Role of Personalized Nutrition in Chronic-Degenerative Diseases
Abstract
1. Introduction
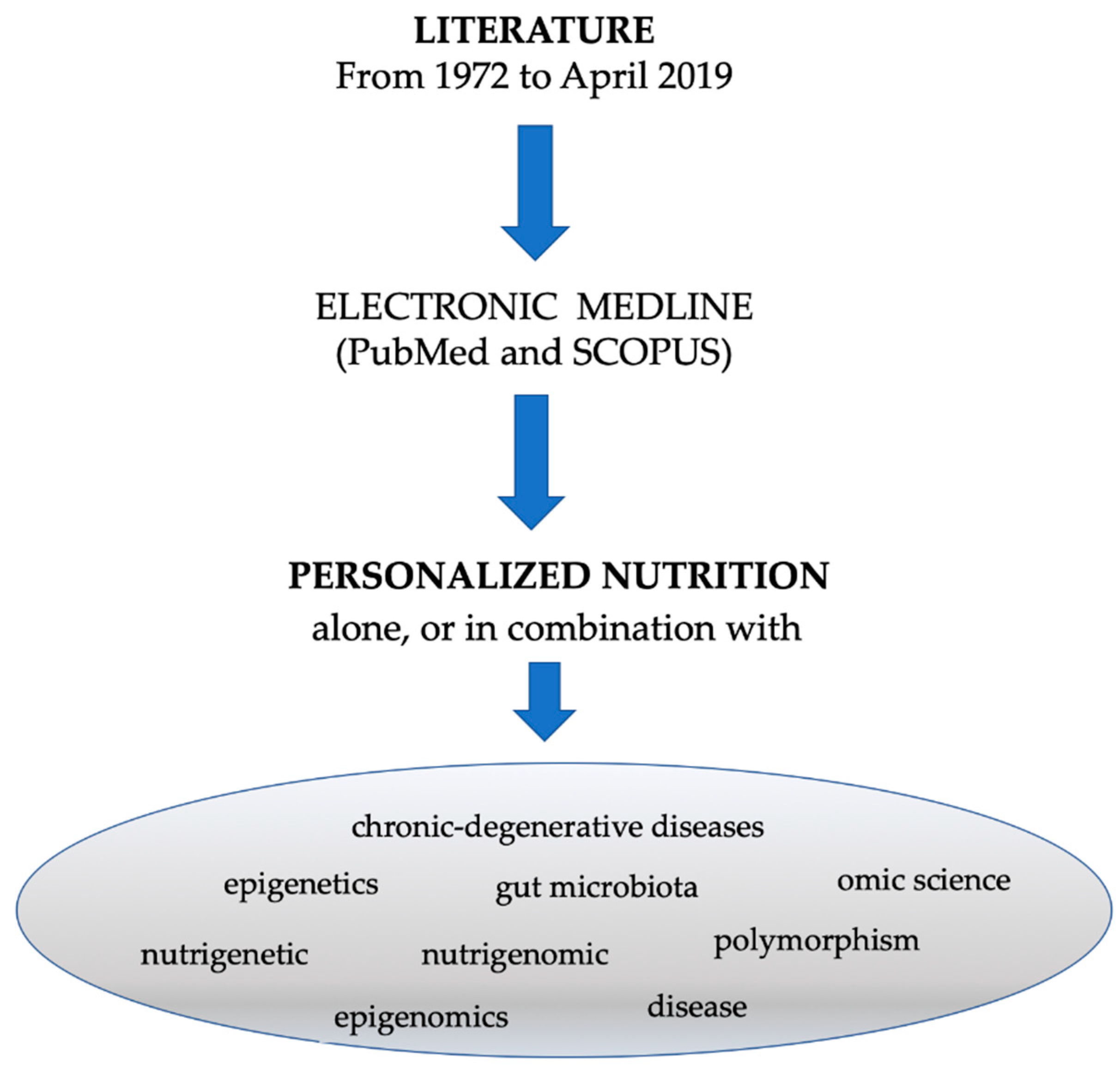
2. Methods
Search Strategy and Study Selection
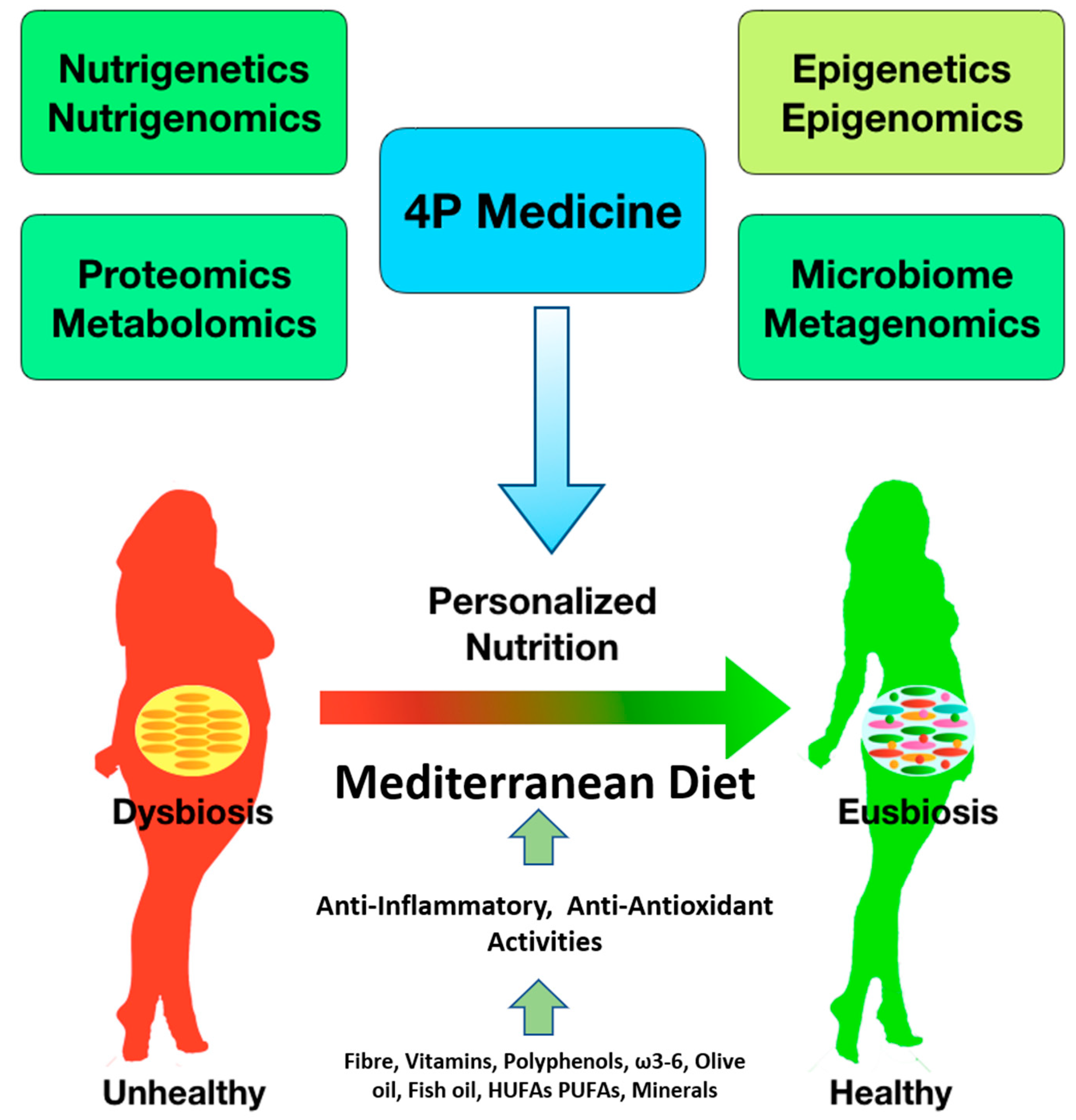
3. From Health to Disease Status
4. The -Omic Science
5. Nutrigenetics and Nutrigenomics
6. Epigenetics and Epigenomics
7. Proteomics and Metabolomics
8. From Disease to Health Status
9. Gut Microbiota and Personalized Nutrition
10. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviation
| 4P | Predictive: Preventive, Personalized and Proactive |
| ACE | Angiotensin-Converting-Enzyme |
| ALA | α-Linoleic Acid |
| BFC | Bioactive Food Components |
| BMI | Body Mass Index |
| CAMC | Cluster Analysis of Multivariate Correlation |
| CDD | Chronic-Degenerative Disease |
| CHD | Coronary Heart Disease |
| CSF | Cerebrospinal Fluid |
| CVD | Cardiovascular Diseases |
| DASH | Dietary Approaches to Stop Hypertension |
| DHA | Docoashexaenoic Acid |
| Dnmt | DNA Methyltransferase |
| EPA | Eicosapentaenoic Acid |
| FTO | Fat Mass and Obesity-Associated |
| GPS | Genome-Wide Polygenic Score |
| GSTM | Glutathione S-transferase Mu |
| GWAS | Genome-Wide Association Study |
| HDAC | Histone Deacetylase |
| HMGCR | 3-Hydroxy-3-Methylglutaryl-Coa Reductase Gene |
| HUFA | n-3 highly unsaturated fatty acids |
| IBD | Inflammatory Bowel Disease |
| IL | Interleukin |
| iNOS | Inducible Nitric Oxide Synthase |
| LAGB | Laparoscopic Adjustable Gastric Banding |
| LDL | Low-Density Lipoprotein |
| LX | Lipoxins |
| MC4R | Melanocortin- 4 Receptor |
| MD | Mediterranean Diet |
| MnSOD | Manganese Superoxide Dismutase |
| MS | Methionine Synthase |
| MTHFR | Methyltetrahydrofolate Reductase |
| MTRR | Methionine Synthase Reductase |
| NFkB | Nuclear Factor Kappa-Light-Chain-Enhancer of Activated B Cells |
| NWO | Normal Weight Obese |
| OmniHeart | Optimal Macro-Nutrient Intake to Prevent Heart Disease |
| PUFA | Polyunsaturated Fatty Acids |
| ROS | Reactive Oxygen Species |
| SAH | S-Adenosylhomocysteine |
| SAM | S-Adenosyl-Methionine |
| SCFA | Short Chain Fatty Acids |
| Se | Selenium |
| SNPs | Single Nucleotide Polymorphisms |
| T2DM | Type 2 Diabetes Mellitus |
| TC | Total Cholesterol |
| TNFR2 | Tumor Necrosis Factor Receptor-2 |
| TNFα | Tumor Necrosis Factor-α |
| WAT | White Adipose Tissue |
References
- Bragazzi, N.L. Situating Nutri-Ethics at the Junction of Nutrigenomics and Nutriproteomics in Postgenomics Medicine. Curr. Pharm. Pers. Med. 2013, 11, 162–166. [Google Scholar] [CrossRef] [PubMed]
- Virgili, F.; Perozzi, G. How does Nutrigenomics impact human health? IUBMB Life 2008, 60, 341–344. [Google Scholar] [CrossRef] [PubMed]
- Wolffe, A.P.; Matzke, M.A. Epigenetics: Regulation through repression. Science 1999, 286, 481–486. [Google Scholar] [CrossRef] [PubMed]
- Juma, S.; Imrhan, V.; Vijayagopal, P.; Prasad, C. Prescribing personalized nutrition for cardiovascular health: Are we ready? J. Nutrigenet. Nutr. 2014, 7, 153–160. [Google Scholar] [CrossRef] [PubMed]
- Kang, J.X. Nutritional problems and solutions for the modern health epidemic. J. Nutrigenet. Nutr. 2014, 7, 188–190. [Google Scholar] [CrossRef]
- Gillies, P.J. Nutrigenomics: The Rubicon of molecular nutrition. J. Am. Diet. Assoc. 2003, 103, S50–S55. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Young, V.R. 2001 W.O. Atwater Memorial Lecture and the 2001 ASNS President’s Lecture: Human nutrient requirements: The challenge of the post-genome era. J. Nutr. 2002, 132, 621–629. [Google Scholar] [CrossRef][Green Version]
- Jones, P.A.; Takai, D. The role of DNA methylation in mammalian epigenetics. Science 2001, 293, 1068–1070. [Google Scholar] [CrossRef]
- Feinberg, A.P. Methylation meets genomics. Nat. Genet. 2001, 27, 9–10. [Google Scholar] [CrossRef]
- Green, M.R.; van der Ouderaa, F. Nutrigenetics: Where next for the foods industry? Pharm. J. 2003, 3, 191–193. [Google Scholar] [CrossRef]
- Ross, J. Control of messenger RNA stability in higher eukaryotes. Trends Genet. 1996, 12, 171–175. [Google Scholar] [CrossRef]
- Emilsson, V.; Thorleifsson, G.; Zhang, B.; Leonardson, A.S.; Zink, F.; Zhu, J.; Carlson, S.; Helgason, A.; Walters, G.B.; Gunnarsdottir, S.; et al. Genetics of gene expression and its effect on disease. Nature 2008, 452, 423–428. [Google Scholar] [CrossRef] [PubMed]
- Simmons, L.A.; Dinan, M.A.; Robinson, T.J.; Snyderman, R. Personalized medicine is more than genomic medicine: Confusion over terminology impedes progress towards personalized healthcare. Per. Med. 2012, 9, 85–91. [Google Scholar] [CrossRef] [PubMed]
- German, J.B.; Roberts, M.A.; Watkins, S.M. Genomics and metabolomics as markers for the interaction of diet and health: Lessons from lipids. J. Nutr. 2003, 133, 2078S–2083S. [Google Scholar] [CrossRef] [PubMed]
- 1000 Genomes Project Consortium; Auton, A.; Brooks, L.D.; Durbin, R.M.; Garrison, E.P.; Kang, H.M.; Korbel, J.O.; Marchini, J.L.; McCarthy, S.; McVean, G.A.; et al. A global reference for human genetic variation. Nature 2015, 526, 68–74. [Google Scholar] [CrossRef] [PubMed]
- Di Renzo, L.; Colica, C.; Carraro, A.; Cenci Goga, B.; Marsella, L.T.; Botta, R.; Colombo, M.L.; Gratteri, S.; Chang, T.F.; Droli, M.; et al. Food safety and nutritional quality for the prevention of non communicable diseases: The Nutrient, hazard Analysis and Critical Control Point process (NACCP). J. Transl. Med. 2015, 13, 128. [Google Scholar] [CrossRef] [PubMed]
- Di Renzo, L.; Marsella, L.T.; Carraro, A.; Valente, R.; Gualtieri, P.; Gratteri, S.; Tomasi, D.; Gaiotti, F.; De Lorenzo, A. Changes in LDL Oxidative Status and Oxidative and Inflammatory Gene Expression after Red Wine Intake in Healthy People: A Randomized Trial. Mediat. Inflamm. 2015, 2015, 317348. [Google Scholar] [CrossRef]
