, J.D.; Lichtenstein, G.R.; Deren, J.J.; Sands, B.E.; Hanauer, S.B.; Katz, J.A.; Lashner, B.; Present, D.H.; Chuai, S.; Ellenberg, J.H.; et al. Rosiglitazone for active ulcerative colitis: A randomized placebo-controlled trial. Gastroenterology 2008, 134, 688–695. [Google Scholar] [CrossRef] [PubMed]
- Bassaganya-Riera, J.; Hontecillas, R. CLA and n-3 PUFA differentially modulate clinical activity and colonic PPAR-responsive gene expression in a pig model of experimental IBD. Clin. Nutr. 2006, 25, 454–465. [Google Scholar] [CrossRef]
- Sanchez-Hidalgo, M.; Martin, A.R.; Villegas, I.; de la Lastra, C.A. Rosiglitazone, a PPARgamma ligand, modulates signal transduction pathways during the development of acute TNBS-induced colitis in rats. Eur. J. Pharm. 2007, 562, 247–258. [Google Scholar] [CrossRef]
- Su, C.G.; Wen, X.; Bailey, S.T.; Jiang, W.; Rangwala, S.M.; Keilbaugh, S.A.; Flanigan, A.; Murthy, S.; Lazar, M.A.; Wu, G.D. A novel therapy for colitis utilizing PPAR-gamma ligands to inhibit the epithelial inflammatory response. J. Clin. Investig. 1999, 104, 383–389. [Google Scholar] [CrossRef]
- Michalik, L.; Wahli, W. PPARs mediate lipid signaling in inflammation and cancer. PPAR Res. 2008, 2008, 134059. [Google Scholar] [CrossRef]
- Wada, K.; Nakajima, A.; Blumberg, R.S. PPARgamma and inflammatory bowel disease: A new therapeutic target for ulcerative colitis and Crohn’s disease. Trends Mol. Med. 2001, 7, 329–331. [Google Scholar] [CrossRef]
- Chawla, A.; Barak, Y.; Nagy, L.; Liao, D.; Tontonoz, P.; Evans, R.M. PPAR-gamma dependent and independent effects on macrophage-gene expression in lipid metabolism and inflammation. Nat. Med. 2001, 7, 48–52. [Google Scholar] [CrossRef] [PubMed]
- Odegaard, J.I.; Ricardo-Gonzalez, R.R.; Goforth, M.H.; Morel, C.R.; Subramanian, V.; Mukundan, L.; Red Eagle, A.; Vats, D.; Brombacher, F.; Ferrante, A.W.; et al. Macrophage-specific PPARgamma controls alternative activation and improves insulin resistance. Nature 2007, 447, 1116–1120. [Google Scholar] [CrossRef] [PubMed]