- Wallace, H. Your Diet Tailored to Your Genes: Preventing Diseases or Misleading Marketing? GeneWatch UK: Derbyshire, UK, 2006. [Google Scholar]
- NCD Risk Factor Collaboration (NCD-RisC). Worldwide trends in body-mass index, underweight, overweight, and obesity from 1975 to 2016: A pooled analysis of 2416 population-based measurement studies in 128.9 million children, adolescents, and adults. Lancet 2017, 390, 2627–2642. [Google Scholar] [CrossRef]
- Barsh, G.S.; Farooqi, I.S.; O’Rahilly, S. Genetics of body-weight regulation. Nature 2000, 404, 644–651. [Google Scholar] [CrossRef]
- Vaisse, C.; Clement, K.; Durand, E.; Hercberg, S.; Guy-Grand, B.; Froguel, P. Melanocortin-4 receptor mutations are a frequent and heterogeneous cause of morbid obesity. J. Clin. Investig. 2000, 106, 253–262. [Google Scholar] [CrossRef]
- Farooqi, I.S.; Keogh, J.M.; Yeo, G.S.; Lank, E.J.; Cheetham, T.; O’Rahilly, S. Clinical spectrum of obesity and mutations in the melanocortin 4 receptor gene. N. Engl. J. Med. 2003, 348, 1085–1095. [Google Scholar] [CrossRef] [PubMed]
- Larsen, L.H.; Echwald, S.M.; Sorensen, T.I.; Andersen, T.; Wulff, B.S.; Pedersen, O. Prevalence of mutations and functional analyses of melanocortin 4 receptor variants identified among 750 men with juvenile-onset obesity. J. Clin. Endocrinol. Metab. 2005, 90, 219–224. [Google Scholar] [CrossRef] [PubMed]
- Golan, D.; Lander, E.S.; Rosset, S. Measuring missing heritability: Inferring the contribution of common variants. Proc. Natl. Acad. Sci. USA 2014, 111, E5272–E5281. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; Bakshi, A.; Vinkhuyzen, A.A.; Hemani, G.; Lee, S.H.; Nolte, I.M.; van Vliet-Ostaptchouk, J.V.; Snieder, H.; LifeLines Cohort, S.; Esko, T.; et al. Dominance genetic variation contributes little to the missing heritability for human complex traits. Am. J. Hum. Genet. 2015, 96, 377–385. [Google Scholar] [CrossRef] [PubMed]
- Chatterjee, N.; Wheeler, B.; Sampson, J.; Hartge, P.; Chanock, S.J.; Park, J.H. Projecting the performance of risk prediction based on polygenic analyses of genome-wide association studies. Nat. Genet. 2013, 45, 400–405. [Google Scholar] [CrossRef] [PubMed]
- Locke, A.E.; Kahali, B.; Berndt, S.I.; Justice, A.E.; Pers, T.H.; Day, F.R.; Powell, C.; Vedantam, S.; Buchkovich, M.L.; Yang, J.; et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 2015, 518, 197–206. [Google Scholar] [CrossRef] [PubMed]
- Di Renzo, L.; Cioccoloni, G.; Falco, S.; Abenavoli, L.; Moia, A.; Sinibaldi Salimei, P.; De Lorenzo, A. Influence of FTO rs9939609 and Mediterranean diet on body composition and weight loss: A randomized clinical trial. J. Transl. Med. 2018, 16, 308. [Google Scholar] [CrossRef]
- Olszewski, P.K.; Fredriksson, R.; Olszewska, A.M.; Stephansson, O.; Alsio, J.; Radomska, K.J.; Levine, A.S.; Schioth, H.B. Hypothalamic FTO is associated with the regulation of energy intake not feeding reward. BMC Neurosci. 2009, 10, 129. [Google Scholar] [CrossRef]
- Cheung, M.K.; Gulati, P.; O’Rahilly, S.; Yeo, G.S. FTO expression is regulated by availability of essential amino acids. Int. J. Obes. (Lond.) 2013, 37, 744–747. [Google Scholar] [CrossRef]
- Tanofsky-Kraff, M.; Han, J.C.; Anandalingam, K.; Shomaker, L.B.; Columbo, K.M.; Wolkoff, L.E.; Kozlosky, M.; Elliott, C.; Ranzenhofer, L.M.; Roza, C.A.; et al. The FTO gene rs9939609 obesity-risk allele and loss of control over eating. Am. J. Clin. Nutr. 2009, 90, 1483–1488. [Google Scholar] [CrossRef]
- Karra, E.; O’Daly, O.G.; Choudhury, A.I.; Yousseif, A.; Millership, S.; Neary, M.T.; Scott, W.R.; Chandarana, K.; Manning, S.; Hess, M.E.; et al. A link between FTO, ghrelin, and impaired brain food-cue responsivity. J. Clin. Investig. 2013, 123, 3539–3551. [Google Scholar] [CrossRef] [PubMed]
- Di Renzo, L.; Marsella, L.T.; Sarlo, F.; Soldati, L.; Gratteri, S.; Abenavoli, L.; De Lorenzo, A. C677T gene polymorphism of MTHFR and metabolic syndrome: Response to dietary intervention. J. Transl. Med. 2014, 12, 329. [Google Scholar] [CrossRef] [PubMed]
- Fan, S.J.; Yang, B.Y.; Zhi, X.Y.; He, M.; Wang, D.; Wang, Y.X.; Wang, Y.N.; Wei, J.; Zheng, Q.M.; Sun, G.F. Are MTHFR C677T and MTRR A66G Polymorphisms Associated with Overweight/Obesity Risk? From a Case-Control to a Meta-Analysis of 30,327 Subjects. Int. J. Mol. Sci. 2015, 16, 11849–11863. [Google Scholar] [CrossRef] [PubMed]
- Bailey, L.B.; Stover, P.J.; McNulty, H.; Fenech, M.F.; Gregory, J.F., 3rd; Mills, J.L.; Pfeiffer, C.M.; Fazili, Z.; Zhang, M.; Ueland, P.M.; et al. Biomarkers of Nutrition for Development-Folate Review. J. Nutr. 2015, 145, 1636S–1680S. [Google Scholar] [CrossRef] [PubMed]
- Fontaine-Bisson, B.; Wolever, T.M.; Chiasson, J.L.; Rabasa-Lhoret, R.; Maheux, P.; Josse, R.G.; Leiter, L.A.; Rodger, N.W.; Ryan, E.A.; Connelly, P.W.; et al. Genetic polymorphisms of tumor necrosis factor-alpha modify the association between dietary polyunsaturated fatty acids and fasting HDL-cholesterol and apo A-I concentrations. Am. J. Clin. Nutr. 2007, 86, 768–774. [Google Scholar] [CrossRef]
- Di Renzo, L.; Sarlo, F.; Petramala, L.; Iacopino, L.; Monteleone, G.; Colica, C.; De Lorenzo, A. Association between -308 G/A TNF-alpha polymorphism and appendicular skeletal muscle mass index as a marker of sarcopenia in normal weight obese syndrome. Dis. Markers 2013, 35, 615–623. [Google Scholar] [CrossRef] [PubMed]
- Hedayati, M.; Sharifi, K.; Rostami, F.; Daneshpour, M.S.; Zarif Yeganeh, M.; Azizi, F. Association between TNF-alpha promoter G-308A and G-238A polymorphisms and obesity. Mol. Biol. Rep. 2012, 39, 825–829. [Google Scholar] [CrossRef]