- Bassaganya-Riera, J.; Reynolds, K.; Martino-Catt, S.; Cui, Y.; Hennighausen, L.; Gonzalez, F.; Rohrer, J.; Benninghoff, A.U.; Hontecillas, R. Activation of PPAR gamma and delta by conjugated linoleic acid mediates protection from experimental inflammatory bowel disease. Gastroenterology 2004, 127, 777–791. [Google Scholar] [CrossRef] [PubMed]
- Klotz, L.; Schmidt, S.; Heun, R.; Klockgether, T.; Kolsch, H. Association of the PPARgamma gene polymorphism Pro12Ala with delayed onset of multiple sclerosis. Neurosci. Lett. 2009, 449, 81–83. [Google Scholar] [CrossRef] [PubMed]
- Regieli, J.J.; Jukema, J.W.; Doevendans, P.A.; Zwinderman, A.H.; van der Graaf, Y.; Kastelein, J.J.; Grobbee, D.E. PPAR gamma variant influences angiographic outcome and 10-year cardiovascular risk in male symptomatic coronary artery disease patients. Diabetes Care 2009, 32, 839–844. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Jiang, J.; Xie, Z.; Guo, J.; Wang, Y.; Liu, C.; Zhang, S.; Tang, W.; Chen, Y. Association of PPARG rs 1801282 C>G polymorphism with risk of colorectal cancer: From a case-control study to a meta-analysis. Oncotarget 2017, 8, 100558–100569. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Chen, Y.; Tang, W.F.; Liu, C.; Zhang, S.; Guo, Z.Q.; Chen, G.; Zheng, X.W. PPARG rs3856806 C>T Polymorphism Increased the Risk of Colorectal Cancer: A Case-Control Study in Eastern Chinese Han Population. Front. Oncol. 2019, 9, 63. [Google Scholar] [CrossRef]
- Yasui, Y.; Hosokawa, M.; Sahara, T.; Suzuki, R.; Ohgiya, S.; Kohno, H.; Tanaka, T.; Miyashita, K. Bitter gourd seed fatty acid rich in 9c,11t,13t-conjugated linolenic acid induces apoptosis and up-regulates the GADD45, p53 and PPAR gamma in human colon cancer Caco-2 cells. Prostaglandins Leukot. Essent. Fatty Acids 2005, 73, 113–119. [Google Scholar] [CrossRef]