- Wang, Y.; Ng, M.C.; So, W.Y.; Ma, R.; Ko, G.T.; Tong, P.C.; Chan, J.C. Association between tumour necrosis factor-alpha G-308A polymorphism and risk of nephropathy in obese Chinese type 2 diabetic patients. Nephrol. Dial. Transplant. 2005, 20, 2733–2738. [Google Scholar] [CrossRef]
- Dalziel, B.; Gosby, A.K.; Richman, R.M.; Bryson, J.M.; Caterson, I.D. Association of the TNF-alpha -308 G/A Promoter Polymorphism with Insulin Resistance in Obesity. Obes. Res. 2002, 10, 401–407. [Google Scholar] [CrossRef]
- Jackson, K.G.; Li, Y.; Ryan, M.F.; Gibney, E.R.; Brennan, L.; Roche, H.M.; Williams, C.M.; Lovegrove, J.A.; Vimaleswaran, K.S. Association of the tumor necrosis factor-alpha promoter polymorphism with change in triacylglycerol response to sequential meals. Nutr. J. 2016, 15, 70. [Google Scholar] [CrossRef]
- Fernandez-Real, J.M.; Vendrell, J.; Ricart, W.; Broch, M.; Gutierrez, C.; Casamitjana, R.; Oriola, J.; Richart, C. Polymorphism of the tumor necrosis factor-alpha receptor 2 gene is associated with obesity, leptin levels, and insulin resistance in young subjects and diet-treated type 2 diabetic patients. Diabetes Care 2000, 23, 831–837. [Google Scholar] [CrossRef] [PubMed]
- Di Renzo, L.; Bertoli, A.; Bigioni, M.; Del Gobbo, V.; Premrov, M.G.; Calabrese, V.; Di Daniele, N.; De Lorenzo, A. Body composition and -174G/C interleukin-6 promoter gene polymorphism: Association with progression of insulin resistance in normal weight obese syndrome. Curr. Pharm. Des. 2008, 14, 2699–2706. [Google Scholar] [CrossRef] [PubMed]
- Garcia, M.D.C.; Pazos, P.; Lima, L.; Dieguez, C. Regulation of Energy Expenditure and Brown/Beige Thermogenic Activity by Interleukins: New Roles for Old Actors. Int. J. Mol. Sci. 2018, 19, 2569. [Google Scholar] [CrossRef] [PubMed]
- Di Renzo, L.; Carbonelli, M.G.; Bianchi, A.; Iacopino, L.; Fiorito, R.; Di Daniele, N.; De Lorenzo, A. Body composition changes after laparoscopic adjustable gastric banding: What is the role of -174G>C interleukin-6 promoter gene polymorphism in the therapeutic strategy? Int. J. Obes. (Lond.) 2012, 36, 369–378. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Di Renzo, L.; Rizzo, M.; Sarlo, F.; Colica, C.; Iacopino, L.; Domino, E.; Sergi, D.; De Lorenzo, A. Effects of dark chocolate in a population of normal weight obese women: A pilot study. Eur. Rev. Med. Pharmacol. Sci. 2013, 17, 2257–2266. [Google Scholar]
- Colica, C.; Avolio, E.; Bollero, P.; Costa de Miranda, R.; Ferraro, S.; Sinibaldi Salimei, P.; De Lorenzo, A.; Di Renzo, L. Evidences of a New Psychobiotic Formulation on Body Composition and Anxiety. Mediat. Inflamm. 2017, 2017, 5650627. [Google Scholar] [CrossRef] [PubMed]
- De Lorenzo, A.; Costacurta, M.; Merra, G.; Gualtieri, P.; Cioccoloni, G.; Marchetti, M.; Varvaras, D.; Docimo, R.; Di Renzo, L. Can psychobiotics intake modulate psychological profile and body composition of women affected by normal weight obese syndrome and obesity? A double blind randomized clinical trial. J. Transl. Med. 2017, 15, 135. [Google Scholar] [CrossRef]
- Khera, A.V.; Chaffin, M.; Wade, K.H.; Zahid, S.; Brancale, J.; Xia, R.; Distefano, M.; Senol-Cosar, O.; Haas, M.E.; Bick, A.; et al. Polygenic Prediction of Weight and Obesity Trajectories from Birth to Adulthood. Cell 2019, 177, 587–596.e589. [Google Scholar] [CrossRef]
- De Lorenzo, A.; Soldati, L.; Sarlo, F.; Calvani, M.; Di Lorenzo, N.; Di Renzo, L. New obesity classification criteria as a tool for bariatric surgery indication. World J. Gastroenterol. 2016, 22, 681–703. [Google Scholar] [CrossRef]
- Kato, H.; Takahashi, S.; Saito, K. Omics and integrated omics for the promotion of food and nutrition science. J. Tradit. Complement. Med. 2011, 1, 25–30. [Google Scholar] [CrossRef]
- Ferguson, L.R.; De Caterina, R.; Gorman, U.; Allayee, H.; Kohlmeier, M.; Prasad, C.; Choi, M.S.; Curi, R.; de Luis, D.A.; Gil, A.; et al. Guide and Position of the International Society of Nutrigenetics/Nutrigenomics on Personalised Nutrition: Part 1—Fields of Precision Nutrition. J. Nutrigenet. Nutr. 2016, 9, 12–27. [Google Scholar] [CrossRef] [PubMed]
- Merra, G.; Miranda, R.; Barrucco, S.; Gualtieri, P.; Mazza, M.; Moriconi, E.; Marchetti, M.; Chang, T.F.; De Lorenzo, A.; Di Renzo, L. Very-low-calorie ketogenic diet with aminoacid supplement versus very low restricted-calorie diet for preserving muscle mass during weight loss: A pilot double-blind study. Eur. Rev. Med. Pharmacol. Sci. 2016, 20, 2613–2621. [Google Scholar] [PubMed]
- Hasin, Y.; Seldin, M.; Lusis, A. Multi-omics approaches to disease. Genome Biol. 2017, 18, 83. [Google Scholar] [CrossRef] [PubMed]
- Weckwerth, W. Metabolomics in systems biology. Annu. Rev. Plant. Biol. 2003, 54, 669–689. [Google Scholar] [CrossRef] [PubMed]
- Kang, J.X. The coming of age of nutrigenetics and nutrigenomics. J. Nutrigenet. Nutr. 2012, 5, I–II. [Google Scholar] [CrossRef] [PubMed]
- Scapagnini, G.; Davinelli, S.; Di Renzo, L.; De Lorenzo, A.; Olarte, H.H.; Micali, G.; Cicero, A.F.; Gonzalez, S. Cocoa bioactive compounds: Significance and potential for the maintenance of skin health. Nutrients 2014, 6, 3202–3213. [Google Scholar] [CrossRef]
- Berna, G.; Oliveras-Lopez, M.J.; Jurado-Ruiz, E.; Tejedo, J.; Bedoya, F.; Soria, B.; Martin, F. Nutrigenetics and nutrigenomics insights into diabetes etiopathogenesis. Nutrients 2014, 6, 5338–5369. [Google Scholar] [CrossRef]
- Bollero, P.; Di Renzo, L.; Franco, R.; Rampello, T.; Pujia, A.; Merra, G.; De Lorenzo, A.; Docimo, R. Effects of new probiotic mouthwash in patients with diabetes mellitus and cardiovascular diseases. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 5827–5836. [Google Scholar] [CrossRef]
- Alfredo Martinez, J. Perspectives on personalized nutrition for obesity. J. Nutrigenet. Nutr. 2014, 7, I–III. [Google Scholar] [CrossRef]