- Akinyeke, T.O.; Stewart, L.V. Troglitazone suppresses c-Myc levels in human prostate cancer cells via a PPARgamma-independent mechanism. Cancer Biol. Ther. 2011, 11, 1046–1058. [Google Scholar] [CrossRef]
- Subbaramaiah, K.; Lin, D.T.; Hart, J.C.; Dannenberg, A.J. Peroxisome proliferator-activated receptor gamma ligands suppress the transcriptional activation of cyclooxygenase-2. Evidence for involvement of activator protein-1 and CREB-binding protein/p300. J. Biol. Chem. 2001, 276, 12440–12448. [Google Scholar] [CrossRef]
- Vandoros, G.P.; Konstantinopoulos, P.A.; Sotiropoulou-Bonikou, G.; Kominea, A.; Papachristou, G.I.; Karamouzis, M.V.; Gkermpesi, M.; Varakis, I.; Papavassiliou, A.G. PPAR-gamma is expressed and NF-kB pathway is activated and correlates positively with COX-2 expression in stromal myofibroblasts surrounding colon adenocarcinomas. J. Cancer Res. Clin. Oncol. 2006, 132, 76–84. [Google Scholar] [CrossRef] [PubMed]
- Heneka, M.T.; Klockgether, T.; Feinstein, D.L. Peroxisome proliferator-activated receptor-gamma ligands reduce neuronal inducible nitric oxide synthase expression and cell death in vivo. J. Neurosci. 2000, 20, 6862–6867. [Google Scholar] [CrossRef] [PubMed]
- Girnun, G.D.; Smith, W.M.; Drori, S.; Sarraf, P.; Mueller, E.; Eng, C.; Nambiar, P.; Rosenberg, D.W.; Bronson, R.T.; Edelmann, W.; et al. APC-dependent suppression of colon carcinogenesis by PPAR gamma. Proc. Natl. Acad. Sci. USA 2002, 99, 13771–13776. [Google Scholar] [CrossRef] [PubMed]
- Fujisawa, T.; Nakajima, A.; Fujisawa, N.; Takahashi, H.; Ikeda, I.; Tomimoto, A.; Yonemitsu, K.; Nakajima, N.; Kudo, C.; Wada, K.; et al. Peroxisome proliferator-activated receptor gamma (PPAR gamma) suppresses colonic epithelial cell turnover and colon carcinogenesis through inhibition of the beta-catenin/T cell factor (TCF) pathway. J. Pharmacol. Sci. 2008, 106, 627–638. [Google Scholar] [CrossRef] [PubMed]
- Ye, J. Regulation of PPARgamma function by TNF-alpha. Biochem. Biophys. Res. Commun. 2008, 374, 405–408. [Google Scholar] [CrossRef] [PubMed]
- Napolitano, M.; Costa, L.; Palermo, R.; Giovenco, A.; Vacca, A.; Gulino, A. Protective effect of pioglitazone, a PPARgamma ligand, in a 3 nitropropionic acid model of Huntington’s disease. Brain Res. Bull. 2011, 85, 231–237. [Google Scholar] [CrossRef] [PubMed]
- Levi, F.; Focan, C.; Karaboue, A.; de la Valette, V.; Focan-Henrard, D.; Baron, B.; Kreutz, F.; Giacchetti, S. Implications of circadian clocks for the rhythmic delivery of cancer therapeutics. Adv. Drug Deliv. Rev. 2007, 59, 1015–1035. [Google Scholar] [CrossRef]
- Yamazaki, S.; Straume, M.; Tei, H.; Sakaki, Y.; Menaker, M.; Block, G.D. Effects of aging on central and peripheral mammalian clocks. Proc. Natl. Acad. Sci. USA 2002, 99, 10801–10806. [Google Scholar] [CrossRef]
- Scarbrough, K.; Losee-Olson, S.; Wallen, E.P.; Turek, F.W. Aging and photoperiod affect entrainment and quantitative aspects of locomotor behavior in Syrian hamsters. Am. J. Physiol. 1997, 272, R1219–R1225. [Google Scholar] [CrossRef]
- Gibson, E.M.; Williams, W.P., 3rd; Kriegsfeld, L.J. Aging in the circadian system: Considerations for health, disease prevention and longevity. Exp. Gerontol. 2009, 44, 51–56. [Google Scholar] [CrossRef]
- Hofman, M.A.; Swaab, D.F. Living by the clock: The circadian pacemaker in older people. Ageing Res. Rev. 2006, 5, 33–51. [Google Scholar] [CrossRef] [PubMed]
- Chang, H.C.; Guarente, L. SIRT1 mediates central circadian control in the SCN by a mechanism that decays with aging. Cell 2013, 153, 1448–1460. [Google Scholar] [CrossRef] [PubMed]
- Sellix, M.T.; Evans, J.A.; Leise, T.L.; Castanon-Cervantes, O.; Hill, D.D.; DeLisser, P.; Block, G.D.; Menaker, M.; Davidson, A.J. Aging differentially affects the re-entrainment response of central and peripheral circadian oscillators. J. Neurosci. 2012, 32, 16193–16202. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Brainard, G.C.; Zee, P.C.; Pinto, L.H.; Takahashi, J.S.; Turek, F.W. Effects of aging on lens transmittance and retinal input to the suprachiasmatic nucleus in golden hamsters. Neurosci. Lett. 1998, 258, 167–170. [Google Scholar] [CrossRef]