- Di Renzo, L.; Di Pierro, D.; Bigioni, M.; Sodi, V.; Galvano, F.; Cianci, R.; La Fauci, L.; De Lorenzo, A. Is antioxidant plasma status in humans a consequence of the antioxidant food content influence? Eur. Rev. Med. Pharmacol. Sci. 2007, 11, 185–192. [Google Scholar]
- Peddireddy, V.; Badabagni, S.P.; Gundimeda, S.D.; Mamidipudi, V.; Penagaluru, P.R.; Mundluru, H.P. Association of CYP1A1, GSTM1 and GSTT1 gene polymorphisms with risk of non-small cell lung cancer in Andhra Pradesh region of South India. Eur. J. Med. Res. 2016, 21, 17. [Google Scholar] [CrossRef] [PubMed]
- Neeha, V.S.; Kinth, P. Nutrigenomics research: A review. J. Food Sci. Technol. 2013, 50, 415–428. [Google Scholar] [CrossRef] [PubMed]
- Kang, J.X. Nutrigenomics and systems biology. J. Nutrigenet. Nutr. 2012, 5, I–II. [Google Scholar] [CrossRef] [PubMed]
- Kang, J.X. Nutrigenomics and cancer therapy. J. Nutrigenet. Nutr. 2013, 6, I–II. [Google Scholar] [CrossRef] [PubMed]
- Andreoli, A.; De Lorenzo, A.; Cadeddu, F.; Iacopino, L.; Grande, M. New trends in nutritional status assessment of cancer patients. Eur. Rev. Med. Pharmacol. Sci. 2011, 15, 469–480. [Google Scholar]
- Lai, C.Q.; Corella, D.; Demissie, S.; Cupples, L.A.; Adiconis, X.; Zhu, Y.; Parnell, L.D.; Tucker, K.L.; Ordovas, J.M. Dietary intake of n-6 fatty acids modulates effect of apolipoprotein A5 gene on plasma fasting triglycerides, remnant lipoprotein concentrations, and lipoprotein particle size: The Framingham Heart Study. Circulation 2006, 113, 2062–2070. [Google Scholar] [CrossRef] [PubMed]
- Merched, A.J.; Chan, L. Nutrigenetics and nutrigenomics of atherosclerosis. Curr. Atheroscler. Rep. 2013, 15, 328. [Google Scholar] [CrossRef]
- Tak, P.P.; Firestein, G.S. NF-kappaB: A key role in inflammatory diseases. J. Clin. Investig. 2001, 107, 7–11. [Google Scholar] [CrossRef]
- Sales, N.M.; Pelegrini, P.B.; Goersch, M.C. Nutrigenomics: Definitions and advances of this new science. J. Nutr. Metab. 2014, 2014, 202759. [Google Scholar] [CrossRef]
- Oltean, S.; Banerjee, R. Nutritional modulation of gene expression and homocysteine utilization by vitamin B12. J. Biol. Chem. 2003, 278, 20778–20784. [Google Scholar] [CrossRef]
- Kolhouse, J.F.; Allen, R.H. Recognition of two intracellular cobalamin binding proteins and their identification as methylmalonyl-CoA mutase and methionine synthetase. Proc. Natl. Acad. Sci. USA 1977, 74, 921–925. [Google Scholar] [CrossRef] [PubMed]
- Ashfield-Watt, P.A.; Moat, S.J.; Newcombe, R.G.; McDowell, I.F. Effect of supplementation with folic-acid on relation between plasma homocysteine, folate, and vitamin B12. Lancet 2002, 360, 172–173. [Google Scholar] [CrossRef]
- Davis, C.D.; Uthus, E.O. Dietary selenite and azadeoxycytidine treatments affect dimethylhydrazine-induced aberrant crypt formation in rat colon and DNA methylation in HT-29 cells. J. Nutr. 2002, 132, 292–297. [Google Scholar] [CrossRef] [PubMed]
- Rotruck, J.T.; Pope, A.L.; Ganther, H.E.; Swanson, A.B.; Hafeman, D.G.; Hoekstra, W.G. Selenium: Biochemical role as a component of glutathione peroxidase. Science 1973, 179, 588–590. [Google Scholar] [CrossRef] [PubMed]
- Majumder, S.; Ghoshal, K.; Summers, D.; Bai, S.; Datta, J.; Jacob, S.T. Chromium(VI) down-regulates heavy metal-induced metallothionein gene transcription by modifying transactivation potential of the key transcription factor, metal-responsive transcription factor 1. J. Biol. Chem. 2003, 278, 26216–26226. [Google Scholar] [CrossRef] [PubMed]
- Bi, Y.; Palmiter, R.D.; Wood, K.M.; Ma, Q. Induction of metallothionein I by phenolic antioxidants requires metal-activated transcription factor 1 (MTF-1) and zinc. Biochem. J. 2004, 380, 695–703. [Google Scholar] [CrossRef] [PubMed]
- Arthur, J.R. The glutathione peroxidases. Cell Mol. Life Sci. 2000, 57, 1825–1835. [Google Scholar] [CrossRef] [PubMed]
- Soldati, L.; Di Renzo, L.; Jirillo, E.; Ascierto, P.A.; Marincola, F.M.; De Lorenzo, A. The influence of diet on anti-cancer immune responsiveness. J. Transl. Med. 2018, 16, 75. [Google Scholar] [CrossRef] [PubMed]
- Ardekani, A.M.; Jabbari, S. Nutrigenomics and cancer. Avicenna J. Med. Biotechnol. 2009, 1, 9–17. [Google Scholar]
- Verma, M.; Dunn, B.K.; Ross, S.; Jain, P.; Wang, W.; Hayes, R.; Umar, A. Early detection and risk assessment: Proceedings and recommendations from the Workshop on Epigenetics in Cancer Prevention. Ann. N. Y. Acad. Sci. 2003, 983, 298–319. [Google Scholar] [CrossRef]
- Alaskhar Alhamwe, B.; Khalaila, R.; Wolf, J.; von Bulow, V.; Harb, H.; Alhamdan, F.; Hii, C.S.; Prescott, S.L.; Ferrante, A.; Renz, H.; et al. Histone modifications and their role in epigenetics of atopy and allergic diseases. Allergy Asthma Clin. Immunol. 2018, 14, 39. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; He, C.; Moore, S.C.; Ausio, J. Effects of histone acetylation on the solubility and folding of the chromatin fiber. J. Biol. Chem. 2001, 276, 12764–12768. [Google Scholar] [CrossRef] [PubMed]
- Xiao, H.; Hasegawa, T.; Isobe, K. p300 collaborates with Sp1 and Sp3 in p21(waf1/cip1) promoter activation induced by histone deacetylase inhibitor. J. Biol. Chem. 2000, 275, 1371–1376. [Google Scholar] [CrossRef] [PubMed]
- Lu, Q.; Yang, Y.T.; Chen, C.S.; Davis, M.; Byrd, J.C.; Etherton, M.R.; Umar, A.; Chen, C.S. Zn2+-chelating motif-tethered short-chain fatty acids as a novel class of histone deacetylase inhibitors. J. Med. Chem. 2004, 47, 467–474. [Google Scholar] [CrossRef] [PubMed]
- Razin, A.; Cedar, H. DNA methylation and genomic imprinting. Cell 1994, 77, 473–476. [Google Scholar] [CrossRef]
- Knowles, L.M.; Milner, J.A. Diallyl disulfide inhibits p34(cdc2) kinase activity through changes in complex formation and phosphorylation. Carcinogenesis 2000, 21, 1129–1134. [Google Scholar] [CrossRef] [PubMed]
- Jin, B.; Li, Y.; Robertson, K.D. DNA methylation: Superior or subordinate in the epigenetic hierarchy? Genes Cancer 2011, 2, 607–617. [Google Scholar] [CrossRef]
- Milner, J.A. Incorporating basic nutrition science into health interventions for cancer prevention. J. Nutr. 2003, 133, 3820S–3826S. [Google Scholar] [CrossRef]
- Jeffery, C.J. Moonlighting proteins. Trends Biochem. Sci. 1999, 24, 8–11. [Google Scholar] [CrossRef]
- Chambers, G.; Lawrie, L.; Cash, P.; Murray, G.I. Proteomics: A new approach to the study of disease. J. Pathol. 2000, 192, 280–288. [Google Scholar] [CrossRef]
- Vogel, G. Can old cells learn new tricks? Science 2000, 287, 1418–1419. [Google Scholar] [CrossRef] [PubMed]
- Noguchi, Y.; Sakai, R.; Kimura, T. Metabolomics and its potential for assessment of adequacy and safety of amino acid intake. J. Nutr. 2003, 133, 2097S–2100S. [Google Scholar] [CrossRef] [PubMed]
- Ozalp, I.; Young, V.R.; Nagchaudhuri, J.; Tontisirin, K.; Scrimshaw, N.S. Plasma amino acid response in young men given diets devoid of single essential amino acids. J. Nutr. 1972, 102, 1147–1158. [Google Scholar] [CrossRef] [PubMed]
- Moundras, C.; Remesy, C.; Demigne, C. Dietary protein paradox: Decrease of amino acid availability induced by high-protein diets. Am. J. Physiol. 1993, 264, G1057–G1065. [Google Scholar] [CrossRef] [PubMed]
- Endo, Y.; Fu, Z.; Abe, K.; Arai, S.; Kato, H. Dietary protein quantity and quality affect rat hepatic gene expression. J. Nutr. 2002, 132, 3632–3637. [Google Scholar] [CrossRef] [PubMed]
- Oz, H.S. Nutrients, Infectious and Inflammatory Diseases. Nutrients 2017, 9, 1085. [Google Scholar] [CrossRef] [PubMed]
- Erdmann, K.; Cheung, B.W.; Schroder, H. The possible roles of food-derived bioactive peptides in reducing the risk of cardiovascular disease. J. Nutr. Biochem. 2008, 19, 643–654. [Google Scholar] [CrossRef]
- Cuervo, A. Calorie restriction and aging: The ultimate “cleansing diet”. J. Gerontol. A Biol. Sci. Med. Sci. 2008, 63, 547–549. [Google Scholar] [CrossRef]
- Mancuso, M.; Coppede, F.; Murri, L.; Siciliano, G. Mitochondrial cascade hypothesis of Alzheimer’s disease: Myth or reality? Antioxid. Redox Signal. 2007, 9, 1631–1646. [Google Scholar] [CrossRef] [PubMed]
- Mancuso, C.; Scapagini, G.; Curro, D.; Giuffrida Stella, A.M.; De Marco, C.; Butterfield, D.A.; Calabrese, V. Mitochondrial dysfunction, free radical generation and cellular stress response in neurodegenerative disorders. Front. Biosci. 2007, 12, 1107–1123. [Google Scholar] [CrossRef]
- Dedoussis, G.V.; Kanoni, S.; Mariani, E.; Cattini, L.; Herbein, G.; Fulop, T.; Varin, A.; Rink, L.; Jajte, J.; Monti, D.; et al. Mediterranean diet and plasma concentration of inflammatory markers in old and very old subjects in the ZINCAGE population study. Clin. Chem. Lab. Med. 2008, 46, 990–996. [Google Scholar] [CrossRef] [PubMed]
- Esposito, K.; Marfella, R.; Ciotola, M.; Di Palo, C.; Giugliano, F.; Giugliano, G.; D’Armiento, M.; D’Andrea, F.; Giugliano, D. Effect of a mediterranean-style diet on endothelial dysfunction and markers of vascular inflammation in the metabolic syndrome: A randomized trial. JAMA 2004, 292, 1440–1446. [Google Scholar] [CrossRef] [PubMed]
- De Lorenzo, A.; Noce, A.; Bigioni, M.; Calabrese, V.; Della Rocca, D.G.; Di Daniele, N.; Tozzo, C.; Di Renzo, L. The effects of Italian Mediterranean organic diet (IMOD) on health status. Curr. Pharm. Des. 2010, 16, 814–824. [Google Scholar] [CrossRef] [PubMed]
- Bazzano, L.A.; Serdula, M.K.; Liu, S. Dietary intake of fruits and vegetables and risk of cardiovascular disease. Curr. Atheroscler. Rep. 2003, 5, 492–499. [Google Scholar] [CrossRef] [PubMed]
- Blekkenhorst, L.C.; Sim, M.; Bondonno, C.P.; Bondonno, N.P.; Ward, N.C.; Prince, R.L.; Devine, A.; Lewis, J.R.; Hodgson, J.M. Cardiovascular Health Benefits of Specific Vegetable Types: A Narrative Review. Nutrients 2018, 10, 595. [Google Scholar] [CrossRef] [PubMed]
- Gammone, M.A.; Riccioni, G.; Parrinello, G.; D’Orazio, N. Omega-3 Polyunsaturated Fatty Acids: Benefits and Endpoints in Sport. Nutrients 2018, 11, 46. [Google Scholar] [CrossRef] [PubMed]
- Richard, D.; Kefi, K.; Barbe, U.; Bausero, P.; Visioli, F. Polyunsaturated fatty acids as antioxidants. Pharmacol. Res. 2008, 57, 451–455. [Google Scholar] [CrossRef]
- Jensen-Urstad, A.P.; Semenkovich, C.F. Fatty acid synthase and liver triglyceride metabolism: Housekeeper or messenger? Biochim. Biophys. Acta 2012, 1821, 747–753. [Google Scholar] [CrossRef]
- Bjorndal, B.; Alteras, E.K.; Lindquist, C.; Svardal, A.; Skorve, J.; Berge, R.K. Associations between fatty acid oxidation, hepatic mitochondrial function, and plasma acylcarnitine levels in mice. Nutr. Metab. (Lond.) 2018, 15, 10. [Google Scholar] [CrossRef]
- Alaswad, K.; Lavie, C.J.; Milani, R.V.; O’Keefe, J.H., Jr. Fish oil in cardiovascular prevention. Ochsner J. 2002, 4, 83–91. [Google Scholar]
- Becerra-Tomas, N.; Blanco Mejia, S.; Viguiliouk, E.; Khan, T.; Kendall, C.W.C.; Kahleova, H.; Rahelic, D.; Sievenpiper, J.L.; Salas-Salvado, J. Mediterranean diet, cardiovascular disease and mortality in diabetes: A systematic review and meta-analysis of prospective cohort studies and randomized clinical trials. Crit. Rev. Food Sci. Nutr. 2019, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Esposito, K.; Maiorino, M.I.; Bellastella, G.; Panagiotakos, D.B.; Giugliano, D. Mediterranean diet for type 2 diabetes: Cardiometabolic benefits. Endocrine 2017, 56, 27–32. [Google Scholar] [CrossRef] [PubMed]
- Manco, M.; Calvani, M.; Mingrone, G. Effects of dietary fatty acids on insulin sensitivity and secretion. Diabetes Obes. Metab. 2004, 6, 402–413. [Google Scholar] [CrossRef] [PubMed]