- Weinert, D. Age-dependent changes of the circadian system. Chronobiol. Int. 2000, 17, 261–283. [Google Scholar] [CrossRef] [PubMed]
- Witting, W.; Mirmiran, M.; Bos, N.P.; Swaab, D.F. The effect of old age on the free-running period of circadian rhythms in rat. Chronobiol. Int. 1994, 11, 103–112. [Google Scholar] [CrossRef] [PubMed]
- Duffy, J.F.; Zeitzer, J.M.; Rimmer, D.W.; Klerman, E.B.; Dijk, D.J.; Czeisler, C.A. Peak of circadian melatonin rhythm occurs later within the sleep of older subjects. Am. J. Physiol. Endocrinol. Metab. 2002, 282, E297–E303. [Google Scholar] [CrossRef] [PubMed]
- Yoon, I.Y.; Kripke, D.F.; Elliott, J.A.; Youngstedt, S.D.; Rex, K.M.; Hauger, R.L. Age-related changes of circadian rhythms and sleep-wake cycles. J. Am. Geriatr. Soc. 2003, 51, 1085–1091. [Google Scholar] [CrossRef] [PubMed]
- Kolker, D.E.; Fukuyama, H.; Huang, D.S.; Takahashi, J.S.; Horton, T.H.; Turek, F.W. Aging alters circadian and light-induced expression of clock genes in golden hamsters. J. Biol. Rhythms 2003, 18, 159–169. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.C. The circadian clock and tumor suppression by mammalian period genes. Methods Enzymol. 2005, 393, 852–861. [Google Scholar] [CrossRef] [PubMed]
- Swindell, W.R. Gene expression profiling of long-lived dwarf mice: Longevity-associated genes and relationships with diet, gender and aging. BMC Genom. 2007, 8, 353. [Google Scholar] [CrossRef] [PubMed]
- Hurd, M.W.; Ralph, M.R. The significance of circadian organization for longevity in the golden hamster. J. Biol. Rhythms 1998, 13, 430–436. [Google Scholar] [CrossRef] [PubMed]
- Hurd, M.W.; Zimmer, K.A.; Lehman, M.N.; Ralph, M.R. Circadian locomotor rhythms in aged hamsters following suprachiasmatic transplant. Am. J. Physiol. 1995, 269, R958–R968. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Satinoff, E. Fetal tissue containing the suprachiasmatic nucleus restores multiple circadian rhythms in old rats. Am. J. Physiol. 1998, 275, R1735–R1744. [Google Scholar] [CrossRef] [PubMed]