- Lepretti, M.; Martucciello, S.; Burgos Aceves, M.A.; Putti, R.; Lionetti, L. Omega-3 Fatty Acids and Insulin Resistance: Focus on the Regulation of Mitochondria and Endoplasmic Reticulum Stress. Nutrients 2018, 10, 350. [Google Scholar] [CrossRef] [PubMed]
- Di Renzo, L.; Rizzo, M.; Iacopino, L.; Sarlo, F.; Domino, E.; Jacoangeli, F.; Colica, C.; Sergi, D.; De Lorenzo, A. Body composition phenotype: Italian Mediterranean Diet and C677T MTHFR gene polymorphism interaction. Eur. Rev. Med. Pharmacol. Sci. 2013, 17, 2555–2565. [Google Scholar] [PubMed]
- Di Daniele, N.; Noce, A.; Vidiri, M.F.; Moriconi, E.; Marrone, G.; Annicchiarico-Petruzzelli, M.; D’Urso, G.; Tesauro, M.; Rovella, V.; De Lorenzo, A. Impact of Mediterranean diet on metabolic syndrome, cancer and longevity. Oncotarget 2017, 8, 8947–8979. [Google Scholar] [CrossRef]
- Pandey, K.B.; Rizvi, S.I. Plant polyphenols as dietary antioxidants in human health and disease. Oxid. Med. Cell Longev. 2009, 2, 270–278. [Google Scholar] [CrossRef]
- Xu, D.P.; Li, Y.; Meng, X.; Zhou, T.; Zhou, Y.; Zheng, J.; Zhang, J.J.; Li, H.B. Natural Antioxidants in Foods and Medicinal Plants: Extraction, Assessment and Resources. Int. J. Mol. Sci. 2017, 18, 96. [Google Scholar] [CrossRef]
- Colica, C.; Di Renzo, L.; Trombetta, D.; Smeriglio, A.; Bernardini, S.; Cioccoloni, G.; Costa de Miranda, R.; Gualtieri, P.; Sinibaldi Salimei, P.; De Lorenzo, A. Antioxidant Effects of a Hydroxytyrosol-Based Pharmaceutical Formulation on Body Composition, Metabolic State, and Gene Expression: A Randomized Double-Blinded, Placebo-Controlled Crossover Trial. Oxid. Med. Cell Longev. 2017, 2017, 2473495. [Google Scholar] [CrossRef]
- Di Daniele, N.; Petramala, L.; Di Renzo, L.; Sarlo, F.; Della Rocca, D.G.; Rizzo, M.; Fondacaro, V.; Iacopino, L.; Pepine, C.J.; De Lorenzo, A. Body composition changes and cardiometabolic benefits of a balanced Italian Mediterranean Diet in obese patients with metabolic syndrome. Acta Diabetol. 2013, 50, 409–416. [Google Scholar] [CrossRef]
- De Lorenzo, A.; Bernardini, S.; Gualtieri, P.; Cabibbo, A.; Perrone, M.A.; Giambini, I.; Di Renzo, L. Mediterranean meal versus Western meal effects on postprandial ox-LDL, oxidative and inflammatory gene expression in healthy subjects: A randomized controlled trial for nutrigenomic approach in cardiometabolic risk. Acta Diabetol. 2017, 54, 141–149. [Google Scholar] [CrossRef] [PubMed]
- Di Renzo, L.; Carraro, A.; Valente, R.; Iacopino, L.; Colica, C.; De Lorenzo, A. Intake of red wine in different meals modulates oxidized LDL level, oxidative and inflammatory gene expression in healthy people: A randomized crossover trial. Oxid Med. Cell Longev. 2014, 2014, 681318. [Google Scholar] [CrossRef] [PubMed]
- Lattimer, J.M.; Haub, M.D. Effects of dietary fiber and its components on metabolic health. Nutrients 2010, 2, 1266–1289. [Google Scholar] [CrossRef] [PubMed]
- Noce, A.; Marrone, G.; Di Daniele, F.; Ottaviani, E.; Wilson Jones, G.; Bernini, R.; Romani, A.; Rovella, V. Impact of Gut Microbiota Composition on Onset and Progression of Chronic Non-Communicable Diseases. Nutrients 2019, 11, 1073. [Google Scholar] [CrossRef] [PubMed]
- Fenech, M.; El-Sohemy, A.; Cahill, L.; Ferguson, L.R.; French, T.A.; Tai, E.S.; Milner, J.; Koh, W.P.; Xie, L.; Zucker, M.; et al. Nutrigenetics and nutrigenomics: Viewpoints on the current status and applications in nutrition research and practice. J. Nutrigenet. Nutr. 2011, 4, 69–89. [Google Scholar] [CrossRef] [PubMed]
- Djuric, Z.; Turgeon, D.K.; Ren, J.; Neilson, A.; Plegue, M.; Waters, I.G.; Chan, A.; Askew, L.M.; Ruffin, M.T.t.; Sen, A.; et al. Effects of a Mediterranean Diet Intervention on Anti- and Pro-Inflammatory Eicosanoids, Epithelial Proliferation, and Nuclear Morphology in Biopsies of Normal Colon Tissue. Nutr. Cancer 2015, 67, 721–729. [Google Scholar] [CrossRef] [PubMed]
- Wojdasiewicz, P.; Poniatowski, L.A.; Szukiewicz, D. The role of inflammatory and anti-inflammatory cytokines in the pathogenesis of osteoarthritis. Mediat. Inflamm. 2014, 2014, 561459. [Google Scholar] [CrossRef]
- Kornman, K.S.; Martha, P.M.; Duff, G.W. Genetic variations and inflammation: A practical nutrigenomics opportunity. Nutrition 2004, 20, 44–49. [Google Scholar] [CrossRef]
- Furst, R.; Zundorf, I. Plant-derived anti-inflammatory compounds: Hopes and disappointments regarding the translation of preclinical knowledge into clinical progress. Mediat. Inflamm. 2014, 2014, 146832. [Google Scholar] [CrossRef]
- Sureda, A.; Bibiloni, M.D.M.; Julibert, A.; Bouzas, C.; Argelich, E.; Llompart, I.; Pons, A.; Tur, J.A. Adherence to the Mediterranean Diet and Inflammatory Markers. Nutrients 2018, 10, 62. [Google Scholar] [CrossRef]
- Zaharieva, E.; Kamenov, Z.; Velikova, T.; Tsakova, A.; El-Darawish, Y.; Okamura, H. Interleukin-18 serum level is elevated in type 2 diabetes and latent autoimmune diabetes. Endocr. Connect. 2018, 7, 179–185. [Google Scholar] [CrossRef] [PubMed]
- Calder, P.C. Omega-3 fatty acids and inflammatory processes. Nutrients 2010, 2, 355–374. [Google Scholar] [CrossRef] [PubMed]
- Goldberg, I.J.; Eckel, R.H.; McPherson, R. Triglycerides and heart disease: Still a hypothesis? Arterioscler. Thromb. Vasc. Biol. 2011, 31, 1716–1725. [Google Scholar] [CrossRef] [PubMed]
- Goel, A.; Pothineni, N.V.; Singhal, M.; Paydak, H.; Saldeen, T.; Mehta, J.L. Fish, Fish Oils and Cardioprotection: Promise or Fish Tale? Int. J. Mol. Sci. 2018, 19, 3703. [Google Scholar] [CrossRef] [PubMed]
- Omega-3 Fatty Acids and Cardiovascular Disease: Current State of the Evidence. In Comparative Effectiveness Review Summary Guides for Clinicians; Agency for Healthcare Research and Quality (US): Rockville, MD, USA, 2007.