- Penev, P.D.; Kolker, D.E.; Zee, P.C.; Turek, F.W. Chronic circadian desynchronization decreases the survival of animals with cardiomyopathic heart disease. Am. J. Physiol. 1998, 275, H2334–H2337. [Google Scholar] [CrossRef]
- Viswanathan, N.; Davis, F.C. Suprachiasmatic nucleus grafts restore circadian function in aged hamsters. Brain Res. 1995, 686, 10–16. [Google Scholar] [CrossRef]
- Van Reeth, O.; Zhang, Y.; Zee, P.C.; Turek, F.W. Grafting fetal suprachiasmatic nuclei in the hypothalamus of old hamsters restores responsiveness of the circadian clock to a phase shifting stimulus. Brain Res. 1994, 643, 338–342. [Google Scholar] [CrossRef]
- Khan, S.; Duan, P.; Yao, L.; Hou, H. Shiftwork-mediated disruptions of circadian rhythms and sleep homeostasis cause serious health problems. Int. J. Genom. 2018, 2018, 8576890. [Google Scholar] [CrossRef]
- Solanas, G.; Peixoto, F.O.; Perdiguero, E.; Jardi, M.; Ruiz-Bonilla, V.; Datta, D.; Symeonidi, A.; Castellanos, A.; Welz, P.S.; Caballero, J.M.; et al. Aged stem cells reprogram their daily rhythmic functions to adapt to stress. Cell 2017, 170, 678–692.e620. [Google Scholar] [CrossRef]
- Knight, J.A. The biochemistry of aging. Adv. Clin. Chem. 2000, 35, 1–62. [Google Scholar]
- Park, S.Y.; Kim, Y.W.; Kim, J.E.; Kim, J.Y. Age-associated changes in fat metabolism in the rat and its relation to sympathetic activity. Life Sci. 2006, 79, 2228–2233. [Google Scholar] [CrossRef] [PubMed]
- Petersen, K.F.; Befroy, D.; Dufour, S.; Dziura, J.; Ariyan, C.; Rothman, D.L.; DiPietro, L.; Cline, G.W.; Shulman, G.I. Mitochondrial dysfunction in the elderly: Possible role in insulin resistance. Science 2003, 300, 1140–1142. [Google Scholar] [CrossRef] [PubMed]
- Kenyon, C. The plasticity of aging: Insights from long-lived mutants. Cell 2005, 120, 449–460. [Google Scholar] [CrossRef] [PubMed]
- Siersbaek, R.; Nielsen, R.; Mandrup, S. PPARgamma in adipocyte differentiation and metabolism–novel insights from genome-wide studies. FEBS Lett. 2010, 584, 3242–3249. [Google Scholar] [CrossRef] [PubMed]
- Chawla, A.; Schwarz, E.J.; Dimaculangan, D.D.; Lazar, M.A. Peroxisome proliferator-activated receptor (PPAR) gamma: Adipose-predominant expression and induction early in adipocyte differentiation. Endocrinology 1994, 135, 798–800. [Google Scholar] [CrossRef] [PubMed]
- Tontonoz, P.; Hu, E.; Spiegelman, B.M. Stimulation of adipogenesis in fibroblasts by PPAR gamma 2, a lipid-activated transcription factor. Cell 1994, 79, 1147–1156. [Google Scholar] [CrossRef]
- Argmann, C.; Dobrin, R.; Heikkinen, S.; Auburtin, A.; Pouilly, L.; Cock, T.A.; Koutnikova, H.; Zhu, J.; Schadt, E.E.; Auwerx, J. Ppargamma2 is a key driver of longevity in the mouse. PLoS Genet. 2009, 5, e1000752. [Google Scholar] [CrossRef]
- Lee, C.K.; Allison, D.B.; Brand, J.; Weindruch, R.; Prolla, T.A. Transcriptional profiles associated with aging and middle age-onset caloric restriction in mouse hearts. Proc. Natl. Acad. Sci. USA 2002, 99, 14988–14993. [Google Scholar] [CrossRef]
- Iemitsu, M.; Miyauchi, T.; Maeda, S.; Tanabe, T.; Takanashi, M.; Irukayama-Tomobe, Y.; Sakai, S.; Ohmori, H.; Matsuda, M.; Yamaguchi, I. Aging-induced decrease in the PPAR-alpha level in hearts is improved by exercise training. Am. J. Physiol. Heart Circ. Physiol. 2002, 283, H1750–H1760. [Google Scholar] [CrossRef]
- Corton, J.C.; Brown-Borg, H.M. Peroxisome proliferator-activated receptor gamma coactivator 1 in caloric restriction and other models of longevity. J. Gerontol. A Biol. Sci. Med. Sci. 2005, 60, 1494–1509. [Google Scholar] [CrossRef]
- Stauber, A.J.; Brown-Borg, H.; Liu, J.; Waalkes, M.P.; Laughter, A.; Staben, R.A.; Coley, J.C.; Swanson, C.; Voss, K.A.; Kopchick, J.J.; et al. Constitutive expression of peroxisome proliferator-activated receptor alpha-regulated genes in dwarf mice. Mol. Pharmacol. 2005, 67, 681–694. [Google Scholar] [CrossRef] [PubMed]