- De Lorenzo, A.; Del Gobbo, V.; Premrov, M.G.; Bigioni, M.; Galvano, F.; Di Renzo, L. Normal-weight obese syndrome: Early inflammation? Am. J. Clin. Nutr. 2007, 85, 40–45. [Google Scholar] [CrossRef]
- Petersen, K.S.; Flock, M.R.; Richter, C.K.; Mukherjea, R.; Slavin, J.L.; Kris-Etherton, P.M. Healthy Dietary Patterns for Preventing Cardiometabolic Disease: The Role of Plant-Based Foods and Animal Products. Curr. Dev. Nutr. 2017, 1. [Google Scholar] [CrossRef] [PubMed]
- Tanzi, R.E.; Bertram, L. New frontiers in Alzheimer’s disease genetics. Neuron 2001, 32, 181–184. [Google Scholar] [CrossRef]
- Ueland, P.M.; Hustad, S.; Schneede, J.; Refsum, H.; Vollset, S.E. Biological and clinical implications of the MTHFR C677T polymorphism. Trends Pharmacol. Sci. 2001, 22, 195–201. [Google Scholar] [CrossRef]
- Mattson, M.P. Will caloric restriction and folate protect against AD and PD? Neurology 2003, 60, 690–695. [Google Scholar] [CrossRef]
- Di Renzo, L.; Merra, G.; Botta, R.; Gualtieri, P.; Manzo, A.; Perrone, M.A.; Mazza, M.; Cascapera, S.; De Lorenzo, A. Post-prandial effects of hazelnut-enriched high fat meal on LDL oxidative status, oxidative and inflammatory gene expression of healthy subjects: A randomized trial. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 1610–1626. [Google Scholar]
- Oltvai, Z.N.; Barabasi, A.L. Systems biology. Life’s complexity pyramid. Science 2002, 298, 763–764. [Google Scholar] [CrossRef] [PubMed]
- Muller, M.; Kersten, S. Nutrigenomics: Goals and strategies. Nat. Rev. Genet. 2003, 4, 315–322. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.K.; Chang, H.W.; Yan, D.; Lee, K.M.; Ucmak, D.; Wong, K.; Abrouk, M.; Farahnik, B.; Nakamura, M.; Zhu, T.H.; et al. Influence of diet on the gut microbiome and implications for human health. J. Transl. Med. 2017, 15, 73. [Google Scholar] [CrossRef] [PubMed]
- Annalisa, N.; Alessio, T.; Claudette, T.D.; Erald, V.; Antonino de, L.; Nicola, D.D. Gut microbioma population: An indicator really sensible to any change in age, diet, metabolic syndrome, and life-style. Mediat. Inflamm. 2014, 2014, 901308. [Google Scholar] [CrossRef] [PubMed]
- Kurokawa, K.; Itoh, T.; Kuwahara, T.; Oshima, K.; Toh, H.; Toyoda, A.; Takami, H.; Morita, H.; Sharma, V.K.; Srivastava, T.P.; et al. Comparative metagenomics revealed commonly enriched gene sets in human gut microbiomes. DNA Res. 2007, 14, 169–181. [Google Scholar] [CrossRef] [PubMed]
- De la Cuesta-Zuluaga, J.; Kelley, S.T.; Chen, Y.; Escobar, J.S.; Mueller, N.T.; Ley, R.E.; McDonald, D.; Huang, S.; Swafford, A.D.; Knight, R.; et al. Age- and Sex-Dependent Patterns of Gut Microbial Diversity in Human Adults. mSystems 2019, 4. [Google Scholar] [CrossRef]
- Hopkins, M.J.; Sharp, R.; Macfarlane, G.T. Variation in human intestinal microbiota with age. Dig. Liver Dis. 2002, 34 (Suppl. 2), S12–S18. [Google Scholar] [CrossRef]
- Mariat, D.; Firmesse, O.; Levenez, F.; Guimaraes, V.; Sokol, H.; Dore, J.; Corthier, G.; Furet, J.P. The Firmicutes/Bacteroidetes ratio of the human microbiota changes with age. BMC Microbiol. 2009, 9, 123. [Google Scholar] [CrossRef]
- Yatsunenko, T.; Rey, F.E.; Manary, M.J.; Trehan, I.; Dominguez-Bello, M.G.; Contreras, M.; Magris, M.; Hidalgo, G.; Baldassano, R.N.; Anokhin, A.P.; et al. Human gut microbiome viewed across age and geography. Nature 2012, 486, 222–227. [Google Scholar] [CrossRef]
- Swiatecka, D.; Narbad, A.; Ridgway, K.P.; Kostyra, H. The study on the impact of glycated pea proteins on human intestinal bacteria. Int. J. Food Microbiol. 2011, 145, 267–272. [Google Scholar] [CrossRef]
- Gelli, C.; Tarocchi, M.; Abenavoli, L.; Di Renzo, L.; Galli, A.; De Lorenzo, A. Effect of a counseling-supported treatment with the Mediterranean diet and physical activity on the severity of the non-alcoholic fatty liver disease. World J. Gastroenterol. 2017, 23, 3150–3162. [Google Scholar] [CrossRef] [PubMed]
- Kim, C.H.; Park, J.; Kim, M. Gut microbiota-derived short-chain Fatty acids, T cells, and inflammation. Immune Netw. 2014, 14, 277–288. [Google Scholar] [CrossRef] [PubMed]
- Jantchou, P.; Morois, S.; Clavel-Chapelon, F.; Boutron-Ruault, M.C.; Carbonnel, F. Animal protein intake and risk of inflammatory bowel disease: The E3N prospective study. Am. J. Gastroenterol. 2010, 105, 2195–2201. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D.; Bibiloni, R.; Knauf, C.; Waget, A.; Neyrinck, A.M.; Delzenne, N.M.; Burcelin, R. Changes in gut microbiota control metabolic endotoxemia-induced inflammation in high-fat diet-induced obesity and diabetes in mice. Diabetes 2008, 57, 1470–1481. [Google Scholar] [CrossRef] [PubMed]
- Caesar, R.; Tremaroli, V.; Kovatcheva-Datchary, P.; Cani, P.D.; Backhed, F. Crosstalk between Gut Microbiota and Dietary Lipids Aggravates WAT Inflammation through TLR Signaling. Cell Metab. 2015, 22, 658–668. [Google Scholar] [CrossRef] [PubMed]
- Suez, J.; Korem, T.; Zeevi, D.; Zilberman-Schapira, G.; Thaiss, C.A.; Maza, O.; Israeli, D.; Zmora, N.; Gilad, S.; Weinberger, A.; et al. Artificial sweeteners induce glucose intolerance by altering the gut microbiota. Nature 2014, 514, 181–186. [Google Scholar] [CrossRef] [PubMed]
- De Vrese, M.; Schrezenmeir, J. Probiotics, prebiotics, and synbiotics. Adv. Biochem. Eng. Biotechnol. 2008, 111, 1–66. [Google Scholar] [CrossRef]
- Keim, N.L.; Martin, R.J. Dietary whole grain-microbiota interactions: Insights into mechanisms for human health. Adv. Nutr. 2014, 5, 556–557. [Google Scholar] [CrossRef]
- Martinez, I.; Lattimer, J.M.; Hubach, K.L.; Case, J.A.; Yang, J.; Weber, C.G.; Louk, J.A.; Rose, D.J.; Kyureghian, G.; Peterson, D.A.; et al. Gut microbiome composition is linked to whole grain-induced immunological improvements. ISME J. 2013, 7, 269–280. [Google Scholar] [CrossRef]
- Kim, M.S.; Hwang, S.S.; Park, E.J.; Bae, J.W. Strict vegetarian diet improves the risk factors associated with metabolic diseases by modulating gut microbiota and reducing intestinal inflammation. Environ. Microbiol. Rep. 2013, 5, 765–775. [Google Scholar] [CrossRef]