- Olsson, B.; Bohlooly, Y.M.; Brusehed, O.; Isaksson, O.G.; Ahren, B.; Olofsson, S.O.; Oscarsson, J.; Tornell, J. Bovine growth hormone-transgenic mice have major alterations in hepatic expression of metabolic genes. Am. J. Physiol. Endocrinol. Metab. 2003, 285, E504–E511. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Xie, Y.; Berglund, E.D.; Coate, K.C.; He, T.T.; Katafuchi, T.; Xiao, G.; Potthoff, M.J.; Wei, W.; Wan, Y.; et al. The starvation hormone, fibroblast growth factor-21, extends lifespan in mice. Elife 2012, 1, e00065. [Google Scholar] [CrossRef] [PubMed]
- Hirao, A.; Tahara, Y.; Kimura, I.; Shibata, S. A balanced diet is necessary for proper entrainment signals of the mouse liver clock. PLoS ONE 2009, 4, e6909. [Google Scholar] [CrossRef] [PubMed]
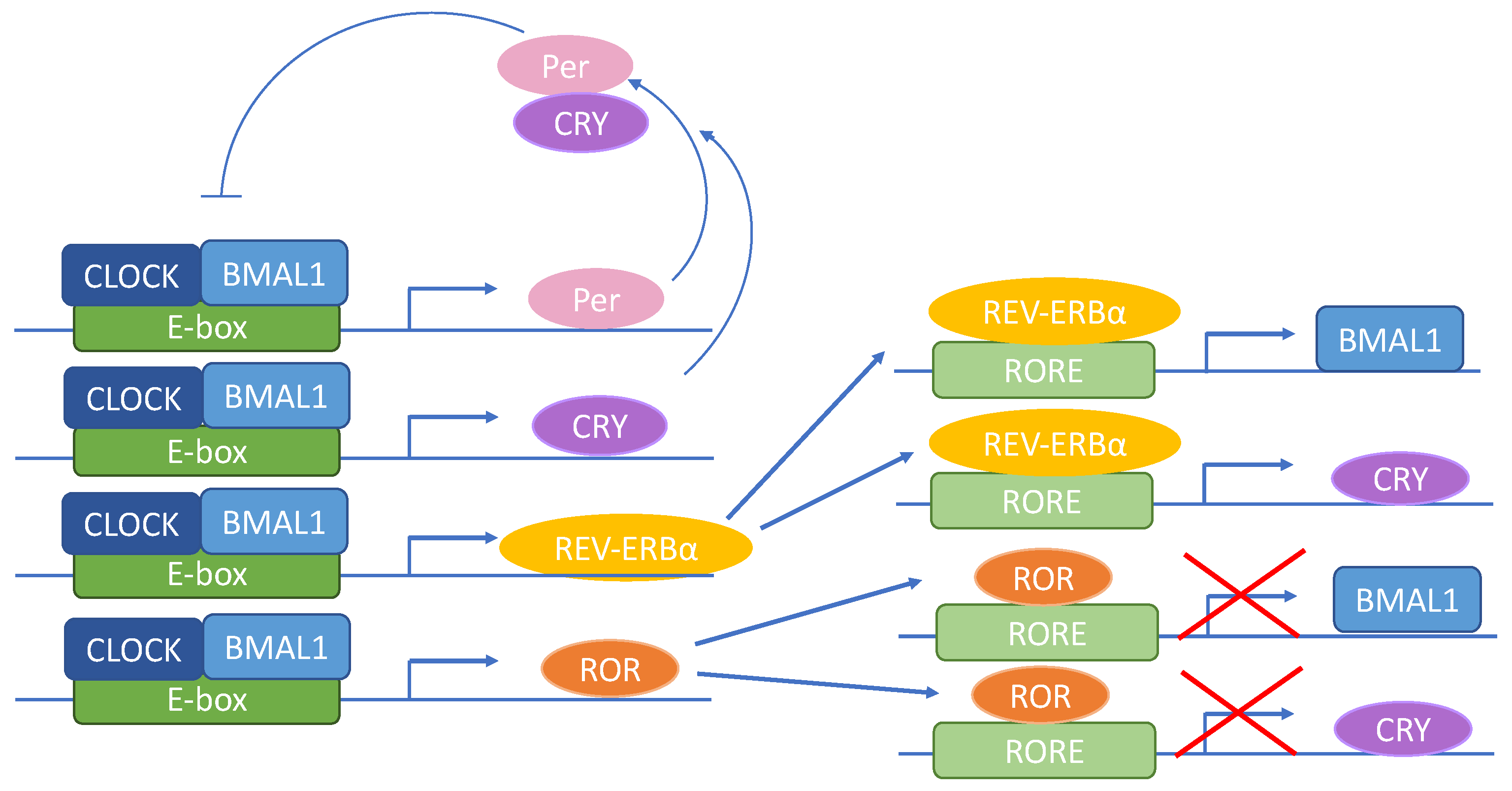
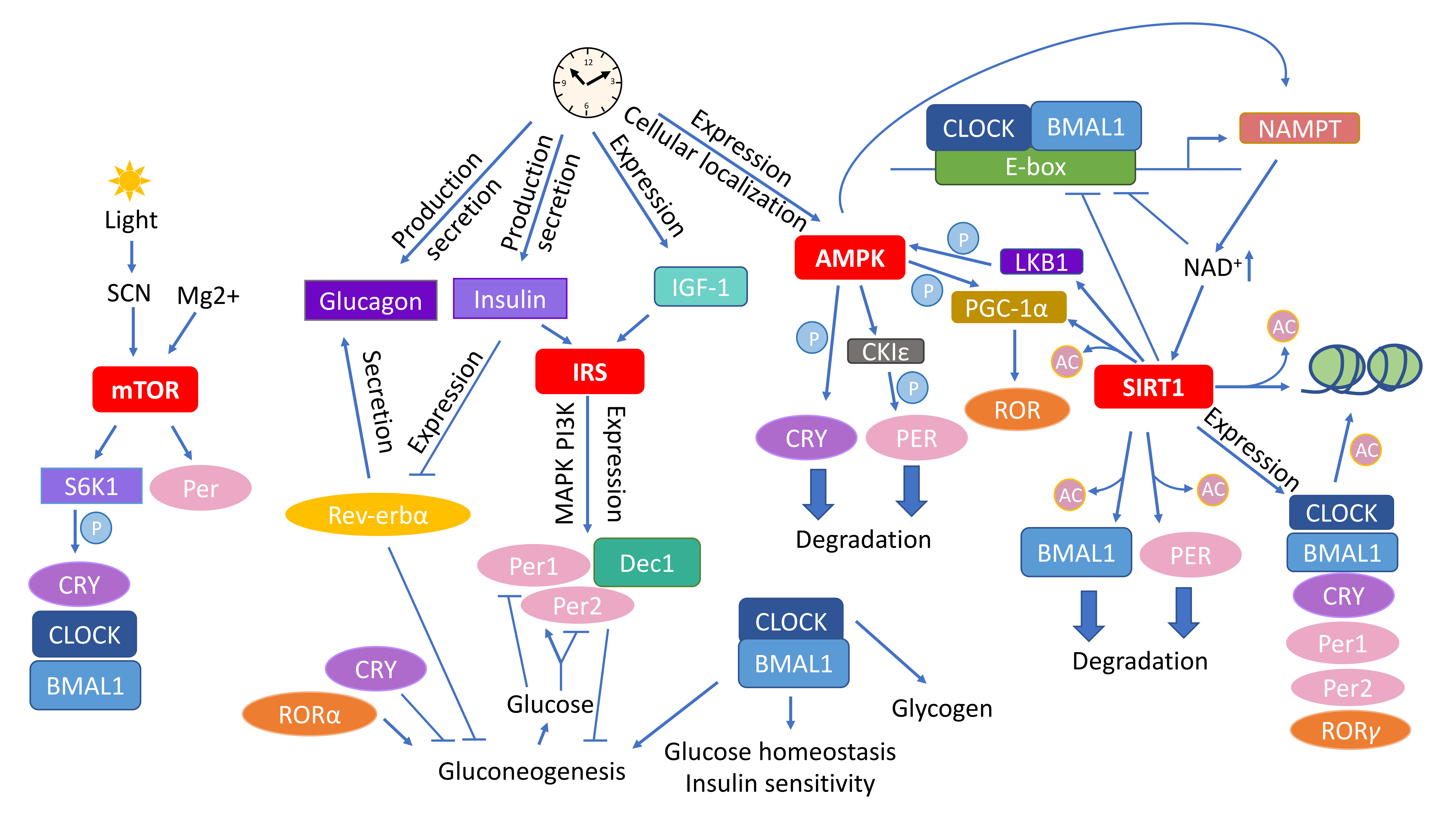
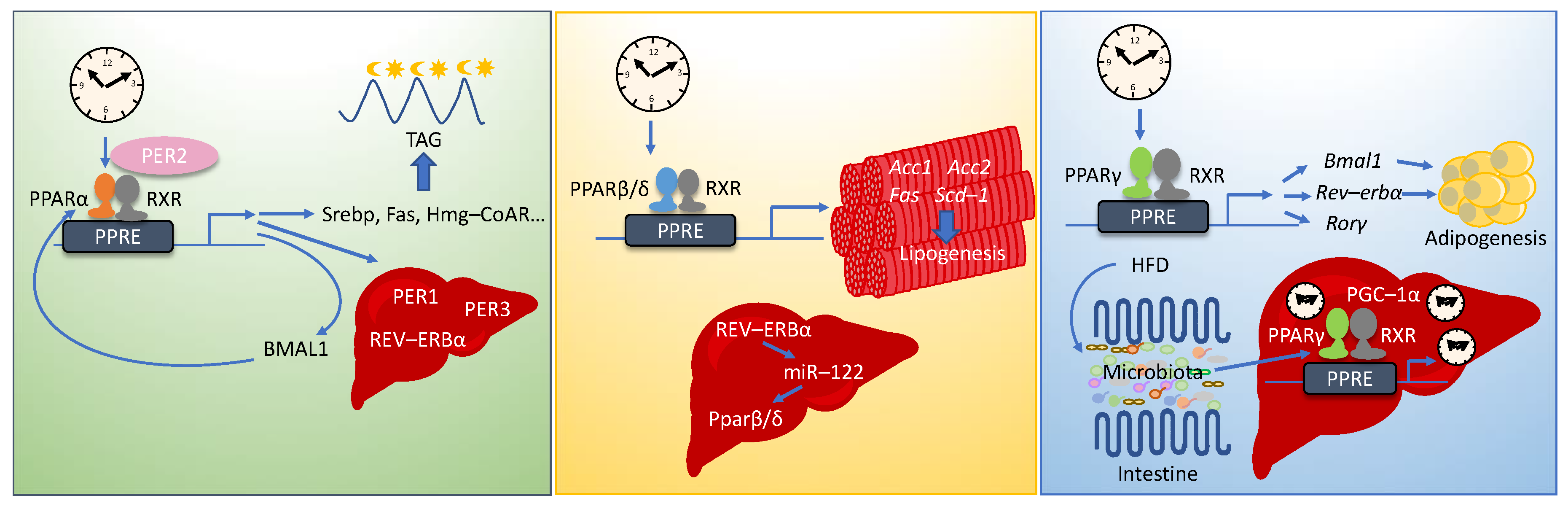
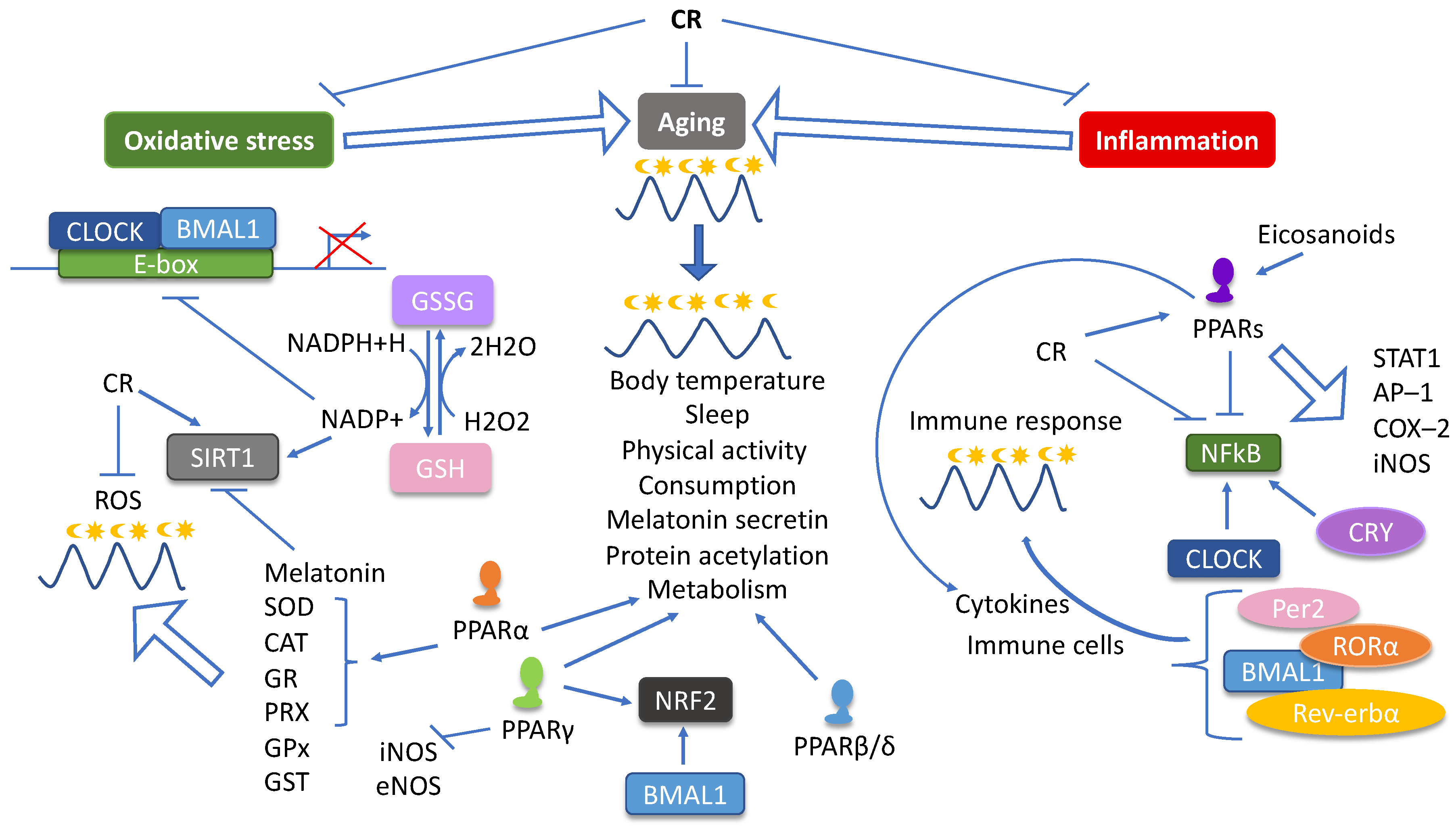
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Duszka, K.; Wahli, W. Peroxisome Proliferator-Activated Receptors as Molecular Links between Caloric Restriction and Circadian Rhythm. Nutrients 2020, 12, 3476. https://doi.org/10.3390/nu12113476
Duszka K, Wahli W. Peroxisome Proliferator-Activated Receptors as Molecular Links between Caloric Restriction and Circadian Rhythm. Nutrients. 2020; 12(11):3476. https://doi.org/10.3390/nu12113476
Chicago/Turabian StyleDuszka, Kalina, and Walter Wahli. 2020. "Peroxisome Proliferator-Activated Receptors as Molecular Links between Caloric Restriction and Circadian Rhythm" Nutrients 12, no. 11: 3476. https://doi.org/10.3390/nu12113476
APA StyleDuszka, K., & Wahli, W. (2020). Peroxisome Proliferator-Activated Receptors as Molecular Links between Caloric Restriction and Circadian Rhythm. Nutrients, 12(11), 3476. https://doi.org/10.3390/nu12113476