- Pujia, A.M.; Costacurta, M.; Fortunato, L.; Merra, G.; Cascapera, S.; Calvani, M.; Gratteri, S. The probiotics in dentistry: A narrative review. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 1405–1412. [Google Scholar] [PubMed]
- Valdes, A.M.; Walter, J.; Segal, E.; Spector, T.D. Role of the gut microbiota in nutrition and health. BMJ 2018, 361, k2179. [Google Scholar] [CrossRef] [PubMed]
- Goodrich, J.K.; Waters, J.L.; Poole, A.C.; Sutter, J.L.; Koren, O.; Blekhman, R.; Beaumont, M.; Van Treuren, W.; Knight, R.; Bell, J.T.; et al. Human genetics shape the gut microbiome. Cell 2014, 159, 789–799. [Google Scholar] [CrossRef] [PubMed]
- Zeevi, D.; Korem, T.; Zmora, N.; Israeli, D.; Rothschild, D.; Weinberger, A.; Ben-Yacov, O.; Lador, D.; Avnit-Sagi, T.; Lotan-Pompan, M.; et al. Personalized Nutrition by Prediction of Glycemic Responses. Cell 2015, 163, 1079–1094. [Google Scholar] [CrossRef] [PubMed]
- Haverson, K.; Rehakova, Z.; Sinkora, J.; Sver, L.; Bailey, M. Immune development in jejunal mucosa after colonization with selected commensal gut bacteria: A study in germ-free pigs. Vet. Immunol. Immunopathol. 2007, 119, 243–253. [Google Scholar] [CrossRef] [PubMed]
- Karlsson, F.; Tremaroli, V.; Nielsen, J.; Backhed, F. Assessing the human gut microbiota in metabolic diseases. Diabetes 2013, 62, 3341–3349. [Google Scholar] [CrossRef]
- Dao, M.C.; Everard, A.; Aron-Wisnewsky, J.; Sokolovska, N.; Prifti, E.; Verger, E.O.; Kayser, B.D.; Levenez, F.; Chilloux, J.; Hoyles, L.; et al. Akkermansia muciniphila and improved metabolic health during a dietary intervention in obesity: Relationship with gut microbiome richness and ecology. Gut 2016, 65, 426–436. [Google Scholar] [CrossRef]
- Merra, G.; Gratteri, S.; De Lorenzo, A.; Barrucco, S.; Perrone, M.A.; Avolio, E.; Bernardini, S.; Marchetti, M.; Di Renzo, L. Effects of very-low-calorie diet on body composition, metabolic state, and genes expression: A randomized double-blind placebo-controlled trial. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 329–345. [Google Scholar]
- Magni, P.; Bier, D.M.; Pecorelli, S.; Agostoni, C.; Astrup, A.; Brighenti, F.; Cook, R.; Folco, E.; Fontana, L.; Gibson, R.A.; et al. Perspective: Improving Nutritional Guidelines for Sustainable Health Policies: Current Status and Perspectives. Adv. Nutr. 2017, 8, 532–545. [Google Scholar] [CrossRef]
| Gene | Activity | Disease | Nutrients to Supply | Reference |
|---|---|---|---|---|
| FTO | Hunger keeper Energy homeostasis Fat mass storage | Obesity | MD | [24] |
| IL-6 | Inflammation | CCD CVD Obesity | MD Polyphenols Dried Fruits | [28] |
| MCR4 | Leptin pathways | Obesity | MD | [23] |
| MTHFR | Homocysteine metabolism Lean mass development | CVD CDD Obesity Sarcopenia | Folate Cobalamin Protein | [33,34] |
| TNFα | Inflammation | CCD CVD Obesity | MD Polyphenols Dried Fruits | [36,37,38] |
| GSTM1 | Oxidative Stress | Non-Small Cell Lung Cancer | Antioxidant compound | [62] |
| MnSOD | Oxidative Stress | Breast Cancer | Antioxidant compound | [63] |
| HMGCR | Metabolism of lipids and carbohydrates | CDD | MD | [63] |
| IL-1β | Metabolism of lipids and carbohydrates | CDD | ω3 long-chain polyunsaturated fatty acids (PUFAs) | [62] |
| NFkB | Inflammation, Proliferation Angiogenesis | CDD Cancer | ω3 ω6 PUFAs | [70] |
| Nutrients | Activity | Effect | Reference |
|---|---|---|---|
| Folic acid (B9) Cobalamin (B12) Pyridoxine (B6) | Cofactor methionine synthase Availability of methyl groups for DNA methylation | Homocysteine detoxification Epigenetics regulation | [72,73,139,140] |
| Selenium | Inhibit of Dntms | Anti-Cancer | [74] |
| Ω-3 and 6 α-linoleic acid | Lowering serum triglyceride levels Ameliorating of endothelial disfunction Anti-inflammatory | Blood pressure normalization Stabilization of atherosclerotic plaques | [68,134,135,136] |
| Zinc | Metal Transcription Factor 1 | Anti-Cancer | [76,77,78] |
| Isothiocyanates | Expression of p21 Inhibit of passage from the G2 to the M phase of the cell cycle | Anti-Cancer | [80] |
| Acid butyrate | Inhibitor of histone deacetylase Inhibition of Sp1/Sp3 Activation of the p21waf1/cip1 | Anti-Cancer | [84] |
| MD (Fiber, Vitamins, Antioxidant compound, Polyphenols, ω3-6, Olive oil, Fish oil, PUFAs, Minerals) | Decreases the biosynthesis of fatty acids and the triglyceride production Levels of fatty acid oxidation | Prevent of CVD and CDDs Prevent damage of skeletal muscles | [104,105,106,107,108,109,110,111,112,113,114,115,116,117] |
| Antioxidant effects Direct scavenging of ROS | Prevent of CVD and CDDs Anti Cancer | [120,125,126] | |
| Plant used in the MD (flavonoids and polymeric flavonoid, carotenoids, monophenolic alcohols, monoterpens, phenolic acids, tannins) | Inhibit inflammatory cytokine stimulation, cell adhesion molecules and iNOS-dependent | Prevent of CVD, CDDs Anti Cancer | [130,131] |
| Plant protein | production of SCFA | High microbial diversity Gut mucosa integrity | [155] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Di Renzo, L.; Gualtieri, P.; Romano, L.; Marrone, G.; Noce, A.; Pujia, A.; Perrone, M.A.; Aiello, V.; Colica, C.; De Lorenzo, A. Role of Personalized Nutrition in Chronic-Degenerative Diseases. Nutrients 2019, 11, 1707. https://doi.org/10.3390/nu11081707
Di Renzo L, Gualtieri P, Romano L, Marrone G, Noce A, Pujia A, Perrone MA, Aiello V, Colica C, De Lorenzo A. Role of Personalized Nutrition in Chronic-Degenerative Diseases. Nutrients. 2019; 11(8):1707. https://doi.org/10.3390/nu11081707
Chicago/Turabian StyleDi Renzo, Laura, Paola Gualtieri, Lorenzo Romano, Giulia Marrone, Annalisa Noce, Alberto Pujia, Marco Alfonso Perrone, Vincenzo Aiello, Carmela Colica, and Antonino De Lorenzo. 2019. "Role of Personalized Nutrition in Chronic-Degenerative Diseases" Nutrients 11, no. 8: 1707. https://doi.org/10.3390/nu11081707
APA StyleDi Renzo, L., Gualtieri, P., Romano, L., Marrone, G., Noce, A., Pujia, A., Perrone, M. A., Aiello, V., Colica, C., & De Lorenzo, A. (2019). Role of Personalized Nutrition in Chronic-Degenerative Diseases. Nutrients, 11(8), 1707. https://doi.org/10.3390/nu11081707








